Abstract
Basic leucine zipper (bZIP) TF family is key regulator of diverse biological functions, including heat stress tolerance. However, its role in response to heat stress is unexplored in Sorghum bicolor, an abiotic stress-tolerant cereal plant. Bio-prospecting of genes for abiotic stress tolerance from elite natural stress-tolerant species is a promising approach for development of abiotic stress-tolerant crops. Therefore, qRT-PCR analysed for heat stress induced bZIP17 gene from S. bicolor, showed 14.7 and 17.3-fold expression at 4 h and 6 h of heat stress respectively at one-month old seedling stage. Further its cDNA sequence was cloned (named as SbbZIP17) for further functional validation in model plant system. In silico analysis showed SbbZIP17 encodes for bZIP polypeptide, an endoplasmic reticulum (ER) type II membrane-tethered transcription factor (type II MMTF), highly conserved, having a single ligand-binding site, interacting with heat stress-responsive proteins (HSP70, NF-Ys), expressed in different tissues and organs. Over-expression of SbbZIP17 in independent events of transgenic tobacco (Nicotiana tabacum) lines (T1) is responsible for activation of genes involved in unfolded protein response (UPR) pathway under heat stress. Transgenic tobacco lines showed enhancement in hydration status, antioxidant activity, reduction in chlorophyll loss, and membrane damage. Our analysis demonstrated that SbbZIP17 plays an important role in regulating heat stress tolerance in plants.
Key message
SbbZIP17 (encoding membrane-tethered transcription factor) from Sorghum bicolor overexpressed in tobacco plants showed heat stress tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rapidly changing global climate conditions result in different abiotic stresses, especially heat stress which is a major constraint for future food security. Heat stress hampers physiological and biochemical processes by negatively affecting crop growth, development, and ultimately yield (Saeed et al. 2023). Plants have evolved a variety of tolerance mechanism in response to heat stress conditions and undergo various physiological, biochemical, and molecular modifications for adaptation (Janni et al. 2020; Saeed et al. 2023). However, many of these modifications are achieved via stress signal transduction and transcriptional modulation of heat stress-responsive genes by different transcription factors (TFs) (Saeed et al. 2023). TFs families, such as HSF, bHLH, bZIP, WRKY ERF, and MYB are well known as crucial parts of transcriptional regulatory network of stress-responsive genes to mitigate heat stress conditions (Janni et al. 2020).
Basic leucine zipper (bZIP) is the most important and largest TF family; there are 75 bZIPs in (A) thaliana, 89 in P. trichocarpa, 89 in O. sativa, 92 in S. bicolor, 96 in (B) distachyon, 121 in M. acuminata, 77 in M. esculenta, and 187 in T. aestivum (Agarwal et al. 2019). Preceding studies have explained that bZIP TFs are key regulators of fundamental biological processes, such as pathogen resistance, hormone signaling, developmental processes, heat, cold, salt, and drought stress tolerance (Sornaraj et al. 2016; Banerjee et al. 2017). The bZIPs protein comprises a conserved bZIP domain having binding affinity to specific DNA sequence along with a less conserved leucine zipper domain for dimerization after binding to dsDNA (preferably to ACGT sequence). The bZIP region contains 40–80 amino acid residues and the basic region consists of sixteen amino acid residues with a nuclear localization signal (NLS) (Sornaraj et al. 2016; Banerjee et al. 2017). The bZIPs are classified as ten different types (Type-I to Type-X) of group in O. sativa (89) and A. thaliana (75) (Nijhawan et al. 2008). Among them, Type-II bZIP membrane-tethered transcription factors (MTTFs) take part in unfolded protein response (UPR) pathway also referred as endoplasmic reticulum (ER) stress response (Howell 2013). The UPR triggers when improperly assembled proteins start accumulating in the ER under abiotic stress conditions (Howell 2013). Type-II bZIP protein consists of site 1 protease (S1P) cleavage position and a transmembrane domain (Sornaraj et al. 2016). ER is the site of biosynthesis for Type-II bZIP MTTFs (Membrane-tethered transcription factor), for instance AtbZIP60, AtbZIP49, AtbZIP28, and AtbZIP17 in A. thaliana. Under normal growth conditions, MTTFs are situated in ER membrane, while during stress conditions they move to Golgi body for proteolytic cleavage accompanied by Golgi-specific proteases i.e., S1P and S2P. The cleaved part of MTTFs is transported to the nucleus and activates stress-responsive genes of UPR pathway including Calreticulin (CRT1), Calnexin (CNX), protein disulphide isomerase (PDIL), and binding protein (BiP) (Liu and Howell 2010). Recent analysis exposed that plant UPR affect cellular processes in many ways and would be one of the reasons for providing abiotic stress tolerance (Hayashi et al. 2013).
In A. thaliana, AtbZIP17 and AtbZIP24 TFs are functionally validated for their role in salt stress tolerance (Liu et al. 2008; Yang et al. 2009). Similarly, overexpression of bZIP gene from O. Sativa, T. aestivum, P. trifoliata, M. hupehensis, and C. annum improved abiotic stress tolerance ability via modulating the genes involved in UPR pathway (Huang et al. 2010; Zhang et al. 2012; Yang et al. 2019; Agarwal et al. 2019; Gai et al. 2020). To date, several genes encode TFs have been cloned from naturally stress-tolerant crop plant species and characterized for enhancement of abiotc stress tolerance (Agarwal et al. 2013; Gupta et al. 2014) but limited studies are available in S. bicolor. Sorghum (S. bicolor L. Moench) is a prominent cereal crop (fodder and grain) having outstanding ability to tolerate drought, heat, and waterlogging conditions. Therefore, it is an appropriate model crop to study the biochemical and molecular mechanism underlying abiotic stress tolerance (Mutava et al. 2011; Tari et al. 2013; Baillo et al. 2020). Even though, S. bicolor is considered as a source of genes liable for abiotic stress tolerance, just few of them (Hsp70, SbP5CS1, and SbP5CS2) are functionally characterized (Su et al. 2011; Mulaudzi-Masuku et al. 2015; Halder et al. 2016).
The genome of S. bicolor encodes 92 bZIP genes, but only a single gene has been cloned i.e., Opaque2 (Z. mays) homologous gene (Vettore et al. 1998). However functional role of bZIP gene family in terms of abiotic stress tolerance is still unexplored in S. bicolor (Wang et al. 2011). Here, we analysed expression of SbbZIP17 under heat stress and after cloning analysed using in-silico based bioinformatic approach. Furthermore, tobacco plants were over-expressed with SbbZIP17 to study its role under heat stress conditions.
Materials and methods
Plant growth conditions and heat stress for S. bicolor
Seeds of S. bicolor genotype “Swati” was surface sterilized using 70% (v/v) alcohol for 1.5 min afterwards treated with 12% sodium hypochlorite (NaOCl) solution for 15 min. Sterilized seeds were sown in a pot containing soilrite under growth chamber condition at SDMVMs College of Agricultural Biotechnology, Aurangabad, India. Conditions of growth chamber were maintained for seed germination and establishment as 16 h light/8 h dark photoperiod, temperature, and light intensity of 24 ± 2 °C and 100 µmol m− 2s− 1 respectively. One-month-old seedlings were subjected to heat stress conditions for 0 h, 2 h, 4 h, and 6 h at 42 °C inside a growth chamber in a sinusoidal manner; the temperature stress was increased from 25 °C (ambient) to 42 °C with 1 °C/5 min (Ngara et al. 2012; Mulaudzi-Masuku et al. 2015; Goswami et al. 2016). The heat stress exposed and control leaf tissues were collected in three replicates for RNA isolation, instantly exposed to liquid nitrogen before storage in deep freezer (-80 °C).
Expression analysis of SbbZIP17
Relative expression level of SbbZIP17 under heat stress conditions was analyzed using qRT-PCR. Isolation of total RNA was carried out from heat stress (2 h, 4 h, and 6 h) and control leaf samples (0 h) using MagMAX™ Plant RNA Isolation Kit (Applied Biosystems, USA). The cDNA was synthesized from total RNA (2 µg) using SuperScript™ III first-strand cDNA synthesis kit (Invitrogen, USA). Integrated DNA Technology (IDT) software (www.idtdna.com) was used to design the primers specific to genes and got synthesized from IDT (Table S1). The level of gene expression was normalized using internal reference S. bicolor GAPDH gene (Glyceraldehyde 3 phosphate dehydrogenase) (Table S1) (Baillo et al. 2020). The total reaction mixture volume of 20 µl contained 10 µl of KAPA SYBR® FAST qPCR Master Mix (KAPA Biosystems, USA), 2 µl of cDNA (200 ng), 0.5 µl each of forward and reverse primers (400 nM of each primer). Nuclease-free water was added to make up the total volume of 20 µl. The qRT-PCR reactions were carried out in biological triplicates for each heat stress (2 h, 4 h, and 6 h) and control leaf samples, and technical triplicates were taken for each biological replicate (Panzade et al. 2021, 2022). Steps of PCR cycles were as follows: 94 °C for 4 min then 32 cycles of 94 °C for 16 s, and 60 °C for 16 s, and 72 °C for 22 s. The 2−ΔΔCt equation was used to evaluate the relative expression level (Livak and Schmittgen 2008). Leaf samples without exposed to heat stress were used as control. The primers and template-specific binding were analyzed by generating the melting curve. Cp values were noted down for each sample to analyze the fold change in gene expression.
Cloning and sequence analysis of SbbZIP17
To amplify the CDS (Coding DNA Sequence) of gene SbbZIP17, primers were designed according to upstream and downstream sites of S. bicolor SbbZIP39 from NCBI (Accession No. LOC8085134). For directional cloning of gene SbbZIP17 in binary vector, restriction sites NdeI and SalI were incorporated in forward and reverse primers respectively (Table S1). The PCR cycles were as follows: 94 °C for 2.5 min, 32 cycles of 94 °C for 55s, 61 °C for 35s, 72 °C for 120s. Proofreading Ex Taq DNA polymerase (Takara, Japan) was used to ensure zero error while amplification. The amplified PCR product was analyzed on agarose gel (1.2%) and then eluted from the agarose gel using GenElute™ Gel Extraction Kit (Sigma, USA). The amplicon of gene SbbZIP17 was ligated to pGEM®-T Easy vector (Promega, USA). The ligation was performed at 4 °C for 16 h and 10 µl of ligated reaction mix was transformed to E. coli strain DH5α, the transformed cells were selected for blue/white screening. The PCR positive clones were picked up for sequencing. The sequences were aligned to obtain entire CDS (Coding DNA Sequence) and submitted to NCBI database.
In silico analysis of SbbZIP17
The SbbZIP17 sequence homology analysis was carried out by the BLAST tool available at National Centre for Biotechnology Information (NCBI). The conserved domains in SbbZIP17 were identified by SMART tool (https://smart.embl-heidelberg.de/). CELLO2GO server (http://cello.life.nctu.edu.tw/cello2go/) was used for the prediction of protein at a subcellular level. The phylogenetic analysis was performed with homologous genes of SbbZIP17 from several species using MEGA 7.0 software tool (https://www.megasoftware.net/). The physio-biochemical features of protein SbbZIP17 were assessed by Protparam tool (https://web.expasy.org/protparam/). MEME (Multiple Em for Motif Elicitation) server (https://meme-suite.org/meme/tools/meme) was used for the identification of conserved motif domains. YinOYang 1.2 software and NetNGlyc 1.0 software was used to analyse O-linked and N-linked glycosylation in SbbZIP. The secondary structure was predicted by PSIPRED tool. For protein 3D structure and ligand-binding site prediction Phyre2 and 3D Ligand binding site software was used respectively. Protein to protein interaction of SbbZIP with closely related proteins was analyzed using the STRING (https://string-db.org/) online tool. Transcript (SbbZIP1) abundance was analyzed with the help of Genevestigator tool (https://genevestigator.com/) using the publicly available mRNA Seq S. bicolor database (mRNA Seq Gene level S. bicolor (ref: Sbicolor v3.1.1) at default parameters. The probeset (Sobic.009G137100) encoding SbbZIP17 was used for the analysis of spatio-temporal expression level of SbbZIP17 at various developmental phases and in 32 anatomical organs of S. bicolor. Further, we analyzed the similar and correlated genes to SbbZIP17 using the probeset Sobic.009G137100, and the comparative gene expression was assessed in 32 different organs of S. bicolor. The number of target genes were set at 50.
Binary vector construction
The pRI101-AN and pGEM-T-SbbZIP17 vectors were restricted with NdeI and SalI restriction enzymes. The purified CDS of SbbZIP17 was ligated to linearized pRI101-AN vector having sites overhang, the reaction mix was incubated at 4 °C for 12 h using Rapid DNA Ligation Kit (Invitrogen, USA). The DH5α strain of E. coli was transformed with the ligation mix using heat shock method and plated on LB agar plates having kanamycin selection (50 mg L− 1). The positive clones were confirmed by PCR using conditions as mentioned above and the resulting recombinant plasmid was named pRI101-AN-SbbZIP17. The pRI101-AN-SbbZIP17 construct was transformed into EHA105 strain of A. tumefaciens competent cells using heat shock (Kaman-Toth et al. 2018).
Development of transgenic tobacco plants
pRI101-AN-SbbZIP17 vector was transformed into 30 days old healthy tobacco (Nicotiana tobaccum cv. ‘Benthamiana’) leaf discs via Agrobacterium-mediated transformation method (Zhou et al. 2011). Cells of A. tumefaciens harboring pRI101-AN-SbbZIP17 were grown in YEM (Yeast Extract Mannitol) broth media containing antibiotics rifampicin (5 mgL− 1) and kanamycin (100 mgL− 1) at 28 °C till Optical Density (O.D., A600) reached 0.6–0.8. The A. tumefaciens cells were pellet down and resuspended in 30 ml ½ strength MS media. Small leaf discs were infected by A. tumefaciens cells in liquid MS for 20 min. Afterward, the leaf discs were air-dried using sterile Whatman filter paper and further placed on MS media for co-cultivation for 48 h at 23 ± 2 °C in dark. After 48 h, the leaf discs were shifted to selection MS media (kanamycin 200 mgL− 1, Timentin 150 mgL− 1, BAP 2 mgL− 1, and NAA 0.2 mgL− 1) and placed under normal light conditions. The leaf discs were subcultured at a regular interval of 10 d. After 1 month, the developed shoots were placed on rooting media (kanamycin 100 mgL− 1 and timentin 300 mgL− 1 in MS medium) for the development of plantlets. Following transformation of tobacco, a total of 11 probable transgenic lines (at T0 stage) were developed on kanamycin selection media. After the roots became stronger, plantlets were transferred from jam bottles to soil for hardening and kept under controlled conditions in growth chamber, and later shifted to greenhouse till maturity.
Putative tobacco transgenic plants confirmation
After screening on antibiotic selection medium, two months old transgenic tobacco T0 plants and wild-type plants were taken for genomic DNA isolation using CTAB method. The genomic DNA was checked on 0.8% agarose gel, Nanodrop spectrophotometer (ThermoScientific, USA) was used to quantify the DNA. The existence of transgene was identified by using PCR with nptII and SbbZIP primers (Table S1).
Southern hybridization of transgenic tobacco SbbZIP17 plants at T1 stage
Southern hybridization was performed for SbbZIP17-T1 transgenic lines. The genomic DNA (25 µg) of WT and transgenic lines was digested with SalI enzyme and thereafter resolved on agarose gel (0.8%). Resolved genomic fragments transferred onto a blotting membrane (Merck, USA) using capillary method (Russell and Sambrook 2001). PCR DIG Probe Synthesis kit (Roche, Germany) was used to prepare probe for SbbZIP17. Pre-hybridization was performed at 45 °C in a hybridization oven for 1 h subsequently followed by hybridization for overnight (16 h). Washing, blocking, incubation steps were carried out as per protocols given in DIG Luminescent Detection Kit (Roche, Germany). Fluorescent signals from the nitrocellulose membrane were exposed on an X-ray film placed in a cassette.
Expression analysis of SbbZIP17 and UPR pathway genes in SbbZIP17-T1 transgenic lines
The qRT-PCR was performed to asses expression of SbbZIP17 transcripts in 30-days-old T1 tobacco transgenic lines. For expression analysis of UPR pathway genes, 30-days-old seedlings of SbbZIP17-T1 transgenic lines and WT exposed to 42 °C of heat stress for 0 h and 6 h. Total RNA was isolated from transgenic and WT plants. The cDNA (complementary DNA) was synthesized from total RNA (5 µg) using SuperscriptIII First-strand cDNA synthesis kit (Invitrogen, USA). The qRT-PCR reaction steps were as follows: 95 °C for 3 min, followed by 32 cycles of 95 °C for 10 s, and 60 °C for 10 s, and 72 °C for 10 s. All the reactions were performed in three replicates and gene expression was estimated using 2−∆∆Ct equation (Schmittgen and Livak 2008). Reference gene Ntactin, was used for normalization (Table S1). The Ct value of transgenic line SbbZIP17-2 was observed to be lowest and considered as 1 (1-fold) for comparing gene expression with another transgenic lines (Fig. 4a).
Schematic representation of qPCR analysis, gene amplification and binary vector construct preparation for SbbZIP17. (a) Expression pattern of S. bicolor SbbZIP17 transcript. Statistical significance difference was indicated with an asterisk (p ≤ 0.05). Error bars represents ± SD from three technical replicates (b) PCR amplification of SbbZIP17 from S. bicolor cDNA. Lane 1 and 2 correspond to SbbZIP17 fragments,–ve corresponds to the negative control. M refers to the molecular marker (1Kb DNA ladder RTU #BM101-R500). (c) T-DNA region of binary vector pRI101-AN carrying SbbZIP17 gene under 35 S enhancer promoter. nptII (Neomycin Phosphotransferase) gene for Kan resistance, NOS (nopaline synthase gene terminator), RB-Right border, and LB-Left border
Physio-biochemical analysis of SbbZIP17-T1 transgenic tobacco lines under stress
Wild type (WT) plants and independent single copy containing southern positive SbbZIP17-T1 transgenic lines (SbbZIP17-2, SbbZIP17-3, and SbbZIP17-4) were analyzed for physiological and biochemical indices. After sterilization, seeds of SbbZIP17-T1 transgenic lines were placed on MS medium having 100 mg L− 1 kanamycin, while seeds of WT plants were placed on plain MS medium under conditions of 16 h light/8 h dark and temperature of 24 ± 1 °C. After 30d of germination, plantlets were exposed to heat stress of 42 °C for 0 h, 2 h, 4 h and 6 h in an incubator chamber, thereafter leaves were collected for physio-biochemical analysis. Following analyses were performed in biological triplicates.
Relative water content (RWC) was estimated as reported by Smart et al. (1973). Extracted and estimated total chlorophyll content as reported by Barnes et al. (1992). The MSI (Membrane Stability Index) was estimated as reported by Sairam et al. (1994) with a conductivity meter. Membrane damage was analyzed by Malondialdehyde (MDA) content, which was estimated by thiobarbituric acid (TBA) reaction as previously reported by Draper et al. (1993). A crude enzyme extracted from leaf tissue was used to assay superoxide dismutase (SOD) activity as reported by Zhang and Kirkham (1994). Total SOD activity was estimated by a capability to hinder reduction of colourless substrate NBT (Nitro Blue Tetrazolium) (Zhang and Kirkham 1995).
Statistical analysis
The estimated data of qRT-PCR and physio-biochemical analysis were applied to a one-way ANOVA through DMRT (Duncan’s Multiple Range Test) to estimate the significant difference among the means (p ≤ 0.05). All given values are means of a minimum of three replicates ± SD. GraphPad Prism 5.0 software was used for statistical analysis. Lower case letters were used to indicate the significant difference.
Results
Expression analysis, cloning and construct preparation for SbbZIP17
The qRT-PCR analysis of S. bicolor SbbZIP17 was evidenced for increased expression level under heat stress. The expression level of SbbZIP17 was induced rapidly at 2 h (14.7-fold) and reduced at 4 h (8.9-fold), then again increased at 6 h (17.3-fold) of heat stress in comparison to control sample (Fig. 1a). The PCR amplification of SbbZIP17 with complete ORF carried out using cDNA from total RNA (2 h) (Fig. 1b). Followed by sequencing and annotation of SbbZIP17, further it was cloned into shuttle vector pRI101-AN (Fig. 1c).
In silico analysis of SbbZIP17
Sequencing analysis confirmed that the length of SbbZIP17 CDS was 1971 bp, which was translated into a protein of 656 aa, having a molecular weight of 69.57 kDa. A BRLZ (basic region leucine zipper) domain (177–240 aa) was predicted in SbbZIP17, along with a nuclear localization signal (NLS) (198–207 aa) within BRLZ domain, a TM (Transmembrane) domain (320–342 aa) and a canonical S1P protease site (557–561 aa) (Fig. 2a). Thus, the presence of NLS at N-terminal (towards cytosol) and a TM (Transmembrane) domain at C-terminal (towards ER lumen), is a property of homologous proteins, which led to point out that SbbZIP17 may be present in the ER membrane. Multiple sequence alignment and motif analysis illustrated that SbbZIP17 is extremely conserved among different plant species (Fig. 2b). SbbZIP17 shared similar motifs while motif 15 present only once in SbbZIP17, while it occurred twice in homologous protein (Fig. S1). Homology analysis of SbbZIP17 revealed 97.71% identity with S. bicolor SbbZIP39 (XP_002441096.1), 84.85% identity with the S. italica SibZIP39 (XP_004962093.1), 85% with Z. mays (NP_001148077.1), 73.52% with O. sativa OsbZIP39 (XP_015640085.1), 38.52% with A. thaliana (NP_565946.1). Physio-biochemical features of SbbZIP17 are given in Table S2. However, a few additional homologous proteins, HvbZiP17 (KAE8789011.1), AcbZIP39 (XP_020086811.1), MabZIP17 (XP_009384980.1), and SobZIP17 (XP_021840232.1) were used to analyse the evolutionary relationship (Fig. S2). The homologous proteins are well recognized to contribute a significant function in the UPR or the ER stress response under abiotic stress conditions. It is predicted to be localized inside nucleus and gene ontology analysis indicated a role in biological processes including response to stress, signal transduction, and biosynthetic process. Cloned CDS was designated as SbbZIP17 (accession no: MW532120).
The protein SbbZIP17 was predicted to have a transmembrane helix (TMH) from a region of 320 to 342 amino acids (Fig. S3a). Further, there were 6 N-glycosylation sites and 28 O-linked glycosylation sites present in the protein (Fig. S3b, c). The secondary structure of SbbZIP17 observed 8 strands and 9 helices (Fig. S4a). 3D ligand binding site was predicted and it showed that only 1 amino acid (i.e., Leucine) was involved in binding located at position 187 (Fig. S4b). Protein-protein interaction showed that SbbZIP17 interacted with stress-responsive proteins and TFs like heat shock 70 (Sb01g010460.1), bZIP transcription factor 60 (Sb02g001970.1), NF-YC4 (Sb07g005060.1), NF-YC3 (Sb07g005540.1) (Fig. S4c). SbbZIP17 also interacted with protease like endoribonuclease IRE1 (Sb02g032960.1), membrane-bound TF site-2 protease (Sb03g029800.1), subtilisin-like protease SBT6.1 (Sb10g004450.1). This analysis further concluded that the SbbZIP17 has a major role under abiotic stress conditions. Based on available mRNA-seq data, we observed expression of SbbZIP17 was higher at stem elongation, heading, milking, and ripening phase (Fig S5a). Similarly, in almost all other organs expression of SbbZIP17 was more than 4-fold including leaf, flag leaf while in case of pollen grains it was down-regulated (Fig S5b). The tissue-specific expression analysis of genes homologous to SbbZIP17 showed diverse differential expression pattern (Fig S6).
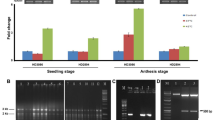
Development of tobacco lines overexpressing SbbZIP17
The T0 seeds of all transgenic lines were harvested to obtain T1 generation lines for further analysis (Fig. S7). Among them, 7 lines were identified as transgenic by using PCR with nptII and SbbZIP17 primers at T0 stage (Fig. 3a). Southern hybridization at T1 stage revealed that 6 out of 7 tobacco PCR positive transgenic lines were independent events having 1 to 2 copies of transgene integrations (Fig. 3b). The transgenic lines SbbZIP17-2, SbbZIP17-3 and SbbZIP17-4 showed single-copy transgene integration (Fig. 3b), the transgenic line SbbZIP17-1, SbbZIP17-6, and SbbZIP17-7 showed two copies of transgene. However, transgenic line SbbZIP17-5 failed to show any transgene integration (Fig. 3b). These analyses showed that SbbZIP17 was integrated into the tobacco genome independent of other lines. Single-copy transgene containing transgenic tobacco lines were used for more analysis.
Putative tobacco transgenic plants confirmation and southern hybridization of transgenic lines. (a) PCR analysis of transgenic tobacco lines at T0, using SbbZIP17 gene (1971 bp) and nptII (750 bp) specific primers. ‘–‘ sign represents negative control i.e., reaction without any template DNA, ‘+’ sign indicates positive control. (b) Southern hybridization of SbbZIP17-T1 transgenic lines using SbbZIP17 probe., ‘+’ indicates digested and linearized pRI101-AN-SbbZIP17 vector backbone (~ 10 kb), -ve represent genomic DNA of wild type tobacco plant
Expression analysis of SbbZIP17 and UPR pathway genes in SbbZIP17-T1 transgenic lines
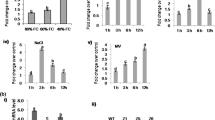
Transgene SbbZIP17 was expressed 26.4 to 52.6 folds in T1 transgenic tobacco lines (Fig. 4a). Further to verify the role of SbbZIP17 in activation of UPR stress responsive genes we performed expression analysis of BiP, PDIL, CNX, and CRT1. Under normal conditions, the genes of UPR pathway expressed approximately at similar level in WT and transgenic lines. However, in response to heat stress all UPR pathway genes were induced in transgenic tobacco than WT plants (Fig. 4b, c, d, e). These analyses showed that SbbZIP17 expression in transgenic tobacco lines, which involved in activation of genes related to UPR pathway under heat stress conditions.
Expression analysis of SbbZIP17 and UPR pathway genes in SbbZIP17-T1 transgenic lines. (a) Expression analysis of SbbZIP17 in SbbZIP17-T1 transgenic lines. The Ct value of SbbZIP17-2 was the lowest and set as 1-fold for comparison. Expression level of UPR pathway genes (b) BiP (c) PDIL and (d) CRT1 in SbbZIP17-T1 transgenic lines
Physio-biochemical analysis of SbbZIP17-T1 transgenic lines in response to heat stress
Biochemical indexes such as Membrane Stability Index (MSI), Relative Water Content (RWC), and Total chlorophyll content was observed to increase in SbbZIP17-T1 transgenic lines as compared to WT plants under heat stress (Fig. 5). The physiological parameters were positively improved in all tobacco transgenic lines in comparison to WT plants.
Changes in lipid peroxidation activity in SbbZIP17-T1 transgenic lines in response to heat stress were studied. Under controlled conditions, WT and transgenic lines demonstrated nearly similar concentrations of MDA (Fig. 5d). However, under heat stress conditions, the quantity of MDA in transgenic lines and WT were considerably increased, yet the mean increased level of the MDA quantity in tobacco transgenic lines was less in comparison to WT (Fig. 5d). This analysis demonstrated that tobacco transgenic lines with less cell membrane damage as compared to the WT under heat stress. Further, analyzed the quantity of antioxidant enzyme (SOD), between WT and SbbZIP17-T1 transgenic lines under heat stress. It revealed that amount of SOD enzyme in SbbZIP17-T1 transgenic lines significantly enhanced, however less increment was observed in WT plants comparatively.
Discussion
The bZIP gene family play important functions in development and adaptation to abiotic stress conditions in plant species. As well, Sorghum bicolor, an elite species naturally tolerant to abiotic stresses and has outstanding potential source for bioprospecting of genes liable for abiotic stress tolerance (Mutava et al. 2011; Tari et al. 2013; Baillo et al. 2020). Therefore, present study was performed for functional analysis of the bZIP gene from S. bicolor.
The qRT-PCR analysis of SbbZIP17 from S. bicolor was showed increased expression level for more than 4-fold under heat stress of 2 h, 4 h and 6 h (Fig. 1a). However, in response to heat stress, genes of UPR pathway (CRT1, BiP, and PDIL) were induced in transgenic tobacco lines overexpressing SbbZIP17 than WT plants (Fig. 4b, c, d, e). An abiotic stress condition disrupts functional protein assembly and promotes misfolding in the ER leads to turn on UPR pathway (Howell 2013). When build-up of misfolded proteins in ER, moves ER located MTTFs in Golgi body for processing by S1P and S2P protease present in it. DNA binding section of MTTFs transported into nucleus upon S1P intramembrane processing and acts as a TF. In nucleus MTTFs increase expression levels of UPR pathway genes, for instance, CRT1, CNX, BiP (ER chaperone), and PDIL, which assist protein folding in ER lumen thereby provide abiotic stress tolerance (Liu 2012). In A. thaliana, four MTTFs AtbZIP members (AtbZIP17, AtbZIP28, AtbZIP49, and AtbZIP60) responsible for the transduction of ER stress signals (Liu and Howell 2010; Bao and Howell 2017).
The cloned SbbZIP17 encodes a type II transmembrane bZIP MTTF, these TFs usually have a cytosolic facing N-terminus and C-terminus with a canonical S1P cleavage site facing the ER lumen (Howell et al. 2013; Yang et al. 2013). Recently abiotic stress-responsive bZIP genes including MTTFs bZIP from different plant species, T. aestivum, and C. annum were isolated and analyzed based on in silico studies (Agarwal et al. 2019; Gai et al. 2020;). TFs reported in A. thaliana such as AtbZIP17, AtbZIP28, and AtbZIP49 are type II MTTFs processed with regulated intramembrane proteolysis (RIP) and moves to nucleus thereby turn on the UPR pathway. In contrast AtbZIP60 was processed by IRE1-dependent splicing under ER stress (Howell et al. 2013). In Zea mays, ZmbZIP60 was identified to be involved in UPR and it belongs to the IRE1 pathway (Howell et al. 2013). Similarly, in our study protein-protein interaction predicted that SbbZIP17 interacted with protease endoribonuclease IRE1 (Sb02g032960.1), membrane-bound TF site-2 protease (Sb03g029800.1), and subtilisin-like protease SBT6.1 (Sb10g004450.1). Even under the absence of major abiotic stress conditions, little quantity of bZIPs move from the Golgi body into nucleus because plants are continuously exposed to different small intensity stresses, for instance, temperature variations and irrigation times (Che et al. 2010). Therefore, we observed in silico expression of SbbZIP17 at the stem elongation, heading, milking, and ripening phase and lowest during germination (Fig S5a). Almost in all analysed anatomical organs expression of SbbZIP17 was observed including flag leaf and other leaves while in pollen grain it was found to be down-regulated (Fig S6b).
Further to analyse the role of SbbZIP17 in response to heat stress, we developed single-copy transgene integrated tobacco lines overexpressing SbbZIP17 (Fig. 3b). Several abiotic stress-responsive members of bZIP family from S. bicolor, P. trifoliate, M. hupehensis, T. aestivum, O. sativa, P. trifoliata, and C. annum were functionally characterized for drought, heat, osmotic and salinity stress tolerance (Huang et al. 2010; Wang et al. 2011; Zhang et al. 2012; Agarwal et al. 2019; Yang et al. 2019; Gai et al. 2020). Earlier experiments demonstrated that the constitutive expression of bZIP MTTFs such as AtbZIP60 and AtbZIP28 in A. thaliana offer tolerance to high temperature and salinity stress, correspondingly (Fujita et al. 2007; Gao et al. 2008). A. thaliana overexpressing maize ZmbZIP17 the ortholog of AtbZIP17 demonstrated that ER acts as stress transducer under stress and optimal growth conditions (Yang et al. 2013). The ZmbZIP17 was upregulated by ER stress-inducing agents and ABA. ZmbZIP17 localized in ER under normal conditions thereafter transported to nucleus due to ER stress-eliciting agents or deletion of transmembrane domain (Yang et al. 2013).
Transgenic tobacco lines overexpressing SbbZIP17 genes showed enhanced hydration status, antioxidant activities, reduced chlorophyll loss, and membrane damage under heat stress conditions (Fig. 5) The RWC is regarded as an index of dehydration tolerance which points out the metabolic activities and measured the status of water in plant cells (Sinclair and Ludlow 1986). Abiotic stresses such as heat or cold can negatively affect Chl biosynthesis and lead to its degradation, therefore Chl content is considered as a parameter of thermo-tolerance (Rossi et al. 2017). Additionally, heat stress damage the photosynthetic machinery, consequently inhibit plant growth and development. Transgenic tobacco overexpressing EcbZIP17 from E. coracana increased chlorophyll and RWC content in the leaves (Ramakrishna et al. 2018). A few reactive oxygen species (ROS) like hydroxyl radical, hydrogen peroxide, and superoxide, would be produced and build up in plant cells under abiotic stress conditions (Gosavi et al. 2014). The build-up of ROS is cytotoxic to cell and lead to increase in cell membrane permeability, inactivate enzymes, and damage cellular components (Karuppanapandian et al. 2011). To reduce oxidative damages in plant cells, antioxidative enzymes including SOD is playing a fundamental role in scavenging harmful ROS (Gill et al. 2010; Hameed et al. 2012). In O. sativa overexpressing OsbZIP62 gene enhanced the oxidative and drought stress tolerance, whereas OsbZIP62 mutants showed contrasting phenotype by modulating levels of SOD, MDA, and chlorophyll content (Huang et al. 2010; Yang et al. 2019). Cell membrane injury and destruction of cell structural components are the characteristic attributes related to heat stress damage in plant species (Xu et al. 2014). Transgenic tobacco over-expressing EcbZIP17 and PtrABF bZIP gene from E. coracana and Poncirus trifoliate respectively strengthen membrane stability (Huang et al. 2010; Ramakrishna et al. 2018). MDA is the ultimate product of membrane peroxidation, higher the peroxidation, the more amount of MDA produced (Hameed et al. 2012). The OsbZIP62 mutant of O. sativa contains significantly greater H2O2 and MDA contents than wild type and reduced tolerance to oxidative damage and drought stress (Fig. 5) (Yang et al. 2019). Similarly, SbbZIP17 expressing transgenic tobacco lines suggested that SbbZIP17 was involved in enhanced heat stress tolerance. The gene SbbZIP17 is a potential genomic resource for breeders and researchers for genetic gains in Sorghum crop especially to provide heat stress tolerance under globally changing climatic conditions.
Conclusion
Expression level of SbbZIP17 from S. bicolor was increased under heat stress condition. Bioinformatics analysis of cloned SbbZIP17 showed that it belongs to type II MTTF of bZIP family. It is highly conserved, having a single ligand-binding site, interacting with heat stress-responsive proteins (HSP70, NF-Ys), expressed in different tissues and organs. Transgenic tobacco lines overexpressing SbbZIP17 responsible for increased hydration status, SOD, and reduces MDA content, chlorophyll loss, and membrane damage by regulating the UPR pathway genes. The SbbZIP17 can be used as a potential candidate for the development of climate-resilient crops. Present work has paved way for further study to understand the function of S. bicolor bZIPs for abiotic stress tolerance.
Data availability
Gene SbbZIP17 sequence data was deposited into NCBI database under accession number MW532120.
References
Agarwal PK, Shukla PS, Gupta K, Jha B (2013) Bioengineering for salinity tolerance in plants: state of the art. Mol Biotechnol 54:102–123
Agarwal P, Baranwal VK, Khurana P (2019) Genome-wide analysis of bZIP transcription factors in wheat and functional characterization of a TabZIP under abiotic stress. Sci Rep 9:1–18
Baillo EH, Hanif MS, Guo Y, Zhang Z, Xu P, Algam SA (2020) Genome-wide identification of WRKY transcription factor family members in sorghum (Sorghum bicolor (L.) moench). PLoS ONE 15, e0236651
Banerjee A, Roychoudhury A (2017) Abscisic-acid-dependent basic leucine zipper (bZIP) transcription factors in plant abiotic stress. Protoplasma 254:3–16
Bao Y, Howell SH (2017) The unfolded protein response supports plant development and defense as well as responses to abiotic stress. Front Plant Sci 8:344
Barnes JD, Balaguer L, Manrique E, Elvira S, Davison AW (1992) A reappraisal of the use of DMSO for the extraction and determination of chlorophylls a and b in lichens and higher plants. Environ Exp Bot 32:85–100
Che P, Bussell JD, Zhou W, Estavillo GM, Pogson BJ, Smith SM (2010) Signaling from the endoplasmic reticulum activates brassinosteroid signaling and promotes acclimation to stress in Arabidopsis. Sci Signal 3:69
Draper HH, Squires EJ, Mahmoodi H, Wu J, Agarwal S, Hadley M (1993) A comparative evaluation of thiobarbituric acid methods for the determination of malondialdehyde in biological materials. Free Rad Biol Med 15:353–363
Fujita M, Mizukado S, Fujita Y, Ichikawa T, Nakazawa M, Seki M (2007) Identification of stress-tolerance-related transcription-factor genes via mini-scale full-length cDNA Over-expressor (FOX) gene hunting system. Biochem Biophy Res Commun 364:250–257
Gai WX, Ma X, Qiao YM, Shi BH, Li QH, Wei AM (2020) Characterization of the bZIP transcription factor family in pepper (Capsicum annuum L.): CabZIP25 positively modulates the salt tolerance. Front Plant Sci 11:139
Gao H, Brandizzi F, Benning C, Larkin RM (2008) A membrane-tethered transcription factor defines a branch of the heat stress response in Arabidopsis thaliana. Proc Natl Acad Sci 105:16398–16403
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Gosavi GU, Jadhav AS, Kale AA, Gadakh SR, Pawar BD, Chimote VP (2014) Effect of heat stress on proline, chlorophyll content, heat shock proteins and antioxidant enzyme activity in sorghum (Sorghum bicolor) at seedlings stage. Indian J Biotechnol 13:356–363
Goswami S, Kumar RR, Dubey K, Singh JP, Tiwari S, Kumar A et al (2016) SSH analysis of endosperm transcripts and characterization of heat stress regulated expressed sequence tags in bread wheat. Front Plant Sci 7:1230
Gupta K, Jha B, Agarwal PK (2014) A dehydration-responsive element binding (DREB) transcription factor from the succulent halophyte Salicornia brachiata enhances abiotic stress tolerance in transgenic tobacco. Mar Biotechnol 16:657–673
Halder T, Agarwal T, Ray S (2016) Isolation, cloning, and characterization of a novel Sorghum dehydrin (SbDhn2) protein. Protoplasma 253:1475–1488
Hameed A, Goher M, Iqbal N (2012) Heat stress-induced cell death, changes in antioxidants, lipid peroxidation, and protease activity in wheat leaves. J Plant Growth Regul 31:283–291
Hayashi S, Wakasa Y, Takaiwa F (2013) Recent advances in understanding the control of secretory proteins by the unfolded protein response in plants. Int J Mol Sci 14:9396–9407
Howell SH (2013) Endoplasmic reticulum stress responses in plants. Annu Rev Plant Biol 64:477–499
Huang XS, Liu JH, Chen XJ (2010) Overexpression of PtrABF gene, a bZIP transcription factor isolated from Poncirus trifoliata, enhances dehydration and drought tolerance in tobacco via scavenging ROS and modulating expression of stress-responsive genes. BMC Plant Biol 10:230
Janni M, Gullì M, Maestri E, Marmiroli M, Valliyodan B, Nguyen HT, Marmiroli N (2020) Molecular and genetic bases of heat stress responses in crop plants and breeding for increased resilience and productivity. J Exp Bot 71(13):3780–3802
Kaman-Toth E, Pogany M, Danko T, Szatmari A, Bozso Z (2018) A simplified and efficient Agrobacterium tumefaciens electroporation method. 3 Biotech 8:148
Karuppanapandian T, Moon JC, Kim C, Manoharan K, Kim W (2011) Reactive oxygen species in plants: their generation, signal transduction, and scavenging mechanisms. Aust J Crop Sci 5:709
Liu JX, Howell SH (2010) Endoplasmic reticulum protein quality control and its relationship to environmental stress responses in plants. Plant Cell 22:2930–2942
Liu JX, Srivastava R, Howell SH (2008) Stress-induced expression of an activated form of AtbZIP17 provides protection from salt stress in Arabidopsis. Plant Cell Environ 31:1735–1743
Liu C, Wu Y, Wang X (2012) bZIP transcription factor OsbZIP52/RISBZ5: a potential negative regulator of cold and drought stress response in rice. Planta 235:1157–1169
Mulaudzi-Masuku T, Mutepe RD, Mukhoro OC, Faro A, Ndimba B (2015) Identification and characterization of a heat-inducible Hsp70 gene from Sorghum bicolor which confers tolerance to thermal stress. Cell Stress Chaperones 20:793–804
Mutava RN, Prasad PVV, Tuinstra MR, Kofoid KD, Yu J (2011) Characterization of sorghum genotypes for traits related to drought tolerance. Field Crops Res 123:10–18
Ngara R, Ndimba R, Borch-Jensen J (2012) Identification and profiling of salinity stress-responsive proteins in Sorghum bicolor seedlings. J Proteom 75:4139–4150
Nijhawan A, Jain M, Tyagi AK, Khurana JP (2008) Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol 146:333–350
Panzade KP, Kale SS, Chavan NR, Hatzade B (2021) Genome-wide analysis of Hsp70 and Hsp100 gene families in Ziziphus jujuba. Cell Stress Chaperon 26(2):341–353
Panzade KP, Kale SS, Manoj ML, Kothawale SP, Damse DN (2022) Genome-wide analysis and expression profile of nuclear factor Y (NF-Y) gene family in Z. Jujuba. Biotechnol Appl Biochem 194(3):1373–1389
Ramakrishna C, Singh S, Raghavendrarao S, Padaria JC, Mohanty S, Sharma TR, Solanke AU (2018) The membrane tethered transcription factor EcbZIP17 from finger millet promotes plant growth and enhances tolerance to abiotic stresses. Sci Rep 8:1–14
Rossi S, Burgess P, Jespersen D, Huang B (2017) Heat-induced leaf senescence associated with chlorophyll metabolism in bentgrass lines differing in heat tolerance. Crop Sci 57:S–169. https://doi.org/10.2135/cropsci2016.06.0542
Russell DW, Sambrook J (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Saeed F, Chaudhry UK, Raza A, Charagh S, Bakhsh A, Bohra A, Varshney RK (2023) Developing future heat-resilient vegetable crops. Funct Integr Genom 23(1):47
Sairam RK (1994) Effect of moisture-stress on physiological activities of two contrasting wheat genotypes. Indian J Exp Biol 32:594–594
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C T method. Nat Protoc 3:1101
Sinclair TR, Ludlow MM (1986) Influence of soil water supply on the plant water balance of four tropical grain legumes. Funct Plant Biol 13:329–341
Smart RE, Bingham GE (1973) Rapid estimates of relative water content. Plant Physiol 53:258–260
Sornaraj P, Luang S, Lopato S, Hrmova M (2016) Basic leucine zipper (bZIP) transcription factors involved in abiotic stresses: a molecular model of a wheat bZIP factor and implications of its structure in function. Biochim Biophys Acta 1860:46–56
Su M, Li XF, Ma XY, Peng XJ, Zhao AG, Cheng LQ, Chen SY, Liu GS (2011) Cloning two P5CS genes from bioenergy sorghum and their expression profiles under abiotic stresses and MeJA treatment. Plant Sci 181:652–659
Tari I, Laskay G, Takacs Z, Poor P (2013) Response of sorghum to abiotic stresses: a review. J Agron Crop Sci 199:264–274
Vettore AL, Yunes JA, Neto GC, Da Silva MJ, Arruda P, Leite A (1998) The molecular and functional characterization of an Opaque2 homologue gene from Coix and a new classification of plant bZIP proteins. Plant Mol Biol 36:249–263
Wang J, Zhou J, Zhang B, Vanitha J, Ramachandran S, Jiang SY (2011) Genome-wide expansion and expression divergence of the Basic Leucine Zipper Transcription Factors in higher plants with an emphasis on Sorghum. J Integr Plant Biol 53:212–231
Xu Q, Xu X, Shi Y, Xu J, Huang B (2014) Transgenic tobacco plants overexpressing a grass PpEXP1 gene exhibit enhanced tolerance to heat stress. PLoS ONE 9:e100792
Yang O, Popova OV, Süthoff U, Lüking I, Dietz KJ, Golldack D (2009) The Arabidopsis basic leucine zipper transcription factor AtbZIP24 regulates complex transcriptional networks involved in abiotic stress resistance. Gene 436:45–55
Yang YG, Lv WT, Li MJ, Wang B, Sun DM, Deng X (2013) Maize membrane-bound transcription factor Zmbzip17 is a key regulator in the cross-talk of ER quality control and ABA signaling. Plant Cell Physiol 54:2020–2033
Yang S, Xu K, Chen S, Li T, Xia H, Chen L (2019) A stress-responsive bZIP transcription factor OsbZIP62 improves drought and oxidative tolerance in rice. BMC Plant Biol 19:1–15
Zhang J, Kirkham MB (1994) Drought-stress-induced changes in activities of superoxide dismutase, catalase, and peroxidase in wheat species. Plant Cell Physiol 35:785–791
Zhang J, Kirkham MB (1995) Water relations of water-stressed, split‐root C4 (Sorghum bicolor; Poaceae) and C3 (Helianthus annuus; Asteraceae) plants. Am J Bot 82:1220–1229
Zhang JY, Qu SC, Du XL, Qiao YS, Cai BH, Guo ZR, Zhang Z (2012) Overexpression of the Malus hupehensis MhTGA2 gene, a novel bZIP transcription factor for increased tolerance to salt and osmotic stress in transgenic tobacco. Int J Plant Sci 173:441–453
Zhou C, Qian Z, Ji Q, Xu H, Chen L, Luo X, Liang M, Kexuan T, Jianbo X, Guoyin K (2011) Expression of the zga agglutinin gene in tobacco can enhance its anti-pest ability for peach-potato aphid (Myzus Persica). Acta Physiol Plant 33:2003–2010
Acknowledgements
Authors are thankful to principal of SDMVM College of Agricultural Biotechnology, Georai Tanda, for providing necessary facilities and technical support.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
KPP Conceptualize, designed methodology, conducted experiments, most of formal analysis, data curation, visualization, wrote manuscript and supervise; HV conducted part of in silico analysis, helped in experiment, edited and reviewed the manuscript; SPK help in experiments, edited, and reviewed the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
All the authors have been agreed to submit it. This article does not contain any studies with animals performed by any of the authors.
Consent to participate
Before the submission of paper, all the author has given the consent to publish.
Consent to Publish
All the authors have given the consent to publish.
Conflict of interest
The authors declare that they have no conflict of interests to this work. We declare that we do not have any commercial or associative interest that represents a conflict of interest in connection with the work submitted.
Additional information
Communicated by Goetz Hensel.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Panzade, K.P., Vishwakarma, H. & Kothawale, S.P. Indexing heat stress induced changes in transgenic tobacco by overexpressing membrane-tethered transcription factor from Sorghum bicolor bZIP17. Plant Cell Tiss Organ Cult 157, 22 (2024). https://doi.org/10.1007/s11240-024-02749-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11240-024-02749-x