Abstract
Phytochelatins chelate heavy metal ions to decrease their toxicity. The chelates are then transferred to, and stored in, the vacuole. Phytochelatin synthase (PCS), which is involved in phytochelatin synthesis, is thought to be a key enzyme for phytoremediation. In this study, a PCS gene encoding phytochelatin synthase was cloned from poplar (Populus tomentosa Carr.), a widely grown model woody plant that accumulates high levels of heavy metals, especially cadmium. Poplar is considered to have potential applications in phytoremediation. The full-length PtPCS cDNA (1512-bp) encoded a polypeptide of 503 amino acid residues. The PtPCS cDNA was transferred into tobacco by Agrobacterium-mediated leaf disk transformation. The transgenic and wild-type (WT) lines of tobacco were subjected to a one time Cd treatment (90 μmol Cd2+) for 30 days, and then evaluated to determine their Cd tolerance. We evaluated morphological and physiological indices including leaf relative electrolyte leakage, malondialdehyde content, total superoxide dismutase activity, chlorophyll content and root activity. Compared with WT plants, the transgenic plants expressing PtPCS grew better in the Cd treatment and showed significantly higher Cd tolerance. Compared with WT plants, the transgenic lines accumulated higher concentrations of Cd (1.7 to 3.0-fold higher Cd concentration in roots; 1.24 to 2.28-fold higher Cd concentration in leaves). However, the transfer coefficient was lower in the transgenic lines than in wild type. We concluded that PtPCS encodes a functional PCS that may be involved in Cd tolerance and accumulation, but not in Cd transport.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With increasing industrialization, there is greater pollution of soils with heavy metals, which negatively affect human health. Phytoremediation refers to the process in which heavy metals are absorbed by plants, and are consequently removed from the soil. This process is regarded as a healthy, environmentally friendly, and low-cost method to control soil heavy metal pollution (Shukla et al. 2013). Several genes and gene families have been identified to play roles in heavy metal enrichment in plants. These genes/gene families include those encoding cation diffusion facilitator (CDF) family proteins (Clemens et al. 2002), natural resistance macrophage associated proteins (Nramp) (Thomine et al. 2000), heavy metal-transporting ATPases (Bernarda et al. 2004; Lee et al. 2007), phytochelatin synthases (PCS) (Clemens et al. 1999), metallothioneins (Hasegawa et al. 1997; Vrbová et al. 2013), the yellow stripe-like (YSL) protein family (Curie et al. 2009), Zn2+ transporters (ZIP) (Pence et al. 2000), ATP-binding cassette (ABC) transporter (Rea 2007; Bhuiyan et al. 2011a, b), and some key enzymes involved in glutathione (GSH) biosynthesis of GSH S-transferases (GSTs) (Polle et al. 2013) and γ-glutamylcysteine synthetase (γ-ECS) (He et al. 2015).
Phytochelatins (PCs) are a family of thiol-rich peptides, with the general structure (γ-Glu-Cys)n-Gly (n = 2–11). These compounds were first identified from Rauvolfia serpentine by Grill et al. (1985). PCs can bind heavy metal ions, especially cadmium ions, forming almost non-toxic heavy metal-protein complexes. This decreases the concentration of free heavy metal ions and decreases their toxicity in plant cells. Also, heavy metal-PC complexes can be transferred to, and stored in, the vacuole, where they are separated from enzymes and other active substances in the cytoplasm (Yadav 2010). This heavy metal response mechanism is ubiquitous in plants (DalCorso et al. 2008; Meyer et al. 2011). The role of PCs in heavy metal detoxification was confirmed in glutathione-deficient mutants (gsh −) of Schizosaccharomyces pombe (Glaeser et al. 1991) and in phytochelatin-deficient mutants (PC −) of Arabidopsis thaliana (Cobbett 2000).
Phytochelatins are not gene-encoded proteins, but are produced from GSH via the activity of phytochelatin synthase (PCS). A transgenic line expressing a PCS showed increased contents of PCs (Gasic and Korban 2007a; Brunetti et al. 2011) and non-protein thiols (NPTs) in cadmium-treated shoots (Gasic and Korban 2007a). Several studies have shown that the PCS gene encoding PCS is constitutively expressed, but is further activated in the presence of heavy metal ions (Vatamaniuk et al. 2004; Rea 2012; Shukla et al. 2012). Several PCS genes have been cloned from various species, such as Arabidopsis (AtPCS1) (Vatamaniuk et al. 1999), Triticum aestivum (TaPCS1) (Clemens et al. 1999; Martínez et al. 2006; Wang et al. 2012), Brassica juncea (BjPCS1) (Heiss et al. 2003), Cynodon dactylon (CdPCS1) (Li et al. 2006), Thlaspi caerulescens (TcPCS1) (Meyer et al. 2011; Liu et al. 2011), Phragmites australis (PaPCS) (Zhao et al. 2014), and Ciona intestinalis (CiPCS) (Franchi et al. 2014). Functional analyses have confirmed their roles in heavy metal resistance and accumulation.
Populus tomentosa Carr. is a model woody plant that plays important roles in ecological and environmental protection. It has been demonstrated that poplar plants can accumulate high concentrations of heavy metals (Zacchini et al. 2009; Polle et al. 2013), especially Cd and zinc (Robinson et al. 2000; Lee et al. 2003). The Cd contents of dry leaves can be as high at 209 μg g−1 when the Cd concentration in soil is >100 μg g−1 (Robinson et al. 2000). The cadmium concentrations in leaf, bark and root of Populus × canescens were about 200, 200 and 700 μg g−1, respectively, when the poplar plant were exposed to 200 μM CdSO4 in sand culture (He et al. 2013). Therefore, poplar has potential applications in the phytoremediation of Cd-polluted soil. The aim of this study was to clone the PCS gene from P. tomentosa Carr., and evaluate whether it could increase Cd-tolerance and Cd accumulation in transgenic lines of Nicotiana tabacum L. The results of this study contribute to our understanding of the molecular mechanism of Cd resistance in woody plants, and provide an important genetic resource for generating phytoremediation via genetic engineering.
Materials and methods
Plant materials and expression vector
The PtPCS gene was cloned from the P. tomentosa cultivar TC1521. Aseptic plantlets of N. tabacum (tobacco) were used for gene transformation. We used the plant expression vector pEZR(K)-LC, which contains two expression units; 35S promoter-GFP gene and 35S promoter-neomycin phosphotransferase II (npt II) gene. The latter confers kanamycin resistance for host cells. Both the expression vector and Agrobacterium tumefaciens strain LBA4404 were stored in our laboratory. We used cadmium chloride hemi-pentahydrate (CdCl2 99.0 %) solution for the Cd treatment.
Cloning of PtPCS gene and construction of expression vector
Total RNA was isolated from the Cd2+-treated (100 mL 0.9 mM CdCl2) poplar leaves by the TRIzol method (Invitrogen, Carlsbad, CA, USA). Based on the PtPCS sequence reported by Liu et al. (2012), the following pair of adaptor primers was designed: PCSLCF: 5′-CGGAAGCTTATGGCGATGGCGGGGTTGTAC-3′ (Hind III site underlined) and PCSLCR: 5′-CGGGAATTCCTAGGAAAGAGGTGCGCCGAG-3′ (EcoR I site underlined). The full-length PtPCS gene was cloned from P. tomentosa samples and inserted into the pEASY-Blunt vector. After digestion with Hind III and EcoR I, the positive gene clone and the plant expression vector pEZR(K)-LC were ligated to form the recombinant expression vector pEZR(K)-LC-PtPCS, which was introduced into A. tumefaciens LBA4404 by electroporation.
Gene transformation of tobacco plants
The Agrobacterium-mediated leaf disk method was used for the genetic transformation of tobacco. Cells of A. tumefaciens LBA4404 harboring the recombinant expression vector pEZR(K)-LC-PtPCS were grown overnight at 28 °C in YEB medium (pH 7.0) containing 50 μg/mL kanamycin. The activated bacteria were transferred to antibiotic-free LB liquid medium and cultivated to an OD600 of 0.4. Young tobacco leaves were cut into small pieces (0.5 × 0.5 cm). The pieces were immersed in the bacterial solution for 5 min, transferred to differentiation medium [MS + 3 mg/L 6-benzyladenine (BA) + 0.2 mg/L naphthaleneacetic acid (NAA) + 3 % (w/v) sucrose + 0.8 % (w/v) agar], and kept in the dark at 28 °C for 3 d. Then, the tobacco pieces were transferred onto MS differentiation medium containing kanamycin (300 mg/L) and sodium cefotaxime (250 mg/L). Regenerated shoots were transferred onto MS rooting medium [MS + 0.1 mg/L NAA + 3 % (w/v) sucrose + 0.8 % (w/v) agar] containing kanamycin (300 mg/L) and sodium cefotaxime (250 mg/L). RNA were extracted from the regenerated tobacco leaves. Successful transformation was confirmed by RT-PCR analyses with the gene-specific primers.
Evaluation of transgenic tobacco lines
The transgenic and wild-type (WT) tobacco plants were transplanted into plastic pots (15 × 20 cm) containing perlite and vermiculite (1:1 v/v). Six replicates for each line were grown in a greenhouse at 25 °C for 3 weeks. For the Cd treatment, 100 mL 0.9 mM CdCl2 was added to each pot. After 1 month, the plants were analyzed to determine their Cd content and Cd resistance based on phenotypic and physiological indices.
To measure electrolyte leakage, 10 leaf discs (1 cm in diameter) were collected from leaves of each tobacco line, immersed in 40 mL deionized water, and shaken overnight. The electrical conductivity of the solution was measured (R1). The solutions containing the leaves were then boiled for 15 min, shaken overnight, and then the total conductivity was determined (R2; maximum conductivity of the tissue). Relative electrolyte leakage (REL) was calculated using the formula REL = R1/R2 × 100 %.
Malondialdehyde (MDA) levels were estimated by the thiobarbituric acid (TBA) method. Briefly, 0.2 g leaf tissue was homogenized in 1 mL of 10 % trichloroacetic acid (TCA). The mixture was centrifuged, and then the supernatant was collected and mixed with 0.6 % TBA. The mixture was boiled for 10 min, immediately cooled on ice, and then centrifuged. Absorbance of the supernatant at 450, 532, and 600 nm was measured. The MDA content was calculated as follows: (6.45(A532−A600)−0.56A450) × total extract volume (mL)/total fresh weight of sample (g).
To estimate chlorophyll content, 0.1 g leaf tissue was immersed in 10 mL dimethylsulfoxide (DMSO) in the dark for 2 days. The absorbance of the solution at 645 and 663 nm was measured. Chlorophyll a and chlorophyll b contents were calculated using the following formulae: Chla = (0.0127 × A663−0.00269 × A645) × total extract volume (mL)/total fresh weight of sample (g); Chlb = (0.0029 × A645−0.00468 × A663) × total extract volume (mL)/total fresh weight of sample (g).
Root activity was determined by TTC (2,3,5-triphenyltetrazolium chloride) method. 0.1 g cleansed roots was incubated in 2.5 mL of 0.4 % TTC for 24 h in the dark. After rinsing with distilled water and drying out with filter paper, the samples were putting into 5 mL 95 % ethanol at 60 °C for 4 h. Absorbance at 490 nm was measured. The tripheny formazan (TTF) content was inferred with the standard curve and computed for the root vitality by TTF content/root dry weight (g).
Total superoxide dismutase activity was estimated with a Total Superoxide Dismutase (SOD) Kit (Jiancheng, Nanjing, China) following the manufacturer’s instructions.
Detection of cadmium content in plant tissues
The Cd content in transgenic and WT tobacco was measured after the 1 month CdCl2 treatment. Leaf or root samples were washed in distilled water three times, and then dried at 105 °C for 30 min and then at 80 °C for 48 h. The dry tissues were ground and digested in HNO3/HClO4 (4:1, v/v) at 100 °C for 20 min, and then heated at 190 °C for 60 min until the liquid evaporated. The digest was dissolved in deionized water and the Cd2+ content was measured using an Agilent 7500 ICP-MS instrument (Agilent Technologies Inc, USA). The following formula was used to calculate Cd content: (C × 0.1L)/M, where C is the Cd concentration detected by the instrument and M is sample dry weight (g). The Cd transfer coefficient was calculated as follows: Cd content in leaves/Cd content in roots.
Statistical analysis
Data were statistically analyzed by single-factor ANOVA using SAS software (SAS Institute, Cary, NC, USA; PROC ANOVA). The gene effect on tested parameters were evaluated by t test and least significant difference (LSD) at 0.05 and 0.01 probability levels.
Results
Cloning of PtPCS and construction of expression vector
The full-length 1512-bp PtPCS cDNA was amplified from poplar (P. tomentosa) cDNA using a pair of adaptor primers incorporating Hind III and EcoR I sites. The cloned fragment was confirmed by sequencing. The pEZR(K)-LC expression plasmid was double-digested with restriction enzymes, and then ligated as the recombinant expression vector pEZR(K)-LC-PtPCS in which the GFP fragment of pEZR(K)-LC was replaced by PtPCS (Fig. 1). A transformed A. tumefaciens clone was identified using the plasmid double-digestion test (Fig. 2a).
Identification of the recombinant expression plasmid pEZR(K)-LC-PtPCS. Lane 1 in a undigested plasmid of pEZR(K)-LC-PtPCS; Lane 2 in a double digested plasmid of pEZR(K)-LC-PtPCS with Hind III and EcoR I; Lane 3 in b PCR amplification of the plasmid pEZR(K)-LC-PtPCS with PtPCS specific primer; M, 1 kb DNA Marker (Thermo Fisher Scientific Inc. USA)
Screening of transgenic plants
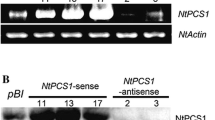
The leaves of tobacco were cut and infected with A. tumefaciens strain LBA4404 containing the recombinant plasmid pEZR(K)-LC-PtPCS. After initial culturing in the dark, differentiation, and regeneration, three regenerated healthy seedlings were obtained and were confirmed to contain PtPCS transcripts by semi-quantitative RT-PCR analysis (Fig. 3).
Phenotyping of transgenic tobacco lines under cadmium stress
After a 1 month Cd treatment, WT plants showed slower growth and wilting of lower leaves, while the transgenic plants showed better growth performance (Fig. 4a). The WT plants also showed poor root performance, embodied in much more brown senile roots, one-third to a half of the biomass when compared to the transgenic plants. The 124-mm root length of WT was just two-thirds that of T12 and T18 (Fig. 4b). Among the transgenic lines, T8 was more sensitive to Cd than were the other two lines, and showed some symptoms of Cd toxicity. All of the transgenic lines were more Cd tolerant than were WT plants.
Physiology of transgenic plants in response of cadmium stress
The physiological and biochemical parameters of the transgenic and WT plants were measured before and after the 1 month Cd treatment. A variance analysis showed that there were no significant differences between transgenic and WT lines before the Cd treatment, while the transgenic lines showed significant differences from WT plants after the Cd treatment in four leaf parameters and root activity tested (Fig. 5). After the Cd treatment, the REL, MDA content, and SOD activity of leaves in transgenic line T12 were 39.19 %, 17.63 nmol g−1, and 679.64 unit g−1, respectively, compared with 45.80 %, 43.90 nmol g−1, and 461.64 unit g−1 in WT (Fig. 5a–d). Compared with root activity after Cd stress treatment, the range of value in transgenic plants was from 463.6 to 657.7 μg TTF g−1 DW h−1, which had a significant difference with 320.2 μg TTF g−1 DW h−1 in WT (Fig. 5e).The results of the physiological analyses indicated that expression of PtPCS strongly enhanced the Cd-tolerance of the transgenic lines, consistent with their morphological appearance.
Physiological analyses of transgenic (T8, T12, T18) and wild-type tobacco plants before and after a 1 month cadmium treatment. The physiological parameters were measured with the leaf or root tissues sampled before or after cadmium treatment (90 μmol CdCl2 per pot) for 1 month. Data are expressed as mean ± SD of six independent experiments. Asterisk and double asterisk indicates values that differ significantly from WT at P < 0.05 and P < 0.01 according to LSD t test (SAS PROC ANOVA). a The leaf Relative electrolyte leakage (REL); b The leaf MDA content; c The leaf SOD content; d The leaf Chlorophyll content; e The root activity
Cadmium content in transgenic lines and wild type
After a 1 month Cd treatment, the Cd content was higher in transgenic lines than in WT (Table 1). This result suggested that expression of PtPCS resulted in enhanced Cd accumulation in tobacco plants. The Cd content in leaves of transgenic lines was 1.24- to 2.28-fold than that in leaves of WT, and the Cd content in roots of transgenic lines was 1.7- to 3-fold than that in roots of WT. In both WT and transgenic plants, the Cd concentration was higher in roots than in leaves. The Cd transfer coefficient was markedly lower in transgenic lines than in WT plants..
Discussion
Phytochelatins can chelate heavy metal ions in the cytoplasm, thereby reducing their toxic effects and protecting normal cellular metabolism (Shukla et al. 2012). Some PC-heavy metal complexes accumulate in the vacuole through an ATP-dependent process (Park et al. 2012; Lin et al. 2012). This effectively stores heavy metals away from sensitive cellular targets (Vögeli-Lange and Wagner 1990; Salt and Rauser 1995; Li et al. 1997; Benavides et al. 2005; Migocka et al. 2011). Consequently, PCS, which encodes the key PC biosynthesis enzyme, is thought to be a key gene in tolerance and accumulation of heavy metal ions in plant tissues. In the present study, transgenic plants overexpressing PtPCS from poplar showed enhanced growth performance under Cd stress in comparison with WT tobacco plants, or higher Cd tolerance based on the physiological analysis. The transgenic plants also showed the higher accumulation of Cd, up to 2.28-fold or 3-fold than that in leaves or roots of WT. Some other reports also provided the evidences for the PCS function. For example, the transgenic tobacco plants expressing the AtPCS gene from Arabidopsis were with 2.2–2.75 times longer roots and double Cd content than those of WT after cadmium treatment (Pomponi et al. 2006). The transgenic tobacco expressing CdPCS1 from Ceratophyllum demersum accumulated Cd to a level 6-fold higher than that in WT when grown medium with Cd (300 μM) (Shukla et al. 2012, 2013). The differences in heavy metal tolerance and accumulation level among various transgenic plants might be due to the sequence variation of PCS genes from various sources (Zhao et al. 2014), and/or the differences in downstream Cd-PC complexes processing (Gasic and Korban 2007b). Recently, a study on Pid3 orthologs in rice revealed that the sequence variation caused the different resistance responses to Magnaporthe oryzae (Xu et al. 2014). It suggested that the extensive mining of PCS orthologs would be valuable for genetic engineering programs aimed at producing effective phytoremediation. Also, the research here mentioned that the transfer coefficient of Cd from underground to ground was markedly lower in transgenic lines than in WT. The similar results were also reported by Pomponi et al. (2006). These findings suggest that PtPCS functions were limited in Cd tolerance and accumulation, but not in Cd transport.
Even all the transgenic lines showed better cadmium tolerance and accumulation in comparison with wild type plants, a little difference was revealed between T8 and the transgenic line. Obviously, T8 accumulated the higher Cd concentration in leaf (about 1.75 folds) and root (about 1.84 folds). It suggested that there might be an extra copy of the gene in the T8 line. RT-PCR results also provide the evidence of that (Fig. 3). The high level of Cd accumulation in the tissues may have negatively affected growth. This could explain why the T8 line was more sensitive to Cd than other transgenic lines, as reflected by its poor growth and physiological appearance. These results suggested a threshold Cd concentration existed in plant cells. Excess value would be adversely harmful for plant growth. Therefore, Cd tolerance and accumulation, as well as Cd transport, should be considered as a whole in genetic engineering programs aimed at producing effective phytoremediation.
Conclusions
The present work showed that the overexpression of PtPCS gene in tobacco not only improves the tolerance but also increases Cd accumulation in plants relative to wild type plants, even excess accumulation of the cadmium might be harmful to plant growth. It suggested PtPCS as a candidate gene for heavy metal bioremediation or phytoremediation.
References
Benavides MP, Gallego SM, Tomaro ML, Braz J (2005) Cadmium toxicity in plants. Braz J Plant Physiol 17:21–34
Bernarda C, Roosensa N, Czernicb P, Lebrunb M, Verbruggen N (2004) A novel CPx-ATPase from the cadmium hyperaccumulator Thlaspi caerulescens. FEBS Lett 569:140–148
Bhuiyan MSU, Min SR, Jeong WJ, Sultana S, Choi KS, Lee Y, Liu JR (2011a) Overexpression of a yeast cadmium factor 1 (YCF1) enhances heavy metal tolerance and accumulation in Brassica juncea. Plant Cell Tissue Organ Cult 105:85–91
Bhuiyan MSU, Min SR, Jeong WJ, Sultana S, Choi KS, Song WY, Lee Y, Lim YP, Liu JR (2011b) Overexpression of AtATM3 in Brassica juncea confers enhanced heavy metal tolerance and accumulation. Plant Cell Tissue Organ Cult 107:69–77
Brunetti P, Zanella L, Proia A, De Paolis A, Falasca G, Altamura MM, Sanità di Toppi L, Costantino P, Cardarelli M (2011) Cadmium tolerance and phytochelatin content of Arabidopsis seedlings over-expressing the phytochelatin synthase gene AtPCS1. J Exp Bot 62:5509–5519
Clemens S, Kim EJ, Neumann D, Schroeder JI (1999) Tolerance to toxic metals by a gene family of phytochelatin synthases from plants and yeast. EMBO J 18:3325–3333
Clemens S, Bloss T, Vess C, Neumann D, Nies DH, Zur Nieden U (2002) A transporter in the endoplasmic reticulum of Schizosaccharomyces pombe cells mediates zinc storage and differentially affects transition metal tolerance. J Biol Chem 277:18215–18221
Cobbett CS (2000) Phytochelatin biosynthesis and function in heavy-metal detoxification. Curr Opin Plant Biol 3:211–216
Curie C, Cassin G, Couch D, Divol F, Higuchi K, Jean ML, Misson J, Schikora A, Czernic P, Mari S (2009) Metal movement within the plant: contribution of nicotianamine and yellow stripe 1-like transporters. Ann Bot-Lond 103:1–11
DalCorso G, Farinati S, Maistri S, Furini A (2008) How plants cope with cadmium: staking all on metabolism and gene expression. J Integr Plant Biol 50:1268–1280
Franchi N, Piccinni E, Ferro D, Basso G, Spolaore B, Santovito G, Ballarin L (2014) Characterization and transcription studies of a phytochelatin synthase gene from the solitary tunicate Ciona intestinalis exposed to cadmium. Aquat Toxicol 152:47–56
Gasic K, Korban SS (2007a) Expression of Arabidopsis phytochelatin synthase in Indian mustard (Brassica juncea) plants enhances tolerance for Cd and Zn. Planta 225:1277–1285
Gasic K, Korban SS (2007b) Transgenic Indian mustard (Brassica juncea) plants expressing an Arabidopsis phytochelatin synthase (AtPCS1) exhibit enhanced As and Cd tolerance. Plant Mol Biol 64:361–369
Glaeser H, Coblenz A, Kruczek R, Ruttke I, Ebert-Jung A, Wolf K (1991) Glutathione metabolism and heavy metal detoxification in Schizosaccharomyces pombe. Curr Genet 19:207–213
Grill E, Winnacker EL, Zenk MH (1985) Phytochelatins: the principal heavy-metal complexing peptides of higher plants. Science 230:674–676
Hasegawa I, Terada E, Sunairi M, Wakita H, Shinmachi F, Noguchi A, Nakajima M, Yazaki J (1997) Genetic improvement of heavy metal tolerance in plants by transfer of the yeast metallothionein gene (CUP1). In: Ando T, Fujita K, Mae T, Matsumoto H, Mori S, Sekiya J (eds) Plant nutrition for sustainable food production and environment. Kluwer Academic Publishers, Dordrecht, pp 391–395
He J, Li H, Luo J, Ma C, Li S, Qu L, Gai Y, Jiang X, Janz D, Polle A, Tyree M, Luo ZB (2013) A transcriptomic network underlies microstructural and physiological responses to cadmium in Populus × canescens. Plant Physiol 162:424–439
He J, Li H, Ma C, Zhang Y, Polle A, Rennenberg H, Cheng X, Luo ZB (2015) Overexpression of bacterial γ-glutamylcysteine synthetase mediates changes in cadmium influx, allocation and detoxification in poplar. New Phytol 205:240–254
Heiss S, Wachter A, Bogs J, Cobbett C, Rausch T (2003) Phytochelatin synthase (PCS) protein is induced in Brassica juncea leaves after prolonged Cd exposure. J Exp Bot 54:1833–1839
Lee S, Moon JS, Ko TS, Petros D, Goldsbrough PB, Korban SS (2003) Overexpression of Arabidopsis phytochelatin synthase paradoxically leads to hypersensitivity to cadmium stress. Plant Physiol 131:656–663
Lee S, Kim YY, Lee Y, An G (2007) Rice P1B-type heavy-metal ATPase, OsHMA9, is a metal efflux protein. Plant Physiol 145:831–842
Li ZS, Lu YP, Zhen RG, Szczypka M, Thiele DJ, Rea PA (1997) A new pathway for vacuolar cadmium sequestration in Saccharomyces cerevisiae: YCF1-catalyzed transport of bis(glutathionato) cadmium. Proc Natl Acad Sci 94:42–47
Li J, Guo J, Xu W, Ma M (2006) Enhanced cadmium accumulation in transgenic tobacco expressing the phytochelatin synthase gene of Cynodon dactylon L. J Integr Plant Biol 48:928–937
Lin YF, Aarts MG (2012) The molecular mechanism of zinc and cadmium stress response in plants. Cell Mol Life Sci 69:3187–3206
Liu GY, Zhang YX, Chai TY (2011) Phytochelatin synthase of Thlaspi caerulescens enhanced tolerance and accumulation of heavy metals when expressed in yeast and tobacco. Plant Cell Rep 30:1067–1076
Liu YX, Wang XT, Su XD, Dai L, Zhang ZY, Xu JC (2012) Cloning and expression analysis of a phytochelatin synthase gene (PtPCS) in Populus tomentosa Carr. Mol Plant Breed 10:174–183 (in Chinese)
Martínez M, Bernal P, Almela C, Vélez D, García-Agustín P, Serrano R, Navarro-Aviñó J (2006) An engineered plant that accumulates higher levels of heavy metals than Thlaspi caerulescens, with yields of 100 times more biomass in mine soils. Chemosphere 64:478–485
Meyer CL, Peisker D, Courbot M, Craciun AR, Cazalé AC, Desgain D, Schat H, Clemens S, Verbruggen N (2011) Isolation and characterization of Arabidopsis halleri and Thlaspi caerulescens phytochelatin synthases. Planta 234:83–95
Migocka M, Papierniak A, Kosatka E, Kłobus G (2011) Comparative study of the active cadmium efflux systems operating at the plasma membrane and tonoplast of cucumber root cells. J Exp Bot 62:4903–4916
Park J, Song WY, Ko D, Eom Y, Hansen TH, Schiller M, Lee TG, Martinoia E, Lee Y (2012) The phytochelatin transporters AtABCC1 and AtABCC2 mediate tolerance to cadmium and mercury. Plant J 69:278–288
Pence NS, Larsen PB, Ebbs SD, Letham DLD, Lasat MM, Garvin DF, Eide D, Kochian LV (2000) The molecular physiology of heavy metal transport in the Zn/Cd hyperaccumulator Thlaspi caerulescens. Proc Natl Acad Sci 97:4956–4960
Polle A, Klein T, Kettner C (2013) Impact of cadmium on young plants of Populus euphratica and P. × canescens, two poplar species that differ in stress tolerance. New For 44:13–22
Pomponi M, Censi V, Di Girolamo V, De Paolis A, di Toppi LS, Aromolo R, Costantino P, Cardarelli M (2006) Overexpression of Arabidopsis phytochelatin synthase in tobacco plants enhances Cd2+ tolerance and accumulation but not translocation to the shoot. Planta 223:180–190
Rea PA (2007) Plant ATP-binding cassette transporters. Annu Rev Plant Biol 58:347–375
Rea PA (2012) Phytochelatin synthase: of a protease a peptide polymerase made. Physiol Plant 145:154–164
Robinson BH, Mills TM, Petit D, Fung LE, Green SR, Clothier BE (2000) Natural and induced cadmium-accumulation in poplar and willow: implications for phytoremediation. Plant Soil 227:301–306
Salt DE, Rauser WE (1995) MgATP-dependent transport of phytochelatins across the tonoplast of oat roots. Plant Physiol 107:1293–1301
Shukla D, Kesari R, Mishra S, Dwivedi S, Tripathi RD, Nath P, Trivedi PK (2012) Expression of phytochelatin synthase from aquatic macrophyte Ceratophyllum demersum L. enhances cadmium and arsenic accumulation in tobacco. Plant Cell Rep 31:1687–1699
Shukla D, Kesari R, Tiwari M, Dwivedi S, Tripathi RD, Nath P, Trivedi PK (2013) Expression of Ceratophyllum demersum phytochelatin synthase, CdPCS1, in Escherichia coli and Arabidopsis enhances heavy metal (loid)s accumulation. Protoplasma 250:1263–1272
Thomine S, Wang R, Ward JM, Crawford NM, Schroeder JI (2000) Cadmium and iron transport by members of a plant metal transporter family in Arabidopsis with homology to Nramp genes. Proc Natl Acad Sci 97:4991–4996
Vatamaniuk OK, Mari S, Lu YP, Rea PA (1999) AtPCS1, a phytochelatin synthase from Arabidopsis: isolation and in vitro reconstitution. Proc Natl Acad Sci 96:7110–7115
Vatamaniuk OK, Mari S, Lang A, Chalasani S, Demkiv LO, Rea PA (2004) Phytochelatin synthase, a dipeptidyltransferase that undergoes multisite acylation with γ-glutamylcysteine during catalysis: stoichiometric and site-directed mutagenic analysis of Arabidopsis thaliana PCS1-catalyzed phytochelatin synthesis. J Biol Chem 279:22449–22460
Vögeli-Lange R, Wagner GJ (1990) Subcellular localization of cadmium and cadmium-binding peptides in tobacco leaves: implication of a transport function for cadmium-binding peptides. Plant Physiol 92:1086–1093
Vrbová M, Kotrba P, Horáček J, Smýkal P, Švábová L, Větrovcová M, Smýkalová I, Griga M (2013) Enhanced accumulation of cadmium in Linum usitatissimum L. plants due to overproduction of metallothionein α-domain as a fusion to β-glucuronidase protein. Plant Cell Tissue Organ Cult 112:321–330
Wang F, Wang Z, Zhu C (2012) Heteroexpression of the wheat phytochelatin synthase gene (TaPCS1) in rice enhances cadmium sensitivity. Acta Biochim Biophys Sin 44:886–893
Xu X, Lv QM, Shang JJ, Pang ZQ, Zhou ZQ, Wang J, Jiang GH, Tao Y, Xu Q, Li XB, Zhao XF, Li SG, Xu JC, Zhu LH (2014) Excavation of Pid3 orthologs with differential resistance spectra to Magnaporthe oryzae in rice resource. PLoS One 9:e93275
Yadav SK (2010) Heavy metals toxicity in plants: an overview on the role of glutathione and phytochelatins in heavy metal stress tolerance of plants. S Afr J Bot 76:167–179
Zacchini M, Pietrini F, Mugnozza GS, Iori V, Pietrosanti L, Massacci A (2009) Metal tolerance, accumulation and translocation in poplar and willow clones treated with cadmium in hydroponics. Water Air Soil Pollut 197:23–34
Zhao C, Xu J, Li Q, Li S, Wang P, Xiang F (2014) Cloning and characterization of a Phragmites australis phytochelatin synthase (PaPCS) and achieving Cd tolerance in tall fescue. PLoS One 9:e103771
Acknowledgments
The authors wish to thank Beijing Natural Science Foundation (#5122019) for funding support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Chen Yongkun and Liu Yuxia have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Chen, Y., Liu, Y., Ding, Y. et al. Overexpression of PtPCS enhances cadmium tolerance and cadmium accumulation in tobacco. Plant Cell Tiss Organ Cult 121, 389–396 (2015). https://doi.org/10.1007/s11240-015-0710-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-015-0710-x