Abstract
Mitochondria play an essential role in intracellular energy metabolism. This study described the involvement of Bombyx mori nucleopolyhedrovirus (BmNPV) GP37 (BmGP37) in host mitochondria. Herein, the proteins associated with host mitochondria isolated from BmNPV-infected or mock-infected cells by two-dimensional gel electrophoresis were compared. One mitochondria-associated protein in virus-infected cells was identified as BmGP37 by liquid chromatography-mass spectrometry analysis. Furthermore, the BmGP37 antibodies were generated, which could react specifically with BmGP37 in the BmNPV-infected BmN cells. Western blot experiments showed that BmGP37 was expressed at 18 h post-infection and was verified as a mitochondria-associated protein. Immunofluorescence analysis demonstrated that BmGP37 localized to the host mitochondria during BmNPV infection. Furthermore, western blot analysis revealed that BmGP37 is a novel component protein of the occlusion-derived virus (ODV) of BmNPV. The present results indicated that BmGP37 is one of the ODV-associated proteins and may have important roles in host mitochondria during BmNPV infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Baculoviridae is a diverse family of large DNA viruses that infect insects. Baculoviruses are divided into four genera; Alphabaculovirus, Betabaculovirus, Gammabaculovirus, and Deltabaculovirus, which are lepidopteran-specific nucleopolyhedroviruses (NPVs), lepidopteran-specific granuloviruses (GVs), hymenopteran-specific NPVs, and dipteran-specific NPVs, respectively [1]. Based on the phylogenetic studies, Alphabaculovirus can be further subdivided into group I and II NPVs [2]. During their life cycle, NPVs produce two morphological forms: the occlusion-derived virus (ODV) and budded virus (BV). ODVs are occluded in a protein matrix that forms polyhedra called occlusion bodies (OBs), which transmit viruses from insect to insect via per os infection. In contrast, BVs spread viruses from cell to cell [3].
It is well known that NPVs regulate and rearrange host cells in different ways during their infection cycles. For example, in a typical host antiviral response, apoptosis is inhibited by NPV-encoded p35 and inhibitor of apoptosis (IAP) genes [4]. The host cell cycle is arrested at the S or G2/M phase by NPV infection [5]. Also, the transcription of many host genes is up- or down-regulated at the RNA level during NPV replication [6,7,8]. As a result of host cell control, a large amount of viral genomic DNA is replicated in the NPV-infected cells, abundant polyhedrin protein is expressed for 30%–50% of the total host protein, and numerous OBs are formed in the host nucleus [9].
Living organisms utilize intracellular adenosine 5´-triphosphate (ATP), derived from cytosolic glycolysis and mitochondrial oxidative phosphorylation, as a primary energy source for most cellular processes. On the other hand, viruses rely entirely on their host for an energy source. Several reports on the relationship between NPV and host cell energy metabolism exist. In NPV-infected cells, an increase in ADP/ATP ratio indicated ATP consumption, activation of the tricarboxylic acid (TCA) cycle involved in ATP production, and increased transcription of host genes involved in glycolysis was observed [6, 10, 11]. However, no correlation between the host mitochondria and NPV-derived proteins has been reported. In this study, the mitochondria of Bombyx mori nucleopolyhedrovirus (BmNPV)-infected cells were isolated. The proteins associated with the mitochondria were analyzed by two-dimensional polyacrylamide gel electrophoresis (2D-PAGE) and liquid chromatography/mass spectrometry (LC/MS). It was confirmed that BmNPV GP37 (BmGP37) is a host mitochondria-associated protein. We further demonstrated that BmGP37 is associated with the occlusion-derived virus (ODV) particles of BmNPV.
Materials and methods
Cells, viruses, and plaque assay
The BmN cells were cultured in TC-100 medium supplemented with 10% fetal bovine serum, as described previously [12]. The BmNPV T3 [13] isolate was propagated in the BmN cells. The cells were infected with BmNPV at a multiplicity of infection of 10. As determined with the plaque assay, the virus titers are expressed as plaque-forming units, as described previously [14].
Isolation and two-dimensional-polyacrylamide gel electrophoresis (2D-PAGE) of mitochondria-associated proteins
The BmN cells (1 × 108) infected with BmNPV were collected at 72 h post-infection (hpi). The mitochondria were isolated from the mock- or virus-infected cells using a mitochondria isolation kit for cultured cells (Thermo Fisher Scientific, Waltham, MA). The mitochondria-associated proteins were separated by isoelectric focusing (IEF) using agarGEL pH5-10 A-C510 (ATTO, Tokyo, Japan) at 300 V for 210 min, followed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE). The excised SDS-PAGE gel fragment corresponding to the protein spot of virus-infected cells-derived mitochondria was subjected to in-gel digestion with trypsin, as described previously [15]. The digested peptides were separated using an InertSustainSwift C18 column (20 × 150 mm, 3 μm pore size; GL Sciences, Tokyo, Japan) and subjected to LC/MS analysis performed with a Triple TOF 5600 mass spectrometer (AB Sciex, Framingham, MA). The MS/MS Ions Search was performed using the Mascot search engine (http://www.matrixscience.com/search_form_select.html) against the Swiss-Prot and NCBI non-redundant protein databases, as described previously [16].
Antibody production
The genomic DNA of BmNPV was extracted from the occlusion-derived virus (ODV) using proteinase K digestion and phenol–chloroform extraction [13]. The fragment of the BmGP37 gene (amino acid residues 97–289) was amplified by PCR using the primers GP37_600_BamHI-F: 5´-ttcgggatccagacgattttgacctaatcaaacaaaggg-3´ and GP37_600_SalI-R: 5´-ttttgtcgactgcacgccaacaagatttttcgtc-3´ and inserted into the pET32b vector (Novagen, Madison, WI). After expressing His-tagged BmGP37 in E. coli strain BL21, a recombinant BmGP37 protein (rBmGP37) was purified using a TALON spin column (Clontech, Palo Alto, CA) under denaturing conditions. Purified rBmGP37 was used to generate polyclonal antibodies in rabbits (Eurofins Genomics, Tokyo, Japan).
Immunodetection of BmGP37
For western blotting, the proteins were electrophoresed, transferred onto nitrocellulose membrane (GVS, Emilia-Romagna, Italy), and probed with antibodies of BmGP37, ATP5A (Santa Cruz Biotechnology, sc-136178, Dallas, TX), HSC70-4 [17], GP64 (Santa Cruz Biotechnology, sc-65499), and VP39 [18]. For immunofluorescence analysis, the virus- or mock-infected BmN cells were fixed, permeabilized, and rehydrated as described previously [17]. After antibody and Mito Tracker (Invitrogen, Carlsbad, CA) treatments, the cells were mounted with SlowFade Antifade Kit (Molecular Probes, Eugene, OR) and analyzed using the Olympus BX50 Fluorescence Microscope (Tokyo, Japan).
Results
Identification of a mitochondria-associated protein from the BmNPV-infected cells
To identify BmNPV-derived proteins associated with host mitochondria, mock- and virus-infected BmN cells-derived mitochondria were isolated and compared using 2D-PAGE. As shown in Fig. 1A, the protein spots were commonly observed in mock- and virus-infected cells. In contrast, a spot (indicated by an arrowhead) was detected in the 72 hpi mitochondrial fraction. This spot was excised and subjected to in-gel digestion and LC/MS analysis, resulting in the identification of five peptide sequences. As shown in Fig. 1B, these peptide sequences corresponded to BmGP37.
BmGP37 is a host mitochondria-associated protein. A 2D-PAGE of host mitochondria-associated proteins derived from BmNPV-infected cells. The BmN cells were infected with BmNPV at a multiplicity of infection of 10. At 72 hpi, host mitochondria were isolated and subjected to IEF followed by SDS-PAGE. B Identification of BmGP37 by LC/MS analysis. The BmNPV-infected cell-specific spot (arrowhead) was subjected to in-gel digestion followed by LC/MS analysis. Underlines indicate the identified amino acid sequences. Ten amino acid residues on the N-terminal side indicate a positively charged amphiphilicity score region, and an asterisk indicates a predicted cleavage site by mitochondrial processing peptidase
BmGP37 expression in the BmNPV-infected cells
To investigate the expression and localization of BmGP37 during viral infection, the polyclonal antiserum raised against E. coli-expressed recombinant BmGP37 (rBmGP37) was generated (Fig. 2A). Western blot analysis using this antiserum revealed the presence of one specific signal in BmNPV-infected cells. In contrast, no signal was detected using the corresponding pre-immune serum (Fig. 2B). Furthermore, to assess the expression profile and glycosylation of BmGP37, western blot analyses were performed with this antiserum. As shown in Fig. 3A, BmGP37 was detected from 18 hpi, and clear signals were observed from 48–96 hpi. On the other hand, the signal of BmHSC70-4, a host molecular chaperone expressed at steady-state level, decreased at 72 and 96 hpi. Also, since tunicamycin treatment shifted the molecular size of BmGP37 in the BmN cells to a lower one, BmGP37 was considered modified by N-linked glycosylation.
Generation of anti- BmGP37 antibody. A Recombinant BmGP37 expressed by E. coli. The recombinant partial BmGP37 was purified under denaturing conditions by TALON spin columns. M and P indicate lanes of molecular weight markers and purified recombinant BmGP37, respectively. B Evaluation of anti-BmGP37 antibody. Mock- or BmNPV-infected BmN cells were subjected to SDS-PAGE followed by western blot analysis using pre-immune or anti-BmGP37, anti-BmHSC70-4 antiserum. M, 1 and 2 indicate lanes of molecular weight markers, mock-infected BmN cells and BmNPV-infected cells at 72 hpi, respectively
BmGP37 is expressed in the late phase of infection and modified with N-linked glycans. A The expression of BmGP37 in BmNPV-infected cells. The BmN cells were inoculated with BmNPV at a multiplicity of infection of 10. At designated time points, the cells were collected and subjected to SDS-PAGE followed by western blotting using anti-BmGP37 or anti-BmHSC70-4 antiserum. M indicates mock infection. B Glycosyl modification of BmGP37. The BmN cells were inoculated with BmNPV after adding 10 μM tunicamycin or DMSO. The cells were collected at 72 hpi and subjected to SDS-PAGE followed by western blot analysis using anti-BmGP37 antiserum. Anti-BmHSC70-4 antiserum was used as a negative control
Distribution of BmGP37 to host mitochondria
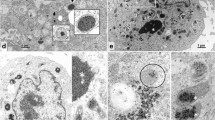
To determine whether virus-derived BmGP37 is associated with the host mitochondria of BmNPV-infected cells, mitochondrial and cytosolic fractions were isolated from mock- or virus-infected cells and subjected to western blot analysis. As shown in Fig. 4A, BmGP37 was detected in the mitochondrial fraction, similar to the signal of ATP5A, a marker protein in the mitochondrial fraction. On the other hand, BmHSC70-4, a well-known molecular chaperone, was detected in both mitochondrial and cytosolic fractions. Furthermore, to identify the intracellular localization of BmGP37, the BmNPV-infected cells were subjected to immunofluorescence analyses. As shown in Fig. 4B, accurate localization of the mitochondrial marker Mito Tracker was observed in mitochondria in mock-infected cells. At 24 hpi, it was observed that the signals of BmGP37 and Mito Tracker coincide in the cytoplasm. In the nucleus, Mito Tracker staining showed sharp, crystalline, and strong foci at 48 and 72 hpi. However, the comparison between Mito Tracker and bright field suggested that Mito Tracker reacted non-specifically with OBs that appeared from 48 hpi. On the other hand, in the cytoplasm, the localization of BmGP37 was observed to be consistent with Mito Tracker. Interestingly, while it was observed that BmGP37 was co-localized with Mito Tracker in the cytoplasm at 72 hpi, weak BmGP37 protein signal was also observed in the nucleus where nucleocapsids and ODV particles are constructed. It was observed that an intranuclear BmGP37 was subtly accumulated, unlike the OB localization.
BmGP37 is a mitochondria-associated protein. A Western blot analysis of mitochondrial fractions derived from BmNPV-infected BmN cells. The mock- or BmNPV-infected BmN cells were enucleated and fractionated into the mitochondrial fraction (M) and the cytosolic fraction (C). Each fraction was subjected to SDS-PAGE followed by western blotting using anti-BmGP37, ATP5A, and BmHSC70-4 antibodies. B Immunofluorescence image of BmNPV-infected cells. The BmN cells were inoculated with BmNPV at a multiplicity of infection of 10. The cells were fixed and subjected to immunofluorescence analysis at 24, 48, and 72 hpi. The localizations of the nucleus, mitochondria, and BmGP37 were identified by 4′,6-diamidino-2-phenylindole (DAPI), Mito Tracker, and anti-BmGP37 antibody, respectively. The scale bar is 10 µm
Confirmation of the association of BmGP37 and ODV particles
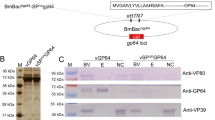
To clarify whether BmGP37 is associated with the viral particles, BV and ODV were purified and subjected to western blot analysis. As shown in Fig. 5, a clear signal of BmGP37 was detected in ODV, but no signal was detected in BV. The signal of GP64, a marker of BV, was observed only in BV. The signals of BmHSC70-4 and VP39, the markers of ODV and BV, were detected in ODV and BV. These results indicated that BmGP37 is an associated protein of BmNPV ODV.
Discussion
Association of BmGP37 with host mitochondria
GP37 is a well-conserved glycoprotein encoded by most alphabaculoviruses and a few betabaculoviruses [3]. For the intracellular localization of GP37, it has been reported that Orgyia pseudotsugata multiple nucleopolyhedrovirus (OpMNPV) GP37 concentrated in the cytoplasmic inclusion bodies [19], and Mamestra brassicae multiple nucleopolyhedrovirus (MbMNPV) GP37 is located in cytoplasmic foci presumed to be the endoplasmic reticulum (ER) [20]. Interestingly, a part of the ER is known to contact the mitochondria in the cytoplasm [21]. This interconnection functions in mitochondrial dynamics [22], inflammasome formation [23], activation of autophagy [24], and redox signaling control [25]. In this study, mitochondria were isolated from the virus-infected cells by centrifugation; a small amount of ER contacted the mitochondria and was not completely removed. It remains to be clarified whether BmGP37 is associated not only with mitochondria but also with ER.
It is exciting how BmGP37 affects mitochondrial ATP production and mitochondrial activity. Cheng et al [26] deleted the gp37 gene from the genome of Autographa californica multiple NPV (AcMNPV). They reported that the inactivation of gp37 had no effect on viral replication in cultured cells and virulence in larvae [26]. It is expected that gp37-deficient BmNPV and wild-type BmNPV will be used to compare the virus proliferation and infectivity. Recently, Zhang et al [27] identified twenty-four host proteins interacting with BmNPV BV particles in the B. mori midgut [27]. In these proteins, sixteen mitochondrial proteins were presumed to be involved in virus transportation, energy metabolism, apoptosis, and viral propagation [27]. However, in the infection cycle of NPV, ODV, not BV, enters the midgut cells first. Also, in this study, we identified that BmGP37 is an ODV-associated protein of BmNPV and that BmGP37 localizes to host mitochondria. Therefore, whether BmGP37 interacts with these host mitochondrial proteins in midgut cells should be clarified. In addition, it has been reported that the amount of RNAs of H+-ATPase and AK2, which are involved in energy metabolism, increases in virus-permissive larvae and decreases in virus-nonpermissive larvae after BmNPV inoculation [27]. Future studies need to investigate how the deletion or the overexpression of gp37 affects the transcription of these genes.
Localization of BmGP37 to host mitochondria
Mitochondrial proteins are divided into two groups: proteins whose N-terminal targeting signal is cleaved and proteins that have a non-cleavable internal targeting signal [28]. The proteins containing an N-terminal targeting signal exhibit a high composition of arginine and a few negatively charged residues. Tom20 and Tom22 in the TOM protein complex, which translocates the target protein to the mitochondria, recognize a local amphiphilic α-helical structure with hydrophobic residues on one face and positively charged residues on the opposite face [28,29,30]. The amino acid sequence of BmGP37 was analyzed using the MitoFates program [31] to search for mitochondrial target signals. As shown in Fig. 1B, although no TOM protein recognition sequence could be found in BmGP37, the N-terminal ten amino acid residues were identified as a positively charged amphiphilicity score region. Also, it was predicted that the serine at position 19 would be cleaved by mitochondrial processing peptidase. However, no α-helical structure was found in this region. In the future, the mitochondrial localization mechanism of the BmGP37 protein may be clarified by deleting the N-terminus or mutating the serine at position 19 of the BmGP37 protein.
Function of GP37 in ODV particles
The homolog of gp37 is entomopoxvirus (EPV)-encoded fusolin [19, 32]. Fusolin expressed in the host cell forms inclusion bodies and is orally internalized by the host larva and spheroid with the occlusion body. Fusolin has a chitin-binding ability and by acting on the peritrophic membrane composed of chitin it assists spheroid-occluded viral particles in passing through the peritrophic barrier [33,34,35].
NPV-derived GP37 has a chitin-binding ability [36]. For oral infection of NPV, GP37, and enhancin, a metalloproteinase, contribute to efficient infection of ODV released from OBs into the midgut cells [37, 38]. Oral inoculation of GP37 derived from Cydia pomonella granulovirus and OBs of Spodoptera exigua multiple nucleopolyhedrovirus (SeMNPV) increases the infectivity of SeMNPV [39]. Therefore, GP37, like fusolin, is thought to facilitate the passage of ODV through the peritrophic membrane. Interestingly, GP37 of OpMNPV and Choristoneura fumiferana nucleopolyhedrovirus form inclusion bodies in the same way as fusolin of EPV, but GP37 of Spodoptera littoralis multiple nucleopolyhedrovirus (SlMNPV) and AcMNPV do not form inclusion bodies. However, it is associated with viral particles and/or occlusion bodies [19, 32, 36, 40]. GP37 of SlMNPV is associated with ODV and BV, and GP37 of AcMNPV is not only a component of BV but is also included in OBs [32, 36]. In this study, GP37 of BmNPV was associated only with ODV (Fig. 5). Also, no signal was observed in BV that share a common nucleocapsid, suggesting that it is a component of the envelope fraction of ODV. Therefore, it is possible that ODV-associated BmGP37 promotes the passage of ODV released from OB through the peritrophic barrier to reach the midgut cells.
In summary, we identified BmGP37 for the first time as a baculovirus-derived protein that localizes to host mitochondria. BmGP37 was expressed at the late phase of viral infection and modified with N-linked glycans. Also, BmGP37 was found to be a protein associated with the ODV of BmNPV. Further analysis of the relationship between BmGP37 and the host’s energy metabolism and the localization mechanism of BmGP37 to host mitochondria will help clarify the infection mechanism of BmNPV.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Jehle JA, Blissard GW, Bonning BC, Cory JS, Herniou EA, Rohrmann GF, Theilmann DA, Vlak JM (2006) On the classification and nomenclature of baculoviruses: a proposal for revision. Arch Virol 151:1257–1266. https://doi.org/10.1007/s00705-006-0763-6
Herniou EA, Luque T, Chen X, Vlak JM, Winstanley D, Cory JS, O’Reilly DR (2001) Use of whole genome sequence data to infer baculovirus phylogeny. J Virol 75:8117–8126. https://doi.org/10.1128/jvi.75.17.8117-8126.2001
Rohrmann GF (2013) Baculovirus Molecular Biology, 4th ed; National Center for Biotechnology Information: Bethesda (MD), USA, 2019, Available from: https://www.ncbi.nlm.nih.gov/books/NBK543458/
Clem RJ (2005) The role of apoptosis in defense against baculovirus infection in insects. Curr Top Microbiol Immunol 289:113–129. https://doi.org/10.1007/3-540-27320-4_5
Ikeda M, Kobayashi M (1999) Cell-cycle perturbation in Sf9 cells infected with Autographa californica nucleopolyhedrovirus. Virology 258:176–188. https://doi.org/10.1006/viro.1999.9706
Iwanaga M, Shimada T, Kobayashi M, Kang W (2007) Identification of differentially expressed host genes in Bombyx mori nucleopolyhedrovirus infected cells by using subtractive hybridization. Appl Entomol Zool 42:151–159. https://doi.org/10.1303/aez.2007.151
Nobiron I, O’Reilly DR, Olszewski JA (2003) Autographa californica nucleopolyhedrovirus infection of Spodoptera frugiperda cells: a global analysis of host gene regulation during infection, using a differential display approach. J Gen Virol 84:3029–3039. https://doi.org/10.1099/vir.0.19270-0
Xue J, Qiao N, Zhang W, Cheng RL, Zhang XQ, Bao YY, Xu YP, Gu LZ, Han JJ, Zhang CX (2012) Dynamic interactions between Bombyx mori nucleopolyhedrovirus and its host cells revealed by transcriptome analysis. J Virol 86:7345–7359. https://doi.org/10.1128/JVI.07217-12
Cha HJ, Dalal NG, Pham MQ, Kramer SF, Vakharia VN, Bentley WE (2002) Monitoring foreign protein expression under baculovirus p10 and polh promoters in insect larvae. Biotechniques 32(5):986–992. https://doi.org/10.2144/02325bm02
Bernal V, Carinhas N, Yokomizo AY, Carrondo MJT, Alves PM (2009) Cell density effect in the baculovirus-insect cells system: A quantitative analysis of energetic metabolism. Biotechnol Bioeng 104:162–180. https://doi.org/10.1002/bit.22364
Lin YH, Tai CC, Brož V, Tang CK, Chen P, Wu CP, Li CH, Wu YL (2020) Adenosine receptor modulates permissiveness of baculovirus (budded virus) infection via regulation of energy metabolism in Bombyx mori. Front Immunol 11:763. https://doi.org/10.3389/fimmu.2020.00763
Iwanaga M, Kurihara M, Kobayashi M, Kang W (2002) Characterization of Bombyx mori nucleopolyhedrovirus orf68 gene that encodes a novel structural protein of budded virus. Virology 297:39–47. https://doi.org/10.1006/viro.2002.1443
Gomi S, Majima K, Maeda S (1999) Sequence analysis of the genome of Bombyx mori nucleopolyhedrovirus. J Gen Virol 80:1323–1337. https://doi.org/10.1099/0022-1317-80-5-1323
Iwanaga M, Takaya K, Katsuma S, Ote M, Tanaka S, Kamita SG, Kang W, Shimada T, Kobayashi M (2004) Expression profiling of baculovirus genes in permissive and nonpermissive cell lines. Biochem Biophys Res Commun 323:599–614. https://doi.org/10.1016/j.bbrc.2004.08.114
Ogata M, Kameshima Y, Hattori T, Michishita K, Suzuki T, Kawagishi H, Totani K, Hiratake J, Usui T (2010) Lactosylamidine-based affinity purification for cellulolytic enzymes EG I and CBH I from Hypocrea jecorina and their properties. Carbohydr Res 345:2623–2629. https://doi.org/10.1016/j.carres.2010.10.015
Tanaka Y, Suzuki T, Nakamura L, Nakamura M, Ebihara S, Kurokura T, Iigo M, Dohra H, Habu N, Konno N (2019) A GH family 28 endo-polygalacturonase from the brown-rot fungus Fomitopsis palustris: Purification, gene cloning, enzymatic characterization and effects of oxalate. Int J Biol Macromol 123:108–116. https://doi.org/10.1016/j.ijbiomac.2018.11.004
Iwanaga M, Shibano Y, Ohsawa T, Fujita T, Katsuma S, Kawasaki H (2014) Involvement of HSC70-4 and other inducible HSPs in Bombyx mori nucleopolyhedrovirus infection. Virus Res 179:113–118. https://doi.org/10.1016/j.virusres.2013.10.028
Katou Y, Ikeda M, Kobayashi M (2006) Abortive replication of Bombyx mori nucleopolyhedrovirus in Sf9 and High Five cells: Defective nuclear transport of the virions. Virology 347:455–465. https://doi.org/10.1016/j.virol.2005.11.043
Gross CH, Wolgamot GM, Russell RL, Pearson MN, Rohrmann GF (1993) A 37-kilodalton glycoprotein from a baculovirus of Orgyia pseudotsugata is localized to cytoplasmic inclusion bodies. J Virol 67:469–475. https://doi.org/10.1128/JVI.67.1.469-475.1993
Phanis CG, Miller DP, Cassar SC, Tristem M, Thiem SM, O’Reilly DR (1999) Identification and expression of two baculovirus gp37 genes. J Gen Virol 80:1823–1831. https://doi.org/10.1099/0022-1317-80-7-1823
Copeland DE, Dalton AJ (1959) An association between mitochondria and the endoplasmic reticulum in cells of the pseudobranch gland of a teleost. J Biophys Biochem Cytol 25:393–396. https://doi.org/10.1083/jcb.5.3.393
Friedman JR, Lackner LL, West M, DiBenedetto JR, Nunnari J, Voeltz GK (2011) ER tubules mark sites of mitochondrial division. Science 334:358–362. https://doi.org/10.1126/science.1207385
Zhou R, Yazdi AS, Menu P, Tschopp J (2011) A role for mitochondria in NLRP3 inflammasome activation. Nature 469:221–225. https://doi.org/10.1038/nature09663
Hamasaki M, Furuta N, Matsuda A, Nezu A, Yamamoto A, Fujita N, Oomori H, Noda T, Haraguchi T, Hiraoka Y, Amano A, Yoshimori T (2013) Autophagosomes form at ER-mitochondria contact sites. Nature 495:389–393. https://doi.org/10.1038/nature11910
Booth DM, Enyedi B, Geiszt M, Várnai P, Hajnóczky G (2016) Redox nanodomains are induced by and control calcium signaling at the ER-mitochondrial interface. Mol Cell 63:240–248. https://doi.org/10.1016/j.molcel.2016.05.040
Cheng XW, Krell PJ, Arif BM (2001) P34.8 (GP37) is not essential for baculovirus replication. J Gen Virol 82:299–305. https://doi.org/10.1099/0022-1317-82-2-299
Zhang SZ, Zhu LB, Yu D, You LL, Wang J, Cao HH, Liu YX, Wang YL, Kong X, Toufeeq S, Xu JP (2020) Identification and functional analysis of BmNPV-interacting proteins from Bombyx mori (Lepidoptera) larval midgut based on subcellular protein levels. Front Microbiol 30:1481. https://doi.org/10.3389/fmicb.2020.01481
Chacinska A, Koehler CM, Milenkovic D, Lithgow T, Pfanner N (2009) Importing mitochondrial proteins: machineries and mechanisms. Cell 138:628–644. https://doi.org/10.1016/j.cell.2009.08.005
Abe Y, Shodai T, Muto T, Mihara K, Torii H, Nishikawa S, Endo T, Kohda D (2000) Structural basis of presequence recognition by the mitochondrial protein import receptor Tom20. Cell 100:551–560. https://doi.org/10.1016/s0092-8674(00)80691-1
Yamano K, Yatsukawa Y, Esaki M, Hobbs AE, Jensen RE, Endo T (2008) Tom20 and Tom22 share the common signal recognition pathway in mitochondrial protein import. J Biol Chem 283:3799–3807. https://doi.org/10.1074/jbc.M708339200
Fukasawa Y, Tsuji J, Fu SC, Tomii K, Horton P, Imai K (2015) MitoFates: improved prediction of mitochondrial targeting sequences and their cleavage sites. Mol Cell Proteomics 14:1113–1126. https://doi.org/10.1074/mcp.M114.043083
Vialard JE, Yuen L, Richardson CD (1990) Identification and characterization of a baculovirus occlusion body glycoprotein which resembles spheroidin, an entomopoxvirus protein. J Virol 64:5804–5811. https://doi.org/10.1128/JVI.64.12.5804-5811.1990
Mitsuhashi W, Sato M, Hirai Y (2000) Involvement of spindles of an entomopoxvirus (EPV) in infectivity of the EPVs to their host insect. Arch Virol 145:1465–1471. https://doi.org/10.1007/s007050070103
Takemoto Y, Mitsuhashi W, Murakami R, Konishi H, Miyamoto K (2008) The N-terminal region of an entomopoxvirus fusolin is essential for the enhancement of peroral infection, whereas the C-terminal region is eliminated in digestive juice. J Virol 82:12406–12415. https://doi.org/10.1128/JVI.01605-08
Wijonarko A, Hukuhara T (1998) Detection of a virus enhancing factor in the spheroid, spindle, and virion of an entomopoxvirus. J Invertebr Pathol 72:82–86. https://doi.org/10.1006/jipa.1998.4756
Li Z, Li C, Yang K, Wang L, Yin C, Gong Y, Pang Y (2003) Characterization of a chitin-binding protein GP37 of Spodoptera litura multicapsid nucleopolyhedrovirus. Virus Res 96:113–122. https://doi.org/10.1016/s0168-1702(03)00179-5
Wang P, Granados RR (1997) An intestinal mucin is the target substrate for a baculovirus enhancin. Proc Natl Acad Sci USA 94:6977–6982. https://doi.org/10.1073/pnas.94.13.6977
Liu X, Fang W, Fan R, Zhang L, Lei C, Zhang J, Nian W, Dou T, An S, Zhou L, Sun X (2019) Granulovirus GP37 facilitated ODVs cross insect peritrophic membranes and fuse with epithelia. Toxins (Basel) 11:145. https://doi.org/10.3390/toxins11030145
Liu X, Ma X, Lei C, Xiao Y, Zhang Z, Sun X (2011) Synergistic effects of Cydia pomonella granulovirus GP37 on the infectivity of nucleopolyhedroviruses and the lethality of Bacillus thuringiensis. Arch Virol 156:1707–1715. https://doi.org/10.1007/s00705-011-1039-3
Li X, Barrett J, Pang A, Klose RJ, Krell PJ, Arif BM (2000) Characterization of an overexpressed spindle protein during a baculovirus infection. Virology 268:56–67. https://doi.org/10.1006/viro.1999.0138
Acknowledgements
This work was supported by grants from JSPS KAKENHI (grant no. 20K06069) to MI. The authors are grateful to Dr. Ikeda for the kind gift of anti-VP39 antibody. The authors would like to thank Enago (www.enago.jp) for the English language review.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. The experiments were conceived and designed by SF and MI. The experiments were performed by SF, KF, TS, SK, and MI. The paper was written by MI and SK.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study was performed in accordance with institutional committee protocols of Utsunomiya University.
Informed consent
There were no human participants in this study.
Research involving human and animal participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Edited by Sassan Asgari.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fujimoto, S., Fujimaki, K., Suzuki, T. et al. Expression and localization of Bombyx mori nucleopolyhedrovirus GP37. Virus Genes 59, 457–463 (2023). https://doi.org/10.1007/s11262-023-01983-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-023-01983-3