Abstract
Heterologous proteins induction by methylotrophic recombinant Pichia pastoris is generally implemented at high cells density condition. Methanol concentration (MeOH) and dissolved oxygen concentration (DO) are two crucial operating parameters controlling proteins production. It is difficult to control MeOH/DO at their desired levels simultaneously due to the extremely high oxygen consumption features. Methanol utilization plus (Mut+) and slow (MutS) strains are the two typical phenotypes of recombinant P. pastoris with quite different dynamic characteristics. Therefore, different MeOH/DO combinational control strategies or sub-optimal induction strategies could be adopted. Environments of “high MeOH/low DO” and “high DO/low MeOH” are the realistic induction strategies. In this study, we summarized our own experimental results (using Mut+/MutS strains to produce human serum albumin-human granulocyte colony stimulating factor—HSA-GCSFm/porcine interferon-α—pIFN-α), and compared to data from the literature using the above mentioned two induction strategies. The results suggested that, heterologous proteins production by Mut+ strains favors “high DO/low MeOH (DO ~ 10%, MeOH ~ 0 g/L)” induction condition, while proteins production by MutS strains prefers “high MeOH/low DO (MeOH 5–10 g/L, DO ~ 0%)” induction environment. Thus, based on the P. pastoris types, the corresponding sub-optimal induction strategies should be applied accordingly. The related metabolic analysis indicating methanol utilizing efficiency and the transcriptional analysis reflecting gene up- or down-regulations involved in several key routes in methanol and sorbitol metabolism were implemented. The analysis results strongly supported the conclusions of using the proposed sub-optimal induction strategies for different heterologous proteins production by Mut+ and MutS strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The methylotrophic P. pastoris is one of the most effective systems for the expression of heterologous proteins. Fed-batch heterologous proteins production by recombinant P. pastoris is basically divided into two phases: a growth phase to accumulate a large amount of functional cells with glycerol as the carbon source, and a subsequent induction phase by feeding methanol to initiate heterologous proteins expression. The induction phase time is much longer compared with that of growth phase (70–80 h vs 20–30 h). Therefore, optimal induction strategies are extremely important in improving heterologous proteins production. The general induction methods used for improving heterologous proteins production are: (1) induction at low temperature. This is an effective induction strategy to relieve the proteolytic activity of P. pastoris proteases and its extracellular secretion, so that the targeted recombinant proteins degradation could be repressed, particularly under conditions of nutrient deprivation (Tan et al. 2011; Wang et al. 2017). (2) methanol feeding rate control. Low methanol levels can reduce proteins productivity by triggering proteolytic degradation of the recombinant protein (Kupcsulik et al. 2001), whereas high methanol levels lead to accumulation of intracellular toxic byproducts such as formaldehyde and peroxide and cell death eventually (Zhang et al. 2000). (3) DO control by aerating pure oxygen. Large amount of oxygen is consumed in oxidation of methanol to formaldehyde (CH3OH → CH2O), thus higher oxygen supply and DO would lead to more efficient recombinant product formation (Jin et al. 2010). (4) sorbitol co-feeding together with methanol. This strategy can improve heterologous protein production by recombinant P. pastoris, since sorbitol acts as both the non-repressing carbon source and energy supplier (Gao et al. 2012; Theron et al. 2018). Co-feeding methanol/sorbitol also presents other advantages, including the lower heat formation rate and the lower oxygen consumption rate for cells growth on sorbitol (Jungo et al. 2007).
Among the above “effective induction strategies”, low temperature induction and utilization of pure oxygen seem not to be industrially realistic in consideration of operation costs and safety. As a result, adequately controlling DO and MeOH under ordinary air-aeration condition seems to be the more practical for improving proteins production. Theoretically, the induction strategies with MeOH and DO as the operating parameters have the following 4 combinations: “high MeOH/high DO”, “low MeOH/low DO”, “high MeOH/low DO” and “high DO/low MeOH”. The strategy of “high MeOH/high DO” almost could not be realized by air-aeration (Ding et al. 2014b; Jia et al. 2017; Yu et al. 2010). The reason is that, methanol induction is generally started at very high cell concentration, when methanol concentration is sufficiently high (5–10 g/L), the extremely high oxygen consuming feature of P. pastoris would decrease DO down to near zero (0) level immediately after initiating methanol induction. On the contrary, to control DO at a moderately high level could only be available by limiting methanol feeding rate or maintaining methanol concentration at very low level. On the other hand, the “low MeOH/low DO” combination is apparently unfavorable for heterologous proteins production. Thus, combinations of “high MeOH/low DO” and “high DO/low MeOH” seem to be the only realistic DO/MeOH control strategies under air-aeration condition.
In this study, we considered various heterologous proteins production processes by the two types P. pastoris (Mut+ and MutS), and investigated the sub-optimal induction strategies (“high MeOH/low DO” and “high DO/low MeOH”) for the two different processes. We summarized our own experimental results (human serum albumin-human granulocyte colony stimulating factor—HSA-GCSFm production by Mut+ P. pastoris and porcine interferon-α—pIFN-α production by MutS P. pastoris). The previous results in our lab [Mut+ strain producing porcine circovirus Cap protein (Ding et al. 2014b) and MutS strain producing human lysozyme (Chen 2018)], as well as the results reported by other related literature (Damasceno et al. 2004; Kim et al. 2013; Ponte et al. 2016; Zhang et al. 2008) under “high MeOH/low DO” and “high DO/low MeOH” induction conditions were also used for comparison purpose to verify the universal ability of the “sub-optimal induction strategies”. Then, the relevant metabolic and transcriptome analysis was carried out attempting to interpret the mechanism accounting for the results of fermentation performance improvement when using different sub-optimal induction strategies for different processes. The results would supply certain useful information in enhancing other heterologous proteins production by P. pastoris.
Materials and methods
Strains
The Mut+ recombinant P. pastoris strain (GS115), expressing the human serum albumin-human granulocyte colony stimulating factor (HSA-GCSFm) fragment (Zhu et al. 2008), was kindly provided by School of Pharmaceutical Sciences, Jiangnan University, China. The MutS recombinant P. pastoris strain (KM71), expressing the porcine interferon-α (pIFN-α) fragment (Ge et al. 2005), was kindly provided by Shanghai Academy of Agricultural Science, China.
Fermentation medium
Seed medium (in g/L, unless otherwise specified): glucose 20, yeast extract 10, peptone 20. The medium for batch fermentation: glycerol 20, (NH4)2SO4 5, H3PO4 2 (%, v/v), MgSO4 1, CaSO4 0.1, K2SO4 1; Pichia trace metals (PTM1) (Englaender et al. 2017) 10 (mL/L), pH 6.0. Feeding medium for cells growth: glycerol 500, (NH4)2SO4 0.5, KH2PO4 0.5, MgSO4 0.03; PTM1 10 (mL/L), pH 6.0. Feeding medium for induction: pure methanol, PTM1 10 (mL/L). Co-feeding medium: sorbitol 500.
Heterologous proteins production by recombinant P. pastoris in 5 L bioreactor
The production of heterologous proteins by recombinant P. pastoris was implemented in a bioreactor (BLBIO-5GJ-3-H, Bailun Bio Co., China). The initial medium was 2.3 L. Inoculation was 13% (v/v) and aeration rate was 3 vvm throughout the fermentations. Fermentation temperature was strictly controlled at 30 °C by circulating tap water or cooling water using a temperature controllable water bath (MP-10, Shanghai Permanent Science and Technology Co., China). pH was maintained at 6.0 by adding 25% (v/v) ammonia water. In cells growth phase, DO was maintained above 10% by consecutively raising agitation rate until 700 rpm. A modified DO-Stat strategy (Ding et al. 2014a) was used for feeding glycerol to allow cells concentration to reach a high density level (about 100 g-DCW/L). Pure oxygen was aerated if DO base line could not be maintained above 10% under the maximum agitation rate. When cells concentration reached the high density level, the subsequent induction phase was then initiated by co-feeding methanol/sorbitol mixture after the residual glycerol was completely used out (at about 30 h). With aid of the induction methods A and B, to be described below, the methanol and sorbitol feeding rates were automatically regulated by two programmable peristaltic pumps (BT00-100 M, Langer Co., China). O2 and CO2 partial pressures in exhaust gas (air aeration) were on-line monitored by a gas analyzer (LKM2000A, Lokas Co., Korea). O2 uptake rate (OUR) and CO2 evolution rate (CER) were determined using the standard calculation formula (Jin et al. 1995).
Induction strategy A: an industrial computer connected with an on-line methanol electrode (FC-2002, Subo Co., China) via A/D-D/A converter was used to collect methanol concentration data, and then to drive a peristaltic pump to control methanol concentration around 5–10 g/L with on–off control manner.
Induction strategy B: DO was controlled around 10% by adjusting methanol feeding rate (FMeOH) as follows:
In this study, F*MeOH and Kc were set at 0.7 mL/min and 0.05, respectively. DOset was the set-point of DO. In strategy A and B, sorbitol was co-fed with methanol at a fixed feeding ratio of 1:4 (g/g).
Measurements of cell, methanol, sorbitol, intermediate metabolites, targeted proteins concentrations and relevant methanol/sorbitol consumption rates
Cells concentration was determined by measuring the optical density at 600 nm (OD600), and then dry cell weight (DCW) was calculated by a consistent calibration curve of DCW vs OD600 (g-DWC/L = 0.25 × OD600) (Jin et al. 2010). Methanol concentration was detected using a gas chromatography (GC112A, FID detector, Shanghai Precision & Scientific Instrument Co., China) with an Alpha-Col AC20 capillary column (SGE Int’l Pty. Ltd., Australia). Sorbitol was determined by a high-performance liquid chromatography (HPLC, Agilent 1100) equipped with an amino column and a differential refractive index detector (RID). The sonication method was used for cell pretreatment to obtain the supernatant containing formaldehyde (HCHO), formate (HCOOH), pyruvate (PYR) and α-ketoglutarate (α-KG), and then the concentrations of those intermediate metabolites were detected with the HPLC (Agilent 1100) at 254 nm using an UV detector and a reverse-phase ZORBAX SBAqC18 column. The measurements details were exactly the same as those reported in the previous study (Gao et al. 2012).
HSA-GCSFm concentration was measured by the kit of trace urinary albumin (MinDian Co., Shanghai) (Guan et al. 2013). As for pIFN-α concentration measurement, an amount of 10 µL sample was placed in each cell of an electrophoretic plate (BIO-RAD Co.). The SDS-PAGE electrophoresis (15% resolving gel) was performed with the molecular weight standards until the bromophenol blue marker had reached the bottom of the gel. After SDS-PAGE analysis, the pIFN-α concentration was quantified with a G:Box Bio Imaging System and GeneTools software (SynGene Co., Cambridge, UK), each band was scanned in triplicate and an average of three readings was obtained. As for measurements of methanol/sorbitol consumption rates, two electronic balances (JA1102, Haikang Instrument Co., China) connecting with the PC via RS232 communication cable were used for on-line monitoring the methanol/sorbitol consumption amounts (g/L), by measuring the weight losses of the methanol/sorbitol feeding reservoirs. The methanol/sorbitol consumption amounts were smoothed by the corresponding quadratic polynomials with (induction) time as the independent variable. The methanol/sorbitol consumption rates at certain instant were determined by differentiating the amounts with regard to time t.
Transcriptome analysis based on RNA-seq technique
Total RNA from four samples (a Mut+ strain producing HSA-GCSFm and a MutS strain producing pIFN-α, each cultured in the environments of “high MeOH/low DO” and “high DO/low MeOH”) was extracted using an RNeasy Mini Kit (Qiagen, Hilden, Germany) and purified using an RNeasy Micro Kit (Qiagen) and RNase-Free DNase Set (Qiagen) after analysis of RNA integrity with an Agilent Bioanalyzer 2100 (Agilent Technologies, Santa Clara, CA). mRNA purification, RNA fragmentation, cDNA synthesis, adapter ligation, and amplification were performed successively to construct a cDNA library. The concentration and quality of cDNA were examined using a Qubit® 2.0 Fluorometer and Agilent Bioanalyzer 2100, respectively. An Illumina HiSeq 2500 was used to sequence the cDNA.
The clean reads obtained after data filtering were collected for genome mapping based on the Mut+ P. pastoris genome sequence (Mattanovich et al. 2009). Gene expression values were calculated using BED Tools (Quinlan and Hall 2010). Fisher’s test was used for the statistical analysis, and the calculated p values were Bonferroni corrected. The differential genes were screened according to |log2(Fold change)| ≥ 0.585 and FDRs (corrected p value) ≤ 0.005.
Results
The sub-optimal induction strategies for HSA-GCSFm and pIFN-α production by Mut+ and MutS recombinant P. pastoris
In this study, the HSA-GCSFm production by a Mut+ strain was carried out under the strategies of “high DO/low MeOH” (run #1) and “high MeOH/low DO” (run #2) respectively, with methanol/sorbitol co-feeding mode. With the “high DO/low MeOH” induction strategy, DO was controlled around 10% and MeOH naturally stayed at ~ 0 g/L during most of the period in induction phase. On the other hand, when using the “high MeOH/low DO” induction strategy, MeOH was controlled around 5 ~ 10 g/L and DO naturally stayed at ~ 0% during most of the period in induction phase (Fig. 1a, b). Sorbitol concentrations were lower than 3 g/L throughout the induction phase in run #1–2 (data not shown). The maximal HSA-GCSFm titer using the “high DO/low MeOH” strategy was 6.56-fold higher than that using the “high MeOH/low DO” strategy (Fig. 1c, d).
Curves of fermentation data and SDS analysis results of HSA-GCSFm and pIFN-α production by Mut+ and MutS strains during induction phase under different induction strategies. a–d The results for HSA-GCSFm production by Mut+ P. pastoris; e–h the results for pIFN-α production by MutS P. pastoris. Filled squares: methanol concentrations under the “high MeOH/low DO” strategy; open squares: methanol concentrations under the “high DO/low MeOH” strategy; filled circles: targeted proteins concentration under the “high MeOH/low DO” strategy; open circles: targeted proteins concentration under the “high DO/low MeOH” strategy; solid line: DO under the “high MeOH/low DO” strategy; dash line: DO under the “high DO/low MeOH” strategy
The pIFN-α production by the MutS P. pastoris was also carried out using the induction strategies of “high DO/low MeOH” (run #3) and “high MeOH/low DO” (run #4), with methanol/sorbitol co-feeding mode. Almost no sorbitol accumulation (~ 0 g/L) in the fermentation broth for both cases was observed. In run #1–4, cell concentrations stayed constantly at high density level (100–130 g-DCW/L) during induction phase. However, higher pIFN-α titer (1.86 g/L) was obtained using the “high MeOH/low DO” strategy, it increased 135% compared with its opposite companion “high DO/low MeOH” (Fig. 1g, h).
It should be addressed that, no any problem was found in creating the “high DO/low MeOH” environment using the methanol/sorbitol co-feeding operation mode. It is true that MutS P. pastoris consumes methanol quickly and in priority, but it also has the ability to utilize sorbitol though the utilization rate is relatively slower, even under the “high DO/low MeOH” environment (Arnau et al. 2010; Gao et al. 2012; Theron et al. 2018). Furthermore, the feeding rate of sorbitol was much lower than that of methanol (1:4, FSor = 1/4⋅FMeOH). The residual concentrations of both methanol and sorbitol were very low (∼ 0 g/L). A sudden rise-up in DO from its set-point implied that both methanol and sorbitol completely exhausted, slightly increasing methanol/sorbitol feeding rate (4:1) would bring DO back down to its set-point once again (Eq. 1). As a result, the existence of sorbitol did not affect the implementation of the “high DO/low MeOH” induction strategy at all. Controlling methanol concentration at low level could be realized by feeding methanol using the proposed DO control strategy (Eq. 1), and it could also be done by feeding methanol with the traditional DO-Stat method. Methanol feeding by DO-Stat method was used in many proteins production by both MutS and Mut+ strains (Jin et al. 2007; Lee et al. 2003; Lim et al. 2003). In this study, we adopted the DO control strategy (Eq. 1) to solve the problems of continuous oscillations and zero (0) base line in DO when using the DO-Stat feeding method.
The universal ability of the sub-optimal induction strategies in porcine circovirus Cap protein and human lysozyme production by Mut+ and MutS recombinant P. pastoris
The effectiveness of the proposed sub-optimal induction strategies should not be only limited in 1–2 specific recombinant proteins production. The strategies should also be applicable for more different proteins production to strength their universal ability. To verify the universal ability of the proposed sub-optimal induction strategies to other heterologous proteins production by Mut+ and MutS strains, the previous results of porcine circovirus Cap protein production by a Mut+ strain (Ding et al. 2014b) and human lysozyme production by a MutS strain (Chen 2018) obtained in our lab were summarized and compared in Table 1. The production efficiency of two other proteins [Rhizopus oryzae lipase (ROL), monoclonal antibody] by Mut+ strains and two proteins [S-adenosylmethionine (SAM), a single-chain Fv antibody (A33AscFv)] by MutS strains reported in the literature (Damasceno et al. 2004; Kim et al. 2013; Ponte et al. 2016; Zhang et al. 2008) using the two sub-optimal induction strategies were also summarized in Table 1. The results indicated that the targeted proteins expression (HSA-GCSFm, porcine circovirus Cap protein, ROL and monoclonal antibody) by Mut+ strains largely enhanced with the “high DO/low MeOH” induction strategy; while the targeted proteins expression (pIFN-α, human lysozyme, SAM, and A33AscFv) by MutS strains significantly improved using the opposite induction strategy of “high MeOH/low DO”.
Transcriptional differences in HSA-GCSFm and pIFN-α production by Mut+ and MutS P. pastoris using different induction strategies
To investigate the underlying molecular basis of heterologous proteins production enhancement using different sub-optimal strategies, global transcriptional differences of the Mut+ P. pastoris producing HSA-GCSFm and the MutS P. pastoris producing pIFN-α with the corresponding sub-optimal induction strategies were analyzed. Transcriptional data indicated that a total of 512 genes showed a significantly differed expression level (fold change > 1.5) in HSA-GCSFm production by Mut+ P. pastoris. The differentially expressed genes (DEGs) contained 235 up-regulated one and 277 down-regulated one when using the “high MeOH/low DO” induction strategy. KEGG enrichment was performed to analyze DEGs. The results implied that DEGs involved in pathways of methanol metabolism, glucolysis, TCA cycle, amino acids and ubiquitin–proteasome biosynthesis, were significantly affected by different induction strategies (“high DO/low MeOH” and “high MeOH/low DO”). As for pIFN-α production by MutS P. pastoris, transcriptional data indicated that a total of 504 genes showed a significantly differed expression level (257 up-regulations and 247 down-regulations with the “high MeOH/low DO” induction strategy). KEGG enrichment was also performed to analyze DEGs. The results suggested that DEGs involved in pathways of methanol metabolism, peroxisome and ribosomal proteins biosynthesis were clearly affected by different induction strategies. The physiological functions of above-mentioned metabolic pathways were shown in Fig. 2.
Transcriptional analysis of enhanced HSA-GCSFm expression by Mut+ P. pastoris with “high DO/low MeOH” induction strategy
Up-regulations of DEGs involved in methanol and sorbitol metabolism using the “high DO/low MeOH” strategy
As shown in Fig. 2, methanol and sorbitol metabolism contains targeted proteins synthesis pathway (pass A), formaldehyde dissimilatory (energizing) pathway (pass B), glucolysis and TCA cycle (pass C). Alcohol oxidase encoded by AOX1 and AOX2 genes mediates the first step of methanol utilization (CH3OH + O2 → HCOH + H2O2). Compared to HSA-GCSFm production with “high MeOH/low DO” strategy, the expression level of AOX1 gene with the “high DO/low MeOH” strategy was largely up-regulated. However, the specific methanol consumption rate (~ 0.027 h−1, run#1 vs ~ 0.033 h−1, run#2) and OUR (~ 130.60 mmol/L/h, run#1 vs ~ 183.09 mmol/L/h, run#2) were reversibly lower (Fig. 3a, b). This was because the “high DO/low MeOH” induction strategy restricted methanol feedings. In this case, the higher expression level of AOX1 gene could consume the fed methanol completely. The expression level of FDH gene involved in the formaldehyde dissimilation (energizing) pathway was also up-regulated with the “high DO/low MeOH” strategy (Fig. 2; Table 2). The higher expression level of FDH gene was beneficial for repressing accumulation of the toxic intermediate metabolites. The expression level of F16P gene involved in the formaldehyde assimilation (targeted proteins synthesis) pathway was also up-regulated using the “high DO/low MeOH” strategy (Fig. 2; Table 2). A higher expression level of F16P gene was favorable for enhancing HSA-GCSFm prodction. Overall, with the “high DO/low MeOH” induction strategy, methanol consumption rate and DO were controlled at moderate levels; expression levels of DEGs involved in methanol metabolism (AOX1, FDH and F16P) were largely up-regulated. We speculated that, up-regulation of DEGs invovled in methanol metabolism could increase the metabolic efficiency so that the HSA-GCSFm production greatly enhanced. On the contrary, intracellular toxic intermediate metabolites (formaldehyde) severely accumulated using the “high MeOH/low DO” induction strategy. The final intracellular formaldehyde concentration was significantly higher than that of using the “high DO/low MeOH” induction strategy (0.041 g/g/-DCW, run #2 vs 0.006 g/g-DCW, run#1).
Time courses of OUR, CER, and specific methanol consumption rate during induction phase under different induction strategies. a, b The results for HSA-GCSFm production by Mut+ P. pastoris; c, d the results for pIFN-α production by MutS P. pastoris. Filled circles: specific methanol consumption rate under the “high MeOH/low DO” strategy; open circles: specific methanol consumption rate under the “high DO/low MeOH” strategy. Black solid line: OUR under the “high MeOH/low DO” strategy; black dash line: CER under the “high MeOH/low DO” strategy; gray solid line: OUR under the “high DO/low MeOH” strategy; gray dash line: CER under the “high DO/low MeOH” strategy
On the other hand, some reports showed that key metabolic genes in glycolysis and TCA cycle are up-regulated with methanol/sorbitol co-feeding mode, suggesting that sorbitol is the main energy supplier (Theron et al. 2018). In this study, expression levels of the PGA, PCKA, HDE, OGDC, IDH2, SUCA, CISY, FUM, SDHA, ACON, and MDHC genes involved in glycolysis and TCA circle were highly up-regulated with the “high DO/low MeOH” induction strategy (Table 2). Functions of above-mentioned DEGs were shown in Fig. 2. Although expression levels of the DEGs involved in glycolysis and TCA were highly up-regulated using “high DO/low MeOH” strategy, specific sorbitol consumption rate with the “high MeOH/low DO” strategy was 22.1% higher than that of using the sub-optimal induction strategy—“high DO/low MeOH” (~ 0.0083 h−1, run #2 vs ~ 0.0068 h−1, run #1). This was due to the fact that, no restriction was exerted on sorbitol feeding in run #2. In this case, sorbitol was consumed to produce extra NADH asssociating with a large amount of CO2 release (Fig. 3a). On the contrary, with the sub-optimal induction strategy (run #1), up-regulations of DEGs invovled in glycolysis and TCA cycle could repress the intracellular accumulation of intermediate metabolites (pyruvate, α-ketoglutarate, etc.). The final intracellular concentrations of pyruvate (PYR) and α-ketoglutarate (α-KG) were low using the “high DO/low MeOH” strategy (PYR 0.021 g/g-DCW, run #1 vs 0.053 g/g-DCW, run #2; α-KG 0.010 g/g-DCW, run #1 vs 0.033 g/g-DCW, run #2).
Down-regulations of DEGs in the ubiquitin-proteasome pathway and up-regulations of DEGs in amino acid biosynthesis with the “high DO/low MeOH” strategy
Degradation of a protein via the ubiquitin–proteasome pathway involves two discrete and successive steps: (1) tagging of the substrate by covalent attachment of multiple ubiquitin molecules; (2) degradation of the tagged protein by the 26S proteasome complex with release of free and reusable ubiquitin (Glickman and Ciechanover 2002). Therefore, to investigate whether HSA-GCSFm was degraded by the ubiquitin–proteasome pathway, expression levels of the genes involved in the ubiquitin–proteasome pathway with different induction strategies were compared. As shown in Table 3, expression levels of the DEGs involved in ubiquitin and proteasome biosynthesis pathways (CDC48, PRN13, PRN11, RPN2, PSB3, PSB5 and PRS6B) were highly up-regulated with the “high MeOH/low DO” strategy. These results suggested that the HSA-GCSFm produced was severely degraded through the ubiquitin–proteasome pathway with the “high MeOH/low DO” strategy. The SEC61 complex is a protein transporting mis-folded proteins. It consists of three subunits encoded by genes of SC61A, SC61B and SC61G. Expression levels of SC61B gene and SC61G gene were also up-regulated using the “high MeOH/low DO” strategy (Table 3). The HSA-GCSFm production efficiency further reduced due to a mass of mis-folded proteins formation. In addition, expression levels of the genes involved in the amino acid biosynthesis pathway with different induction strategies were also investigated. Compared with the “high MeOH/low DO” strategy, the up-regulations of DEGs involved in amino acid biosynthesis pathway using the sub-optimal strategy—“high DO/low MeOH” could further enhance HSA-GCSFm production (Table 3).
Transcriptional analysis of enhanced pIFN-α expression by MutS P. pastoris with the “high MeOH/low DO” induction strategy
Up-regulations of DEGs involved in peroxisomes biosynthesis and methanol metabolism with the “high MeOH/low DO” strategy
As shown in Table 4, expression levels of PEX3, PEX13, PEX5, PEX2, and PMP34 genes involved in the peroxisomes biosynthesis pathway were up-regulated with the “high MeOH/low DO” strategy. The pathway of peroxisomes biosynthesis is described as follows: peroxisomal membrane proteins (PMPs) traffic from the endoplasmic reticulum to the peroxisome by a process of vesicular budding, targeting and fusion. At the same time, peroxisomal matrix proteins are imported into the organelle by the mechanism of autonomous and posttranslational (Agrawal and Subramani 2016). PEX3 serves as the receptor-mediated protein in inserting PMPs into peroxisomal membranes (Smith and Aitchison 2013). PEX13, PEX5 and PEX2 proteins are the key proteins in the process of peroxisomal matrix proteins import (Bonnet et al. 2006; Smith and Aitchison 2013). Up-regulations of these DEGs with the “high MeOH/low DO” induction strategy suggested that the peroxisomes biosynthesis was improved. Figure 2 depicted that AOX, catalase (CAT) and dihydroxyacetone synthase (DHAS) are sequestered within the peroxisomes. Since a large amount of peroxisomes were formed using the “high MeOH/low DO” strategy, higher expression levels of AOX2, DHAS and CAT genes could be obtained (Table 4). It should be noted that in MutS strains, alcohol oxidase is only encoded by AOX2 gene, leading to the comparatively low alcohol oxidase activity and O2 consumption. Meanwhile in Mut+ strains, alcohol oxidase is encoded by two functional genes (AOX1 and AOX2). This is one of the most important differences between MutS and Mut+ strains. Table 4 also indicated that exprssion levels of FLD and F16P genes involved in methanol dissimilation and assimilation pathways, were up-regulated with the “high MeOH/low DO” strategy. As shown in Fig. 2, AOX2 is the gene encoding alcohol oxidase (AOX) which dominates the entire methanol metabolism; CAT is the gene encoding catalase which could prevent the entering of toxic by-metabolite H2O2 into cytosol; FLD is the gene encoding formaldehyde dehydrogenase which contributes to NADH regeneration and detoxification of toxic intermediate metabolites in the dissimilation pathway; DHSA and F16P are the genes encoding key enzymes in methanol assimilation pathway, which is closely assocoated with methanol utilization rate in the assimilation pathway. As shown in Fig. 3d and Table 4, using the “high MeOH/low DO” strategy, both methanol consumption rates and carbon fluxes towards formaldehyde assimilation and dissimilation pathways were increased. At the same time, the toxic by-metabolite H2O2 in peroxisomes stayed at low level. As a result, pIFN-α titer significantly enhanced with this sub-optimal induction strategy. On the contrary, pIFN-α titer stayed at a very low level with the “high DO/low MeOH” induction strategy. In this case, the shortage in carbon sources could not satisfy the lowest requirements for cell maintenance and energy supply. This could also be reflected by the very low levels of the methanol consumption rate, OUR and CER (Fig. 3).
Up-regulations of DEGs involved in the ribosomal proteins biosynthesis pathway using the “high MeOH/low DO” strategy
Table 5 showed that expression levels of DEGs involved in ribosomal protein biosynthesis were largely up-regulated with the “high MeOH/low DO” strategy. Ribosomal proteins are the important structural constituents of ribosomes (Graifer and Karpova 2015). Moreover, it has been discovered that ribosomal proteins play an important role in ribosomal biogenesis, transcription, translation, DNA repair, replicative life span, adhesive growth and morphological transformation (Lu et al. 2015). Thus, we speculated that up-regulations of the DEGs involved in ribosomal proteins biosynthesis pathway significantly improved the translation efficiency of pIFN-α using this sub-optimal induction strategy.
Discussion
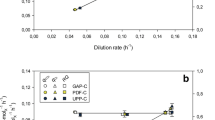
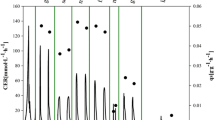
Methanol is the inducer of AOX promoter, and its concentration greatly affects the performance of heterologous proteins production (Arnau et al. 2010; Mayson et al. 2003; Wu et al. 2011). The reported sub-optimal methanol concentrations for heterologous proteins production by various Mut+ and MutS P. pastoris strains vary largely case by case without clarified conclusions. Mut+ clones are grown under methanol limiting conditions (Lim et al. 2003). On the other hand, a report showed that, the maximum 2N-transferrin concentration of 450 mg/L was obtained in the cultures grown at methanol concentration of 7 g/L using a MutS P. pastoris strain (Mayson et al. 2003). In this study, we deeply investigated the underlying molecular basis of heterologous proteins production enhancement using the corresponding sub-optimal induction strategy for Mut+ and MutS P. pastoris strain, with the HSA-GCSFm and pIFN-α producing strains as prototype. As shown in Fig. 2, both methanol dissimilation pathway and TCA cycle could supply energy (NADH) for methanol assimilation pathway in the case of using methanol/sorbitol co-feeding based induction. In the previous report (Gao et al. 2012), a parameter (η) representing methanol utilization efficiency was defined (η = methanol flux toward the assimilation pathway/the total NADH formation rate). Theoretically, a higher η implied a higher methanol-utilizing efficiency. Generally, higher targeted protein titer could be achieved when η was higher. Figure 4 depicted the methanol utilization efficiency (η) along with the induction time in runs #1–4. In HSA-GCSFm production by a Mut+ strain with the “high DO/low MeOH” strategy, η increased 188% (~ 0.08, run #2 vs ~ 0.23, run #1) as compared with its opposite companion (“high MeOH/low DO”). While in pIFN-α production by a MutS strain using the “high MeOH/low DO” strategy, η was 1.83-fold (~ 0.44, run #4 vs ~ 0.24, run #3) of that using the “high DO/low MeOH” strategy.
Time courses of methanol utilization efficiency (η) during induction phase under different induction strategies. a The variation of η for HSA-GCSFm production by Mut+ P. pastoris; b the variation of η for pIFN-α production by MutS P. pastoris. Filled squares: η under the “high MeOH/low DO” strategy; open squares: η under the “high DO/low MeOH” strategy
Major fermentation performance index such as HSA-GCSFm titer, η and transcriptional data in HSA-GCSFm production by a Mut+ strain were extensively analyzed for theoretical interpretation. The results clearly indicated that the “high DO/low MeOH” strategy was the sub-optimal induction strategy for HSA-GCSFm production. The reasons could be summarized as follows: (1) with the sub-optimal induction strategy, expression levels of DEGs involved in methanol and sorbitol metabolism (Fig. 2; Table 2) were greatly up-regulated, and methanol/sorbitol consumption rates (Fig. 3) were maintained at moderate levels. The higher expression levels of DEGs in methanol/sorbitol metabolism contributed to highly-efficient methanol/sorbitol consumption and eliminated the accumulation of toxic intermediate metabolites; (2) Compared with the “high MeOH/low DO” strategy, down-regulations of DEGs involved in ubiquitin–proteasome biosynthesis pathway and up-regulations of DEGs involved in amino acid biosynthesis pathway were obtained simultaneously using the sub-optimal induction strategy of “high DO/low MeOH” (Table 3), leading to a higher HSA-GCSFm production efficiency; (3) With the sub-optimal induction strategy, the methanol utilization efficiency (η, 0.08 → 0.23) and expression levels of AOX1, F16P genes in methanol assimilation pathway (Fig. 4; Table 2) were largely increased, more methanol flux was directed into HSA-GCSFm synthesis route leading to a higher HSA-GCSFm titer.
Major fermentation performance data in pIFN-α production by a MutS strain were also extensively analyzed for theoretical interpretation. The results suggested that pIFN-α production enhancements under the sub-optimal induction strategy of “high MeOH/low DO” were due to following reasons: (1) under the sub-optimal induction strategy, expression levels of AOX2, DHAS, FLD, F16P and CAT genes (Fig. 2; Table 4) involved in methanol metabolism were highly up-regulated. The higher expression levels of DEGs involved in methanol metabolism created a highly-efficient methanol metabolism environment and the accumulation of toxic intermediate metabolites could be significantly repressed. (2) Compared with the “high DO/low MeOH” strategy, expression levels of DEGs (Table 5) involved in ribosomal proteins biosynthesis pathway were largely up-regulated with the sub-optimal induction strategy of “high MeOH/low DO”. In this case, higher translation efficiency could also be achieved. (3) Compared with “high DO/low MeOH” strategy, methanol utilization efficiency (η, ~ 0.24 vs ~ 0.44) was greatly enhanced and expression levels of AOX2, F16P and DHAS genes involved in the methanol assimilation pathway were up-regulated, more methanol flux was directed into pIFN-α synthesis route. The highest pIFN-α titer reached a level of 1.86 g/L, which was 2.35-fold of that obtained using the “high DO/low MeOH” strategy.
References
Agrawal G, Subramani S (2016) De novo peroxisome biogenesis: evolving concepts and conundrums. Biochim Biophys Acta 5:892–901
Arnau C, Ramon R, Casas C, Valero F (2010) Optimization of the heterologous production of a Rhizopus oryzae lipase in Pichia pastoris system using mixed substrates on controlled fed-batch bioprocess. Enzyme Microb Technol 46:494–500
Bonnet C, Espagne E, Zickler D, Boisnard S, Bourdais A, Berteaux-Lecellier V (2006) The peroxisomal import proteins PEX2, PEX5 and PEX7 are differently involved in Podospora anserina sexual cycle. Mol Microbiol 62:157–169
Chen SS (2018) High-level expression of human lysozyme in Pichia pastoris. Dissertation, Jiangnan University
Damasceno LM, Pla I, Chang HJ, Cohen L, Ritter G, Old LJ, Batt CA (2004) An optimized fermentation process for high-level production of a single-chain Fv antibody fragment in Pichia pastoris. Protein Expr Purif 37:18–26
Ding J, Gao MJ, Hou GL, Liang KX, Yu RS, Li Z, Shi ZP (2014a) Stabilizing porcine interferon-α production by Pichia pastoris with an ethanol on-line measurement based DO-Stat glycerol feeding strategy. J Chem Technol Biotechnol 89:1948–1953
Ding J, Zhang CL, Gao MJ, Hou GL, Liang KX, Li CH, Ni JP, Li Z, Shi ZP (2014b) Enhanced porcine circovirus Cap protein production by Pichia pastoris with a fuzzy logic DO control based methanol/sorbitol co-feeding induction strategy. J Biotechnol 137:35–44
Englaender JA, Zhu Y, Shirke AN, Lin L, Liu XY, Zhang FM, Gross RA, Koffas MAG, Linhardt RJ (2017) Expression and secretion of glycosylated heparin biosynthetic enzymes using Komagataella pastoris. Appl Microbiol Biotechnol 101:2843–2851
Gao MJ, Li Z, Yu RS, Wu JR, Zheng ZY, Shi ZP, Zhan XB, Lin CC (2012) Methanol/sorbitol co-feeding induction enhanced porcine interferon-α production by P. pastoris associated with energy metabolism shift. Bioprocess Biosyst Eng 35:1125–1136
Ge L, Li Z, Yu RS, Liu HL, Zhang DF, Zhou ZA, Yin XH (2005) Secretive expression of porcine IFN-α in yeast Pichia pastoris. Chin J Vet Sci 25:289–292
Glickman MH, Ciechanover A (2002) The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev 82:373–428
Graifer D, Karpova G (2015) Roles of ribosomal proteins in the functioning of translational machinery of eukaryotes. Biochimie 109:1–17
Guan B, Chen FX, Lei JY, Li YH, Duan ZY, Zhu RY, Chen Y, Li HZ, Jin J (2013) Constitutive expression of a rhIL-2-HSA fusion protein in Pichia pastoris using glucose as carbon source. Appl Biochem Biotechnol 171:1792–1804
Jia LQ, Tu TY, Huai QQ, Sun JW, Chen SS, Li X, Shi ZP, Ding J (2017) Enhancing monellin production by Pichia pastoris at low cell induction concentration via effectively regulating methanol metabolism patterns and energy utilization efficiency. PLoS ONE 10:e0184602
Jin S, Ye KM, Shimizu K (1995) Metabolic pathway analysis of recombinant Saccharomyces cerevisiae with a galactose-inducible promoter based on a signal flow modeling approach. J Ferment Bioeng 80:541–551
Jin H, Zheng ZY, Gao MJ, Duan ZY, Shi ZP, Wang ZX, Jin J (2007) Effective induction of phytase in Pichia pastoris fed-batch culture using an ANN pattern recognition model-based on-line adaptive control strategy. Biochem Eng J 37:26–33
Jin H, Liu GQ, Ye XF, Duan ZY, Li Z, Shi ZP (2010) Enhanced porcine interferon-α production by recombinant Pichia pastoris with a combinational control strategy of low induction temperature and high dissolved oxygen concentration. Biochem Eng J 52:91–98
Jungo C, Schenk J, Pasquier MM, Marison IW, von Stockar U (2007) A quantitative analysis of the benefits of mixed feeds of sorbitol and methanol for production of recombinant avidin with Pichia pastori. J Biotechnol 131:57–66
Kim S, Warburton S, Boldogh I, Svensson C, Pon L, d’Anjou M, Stadheim TA, Choi BK (2013) Regulation of alcohol oxidase 1 (AOX1) promoter and peroxisome biogenesis in different fermentation processes in Pichia pastoris. J Biotechnol 166:174–181
Kupcsulik B, Sevella B, Ballagi A, Kozma J (2001) Evaluation of three methanol feed strategies for recombinant Pichia pastoris MutS fermentation. Acta Aliment 30:99–111
Lee CY, Lee SJ, Jung KH, Katoh S, Lee EK (2003) High dissolved oxygen tension enhances heterologous protein expression by recombinant Pichia pastoris. Process Biochem 38:1147–1154
Lim HK, Choi SJ, Kim KY, Jung KH (2003) Dissolved-oxygen-stat controlling two variables for methanol induction of rGuamerin in Pichia pastoris and its application to repeated fed-batch. Appl Microbiol Biotechnol 62:342–348
Lu H, Zhu YF, Xiong J, Wang R, Jia ZP (2015) Potential extra-ribosomal functions of ribosomal proteins in Saccharomyces cerevisiae. Microbiol Res 177:28–33
Mattanovich D, Callewaert N, Rouzé P (2009) Open access to sequence: browsing the Pichia pastoris genome. Microb Cell Fact 8:1–4
Mayson BE, Kilburn DG, Zamost BL, Raymond CK, Lesnicki GJ (2003) Effects of methanol concentration on expression levels of recombinant protein in fed-batch cultures of Pichia methanolica. Biotechnol Bioeng 81:291–298
Ponte X, Luis Montesinos-Segui J, Valero F (2016) Bioprocess efficiency in Rhizopus oryzae lipase production by Pichia pastoris under the control of PAOX1 is oxygen tension dependent. Process Biochem 51:1954–1963
Quinlan AR, Hall IM (2010) BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26:841–842
Smith JJ, Aitchison JD (2013) Peroxisomes take shape. Nat Rev Mol Cell Biol 14:803–817
Tan ZB, Li JF, Wu MC, Tang CD, Zhang HM, Wang JQ (2011) High-level heterologous expression of an alkaline lipase gene from Penicillium cyclopium PG37 in Pichia pastoris. World J Microbiol Biotechnol 27:2767–2774
Theron CW, Berrios J, Delvigne F, Fickers P (2018) Integrating metabolic modeling and population heterogeneity analysis into optimizing recombinant protein production by Komagataella (Pichia) pastoris. Appl Microbiol Biotechnol 102:63–80
Wang XD, Jiang T, Yu XW, Xu Y (2017) Effects of UPR and ERAD pathway on the prolyl endopeptidase production in Pichia pastoris by controlling of nitrogen source. J Ind Microbiol Biotechnol 44:1053–1063
Wu D, Yu XW, Wang TC, Wang R, Xu Y (2011) High yield Rhizopus chinenisis prolipase production in Pichia pastoris: impact of methanol concentration. Biotechnol Bioprocess Eng 16:305–311
Yu RS, Dong SJ, Zhu YM, Jin H, Gao MJ, Duan ZY, Zheng ZY, Shi ZP, Li Z (2010) Effective and stable porcine interferon-α production by Pichia pastoris fed-batch cultivation with multi-variables clustering and analysis. Bioprocess Biosyst Eng 33:473–483
Zhang WH, Bevins MA, Plantz BA, Smith LA, Meagher MM (2000) Modeling Pichia pastoris growth on methanol and optimizing the production of a recombinant protein, the heavy-chain fragment C of Botulinum Neurotoxin, serotype A. Biotechnol Bioeng 70:1–8
Zhang JG, Wang XD, Su EZ, Fang GC, Ren YH, Wei DZ (2008) A new fermentation strategy for S-adenosylmethionine production in recombinant Pichia pastoris. Biochem Eng J 41:74–78
Zhu SF, Zhang LF, Chen Y, Chu M, Li Y, Jin J (2008) Expression of mutant HSA-hGCSF in Pichia pastoris. Chin J Biochem Pharm 29:289–293
Acknowledgements
The authors thank the financial supports from Natural Science Foundation of China (#21606106), Natural Science Foundation of Jiangsu Province (#BK20150127), the Fundamental Research Funds for the Central Universities (#JUSRP51632A, #JUSRP11536), Industry-Education-Research Cooperation Project of Jiangsu Province (#BY2016022-15), National first-class discipline program of Light Industry Technology and Engineering (#2018-17) and Open Funding Project of the State Key Laboratory of Bioreactor Engineering.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Research involving human and animal participants
This article does not contain any studies with human participants performed by any of the authors.
Rights and permissions
About this article
Cite this article
Jia, L., Gao, M., Yan, J. et al. Evaluation of the sub-optimal induction strategies for heterologous proteins production by Pichia pastoris Mut+/MutS strains and related transcriptional and metabolic analysis. World J Microbiol Biotechnol 34, 180 (2018). https://doi.org/10.1007/s11274-018-2562-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-018-2562-0