Abstract
Flowering phenology can be one of the most important factors mediating the temporal dynamics of plant–pollinator networks. However, most studies do not explicitly incorporate the effect of flowering phenology, which may bias conclusions about the structuring of plant–pollinator networks, obscuring our understanding of factors that explain the temporal variation of these networks. By analyzing co-flowering networks, floral traits similarity and pollinator sharing, in this study we aim to uncover the temporal dynamics of plant–pollinator network structure in two coastal communities. We recorded the flowering phenology of each plant species to construct co-flowering networks and the identity and frequency of floral visitors. We estimated and tested for differences in floral trait similarity and pollinator sharing across co-flowering modules. To disentangle the phenological effect of flowering on the structure of plant–pollinator networks, we constructed plant–pollinator subnetworks for each co-flowering modules and analyzed the role of the pollinators in each subnetwork. Floral trait similarity and pollinator sharing were related to changes in the structure of plant–pollinator networks, but these changes were community-dependent. The modular structure and network specialization index of plant–pollinator subnetworks were statistically persistent in both communities, suggesting the prevalence of specialized interactions throughout the flowering season. This result was consistent with the predominant peripheral role of most pollinator species across co-flowering modules in both communities. Our results highlight the importance of explicitly considering flowering phenology to advance our understanding of the mechanisms that explain temporal changes in the structure of plant–pollinator networks.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The study of interaction networks between plants and pollinators has helped understand how these communities are assembled (Bascompte et al. 2003; Bascompte and Jordano 2007; Heleno et al. 2014; Tylianakis and Morris 2017). However, the structure of plant–pollinator networks typically vary in space and time (e.g., Petanidou et al. 2008; Trøjelsgaard et al. 2015; Ramos-Robles et al. 2016; Biella et al. 2017; Wang et al. 2020), which has led to attempts to understand the factors mediating their structure and stability (e.g., Olesen et al. 2002; Stang et al. 2006; Vázquez et al. 2009; Olito and Fox 2015; Parra-Tabla et al. 2019; CaraDonna et al. 2021). Flowering phenology (i.e., temporal sequence of species flowering) can be one of the most important factors mediating the temporal dynamic of plant–pollinator networks. This is because it determines the phenological match between pairs of interacting plant and pollinator species and the level of pollinator sharing between co-flowering species (Stang et al. 2006; Aizen and Vázquez 2006; Elzinga et al. 2007; Valdovinos 2019; Peralta et al. 2020; and see Waser and Real 1979; Rathcke and Lacey, 1985). Pollinator sharing is important because it defines the level of generalization in plant–pollinator interactions (Ghazoul 2006; Sargent and Ackerly 2008). In fact, pollinator sharing has been reported to be high in co-flowering communities and is therefore considered one of the main factors explaining the generalist structure of pollination networks (i.e., nestedness structure; Bascompte et al. 2003; Bascompte and Jordano 2007). However, this pattern of generalized interactions has been based on estimates that pool all interactions occurring throughout the flowering season. This approach however likely fails to account for temporal changes in the identity and flowering intensity of plant species, as well as it can overestimate pollinator sharing (Biella et al. 2017; Valdovinos 2019; CaraDonna et al. 2021; Guzman et al. 2021). This in turn may lead to biased conclusions about the level of generalization in plant–pollinator networks.
Recent attempts have been made to incorporate the effect of flowering phenology on the structure of plant–pollinator networks by using “snapshots” at different time scales (e.g., days, weeks, months, seasons; Rasmussen et al. 2013; Wang et al. 2020; Schwarz et al. 2020; CaraDonna et al. 2021). This approach has not only shown continuous changes in the structure of plant–pollinator networks, but has also revealed that generalization is not a persistent property of the networks (e.g., Rasmussen et al. 2013; Biella et al. 2017; Schwarz et al. 2020). However, the arbitrary selection of different time scales still overlooks the natural variation in flowering overlap between plant species and it may obscure understanding of other mechanisms that explain plant–plant interactions via pollinator sharing. For example, it is known that the level of pollinator sharing also depends on the degree of floral trait similarity between co-flowering plants (e.g., Moeller 2004; Ghazoul 2006; Gibson et al. 2012; Sargent and Ackerly 2008). Thus, increases in the level of flowering overlap and floral trait similarity should increase pollinator sharing, resulting in a predominant generalized structure of plant–pollinator networks (Lázaro et al. 2020; Suárez-Mariño et al. 2022). However, it is also possible that even at high flowering overlap, low levels of floral trait similarity reduce the level of generalization due to a decrease in pollinator sharing, thus maintaining pollinator specialization (Albor et al. 2020; Suárez-Mariño et al. 2022). Therefore, considering both, the level of flowering overlap and the level of floral trait similarity in co-flowering communities, it is important to better understand the mechanisms involved in the structure of plant–pollinator networks over time.
A more unbiased approach to test the level of flowering overlap and the level of floral trait similarity could be grounded on the fact that flowering can be organized as co-flowering module within networks, which consists of the grouping of plant species (i.e., co-flowering modules Arceo-Gómez et al. 2018, or pheno-clusters sensu Biella et al. 2017) that showed a greater flowering overlap respect to other groups of plants in the community (Arceo-Gómez et al. 2018; Albor et al. 2020; and see Waser and Real 1979; Rathcke and Lacey 1985). Therefore, plants of the same co-flowering module are expected to interact more intensively due to the temporal coincidence in pollinator sharing, although it could be constrained by the level of floral trait similarity (Arceo-Gómez et al. 2018; Albor et al. 2020). For example, Albor et al. (2020) found in sand dune coastal communities that pollinator sharing varied within and between co-flowering modules depending on differences in floral trait similarity. Thus, it can be suggested that plant–pollinator subnetworks resulting from flowering grouping species should emerge throughout the flowering season, revealing more precisely the role of natural variation of flowering phenology and floral trait similarity on pollinator sharing and finally, in the temporal changes of plant–pollinator networks. For example, Biella et al. (2017) considered differences in flowering overlap in two semi-dry grassland communities to construct plant–pollinator subnetworks and found significant changes in the generalized but not in the specialized structure (i.e., nestedness and modularity, respectively) of the subnetworks. Interestingly, Biella et al. (2017) also found that the changes in network structure were associated with changes in the role played by pollinator within the networks (e.g., network-hub species or peripherals species). These results suggest that changes in the role of pollinator species over the flowering season depends on the identity and floral traits of the flowering species, which influence the level of plant generalization/specialization (see Junker et al. 2010; Coux et al. 2016; Wang et al. 2020).
In this study, we analyzed the co-flowering structure that arise naturally due to interspecific differences in the flowering phenology of plant species and the plant–pollinator subnetworks that result when considering such co-flowering structure, in two coastal communities (i.e., sand dune and scrubland), to better understand the mechanisms that guide the structuring of plant–pollinator networks. Specifically, we aimed to answer the following questions: (a) do flower trait similarity and pollinator sharing in co-flowering modules explain temporal changes in the generalist or specialist structure of plant–pollinator networks? and, (b) do interspecific differences in flowering phenology and floral trait similarity affect the role of pollinators in plant–pollinator subnetworks? We expected that co-flowering modules with high floral trait similarity would show greater pollinator sharing and consequently a generalized structure of plant–pollinator networks. In contrast, co-flowering modules with low floral trait similarity would show low pollinator sharing, contributing to a more specialized network structure. Similarly, we expected that the role played by pollinators in plant–pollinator networks will vary among co-flowering modules. Our results provide a better understanding of the temporal changes and mechanisms involved in the structuring of plant–pollinator networks.
Methods
Study site
We recorded flowering phenology of each plant species and plant–pollinator interactions in a dune and a coastal scrubland community near the town of Telchac in the Yucatan Peninsula, Mexico (21° 20′ 11.7″ N, 89° 20′ 12.5″ W; 0 to 8 m a.s.l.). The climate is hot and dry, with a seasonal rainfall and annual precipitation of 760 mm and a mean annual temperature of 26 °C (Orellana et al. 2009). Both communities are exposed to adverse abiotic conditions (e.g., low rainfall and high temperatures) and are characterized by halophyte and xerophytic vegetation (Espejel 1987) (Fig. S1 a, b). While in the dune, the species grow on a mobile substrate with scarce nutrients, being affected by the wind and the increase in salinity, in the scrubland the species grow on a more stable soil (due to the accumulation of organic matter) and are more tolerant to strong winds (Espejel 1987; Parra-Tabla et al. 2018). Both communities are adjacent and share some plant species (e.g., Bidens pilosa, Melanthera nivea, Porophyllum punctatum, and Okenia hypogaea) but differ significantly in their plant composition (PERMANOVA: F1,18 = 4.29, p < 0.005; Suárez-Mariño et al. 2022). The pollinator composition of these communities is characterized by a large group of insects, mainly Hymenoptera species (Campos-Navarrete et al. 2013; Parra-Tabla et al. 2019; Albor et al. 2020). A previous study of the insect community showed no significant differences in the composition of floral visitors between dune and scrubland communities (PERMANOVA: F1,18 = 1.35, p > 0.05; Suárez-Mariño et al. 2022).
Co-flowering networks
In 2019, we recorded the number of open flowers and the duration of the flowering phenology of each plant species during the flowering season which corresponded to the rainy season (August-December) (Table S1). Previous studies have shown that during this period more than 70% of the species in these communities produce flowers (Campos-Navarrete et al. 2013; Albor et al. 2019; Parra-Tabla et al. 2019). Each community was visited twice per month (10 days in total per community), covering the entire flowering season of most plant species. In each visit, the plant identity and number of open flowers were recorded in ten 20 m2 (10 × 2 m) plots spaced 5 m apart. This sampling effort in these communities has proven to be sufficient to have a good representation of the number of flowering species and the intensity of flowering (i.e., number of flowers produced per species) (Parra-Tabla et al. 2021; Suárez-Mariño et al. 2022).
The co-flowering networks for both communities were constructed using Schoener’s niche overlap Index (Schoener 1970), SI = 1– (1/2) ∑k | Pik–Pjk |, where Pik y Pjk are the proportion of flowering of species i and j respectively, occurring on day k (Forrest et al. 2010; Arceo-Gómez et al. 2018), then the degree of temporal flowering overlap between each pair of plant species was calculated (Arceo-Gómez et al. 2018). The SI index considers the intensity (i.e., number of flowers produced per species) and frequency (i.e., number of samplings in which each species showed flowers) of temporal flowering overlap between each plant species pair. Therefore, species pairs with a greater SI overlap not only flower simultaneously for longer periods of time, but also do so with greater intensity. Following Arceo-Gómez et al. (2018) we used the SI to construct unidirectional co-flowering network for each community with the program Gephi version 9.3 (Bastian et al. 2009). Thus for each co-flowering network, we identified species that interact more strongly with each other than with other species when flowering at the same time and with the same intensity (i.e., co-flowering modules; see Arceo-Gómez et al. 2018). The co-flowering network modularity (Q) was estimated following Emer et al. (2015) by transforming the flowering networks into bipartite quantitative matrices of the form m × n, where m and n are flowering species in the same community and flowering season, and then estimating modularity by means of the ‘QuanBiMo’ algorithm (Dormann and Strauss 2014). To test the statistical significance of co-flowering modularity (Q) in each community, we used a null model analysis where we compared the observed modularity in the co-flowering networks against the expectation of 1000 randomly constructed co-flowering networks using the r2dtable algorithm (“nullmodel” function; Bipartite in R; Dormann and Strauss 2014). Co-flowering modularity was standardized by calculating the Z-score of Q as: ZQ = (Q observed–Q null)/SD Q null. The Z score measures the number of standard deviations that Q of the empirical co-flowering network deviates from the average modularity based on 1000 random networks (see Albor et al. 2020). When Z values are ≥ 2 the co-flowering networks are considered significantly modular (Dormann and Strauss 2014; Dormann 2020).
Floral trait similarity
To calculate the similarity of floral traits we used the following characters that have been associated with pollinator attraction or level of specialization (see Suárez-Mariño et al. 2022): floral length (distance between the calyx and the tip of the corolla), corolla diameter (corolla width), corolla tube opening (internal diameter of the corolla), and flower color (Faegri and Van der Pijl 1979; Caruso 2000; Spaethe et al. 2001; Hirota et al. 2012; Zhao et al. 2016). Morphological characters were measured with a caliper (± 0.1 mm) on 1–5 flowers per plant on at least five plants per species. To estimate the color of the flowers, the flower reflectance spectrum was measured (300–700 nm) from the dominant corolla color in 1–3 flowers per species, with a spectrophotometer (StellarNet INC) and a Tungsten Halogen lamp as an artificial light source (see Albor et al. 2020). With this data we estimated flower color using the hexagonal color vision model which considers the chromatic coordinates (x and y) of the Hymenopteran vision model, based on Apis mellifera (Chittka 1992; Chittka and Raine 2006; and see Albor et al. 2020). We used this vision model because Hymenoptera are reported to be the most abundant floral visitors in these communities (Campos-Navarrete et al. 2013; Albor et al. 2019, 2020; Parra-Tabla et al. 2019; Suárez-Mariño et al. 2022).
To test the degree of floral trait similarity between pairs of species within- and among the co-flowering modules detected (see results), a trait matrix was constructed using the average value of each trait (i.e., flower size, total corolla diameter, corolla tube opening, and color) for each species. Then, trait distances between species pairs and the average similarity of species floral traits (for all traits) were calculated using Gower's pairwise distance (Albor et al. 2020). Gower's distance was used because it is appropriate when descriptors are not dimensionally homogeneous (Gower 1971). The Gower distance index (1–average dissimilarity) is constrained between values of 0 and 1, where values close to 1 indicate high similarity and values close to 0 indicate low similarity.
Pollinator sharing
We calculated the degree of pollinator sharing between pairs of species in each co-flowering module using the Pianka overlap index (Pianka 1973): \({O}_{jk}=\left(\sum {P}_{ij}{P}_{ik}\right)/\sqrt{\left(\sum {P}_{ij}2/{P}_{ik}2\right)},\) where Ojk represents the sharing of pollinators between plant species j and k; and Pij and Pik represent the number of floral visits made by pollinator i to species j and k, respectively. The Pianka index has been used to estimate pollinator sharing, because it considers the identity of the different pollinators, as well as their relative frequency of visits (Muñoz and Cavieres 2008; Suárez-Mariño et al. 2022). This index is bounded between values of 0 (low pollinator sharing) and 1 (high pollinator sharing). We calculated pollinator sharing between pairs of species within-co-flowering module and among-co-flowering module in both communities averaging over each unique species pair (see Albor et al. 2019).
Pollinator visits and plant–pollinator subnetworks
To estimate the frequency of plant–pollinator interactions, we monitored insect visits covering the flowering season of both communities during 2019 (August to December). Observations were carried out twice per month (10 days in total per community), in ten 20 m2 (10 × 2 m) plots parallel to the coastline and 5 m apart. In each visit, two rounds of observation were conducted between 8:30 and 10:30 AM, observing each plot for twenty minutes for a total of 200 min per day. Previous studies in these communities have shown that the higher activity of pollinating insects occurs during this period (Campos-Navarrete et al. 2013; Albor et al. 2019; Parra-Tabla et al. 2019). We recorded the activity and identity of the floral visitors considering a visit to be legitimate when there was contact between the insect and the reproductive structures of the flowers. The identification of pollinators was defined at the species or morphospecies level with the support of field identification guides (Campos-Navarrete et al. 2013; Parra-Tabla et al. 2019).
With the floral visit data, we constructed plant–pollinator subnetworks for each co-flowering module (see results) following the methodology described by Bascompte and Jordano (2007). In short, we constructed an interaction frequency matrix for each co-flowering module using the number of times every floral visitor was observed visiting flowers of a particular plant species. The interaction matrixes were used to estimate the following network metrics for each co-flowering module using the ‘bipartite’ package in R (Dormann and Strauss 2014; Oksanen et al. 2015): (a) nestedness (N): specialist species interacting with subsets of species interacting with generalists. N ranges from 0 to 1, indicating a completely random distribution of interactions (0) or a perfect nestedness (1); (b) modularity (Q): estimates the degree to which the network is organized into groups or modules of plant and pollinator species that interact more within their module than between modules, Q ranges from 0 (the network does not have more links within modules than expected by chance) to a maximum value of 1 (all links are distributed within modules) and (c) network specialization index (H2′): describes the degree of specialization among plants and pollinators across an entire network. H2′ ranges between 0 and 1, indicating extreme generalization and specialization, respectively. The metrics used for the plant–pollinator subnetworks were estimated with the “bipartite” package in R (Dormann et al. 2009), with the exception of the co-flowering module four of the scrubland community because of the low number of plant species (see results).
To test the statistical significance of modularity (Q), nestedness (N) and specialization (H2′) for each plant–pollinator subnetwork, we estimate the significance level of each metric using a null model analysis, where we compare the nestedness, modularity and specialization observed in the subnetworks against the expectation of 1000 randomly constructed networks using the r2dtable algorithm (Bipartite “nullmodel” function in R; Dormann and Strauss 2014). The three metrics were standardized by calculating the Z-score of Q, N and H2′ as: Z Q/N/H2´ = (Q/N/H2′ observed—Q/N/ H2′ null)/SD Q/N/ H2´ null, respectively. The Z score measures the number of standard deviations that Q, N and H2′ of the empirical network deviate from the average modularity, nestedness and specialization based on 1000 random networks. When Z values are ≥ 2 the subnetworks are considered significantly modular, nestedness, or specialized (Dormann and Strauss 2014; Dormann 2020).
Finally, to define the role of each pollinator species within the subnetworks, we used the categories suggested by Olesen et al. (2007). These categories assign a topological role to each species in the subnetwork based on the values of z and c, which estimate interactions within modules and interactions between modules, respectively. Thus, we classified each species as: (a) peripheral species (i.e., generally interacting species within their own module) (z ≤ 2.5 and c ≤ 0.62); (b) module hub (i.e., highly connected species within their own module) (z > 2.5 and c ≤ 0. 62); (c) connector species (i.e., species that link modules) (z ≤ 2.5 and c > 0.62); and (d) network hub (z > 2.5 and c > 0.62) (i.e., species that maintain connection not only to their own module but also to other modules) (Olesen et al. 2007; Donatti et al. 2011). Values of c and z for each subnetwork were estimated with the czvalues function of the “bipartite” package in R (Dormann et al. 2009).
Statistical analysis
To test the effect of co-flowering module, interaction type (i.e., “within-co-flowering module" vs. “among-co-flowering module”) and the effect of their interaction (co-flowering module × interaction type) on floral trait similarity (log-transformed) and pollinator sharing, we applied linear models (LMs). For all models we used a normal error distribution and the link function “identity”. Residuals for models were normally distributed (Shapiro–Wilks test, p > 0.05). Post hoc tests (Tukey HSD) were used for multiple comparisons when LMs revealed significant differences. Finally, we performed a permutational multivariate analysis of variance (PERMANOVA with 999 permutations) to test differences in the composition of floral visitors between co-flowering modules in both communities (Anderson 2001). The analyses were performed with the “lme4” package, the post hoc tests were performed with the emmeans function in the “EMMEANS” package (Lenth et al. 2019) and PERMANOVA were conducted using the adonis function in the “vegan” package in R v4.1.2 (R Core Team 2022).
Results
Co-flowering networks
A total of 74,965 flowers of 40 plant species were recorded during the entire flowering season (dune: 28 species and 29,065 flowers; scrubland: 35 species and 45,900 flowers). The analysis of co-flowering networks showed a significant modular flowering structure in both communities (Z = 86.2, p < 0.05 and Z = 143.9, p < 0.05; dune and scrubland, respectively). Three co-flowering modules were identified in the dune community (Fig. 1a), and four in the scrubland (Fig. 2a). The maximum number of plant species observed in a co-flowering module was 13 (Fig. 1a) and the minimum was three in the dune and scrubland community (Fig. 2a) respectively. However, metrics for co-flowering module four in the scrubland community, were not estimated because of the low number of plant species.
Co-flowering network and plant–pollinator subnetworks for the dune community. In co-flowering network (a), plant species with high phenological overlap (i.e., species that interact more strongly with each other than with other species) are shown within the same co-flowering module and with the same node color. Node size reflects the number of co-flowering interactions. The thickness of the lines connecting the nodes (i.e., links) reflects the magnitude of the phenological overlap (Schoener's index value). Plant–pollinator subnetworks (b), nodes at the bottom part represent plant species and nodes at the top insect species. The thickness of the lines connecting the nodes (i.e., links) represent of interactions degree between plants and pollinators. See Table S1 and Table S2 for a complete list of plant and floral visitors and their codes
Co-flowering network and plant–pollinator subnetworks for the scrubland community. In co-flowering network (a), plant species with high phenological overlap (i.e., species that interact more strongly with each other than with other species) are shown within the same co-flowering module and with the same node color. Node size reflects the number of co-flowering interactions. The thickness of the lines connecting the nodes (i.e., links) reflects the magnitude of the phenological overlap (Schoener's index value). Plant–pollinator subnetworks (b), nodes at the bottom part represent plant species and nodes at the top insect species. The thickness of the lines connecting the nodes (i.e., links) represent of interactions degree between plants and pollinators. The plant–pollinator subnetwork for co-flowering module four (yellow), was not estimated because of the low number of plant species. The size of the co-flowering networks and plant–pollinator subnetworks were not the same considering that not all plant species were visited (See Table S1 and Table S2 for a complete list of plant and floral visitors and their codes)
Floral traits similarity
Floral trait similarity was high in both communities (dune: 0.74 ± 0.15, scrubland: 0.73 ± 0.15, mean ± SE). The statistical analysis showed a similar pattern in both communities where significant differences in floral trait similarity due to co-flowering module and interaction type (i.e., within and among co-flowering modules) were observed (Table 1). In the sand dune, the co-flowering module one showed a significantly lower floral trait similarity than co-flowering modules two and three (t ≥ −2.72, p ≤ 0.01), which did not differ from each other (t = 0.065, p = 0.78). In the scrubland, the co-flowering module three showed a significantly lower floral trait similarity than co-flowering modules one and two (t ≥ 2.44, p ≤ 0.03), which did not differ from each other (t = 1.09, p = 0.51). In addition, in both communities, a significant effect of co-flowering module × interaction type interaction was observed (Table 1). In all cases, floral trait similarity was higher among co-flowering modules than within co-flowering modules (Fig. 3).
Mean (± SD; untransformed data) of floral trait similarity within and among co-flowering modules for the dune community (a) and coastal scrubland (b). Floral trait similarity between pairs of plant species was calculated using Gower's pairwise distance (1–dissimilarity). Different letters indicate significant differences (p < 0.05) of floral trait similarity between each co-flowering module through Tukey's post-hoc comparisons
Pollinator sharing
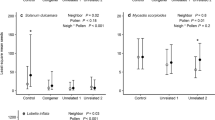
The analysis of pollinator sharing showed a contrasting pattern between communities (Table 1). While in the dune community only a marginal effect of co-flowering module was observed (Table 1; Fig. 4a), in the scrubland, significant effects of co-flowering module, interaction type, and the interaction co-flowering module × interaction type were observed (Table 1). In this latter community, the co-flowering module two showed significantly lower pollinator sharing than co-flowering modules one and three (t ≥ −3.03, p ≤ 0.01; Fig. 4b), which did not differ from each other (t = 1.61, p = 0.24). This difference seem to be driven by the very low pollinator sharing of some particular species of this co-floral module such as Gossypium hirsutum and Malvaviscus arboreus, Fig. 2b; code Gohi y Maar), that shared less than 2% of pollinators. Although on average pollinator sharing was higher within than among co-flowering modules (Fig. 4b), the interaction co-flowering module × interaction type, showed that pollinator sharing was significantly higher within than among co-flowering modules in the co-flowering modules one and three, but in the co-flowering module two pollinator sharing was higher among than within the co-flowering module (Fig. 4b). The statistical analysis also showed that in the scrubland community an increase in floral trait similarity increases pollinator sharing significantly (Table 1; β = 0.55 ± 0.11, p < 0.01).
Mean (± SD) pollinator sharing within and among co-flowering modules for the dune community (a) and coastal scrubland (b). Pollinator sharing between pairs of plant species was calculated using Pianka overlap index. Different letters indicate significant differences (p < 0.05) of pollinator sharing between each co-flowering module through Tukey's post hoc comparisons
Pollinator visits and plant–pollinator subnetworks
A total of 4,302 plant–pollinator interactions were recorded in the dune community and 3,398 in the scrubland community. The number of plant species visited varied among co-flowering modules (mean ± SD; dune 7 ± 4 and scrubland 8 ± 1.7; Table S2). The higher number of visits in both communities was from Hymenoptera (70.08%), followed by Lepidoptera (17.25%) and Diptera (12.65%) (see Table S2). Although the richness of pollinator visitors varied among co-flowering modules (dune: 23.6 ± 15.8 and scrubland: 26.6 ± 11.5; Table S2), no significant changes in floral visitor composition were observed among co-flowering modules in both communities (PERMANOVA: F1,2 = 0.60, p > 0.05 and F1,2 = 4.1, p > 0.05, dune and scrubland respectively). However, plant–pollinator subnetworks showed changes in the frequency of pollinator interactions in both communities (Fig. 1b, 2b). For example, Apis mellifera (Apidae), the species with the higher abundance in both communities, showed differences in the frequency of visits between co-flowering modules (Fig. 1b, 2b; figure code Am). Moreover, some pollinator species were not present in all co-flowering modules (e.g., Xylocopa mexicanorum Apidae in the dune or Condylostylus longicornis Diptera, in the scrubland, see Fig. 1b, 2b; figure codes Xm and Condy, respectively), or showed a very low frequency in some co-flowering modules but a higher frequency in others (e.g., Ceratina capitosa Apidae, see Fig. 2b; figure code Cera).
The plant–pollinator subnetworks of each co-flowering module showed changes in nestedness in both communities (Table 2; Fig. 1b, 2b). While in the dune community two of the three co-flowering modules showed no significant nestedness (co-flowering module 1 and 3, Table 2), in the scrubland only one of the co-flowering modules did not show significant nestedness (co-flowering module 2; Table 2). In contrast, in both communities the modular structure of plant–pollinator subnetworks were significant across all the co-flowering modules (Table 2). Likewise, the specialization values (H2´) were significant in all co-flowering modules in both communities (Table 2).
Finally, the analysis of the role of pollinator species showed that in both communities, plant–pollinator subnetworks were mainly composed by peripheral species across co-flowering modules (≥ 85% and ≥ 78.9%, sand dune and scrubland community respectively; see Fig. S2 a, b). Additionally, while “module hub” species were slightly less represented in the dune than in the scrubland across all co-flowering modules (< 15% and < 21%, dune and scrubland respectively), the percentage of network hub and connector species was very low in the scrubland. Specifically, only Geron sp. was identified as network hub in the subnetwork 1 and Apis mellifera as connector in the subnetwork 2 (see Fig. S2 b; code D3 and Am, respectively).
Discussion
Overall, our results stress the importance of considering the consequences of the temporal organization of flowering within communities to better understand the subjacent mechanisms mediating the structure of plant–pollinator networks. The analysis of co-flowering networks showed that in the studied communities the organization of flowering phenology was not random. On the contrary, it showed the existence of a modular organization based on the distribution of the frequency and intensity of flowering overlap among species (i.e., co-flowering modules). In these co-flowering modules, we expected that plant–plant interactions via pollinator sharing, would be “regulated” by the level of floral trait similarity (Moeller 2004; Ghazoul 2006; Sargent and Ackerly 2008; Arceo-Gómez et al. 2018; Albor et al. 2020) with consequences for the structure of plant–pollinator subnetworks (Junker et al. 2013; Chamberlain et al. 2014; Maruyama et al. 2014; Suárez-Mariño et al. 2022). Specifically, we expected that greater floral trait similarity within co-flowering modules would result in greater pollinator sharing and consequently in a higher level of generalization in plant–pollinator subnetworks. However, our analysis showed first, that floral trait similarity was always significantly lower within co-flowering modules when compared to between co-flowering modules, although this was dependent on the identity of each module (co-flowering module × interaction type). Second, that floral trait similarity was not reflected in a clear pattern of pollinator sharing. Furthermore, although these results suggested that lower floral trait similarity would promote a decrease in pollinator sharing, the results showed that in the sand dune we did not observe significant differences, and in the scrubland the effect was dependent on the identity of each co-flowering module.
However, although the patterns of floral trait similarity and pollinator sharing apparently were not consistent in both communities, the results may help to explain the changes in the structure of plant–pollinator subnetworks. In the scrubland we observed that in the co-flowering modules where pollinator sharing was significantly higher (co-flowering modules one and three), the nestedness and the modular structure of plant–pollinator subnetworks were significant, and in the co-flowering module where pollinator sharing was the lowest observed (< 40%; see Fig. 3), the corresponding subnetwork showed no significant nestedness structure, but significant modularity. In contrast, in the sand dune the subnetworks of two co-flowering modules did not show a significant nested structure (co-flowering modules one and three). Moreover, in these co-flowering modules, pollinator sharing was significantly lower than in the co-flowering module that did show significant nestedness. These results suggest that the high floral trait similarity observed in both communities, accompanied by variation in pollinator sharing across flowering phenology, could shape the structure of plant–pollinator subnetworks although these effects appear to be community-dependent.
Different studies have pointed out the importance of considering not only flowering phenology, or the phenological coincidence between plants and pollinators, but also other factors such as floral and morphological traits of pollinators to better understand the mechanisms involved in the structuring and spatial and temporal variation of plant–pollinator networks (e.g., Kaiser-Bunbury et al. 2010; Bergamo et al. 2017; Valdovinos 2019; Lázaro et al. 2020; Peralta et al. 2020; Suárez-Mariño et al. 2022). In this work we proposed the existence of a “controlling” effect of floral trait similarity on pollinator sharing with consequences for the structuring of plant–pollinator subnetworks emerging from the co-flowering modules. However, our results suggest that such an effect may be minor or variable across flowering phenology in co-flowering communities. It is likely that the high level of floral trait similarity observed in our coastal communities (74 ± 0.15% and 73% ± 0.15%; sand dune and scrubland, respectively) is limiting the discrimination capacity of floral visitors, eliminating or attenuating their effect on pollinator sharing as has been suggested in these and other communities (Gibson et al. 2012; Parra-Tabla et al. 2019; Albor et al. 2020; Suárez-Mariño et al. 2022). However, it is also possible that although we used floral traits that have been widely described as relevant for pollinator attraction (e.g., Caruso 2000; Spaethe et al. 2001; Hirota et al. 2012; Zhao et al. 2016), we may have omitted other important floral traits. For example, it has been documented that traits such as the quantity or quality of nectar or floral scents can determine not only the identity of pollinators but even explain the level of generalization/specialization of plant–pollinator networks (e.g., Knudsen y Tollsten 1993; Ornelas et al. 2007; Junker et al. 2010; Prieto-Benítez et al. 2016; Kantsa et al. 2018; Burkle and Runyon 2019). For example, Burkle and Runyon (2019) found that floral volatile organic compounds (VOCs) influenced strongly the level of generalized plant–pollinator interactions, attracting more pollinators and contributing importantly to the nested structure of plant–pollinator network. In our communities, the high proportion of peripheral species suggests the existence of other floral traits that may be helping to maintain an important level of specialization.
On the other hand, it is also likely that the effect of floral traits depends on the identity of pollinators across the co-flowering modules (Biella et al. 2017; CaraDonna et al. 2017). However, in both communities the analysis of pollinator species composition between co-flowering modules showed no significant differences, suggesting that variation on pollinator sharing depends on plant species composition in each co-flowering module. Thus, the effect of co-flowering species composition could also have driven interaction rewiring between plants and pollinators (Olesen et al. 2011; Campos-Navarrete et al. 2013; Cuartas-Hernández y Medel 2015; Biella et al. 2017). Interaction rewiring is relevant not only because it shapes the structure of plant–pollinator networks but also because it defines the role played by each pollinator species in the networks (e.g., Campos-Navarrete et al. 2013; Watts et al. 2016, Biella et al. 2017, CaraDonna et al. 2017; CaraDonna y Waser 2020).
Plant–pollinator subnetworks constructed for each co-flowering module in both communities showed a significant level of specialization throughout the flowering season. This specialization was revealed by the specialization (H2′) and modularity (Q) estimators, which corresponded to the high proportion of specialized (peripheral) species detected in the subnetworks by the analyses of the role of pollinators (Olesen et al. 2011; and see Jacquemin et al. 2020; Hinton and Peters 2021). Specifically, these analyses showed that in both communities ca. 80% of all pollinators can be considered as specialists, characterized by a low number of links with plant species from the same module of the plant–pollinator subnetworks (Olesen et al. 2011). Moreover, in scrubland only Geron sp. (Diptera; figure code D3) and Apis mellifera (Apidae; figure code Am) were identified as super generalist and highly connected species (i.e., “network hub” and “connector” species; see Fig. S2 a, b, respectively). The role of the pollinators within the networks is defined both by their use of floral resources (frequency of visits and richness of pant species visited), and the way in which they distribute their visits among the plant–pollinator modules within the plant–pollinator networks. Thus, while Geron sp. was identified as a super generalist because it visited all plant species in the subnetwork of the co-flowering module 1, A. mellifera was identified as a connector because it participated in most of the modules (5/7) of the subnetwork 2. Both species have been reported as frequent species in coastal co-flowering communities in the Yucatan, and in the case of A. mellifera has been typically reported as the species with the highest number of pollinator interactions (Campos-Navarrete et al. 2013; Parra-Tabla et al. 2019; Albor et al. 2019; Suárez-Mariño et al. 2022). Interestingly, the analysis of the role of pollinators also showed that the species can modify their role across co-flowering modules, underling the importance of plant species composition within co-flowering modules in the process of plant–pollinator subnetworks rewiring, and in the definition of the role of pollinators. For example, in the sand dune A. mellifera could be identified as a highly connected or specialist species (Fig. S2 a), and in the scrubland Geron sp. was identified as highly generalist, generalist or even peripheral species (see Fig. S2 b).
Overall, our results support other studies by showing that generalization is not a persistent feature of plant–pollinator networks (e.g., Rasmussen et al. 2013; Biella et al. 2017; Wang et al. 2020; Schwarz et al. 2020). Likewise, our data supports the idea put forward by several studies that suggest that aggregation of flowering and flower visitation data, as traditionally analyzed in plant–pollinator networks (Bascompte and Jordano et al. 2007), may obscure our understanding of the temporal dynamics that exist in these networks and underestimate the importance of specialization in co-flowering communities (e.g., Petanidou et al. 2008; Rasmussen et al. 2013; Biella et al. 2017; CaraDonna et al. 2017; Sajjad et al. 2017; Schwarz et al. 2020).
However, in contrast to other studies cited, our work highlights the importance of explicitly considering the organization of flowering phenology (and see Biella et al. 2017; Arceo-Gómez et al. 2018), as well as factors such as floral trait similarity and pollinator sharing to advance our understanding of the mechanisms explaining temporal changes in the structure of plant–pollinator networks.
Data availability
The datasets generated and analysed during the current study are available from the corresponding author on reasonable request.
References
Aizen MA, Vázquez DP (2006) Flowering phenologies of hummingbird plants from the temperate forest of southern South America: is there evidence of competitive displacement? Ecography (cop) 29:357–366. https://doi.org/10.1111/j.2006.0906-7590.04552.x
Albor C, García-Franco JG, Parra-Tabla V, Díaz-Castelazo C, Arceo-Gómez G (2019) Taxonomic and functional diversity of the co-flowering community differentially affect Cakile edentula pollination at different spatial scales. J Ecol 107:2167–2181. https://doi.org/10.1111/1365-2745.13183
Albor C, Arceo-Gómez G, Parra-Tabla V (2020) Integrating floral trait and flowering time distribution patterns help reveal a more dynamic nature of co-flowering community assembly processes. J Ecol 108:2221–2231. https://doi.org/10.1111/1365-2745.13486
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46. https://doi.org/10.1111/j.1442-9993.2001.01070.pp.x
Arceo-Gómez G, Kaczorowski RL, Ashman TL (2018) A network approach to understanding patterns of coflowering in diverse communities. Int J Plant Sci 179:569–582. https://doi.org/10.1086/698712
Bascompte J, Jordano P (2007) Plant-animal mutualistic networks: the architecture of biodiversity. Annu Rev Ecol Evol Syst 38:567–593. https://doi.org/10.1146/annurev.ecolsys.38.091206.095818
Bascompte J, Jordano P, Melián CJ, Olesen JM (2003) The nested assembly of plant-animal mutualistic networks. Proc Natl Acad Sci USA 100:9383–9387. https://doi.org/10.1073/pnas.1633576100
Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. Proc Int AAAI Conf Web Soc Media 3:361–362
Bergamo PJ, Wolowski M, Maruyama PK, Vizentin-Bugoni J, Carvalheiro LG, Sazima M (2017) The potential indirect effects among plants via shared hummingbird pollinators are structured by phenotypic similarity. Ecology 98:1849–1858. https://doi.org/10.1002/ecy.1859
Biella P, Ollerton J, Barcella M, Assini S (2017) Network analysis of phenological units to detect important species in plant-pollinator assemblages: can it inform conservation strategies? Community Ecol 18:1–10. https://doi.org/10.1556/168.2017.18.1.1
Burkle LA, Runyon JB (2019) Floral volatiles structure plant–pollinator interactions in a diverse community across the growing season. Funct Ecol 33:2116–2129. https://doi.org/10.1111/1365-2435.13424
Campos-Navarrete MJ, Parra-Tabla V, Ramos-Zapata J, Díaz-Castelazo C, Reyes-Novelo E (2013) Structure of plant-Hymenoptera networks in two coastal shrub sites in Mexico. Arthropod Plant Interact 7:607–617. https://doi.org/10.1007/s11829-013-9280-1
CaraDonna PJ, Waser NM (2020) Temporal flexibility in the structure of plant–pollinator interaction networks. Oikos 129:1369–1380. https://doi.org/10.1111/oik.07526
CaraDonna PJ, Petry WK, Brennan RM, Cunningham JL, Bronstein JL, Waser NM, Sanders NJ (2017) Interaction rewiring and the rapid turnover of plant–pollinator networks. Ecol Lett 20:385–394. https://doi.org/10.1111/ele.12740
CaraDonna PJ, Burkle LA, Schwarz B, Resasco J, Knight TM, Benadi G, Blüthgen N, Dormann CF, Fang Q, Fründ J, Gauzens B, Kaiser-Bunbury CN, Winfree R, Vázquez DP (2021) Seeing through the static: the temporal dimension of plant–animal mutualistic interactions. Ecol Lett 24:149–161. https://doi.org/10.1111/ele.13623
Caruso CM (2000) Competition for pollination influences selection on floral traits of Ipomopsis aggregata. Evolution (NY) 54:1546–1557. https://doi.org/10.1111/j.0014-3820.2000.tb00700.x
Chamberlain SA, Cartar RV, Worley AC, Semmler SJ, Gielens G, Elwell S, Evans ME, Vamosi JC, Elle E (2014) Traits and phylogenetic history contribute to network structure across Canadian plant–pollinator communities. Oecologia 176:545–556. https://doi.org/10.1007/s00442-014-3035-2
Chittka L (1992) The colour hexagon: a chromaticity diagram based on photoreceptor excitations as a generalized representation of colour opponency. J Comp Physiol A 170:533–543. https://doi.org/10.1007/BF00199331
Chittka L, Raine NE (2006) Recognition of flowers by pollinators. Curr Opin Plant Biol 9:428–435. https://doi.org/10.1016/j.pbi.2006.05.002
Coux C, Rader R, Bartomeus I, Tylianakis JM (2016) Linking species functional roles to their network roles. Ecol Lett 19:762–770. https://doi.org/10.1111/ele.12612
Cuartas-Hernández S, Medel R (2015) Topology of plant—flower-visitor networks in a tropical mountain forest: insights on the role of altitudinal and temporal variation. PLoS ONE 10:e0141804. https://doi.org/10.1371/journal.pone.0141804
Donatti CI, Guimarães PR, Galetti M, Pizo MA, Marquitti FMD, Dirzo R (2011) Analysis of a hyper-diverse seed dispersal network: modularity and underlying mechanisms. Ecol Lett 14:773–781. https://doi.org/10.1111/j.1461-0248.2011.01639.x
Dormann CF (2020) Using bipartite to describe and plot two-mode networks in R. R Package Version 4:1–28
Dormann CF, Strauss R (2014) A method for detecting modules in quantitative bipartite networks. Methods Ecol Evol 5:90–98. https://doi.org/10.1111/2041-210X.12139
Dormann CF, Frund J, Bluthgen N, Gruber B (2009) Indices, graphs and null models: analyzing bipartite ecological networks. Open Ecol J 2:7–24. https://doi.org/10.2174/1874213000902010007
Elzinga JA, Atlan A, Biere A, Gigord L, Weis AE, Bernasconi G (2007) Time after time: flowering phenology and biotic interactions. Trends Ecol Evol 22:432–439. https://doi.org/10.1016/j.tree.2007.05.006
Emer C, Vaughan IP, Hiscock S, Memmott J (2015) The impact of the invasive alien plant, Impatiens glandulifera, on pollen transfer networks. PLoS ONE 10:e0143532. https://doi.org/10.1371/journal.pone.0143532
Espejel I (1987) A Phytogeographical analysis of coastal vegetation in the yucatan peninsula. J Biogeogr 14:499–519. https://doi.org/10.2307/2844877
Faegri K, Van der Pijl L (1979) The principles of Pollination Ecology. Pergamon Press, Oxford
Forrest J, Inouye DW, Thomson JD (2010) Flowering phenology in subalpine meadows: does climate variation influence community co-flowering patterns? Ecology 91:431–440. https://doi.org/10.1890/09-0099.1
Ghazoul J (2006) Floral diversity and the facilitation of pollination. J Ecol 94:295–304. https://doi.org/10.1111/j.1365-2745.2006.01098.x
Gibson MR, Richardson DM, Pauw A (2012) Can floral traits predict an invasive plant’s impact on native plant-pollinator communities? J Ecol 100:1216–1223. https://doi.org/10.1111/j.1365-2745.2012.02004.x
Gower JC (1971) A general coefficient of similarity and some of its properties. Biometrics 27:857–871. https://doi.org/10.2307/2528823
Guzman LM, Chamberlain SA, Elle E (2021) Network robustness and structure depend on the phenological characteristics of plants and pollinators. Ecol Evol 11:13321–13334. https://doi.org/10.1002/ece3.8055
Heleno R, Garcia C, Jordano P, Traveset A, Gómez JM, Blüthgen N, Memmott J, Moora M, Cerdeira J, Rodríguez-Echeverría S, Freitas H, Olesen JM (2014) Ecological networks: delving into the architecture of biodiversity. Biol Lett. https://doi.org/10.1098/rsbl.2013.1000
Hinton CR, Peters VE (2021) Plant species with the trait of continuous flowering do not hold core roles in a Neotropical lowland plant-pollinating insect network. Ecol Evol 11:2346–2359. https://doi.org/10.1002/ece3.7203
Hirota SK, Nitta K, Kim Y, Kato A, Kawakubo N, Yasumoto AA, Yahara T (2012) Relative role of flower color and scent on pollinator attraction: Experimental tests using F1 and F2 hybrids of daylily and nightlily. PLoS ONE 7:e39010. https://doi.org/10.1371/journal.pone.0039010
Jacquemin F, Violle C, Munoz F, Mahy G, Rasmont P, Roberts SPM, Vray S, Dufrêne M (2020) Loss of pollinator specialization revealed by historical opportunistic data: Insights from network-based analysis. PLoS ONE 15:e0235890. https://doi.org/10.1371/journal.pone.0235890
Junker RR, Höcherl N, Blüthgen N (2010) Responses to olfactory signals reflect network structure of flower-visitor interactions. J Anim Ecol 79:818–823. https://doi.org/10.1111/j.1365-2656.2010.01698.x
Junker RR, Blüthgen N, Brehm T, Binkenstein J, Paulus J, Martin Schaefer H, Stang M (2013) Specialization on traits as basis for the niche-breadth of flower visitors and as structuring mechanism of ecological networks. Funct Ecol 27:329–341. https://doi.org/10.1111/1365-2435.12005
Kaiser-Bunbury CN, Muff S, Memmott J, Müller CB, Caflisch A (2010) The robustness of pollination networks to the loss of species and interactions: a quantitative approach incorporating pollinator behaviour. Ecol Lett 13:442–452. https://doi.org/10.1111/j.1461-0248.2009.01437.x
Kantsa A, Raguso RA, Dyer AG, Olesen JM, Tscheulin T, Petanidou T (2018) Disentangling the role of floral sensory stimuli in pollination networks. Nat Commun 9:1041. https://doi.org/10.1038/s41467-018-03448-w
Knudsen JT, Tollsten L (1993) Trends in floral scent chemistry in pollination syndromes: floral scent composition in moth-pollinated taxa. Bot J Linn Soc 113:263–284. https://doi.org/10.1111/j.1095-8339.1993.tb00340.x
Lázaro A, Gómez-Martínez C, Alomar D, González-Estévez MA, Traveset A (2020) Linking species-level network metrics to flower traits and plant fitness. J Ecol 108:1287–1298. https://doi.org/10.1111/1365-2745.13334
Lenth R, Singmann H, Love J, Buerkner P, Herve M (2019) Estimated marginal means, aka least-squares means. Package ‘emmeans’. https://cran.r-project.org/web/packages/emmeans/emmeans.pdf
Maruyama PK, Vizentin-Bugoni J, Oliveira GM, Oliveira PE, Dalsgaard B (2014) Morphological and spatio-temporal mismatches shape a neotropical savanna plant-hummingbird network. Biotropica 46:740–747. https://doi.org/10.1111/btp.12170
Moeller DA (2004) Facilitative interactions among plants via shared pollinators. Ecology 85:3289–3301. https://doi.org/10.1890/03-0810
Muñoz AA, Cavieres LA (2008) The presence of a showy invasive plant disrupts pollinator service and reproductive output in native alpine species only at high densities. J Ecol 96:459–467. https://doi.org/10.1111/j.1365-2745.2008.01361.x
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2015) Vegan: community Ecology Package. R Packag version 2.6–2. https://CRAN.R-project.org/package=vegan
Olesen JM, Eskildsen LI, Venkatasamy S (2002) Invasion of pollination networks on oceanic islands: Importance of invader complexes and endemic super generalists. Divers Distrib 8:181–192. https://doi.org/10.1046/j.1472-4642.2002.00148.x
Olesen JM, Bascompte J, Dupont YL, Jordano P (2007) The modularity of pollination networks. Proc Natl Acad Sci USA 104:19891–19896. https://doi.org/10.1073/pnas.0706375104
Olesen JM, Stefanescu C, Traveset A (2011) Strong, long-term temporal dynamics of an ecological network. PLoS ONE 6:e26455. https://doi.org/10.1371/journal.pone.0026455
Olito C, Fox JW (2015) Species traits and abundances predict metrics of plant–pollinator network structure, but not pairwise interactions. Oikos 124:428–436. https://doi.org/10.1111/oik.01439
Orellana R, Espadas C, Conde C, Gay C (2009) Atlas escenarios de cambio climático en la Península de Yucatán. Cent Investig Científica Yucatán, Mérida
Ornelas JF, Ordano M, De-Nova AJ, Quintero ME, Garland JRT (2007) Phylogenetic analysis of interspecific variation in nectar of hummingbird-visited plants. J Evol Biol 20:1904–1917. https://doi.org/10.1111/j.1420-9101.2007.01374.x
Parra-Tabla V, Albor-Pinto C, Tun-Garrido J, Angulo-Pérez D, Barajas C, Silveira R, Ortíz-Díaz JJ, Arceo-Gómez G (2018) Spatial patterns of species diversity in sand dune plant communities in Yucatan, Mexico: importance of invasive species for species dominance patterns. Plant Ecol Divers 11:157–172. https://doi.org/10.1080/17550874.2018.1455232
Parra-Tabla V, Angulo-Pérez D, Albor C, Campos-Navarrete MJ, Tun-Garrido J, Sosenski P, Alonso C, Ashman TL, Arceo-Gómez G (2019) The role of alien species on plant-floral visitor network structure in invaded communities. PLoS ONE 14:e0218227. https://doi.org/10.1371/journal.pone.0218227
Parra-Tabla V, Alonso C, Ashman T, Raguso RA, Albor C, Sosenski P, Carmona D, Arceo-Gómez G (2021) Pollen transfer networks reveal alien species as main heterospecific pollen donors with fitness consequences for natives. J Ecol 109:939–951. https://doi.org/10.1111/1365-2745.13520
Peralta G, Vázquez DP, Chacoff NP, Lomáscolo SB, Perry GLW, Tylianakis JM (2020) Trait matching and phenological overlap increase the spatio-temporal stability and functionality of plant–pollinator interactions. Ecol Lett 23:1107–1116. https://doi.org/10.1111/ele.13510
Petanidou T, Kallimanis AS, Tzanopoulos J, Sgardelis SP, Pantis JD (2008) Long-term observation of a pollination network: fluctuation in species and interactions, relative invariance of network structure and implications for estimates of specialization. Ecol Lett 11:564–575. https://doi.org/10.1111/j.1461-0248.2008.01170.x
Pianka ER (1973) The structure of lizard communities. Annu Rev Ecol Syst 4:53–74. https://doi.org/10.1146/annurev.es.04.110173.000413
Prieto-Benítez S, Millanes AM, Dötterl S, Giménez-Benavides L (2016) Comparative analyses of flower scent in Sileneae reveal a contrasting phylogenetic signal between night and day emissions. Ecol Evol 6:7869–7881. https://doi.org/10.1002/ece3.2377
R Core Team (2022) R: A language and environment for statistical computing. 2022: version 4.1.2. R Foundation for Statistical Computing, Vienna
Ramos-Robles M, Andresen E, Díaz-Castelazo C (2016) Temporal changes in the structure of a plant-frugivore network are influenced by bird migration and fruit availability. PeerJ 4:e2048. https://doi.org/10.7717/peerj.2048
Rasmussen C, Dupont YL, Mosbacher JB, Trøjelsgaard K, Olesen JM (2013) Strong impact of temporal resolution on the structure of an ecological network. PLoS ONE 8:e81694. https://doi.org/10.1371/journal.pone.0081694
Rathcke B, Lacey EP (1985) Phenological patterns of terrestrial plants. Annu Rev Ecol Syst 16:179–214. https://doi.org/10.1146/annurev.es.16.110185.001143
Sajjad A, Saeed S, Ali M, Khan FZA, Kwon YJ, Devoto M (2017) Effect of temporal data aggregation on the perceived structure of a quantitative plant–floral visitor network. Entomol Res 47:380–387. https://doi.org/10.1111/1748-5967.12233
Sargent RD, Ackerly DD (2008) Plant-pollinator interactions and the assembly of plant communities. Trends Ecol Evol 23:123–130. https://doi.org/10.1016/j.tree.2007.11.003
Schoener TW (1970) Nonsynchronous Spatial overlap of lizards in patchy habitats. Ecology 51:408–418. https://doi.org/10.2307/1935376
Schwarz B, Vázquez DP, CaraDonna PJ, Knight TM, Benadi G, Dormann CF, Gauzens B, Motivans E, Resasco J, Blüthgen N, Burkle LA, Fang Q, Kaiser-Bunbury CN, Alarcón R, Bain JA, Chacoff NP, Huang S-Q, LeBuhn G, MacLeod M, Petanidou T, Rasmussen C, Simanonok MP, Thompson AH, Fründ J (2020) Temporal scale-dependence of plant–pollinator networks. Oikos 129:1289–1302. https://doi.org/10.1111/oik.07303
Spaethe J, Tautz J, Chittka L (2001) Visual constraints in foraging bumblebees: flower size and color affect search time and flight behavior. Proc Natl Acad Sci USA 98:3898–3903. https://doi.org/10.1073/pnas.071053098
Stang M, Klinkhamer PGL, van der Meijden E, Memmott J (2006) Size constraints and flower abundance determine the number of interactions in a plant-flower visitor web. Oikos 112:111–121. https://doi.org/10.1111/j.0030-1299.2006.14199.x
Suárez-Mariño A, Arceo-Gómez G, Albor C, Parra-Tabla V (2022) Flowering overlap and floral trait similarity help explain the structure of pollination network. J Ecol 110:1790–1801. https://doi.org/10.1111/1365-2745.13905
Trøjelsgaard K, Jordano P, Carstensen DW, Olesen JM (2015) Geographical variation in mutualistic networks: similarity, turnover and partner fidelity. Proc R Soc B Biol Sci 282:20142925. https://doi.org/10.1098/rspb.2014.2925
Tylianakis JM, Morris RJ (2017) Ecological networks across environmental gradients. Annu Rev Ecol Evol Syst 48:25–48. https://doi.org/10.1146/annurev-ecolsys-110316-022821
Valdovinos FS (2019) Mutualistic networks: moving closer to a predictive theory. Ecol Lett 22:1517–1534. https://doi.org/10.1111/ele.13279
Vázquez DP, Blüthgen N, Cagnolo L, Chacoff NP (2009) Uniting pattern and process in plant-animal mutualistic networks: a review. Ann Bot 103:1445–1457. https://doi.org/10.1093/aob/mcp057
Wang X, Zeng T, Wu M, Zhang D (2020) Seasonal dynamic variation of pollination network is associated with the number of species in flower in an oceanic island community. J Plant Ecol 13:657–666. https://doi.org/10.1093/jpe/rtaa054
Waser NM, Real LA (1979) Effective mutualism between sequentially flowering plant species. Nature 281:670–672. https://doi.org/10.1038/281670a0
Watts S, Dormann CF, Martín González AM, Ollerton J (2016) The influence of floral traits on specialization and modularity of plant-pollinator networks in a biodiversity hotspot in the Peruvian Andes. Ann Bot 118:415–429. https://doi.org/10.1093/aob/mcw114
Zhao YH, Ren ZX, Lázaro A, Wang H, Bernhardt P, Li HD, Li DZ (2016) Floral traits influence pollen vectors’ choices in higher elevation communities in the Himalaya-Hengduan Mountains. BMC Ecol 16:1–8. https://doi.org/10.1186/s12898-016-0080-1
Acknowledgements
The authors thank L. Abdala-Roberts, P. Sosenski and J. Tun made valuable comments to a previous version of this manuscript and to B. Suárez, E. Soltero, F. Torres and R. Silveira for their help during fieldwork. We would like to thank the editor and one anonymous reviewer for their comments on an earlier version of this manuscript.
Funding
This work was supported by CONACyT (248406) to V.P.-T. G.A.-G. was supported by National Science Foundation DEB (1931163). A.S.-M. was supported by a fellowship grant form CONACyT.
Author information
Authors and Affiliations
Contributions
AS-M, VP-T. and GA-G. formulated the idea and conceptualized the study; AS-M and CA. collected and analyzed the data; the manuscript was drafted by AS-M and VP-T. The final MS was edited by all the co-authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Communicated by Simon Pierce.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (PDF 436 KB)
Study site: sand dune (a) and coastal scrubland (b) communities in Telchac, Yucatan, Mexico
Supplementary file2 (PDF 281 KB)
Role of each insect species in the plant–pollinator subnetworks for dune (a) and scrubland (b). Each symbol describes the within-module degree (z) and the participation coefficient (c) of each species. Vertical and horizontal lines represent 90% quantiles of null model coefficients and delimit groups of species with different topological roles in networks. Peripheral species (bottom left), module hub (top left), network hub (top right) or connector (bottom right). See Table S2 for a complete list of floral visitors and their codes
Supplementary file3 (PDF 161 KB)
List of plant species, codes and the number of open flowers per census for the dune and scrubland community
Supplementary file4 (PDF 165 KB)
List of insect species, codes and number of visits per module for the dune and scrubland community
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Suárez-Mariño, A., Arceo-Gómez, G., Albor, C. et al. Co-flowering modularity and floral trait similarity help explain temporal changes in plant–pollinator network structure. Plant Ecol 223, 1289–1304 (2022). https://doi.org/10.1007/s11258-022-01275-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11258-022-01275-0