Abstract
The current study aimed to address gaps in general understanding of the biodegradation of persistent organic pollutants. POP-contaminated soils were collected from the vicinity of former obsolete pesticide storage facilities in four villages, namely, Amangeldy, Belbulak, Beskainar, and Kyzylkairat, in Talgar district, Almaty region, Kazakhstan. The study soils were heavily contaminated with 24 organochlorine pesticides, 15 of which belong to POPs. Microbial diversity assessment allowed the isolation of 40 strains, which were analyzed for their potential to degrade 4.4-DDE, 4.4-DDT, α-HCH, β-HCH, and γ-HCH. Pseudomonas plecoglossicida K2 (OK217230), Bacillus aryabhattai K3 (MW866565), Solibacillus isronensis KS1 (OK236011), Pseudomonas sp. KS2 (OL348382), Bacillus pumilus B1 (OL348383), Bacillus amyloliquefaciens B2 (OL348394), Bacillus subtilis AK5 (MW866566), Pseudomonas koreensis AK1 (OL348403), Bacillus megaterium AS1 (OL348404), and Bacillus paramycoides SA1 (OL348439) effectively degraded POP-pesticides. Promising elimination was observed for a consortium Pseudomonas plecoglossicida K2 and Bacillus aryabhattai K3, with the highest degradation activity of 52.9 and 31.9% for 4.4-DDT and γ-HCH, respectively, and a single strain Bacillus aryabhattai K3, with a degradation rate of 55.2, 13.8, and 25.7% for 4.4-DDE and α- and β-isomers, respectively.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Currently, large-scale industries and anthropological activities severely damage the biosphere (Akhtar & Mannan, 2020; Gupta et al., 2020). Persistent organic pollutants (POPs) are xenobiotics with a long half-life in the environment and can pass down the food chain, causing adverse effects on human health. More than twenty organochlorine pesticides (OCPs) are currently listed as POPs (UNEP, 2007). As POPs are mostly hydrophobic and lipophilic chemicals, they have a strong affinity for lipid membranes and accumulate mainly in fatty tissues (Jones & de Voogt, 1999). A high threat from these compounds can be explained by biomagnification, i.e., the concentration of POP increases through the links of the food chain (Vasseur & Cossu-Leguille, 2006). In addition, the hotspots of this contamination are considered dangerous because POPs are highly mobile in warm latitudes and can therefore be deposited at great distances from contamination sources (Schmidt, 2010).
Soil is the environmental matrix considered the main sink for pollutants. Consequently, the composition of the soil microbial community, and in particular bacterial activity, plays a crucial role in the fate of POPs, as they form strong bonds with soil organic matter and persist in nonextractable forms (Komprda et al., 2013). The study of microbial diversity in soils contaminated with POPs is gaining popularity because some microorganisms can potentially degrade, transform, or at least alter their hydrophobicity and resistance to microbial degradation, affecting their toxicity and mobility (Bhatt et al., 2020; Burgess, 2013; Huang et al., 2020; Topp, 2003). Such microorganisms utilize POPs as the main sources of carbon, nitrogen, or auxiliary nutrients for growth (Bourtzis et al., 2016; Cycoń et al., 2017; Mishra et al., 2021). Such microbial strains can be used in bioremediation to increase cost-effectiveness and accelerate the performance of environmental remediation.
Numerous studies have shown that microorganisms of the classes Gammaproteobacteria (Pseudomonas, Aerobacter, Acinetobacter, Moraxella, and Plesiomonas), Betaproteobacteria (Burkholderia and Neisseria), Alphaproteobacteria (Sphingomonas), Actinobacteria (Micrococcus), and Flavobacteria (Flavobacterium) can degrade POPs (Geetha & Fulekar, 2008; Matsumoto et al., 2008; Rao & Wani, 2015).
Microbial degradation of POPs occurs through enzymatic activity. The known enzymes involved in the degradation process are cytochrome P450 and A-esterase, which catalyze the oxidation, reduction, or oxidative degradation reactions of xenobiotics (Bernhardt & Urlacher, 2014; Cools et al., 2011; Gonzalez & Lee, 1996; Lamb et al., 2009; Leitão, 2009; Pelkonen & Raunio, 1997; Yang et al., 2010). The bacterial strains gain energy from the final product during the degradation process. Furthermore, the degradation efficiency depends on atmospheric conditions such as temperature, pH, and soil moisture (Javaid et al., 2016).
Most current research is devoted to examining the impact of pesticides on populations of microorganisms in agrocenoses soils, but the challenges of studying soil microbial complexes in pesticide burial sites are not adequately addressed (Damstra, 2002). At the same time, microbes isolated from pesticide-contaminated ecosystems can degrade these compounds more rapidly. As a result, it is vital to investigate the microbial communities of pesticide-contaminated soils to assess the biological risk and select suitable decomposers—microorganisms for the bioremediation of natural objects (Rodríguez-Eugenio et al., 2018). Cleaning pesticide-contaminated soils using microbial destructors is undeniably effective and cost-efficient (Abraham & Silambarasan, 2016). The economic benefits of biological approaches for pesticide-contaminated soil bioremediation stem from the cheaper cost of destructors per unit area of farmed land compared to chemical agents, as well as reduced cleaning expenses compared to other methods. A cost-benefit analysis of several methods of bioremediation of contaminated soils reveals that solvent extraction costs 35–100%, soil exchange costs 10–60%, soil washing costs 10–35%, thermal desorption costs 5–20%, and bioremediation costs 4–15% (Phale et al., 2019).
According to the data presented, bioremediation is the least expensive approach to soil neutralization. The limits of biological approaches include the instability of damaging strains in the soil, the difficulty of adapting indigenous and inoculated microorganisms and plants in contaminated soils, and the necessity to develop and maintain optimal conditions for microorganism vital activity (Alori et al., 2017). As a result, the current state of research on xenobiotic microbiological destruction is characterized by a strong interest in the physiological, biochemical, and genetic aspects of the destructor strains, as well as the examination of the biotransformation routes of these substances.
Microbiological destruction is one of the most environmentally friendly, efficient, and cost-effective methods of soil remediation because microorganisms are represented by a wide variety of species and strains, having unique abilities to rapidly restructure metabolic mechanisms, the ability to exchange genetic material, and the adaptability to survive in fairly harsh environmental conditions (Chakraborty & Das, 2016). As a rule, microorganisms in the environment are involved in the decomposition and mineralization of organic compounds. They convert pollutants into less toxic compounds (metabolites or intermediates) with the help of microbial enzymes.
Microorganisms can mineralize an organic substance that is utilized as the sole source of carbon and energy, or they can utilize alternate carbon and energy sources, with the biotransformation of these contaminants occurring concurrently (cometabolism). Because incomplete degradation can often result in the creation and accumulation of more hazardous metabolites than the main substrates, the results of these catabolic events can be mineralized and then fed into the biogeochemical cycle (Patel et al., 2022).
Studies to explore the microbial landscape of pesticide-contaminated soils, screen and select potential microorganism cultures, and build a consortium based on strains that degrade organochlorine pesticides to repair damaged ecosystems appear to be useful in this context.
The current study aimed to investigate the microbial diversity of POP-contaminated soils collected near the destroyed obsolete pesticide warehouses in the villages Amangeldy, Belbulak, Beskainar, and Kyzylkairat, Talgar district, Almaty region, Kazakhstan. The isolated microbial strains were screened for the potential to degrade POP and assessed for the extent of their destructive activity.
2 Materials and Methods
2.1 Soil Collection
The research soils were collected around five former obsolete pesticide warehouses located in four villages of Talgar district, Almaty region, Kazakhstan: Amangeldy (GPS 43°17′57.98″ N 77°11′30.72″ E), Belbulak (GPS 43°19′25.95″ N 77°06′19.63″ E), Beskainar (GPS 43°13′16.36″ N 77°06′49.78″ E), and Kyzylkairat (GPS 43°17′58.78″ N 77°11′39.60″ E). Soil collected in Basshy village (GPS 44°09′03.00″ N 78°46′06.00″ E), Talgar district, Almaty region, Kazakhstan, was used as control/background soil. Soil sampling was carried out according to GOST 17.4.3.01-2017 (2019) standard using the envelope method: five soil samples were collected from the 5 × 5 m test square at a depth of 0–0.6 m and mixed. The collected soil samples were brought to the laboratory in a sterile container and stored at a temperature of 4–5°C for no longer than 24 hours (GOST 17.4.3.01-2017, 2019) until the start of the experiment.
2.2 Soil Preparation for Microbiological Analysis
Soil samples from a village were poured onto a clean, thick sheet of paper, and small stones, plant parts, and other inclusions were removed. Larger clumps of soil were then ground in a porcelain mortar and mixed with the main part of the soil sample. The thoroughly mixed soil was then placed in a square on the clean paper and divided into four equal parts (triangles) with a spatula. The soil from two opposite triangles was removed and two more were combined, remixed, and divided into four equal parts. This process was continued until 0.5 kg of soil was left. Before inoculation, the soil was dispersed by sieving (d = 3 mm). During the process, the top of the sieve was covered with sterile paper.
For the quantitative determination of microorganisms in the soil, 1–10 g of soil was required. The first dilution of a soil sample was done in a sterile container by adding distilled water (dH2O) in a ratio of 1:10 (1 g soil was diluted in 10 mL dH2O). After sample preparation, the soil was pretreated accordingly, depending on the type of microorganism and species under consideration. The process was continued to adjust the soil dilution to 0.1–0.01 mg mL−1.
The prepared dilutions were used to inoculate soil suspension on different culture media and to count microorganisms by direct microscopy.
2.3 Assessment of Microbial Diversity
The microbial diversity of research soils was analyzed by repeated dilution of soil suspensions on dense culture media to identify physiological groups capable of degrading POP-pesticides and compare microbial community composition. The number of colonies was determined by the Koch method, i.e., certain test samples were inoculated on a dense medium in Petri dishes and the colonies were counted. Different nutrient media were used to determine the number of different physiological groups of microorganisms. The composition of the media used is shown in Table 1. Nutrient agar medium was used to determine the total number of microorganisms; Wort agar medium—the fungal content; the Czapek–Dox agar medium—mold fungi; SNA medium—ammonifying bacteria; Ashby medium—nitrogen-fixing (N2-fixing) bacteria; Hutchinson and Clayton dense culture medium—the cellulolytic aerobic bacteria (Hutchinson & Clayton, 1919).
Cultures were grown in a thermostat at a temperature of 28°C for 2 days (48 h) for the growth of heterotrophic bacteria, for 5–7 days—actinomycetes, N2-fixing bacteria, and molds, and for 7–9 days—cellulolytic bacteria. After incubation, the colonies were quantified and the number of CFUs in 1 g of soil was determined (Netrusov et al., 2005).
2.4 Screening for POP-Degrading Strains
The strains of the dominant bacterial populations were examined for the potential to degrade POPs. Strains were inoculated on M9 agar medium to which POP-pesticides, namely, 4.4-DDE, 4.4-DDT, α-HCH, β-HCH, and γ-HCH, were added as a carbon source and 2,3,5-triphenyl tetrazolium chloride (TTC) as an indicator of degradation process (Table 2). The strains were cultured at 28°C for 6 days (144 h) (Vasnetsova et al., 2016). POP-pesticide degradation under aerobic conditions starts with oxidative reactions catalyzed by oxidoreductases, specifically dehydrogenases; thus, their detection ensures the destructive properties. For the detection of dehydrogenases, a 5% aqueous TTC solution added to the M9 medium was used. The POP-degrading potential of the strain was determined by the appearance of a reddish-pink coloration of the colonies, indicating the formation of reduced triphenyl formazan (TFF) (Granatskaya et al., 1992).
2.5 16S rRNA Sequencing
Molecular identification of the microbial strains was performed using the Sanger sequencing method. Genomic DNA from 1–2-day cultures was isolated using the phenol-chloroform extraction method (Green & Sambrook, 2017). The DNA concentration in the samples was determined using a Qubit fluorometer (Invitrogen, USA) on a scale for dsDNA.
The 16S rRNA gene region was used as a marker, 8F (5′-AGAGTTGATCCTGGCTCAG-3′) and 806R (5′-GGACTACCAGGGTATCTAAT-3′)—as universal primers (Edwards et al., 1989). The reaction mixture (30 μL) contained 3 μL 10×Reaction buffer (Fermentas, Lithuania), 2.5 mM MgCl2, 0.2 mM of each deoxyribonucleoside triphosphate (dNTP), 10 pmol of each primer, and 1 unit of Taq polymerase Maxima Hot Start Taq DNA polymerase (Thermo Fisher Scientific, USA). Amplification was performed in a Mastercycler pro S Thermocycler (Eppendorf, Germany) according to the following scheme: denaturation at 95°C for 7 min; annealing (30 cycles): 95°C for 30 sec, 55°C for 40 sec, 72°C for 1 min; elongation at 72°C for 10 min. The PCR product was separated on a 1.5% agarose gel, and the bands were stained with ethidium bromide and visualized in a UV transilluminator. 1×TBE buffer was used as electrode buffer. The PCR product was purified with CleanSweep™ purification reagent (Thermo Fisher Scientific, USA).
Bacterial 16S rRNA gene fragments were sequenced using the Big Dye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems, USA) according to the manufacturer’s protocol. Obtained products were purified using the BigDye® XTerminator™ Purification Kit following the manufacturer’s protocol. Capillary electrophoresis was performed using an ABI 3500 DNA analyzer (Applied Biosystems, USA). Sequencing results were processed using SeqA software (Applied Biosystems, USA).
Homologous nucleotide sequences of the 16S rRNA genes were selected using the BLAST program in the GenBank database of the US National Centre for Biotechnology Information (http://www.ncbi.nlm.nih.gov) and Ribosomal Database Project (RDP-II) (http://rdp.cme.msu.edu/html/). Phylogenetic analysis was performed using the software MEGA6. Alignment of nucleotide sequences was performed using the ClustalW algorithm. The neighbor-joining method (NJ) was used to construct the phylogenetic trees (Clayton et al., 1995).
2.6 Chemical Analysis
Chemical analysis to determine the POP-pesticide concentrations in soil samples was carried out using a Gas Chromatograph with an Electron Capture Detector (Gas Chromatography Agilent Technologies 6890N) equipped with the Combi-PAL autosampler (CTC Analytics AG, Switzerland) following governmental standard ST RK 2131-2011 (2012) for research soils.
POP-pesticide concentrations in culture media were determined by the chromatographic method according to GOST 31481-2012 (2013) on a TRACE 1310 GC gas chromatograph with a TSQ 8000 EVO triple quadrupole mass spectrometry detector with a Thermo Scientific GC Column (Thermo Fisher Scientific, USA). TG-5SILMS 30 m × 0.25 mm × 0.25 μm with low polarity 5% diphenyl/95% dimethylpolysiloxane. The conditions were as follows: injection volume—1 μL, separation ratio—1:100, flow rate—1 mL min−1, column heating rate—from 70 to 285°C at 10°C min−1, analysis time—34 min, injector temperature—210°C, and source temperature detector—285°C.
A 1000 mL M9 agar medium was placed in a separatory funnel along with 30 mL of n-hexane and then shaken vigorously for 3 min. Ethyl alcohol was added to the mixture during extraction to form an emulsion. After separation, the hexane layer was poured into an Erlenmeyer flask and the extraction with the M9 fraction was repeated twice more with 20 mL n-hexane. The extracts were combined. The resulting extract was transferred to a 100 mL beaker, covered with 10 mL Na2SO4 saturated with anhydrous sulfuric acid, and shaken gently. The purified hexane extract was washed in a long beaker with dH2O (~10 mL) until the purified M9 became neutral. The extract was dehydrated by infusion with anhydrous Na2SO4. The separator was rinsed with n-hexane and then added to the combined extract. The solvent was reduced to a volume of 0.1–0.2 mL in a circulating evaporator and then air-dried. The dry residue was dissolved in 1 mL n-hexane and used for chromatography (GOST 31858-2012, 2014).
2.7 Statistical Analysis
The degree of degradation was assessed by the difference between the POP-pesticide residues in the test and noninoculated (control) samples (in %) (Triola et al., 2017). Means and standard deviations for 3 replicates were calculated using the data analysis tool in MS Excel 2019. Means were compared using the Least Significant Difference (LSD) tests with significance declared at p < 0.05 using MStat 6.1 (Michigan State University, East Lansing, Michigan, USA).
3 Results and Discussion
3.1 POP Concentrations in Research Soils
POPs are a varied set of poisonous, semivolatile, and variable mobile chemicals in the environment that may accumulate over long distances in the air on dust particles, in abiotic matrices, and through bioaccumulation in living organisms (Alharbi et al., 2018). This class of organic compounds of natural or anthropogenic origin has a unique combination of physical and chemical properties that allow them to survive in the environment for an unusually long time because they are resistant to photolysis, chemical degradation, and, in some cases, biodegradation (Babut et al., 2013; Cachada et al., 2012). Organochlorine pesticides (OCPs), polychlorinated biphenyls (PCBs), and several polycyclic aromatic hydrocarbons are included in this category. OCPs are organic compounds that have bonded chlorine atoms, are highly lipophilic, and have high neurotoxicity.
The soils studied were analyzed for 24 OCPs (Table S1). However, for the current study, only widely known POP-pesticides were considered: 4.4-DDE, 4.4-DDT, α-HCH, β-HCH, and γ-HCH (Table 3). The total concentration of OCPs in the studied soils was in the following descending order: Kyzylkairat—121,054 μg kg−1, Beskainar—47,334 μg kg−1, Amangeldy 1—5382 μg kg−1, Amangeldy 2—1032 μg kg−1, Belbulak—1025 μg kg−1, and Basshy—146 μg kg−1 (Table S1, Fig. S1). Generally, this trend remained for the total concentrations of 5 POP-pesticides, with one exception: Beskainar soil was more contaminated than Kyzylkairat soil (Table 3).
Interestingly, residues of 7 POP-pesticides were detected in the control soil (Basshy), with levels for aldrin slightly exceeded (by 2.9 times) (Table S1). Among the soils collected near two warehouses in Amangeldy village, Amangeldy 1 was more contaminated: all POP-pesticides were detected except heptachlor; moreover, concentrations of 2.4-DDD, 4.4-DDD, 4.4-DDE, 4.4-DDT, aldrin, dieldrin, and endrin were 1.3, 1.5, 11.6, 12.4, 2.2, 44.2, and 1289 times higher than the MPC values, respectively. Amangeldy 2 contained only 6 POP-pesticides, and the concentrations of 4.4-DDE, dieldrin, endrin, and β-HCH exceeded MPC by 4.5, 26.2, 182, and 1.1 times, respectively (Table S1). In the Belbulak soil, 8 POP-pesticides were detected and the concentrations of 2.4-DDD, 4.4-DDD, 4.4-DDE, dieldrin, and endrin were higher MPC by 1.8, 2.0, 5.8, 44.0, and 11.9 times, respectively. Beskainar soil contained 13 POP-pesticides, and the concentrations of 2.4-DDD, 4.4-DDD, 4.4-DDE, 4.4-DDT, aldrin, dieldrin, endrin, and β-HCH exceeded the MPC by 8.5, 8.3, 342, 62.7, 25.6, 388, 181, and 2.5 times, respectively. Kyzylkairat soil was the most contaminated and contained all 14 POP-pesticides. The concentrations of 2.4-DDD, 4.4-DDD, 4.4-DDE, 4.4-DDT, aldrin, dieldrin, endrin, heptachlor, and β-HCH were 141, 114, 7.8, 100, 92.0, 266, 44 085, 4.3, and 4.9 times higher than the MPC, respectively (Table S1).
Thus, all six research soils contained 4.4-DDD, 4.4-DDE, dieldrin, and β-HCH, while 4.4-DDT was found in five soils.
3.2 Microbial Diversity of POP-Contaminated Soils
Soil microorganisms are involved in numerous biogeochemical processes being responsible for organic matter mineralization, element cycling, phosphorus transformation, and protein and nucleic acid synthesis (Abigail et al., 2005; Nannipieri et al., 2003). Agricultural biodiversity includes indigenous bacteria, plants, fungi, and animals (Arias et al., 2005; UNEP, 2013). The assessment of microbial diversity in soil includes not only quantitative and qualitative characteristics but also the functions and role in the ecosystem and the impact on living organisms.
Soil biological activity depends on the diversity, composition, and enzymatic activity of microorganisms. Microorganisms are involved in 80–90% of all processes in the soil (Nannipieri & Badalucco, 2003). They create favorable conditions for seed germination and the development of the plant root system, positively affecting biomass yield (Girvan et al., 2003). Plant roots release various chemical compounds into the soil influencing the microbial composition. The plant rhizosphere serves as a habitat for bacteria and mycorrhizal fungi as they utilize the root exudates as a nutrition source. In addition, microorganisms can produce antibiotics as well as substances stimulating plant growth (e.g., ethylene, auxins, and cytokines) and increase the bioavailability of nutrients (e.g., phosphates). Pseudomonas sp. can produce various biologically active compounds such as antibiotics, lytic enzymes, ethylene, auxins, and gibberellins (Sørensen & Nybroe, 2004). Furthermore, Pseudomonas sp. compete with pathogenic microorganisms for nutrients, e.g., for Fe via siderophore formation (Gałązka et al., 2015).
Therefore, POP-contaminated soils were analyzed on the dominant physiologically active microbial groups. The number of ammonifying, cellulolytic, N2-fixing, and heterotrophic bacteria and molds was determined, as these groups provide the soil self-cleaning ability and participate in the soil-forming processes. After sorption/desorption, organic pollutants are usually transported to the cell membrane and taken up by the microorganisms (Ehlers & Luthy, 2003). Sorption and transport are important processes for improving POP bioavailability, and microbial adaptation mechanisms play a crucial role in this process (Fester et al., 2014; Johnsen et al., 2005). The microbial richness of research soils contaminated with POPs was investigated (Fig. 1).
As shown in Fig. 1, the studied soils contained the microorganisms in the following amounts: ammonifying bacteria—2.1–4.7 × 107 CFU g−1, heterotrophs—3.2–7.2 × 107 CFU g−1, mold fungi—1.3–9.5 × 107 CFU g−1, and cellulolytic aerobic bacteria—1.7–7.1 × 107 CFU g−1. Basshy soil contained mainly mold fungi, cellulolytic bacteria, and yeasts being the least contaminated. Amangeldy 2 and Belbulak soils, contaminated to the same extent, differed significantly in microbial richness: Amangeldy 2 consisted mainly of mold fungi, cellulolytic, and N2-fixing bacteria, while Belbulak—cellulolytic bacteria, yeasts, and heterotrophs. The microbial community of Amangeldy 1 soil consisted mainly of N2-fixing bacteria, while heterotrophs and mold fungi were present in almost equal proportions. Beskainar soil contained heterotrophs, mold fungi, and ammonifying bacteria. The most contaminated Kyzylkairat soil contained mainly heterotrophs, N2-fixing bacteria, and mold fungi. The total amount of mesophilic aerobic and facultative anaerobic microorganisms in the control soil was 4.3 × 107 CFU g−1. Thus, the scope of cellulolytic bacteria and yeast decreased with increasing levels of contamination in soil, an opposite trend was found for N2-fixing bacteria (Fig. S2).
3.3 Screening for POP-Degrading Strains
The degradation of pollutants is regulated by abiotic reactions as well as biochemical processes of microorganisms under favorable conditions into less toxic by-products. Thus, studies are currently focused on the investigation of physiological, biochemical, and genetic characteristics of destructor strains and the determination of the xenobiotics’ biotransformation pathways (Clarke, 2013; Collin, 2010; Perelo, 2010; Sarkar et al., 2005; Scragg, 2005; Varjani & Upasani, 2017). Forty strains were isolated from research POP-contaminated soils. The half of strains showed no growth on the medium enriched with POP-pesticides as the sole carbon source, indicating the absence of destructive activity. The screening revealed 10 promising strains capable of active growth in POP-enriched medium, namely, K2, K3, KS1, KS2, B1, B2, AK5, AK1, AS1, and SA1 (Fig. 2).
As shown in Fig. 2, all colonies grown on the M9 medium were smooth, shiny red, and S-shaped. Feng et al. (1998) reported that colonies changed the initial white-dull color to red during growth in the M9 medium enriched with DDT as the sole carbon source.
The search for destructors was conducted among microorganism cultures from dominating populations. On solid medium M9, the presence of destructive strains in the composition of the entire soil microbiota was determined using a pesticide as a carbon supply 0.01% and 2,3,5-triphenyl tetrazolium chloride (TTCH) as an indicator of bacterial dehydrogenase activity. The destructive activity of the cultures was measured by the activity of cell growth and viability preservation in the presence of organochlorine chemicals (Malik et al., 2019).
3.4 Molecular Identification of POP-Degrading Strains
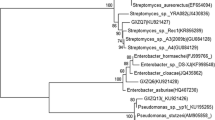
Considering the possible errors in GenBank and RDP-II, which were reported in a few studies (Rifaie et al., 2022; Song et al., 2003; Turenne et al., 2001), the construction of phylogenetic trees with 16S rRNA nucleotide sequences of the gene was additionally performed (http://www.bacterio.net). To construct a phylogenetic tree for strain K2, 16S rRNA nucleotide sequences of reference strain belonging to the Pseudomonas plecoglossicida group were used. As seen in Fig. 3, strain K2 was on the same branch as Pseudomonas plecoglossicida FPC951. Due to the high affinity (100%) of 16S rRNA gene fragments, reliable identification required analysis of the gene nucleotide sequence encoding proteins or phenotypic analysis (Table S2).
Figure 4 shows a phylogenetic analysis of the genetically related species Bacillus aryabhattai B8W22 that results in a high affinity (100%) of the strain K3 to the reference strain (Table S2).
Ten promising strains were identified as follows: Pseudomonas plecoglossicida K2 (OK217230), Bacillus aryabhattai K3 (MW866565), Solibacillus isronensis KS1 (OK236011), Pseudomonas sp. KS2 (OL348382), Bacillus pumilus B1 (OL348383), Bacillus amyloliquefaciens B2 (OL348394), Bacillus subtilis AK5 (MW866566), Pseudomonas koreensis AK1 (OL348403), Bacillus megaterium AS1 (OL348404), and Bacillus paramycoides SA1 (OL348439) (Table S2). The growth dynamic of identified active POP-degrading strains is demonstrated in Fig. 5 and Table S4.
Cell growth of Bacillus megaterium AS1 and Solibacillus isronensis KS1 peaked on the fourth day (96 h) and then declined by 43 and 21%, respectively. The growth peak for other strains was observed on the fifth day (120 h). The highest cell number was observed for Pseudomonas sp. KS2 (12.6 × 107 CFU g−1). Interestingly, the growth of Bacillus paramycoides SA1 was approximately the same during the first 48 h and then increased drastically (Fig. 5). Another interesting point was that Pseudomonas plecoglossicida К2 grew actively for 48 h, but then remained unchanged for 24 h, and further continued to grow until the peak (120 h).
3.5 Construction of Consortium and Its Destructive Activity Investigation
Many traditional and sustainable methods have been invented for the biodegradation of organic pollutants (Gomes et al., 2013). For example, gram-negative (Achromobacter, Alcaligenes, Burkholderia, Comamonas, and Pseudomonas) and gram-positive (Bacillus, Corynebacterium, and Rhodococcus) bacteria can destroy some POPs (Dercová et al., 2013; Kafilzadeh et al., 2015; Murínová et al., 2014; Murínová & Dercová, 2014a). Bioremediation can be carried out as natural attenuation or an additional method: biostimulation (addition of nutrients and inducers to promote the growth of native microorganisms) and bioaugmentation (introduction of native or appropriate exogenous bacteria to promote biodegradation) (Dudasova et al., 2017; Laszlova et al., 2016). However, successful soil bioaugmentation requires not only the use of a dedicated bacterial strain but also the construction of a bacterial consortium with the targeted degradation capacity to survive in an adverse environment (Horváthová et al., 2018; Mrozik et al., 2010; Murínová et al., 2013; Murínová & Dercová, 2014b). Furthermore, the successful degradation usually depends not only on the nature of the biochemical pathway but the overcoming of endogenous and exogenous stress associated with POPs. For example, high concentrations of POPs act as stress and hinder cell survival thereby the ability of a microorganism to degrade them. Therefore, microbial strains must develop effective adaptation mechanisms in unfavorable conditions (de Lorenzo & Loza-Tavera, 2010; Zorádová et al., 2011).
The strains Pseudomonas plecoglossicida K2 and Bacillus aryabhattai K3 showed the greatest activity against POP-pesticides. A consortium was formed based on these bacterial strains. Three different variants were investigated: Bacillus aryabhattai K3, a bacterial consortium developed using Pseudomonas plecoglossicida K2 and Bacillus aryabhattai K3, and the control (M9 medium enriched with POP-pesticides).
The general trend for the experimental treatments was as follows: the examined strain and microbial consortium degraded DDT and its metabolite more actively compared to the HCH isomers (Fig. 6). Bacillus aryabhattai K3 and the consortium Pseudomonas plecoglossicida K2 + Bacillus aryabhattai K3 degraded 4.4-DDT almost equally after both time points, while the degradation of 4.4-DDE differed: after 96 h, the consortium degraded the POP-pesticide 2.5 times more intensively than the single strain. This suggests that Bacillus aryabhattai K3 was able to decompose 4.4-DDT, while Pseudomonas plecoglossicida K2—4.4-DDE. When analyzing the degradation of HCH isomers, Bacillus aryabhattai K3 was more effective in the case of α- and β-HCH compared to the consortium. In the case of γ-HCH, the consortium degraded 31.9%, which was 1.8 times higher than a single strain. Thus, the consortium of Pseudomonas plecoglossicida K2 and Bacillus aryabhattai K3 can be effectively used to degrade 4.4-DDE and γ-HCH, while the single strain Bacillus aryabhattai K3—for 4.4-DDT and α and β isomers.
The advantages of microbiological decontamination methods over other conventional methods are that microorganisms produce different enzymes having high metabolic lability that enables them to degrade POPs. The most effective way to degrade POPs is to use an adapted consortium of microorganisms rather than a monoculture. A consortium has higher and more diverse metabolic capabilities than a single organism. The joint activity of the microorganisms in the consortium enables the complete mineralization of POPs.
In the development of technologies for pesticide-contaminated soil bioremediation using destructors, a purely empirical approach is used in each case in the selection of bioactivation conditions and, where appropriate, the selection of destructor plants for inoculation into the soil system. Pesticide degradation rates are affected not only by the type of microorganisms and substrate but also by soil moisture and the availability of trace elements, nitrogen and phosphorus sources, free oxygen, pH and medium buffering, and temperature. The success of bioremediation technologies is generally determined by the level of microbial activity in the contaminated area. In this regard, the employment of bioremediation technologies for soil remediation entails the use of agricultural practices aimed at generating optimal conditions for speeding the development and activating the metabolism of the damaging microorganisms’ cells (Sharma, 2020). The study of soil microbial complexes in pesticide burial areas has the potential to accelerate the decomposition of these compounds, making it necessary to investigate the microbial communities of pesticide-contaminated soils, both for assessing biological risk and selecting promising agents for bioremediation technology of natural objects.
4 Conclusion
An important prerequisite for this research was to determine the diversity of microorganisms in soils contaminated with POPs and to isolate pure microbial cultures to investigate their destructive properties. Promising strains were isolated and identified. The active POP-degrading strains can be recommended as a compound of commercial biological products intended for the remediation of soils contaminated with POPs. The application of consortia constructed based on POP-degrading strains in bioremediation of contaminated sites in natural conditions is a perspective approach to accelerate the restoration of marginal and contaminated soils for agricultural use.
Data Availability
The datasets generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Abigail, A., Salyers, D., & Whitt, D. (2005). Ziemia: planeta mikroorganizmów. In Mikrobiologia. Różnorodność, chorobotwórczość, środowisko (pp. 3–5). PWN.
Abraham, J., & Silambarasan, S. (2016). Biodegradation of chlor-pyrifos and its hydrolysis product 3, 5, 6-trichloro-2-pyridinol using a novel bacterium Ochrobactrum sp. JAS2:a proposal of its metabolic pathway. Pesticide Biochemistry and Physiology, 126, 13–21.
Akhtar, N., & Mannan, M. (2020). Mycoremediation: expunging environmental pollutants. Biotechnology. Reports, 26, e00452. https://doi.org/10.1016/j.btre.2020.e00452
Alharbi, O., Arsh, B. A., Khattab, R., & Ali, I. (2018). Health and environment al effects of persistent organic pollutants. Journal of Molecular Liquids, 263, 442–453. https://doi.org/10.1016/j.molliq.2018.05.029
Alori, E. T., Dare, M. O., & Babalola, O. O. (2017). Microbial inoculants for soil quality and plant health. In E. Lichtfouse (Ed.), Sustainable agriculture review (Vol. 22, pp. 281–308). Springer Cham. https://doi.org/10.1007/978-3-319-48006-0_9
Arias, M. E., González-Pérez, J. A., González-Vila, F. J., & Ball, A. S. (2005). Soil health: a new challenge for microbiologists and chemists. International Microbiology, 8(1), 13–21.
Babut, M., Arts, G. H., Caracciolo, A. B., Carluer, N., Domange, N., & Friberg, N. (2013). Pesticide risk assessment and management in a globally changing world—report from a European interdisciplinary workshop. Environmental Science and Pollution Research International, 20, 8298–8312. https://doi.org/10.1007/s11356-013-2004-3
Bernhardt, R., & Urlacher, V. B. (2014). Cytochromes P450 as promising catalysts for biotechnological application: chances and limitations. Applied Microbiology and Biotechnology, 98(14), 6185–6203. https://doi.org/10.1007/s00253-014-5767-7
Bhatt, P., Rene, E. R., Kumar, A. J., Zhang, W., & Chen, S. (2020). Binding interaction of allethrin with esterase: bioremediation potential and mechanism. Bioresource Technology, 315, 123845. https://doi.org/10.1016/j.biortech.2020.123845
Bourtzis, K., Lees, R., Hendrichs, J., & Vreysen, M. (2016). More than one rabbit out of the hat: radiation, transgenic and symbiont-based approaches for sustainable management of mosquito and tsetse fly populations. Acta Tropica, 157, 115–130. https://doi.org/10.1016/j.actatropica.2016.01.009
Burgess, L. C. (2013). Chapter 4 - organic pollutants in soil. In E. C. Brevik & L. C. Burgess (Eds.), Soils and Human Health. CRC Press.
Cachada, A., Pereira, M., Da Silva, E., & Duarte, A. (2012). Sources of potentially toxic elements and organic pollutants in an urban area subjected to an industrial impact. Environmental Monitoring and Assessment, 184, 15–32. https://doi.org/10.1007/s10661-011-1943-8
Chakraborty, J., & Das, S. (2016). Molecular perspectives and recent advances in microbial remediation of persistent organic pollutants. Environmental Science and Pollution Research, 23(17), 16883–16903. https://doi.org/10.1007/s11356-016-6887-7
Clarke, K. G. (2013). Bioprocess engineering: an introductory engineering and life science approach. Elsevier.
Clayton, R. A., Sutton, G., Hinkle, P. S., Bult, C., & Fields, C. (1995). Intraspecific variation in small-subunit rRNA sequences in GenBank: why single sequences may not adequately represent prokaryotic taxa. International Journal of Systematic and Evolutionary Microbiology, 45(3), 595–599. https://doi.org/10.1099/00207713-45-3-595
Collin, P. (2010). Dictionary of environment and ecology: over 7,000 terms clearly defined. A&C Black.
Cools, H. J., Mullins, J. G. L., Fraaije, B. A., Parker, J. E., Kelly, D. E., Lucas, J. A., & Kelly, S. L. (2011). Impact of recently emerged sterol 14α-demethylase (CYP51) variants of Mycosphaerella graminicola on azole fungicide sensitivity. Applied and Environmental Microbiology, 77(11), 3830–3837. https://doi.org/10.1128/AEM.00027-11
Cycoń, M., Mrozik, A., & Piotrowska-Seget, Z. (2017). Bioaugmentation as a strategy for the remediation of pesticide-polluted soil: a review. Chemosphere, 172, 52–71. https://doi.org/10.1016/j.chemosphere.2016.12.129
Damstra, T. (2002). Potential effects of certain persistent organic pollutants and endocrine disrupting chemicals on health of children. Clinical Toxicology, 40(4), 457–465. https://doi.org/10.1081/clt-120006748
de Lorenzo, V., & Loza-Tavera, H. (2010). Microbial bioremediation of chemical pollutants: How bacteria cope with multi-stress environmental scenarios. In Bacterial Stress Responses (pp. 481–492). John Wiley & Sons, Ltd.. https://doi.org/10.1128/9781555816841.ch30
Dercová, K., Dudášová, H., Lukáčová, L., Murínová, S., Hucko, P., Tóthová, L., & Škarba, J. (2013). Bioremediation of PCB-contaminated sediments and adaptive mechanisms of bacterial degraders exposed to polychlorinated biphenyls (PCBs). In R. K. Salar, S. K. Gahlawat, P. Siwach, & J. S. Duhan (Eds.), Biotechnology: Prospects and Applications (pp. 155–181). Springer India. https://doi.org/10.1007/978-81-322-1683-4_13
Dudasova, H., Derco, J., Sumegova, L., Dercova, K., & Laszlova, K. (2017). Removal of polychlorinated biphenyl congeners in mixture Delor 103 from wastewater by ozonation vs/and biological method. Journal of Hazardous Materials, 321, 54–61. https://doi.org/10.1016/j.jhazmat.2016.08.077
Edwards, U., Rogall, T., Blöcker, H., Emde, M., & Böttger, E. C. (1989). Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Research, 17(19), 7843–7853. https://doi.org/10.1093/nar/17.19.7843
Ehlers, L. J., & Luthy, R. G. (2003). Peer reviewed: contaminant bioavailability in soil and sediment. Environmental Science & Technology, 37(15), 295A–302A.
Feng, Y., Minard, R. D., & Bollag, J.-M. (1998). Photolytic and microbial degradation of 3,5,6-trichloro-2-pyridinol. Environmental Toxicology and Chemistry, 17(5), 814–819. https://doi.org/10.1002/etc.5620170508
Fester, T., Giebler, J., Wick, L. Y., Schlosser, D., & Kästner, M. (2014). Plant–microbe interactions as drivers of ecosystem functions relevant for the biodegradation of organic contaminants. Current Opinion in Biotechnology, 27, 168–175. https://doi.org/10.1016/j.copbio.2014.01.017
Gałązka, A., Bigos, J., & Siebielec, S. (2015). Plant growth-promoting bacteria of the genus Azospirillum and their application in agriculture. Polish Journal of Agronomy, 23, 48–62.
Geetha, M., & Fulekar, M. H. (2008). Bioremediation of pesticides in surface soil treatment unit using microbial consortia. African Journal of Environmental Science and Technology, 2(2), 36–45. https://doi.org/10.4314/ajest.v2i2
Girvan, M. S., Bullimore, J., Pretty, J. N., Osborn, A. M., & Ball, A. S. (2003). Soil type is the primary determinant of the composition of the total and active bacterial communities in arable soils. Applied and Environmental Microbiology, 69(3), 1800–1809. https://doi.org/10.1128/AEM.69.3.1800-1809.2003
Gomes, H. I., Dias-Ferreira, C., & Ribeiro, A. B. (2013). Overview of in situ and ex situ remediation technologies for PCB-contaminated soils and sediments and obstacles for full-scale application. Science of the Total Environment, 445–446, 237–260. https://doi.org/10.1016/j.scitotenv.2012.11.098
Gonzalez, F. J., & Lee, Y.-H. (1996). Constitutive expression of hepatic cytochrome P450 genes. The FASEB Journal, 10(10), 1112–1117. https://doi.org/10.1096/fasebj.10.10.8751713
GOST 17.4.3.01-2017. (2019). Nature protection. Soils.
GOST 31481-2012 (2013). Mixed fodders, raw stuff for mixed fodders. Method for determining the residual quantities of organochlorine pesticides
GOST 31858-2012 (2014). Drinking water. Method for determination of organochlorine pesticides by gas-liquid chromatography
Granatskaya, T. A., Placynda, V. A., Dvornikova, T. P., & Sirecanu, L. F. (1992). C12N1/00, C12N1/14, C12N1/20, C12Q1/00, C12Q1/32. Method to detect microorganims-destructors of xenobiotics
Green, M. R., & Sambrook, J. (2017). Isolation of high-molecular-weight DNA using organic solvents. Cold Spring Harbor Protocols, 2017(4), 356–359. https://doi.org/10.1101/pdb.prot093450
Gupta, A., Patel, A., Gupta, D., Singh, G., & Mishra, V. (2020). Rhizospheric remediation of organic pollutants from the soil; a green and sustainable technology for soil clean up. In P. Singh, A. Kumar, & A. Borthakur (Eds.), Abatement of Environmental Pollutants, Chap. 13 (pp. 263–286). Elsevier). https://doi.org/10.1016/B978-0-12-818095-2.00013-8
Horváthová, H., Lászlová, K., & Dercová, K. (2018). Bioremediation of PCB-contaminated shallow river sediments: the efficacy of biodegradation using individual bacterial strains and their consortia. Chemosphere, 193, 270–277. https://doi.org/10.1016/j.chemosphere.2017.11.012
Huang, Y., Lin, Z., Zhang, W., Pang, S., Bhatt, P., Rene, E. R., et al. (2020). New insights into the microbial degradation of D-cyphenothrin in contaminated water/soil environments. Microorganisms, 8, 473. https://doi.org/10.3390/microorganisms8040473
Hutchinson, H. B., & Clayton, J. (1919). On the decomposition of cellulose by an aerobic organism (Spirochaeta cytophaga, n. sp.). The Journal of Agricultural Science, 9(2), 143–172. https://doi.org/10.1017/S0021859600004755
Javaid, M. K., Ashiq, M., & Tahir, M. (2016). Potential of biological agents in decontamination of agricultural soil. Scientifica, 2016, e1598325. https://doi.org/10.1155/2016/1598325
Johnsen, A. R., Wick, L. Y., & Harms, H. (2005). Principles of microbial PAH-degradation in soil. Environmental Pollution, 133(1), 71–84. https://doi.org/10.1016/j.envpol.2004.04.015
Jones, K. C., & de Voogt, P. (1999). Persistent organic pollutants (POPs): state of the science. Environmental Pollution, 100(1), 209–221. https://doi.org/10.1016/S0269-7491(99)00098-6
Kafilzadeh, F., Ebrahimnezhad, M., & Tahery, Y. (2015). Isolation and identification of endosulfan-degrading bacteria and evaluation of their bioremediation in Kor river Iran. Osong Public Health and Research Perspectives, 6(1), 39–46. https://doi.org/10.1016/j.phrp.2014.12.003
Komprda, J., Komprdová, K., Sáňka, M., Možný, M., & Nizzetto, L. (2013). Influence of climate and land use change on spatially resolved volatilization of persistent organic pollutants (POPs) from background soils. Environmental Science & Technology, 47(13), 7052–7059. https://doi.org/10.1021/es3048784
Lamb, D. C., Lei, L., Warrilow, A. G. S., Lepesheva, G. I., Mullins, J. G. L., Waterman, M. R., & Kelly, S. L. (2009). The first virally encoded cytochrome P450. Journal of Virology, 83(16), 8266–8269. https://doi.org/10.1128/JVI.00289-09
Laszlova, K., Dercova, K., Horvathova, H., Murinova, S., Skarba, J., & Dudasova, H. (2016). Assisted bioremediation approaches - biostimulation and bioaugmentation - used in the removal of organochlorinated pollutants from the contaminated bottom sediments. International Journal of Environmental Research, 10(3), 367–378. https://doi.org/10.22059/ijer.2016.58756
Leitão, A. L. (2009). Potential of Penicillium species in the bioremediation field. International Journal of Environmental Research and Public Health, 6(4), 1393–1417. https://doi.org/10.3390/ijerph6041393
Malik, A., Abdieva, G., Ualieva, P., & Akimbekov, N. (2019). Study of the destructive activity of microorganisms isolated from soil contaminated by pesticides. E3S Web of Conferences, 122, 2–3. https://doi.org/10.1051/e3sconf/201912205007
Mamirova, A., Pidlisnyuk, V., Amirbekov, A., Ševců, A., & Nurzhanova, A. (2021). Phytoremediation potential of Miscanthus sinensis And. in organochlorine pesticides contaminated soil amended by Tween 20 and activated carbon. Environmental Science and Pollution Research, 28(13), 16092–16106. https://doi.org/10.1007/s11356-020-11609-y
Matsumoto, E., Kawanaka, Y., Yun, S.-J., & Oyaizu, H. (2008). Isolation of dieldrin- and endrin-degrading bacteria using 1,2-epoxycyclohexane as a structural analog of both compounds. Applied Microbiology and Biotechnology, 80(6), 1095–1103. https://doi.org/10.1007/s00253-008-1670-4
MHRK and MEPRK (2004). Standards for maximum permissible concentrations of harmful substances, pests and other biological substances polluting the soil, approved by a joint order of the Ministry of Health of the Republic of Kazakhstan dated January 30, 2004 No. 99 and the Ministry of Environmental Protection of the Republic of Kazakhstan dated January 27, 2004 No. 21-P
Mishra, S., Pang, S., Zhang, W., Lin, Z., Bhatt, P., & Chen, S. (2021). Insights into the microbial degradation and biochemical mechanism of carbamates. Chemosphere, 279, 130500. https://doi.org/10.1016/j.chemosphere.2021.130500
Mit, N., Cherednichenko, O., Mussayeva, A., Khamdiyeva, O., Amirgalieva, A., Begmanova, M., et al. (2021). Ecological risk assessment and long-term environmental pollution caused by obsolete undisposed organochlorine pesticides. Journal of Environmental Science and Health - Part B Pesticides, Food Contaminants, and Agricultural Wastes, 56(5), 490–502. https://doi.org/10.1080/03601234.2021.1913931
Mrozik, A., Cycoń, M., & Piotrowska-Seget, Z. (2010). Changes of FAME profiles as a marker of phenol degradation in different soils inoculated with Pseudomonas sp. CF600. International Biodeterioration & Biodegradation, 64(1), 86–96. https://doi.org/10.1016/j.ibiod.2009.11.002
Murínová, S., & Dercová, K. (2014a). Ochrobactrum anthropi: a promising biocatalyst for degradation of polychlorinated biphenyls in contaminated sediments. Water, Air, and Soil Pollution, 225(6). https://doi.org/10.1007/s11270-014-1980-3
Murínová, S., & Dercová, K. (2014b). Response mechanisms of bacterial degraders to environmental contaminants on the level of cell walls and cytoplasmic membrane. International Journal of Microbiology, 2014, e873081. https://doi.org/10.1155/2014/873081
Murínová, S., Dercová, K., & Dudášová, H. (2014). Degradation of polychlorinated biphenyls (PCBs) by four bacterial isolates obtained from the PCB-contaminated soil and PCB-contaminated sediment. International Biodeterioration & Biodegradation, 91, 52–59. https://doi.org/10.1016/j.ibiod.2014.03.011
Murínová, S., Dercová, K., & Murínová, S. (2013). Bacterial cell membrane adaptation responses on stress caused with the environmental pollutants. Acta Chimica Slovaca, 6(1), 106–114. https://doi.org/10.2478/acs-2013-0017
Nannipieri, P., Ascher, J., Ceccherini, M. T., Landi, L., Pietramellara, G., & Renella, G. (2003). Microbial diversity and soil functions. European Journal of Soil Science, 54(4), 655–670. https://doi.org/10.1046/j.1351-0754.2003.0556.x
Nannipieri, P., & Badalucco, L. (2003). Biological processes. In R. Nieder & D. Benbi (Eds.), Handbook of Processes and Modeling in the Soil-Plant System (pp. 57–82). CRC Press.
Netrusov, A. I., Egorova, M. A., & Zakharchuk, L. M. (2005). In A. I. Netrusov (Ed.), Practicum on microbiology: textbook for university students. Academia.
Patel, A. K., Singhania, R. R., Albarico, F. P. J. B., Pandey, A., Chen, C.-W., & Dong, C.-D. (2022). Organic wastes bioremediation and its changing prospects. Science Total Environment, 824, 153889. https://doi.org/10.1016/j.scitotenv.2022.153889
Pelkonen, O., & Raunio, H. (1997). Metabolic activation of toxins: tissue-specific expression and metabolism in target organs. Environmental Health Perspectives, 105(suppl 4), 767–774. https://doi.org/10.1289/ehp.97105s4767
Perelo, L. W. (2010). Review: in situ and bioremediation of organic pollutants in aquatic sediments. Journal of Hazardous Materials, 177(1), 81–89. https://doi.org/10.1016/j.jhazmat.2009.12.090
Phale, P. S., Sharma, A., & Gautam, K. (2019). Microbial degradation of xenobiotics like aromatic pollutants from the terrestrial environments. In Pharmaceuticals and Personal Care Products: Waste Management and Treatment Technology. Elsevier.
Rao, R. J., & Wani, K. A. (2015). Bioremediation of pesticides under the influence of bacteria and fungi. In Handbook of research on uncovering new methods for ecosystem management through bioremediation (pp. 51–72). IGI Global. https://doi.org/10.4018/978-1-4666-8682-3.ch003
Rifaie, S., Patil, V., & Jangid, K. (2022). Chapter 15 - advances in sequencing technology, databases, and analyses tools for the assessment of microbial diversity. In A. Gunjal & S. Shinde (Eds.), Microbial Diversity in Hotspots (pp. 317–347). Academic Press. https://doi.org/10.1016/B978-0-323-90148-2.00003-1
Rodríguez-Eugenio, N., McLaughlin, M., & Pennock, D. (2018). Soil Pollution: a hidden reality (p. 142). FAO.
Sarkar, D., Ferguson, M., Datta, R., & Birnbaum, S. (2005). Bioremediation of petroleum hydrocarbons in contaminated soils: comparison of biosolids addition, carbon supplementation, and monitored natural attenuation. Environmental Pollution, 136(1), 187–195. https://doi.org/10.1016/j.envpol.2004.09.025
Schmidt, C. (2010). How PCBs are like grasshoppers. Environmental Science & Technology, 44(8), 2752–2752. https://doi.org/10.1021/es100696y
Scragg, A. H. (2005). Environmental biotechnology. OXFORD university press.
Sharma, I. (2020). Bioremediation techniques for polluted environment: concept, advantages, limitations, and prospects. In M. A. Murillo-Tovar, H. A. Saldarriaga-Norena, & A. Saeid (Eds.), Trace metals in the environment: New approaches and recent advances. IntechOpen. https://doi.org/10.5772/intechopen.90453
Song, Y., Liu, C., McTeague, M., & Finegold, S. M. (2003). 16S ribosomal DNA sequence-based analysis of clinically significant gram-positive anaerobic cocci. Journal of Clinical Microbiology, 41(4), 1363–1369. https://doi.org/10.1128/JCM.41.4.1363-1369.2003
Sørensen, J., & Nybroe, O. (2004). Pseudomonas in the soil environment. In J.-L. Ramos (Ed.), Pseudomonas: genomics, life style and molecular architecture (pp. 369–401). Springer US. https://doi.org/10.1007/978-1-4419-9086-0_12
ST RK 2131-2011 (2012). Soil quality. Determination of organochlorine pesticides and polychlorinated biphenyls content. Gas chromatographic method with electron capture detection
Topp, E. (2003). Bacteria in agricultural soils: diversity, role and future perspectives. Canadian Journal of Soil Science, 83(Special Issue), 303–309. https://doi.org/10.4141/S01-065
Triola, M. M., Triola, M. F., & Roy, J. (2017). Biostatistics for the biological and health sciences (2nd ed.). Pearson Addison-Wesley Boston.
Turenne, C. Y., Tschetter, L., Wolfe, J., & Kabani, A. (2001). Necessity of quality-controlled 16S rRNA gene sequence databases: identifying nontuberculous Mycobacterium species. Journal of Clinical Microbiology, 39(10), 3637–3648. https://doi.org/10.1128/JCM.39.10.3638-3648.2001
UNEP. (2007). Listing of POPs in the Stockholm convention. http://chm.pops.int/TheConvention/ThePOPs/ListingofPOPs/tabid/2509/Default.aspx. Accessed 15 June 2022
UNEP. (2013). International environmental law-making and diplomacy review 2012 (No. CBD/SBSTTA/7/12 (p. 242). University of Eastern Finland.
Varjani, S. J., & Upasani, V. N. (2017). A new look on factors affecting microbial degradation of petroleum hydrocarbon pollutants. International Biodeterioration & Biodegradation, 120, 71–83. https://doi.org/10.1016/j.ibiod.2017.02.006
Vasnetsova, E. V., Ksenofontova, O. Y., Tikhonova, D. A., Filimonova, E. A., & Savina, K. V. (2016). Search of bacteria destructors of pesticides prometrin, hexachlorocyclohexane (HCH), dihlordifeniltrihlormetilmetan (4,4-DDT) in soil with pesticides burial places in the Saratov region. News of the Saratov University. New series. Series Chemistry. Biology. Ecology, 16(3), 349–354.
Vasseur, P., & Cossu-Leguille, C. (2006). Linking molecular interactions to consequent effects of persistent organic pollutants (POPs) upon populations. Chemosphere, 62(7), 1033–1042. https://doi.org/10.1016/j.chemosphere.2005.05.043
Yang, W., Bell, S. G., Wang, H., Zhou, W., Hoskins, N., Dale, A., et al. (2010). Molecular characterization of a class I P450 electron transfer system from Novosphingobium aromaticivorans DSM12444*. Journal of Biological Chemistry, 285(35), 27372–27384. https://doi.org/10.1074/jbc.M110.118349
Zorádová, S., Dudášová, H., Lukáčová, L., Dercová, K., & Čertík, M. (2011). The effect of polychlorinated biphenyls (PCBs) on the membrane lipids of Pseudomonas stutzeri. International Biodeterioration & Biodegradation, 65(7), 1019–1023. https://doi.org/10.1016/j.ibiod.2011.03.012
Funding
This research was supported by Program ВR05236379 “Comprehensive assessment of the impact of unused and banned pesticides on the genetic status and health of the population of Almaty region (Determination of microbial diversity of environmental objects in the vicinity of pesticide disposal sites, screening of microorganisms-destructors of chemical pollutants)”, MES RK.
Author information
Authors and Affiliations
Contributions
AM conceived the presented idea. AM and GA contributed to the writing and prepared the figures and tables. PU participated in revising the manuscript. All authors approved it for publication.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 554 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Malik, A., Abdieva, G. & Ualieva, P. Microbial Diversity of Soils Contaminated with Persistent Organic Pollutants and POP-Degrading Strains. Water Air Soil Pollut 234, 290 (2023). https://doi.org/10.1007/s11270-023-06193-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-023-06193-z