Abstract
Bovine leukemia virus (BLV) is the etiological agent of enzootic bovine leucosis (EBL), which affects cattle globally. In Egypt, BLV control strategies have been ignored because of the shortage of BLV research studies and the silent infection in most animals. This study aimed to identify the risk factors associated with the prevalence of BLV among dairy and beef cattle from six different geographic and climatic provinces in Egypt. Additionally, risk factors affecting the BLV proviral load (PVL) among the positive cattle were targeted. The total BLV prevalence in cattle from six investigated Egyptian provinces was 24.2% (105/433), while the mean PVL (8651.6 copies /105 white blood cells) was absolutely high as estimated by the BLV-CoCoMo-quantitative polymerase chain reaction (qPCR)-2 assay. Analysis of the influence of risk factors (age, sex, breed, production type, farm size, and location) on BLV prevalence indicated that the Holstein breed (OR = 1.582, p = 0.007), beef cattle (OR = 1.088, p = 0.0001), large-size farms (OR = 1.26, p = 0.0001), and cattle from Damietta (OR = 1.43, p = 0.0001) and Cairo (OR = 1.16, p = 0.0001) were ultimately proven the most important risks for BLV infection. The risk factors were analyzed considering the BLV PVL levels in the BLV-positive cases. Significantly high PVL (HPVL) levels were observed in cattle > 5 years old (p < 0.0001), females (p = 0.0008), Holstein (p < 0.0001), dairy cows (p = 0.0053), large-size farms (p < 0.0001), and cattle from Damietta (p < 0.0001) compared to other categories. Contrary, no significant differences in PVL levels were reported between the Native and Mixed cattle breeds (p = 0.13). Ultimately, the logistic regression model indicated that the probability of carrying HPVL in cattle > 5 years is 1.27 (95% CI: 1.03–2.09, p < 0.001) times more likely compared to cattle < 2 years old. In conclusion, the findings were valuably correlating the BLV prevalence with PVL as an indicator of the risk of BLV infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bovine leukemia virus (BLV), a virus of the Retroviridae family, Deltaretrovirus genus, is closely related to human T-lymphocyte leukemia virus 1 (HTLV-1) (Schwartz et al. 1994; Aida et al. 2013). BLV has a widespread prevalence and severe economic losses in the cattle industry worldwide (Bartlett et al. 2020). It is the etiological agent of Enzootic Bovine Leucosis (EBL), a disease that causes neoplastic lesions in cattle (Gillet et al. 2007; Aida et al. 2013). EBL has been classified by the World Organization for Animal Health (WOAH) as one of the diseases that could significantly affect global trade (Polat et al. 2017). As soon as BLV infects a host, it integrates a DNA intermediate as a provirus, both randomly and permanently into the host cell’s genome (Schwartz et al. 1994; Saito et al. 2022). BLV infection is characterized by a long incubation period and a variety of clinical outcomes. The majority of BLV-infected cattle are subclinical in the aleukemic stage, although, digestive disturbance, weight loss, weakness, reduced meat, and milk production, shortened longevity, and immune suppression may occur (Aida et al. 2013; Bartlett et al. 2013; Frie and Coussens 2015; Norby et al. 2016; Polat et al. 2017). About one-third of infected cattle can induce the persistent lymphocytosis stage, which is characterized by non-malignant polyclonal expansion of B-lymphocytes. However, only 1–5% develop B-cell lymphoma within different organs such as the spleen, liver, heart, abomasum, uterus, lymph nodes, and spinal cord resulting in wide varieties of clinical signs after a long latency (Burny et al. 1988; Gillet et al. 2007; Aida et al. 2013). Carcass condemnation at slaughter commonly occurs in cattle with lymphosarcoma in some countries such as Japan as it is judged unfit for human consumption (White and Moore 2009; Kobayashi et al. 2014).
BLV proviral load (PVL) is represented by the number of integrated copies of the retroviral genome into the host genome (Kettmann et al. 1979; Tajima et al. 1998). The level of BLV PVL is strongly correlated with BLV infection capacity (Jimba et al. 2010; Sato et al. 2018) and EBL progression (Somura et al. 2014; Ohno et al. 2015; Lo et al. 2020). BLV-infected cattle with high PVL (HPVL) are suggested to increase the risk of BLV transmission via direct contact with BLV-free animals. However, cows with low PVL (LPVL) are at low risk of BLV infection for BLV-free animals (Juliarena et al. 2016). There is evidence for the presence of BLV provirus in the milk, nasal mucus, and saliva of dairy cows with PVL > 10,000, 14,000, and 18,000 copies/105 blood cells, respectively (Yuan et al. 2015; Watanuki et al. 2019). These cattle could spread BLV through milk, nasal discharge, and saliva (Hopkins and DiGiacomo 1997; Yuan et al. 2015; Watanuki et al. 2019; Barzegar et al. 2021; Nakatsuchi et al. 2022). BLV transmission varies greatly by geography and agricultural system (Bartlett et al. 2020). It can be transmitted by blood-sucking insects in addition to the other iatrogenic procedures involving the transfer of infected blood between animals (i.e., dehorning, ear tattooing, rectal palpation, and needle reuse), and is responsible for disease propagation in a herd (Hopkins and DiGiacomo 1997). Transmission may occur transplacentally from an infected dam to the fetus, intrapartum by contact with infected blood, or postpartum from the dam to the calf through ingestion of infected milk (Hopkins and DiGiacomo 1997).
BLV genome consists of four structural and enzymatic genes namely; gag, pro, pol, and env, regulatory genes tax and rex, and accessory genes R3 and G4, and is flanked by two identical long terminal repeats (LTRs). BLV env gene is transcribed as an envelope glycoprotein (Env) complex that comprises, gp30 transmembrane (TM) protein and gp51 surface glycoprotein (SU), which is essential in the virus entry into the host cell (Gillet et al. 2007; Matsuura et al. 2019). Up to date, 12 BLV genotypes have been identified globally based on the phylogenetic analysis of env-gp51 sequences after a recent detection of BLV genotype-12 in Kazakhstan (Sultanov et al. 2022). BLV has invaded all continents and is currently distributed everywhere (Polat et al. 2017). Recent investigations in most nations outside of Europe have reported continuous increases in BLV prevalence in both beef and dairy cattle (Bartlett et al. 2020), although, about 20 European countries have successfully eradicated the BLV and achieved the BLV-free status (More et al. 2017).
In Egypt, few studies identifying BLV provirus in cattle were conducted. Our group has characterized the BLV genotypes 1 and 4 circulating in Egyptian dairy (Hamada et al. 2020) and beef cattle (Metwally et al. 2020) for the first time. Another study detected BLV genotype-1 in one sample from cattle blood (Selim et al. 2021). Furthermore, two serological surveys of anti-BLV antibodies were performed in cattle samples from a limited geographical region of Egypt (Zaher and Ahmed 2014; Selim et al. 2021). BLV national control strategies have been neglected in Egypt because the disease is present in an asymptomatic form in most animals (Selim et al. 2021). The BLV distribution throughout a wide area of Egypt is poorly investigated and the level of BLV PVL as a potential risk factor among Egyptian cattle has not been previously analyzed. Therefore, this study aimed to detect the BLV PVL among dairy and beef cattle from Southern, Middle, and Northern areas in Egypt using the BLV-CoCoMo-quantitative polymerase chain reaction (qPCR)-2 method. We also aimed to analyze the influence of some risk factors on BLV prevalence among Egyptian cattle. Moreover, the correlations between the BLV PVL levels in the infected animal’s blood and the investigated risk factors were targeted.
Materials and methods
Ethical approval
All animals were handled by the regulation of the Animal Ethics Committee at the Faculty of Veterinary Medicine, Damanhour University, Damanhour, Egypt, and by the regulation of RIKEN, Japan in strict accordance with good animal practice following the guidelines of RIKEN. Permission of the farm owners was obtained for sample collection. The study was reviewed and approved by the Research Code of Ethics (RCOE-DMU) at the Damanhour University (DMU/VetINF-2022-/0151), and by the RIKEN Animal Experiments Committee (approval number H29-2-104).
Cattle population
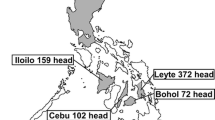
A total number of 433 (266 dairies and 167 beef) apparent healthy cattle from 24 cattle farms and four abattoirs located in six Egyptian provinces, distributed in Southern (Qena and Luxor), Middle (Cairo and Fayoum), and Northern (Beheira, and Damietta) areas of Egypt were randomly selected (Fig. 1). The sample size from each farm and/or abattoir was detected by Thrusfield’s formula (Thrusfield 2018) considering the expected BLV prevalence of 15.8% as reported by Zaher and Ahmed (Zaher and Ahmed. 2014). The blood samples from cattle in each farm and/or abattoir (about 10%) were randomly collected for investigation, while the total samples’ number from each province (Table 1) was based on the owner’s permission. These animals included various cattle breeds such as the Egyptian Native breed (n = 28), Holstein breed (n = 125), and the Mixed breed (n = 280), the crossbreed between foreign breeds and Native cattle, and included cattle of both sexes (188 males and 245 females) of different ages, < 2 years (n = 48), 2–5 years (n = 210), and > 5 years (n = 175). Regarding farm size, a number of 141 blood samples were collected from cattle in 21 small-size farms (< 200 heads per farm), and 125 blood samples were collected from cattle in three large-size farms (> 200 heads per farm), while 167 bulls were samples from abattoirs of unknown farm size. The investigated cattle appeared in good health, and none of them had any obvious signs of lymphadenitis or tumors as examined by the veterinarians in different farms and abattoirs.
The geographical map of Egypt shows the location of the six provinces, where the samples were collected in this study. In the North, Damietta is indicated by purple color and Beheira by blue. In the Middle, Cairo is indicated by green color and Fayom by brown. In the South, Qena is indicated by yellow color and Luxor by red. Numbers on the left side indicate the climatic conditions [Average annual temperature °C (T) and humidity % (H)] in the North (Blue), Middle (Yellow), and South (Red), which obtained from https://jssa.journals.ekb.eg/ (Accessed, 6 June 2023)
Blood sampling and DNA extraction
Blood samples were collected into a glass tube containing K2 EDTA anticoagulant using the coccygeal vein puncture procedure and then transported immediately into the laboratory. Genomic DNA was extracted from 300 µl whole blood using the Wizard Genomic DNA Purification Kit (Promega; Madison, WI, USA), according to the manufacturer’s instructions. The concentration, quality (A260/280), and purity (A260/230) of extracted DNA samples were measured using NanoDrop One Spectrophotometer (Thermo Fisher Scientific; Waltham, MA, USA). DNA samples of concentration > 200 ng/µl, quality > 1.8, and purity > 2 were subjected to investigation, while samples of low quality were re-extracted. DNA samples were diluted in nuclease-free water to a final concentration of 30 ng/µl for PCR experiments.
Evaluation of BLV PVL using CoCoMo-qPCR-2
BLV PVL was estimated using the CoCoMo-qPCR-2 method (RIKEN Genesis; Kanagawa, Japan), as described previously (Jimba et al. 2010; Takeshima et al. 2015). This study used a commercial CoCoMo-qPCR-2 kit (RIKEN Genesis; Kanagawa, Japan) to detect the PVL according to the manufacturer’s instructions. Briefly, a 183-bp sequence of the BLV LTR regions was amplified in a reaction mixture containing THUNDERBIRD Probe qPCR Mix (Toyobo; Tokyo, Japan), using the degenerate primer pairs, CoCoMo FRW and CoCoMo REV, and a 15 bp 6-carboxyfluorescein (FAM)-labeled LTR probe. To normalize the viral genomic DNA level within the host cellular genome, a 151-bp sequence of BoLA-DRA was amplified using the primer pairs, DRA-FW and DRA-RW, and a FAM-labeled DRA probe, as previously described (Takeshima et al. 2015). The reaction was carried out in 96-well plates according to the manufacturer’s guidelines on a LightCycler® 480 Instrument II (Roche Diagnostics; Basel, Switzerland). The PVL was calculated using the equation (number of BLV-LTR copies/number of BoLA-DRA copies) × 105 cells.
Statistical analysis
BLV prevalence among the tested cattle was determined by counting. The statistical analysis was performed using SPSS version 25 (IBM, New York, USA). The significanceof the investigated risk factors (age, sex, breed, production type, farm size, and location) and the differences in BLV PVL between the cattle groups were analyzed, and the Odds ratios (ORs) and 95% confidence intervals (CI) were calculated. BLV PVL was analyzed by the Kruskal-Wallis test for age, breed, farm size, and location, while the Mann-Whitney U test was used for sex and production type. Additionally, a logistic regression (LR) model analysis was performed to identify the most important risk factor (s) related to BLV infection. The investigated risk factor’s P-values for entry into or removal from the LR model were < 0.20. The LR model fitted with BLV infection as the outcome variable (negative: 0, positive: 1), and included effects of four variables; breed (3 categories: Native, Mixed, and Holstein), production type (2 categories: beef and dairy), farm size (3 categories: small, large, and abattoir), and location (6 categories: Qena, Damietta, Beheira, Fayoum, Luxor, and Cairo). The fit of the LR model was estimated using the Hosmer–Lemeshow goodness-of-fit test. Moreover, the risk influence of age (3 categories: > 2, 2–5, and < 5 years) on the occurrence of HPVL (2 categories: LPVL < 1000: 0, and HPVL ≥ 1000: 1) was evaluated using the LR model in BLV-positive cases which were dichotomized at a level of 1000 copies/105 cells (Farias et al. 2017). A p-value of < 0.05 was considered statistically significant. An OR greater than 1 indicated an increased risk of outcomes (BLV infection or occurrence of HPVL), and an OR less than 1 indicated a decreased risk of outcomes (BLV infection or occurrence of HPVL). The figures were constructed in a Graph Pad Prism version 7 (GraphPad Software, San Diego, California USA).
Results
Regarding the prevalence of BLV, samples were collected from different provinces representing the various Egyptian geographic and climatic regions. Cattle samples were collected either from animal farms or abattoirs, seeking more inclusive results and interpretation. The total BLV prevalence in cattle from the six investigated Egyptian provinces was 24.2% (105/433) (Table 1). In the case of the southern region, positive samples were recorded only in cattle from Qena province (12.9%; 23/178), while no positive samples have been detected from Luxor (0/16) (Table 1). Samples from Fayoum and Cairo provinces representing the middle region demonstrated a prevalence of 7.7% (1/13) and 77.4% (41/53), respectively (Table 1). Herein, the highest prevalence was detected in Damietta (100%; 40/40) with no positive cases in Beheira province (0/133), both provinces are located in northern Egypt (Table 1). These results demonstrated the distribution of BLV among cattle from various geographical areas in Egypt.
Risk factors analysis was conducted to assess the influence of age, sex, breed, production type, farm size, and location on the prevalence of BLV in cattle in Egypt. The findings indicated that the tested variables were not identified as risk factors except for the breed, farm size, and location factors (Table 2). The prevalence of BLV in cattle < 2 years old (18.7%), set as a reference group was lower than that in cattle 2–5 years old (23.3%, OR = 1.3, 95% CI: 0.6–2.9, p = 0.492), and also than the prevalence in older cattle > 5 years old (26.9%, OR = 1.6, 95% CI: 0.7–3.5, p = 0.251) but not significant in comparison. Moreover, the positivity of BLV was higher in bulls (26.1%) than that in cows (22.9%, OR = 0.8, 95% CI: 0.5–1.3, p = 0.440). While in the case of breed-wise effect, the Mixed cattle breed, set as reference the prevalence was significantly lower against the Holstein breed (36.0%, OR = 2.4, 95% CI: 1.5–3.8, p = 0.0003), but not against the Native breed of cattle tested (21.4%, OR = 1.1, 95% CI: 0.4-3, p = 0.785). Also, the type of production was investigated without recording statistically significant differences among the dairy cows (21.8%) and beef cattle (28.1%, OR = 1.4, 95% CI: 0.9–2.2, p = 0.134). On the contrary, the size of the farm was recorded as a potential risk factor as the highest significant prevalence was found in large-size farms (> 200 heads per farm) (36.0%, OR = 5.3, 95% CI: 2.7–10.4, p < 0.0001), followed by cases from abattoirs (28.1%, OR = 3.9, 95% CI: 1.9–7.5, p < 0.0001) compared to the reference small-size farms (< 200 heads per farm) (9.2%). Regarding the location of the cattle tested, Qena province, where the highest number of samples showed a prevalence of 12.9%, was taken as a reference. A significantly higher prevalence was detected in cattle from Cairo (77.4%, OR = 23, 95% CI: 10.6–50, p < 0.0001), and Damietta (100%, OR = Infinity, p < 0.0001), while a non-significant decrease in prevalence was reported in cattle from Fayoum (7.7%, OR = 0.5, 95% CI: 0.07–4.5, p = 0.7).
The results of the LR model showed the risk effect of four significant variables (breed, type of production, farm size, and location) on the BLV infection (Table 3). Considering the OR, an OR equal to 1 shows no effect; an OR greater than 1 shows the variable in question increases the odds of the outcome (BLV infection); and an OR less than 1 indicates the variable in question decreases the odds of the outcome (BLV infection). Based on the OR evaluation, the Holstein cattle are predicted to be infected with BLV 1.582 (95% CI:1.041–3.04, p = 0.007) times more likely compared to the Native cattle, although, the Mixed breed was not significant (95% CI: 0.106–2.206, p = 0.40). However, beef cattle are predicted to be infected with BLV 1.088 (95% CI: 1.035–3.327, p = 0.0001) times more likely compared to dairy cattle. Additionally, the probability of BLV infection in cattle kept in large farms is 1.26 (95% CI: 1.134–1.503, p = 0.0001) times more likely compared to cattle kept in small farms, while the cattle samples from abattoirs were of non-significant values (OR = 1.44, 95% CI: 0.174–2.361, p = 0.154) when compared to cattle from small farms. Regarding location, cattle from Damietta and Cairo are predicted to be infected with BLV 1.43 (95% CI: 1.005–2.043, p = 0.0001) and 1.16 (95% CI: 1.01–1.41, p = 0.0001) more likely compared to cattle from Qena, respectively. However, cattle from Beheira, Fayoum, and Luxor were of non-significant values (p = 0.994, 0.998, and 0.998 respectively) when compared to cattle from Qena.
The frequency of cattle carrying LPVL or HPVL levels which were classified at a level of 1000 copies per 105 white blood cells was estimated for each category of the risk variables (age, sex, breed, type of production, farm size, and location) (Table 4). In total, the 105 positive cattle samples, in which even one copy is considered positive during interpretation were included. The data indicated HPVL in 21% (22/105), while 79% (84/105) of the tested samples showed LPVL. The mean PVL for all positive cattle was 8651.6 copies / 105 cells. However, in this study, the correlation of PVL with the clinical picture could not be established. Cattle aged < 2 years showed a percentage of 11.1% were carrying HPVL (mean PVL = 138.2), and 2.1% of cattle aged 2–5 years showed HPVL (mean PVL = 140.2), while 42.6% of cattle > 5 years were carrying HPVL (mean PVL = 19,177). Females showed a higher frequency of HPVL (35.7%; mean PVL = 16,099) than males (4.1%; mean PVL = 160.1). The Native cattle showed 16.7% with HPVL (mean PVL = 195.5), and only 1.9% of Mixed cattle showed HPVL (mean PVL = 130.9), while 44.4% of Holstein cattle were carrying HPVL (mean PVL = 20,027). Dairy cows showed a higher frequency of HPVL (34.5%; mean PVL = 15,545) than beef cattle (4.3%; mean PVL = 166.1). Regarding farm size, none of the animals from small farms showed HPVL (0%; mean PVL = 31.08), while cattle from large farms showed the highest frequency of HPVL (44.4%; mean PVL = 20,027), and samples from abattoirs showed HPVL in 4.3% (mean PVL = 166.1). Additionally, cattle from Damietta showed the highest frequency of HPVL (50%; mean PVL = 22,522) compared to cattle from Cairo showed HPVL (4.9%, mean PVL = 173.5), while none of the cattle from Qena showed HPVL (0%, mean PVL = 43.8). Moreover, only one positive sample from Fayoum was detected and carried LPVL (433 copies/105 cells).
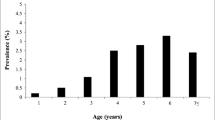
In addition, the above-mentioned factors (age, sex, breed, type of production, farm size, and location) have been analyzed as risk factors considering the PVL instead of the BLV prevalence (Fig. 2). A significantly higher PVL was recorded in older cattle > 5 years old than those of 2–5 years old (p < 0.0001) and cattle < 2 years old (p < 0.0001). Moreover, the lower PVL in cattle < 2 years old compared to 2–5 years old cattle was statistically significant (p = 0.04) (Fig. 2A). Regarding sex, cows were more vulnerable to BLV than bulls as indicated by statistically higher PVL in female cattle than males (p = 0.0008) (Fig. 2B). Similarly, to those obtained using prevalence, a breed-wise effect significantly affected the PVL in the current study. The Holstein breed demonstrated a higher PVL than those in the Mixed cattle breed (p < 0.0001) and Native cattle (p < 0.0001), while no significant difference has been reported between the Native and Mixed breeds (p = 0.13) (Fig. 2C). Consistently to those recorded to sex-wise effect, a highly significant difference was obtained among dairy (cows) and beef cattle (bull) as dairy cattle had a higher PVL (p = 0.0053) (Fig. 2D). Also, cattle housed in large-size farms showed significantly higher PVL than those bred in small-size farms (p < 0.0001), and sporadic cases from abattoirs (p < 0.0001), and also significantly lower in small-size farms compared to cases from abattoirs (p = 0.02) (Fig. 2E).
The influence of the risk factors, (A) age, (B) sex, (C) breed, (D) production type, (E) farm size and (F) location on the BLV PVL (log10 copies / 105 cells) among the infected Egyptian cattle. Black bars show the median of log10PVL and 95% confidence interval values. Significant differences within the different risk factors were calculated using SPSS version 25, then the figure was constructed using Graph Pad Prism version 7. A P < 0.05 were considered statistically significant. The comparison of p-values of BLV PVL between the groups was estimated by Kruskal-Wallis with Dunn’s multiple comparisons test (age, breed, farm size, and location) and Mann-Whitney U Test (sex and production type), and was indicated above the error bars
Regarding PVL levels in the cattle tested from different locations (Fig. 2F), samples from Damietta showed the highest PVL, while a significant decrease in PVL in cattle from Cairo (p < 0.0001), and Qena (p < 0.0001) in comparison was recorded. Furthermore, a significant increase in PVL levels in cattle from Cairo compared to Qena (p = 0.018) was shown.
The estimates from the LR model which analyzed the risk of age on the occurrence of HPVL among the infected cattle were shown (Table 5). The data revealed that the probability of carrying HPVL in cattle > 5 years is 1.27 (95% CI: 1.03–2.09, p < 0.001) times more likely compared to cattle < 2 years old. However, the probability of carrying HPVL in cattle 2–5 years compared to cattle < 2 years old was not significant (p = 0.999). Such results regarding PVL have made a valuable addition by correlating the BLV prevalence with PVL as an indicator of the severity of the infection.
Discussion
The geography of Egypt relates to two continents, North Africa and Southwest Asia. It has an area of 1,002,450 km2 which is divided into 28 provinces. Approximately, 5% of the total area is cultivable regions, which support more than 100 million humans and 18 million animals, of them about 5 million cattle heads (FAO stat. 2018). The majority of Egyptian cattle are reared for dairy purposes however male calves and infertile females are reared for beef production (Abdi et al. 2020). The updated statistical reports recorded a gradual decrease in the number of cattle in Egypt due to outbreaks of infectious diseases (FAO stat, 2020). Little is known about the impact of BLV infection in both the dairy and beef cattle industries due to the lack of BLV screening on a large scale and its asymptomatic form in most animals (Suarez Archilla et al. 2022). In this study, firstly, we performed a random screening of BLV among dairy and beef animals from six Egyptian provinces distributed in different climatic and geographical regions. Secondly, the risk factors influencing BLV infection were analyzed. Finally, the correlations between the BLV PVL levels in the infected animal’s blood and the analytic risk factors were estimated.
This study reported a 24.2% total BLV prevalence among the cattle tested by using the BLV-CoCoMo- qPCR-2 assay. Noticeably, this BLV molecular prevalence was relatively high compared to the previous serological evidences that reported a 15.8% infection rate in cattle from Alexandria, Kafr-Elsheikh, and Menofia (Zaher and Ahmed. 2014), and 20.8% in Menofia and Qalyubia (Selim et al. 2019). This data is in accordance with the theory that BLV prevalence is increasing in many countries all over the world (Bartlett et al. 2020). Another reason for the higher prevalence in this study could be the method of BLV diagnosis as we used the BLV-CoCoMo- qPCR-2 method which is able to detect even one copy of the integrated BLV provirus in the host genome (Takeshima et al. 2015). This method was highly effective in the detection of BLV in cattle from different countries (Polat et al. 2016; Moe et al. 2020; Lo et al. 2020), and it can detect new infections, before the development of antibodies to BLV (Takeshima et al. 2015). It was proven to be an effective method for monitoring the progression of BLV-induced disease and allowed us to evaluate the BLV infection from a different viewpoint compared with classical serological methods (Jimba et al. 2012). Additionally, the study area and sample population of cattle investigated in this study were different from the previous studies, which included animals only from the north of Egypt. Our study was more comprehensive and represented more different geographical and climatic localities of Egypt in the Southern, Middle, and Northern. The geography might be also a reason for a highly variable BLV prevalence between the examined provinces. The climatic changes could play an important role in the BLV distribution everywhere as the blood-sucking insects are important route of BLV transmission in some parts of the world in addition to iatrogenic routes in other regions (Bartlett et al. 2020). In this context, the climate of Egypt is very confusing as in the southern regions, the climate is very dry, hot, and rarely rains, however, humid in the middle, and cold and rainy in the northern regions. This could be a reason for the high prevalence of BLV in Damietta which is located on the branch of the River Nile and includes many ponds, canals, and rice fields where the flies prefer to inhabit (Selim et al. 2020b). Furthermore, Damietta is a coastal province, where an important seaport for the entry of imported animals from worldwide to Egypt is present. However, this study did not investigate whether the entry of cattle could be a risk factor for BLV infection in this region because in this province we investigated only a single large dairy farm. Therefore, we think about a future thorough screening of BLV in such an important area. In contrast, Selim and colleagues did not find significant associations between the BLV prevalence and the studied geographic regions in the Nile Delta of Egypt because of the similar geographic nature (Selim et al. 2020a, 2021).
Regarding risk factors analysis in this study, the BLV prevalence was greatly influenced by the cattle breeds and the size of comprising farm. Interestingly, higher prevalence was reported among the imported Holstein cattle which reared in large-size farms than Mixed and Native cattle which reared in small farms. Because of the low productivity of milk in the Egyptian Native and Mixed cattle, Egypt imports Holstein cattle from Germany, the Netherlands, and the United States (US) to produce higher quantities of milk for consumers and to improve the genetic characteristics of the Egyptian Native and Mixed cattle via breeding (Abdi et al. 2020). The imported cattle are usually kept in large-size farms comprising high numbers of animals. In this study, Holstein cows tested were imported as pregnant heifers from the US. To our knowledge, the veterinarians in the Egyptian quarantines randomly investigate some imported cattle for the presence of antibodies to BLV using serological methods. However, it might be a new infection before the appearance of antibodies, since the BLV prevalence in cattle in the US is approximately 46.5% as reported in a wide national survey (LaDronka et al. 2018). These findings were supported by Zaghawa and colleagues in 2002, who reported a 50.3% of BLV infection rate after the appearance of BLV symptoms in a dairy herd of Holstein Friesian cattle imported from the US (Zaghawa et al. 2002). Therefore, we believe that a comprehensive study to detect BLV among imported cattle in quarantines based on various techniques could shed more light on the risk of imported cattle from everywhere to spread BLV infection in Egypt. Selim and colleagues indicated that the herd size and housing were potential risk factors for BLV prevalence after a serological screening of cattle from the Nile Delta in Egypt (Selim et al. 2020a, 2021). Additionally, our data agree with the studies that found increased within-herd prevalence of BLV in larger herds compared to small herds (Baumgartener et al. 1975; Haghparast et al. 2012). However, conversely to our data, some studies reported age as another risk factor influencing BLV infection (Nagy et al. 2007; Erskine et al. 2012; Selim et al. 2020a). We think that the influence of age on the BLV infection in this study might be affected by the testing of a low number of samples from cattle < 2 years (n = 48) compared to the previous studies (Nagy et al. 2007; Erskine et al. 2012; Selim et al. 2020a). With regards to beef cattle showed a higher BLV prevalence than dairy cattle in this study, it might be one of the reasons that the beef animals were young ages and originally isolated from dairy mothers to be kept in overcrowded feedlots in Egypt. Furthermore, calves probably get the infection from the milk of infected mothers with high PVL and/or by other horizontal routes of transmission (Yuan et al. 2015; Watanuki et al. 2019). This result supports the reports of the global increase of BLV prevalence everywhere (Bartlett et al. 2020) and predicts a future increase in BLV infection in Egyptian cattle.
BLV-positive samples were dichotomized at a level of 1000 copies/105 cells into LPVL and HPVL as previously described (Farias et al. 2017). In total, our results reported a high BLV PVL in the blood of the BLV-positive cattle (mean = 8651.6 copies/105 cells). However, we did not notice any clinical symptoms of BLV such as lymphadenitis and tumors among the examined animals. This in agreement with the theory proved that the majority of infected animals are asymptomatic carriers (Aida et al. 2013; Polat et al. 2017). Our findings were supported by the results of Lo and colleagues who reported a strong association between the BLV PVL and occurrence of lymphoma in cattle (Lo et al. 2020). They found that asymptomatic cows carried a mean PVL of 9,401 copies/105 cells, while cows induced-lymphoma carried a mean PVL of 99,522 copies/105 cells. Similarly, Kobayashi and colleagues reported a higher PVL in EBL cattle compared to asymptomatic cattle (Kobayashi et al. 2020). In this context, the HPVL reported in Egyptian cattle investigated in this study could be a potential risk factor for disease transmission within the infected farms via multiple routes of BLV transmission (Juliarena et al. 2016). Additionally, previous studies detected the BLV provirus in milk, saliva, and nasal discharge increasing the possibility of BLV transmission horizontally by licking, sneezing, rubbing the nose, or milking (Yuan et al. 2015; Watanuki et al. 2019). Therefore, we think that HPVL reported in cattle from the Damietta dairy farm (mean PVL = 22,522) could be a reason for 100% BLV prevalence in the samples tested.
The association between the risk factors (age, sex, breed, production type, farm size and location) and the BLV PVL levels revealed significant differences within all categories of the infected cattle. Moreover, the effect of age on the occurrence of HPVL was analyzed in detail by the LR model. The older cattle group > 5 years old showed a higher BLV PVL, and they are predicted to carry HPVL 1.27 times more likely than the youngest group. However, no significant difference was ultimately proven in BLV PVL levels between the younger groups of infected cattle. This data is consistent with studies linked the PVL to the longevity of the infected animals as old cows are more likely to develop the malignant form of B-cell lymphoma after a latency (Zaghawa et al. 2002; Gillet et al. 2007; Radostits et al. 2007; Aida et al. 2013; Gutiérrez et al. 2014; Lo et al. 2020). Furthermore, our findings agree with Lo and colleagues who indicated that age was significantly associated with BLV susceptibility and PVL levels (Lo et al. 2020). A combination of sex, breed, production type, and the farm size risk factors effects on the BLV PVL revealed a significant increase in PVL in infected female Holstein dairy cows that were kept in large-size farms than PVL in infected males, Native and Mixed beef cattle that kept in small farms and sporadic cases from abattoirs. However, no significant difference was observed in BLV PVL levels in infected Native and Mixed cattle breeds. As previously reported BLV is likely to spread more in large-sized than small-sized herds (Kobayashi et al. 2010; Haghparast et al. 2012; Hamada et al. 2020; Selim et al. 2020a). Therefore, the importation of cows with high BLV PVL from BLV-endemic countries is considered a potential risk for BLV as it could transmit the infection to BLV-free cows. However, the Native and Mixed cattle that carry LPVL are possibly at fewer risks of BLV transmission (Juliarena et al. 2016). Additionally, other genetic factors in the Native and Mixed cattle could be associated with BLV susceptibility and PVL levels. In this context, some studies referred the BLV susceptibility and the PVL levels to the polymorphism in the bovine leukocyte antigen (BoLA)-DRB3 in the Holstein and Japanese black cattle (Juliarena et al. 2016; Lo et al. 2020). However, it is poorly investigated in the Egyptian breeds.
Despite this study being the first to evaluate the BLV PVL levels in Egyptian cattle, it had two limitations that should be mentioned. First, it was carried out on a few samples from some provinces because of difficulties with the owner’s permission for sampling. Second, some risk factors related to cattle management and BLV transmission were not included because of inadequate information in the farms’ records. Therefore, we believe that a comprehensive nationwide study of BLV infection and PVL across various Egyptian provinces and an analysis of more risk factors could shed more light on the disease’s epidemiology.
Conclusion
In conclusion, this study gives insight into the effect of the risk factors, age, sex, breed, production type, farm size and location of the cattle tested on the prevalence of BLV among Egyptian dairy and beef cattle from different geographical and climatic regions. Holstein breed, beef cattle, large-size farms and cattle from Damietta and Cairo were ultimately proven as the most important risk for BLV infection in Egyptian cattle. Additionally, the BLV PVL levels were absolutely high and greatly variable within the different categories of tested cattle. Females, Holstein, dairy cows, older than 5 years, that were kept in large farms, and cattle from Damietta have maintained the highest BLV PVL levels and could act as a source of BLV transmission to BLV-free animals. Ultimately, older cattle are predicted to carry HPVL 1.27 times more likely compared to younger age. However, the association between the BLV prevalence as well as the BLV PVL and the clinical picture of EBL in Egyptian cattle is needed.
Data Availability
All data included in this study are available on request from the corresponding author.
References
Abdi A, Roushdy S, Beillard M et al (2020) Egypt Livestock and Products Annual 2019 egyptian beef prices stable, U. S. Beef Imports Challenged. 1–12
Aida Y, Murakami H, Takahashi M, Takeshima SN (2013) Mechanisms of pathogenesis induced by bovine leukemia virus as a model for human T-cell leukemia virus. Front Microbiol 4:1–11. https://doi.org/10.3389/fmicb.2013.00328
Bartlett PC, Norby B, Byrem TM et al (2013) Bovine leukemia virus and cow longevity in Michigan dairy herds. J Dairy Sci 96:1591–1597
Bartlett PC, Ruggiero VJ, Hutchinson HC et al (2020) Current developments in the epidemiology and control of enzootic bovine leukosis as caused by bovine leukemia virus. Pathogens 9:1058
Barzegar H, Mirshahabi H, Motamed N et al (2021) Identification of bovine leukemia virus in raw milk samples in North-West of Iran. Veterinary Res Forum Vol 12(2):223
Baumgartener LE, Olson C, Miller JM, Van Der Maaten MJ (1975) Survey for antibodies to leukemia (C-type) virus in cattle. J Am Vet Med Assoc 166:249–251
Burny A, Cleuter Y, Kettmann R et al (1988) Bovine leukaemia: facts and hypotheses derived from the study of an infectious cancer. Virus Infect Dev Nerv Syst 37–56
Erskine RJ, Bartlett PC, Byrem TM et al (2012) Using a herd profile to determine age-specific prevalence of bovine leukemia virus in Michigan dairy herds. Vet Med Int 2012:1–5
Farías M, Caffaro M, Lendez P et al (2017) A novel association of BoLA DRB3 alleles with BLV infected cattle with different proviral loads. Braz J Vet Res Anim Sci 54:3:215–224. https://doi.org/10.11606/issn.1678-4456.bjvras.2017.123769
Food Agriculture Organization (2020) World Food and Agriculture - Statistical Yearbook 2020
Food Agriculture Organization (2018) Agency US, Development I livestock production systems spotlight cattle and buffaloes and poultry sectors livestock production systems spotlight cattle and buffaloes, and poultry sectors in Egypt. http://www.fao.org/ag/againfo/programmes/en/ASL2050.html
Frie MC, Coussens PM (2015) Bovine leukemia virus: a major silent threat to proper immune responses in cattle. Vet Immunol Immunopathol 163:103–114
Gillet N, Florins A, Boxus M et al (2007) Mechanisms of leukemogenesis induced by bovine leukemia virus: prospects for novel anti-retroviral therapies in human. Retrovirology 4:1–32
Gutiérrez G, Rodríguez SM, De Brogniez A et al (2014) Vaccination against δ-retroviruses: the bovine leukemia virus paradigm. Viruses 6:2416–2427. https://doi.org/10.3390/v6062416
Haghparast A, Tabatabaei Zadeh SE, Mohammadi GR (2012) Prevalence of bovine leukemia virus (BLV) antibodies in bulk tank milk of dairy cattle herds of Mashhad area, North East of Iran. J Anim Vet Adv 11
Hamada R, Metwally S, Polat M et al (2020) Detection and molecular characterization of bovine leukemia virus in egyptian dairy cattle. Front Vet Sci 7:608
Hopkins SG, DiGiacomo RF (1997) Natural transmission of bovine leukemia virus in dairy and beef cattle. Vet Clin North Am Food Anim Pract 13:107–128
Jimba M, Takeshima SN, Matoba K et al (2010) BLV-CoCoMo-qPCR: quantitation of bovine leukemia virus proviral load using the CoCoMo algorithm. Retrovirology 7:91. https://doi.org/10.1186/1742-4690-7-91
Jimba M, Takeshima S, Murakami H et al (2012) BLV-CoCoMo-qPCR: a useful tool for evaluating bovine leukemia virus infection status. BMC Vet Res 8:1–12
Juliarena MA, Barrios CN, Ceriani MC, Esteban EN (2016) Hot topic: bovine leukemia virus (BLV)-infected cows with low proviral load are not a source of infection for BLV-free cattle. J Dairy Sci 99:4586–4589
Kettmann R, Meunier-Rotival M, Cortadas J et al (1979) Integration of bovine leukemia virus DNA in the bovine genome. Proc Natl Acad Sci 76:4822–4826
Kobayashi S, Tsutsui T, Yamamoto T et al (2010) Risk factors associated with within-herd transmission of bovine leukemia virus on dairy farms in Japan. BMC Vet Res 6:1–6. https://doi.org/10.1186/1746-6148-6-1
Kobayashi S, Hidano A, Tsutsui T et al (2014) Analysis of risk factors associated with bovine leukemia virus seropositivity within dairy and beef breeding farms in Japan: a nationwide survey. Res Vet Sci 96:47–53
Kobayashi T, Inagaki Y, Ohnuki N et al (2020) Increasing bovine leukemia virus (BLV) proviral load is a risk factor for progression of enzootic bovine leucosis: a prospective study in Japan. Preventive veterinary medicine. 178:104680
LaDronka R, Ainsworth S, Wilkins M et al (2018) Prevalence of bovine leukemia virus antibodies in US dairy cattle. Vet Med Int 1–8. https://doi.org/10.1155/2018/5831278
Lo C-W, Borjigin L, Saito S et al (2020) BoLA-DRB3 polymorphism is associated with differential susceptibility to bovine leukemia virus-induced lymphoma and proviral load. Viruses 12:352
Matsuura R, Inabe K, Otsuki H et al (2019) Three YXXL sequences of a bovine leukemia virus transmembrane protein are independently required for fusion activity by controlling expression on the cell membrane. Viruses 11:1–22. https://doi.org/10.3390/v11121140
Metwally S, Hamada R, Ali AO et al (2020) Detection and molecular characterization of bovine leukemia virus in beef cattle presented for slaughter in Egypt. J Vet Med Sci 20–477
Moe K, Polat M, Borjigin L et al (2020) New evidence of bovine leukemia virus circulating in Myanmar cattle through epidemiological and molecular characterization. PLoS ONE 15(2):e0229126
More S, Bøtner A, Butterworth A et al (2017) Assessment of listing and categorisation of animal diseases within the framework of the Animal Health Law (Regulation (EU) no 2016/429): enzootic bovine leukosis (EBL). EFSA J. https://doi.org/10.2903/j.efsa.2017.4956. 15:
Nagy DW, Tyler JW, Kleiboeker SB (2007) Decreased periparturient transmission of bovine Leukosis virus in colostrum-fed calves. J Vet Intern Med 21:1104–1107
Nakatsuchi A, Watanuki S, Borjigin L et al (2022) BoLA-DRB3 polymorphism controls proviral load and infectivity of bovine leukemia virus (BLV) in milk. Pathogens 11(2):210
Norby B, Bartlett PC, Byrem TM, Erskine RJ (2016) Effect of infection with bovine leukemia virus on milk production in Michigan dairy cows. J Dairy Sci 99:2043–2052
Ohno A, Takeshima S, Matsumoto Y, Aida Y (2015) Risk factors associated with increased bovine leukemia virus proviral load in infected cattle in Japan from 2012 to 2014. Virus Res 210:283–290
Polat M, Takeshima S, Hosomichi K et al (2016) A new genotype of bovine leukemia virus in South America identified by NGS-based whole genome sequencing and molecular evolutionary genetic analysis. Retrovirology 13(1):1–23
Polat M, Takeshima SN, Aida Y (2017) Epidemiology and genetic diversity of bovine leukemia virus. Virol J 14:1–16. https://doi.org/10.1186/s12985-017-0876-4
Radostits OM, Gay CC, Hinchcliff KW, Constable PD (2007) A textbook of the diseases of cattle, horses, sheep, pigs and goats. Vet Med 10:2045–2050
Saito S, Hosomichi K, Yamanaka MP et al (2022) Visualization of clonal expansion after massive depletion of cells carrying the bovine leukemia virus (BLV) integration sites during the course of disease progression in a BLV naturally-infected cow: a case report. Retrovirology 19:1–11
Sato H, Watanuki S, Murakami H et al (2018) Development of a luminescence syncytium induction assay (LuSIA) for easily detecting and quantitatively measuring bovine leukemia virus infection. Arch Virol 163:1519–1530
Schwartz I, Lévy D, Schwartz I et al (1994) Pathobiology of bovine leukemia virus to cite this version: HAL id : hal-00902257. Rev article 25:521–536
Selim A, Marawan MA, Ali AF et al (2019) Seroprevalence of bovine leukemia virus in cattle, buffalo, and camel in Egypt. Trop Anim Health Prod 3–6. https://doi.org/10.1007/s11250-019-02105-8
Selim A, Megahed AA, Kandeel S, Abdelhady A (2020a) Risk factor analysis of bovine leukemia virus infection in dairy cattle in Egypt. Comp Immunol Microbiol Infect Dis 72:101517
Selim A, Radwan A, Arnaout F, Khater H (2020b) The recent update of the Situation of West Nile Fever among Equids in Egypt after three decades of missing information. Pak Vet J 40
Selim A, Manaa EA, Alanazi AD, Alyousif MS (2021) Seroprevalence, risk factors and molecular identification of bovine leukemia virus in egyptian cattle. Animals 11:319
Somura Y, Sugiyama E, Fujikawa H, Murakami K (2014) Comparison of the copy numbers of bovine leukemia virus in the lymph nodes of cattle with enzootic bovine leukosis and cattle with latent infection. Arch Virol 159:2693–2697
Suarez Archilla G, Gutierrez G, Camussone C et al (2022) A safe and effective vaccine against bovine leukemia virus. Front Immunol 13
Sultanov A, Rola-Łuszczak M, Mamanova S et al (2022) Molecular characterization of bovine leukemia virus with the evidence of a new genotype circulating in cattle from Kazakhstan. Pathogens 11:180
Tajima S, Ikawa Y, Aida Y (1998) Complete bovine leukemia virus (BLV) provirus is conserved in BLV-infected cattle throughout the course of B-cell lymphosarcoma development. J Virol 72:7569–7576
Takeshima S, Kitamura-Muramatsu Y, Yuan Y et al (2015) BLV-CoCoMo-qPCR-2: improvements to the BLV-CoCoMo-qPCR assay for bovine leukemia virus by reducing primer degeneracy and constructing an optimal standard curve. Arch Virol 160:1325–1332
Thrusfield M (2018) Veterinary epidemiology. John Wiley & Sons, pp 1–864
Watanuki S, Takeshima SN, Borjigin L et al (2019) Visualizing bovine leukemia virus (BLV)-infected cells and measuring BLV proviral loads in the milk of BLV seropositive dams. Vet Res 50:1–12. https://doi.org/10.1186/s13567-019-0724-1
White TL, Moore DA (2009) Reasons for whole carcass condemnations of cattle in the United States and implications for producer education and veterinary intervention. J Am Vet Med Assoc 235:937–941
Yuan Y, Kitamura-Muramatsu Y, Saito S et al (2015) Detection of the BLV provirus from nasal secretion and saliva samples using BLV-CoCoMo-qPCR-2: comparison with blood samples from the same cattle. Virus Res 210:248–254
Zaghawa A, Beier D, Abd El-Rahim IHA et al (2002) An outbreak of enzootic bovine leukosis in upper Egypt: clinical, laboratory and molecular-epidemiological studies. J Vet Med Ser B 49:123–129. https://doi.org/10.1046/j.1439-0450.2002.00517.x
Zaher K, Ahmed W (2014) Bovine leukemia virus infection in dairy cows in Egypt. Acad J Cancer Res 7:126–130. https://doi.org/10.5829/idosi.ajcr.2014.7.2.83265
Acknowledgements
We thank the veterinarians and our collaborators for kindly assisting with sampling from many farms and slaughterhouses in Egypt. Proviral load analysis was performed in the Viral Infectious Diseases Field, RIKEN, Japan under the supervision of Prof. Yoko AIDA and we are grateful to Prof. Y. AIDA and members for their technical assistance, help, and suggestions.
Funding
This work was supported by a Grant-in-Aid for Scientific Research (A) from the Japan Society for the Promotion of Science (JSPS) (Grant no. 16H02590) and by a grant from the Embassy of Egypt Culture, Education and Science Bureau.
Author information
Authors and Affiliations
Contributions
Conceptualization and design; SM, RH., Experiments, formal analysis, investigation; RH, SM, RF; Resources and shared materials; RH, SM., Writing—original draft, RH, SM, RF; Writing—review and editing; SM, RH.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hamada, R., Fereig, R.M. & Metwally, S. The influence of risk factors on bovine leukemia virus infection and proviral load in egyptian cattle. Vet Res Commun 48, 191–202 (2024). https://doi.org/10.1007/s11259-023-10198-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11259-023-10198-8