Abstract
Staphylococcus aureus is one of the most dreadful infectious agents, responsible for high mortality and morbidity in both humans and animals. The increased prevalence of multidrug-resistant (MDR) Staphylococcus aureus strains has limited the number of available treatment options, which calls for the development of alternative and effective modalities against MDR S. aureus. Endolysins are bacteriophage-derived antibacterials, which attack essential conserved elements of peptidoglycan that are vital for bacterial survival, making them promising alternatives or complements to existing antibiotics for tackling such infections. For developing endolysin lysin-methicillin-resistant-5 (LysMR-5) as an effective antimicrobial agent, we evaluated its physical and chemical characteristics, and its intrinsic antibacterial activity against staphylococcal strains, including methicillin-resistant Staphylococcus aureus (MRSA). In this study, we cloned, expressed, and purified LysMR-5 from S. aureus phage MR-5. In silico analysis revealed that LysMR-5 harbors two catalytic and one cell wall-binding domain. Biochemical characterization and LC–MS analysis showed that both catalytic domains were active and had no dependence on divalent ions for their action, Zn2+ exerted a negative effect. The optimal lytic activity of the endolysin was at 37 °C/pH 7.0 and in the presence of ≥ 300 mM concentration of NaCl. Circular dichroism (CD) demonstrated a loss in secondary structure with an increase in temperature confirming the thermosensitive nature of endolysin. Antibacterial assays revealed that LysMR-5 was active against diverse clinical isolates of staphylococci. It showed high lytic efficacy against S. aureus ATCC 43300, as an endolysin concentration as low as 15 µg/ml was sufficient to achieve maximum lytic activity within 30 min and it was further confirmed by scanning electron microscopy. Our results indicate that rapid and strong bactericidal activity of LysMR-5 makes it a valuable candidate for eradicating multidrug-resistant S. aureus.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Staphylococcus aureus has long been considered as a major nosocomial pathogen, associated with various infections ranging from skin and soft tissue to other insidious infections like meningitis, osteomyelitis, and pneumonia [1, 2]. Further, rapid development of resistance by S. aureus strains against broad spectrum of antibiotics has posed additional limits to treat these infections. Besides controlling misuse of antibiotics, development of novel antimicrobials that are refractory to resistance generation is essential to eradicate MDR pathogens [3,4,5,6]. In recent years maximum attention has been directed towards bacteriophage endolysins, which are being investigated as a valuable alternative antimicrobial strategy for containing multidrug-resistant Gram-positive bacteria [7,8,9,10]. Endolysins are produced by phages in a bid to escape from their host at the end of phage lytic cycle, by targeting specific bonds in peptidoglycan, resulting in osmotic lysis of the cell and release of phage progeny [11,12,13]. These hydrolases can rapidly kill targeted Gram-positive bacteria even when applied exogenously, since they are able to make direct contact with the cell wall carbohydrates and peptidoglycan layers, which are inaccessible in case of Gram-negative bacteria due to the presence of an outer membrane. Despite repeated attempts, resistance towards specific endolysin has not been reported as yet [14,15,16]. These characteristics have prompted scientists to use them as an antimicrobial agent against antibiotic-resistant bacteria [17, 18].
Endolysins against Gram-positive bacteria are modular in nature, often with one or more enzymatically active domain (EAD) and a cell wall-binding domain (CBD) that recognizes epitopes on the surface of susceptible organisms, contrary to globular architecture of Gram-negative endolysins, which comprises of EAD only and lacks CBD [19, 20]. An enzymatically active catalytic domain can be classified into three basic groups on the basis of specific bonds which they cleave in peptidoglycan: (1) glycosidases, cleave between sugar backbone of peptidoglycan, (2) endopeptidase, cleave bond between two adjacent amino acids of either stem peptide or interpeptide bridge, and (3) amidases, hydrolyze the bond between the sugar (MurNac) and the peptide (L-Ala) [21]. One of the reasons that endolysins are considered as good antimicrobial candidates is that they offer many advantages over antibiotics. Firstly, their genus or species specificity protects commensal flora as opposed to broad range antibiotics which are detrimental to normal flora along with targeted organism. This specificity of endolysin activity reduces the chances of development of resistance, which is generally seen with antibiotics due to their broad host range activity. Another advantage is lesser likelihood of resistance development against endolysins, which could be attributed to co-evolution of phages with their host bacteria, thus requiring endolysin to target highly conserved bonds within bacterial peptidoglycan. Modification of these bonds is believed to be detrimental to the host organism [11, 22]. Another reason for low resistance development is their mode of action (lysis from without) as endolysins are able to attack their target peptidoglycan from outside only, and this in turn decreases the possibility of development of resistance mechanisms [7, 17, 23]. Endolysins are also considered safe, as they do not attack mammalian cells [17, 24,25,26]. Many endolysins against Gram-positive pathogens have been reported during the last few years and shown to be successful in combating infections, in vitro and in vivo, however, only a few of these have been fully characterized [19, 24, 25, 27,28,29]. To apply LysMR-5 and other endolysins for the management of drug-resistant infections, investigation of safety, physical, biochemical and antibacterial properties in an experimental setting is required. Previously, we have reported the isolation, characterization and in vitro and in vivo therapeutic efficacy of a novel S. aureus phage MR-5. Phage MR-5 belongs to Order Caudovirales and Family Herelleviridae [30]. Thus, the aim of the present study was to understand the properties of LysMR-5, the endolysin of phage MR-5, and to demonstrate its therapeutic utility for biomedical applications. To this end, we cloned, over-expressed, purified, and evaluated the intrinsic antibacterial efficacy of LysMR-5 towards MRSA.
Materials and methods
Bacterial strains, bacteriophage, and susceptibility testing
Staphylococcus aureus ATCC 43300 (MRSA) was used for all the antibacterial assays performed in this study. Reference strains of Staphylococcus aureus were from ATCC, Mannasse. These included: S. aureus ATCC 43300 (MRSA), S. aureus ATCC 33591 (MRSA), S. aureus ATCC 25923 (MSSA), and S. aureus ATCC 29213 (MSSA). Staphylococcus epidermidis MTCC 3810 was from MTCC, Chandigarh, India. Clinical isolates of S. aureus (MRSA) and S. epidermidis procured from the Postgraduate Institute of Medical Education and Research (PGIMER), Chandigarh, India, were used. Sensitivity of these S. aureus strains was tested using battery of six lytic phages (MR-1, MR-2, MR-3, MR-4, MR-5, and MR-6) that were isolated previously in our laboratory using S. aureus ATCC 43300 as the primary host strain [30] and were stored as lab phage stock. These phages were lytic DNA phages belonging to tailed Herelleviridae family. Sensitivity of the S. aureus clinical strains towards these phages was determined by spot assay. Briefly, all S. aureus strains were grown up to mid-exponential phase in Brain Heart Infusion (BHI) broth, harvested and spread on nutrient agar plates to form a lawn. On freshly prepared bacterial lawn, 10 µl of each phage suspension (109 PFU/ml) was spotted, plates were air-dried for 10 min, incubated at 37 °C for 24 h and were observed for the presence of lytic zones after incubation. Streptococcus pneumoniae D39 was procured from MTCC IMTECH, Chandigarh. Gram-negative bacteria used to observe host range of endolysin lytic activity were, Klebsiella pneumoniae 43816, Klebsiella pneumoniae ATCC B5055, Pseudomonas aeruginosa strain PAO1 and Salmonella enterica serotype Typhi. S. aureus specific phage, MR-5 which had been previously isolated and characterized in our laboratory was used [30]. Escherichia coli strains DH10β and BL21 (DE3) were used for molecular cloning and protein overexpression, respectively. Staphylococcal and E. coli strains were grown at 37 °C in Brain Heart Infusion broth (BHI, Hi-media, Ref. #M210) and Luria broth (LB, Hi-media,Ref. #M575), respectively. The LB medium was supplemented with kanamycin (50 µg/ml; SRL,India, Ref. #99311) when necessary.
Cloning of the endolysin gene from phage MR-5 genome
Genomic DNA of phage MR-5 was isolated by the method of Sambrook et al. [31] and was used as a template for performing polymerase chain reaction (PCR). The forward primer AAAGGATCCATGGCTAAGACTCAAGCA and the reverse primer AAACTCGAGCTATTTGAATACTCCCCAGGC were designed according to the LysGH15 gene sequence (GenBank accession number HMO15284) using an online tool “OligoEvaluator™”. BamHI and XhoI were the restriction sites added to forward and reverse primers, respectively. Pfu DNA polymerase (Thermo Scientific) was used for PCR amplification. The resulting PCR product was sequenced (BioServe Biotechnologies (India) Pvt. Ltd.) and BLAST analysis was performed to confirm the amplification of complete endolysin gene. To clone the endolysin gene (LysMR-5 gene), the PCR amplified product was digested with BamHI and XhoI restriction enzymes and ligated into the pET28a expression vector. E. coli BL21 (DE3) competent cells were prepared by calcium chloride method and the resulting recombinant plasmid (pET28a-LysMR-5) was transformed into these cells using heat-shock method for expression of recombinant endolysin as N-terminal fusion to His-tag. Colony PCR, double digestion and nucleotide sequencing were performed (using T7 promoter primer and endolysin LysMR-5 specific reverse primer) to confirm gene cloning [32].
Overexpression and purification of LysMR-5 endolysin
The E. coli BL21 (DE3) cells, harboring recombinant plasmid pET28a-LysMR-5, were grown in LB medium containing kanamycin (50 µg/ml) at 37 °C until the OD600 reached 0.5. To express the recombinant endolysin, culture was induced with 1 mM isopropyl-β-d-thiogalactoside for 6 h at 25 °C. Cells were harvested by centrifugation and suspended in lysis buffer [25 mM HEPES buffer, 300 mM NaCl, 10 mM imidazole, 10% (vol/vol) glycerol, 1 mM PMSF]. These were lysed by sonication (8 s on, 15 s off, at an amplitude of 60 for 15 min). The cell debris were removed by centrifugation (10,000×g, 30 min, 4 °C), and supernatant containing soluble His-tagged recombinant protein purified with nickel-nitrilotriacetic acid (Ni–NTA) resin column (Qiagen,India, Cat. #30,210). Washing of the column was done with 40 ml wash buffer [25 mM HEPES buffer, 300 mM NaCl, 10% (vol/vol) glycerol and 35 mM imidazole] and finally the protein was eluted with HEPES buffer [25 mM HEPES buffer, 300 mM NaCl, 10% (vol/vol) glycerol] containing 250 mM imidazole. Eluted protein was dialyzed against HEPES buffer (6L) at 4 °C. Buffer was replaced every 6 h thrice, dialyzed protein was concentrated using Amicon ultra centrifugal filters (30 kDa MWCO, Ref.#UFC903008) and stored at – 20 °C. The purity of the protein was confirmed on 12% SDS-PAGE gel [33] and protein concentration was determined by bicinchoninic acid (BCA) protein estimation kit (G-Biosciences (India) Pvt. Ltd, Cat. #786-570).
In silico analysis of endolysin sequence and structure modeling
For structural analysis of LysMR-5, the nucleotide sequence of endolysin was translated to amino acid sequence by Protparam server (https://web.expasy.org/protparam/). The theoretical isoelectric point of the protein was calculated using IPC (https://isoelectric.org/calculate.php). Further, BLASTp analysis of amino acid sequence of LysMR-5 was done with National Center for Biotechnology Information (NCBI) database (https://blast.ncbi.nlm.nih.gov/Blast.cgi) to check the similarity of LysMR-5 with other reported endolysins. The endolysin sequences showing more than 90% identity with LysMR-5 were selected and multiple alignment of these sequences was performed using MULTALIN software [34]. Presence of conserved domains in endolysin structure was analyzed by conserved domain database server at NCBI (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi). Secondary structure prediction was done using CFSSP server (Chou and Fasman Secondary structure prediction server) and SOPMA server [35]. Protein structures with high identity to LysMR-5 were extracted from PDB database using HHpred and secondary structure analysis was performed. Analysis of secondary structure elements was performed by circular dichroism (CD) spectroscopy also. LysMR-5 (5.45 µM) was dialyzed using buffer containing 5 mM HEPES (pH 7.4), 100 mM NaCl and 2% glycerol. A far UV CD spectrum was recorded on V100 chirascan spectrometer (Applied Photophysics) from 200 to 250 nm at a scanning rate of 10 nm/min using 1 mm path length cuvette at 25 °C and averaged over five scans. Raw data were converted to mean residue ellipticity (MRE) and plotted against wavelength and temperature, respectively.
where Φobs is the observed ellipticity (in degrees), d is path length (in centimeters), C is protein concentration (in molar), and n is the total number of amino acid residues in the protein.
For constructing three-dimensional (3D) structure of all three domains of LysMR-5 (CHAP, Amidase-2, and Sh3b), homologous proteins were searched in the PDB database against LysMR-5 sequence. Homology modeling on the basis of these selected templates was done using the SWISS-MODEL server [36]. The cartoon representation was done using Chimera [37]. Active site prediction of the individual domain structure was done using model.pdb files in CASTp software [38]. To validate the stability and quality of generated structures, 3D PDB models were subjected to evaluation by PROCHECK (Ramachandran plot) [39].
Antibacterial activity and zymogram assay
The lytic activity of endolysin (LysMR-5) was checked against Staphylococcus aureus ATCC 43300, Staphylococcus epidermidis 3810, Streptococcus pneumoniae strain D39, a panel of clinical strains of Staphylococcus aureus and Staphylococcus epidermidis via plate lysis assay. All the strains were grown up to mid-exponential phase (OD600 0.6) and 100 µl of the suspension was spread plated on nutrient agar. 20 µl (10 µg/spot) of endolysin or 20 µl of HEPES buffer (negative control) was spotted onto the plates. The plates were air-dried and incubated overnight at 37 °C. Cleared spots on the bacterial lawn depicting cell lysis were scored next day. To further check the host range of endolysin, lytic activity against some Gram-negative bacteria was also assessed. These included, Klebsiella pneumoniae 43816, Klebsiella pneumoniae ATCC B5055, Pseudomonas aeruginosa strain PAO1 and Salmonella enterica serotype Typhi. All bacterial strains were grown till mid-exponential phase, pelleted, washed, and resuspended to have a cell density of 108 CFU/ml in activity buffer (50 mM phosphate buffer, 300 mM NaCl, pH 7.2) containing 100 mM EDTA. The latter was added in order to permeabilize outer membrane of Gram-negative bacteria by incubating for 5 min at 37 °C. Cells were washed three times to remove residual EDTA, resuspended in 950 μl of activity buffer [40]. The endolysin was added (25 µg/ml) to each reaction and incubated at 37 °C for 30 min, treatment with HEPES buffer served as negative control. Serial dilutions of each reaction were plated, incubated at 37 °C and CFU counted next day. Antibacterial activity was calculated, as the relative decrease in CFU count in the test sample in comparison with negative control. Zymogram assay was performed to check the activity of LysMR-5. Briefly, the SDS-PAGE was performed, in which 0.2% of autoclaved S. aureus 43300 cells were embedded in 12% polyacrylamide stacking gel during polymerization and lanes were loaded with 10 µg of purified endolysin along with a protein marker (BLUeye prestained protein ladder, GeneDireX, Inc., Cat. #PM007-0500). After electrophoresis, gel was soaked in distilled water (30 min) to remove SDS. Renaturation of endolysin was performed in 50 mM Tris–HCl, pH 8.0, 1% Triton X-100 and incubated overnight at 37 °C. To detect the areas of lytic activity, gels were stained with 0.1% methylene blue in 0.01% KOH and subsequently de-stained with distilled water [41].
Biochemical characterization of LysMR-5
The optimum temperature for activity and thermostability of endolysin was evaluated at different temperatures (25, 30, 37, 45, 50 °C). Effect of temperature on endolysin stability was evaluated in terms of its effect on lytic activity and secondary structure, as these two parameters are interrelated, and any change in endolysin structure can alter its catalytic activity. To check the optimum temperature for its activity, an antibacterial assay was performed at different temperatures. Briefly, S. aureus cells grown up to mid-exponential phase were suspended in activity buffer to achieve a cell count of approx. 108 CFU/ml and the reaction was initiated by addition of endolysin (25 µg/ml) or buffer (control) and incubated for 30 min. Samples were processed and lytic activity was analyzed on the basis of decrease in CFU count as described in previous section for antibacterial assay against Gram-negative bacteria. For evaluating thermostability of endolysin, it was pre-incubated at different temperatures for 30 min in the absence of bacteria, and the residual lytic activity was assessed at 37 °C under the conditions described for antibacterial assay to check optimum temperature for endolysin activity. Control samples were not pre-incubated. Thermal stability of endolysin was also assessed by CD spectropolarimetry. Secondary structure spectra was collected at various temperatures (25 °C to 70 °C) under conditions described above for secondary structure analysis. For thermal denaturation, molar ellipticity at wavelength 222 nm was monitored from 25 to 70 °C at a denaturation rate of 1 °C/min. For renaturation, same parameters were used as mentioned above and temperature ranged from 70 to 25 °C. Effect of pH on endolysin activity was determined by performing antibacterial assay in buffers with different pH values [50 mM sodium acetate buffer (pH 4.0–6.0), 50 mM Phosphate buffer (pH 7 and 8) and 50 mM Tris–HCl buffer for pH 9]. For stability assays, endolysin (250 µg/ml) was pre-incubated for 30 min in buffer of respective pH and antibacterial assay performed at pH 7.0 at 37 °C. The effect of addition of different concentrations of NaCl (0–800 mM) in activity buffer (phosphate buffer pH 7.2) was checked on lytic activity of LysMR-5. The data for biochemical characterization is represented in antibacterial activity percentage, in comparison with highest decrease in bacterial log units (after treatment with LysMR-5), taken as 100% activity. The effect of divalent metal ions on the lytic activity of endolysin was evaluated. Endolysin LysMR-5 was incubated either with 5 mM EDTA for 30 min or 100 mM for 10 min at 37 °C in order to chelate any bound metal ions and dialyzed overnight against activity buffer. The activity was assayed using antibacterial assay after addition of either Ca2+, Zn2+, Mg2+ or Mn2+ at a final concentration of 0, 0.1, 1, and 5 mM to 5 mM EDTA-treated endolysin and compared with lytic activity of EDTA untreated endolysin. All the experiments were performed in triplicate.
Lytic activity of LysMR-5 against MRSA
To determine dose-dependent effect of endolysin, S. aureus 43300 cells were prepared as described for evaluation of optimum temperature using antibacterial assay (108 CFU/ml). Different endolysin concentrations (0, 1, 5, 10, 15, 20, 30, 40, and 50 µg/ml) were added to separate tubes and incubated at 37 °C for 30 min. The samples were serially diluted and plated. The lytic activity was assessed on the basis of decrease in bacterial count after treatment with endolysin. Time-dependent bacterial killing by endolysin was also analyzed. Briefly, 108 CFU/ml of cells were taken in different tubes and 15 µg/ml of endolysin was added to each tube, an equal volume of buffer (HEPES buffer) (no endolysin) was added to the control tube. Samples were taken out at 1, 5, 15, 30, 45, and 60 min, serially diluted and plated to assess the bacterial killing of endolysin treated and untreated cells. Similarly, the potential of LysMR-5 to kill S. aureus 43300 cells at different stages of the growth cycle was also tested [42]. S. aureus cells were harvested at early (OD600 0.2), mid-exponential (OD600 0.55), late-exponential (OD600 2), and stationary phase (overnight culture), resuspended to achieve a cell density of 108 CFU/ml in activity buffer and treated with 15 µg/ml of endolysin for 30 min at 37 °C. The antibacterial activity was assessed on the basis of decrease in CFU/ml. The highest decrease in viable cell count was taken as 100% activity. All the above experiments were performed in triplicate. Effect on bacterial viability and morphology after endolysin treatment was also observed using field emission scanning electron microscope (FESEM). Briefly, S. aureus 43300 cells were grown to mid-log phase, harvested, and resuspended in activity buffer (approx. 108 cells/ml). Endolysin (15 μg/ml) treatment was given for 3 min at 37 °C. Control was treated similarly without the addition of endolysin. The reaction was stopped with addition of 2.5% (wt/vol) glutaraldehyde, cells washed with PBS and dehydrated with graded ethanol series. Samples were critical-point dried, gold-coated, and observed at SAIF-CIL, Panjab University, Chandigarh, India.
Enzymatic specificity of LysMR-5
Three different assays (endopeptidase, amidase, and glycosidase) were performed to analyze the type of activities present in the endolysin’s catalytic domains. For measuring endopeptidase activity, free amino groups of purified S. aureus peptidoglycan (Sigma-Aldrich, Cat. #77140) were acetylated as described by Pritchard et al. [43]. Briefly, to S. aureus peptidoglycan in water (1 mg/ml) an equal volume of saturated aqueous sodium acetate solution was added and acetylation was done by adding 7 × 40 µl aliquots of acetic anhydride over a period of 1 h with constant stirring. The acetylated peptidoglycan was washed three times with water and resuspended in 1 ml activity buffer. Reaction was initiated by the addition of 25 μg/ml endolysin and incubated for 4 h at 37 °C. The controls (buffer+ peptidoglycan, buffer+ endolysin) were included for comparison. Supernatant was collected and passed through a 10,000 MW filter (Amicon, Merck). Presence of free amino groups in the filtrate by endolysin digestion were assessed by trinitrobenzene sulfonic acid (TNBS) method [44], Serine (0.02 to 5 mM) (Sigma-Aldrich, Cat #S4500) was used as standard. Amidase assay was performed by method of Hazenberg and de Visser [45]. Briefly, to peptidoglycan dissolved in activity buffer (500 µg/ml), endolysin (25 μg/ml) was added and reaction was incubated at 37 °C for 4 h. Pure peptidoglycan suspension and endolysin in activity buffer were used as negative controls. In 100 μl of supernatant, muramic acid content was determined by adding 1 ml of conc. H2SO4, boiled for 5 min and cooled in ice water. Then 10 µl of 0.16 M CuSO4·5H2O and 20 µl of 0.09 M p-hydroxydiphenyl (Sigma-Aldrich, Cat. #134341) in ethanol were added, incubated at 30 °C for 30 min. Absorbance was taken at 570 nm and compared with standard curve prepared with muramic acid (Sigma-Aldrich, Cat. #M2503) using various concentrations 50, 100, 200, 300, 400, 500, 600, 700, 800, and 1000 µg/ml. Glycosidase assay for reducing sugar analysis was performed according to Park- Jhonson method [46].
Cleavage site determination of digested peptidoglycan through ESI–MS analysis
As previously described for endopeptidase assay, acetylated peptidoglycan of S. aureus was digested with LysMR-5 (25 µg/ml) for 4 h at 37 ºC. Undigested peptidoglycan was pelleted by centrifugation and the supernatant was passed through a 10,000 MW filter (Amicon, Merck, Ref. # UFC201024). Filtrate so obtained was then analyzed for the presence of muropeptides generated by endolysin digestion. Separation of soluble muropeptides was done by rp-HPLC using C18 cartridge. Peaks were analyzed by Liquid Chromatography-Mass Spectrometry (LTQ XL Linear Ion Trap Mass Spectrometer, Thermo Fisher Scientific, India). Undigested peptidoglycan samples were treated similarly, analyzed, and served as a negative control [47].
Quantification of LysMR-5 activity by Turbidity reduction assay (TRA)
To quantify the endolysin activity (enzyme units) against S. aureus 43300, turbidity reduction assay was performed as described previously [48]. Briefly, mid-exponential-phase cells were centrifuged, washed and resuspended in activity buffer to achieve a cell density of OD600 1.0. The reaction was initiated by adding 100 μl of bacterial suspension to 100 μl of endolysin (concentration range of 0, 0.15, 0.31, 0.62, 1.25, 2.5, 5, and 10 μg) in a 96-well microtiter plate (all the endolysin concentrations were added in triplicate). Bacterial suspension in buffer (no endolysin) served as a negative control. The decrease in OD600 of bacterial suspension was measured every 40 s spectrophotometrically in a BioTek Synergy H1 microplate reader for up to 30 min at 37 °C. The experiment was performed three times in triplicate. The enzyme activity was calculated on the basis of steepest slope of the lysis curve, calculated from the linear regression of the saturation curve (relationship between lysis activity and enzyme concentration) as suggested by Briers et al. [48]. Enzyme activity units were calculated and expressed as enzyme unit per mg using the following formula.
Endolysin enzyme unit was also calculated using another TRA for comparison. Various concentrations of endolysin (15, 10, 5, 1, 0.3, and 0.1 µg) were added in bacterial suspension having an OD600 of 1.2, decrease in turbidity was measured up to 30 min at 37 °C in microplate reader relative to control wells (no endolysin added). The enzyme unit was defined as the highest dilution that caused a 50% decrease in absorbance at OD600 after 15 min at 37 °C relative to control wells and calculated as described previously [49].
Results
Sequence and structural analysis of LysMR-5
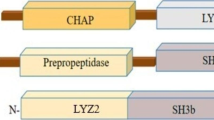
BLASTp and multiple sequence alignment of endolysin LysMR-5 gene showed that it had higher similarity with other fully characterized Staphylococcus endolysins like LysGH15 (99%), LysK (99%) and endolysins from phage JD007, qdsa002 (99%), S25-4, Sb-1, P108, phiIPLA-RODI, MCE-2014, vB_Sau_CG, phiSA039 (98%), vB_Sau_S24 (97%), and vB_Sau_Clo6 (97%) (Fig. 1). Conserved domain analysis using the CDD search system at NCBI predicted that LysMR-5, a 529 amino acid long endolysin is a modular protein, having two catalytic domains, a cysteine, histidine-dependent amidohydrolase/peptidase or CHAP domain(45–136 aa), a Zn2+-dependent amidase domain or Ami_2 (204–335 aa) and one cell wall-binding Sh3B_5 domain (409-475aa) (Fig S.1A). Predicted secondary structure by CFSSP showed that protein consisted of mostly alpha-helix (52.9%) and beta-sheet regions (57.4%) separated by turns and coils (Fig S.1B). SOPMA-based analysis (Fig S.1C) revealed the presence of alpha-helix (18.79%), beta-turns (9.09%), extended strand (25.45%), and coils (46.67%). Circular dichroism (CD) spectropolarimetry was performed to explore the secondary structure of LysMR-5 and to validate the results of predicted structures (Fig. 2a). Since a protein with alpha-helix secondary structure shows negative minima at 208 and 222 nm and a peak at 193 nm whereas a protein consisting predominantly of beta-sheet secondary structure displays a negative minima at 218 nm and a peak at 195 nm. The CD spectrum of LysMR-5 showed two dichroic minima at 210 nm and 222 nm with negative ellipticity of − 1300, this shift in dichroic minima and ellipticity are slightly lower than expected from a completely α-helical protein, suggested that LysMR-5 could be a mixture of both α-helix and ß-sheets in a well-folded native state (Fig. 2A). Deconvolution of the spectrum using K2D3 program also showed that LysMR-5 contained 27% of α-helix and 21% of ß-sheets. This data endorses the predicted structure models for LysMR-5, since these structures also consisted of both α-helices and β-sheets. The CHAP domain of endolysin LysGH15 (PDB ID 4OLK A) has 100% identity with LysMR-5 CHAP domain and was used as a template for 3D structure generation of LysMR-5 CHAP domain. Predicted model showed the presence of Ca2+ inside the domain, consisting of three ɑ-helices and six β-sheets (Fig. 2b). For Amidase-2 domain of the LysGH15, 99% identity (PDB ID 4OLS A) was used as a template for building 3D model of Ami-2 domain. It showed that Zn was bound to the center of the domain with seven ɑ-helices and four β-sheets (Fig. 2C). LysF1 sh3b domain of Lys K, 99% identity (PDB ID 5O1Q A) was used as a template for Sh3b, predicted structure consisted of nine β-sheets and no ɑ-helix (Fig. 2d). Amino acid residues present in the active site of all three domains are listed in Table 1. Stability of the generated 3D models was examined by Ramachandran plot analysis (Fig S.2). More than 90% of residues of all three domains were present in most favored and additional allowed regions. These characteristics indicated that built models are highly stable and of good quality.
Multiple Sequence alignment analysis of LysMR-5. Multiple sequence alignment of LysMR-5 sequence compared to Staphylococcal phage endolysins having > 90% amino acid sequence identity. Coloring of letters indicates level of amino acid conservation between sequences (red, completely conserved; blue, partially conserved; black, not conserved)
CD spectra, tertiary structure and active site prediction of LysMR-5. a Secondary structure of LysMR-5 recorded as mean residue ellipticity from 200 to 250 nm at 25 ºC. b, c, and d 3D structure of CHAP, Amidase-2, and Sh3B_5 domain of LysMR-5. Along with the respective 3D structures, active sites are also shown (1, 2, 3). The models were generated using the SWISS-MODEL server, cartoon representation of the model was done using Chimera and the active site prediction by CASTp software. The colors pink, yellow, and red represent secondary structure elements alpha-helices, beta-strands, and loops, respectively. The Zn2+ in the amidase-2 domain and Ca2+ in the CHAP domain are shown in green and blue color, respectively
Antimicrobial activity of LysMR-5 against S. aureus and other bacteria
Expression of recombinant LysMR-5 was induced in E. coli BL21 (DE3) cells harboring recombinant plasmid (LysMR-5 gene+ pET 28a) at 25 °C. Recombinant protein was purified and found to be of 58 kDa [corresponding to the calculated mass (58.2), 529 aa] as assessed by SDS-PAGE analysis (Fig. 3a). Purified protein was found to be active in zymogram assay. A clear band indicating bacterial lysis was observed in the lane containing endolysin, corresponding to the calculated molar mass and position of purified LysMR-5 in SDS-PAGE (Fig. 3b). In plate lysis assay, clear zones of lysis on a lawn of S. aureus ATCC 43300 cells, indicated bacterial sensitivity towards LysMR-5 (Fig S.3). In another experiment approx. 4 log reduction (a decrease from 8.52 ± 0.14 log units to 4.32 ± 0.267 log units) in the number of S. aureus ATCC 43300 cells, was observed within 30 min of incubation at 37 °C. Lytic spots on the bacterial lawn of all tested strains of Staphylococcus aureus and 95% strains of S. epidermidis were observed. However, it was seen that LysMR-5 was unable to kill Streptococcus pneumoniae D39 and any of the Gram-negative bacteria tested (Table S.3). When tested against S. aureus cells in different stages of growth cycle, LysMR-5 was found to be highly active against cells in early (100% activity), mid (98 ± 4.2% activity), and late-exponential phase (89 ± 6% activity). However, in case of stationary-phase cells, activity of enzyme decreased to 45% ± 5.4% as compared to 100% activity on early phase cells (Fig S.4).
SDS-PAGE and Zymogram analysis of LysMR-5. a N-terminal His-6 tagged LysMR-5 purified by Ni-NTA chromatography was electrophoresed on 12% SDS gel and stained. Coomassie-blue stained SDS gel showing lane 1; marker, lane 2; supernatant of IPTG induced E. coli Bl21 cells containing pET28a-LysMR-5, lane 3; column flow through, lane 4 and 5; column washes, lane 6, 7, 8 and 9; purified recombinant LysMR-5 (approx. 58 kDa), lane 10; marker. b Zymogram analysis of purified LysMR-5. Zymogram containing S. aureus 43300 cells were run, renaturation of LysMR-5 in 50 mM Tris buffer pH 7.2 and 0.1% Triton X-100, incubated overnight and stained. Clear zone in lane 1 and 2 containing endolysin indicates active enzyme
Physicochemical properties of LysMR-5
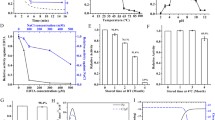
It is important to investigate the optimum conditions for maximum activity and stability of a lytic endolysin from the viewpoint of its clinical use. The effect of different temperatures on endolysin lytic activity was assessed to obtain optimum temperature for its activity (Fig. 4a). The effect on endolysin stability was determined using two different assays: the antibacterial assay to evaluate the effect of temperature on endolysin stability in terms of its lytic activity and the CD analysis at various temperatures to test the impact of temperature on the endolysin’s secondary structure, related to endolysin’s activity. LysMR-5 retained activity at temperatures from 25 °C to 45 °C (maintaining 93.5 ± 6% at 25 °C, 94 ± 5% at 30 °C, and 34 ± 5.9% at 45 °C). Highest activity was observed at 37 °C, making it optimum temperature for endolysin functioning. In stability assays, it was observed that LysMR-5 was highly sensitive towards high temperature, as activity was completely abolished at 50 °C. Endolysin was completely stable at 25 and 30 °C as no effect on the lytic activity was observed after pre-incubation for 30 min. After incubation at 45 °C for 30 min, 50 ± 5% and 87 ± 3.8% activity was lost as compared to the activity of native endolysin (not pre-incubated for 30 min) at 45 °C and activity of native endolysin at 37 °C, respectively. After pre-incubation of enzyme for ½, 1, 2, 3, 6, 24, and 48 h at 37 °C the residual activity observed was 76 ± 3.9%, 65 ± 2.4%, 53.88 ± 1.2%, 34.76 ± 5.1%, 23.58 ± 3.6%, 10.33 ± 0.83%, 4 ± 1.3%, respectively, indicating that enzyme became unstable on extended incubation at 37 °C. Hence, it should be stored at lower temperatures for long term use. Only 20% activity was lost after storage of LysMR-5 for 3 weeks at 4 °C, it was stable for 2 months at -20 °C. To explore the thermal stability and reason behind loss of endolysin activity with increasing temperature, CD spectra of LysMR-5 was measured at different temperatures (25, 40, 45, 50, 55, 60, and 70 °C). LysMR-5 dissipates its α-helix structure beyond 40 ºC and showed complete loss in overall secondary structure at 70 ºC (Fig. 4b). Loss in ellipticity was observed upon increase in temperature from 25 ºC to 70 ºC, showing two state thermal unfolding, with a midpoint of 47 ºC. Further LysMR-5 was unable to regain is structure upon cooling from 70 to 25 ºC, suggesting irreversible thermal unfolding (Fig. 4c). Similarly, pH dependence of lytic activity was checked and LysMR-5 was found to be active over a broad range of pH 5–9 (maintaining 52 ± 2.5% and 90.7 ± 6.2% of its activity at pH 5 and 9, respectively). However, optimum pH was 7.0, as endolysin was stable and showed the highest activity at this pH (Fig. 4d). After incubation of endolysin for 30 min at pH 8 and 9, 31.24 ± 6.4%, and 59.47 ± 3% a decrease in activity was observed, respectively. It was completely stable at pH 4, 5, 6, and 7 as no appreciable reduction in lytic activity was observed. Endolysin lytic activity was observed at various salt concentrations (Fig. 4e). Endolysin activity increased with an increase in NaCl concentrations, 44 ± 2.1% activity achieved at 50 mM NaCl which became 100% at 300 mM NaCl. The endolysin activity was even stable at higher NaCl concentrations (up to 800 mM). Like several other reports of staphylococcal endolysins, bioinformatic analysis of LysMR-5 sequence predicted association of Zn2+ and Ca2+ with amidase and CHAP domain, respectively [50,51,52]. In order to determine the role of divalent metal ions on LysMR-5 activity, various concentrations of metal ions (1 µM, 100 µm, 1 mM, and 5 mM of Ca, Zn, Mg, and Mn ions) were added to the 5 mM EDTA-treated endolysin (excess EDTA was removed by dialysis). The activity of the endolysin was not abolished by EDTA either 5 mM or 100 mM, as activity remained similar to untreated endolysin control. Addition of Ca, Mg, and Mn ions did not increase or decrease the endolysin activity at all the concentrations tested as compared to control, but addition of Zn2+ at 100 µm or more led to inhibition of activity (80% at 100 µm and 100% at higher concentrations) (Fig. S.5).
Optimization of activity conditions for LysMR-5 and thermal stability analysis of LysMR-5 by CD spectropolarimetry. The relative antibacterial activity was measured in terms of decrease in a number of viable S. aureus 43300 cells by cell counting. a Effect of temperature (25, 30, 37, 45, 50 °C) was measured on the activity of LysMR-5 by adding lysin to bacterial suspension and incubating for 30 min at various temperatures. The optimum temperature for endolysin activity was 37 °C, with good activity within a temperature range of 25 °C to 37 °C. b CD spectra of LysMR-5 at various temperatures. The loss of secondary structure (protein ellipticity) was observed with an increase in temperature. c Thermal denaturation curves. Blue shows loss in molar ellipticity of LysMR-5 monitored at 222 nm from 25 ºC to 70 ºC. Red depicts the renaturation profile upon decreasing temperature from 70 ºC to 25 ºC. d The influence of pH on bactericidal activity of LysMR-5 was assessed by adding lysin to bacterial suspensions prepared in buffers of different pH (4 to 9), incubated at 37 °C for 30 min. Endolysin showed good activity at pH 6–9, but optimum pH was 7. e Effect of NaCl concentrations was assessed and maximum activity was achieved at 300 mM NaCl, beyond which no further increase in lytic activity occurred
LysMR-5 antibacterial activity visualization and its concentration and time dependence
The effect of different endolysin concentrations on its lytic activity was analyzed by performing antibacterial assay. The antibacterial activity increased with increasing endolysin concentration till endolysin saturation was attained (Fig. 5a). Even 1 µg/ml of endolysin was able to show 52 ± 4.8% of lytic activity and 15 µg/ml of endolysin was enough to achieve maximum lytic activity (100%). The bacterial log units decreased from 8.54 ± 0.1 log units to 4.6 ± 0.08 log units in 30 min at optimum conditions. No further decrease in bacterial viability was seen by increasing the endolysin concentration (up to 50 µg/ml). Time-kill assay was performed to determine the maximum time required for the endolysin to exhibit maximum killing of bacterial cells (Fig. 5B). The activity observed after 30 min from sample incubation was taken as 100% activity, since no increase in bacterial lysis or antibacterial activity was observed upon further incubation of bacterial samples with endolysin for 45 or 60 min at 37 °C. In this relation 41 ± 4.4% activity was seen within 1 min of reaction initiation, after 5 min 86 ± 4% activity, after 15 min 95 ± 6.3% activity, and after 30 min 100% activity was executed. The killing effect of LysMR-5 (15 µg/ml) on S. aureus 43300 cells was visualized using FESEM (Fig. 6). Cells not treated with endolysin showed no change in cell surface morphology, no lysis, and no presence of extracellular cytoplasmic content (Fig. 6a, b, and c). On the contrary, cells exposed to LysMR-5 showed distorted surface morphology due to degradation of cell wall, cell debris (cell ghost), and cytoplasmic content present around the bacterial cells. The localized degradation of the cell wall led to extrusion, rupture of cell membrane, and release of cytoplasmic content (Fig. 6d, e, f, and g).
Enzyme concentration and time dependence of endolysin lytic activity. a Enzyme concentration dependence was observed by incubating bacterial suspension (108 CFU/ml) with different enzyme concentrations (0, 1, 5, 10, 15, 20, 30, 40, 50 µg/ml) for 30 min at 37 °C and decrease in viable bacterial count was assessed. An enzyme concentration of 15 µg/ml was sufficient for maximum bacterial killing. b For assessing effect of incubation time on lytic activity, decrease in bacterial viability after addition of equal amount of endolysin (15 µg/ml) for different time period was calculated. 30 min were required for endolysin to achieve maximum antibacterial activity at optimum activity conditions. Error bars represent means ± standard deviations of each experiment repeated in triplicate
Field emission scanning electron microscopy images of LysMR-5 treated and untreated S. aureus 43300 cells. S. aureus cells in the mid-exponential phase were harvested, resuspended in activity buffer and treated with either endolysin or activity buffer for 3 min at 37 °C. a, b, and c Untreated (only buffer) cells, bar 5 µm, 1 µm, and 500 nm, respectively. d, e, and f LysMR-5 treated cells, bar 5 µm, 500 nm, and 500 nm, respectively. g Lysed cells (cell ghost), bar 1 µm
LysMR-5 possesses active amidase and endopeptidase domains
Purified S. aureus peptidoglycan was digested with LysMR-5 for 4 h at 37 °C and supernatant subjected to different assays. High concentration (300 \(\upmu \mathrm{M}\)) of free amino groups in LysMR-5 treated sample in TNBS assay suggested presence of active endopeptidase domain. No free amino group was detected in undigested peptidoglycan since all the pre-existing free groups were removed by acetylation (Fig. 7a). In amidase assay, a significantly high amount of free muramic acid (600 µg) was detected as compared to other control groups (Fig. 7b). No reducing group was detected by Park-Johnson assay in the digested sample, suggesting that LysMR-5 does not exhibit glycosidase activity. To further confirm these results and determine the specific cleavage site of LysMR-5, ESI–MS of the digested S. aureus peptidoglycan was performed (Fig. 8). Analysis of the digest showed a major peak with m/z of 702.47 MS, which corresponded closely to the structure of peptidoglycan cleavage fragment NH2-l-Ala-d-Instability-l-Lys-(NH2-Gly5)-d-Ala-COOH (calculated MH+702.41). These results suggest that LysMR-5 acts as N-acetylmuramoyl-l-alanine amidase (hydrolyze amide bond b/w N-acetylmuramic acid and l-alanine) and endopeptidase (cleaves between stem peptide and pentaglycine cross-bridge), as these two enzymatic activities are needed for the generation of above mentioned fragment. These findings are consistent with conserved domains of LysMR-5 identified with conserved domain database server (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi), as well as observations made in amidase and endopeptidase assay.
Identification of endopeptidase and amidase activities of LysMR-5. Purified S. aureus peptidoglycan (500 µg/ml) were digested (except controls) with LysMR-5 (25 µg/ml) for 4 h at 37 °C. a To detect endopeptidase activity, release of free amino groups was measured calorimetrically at 345 nm. Pre-existing free amine groups were removed by acetylation to avoid false positive results. A high concentration of amino groups (300 µm) was detected in digested samples as compared to control groups. bN-Acetylmuramoyl-l-alanine amidase activity of LysMR-5 was detected (570 nm) by the presence of free muramic acid in digests by calorimetric assay. PGN; peptidoglycan only, LysMR-5; LysMR-5+ activity buffer, LysMR-5+ PGN; LysMR-5 digested peptidoglycan. Error bars represent the mean and standard deviation of triplicate assays
LysMR-5 peptidoglycan cleavage site determination using ESI–MS analysis. ESI–MS spectra (m/z range 400–800) of LysMR-5 digested purified S. aureus peptidoglycan. A major peak at m/z = 702.47 was observed, which is speculated to be the digestion product of functionally active amidase and d-Ala-Gly endopeptidase domains of endolysin
Quantification of endolysin activity using turbidity reduction assays
The lytic activity of LysMR-5 was determined using two different turbidity reduction assays. These methods are most commonly used to quantify endolysin lytic activity in terms of enzyme units. Various concentrations of endolysin were added to S. aureus 43300 live cells in a 96-well plate and change in turbidity was measured over a period of time as compared to control (no endolysin) (Fig. 9). Strong lytic activity was observed, as sharp drop in the OD600 was observed within 15 min after addition of endolysin at a concentration of 15, 10, and 5 µg per reaction. Even 0.3 ± 0.05 µg of enzyme was able to decrease the initial OD600 to its half in 15 min, which corresponds to 3333 ± 500 U/mg. Lytic activity of LysMR-5 against targeted bacteria (S. aureus 43300) was also quantitated by TRA previously described by Briers et al. [48]. Using this method, LysMR-5 was found to have an activity of 3800 units/mg with an R2 of 0.960.
Turbidity reduction assay. After addition of buffer (control) or different amount of endolysin to S. aureus cells, change in OD600 of bacterial suspension was measured. A small amount of endolysin (0.3 µg) was able to decrease bacterial OD600 to its half within 15 min. Increase in enzyme activity was proportional to the enzyme concentration until saturation was achieved (5 µg endolysin within 15 min of assay). Error bars represent means ± standard deviations of each experiment repeated in triplicate
Discussion
In the past two decades, phage endolysins have been used against multidrug-resistant bacteria because these are highly effective, specific and refractory to resistance generation, making them eminent antibacterial candidates [11, 17, 24, 25, 49, 53,54,55]. Only 25% of the phage population belongs to Herelleviridae family [56] and few endolysins from Herelleviridae phages like phi twort, LysK, and LysGH15 have been well characterized [19, 25, 27]. The present study demonstrates the in silico analysis, expression, purification, physicochemical, and antibacterial properties of novel endolysin (LysMR-5) from staphylococcal phage (MR-5) belonging to Herelleviridae family. In silico analysis of LysMR-5 showed that it has high amino acid sequence similarity (99%) with other reported staphylococcal endolysins, LysK, LysGH15, Lys-phiSA012, etc. [25, 27, 52]. It was predicted to have a modular architecture similar to previously described endolysins [19, 20, 25, 28, 57], which consisted of three domains N-terminal CHAP, amidase-2, and C-terminal SH3b_5 domain. This architecture is generally found in mycobacterial and staphylococcal endolysins but is more common in mycobacterial endolysins. Modular organization of endolysin has been proposed to be an evolutionary mechanism and is suggested to occur through domain swapping and their co-evolution with host autolysins [58]. In silico predictions of the secondary and tertiary structure of LysMR-5 showed that it consisted of α-helical and β-sheets. These observations were supported by the spectra collected through the CD spectropolarimetry of LysMR-5.
It is reported that CHAP domain can either have a peptidase or amidase activity. The CHAP domain of LysMR-5 had endopeptidase activity as it resulted in the release of high concentration of muramic acid in peptidoglycan digest in an endopeptidase assay. Presence of the active amidase domain in LysMR-5 was affirmed by amidase assay, in which high concentration of free alanine amino groups was detected as compared to control. The results are consistent with other dual lytic staphylococcal endolysins, endolysin of phage phi11 has d-alanyl-glycyl endopeptidase activity [28, 59] and Lys-phi K has active CHAP and amidase domains [47, 60]. Further, analysis of purified peptidoglycan digests by ESI–MS detected a major peak with m/z ratio of 702.47 which again corroborates the results of this study that LysMR-5 harbors both N-acetylmuramoyl-l-analine amidase and endopeptidase activity [61]. The results are concordant with the ESI–MS spectra given for endolysins that harbor similar catalytic domains, i.e., phi11, LysK, Twort, and LysH5 [47, 57, 62]. The presence of two putative antimicrobial activities in LysMR-5 (endopeptidase and amidase), might make it refractory to resistance generation as the bacteria would require two simultaneous compensatory mutations [15, 59]. In the era of drug resistance, this trait is very useful for an antimicrobial candidate. LysMR-5 demonstrated lytic activity against many diverse clinical isolates of S. aureus and S. epidermidis, which were collected from different patients, samples (blood, pus, soft tissue and wound swabs) and also had different antibiotic susceptibility and phage sensitivity patterns. These observations promote the use of LysMR-5 as a valuable antibacterial agent against staphylococcal infections. The results are consistent with previous findings, staphylococcal phage lysins like LysK, phi11 and phage-associated peptidoglycan hydrolase HydH5 also had broad lytic activity against various MRSA and VRSA CoNS strains [19, 28, 63]. This to some extent has been correlated with the amidase activity of these enzymes, as MurNAc-l-alanine linkages are ubiquitously present in peptidoglycan of bacterial cells. Due to this reason, lysins are able to act against a wide variety of bacterial species [57]. As observed in case of LysGH15, LysK, and SAL-1, no activity was observed against Gram-negative bacteria in this study as well [25, 64]. The genus specificity of most endolysins is an important advantage over classical broad range antibiotics, as it decreases the risk of resistance development. LysMR-5 was found to be highly active on actively dividing S. aureus cells in early, mid and late-exponential phase, no significant difference in enzyme activity (p > 0.05) was seen against cells in different exponential phase. However, a significant decrease (p < 0.05) in LysMR-5 activity against stationary-phase cells was observed. Similar results have been reported in previous studies on Streptococcus, Bacillus, and Staphylococcus aureus cells [42, 63, 65, 66]. These endolysins were more active against early and mid-exponential cells, but LysMR-5 was equally active against late-exponential cells also, which is an added advantage. Its inability to act against stationary bacteria might be due to increased cross-linking of peptidoglycan layer and other modifications in the cell wall during this phase [66].
The biochemical properties of endolysin LysMR-5 showed that it is active over a pH range of 5–9 (optimum 7) and a temperature range of 25 to 45 ºC (optimum 37 ºC). The optimum conditions for maximum enzyme activity are similar to physiological conditions, making it an attractive antimicrobial candidate. These results are consistent with previous reports, for LysK and LysGH15 [24, 67]. The isoelectric point of LysMR-5 was predicted to be 8.9, the enzyme was stable at pH 4–7 and instability of endolysin at pH 8 and 9 might be due to close proximity of these pH values to pI, which leads to inactivation of enzyme, most probably due to aggregation [68]. The biochemical characterization showed that the enzyme is highly sensitive towards higher temperature. This is corroborated on the basis of the observations made on stability and changes in the secondary structure analysis of LysMR-5 at different temperatures. CD spectrum of LysMR-5 showed loss in secondary structure at temperatures above 37 ºC with an apparent Tm of 47 ºC. At higher temperatures, complete loss of secondary structure could be seen. Inability of LysMR-5 to regain its secondary structure upon renaturation depicts its irreversible nature. These observations suggest that instability of LysMR-5 and loss of its enzyme activity could be attributed to the gradual loss of secondary structure. Only few similar studies have been conducted, and similar phenomenon has been reported. Loss of secondary structure at 40 ºC in case of staphylococcal endolysin LysK resulted in enzyme inactivation [68]. On the contrary a Salmonella phage endolysin Lys68 was thermostable and had stable CD spectra at temperatures up to 40 ºC (Tm 44 °C). Only small changes in ellipticity were observed up to 80 ºC [69]. Another Acinetobacter phage endolysin ABgp46 had an α-helical structure and a high melting temperature of 52 °C [70]. In this study highest lytic activity of LysMR-5 was observed in the presence of 300 mM of NaCl. Quite similar observations were made for two other staphylococcal endolysins LysGH15 and LysK, which were structurally also similar to LysMR-5. These endolysins showed maximum lytic activity at 400 mM NaCl [25, 70]. On the contrary, endolysins like HydH5 and LysB4 showed high lytic activities at lower NaCl concentrations (< 200 mM). The domain architecture of these endolysins was also different from LysMR-5, as HydH5consists of CHAP and LYZ2 domains and LysB4 has VanY and SH3_5 domain [8, 63].
The predicted structure model of CHAP domain had three α-helices packed against six β-sheets, this conformation is considered to be typical of NplC/P60 papain-related peptidases. The 3D structure of amidase_2 domain consisted predominantly of seven α-helices and four beta-sheets. This alpha/beta/alpha structure is a characteristic of amidase family [71]. Predicted 3D models of CHAP and Amidase_2 domain showed the presence of bound Ca2+ and Zn2+, so dependence of the lytic activity on these metal ions was analyzed. EDTA treatment did not abolish the endolysin activity even though 3 Zn2+-binding residues in the PGRP domain were predicted in LysMR-5. This could be due to the fact that either the metal ions are too tightly bound to endolysin and EDTA is unable to chelate them or that the endolysin does not require the metal ions for its lytic activity. The addition of Ca2+, Mg2+ or Mn2+ (1, 10, 100 µM, 1 and 5 mM) did not show any effect on LysMR-5 lytic activity, but addition of Zn2+ 100 µM onwards inhibited its lytic activity. LysBPS13 from Bacillus cereus phage and PGH from P. acidilactici ATCC 8042 have also been shown to be unaffected by EDTA treatment and metal ion addition [72, 73]. The inhibitory effect of Zn2+ on lytic activity has previously been reported for staphylococcal endolysin LysK, LysGH15, PlyTW, and HydH5 [51, 63, 68, 74]. Another Zn2+-independent amidase was reported from E. faecalis bacteriophage EF24C [75], and endolysin LysBPS13 [72]. On the contrary some of the reports have shown that Zn ion exerts a positive effect on enzyme’s lytic activity [8, 52, 76, 77]. Increase in lytic activity with addition of Ca2+ has been reported by several groups, but in case of LysK, similar to LysMR-5, no effect on lytic activity was observed. It is hypothesized that these ions might not be involved in catalytic reaction at the active site but may be of great importance in supporting an active conformation or stability of LysK.
It is known that endolysins, when tested in various antibacterial assays, give different results. This may be due to the differences in the sensitivity of every assay and its inherent biases. Hence, lytic activity of LysMR-5 against S. aureus cells was characterized with the help of different activity assays. An assay was performed to check the dependence of lytic activity on endolysin concentration. The results showed that LysMR-5 was highly antibacterial even at 15 µg/ml as it was able to kill 4 log units of bacteria within 30 min of incubation at 37 ºC. Similar results have been shown in case of P. aeruginosa phage endolysin LysPA26 [78], endolysin LysB4 [8], ClyS and HydH5 [29, 63]. In a time-kill assay, LysMR-5 showed rapid bacterial killing as within 1 min of endolysin incubation with S. aureus, 2 log unit reduction was seen, though 30 min of incubation resulted in 100% antibacterial activity (approx. 4 log units decrease). Exceptionally high enzyme activity was reported by Nelson and co-workers [49] for Streptococcal lysin (PlyC), as 100 units (1 ng enzyme) of this lysin was able to decrease 3 log units of streptococcus in 5 s.
Different laboratories have used different types of assays, conditions and unit definitions to quantitate endolysin activity [17]. In order to compare the activity of LysMR-5 with other reported endolysins, we determined the enzyme units by employing two different assays. The results obtained with both the methods showed compatibility. In a turbidity reduction assay, to achieve maximum activity (OD600 dropped to 0.06 from 1.1) only 5 µg of protein was required, a similar assay performed by Son et al. [8] also showed that maximum activity and enzyme saturation was achieved at 5 µg of protein. It was observed that minimum of 0.3 ± 0.05 µg enzyme was required to drop the OD600 to its half, which translated to 3333 Units/mg ± 500. In the case of purified LysGH15, comparatively higher concentration (approximately 0.8 μg) of enzyme was required to bring about similar decrease [25]. Equivalent amount of enzyme units were quantified for LysMR-5, i.e., 3800 U/mg in another TRA [48]. The antibacterial nature of LysMR-5 is further supported on the basis of results of FESEM analysis, as major alteration in the morphology of S. aureus cells was observed even after an exposure of 3 min. Treatment resulted in distorted cell shape, blebbing due to extrusion of cell contents from degraded cell wall of bacteria. Similar observations were made after addition of 250 μg of ClyS for 1 to 3 min to S. aureus 8325-4 cells and 500 µg of LysAB2 for 60 min to Acinetobacter baumannii cells by SEM and TEM analysis, respectively [29, 79]. Most of the structural and biochemical properties of the LysMR-5 are similar to the endolysins that share high sequence identity with it. Few of the observed variations in the properties of LysMR-5, like effect of Ca2+ ions and difference in the endolysin concentration required for achieving similar bactericidal effect, might be due to the minor difference (4 amino acids) observed between the sequences of these endolysins. Previously it has been reported that minor changes in the endolysin sequence can affect its antibacterial activity [64, 80].
In conclusion, LysMR-5 exhibits highest lytic activity at conditions similar to the physiological environment, shows rapid bactericidal kinetics and exhibits a strong bacteriolytic effect against staphylococcal strains. Taken together these findings suggest that, LysMR-5 has features conducive for a promising antimicrobial agent, making it an attractive addition to the alternative drug arsenal and potent enzybiotic against MRSA infections. As the in vitro conditions that are used to evaluate antibacterial efficacy of enzymes vary greatly from the in vivo scenario, work on animals will further validate the endolysin potential as a therapeutic agent to treat human infections. To implement LysMR-5 as a clinically viable treatment option, currently we are working to improve the delivery of LysMR-5 which could further enhance its therapeutic efficacy by various means and on the in vivo evaluation of its antibacterial efficacy in treating S. aureus mediated infection.
References
Lowy FD (1998) Staphylococcus aureus infections. N Engl J Med 339:520–532
Ferry T, Perpoint T, Vandenesch F, Etienne J (2005) Virulence determinants in Staphylococcus aureus and their involvement in clinical syndromes. Curr Infect Dis Rep 7:420–428
Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, Harbarth S, Hindler JF, Kahlmeter G, Olsson-Liljequist B, Paterson DL, Rice LB, Stelling J, Struelens MJ, Vatopoulos A, Weber JT, Monnet DL (2012) Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect 18:268–281
Joshi S, Ray P, Manchanda V, Bajaj J, Chitnis DS, Gautam V, Goswami P, Gupta V, Harish BN, Kagal A, Kapil A, Rao R, Rodrigues C, Sardana R, Devi S, Sharma A (2013) Veeragaghavan Balaji Indian Network for Surveillance of Antimicrobial Resistance (INSAR) group, India. Methicillin resistant Staphylococcus aureus (MRSA) in India: prevalence & susceptibility pattern. Indian J Med Res 137:363–369
Boswihi SS, Udo EE (2018) Methicillin-resistant Staphylococcus aureus: An update on the epidemiology, treatment options and infection control. Curr Med Res Pract 8:18–24
Wu M, Hu K, Xie Y, Liu Y, Mu D, Guo H, Zhang Z, Zhang Y, Chang D, Shi Y (2019) A Novel phage PD-6A3, and its endolysin Ply6A3, with extended lytic activity against Acinetobacter baumannii. Front Microbiol. https://doi.org/10.3389/fmicb.2018.03302
Donovan DM, Becker SC, Dong S, Baker JR, Foster-Frey J, Pritchard DG (2009) Peptidoglycan hydrolase enzyme fusions for treating multi-drug resistant pathogens. Biotech Int 21:6–10
Son B, Yun J, Lim JA, Shin H, Heu S, Ryu S (2012) Characterization of LysB4, an endolysin from the Bacillus cereus-infecting bacteriophage B4. BMC Microbiol 12:33
Haddad KH, Schmelcher M, Sabzalipoor H, Seyed HE, Moniri R (2018) Recombinant endolysins as potential therapeutics against antibiotic-resistant Staphylococcus aureus: current status of research and novel delivery strategies. Clin Microbiol Rev 31:e00071–e00117
Melo LDR, Brandao A, Akturk E, Santos SB, Azeredo J (2018) Characterization of a new Staphylococcus aureus kayvirus harboring a lysin active against biofilms. Viruses 10:182. https://doi.org/10.3390/v10040182
Loessner MJ (2005) Bacteriophage endolysins: current state of research and applications. Curr Opin Microbiol 8:480–487
Fischetti VA (2010) Bacteriophage endolysins: a novel anti-infective to control Gram-positive pathogens. Int J Med Microbiol 300:357–362
Oliveira H, São-José C, Azeredo J (2018) Phage-derived peptidoglycan degrading enzymes: challenges and future prospects for in vivo therapy. Viruses 10:E292. https://doi.org/10.3390/v10060292
Loeffler JM, Nelson D, Fischetti VA (2001) Rapid killing of Streptococcus pneumoniae with a bacteriophage cell wall hydrolase. Science 294:2170–2172
Fischetti VA (2005) Bacteriophage lytic enzymes: novel anti-infectives. Trends Microbiol 13:491–496
Gutiérrez DFL, Rodríguez A, García P (2018) Are phage lytic proteins the secret weapon to kill Staphylococcus aureus? mBio 9:e01923–e02017
Schmelcher M, Donovan DM, Loessner MJ (2012) Bacteriophage endolysins as novel antimicrobials. Future Microbiol 7:1147–1171
Jun SY, Jung GM, Yoon SJ, Oh MD, Choi YJ, Lee WJ, Kong JC, Seol JG, Kang SH (2013) Antibacterial properties of a pre-formulated recombinant phage endolysin, SAL-1. Int J Antimicrob Agents 41:156–161
O'Flaherty S, Coffey A, Meaney W, Fitzgerald GF, Ross RP (2005) The recombinant phage lysin LysK has a broad spectrum of lytic activity against clinically relevant staphylococci, including methicillin-resistant Staphylococcus aureus. J Bacteriol 187:7161–7164
Rashel M, Uchiyama J, Ujihara T, Uehara Y, Kuramoto S, Sugihara S, Yagyu K, Muraoka A, Sugai M, Hiramatsu K, Honke K, Matsuzaki S (2007) Efficient elimination of multidrug-resistant Staphylococcus aureus by cloned lysin derived from bacteriophage phi MR11. J Infect Dis 196:1237–1247
Drulis-Kawa Z, Majkowska-Skrobek G, Maciejewska B (2015) Bacteriophages and phage-derived proteins–application approaches. Curr Med Chem 22:1757–1773
Heselpoth RD, Swift SM, Linden SB, Mitchell MS, Nelson DC (2018) Enzybiotics: endolysins and bacteriocins. In: Harper D, Abedon ST, Burrowes B, McConvill M (eds) Bacteriophages. Springer, Cham, pp 1–42
Fernández-Ruiz I, Coutinho FH, Rodriguez-Valera F (2018) Thousands of novel endolysins discovered in uncultured phage genomes. Front Microbiol 9:1033. https://doi.org/10.3389/fmicb.2018.01033
Loeffler JM, Fischetti VA (2003) Synergistic lethal effect of a combination of phage lytic enzymes with different activities on penicillin-sensitive and -resistant Streptococcus pneumoniae strains. Antimicrob Agents Chemother 47:375–377
Gu J, Xu W, Lei L, Huang J, Feng X, Sun C, Du C, Zuo J, Li Y, Du T, Li L, Han W (2010) Lysgh15, a novel bacteriophage lysin, protects a murine bacteremia model efficiently against lethal methicillin-resistant Staphylococcus aureus infection. J Clin Microbiol 49:111–117
Jun SY, Jung GM, Yoon SJ, Choi YJ, Koh WS, Moon KS, Kang SH (2014) Preclinical safety evaluation of intravenously administered SAL200 containing the recombinant phage endolysin SAL-1 as a pharmaceutical ingredient. Antimicrob Agents Chemother 58:2084–2088. https://doi.org/10.1128/AAC.02232-13
Loessner MJ, Gaeng S, Wendlinger G, Maier SK, Scherer S (1998) The two-component lysis system of Staphylococcus aureus bacteriophage Twort: a large TTG-start holin and an associated amidase endolysin. FEMS Microbiol Lett 162:265–274
Sass P, Bierbaum G (2007) Lytic activity of recombinant bacteriophage phi11 and phi12 endolysins on whole cells and biofilms of Staphylococcus aureus. Appl Environ Microbiol 73:347–352
Daniel A, Euler C, Collin M, Chahales P, Gorelick KJ, Fischetti VA (2010) Synergism between a novel chimeric lysin and oxacillin protects against infection by methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother 54:1603–1612
Kaur S, Harjai K, Chhibber S (2012) Methicillin-resistant Staphylococcus aureus phage plaque size enhancement using sublethal concentrations of antibiotics. Appl Environ Microbiol 78:8227–8233
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual. Cold Spring Harbor, New York
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor, New York
He F (1997) Laemmli-SDSPAGE Bio-protocol. Bio101. https://doi.org/10.21769/BioProtoc.80
Corpet F (1988) Multiple sequence alignment with hierarchical clustering. Nucl Acids Res 16:10881–10890
Geourjon C, Deleage G (1995) SOPMA: significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments. Comput Appl Biosci 11:681–684. https://doi.org/10.1093/bioinformatics/11.6.681
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723. https://doi.org/10.1002/elps.1150181505
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Binkowski TA, Naghibzadeh S, Liang J (2003) CASTp: Computed Atlas of Surface Topography of proteins. Nucleic Acids Res 31:3352–3355. https://doi.org/10.1093/nar/gkg512
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291. https://doi.org/10.1107/S0021889892009944
Leive L (1968) Studies on the permeability change produced in coliform bacteria by ethylenediaminetetraacetate. J Biol Chem 243:2373–2380
Lepeuple AS, Gemert EV, Chapot-Chartier MP (1998) Analysis of the bacteriolytic enzymes of the autolytic Lactococcus lactis subsp. cremoris strain AM2 by renaturing polyacrylamide gel electrophoresis: identification of a prophage-encoded enzyme. Appl Environ Microbiol 64:4142–4148
Lavigne R, Briers Y, Hertveldt K, Robben J, Volckaert G (2004) Identification and characterization of a highly thermostable bacteriophage lysozyme. Cell Mol Life Sci 61:2753–2759
Pritchard DG, Dong S, Kirk MC, Cartee RT, Baker JR (2007) LambdaSa1 and LambdaSa2 prophage lysins of Streptococcus agalactiae. Appl Environ Microbiol 73:7150–7154
Mokrasch LC (1967) Use of 2,4,6-trinitrobenzenesulfonic acid for the coestimation of amines, amino acids, and proteins in mixtures. Anal Biochem 18:64–71
Hazenberg MP, de Visser H (1992) Assay for N-acetylmuramyl-l-alanine amidase in serum by determination of muramic acid released from the peptidoglycan of Brevibacterium divaricatum. Eur J Clin Chem Clin Biochem 30:141–144
Park JT, Johnson MJ (1949) A submicrodetermination of glucose. J Biol Chem 181:149–151
Becker SC, Dong S, Baker JR, Foster-Frey J, Pritchard DG, Donovan DM (2009) LysK CHAP endopeptidase domain is required for lysis of live staphylococcal cells. FEMS Microbiol Lett 294:52–60
Briers Y, Lavigne R, Volckaert G, Hertveldt K (2007) A standardized approach for accurate quantification of murein hydrolase activity in high-throughput assays. J Biochem Biophys Methods 70:531–533
Nelson D, Loomis L, Fischetti VA (2001) Prevention and elimination of upper respiratory colonization of mice by group A streptococci by using a bacteriophage lytic enzyme. Proc Natl Acad Sci USA 98:4107–4112
Fenton M, Cooney JC, Ross RP, Sleator RD, McAuliffe O, O’Mahony J, Coffey A (2011) In silico modeling of the staphylococcal bacteriophage-derived peptidase CHAPK. Bacteriophage 1:198–206
Gu J, Feng Y, Feng X, Sun C, Lei L, Ding W, Niu F, Jiao L, Yang M, Li Y, Liu X, Song J, Cui Z, Han D, Du C, Yang Y, Ouyang S, Liu ZJ, Han W (2014) Structural and biochemical characterization reveals LysGH15 as an unprecedented “EF-Hand-Like” calcium-binding phage lysin. PLOS Pathog 10:e1004109
Fujiki J, Nakamura T, Furusawa T, Ohno H, Takahashi H, Kitana J, Usui M, Higuchi H, Tanji Y, Tamura Y, Iwano H (2018) Characterization of the lytic capability of a LysK-like endolysin, Lys-phiSA012, derived from a polyvalent Staphylococcus aureus bacteriophage. Pharmaceuticals (Basel) 11:E25. https://doi.org/10.3390/ph11010025
Fischetti VA (2008) Bacteriophage lysins as effective antibacterials. Curr Opin Microbiol 11:393–400
Fenton M, Casey PG, Hill C, Gahan CG, Ross RP, McAuliffe O, O'Mahony J, Maher F, Coffey A (2010) The truncated phage lysin CHAP(k) eliminates Staphylococcus aureus in the nares of mice. Bioeng Bugs 1:404–407
Singh PK, Donovan DM, Kumar A (2014) Intravitreal injection of the chimeric phage endolysin Ply187 protects mice from Staphylococcus aureus endophthalmitis. Antimicrob Agents Chemother 58:4621–4629
Oliveira H, Melo LD, Santos SB, Nóbrega FL, Ferreira EC, Cerca N, Azeredo J, Kluskens LD (2013) Molecular aspects and comparative genomics of bacteriophage endolysins. J Virol 87:4558–4570
Navarre WW, Schneewind O (1999) Surface proteins of Gram-positive bacteria and mechanisms of their targeting to the cell wall envelope. Microbiol Mol Biol Rev 63:174–229
Díaz E, López R, García JL (1990) Chimeric phage-bacterial enzymes: a clue to the modular evolution of genes. Proc Natl Acad Sci USA 87:8125–8129
Donovan DM, Lardeo M, Foster-Frey J (2006) Lysis of staphylococcal mastitis pathogens by bacteriophage phi11 endolysin. FEMS Microbiol Lett 265:133–139
Horgan M, O’Flynn G, Garry J, Cooney J, Coffey A, Fitzgerald GF, Ross RP, McAuliffe O (2009) Phage lysin LysK can be truncated to its CHAP domain and retain lytic activity against live antibiotic-resistant staphylococci. Appl Environ Microbiol 75:872–874
Rodríguez-Rubio L, Martínez B, Rodríguez A, Donovan DM, Götz F, García P (2013) The phage lytic proteins from the Staphylococcus aureus bacteriophage vB_SauS-phiIPLA88 display multiple active catalytic domains and do not trigger staphylococcal resistance. PLoS ONE 8:e64671. https://doi.org/10.1371/journal.pone.0064671
Schmelcher M, Shen Y, Nelson DC, Eugster MR, Eichenseher F, Hanke DC, Loessner MJ, Dong S, Pritchard DG, Lee JC, Becker SC, Foster-Frey J, Donovan DM (2015) Evolutionarily distinct bacteriophage endolysins featuring conserved peptidoglycan cleavage sites protect mice from MRSA infection. J Antimicrob Chemother 70:1453–1465
Rodríguez-Rubio L, Martínez B, Zhou Y, Rodríguez A, Donovan DM, García P (2011) Lytic activity of the virion-associated peptidoglycan hydrolase HydH5 of Staphylococcus aureus bacteriophage vB_SauS-phiIPLA88. BMC Microbiol 11:138
Jun SY, Jung GM, Son JS, Yoon SJ, Choi YJ, Kang SH (2011) Comparison of the antibacterial properties of phage endolysins SAL-1 and LysK. Antimicrob Agents Chemother 55:1764–1767
Celia LK, Nelson D, Kerr DE (2008) Characterization of a bacteriophage lysin (Ply700) from Streptococcus uberis. Vet Microbiol 130:107–117
Pritchard DG, Dong S, Baker JR, Engler JA (2004) The bifunctional peptidoglycan lysin of Streptococcus agalactiae bacteriophage B30. Microbiology 150:2079–2087
Becker SC, Foster-Frey J, Donovan DM (2008) The phage K lytic enzyme LysK and lysostaphin act synergistically to kill MRSA. FEMS Microbiol Lett 287:185–191
Filatov LY, Becker SC, Donovan DM, Gladilin AK, Klyachko NL (2010) Lysk, the enzyme lysing Staphylococcus aureus cells: specific kinetic features and approaches towards stabilization. Biochimie 92:507–513
Oliveira H, Thiagarajan V, Walmagh M, Sillankorva S, Lavigne R, Neves-Petersen MT, Kluskens LD, Azeredo J (2014) A thermostable Salmonella phage endolysin, Lys68, with broad bactericidal properties against gram-negative pathogens in presence of weak acids. PLoS ONE 9:e108376. https://doi.org/10.1371/journal.pone.0108376
Oliveira H, Vilas BD, Mesnage S, Kluskens LD, Lavigne R, Sillankorva S, Secundo F, Azeredo J (2016) Structural and enzymatic characterization of ABgp46, a novel phage endolysin with broad anti-gram-negative bacterial activity. Front Microbiol 7:208. https://doi.org/10.3389/fmicb.2016.00208
Cheng X, Zhang X, Pflugrath JW, Studier FW (1994) The structure of bacteriophage T7 lysozyme, a zinc amidase and an inhibitor of T7 RNA polymerase. Proc Natl Acad Sci USA 9:4034–4038
Park J, Yun J, Lim JA, Kang DH, Ryu S (2012) Characterization of an endolysin, LysBPS13, from a Bacillus cereus bacteriophage. FEMS Microbiol Lett 33:76–83
Cano IG, Campos-Gómez M, Contreras-Cruz M et al (2015) Expression, purification, and characterization of a bifunctional 99-kDa peptidoglycan hydrolase from Pediococcus acidilactici ATCC 8042. Appl Microbiol Biotechnol 99:8563–8573
Becker SC, Swift S, Korobova O, Schischkova N, Kopylov P, Donovan DM, Abaev I (2015) Lytic activity of the staphylolytic Twort phage endolysin CHAP domain is enhanced by the SH3b cell wall binding domain. FEMS Microbiol Lett 362:1–8
Uchiyama J, Takemura I, Hayashi I, Matsuzaki S, Satoh M, Ujihara T, Murakami M, Imajoh M, Sugai M, Daibata M (2011) Characterization of lytic enzyme open reading frame 9 (ORF9) derived from Enterococcus faecalis bacteriophage phiEF24C. Appl Environ Microbiol 77:580–585
Plotka M, Kaczorowska AK, Morzywolek A, Makowska J, Kozlowski LP, Thorisdottir A, Skírnisdottir S, Hjörleifsdottir S, Fridjonsson OH, Hreggvidsson GO, Kristjansson JK, Dabrowski S, Bujnicki JM, Kaczorowski T (2015) Biochemical characterization and validation of a catalytic site of a highly thermostable Ts2631 endolysin from the Thermus scotoductus Phage vB_Tsc2631. PLoS ONE 10:e0137374
Plotka M, Sancho-Vaello E, Dorawa S, Kaczorowska AK, Kozlowski LP, KaczorowskI T, Zeth K (2019) Structure and function of the Ts2631 endolysin of Thermus scotoductus phage vB_Tsc2631 with unique N-terminal extension used for peptidoglycan binding. Sci Rep. https://doi.org/10.1038/s41598-018-37417-6
Guo M, Feng C, Ren J, Zhuang X, Zhang Y, Zhu Y, Dong K, He P, Guo X, Qin J (2017) A novel antimicrobial endolysin, LysPA26, against Pseudomonas aeruginosa. Front Microbiol 8:293
Lai MJ, Lin NT, Hu A, Soo PC, Chen LK, Chen LH, Chang KC (2011) Antibacterial activity of Acinetobacter baumannii phage ϕAB2 endolysin (LysAB2) against both gram-positive and gram-negative bacteria. Appl Microbiol Biotechnol 90:529–539
Walmagh M, Boczkowska B, Grymonprez B, Briers Y, Drulis-Kawa Z, Lavigne R (2013) Characterization of five novel endolysins from Gram-negative infecting bacteriophages. Appl Microbiol Biotechnol 97:4369–4375
Acknowledgments
We gratefully acknowledge the University Grant Commission, New Delhi, India, and DST-PURSE grant, New Delhi for the financial support for the purchase of consumables and Ashutosh Kumar for assistance with the cloning of the endolysin gene in this project.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: S.C., J.K. Performed the experiments: J.K., P.S. Analyzed the data: S.C., J.K., P.S., D.S., K.H. Contributed reagents/materials/analysis tools: S.C., D.S., K.H. Wrote the paper: S.C., J.K., P.S., D.S., KH.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual authors included in this article.
Additional information
Edited by Andrew Millard.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kaur, J., Singh, P., Sharma, D. et al. A potent enzybiotic against methicillin-resistant Staphylococcus aureus. Virus Genes 56, 480–497 (2020). https://doi.org/10.1007/s11262-020-01762-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-020-01762-4