Abstract
In the present study, it is attempted to scrutinize the hydrogen bonding interaction between Carmustine drug and DNA pyrimidine bases by means of density functional theory calculations regarding their geometries, binding energies, vibrational frequencies, and topological features of the electron density in the gas phase and the water solution. Based on the density functional theory results, it is found that the process of intermolecular interaction between Carmustine drug and nucleobases is exothermic and all of the optimized configurations are stable. Furthermore, the negative stability energy represented by a polarizable continuum model shows the significant increase in the solubility of the nucleobase after hydrogen bonding intermolecular interaction in the presence of water solvent. It is also found that the intermolecular hydrogen bonds between drug and the nucleobases play the significant role in the stability of the physisorption configurations. Hydrogen bond energies for hydrogen-bonded complexes are obtained from Espinosa method and the atoms-in-molecules theory are also applied to get a more precise insight into the nature of the intermolecular hydrogen bond interactions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the recent years, the interaction of different kinds of anticancer drugs and nucleic acid base pairs of ribonucleic acid (RNA) and deoxyribonucleic acid (DNA) is a subject of research interest for the rational development of new diagnostics and chemotherapeutics. It is known that the interaction between drugs and DNA and RNA cases the changes in the conformation of the DNA and RNA backbone [1,2,3,4] as well as it affects the biological activity of antitumor, antiviral drugs, and some carcinogenic compounds. Various computational studies on the intermolecular interaction of the nucleobases with small molecule such as cis-Platine [5,6,7,8,9,10], 5-fluorouracil [11,12,13,14,15], ions [16,17,18,19,20,21,22,23,24], and cluster [25,26,27,28] have been reported.
The nucleotides bases such as adenine (A), thymine (T), cytosine (C), guanine (G), and uracil (U) as the pyrimidine derivatives are the standard building blocks for DNA molecules. Pyrimidine is an aromatic heterocyclic organic compound considered as a fundamental unit for all of living cells. The nucleotides bases contain many consecutive hydrogen bond donor and acceptor groups, which make them ideal for studying hydrogen bond (HB) interactions [29].
Currently, the investigation of hydrogen bond in the areas of medicinal chemistry can provide insight into the antioxidant and antiradical activities of some compounds and help sophisticated drug and material design [30,31,32,33,34].
In the current study, we analyzed Carmustine (BNU) drug interaction with the DNA pyrimidine bases, namely cytosine, thymine, and uracil using density functional theory (DFT) calculations. BNU is an alkylating agent which broadly applied in the cure of brain tumors [35,36,37]. Carmustine drug is a non-planar molecule with the amino and hydroxyl groups that is expected to establish effective HB interaction with the nucleobases. We are interested in demonstrating the change of the geometry structures of the monomers as well as the structural stability and energy properties of the base-BNU complexes.
Computational methods
Using density function theory as implemented in Gaussian 03 [38], the mechanism of intermolecular interaction between BNU drug and pyrimidine bases, i.e., cytosine, thymine, and uracil, is investigated by employing the coulomb attenuating method (CAM-B3LYP) functional [39, 40], along with 6-31++G(d,p) basis set. CAM-B3LYP is a long-range corrected version of B3LYP, which includes exact Hartree–Fock exchange at long distances. The calculations are performed in both gas phase and water solution, which the water environment is designed by using the polarizable continuum model [41, 42]. Harmonic vibrational frequencies are estimated to evaluate the zero-point vibrational energy (ZPVE) corrections and also to confirm the nature of the stationary points.
The interaction characteristics of BNU with the pyrimidine bases are obtained by determining the preferred interaction geometries and their corresponding binding energies (ΔE) calculated from the below relation:
where EBNU/B is the total energy of the pyrimidine bases with the drug molecule, and EBNU and EB are the energies of free BNU drug and the pyrimidine bases, respectively.
If the value of ∆E is negative, the complex is more stable relative to the precursors. The binding energies of the complexes are corrected separately, for both zero-point energy and basis set superposition error (BSSE) using the counterpoise correction scheme outlined by the Boys–Bernardi counterpoise technique [43].
In this study, the hydrogen bond energy (EHB) is evaluated by means of Espinosa method. In this method, the HB energies could be estimated from the properties of bond critical points (BCPs). In order to investigate the properties of the bond critical points, their electron densities (ρ), and their Laplacians (∇2ρ), atoms-in-molecule (AIM) calculations are carried out using the AIM2000 program [44]. The simple relationship between HB energy and the potential energy density VBCP at the critical point corresponding to the intermolecular hydrogen bond contacts is assigned to be EHB = 1/2 VBCP [45,46,47,48,49,50].
Results and discussion
Structure and stability of base-BNU complexes
The molecular electrostatic potential (MEP) [51] as a helpful criterion in research of molecular structure with its physiochemical property relationship is evaluated based on the potentials created in the space around a molecule by its nuclei and electrons. The MEP surfaces of the single pyrimidine bases and drug molecule are shown in Fig. 1. It is obvious that there is an extensive region of negative electrostatic potential around the oxygen and nitrogen atoms of the bases, whereas the regions around hydrogen atoms in the case of the considered bases have positive electrostatic potential (see Fig. 1). On the other hand, as shown by the MEP plot in Fig. 1, in BNU molecule, the oxygen atoms are negatively charged (red colors), while the hydrogen atoms are positively charged (blue colors). The negative regions of MEP shown in red correspond to the electrophilic reactivity and the positive regions shown in blue are responsible for nucleophilic reactivity. Therefore, hydrogen bonding in complexes formed between nucleobases and BNU drug is expectable.
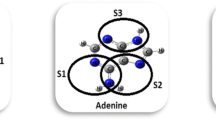
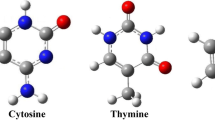
To obtain the most appropriate structure of each complex, different initial orientations of the nucleobases with respect to the BNU drug are studied and after these initial optimizations and extracting their energies, the most stable complexes are considered. For the notations of the intermolecular interaction of drug molecule with the selected bases, we use Tn-BNU, Cn-BNU, and Un-BNU (where n offering multiple sites for hydrogen bond formation: S1, S2, S3, S4 for BNU, and S1, S2, S3 for C, U, and T as shown in Fig. 2).
The results of the optimized structural parameters for the studied complexes in the gas phase are given in Tables 1, 2, and 3. Table 4 illustrates the interaction (Eint), deformation (Edef), and binding energies as well as the stretching frequencies (∆ν) for the studied structures. Also, the actual calculations are performed for water solution and the calculated results are collected in ST1–ST4, supplementary material. For solvation studies, water with dielectric constant of 78.4 is taken as solvating media as it mimics human biological system.
The results indicate that the binding energy values for U-BNU, C-BNT, and T-BNU complexes in the gas phase are ranging from − 14.13 to − 43.80 kJ/mol, from − 22.25 to − 51.06 kJ/mol, and from − 15.97 to − 43.20 kJ/mol at CAM-B3LYP/6-311G++(d,p) level of theory, respectively. Moreover, the equilibrium distances between the drug molecule and bases are in the range of 1.844–2.564 Å, 1.884–2.521 Å, and 1.873–2.545 Å for U-BNU, C-BNT, and T-BNU complexes, respectively.
The negative binding energy values indicate that the intermolecular interaction of BNU drug with pyramidine bases is energetically favorable. According to the obtained results, it is found that C-BNU complexes presented more stability in comparison to U-BNU and T-BNU complexes. In most cases, stronger interactions well correlate with shorter bonds.
Based on the obtained results, it can be stated that the stability of the studied base-BNU complexes in the water solution is decreased in comparison to the gas phase (see Tables 4 and ST4, supplementary material). Close inspection of the calculated equilibrium distances in the intermolecular interaction sites shows that the intermolecular distances between drug and the base molecules increased from gas phase to the water solution (see Tables 1, 2, and 3 and ST1–ST3, supplementary material). These results confirm that the intermolecular bond strength in solution phase is weaker than that in the gas phase. Our computational study also reveals that the geometries of the base-BNU complexes do not change considerably when solvent effects are taken into account (see Tables ST1–ST3, supplementary material).
Furthermore, the interaction energy values of base-BNU complexes in the gas phase are in the range of − 28.21 to − 54.29 kJ/mol, − 27.07 to − 64.25 kJ/mol, and − 20.16 to − 53.40 kJ/mol for the U-BNU, C-BNU, and T-BNU complexes, respectively. The obtained interaction energy values of the HB formed complexes between BNU and bases confirm that there is the weak intermolecular interaction between pyramidine bases and drug molecule. It is also found that by increasing the intermolecular interactions, Eint value increases from − 20.16 kJ/mol to the maximum value of − 64.25 kJ/mol. Besides, the deformation energies are also analyzed as the significant factor in determining the stabilities of the complexes. It is observed no significant perturbation in the relaxed geometry of the considered segments during the intermolecular interactions (see Table 4).
Close inspection of Table 4 reveals that the relative stability of the studied complexes in the gas phase is
-
C2-BNU > C3-BNU > C1-BNU > C4-BNU
-
U2-BNU > U4-BNU > U3-BNU > U5-BNU > U1-BNU > U6-BNU
-
T2-BNU > T4-BNU > T3-BNU > T1-BNU > T5-BNU
The obtained results show that the intermolecular hydrogen bond lengths and the stability of the considered structures are greatly dependent on the nature of the different multiple sites of BNU and base monomers for the intermolecular hydrogen bond formation. Also, since uracil base presents more symmetry constraint with respect to thymine and cytosine, this base has more freedom to choose proper interaction site; therefore, more stable structures are observed for the interaction of uracil with BNU drug molecule, as shown in Fig. 3.
The stability energy (Estab) for all studied geometries has been calculated according to the following equation:
where Ein solvent and Ein gas are the total energies of the considered base-BNU complexes in the water solution and the gas phase, respectively.
According to Table ST4, supplementary material, negative values of the stability energies show that the salvation is a spontaneous process and also signifies the stability of the considered complexes in the aqueous medium.
According to the obtained computational results, it is found that the most negative binding energy value belongs to the C2-BNU complex, so it is the most stable complex among the base-BNU complexes. In this structure, the hydrogen and oxygen atoms of cytosine base are participated as the donor and acceptor to form two intermolecular hydrogen bonds with BNU drug, consisting of H17…O31 and N7…H25 contacts with equilibrium distances of 1.825 and 2.156 Å, respectively. The comparison between the geometrical parameters of monomer and base-BNU complex shows that the HBs formed between cytosine nucleic acid base and the pharmaceutical agent BNU change the monomer bond lengths and angles. The bond length elongations at C2-BNU structure are 1.224 and 1.012 Å for carbonyl group and H25-N29 terminal group of cytosine base, respectively. Similarly, enlargement of N7=O4 and N6-H17 bond lengths of BNU drug indicates the possibility of formation of the intermolecular HBs in this complex. With respect to the geometrical results, the negligible decrease in the bond angles of the base and drug monomers is observed after complexation. For example, the average decreases ∠N28-C30=O31 and ∠C22-N29-H25 at cytosine base and ∠H17-N6-C10 and ∠O4-N7-N5 for BNU drug are 1.4°, 0.9°, 4.2°, and 1.1°, respectively.
The next closest binding energy is achieved with C3-BNU complex with − 50.94 kJ/mol. The difference between the binding energies of C2-BNU and C3-BNU geometries is only 0.12 kJ/mol. As can be seen from Fig. 3, this minimum energy structure is mainly dominated by the three conventional hydrogen bonds (N7-H34…N32, N28…H17-N6, and O4….H34-N32) as well as one nonconventional HB (C9-H16….O31). The hydrogen bond angles are 169.5°, 157.5°, 155.5°, and 152.3° for N7-H34…N32, N28…H17-N6, O4….H34-N32, and C9-H16….O31, respectively. Close inspection of Table 3 reveals that N7…H34-N32 intermolecular interaction has larger angle and shorter HB length (169.5° and 2.113 Å) compared with other intermolecular hydrogen bonds in C3-BNU complex. In this intermolecular interaction, the hydrogen atom of cytosine plays the role of the acceptor of the electron pair, whereas the nitrogen atom of BNU drug is an electron donor.
Additionally, the most stable uracil-BNU complex is U2-BNU system which the oxygen atom of uracil as a Lewis base interacts with the H atom of the drug molecule as a Lewis acid. Furthermore, it is observed that from the interaction of the oxygen atom of BNU drug as a proton acceptor with the hydrogen atom of uracil base as a proton donor, the O…H containing HB complex is obtained. It is found that U2-BNU structure is stabilized by two H-bonds of the O…H-N type, i.e., N6-H17…O31 and N29-H28…O4 with the O…H distances of 1.917 and 2.055 Å as well as the NHO bond angles of 155.7° and 173.6°, respectively.
The structural analysis of the T2-BNU complex indicates two hydrogen bonds which are observed between BNU and thymine nucleobase: one is formed by the O19 atom of N=O terminal group of the drug molecule with H7 atom of thymine base, with O···H distance being 2.115 Å and the other one is formed between O13 atom of thymine and H32 atom of BNU molecule, with O···H distance being equal to 1.903 Å. Based on these results, it is found that BNU drug molecule form a strong binary H-bonded complex with thymine base which are not only a proton donor but also a proton acceptor. It is noted that the N21-H32…O13 and N14-H7…O19 intermolecular HB angles of T2-BNU complex equal to 161.7° and 168.5°, respectively. The binding energy obtained for the most stable T-BNU and U-BNU complexes is about − 43 kJ/mol. It is noted that uracil and thymine have the similar structures. In other words, thymine nucleobase is C5-methyl derivative of uracil.
A closer look at the results of Tables 1, 2, and 3 shows that the intermolecular OB…HBNU (H-bond between bases and drug) distances in the most stable base-BNU complexes are clearly shorter than the OBNU…HB ones; as a result, in these complexes, OB…HBNU H-bonds are much stronger than the OBNU…HB ones. The shortest OB…HBNU distance is observed in the C2-BNU complex; therefore, the outcome is a stronger interaction. This result is well agreement with the most negative binding energy value for C2-BNU structure.
The calculated values of electron density, ρ, Laplacian of electron density, ∇2ρ, and total electronic energy density (H) as well as the HB energy values at CAM-B3LYP/6-311G++(d,p) level of theory are listed in Table ST5, supplementary material. As can be seen from Table ST5, supplementary material, there are the significant accumulation of electron density in the regions of OB…HBNU H-bonds interaction in comparison to OBNU…HB ones at the U2-BNU, C2-BNU, and T2-BNU complexes. Furthermore, the values of the obtained HB energy for the OB…HBNU interactions are greater in comparison to OBNU…HB intermolecular contacts (see Table ST5, supplementary material). Close inspection of Table ST5, supplementary material, reveals that the greatest HB energy value belongs to the conjunction between oxygen atom of cytosine base with hydrogen atom of the drug molecule at the C2-BNU complex.
Based on the results of Tables 1, 2, 3, and 4, it is observed that after the intermolecular interaction between bases and BNU drug, the N–H and C=O bond distances involved in H-bond interactions have been slightly increased in comparison with the corresponding values of monomer for all base-BNU complexes. Furthermore, the largest change in N–H and C=O bond length is observed for the most stable structure, i.e., C2-BNU complex, which is not unexpected because there is a stronger interaction between BNU molecule and cytosine base (see Tables 1, 2, and 3). Table 4 presents the shift of stretching frequencies (∆ν) of the N–H bond as a difference between the frequency of the complex and the frequencies of isolated molecules for base-BNU complexes. Inspection of the vibrational frequency shifts shows that complex formation results in a change in vibrational frequency of N–H bonds involved in H-bonding. Generally, it is found that the interaction of base molecules with BNU drug causes the bond length elongation and the frequency shifting for the N–H bonds involved in H-bond interactions. Our theoretical results show the significant shifts are observed for C2-BNU (− 215.15 cm−1), T2-BNU (− 199.33 cm−1), and U2-BNU (− 194.14 cm−1) complexes. Compared with the N-H stretching, vibrational frequency shift in the most stable base-BNU complexes reveals that the greater shift of the N–H vibrational frequency in C2-BNU model implies the high interaction ability between cytosine base and BNU drug molecule.
In order to investigate the stability of the studied base-BNU complexes, the thermodynamic properties of the considered systems are calculated in the gas phase and water solution and the obtained results are listed in Tables 4 and ST4, supplementary material. It is observed that the formation of the most base-BNU complexes is enthalpically favored in both phases, as shown by negative values of the standard enthalpy changes (ΔH). More negative signs for the ΔH of C2-BNU and C3-BNU complexes prove their more exothermic intermolecular interaction nature in the gas phase. However, free Gibbs energy changes (∆G) values are much less negative than the standard enthalpy changes and in most cases have the opposite sign in the gas phase. Furthermore, the large entropy changes (TΔS) are observed during the formation of the hydrogen-bonded complexes (see Tables 4 and ST4, supplementary material). Close inspection of Table 4 reveals that the high negative values of TΔS determine the positive values of ΔG in the most studied complexes in the gas phase. It is observed that the formation of C2-BNU and C3-BNU complexes requires the larger enthalpy than the entropy changes (in other words | ΔH | > |TΔS|). On the other hand, the ΔG values are negative in the gas phase only for C2-BNU and C3-BNU configurations as more stable base-BNU complexes. These results indicate that the formation of hydrogen-bonded complexes between BNU and cytosine base in these structures is spontaneous and thermodynamically favorable (ΔG < 0, ΔS < 0). Therefore, the entropic factor controls the stability of these complexes.
Conclusion
In this work, we performed density functional theory calculations to study the structural and electronic effects of the interaction of BNU molecule with pyrimidine bases (C, T, and U) in the gas phase and water solution. We found that for various interaction structures, the binding energy of studied configurations is negative, showing that the interaction process is exothermic. Also, it is found that the intermolecular hydrogen bond interactions play a serious role in the stabilization of the considered base-BNU complexes. The enhancement in the stability of the considered configurations reflected by the negative values of Estab in water solvent. The estimation of HBs strength derived from the theory of Bader confirms the geometrical and energetics conclusions.
References
Li H, Mei WJ, Xu ZH, Pang DW, Ji LN, Lin ZH (2007) Electrochemistry of a novel monoruthenatedporphyrin and its interaction with DNA. J Electroanal Chem 600:243–250
Sirajuddin M, Ali S, Badshah A (2013) Drug-DNA interactions and their study by UV-visible, fluorescence spectroscopies and cyclic voltammetry. J Photochem Photobiol B 124:1–19
Zhou YL, Li YZ (2004) Studies of interaction between poly (allylamine hydrochloride) and double helix DNA by spectral methods. Biophys Chem 107:273–281
Rubinson MA, Parkinson JA, Evstigneev MP (2015) Entropic binding mode preference in cooperative homo-dimeric drug–DNA recognition. Chem Phys Lett 624:12–14
Baik M-H, Friesner RA, Lippard SJ (2003) Theoretical study of cisplatin binding to purine bases: why does cisplatin prefer guanine over adenine? J Am Chem Soc 125:14082–14092
Chiavarino B, Crestoni ME, Fornarini S, Scuderi D, Salpin J-Y (2013) Interaction of cisplatin with adenine and guanine: a combined IRMPD, MS/MS, and theoretical study. J Am Chem Soc 135:1445–1455
Kothandapani A, Sawant A, Dangeti VSMN, Sobol RW, Patrick SM (2013) Epistatic role of base excision repair and mismatch repair pathways in mediating cisplatin cytotoxicity. Nucleic Acids Res 41:7332–7343
Truong TB, Pham VN (2017) A DFT investigation on interactions between asymmetric derivatives of cisplatin and nucleobase guanine. Chem Phys Lett 680:44–50
Sarmah A, Roy RK (2012) Understanding the preferential binding interaction of aqua-cisplatins with nucleobase guanine over adenine: a density functional reactivity theory based. RSC Adv 3:2822–2830
Jiang B, Zhou L (2011) Theoretical study of anticancer drug trans-[Pd(dmnp)2Cl2] binding to DNA purine bases, phosphate group and amino acid residues. Struct Chem 22:1353–1364
Yamazaki S, Taketsugu T (2012) Nonradiative deactivation mechanisms of uracil, thymine, and 5-fluorouracil: a comparative ab initio study. J Phys Chem A 116:491–503
Deepa P, Kolandaivel P, Senthil K (2008) Interactions of anticancer drugs with usual and mismatch base pairs—density functional theory studies. Biophys Chem 136:50–58
Gustavsson T, Sarkar N, Banyasz Á, Markovitsi D, Improta R (2007) Solvent effects on the steady-state absorption and fluorescence spectra of uracil, thymine and 5-fluorouracil. J Photochem Photobiol 83:595–599
Hokmabady L, Raissi H, Khanmohammadi A (2016) Interactions of the 5-fluorouracil anticancer drug with DNA pyrimidine bases: a detailed computational approach. Struct Chem 27:487–504
Gester RM, Bistafa C, Georg HC, Coutinho K, Canuto S (2014) Theoretically describing the 17O magnetic shielding constant of biomolecular systems: uracil and 5-fluorouracil in water environment. Theor Chem Accounts 133:1424–1432
Kong H, Sun Q, Wang L, Tan Q, Zhang C, Sheng K, Xu W (2014) Atomic-scale investigation on the facilitation and inhibition of guanine tautomerization at Au(111) surface. ACS Nano 8:1804–1808
Salvatore P, Nazmutdinov R, Ulstrup J, Zhang J (2015) DNA bases assembled on the Au(110)/electrolyte interface: a combined experimental and theoretical study. J Phys Chem B 119:3123–3134
Sponer J, Sabat M, Burda JV, Leszczynski J, Hobza P, Lippert B (1999) Metal ions in non-complementary DNA base pairs: an ab initio study of Cu(I), Ag(I), and Au(I) complexes with the cytosine-adenine base pair. J Biol Inorg Chem 4:537–545
Zhao H, Zhou L (2012) A theoretical study on transition state of the antitumor drug: gold(III)dithiocarbamate derivative interaction with cysteine and DNA purine bases. Comput Theor Chem 979:22–32
Zhu W, Luo X, Puah CM, Tan X, Shen J, Gu J, Chen K, Jiang H (2004) The multiplicity, strength, and nature of the interaction of nucleobases with alkaline and alkaline earth metal cations: a density functional theory investigation. J Phys Chem A 108:4008–4018
Shakourian-Fard M, Fattahi A (2012) Theoretical investigation on the structural and electronic properties of complexes formed by thymine and 2′-deoxythymidine with different anions. Struct Chem 23(1):7–28
Rosa M, Corni S, Di Felice R (2012) A density functional theory study of cytosine on Au(111). J Phys Chem C 116:21366–21373
Nakanishia Y, Kitagawaa Y, Shigetab Y, Saitoa T, Matsuic T, Miyachic H, Kawakamia T, Okumuraa M, Yamaguchid K (2009) Theoretical studies on magnetic interactions between Cu(II) ions in salen nucleobases. Polyhedron 28:1945–1949
Bacchus-Montabonel MC (2012) Theoretical study of charge transfer dynamics in collisions of C6+ carbon ions with pyrimidine nucleobases. Eur Phys J D 66:3002–3008
Sarmah A, Kinkar Roy R (2015) Interaction between small gold clusters and nucleobases: a density functional reactivity theory based study. Phys Chem C 119:17940–17953
Cao GJ, Xu HG, Re L, Zheng W (2012) Hydrogen bonds in the nucleobase, gold complexes: photoelectron spectroscopy and density functional calculations. J Chem Phys 136:014305
Benda Z, Szalay PG (2016) Characterization of the excited states of DNA building blocks: a coupled cluster computational study. Phys Chem Chem Phys 18:23596–23606
Wang H, Zeng X, Zhou R, Zhao C (2013) A comparative DFT study on aquation and nucleobase binding of ruthenium (II) and osmium (II) arene complexes. J Mol Model 19:4849–4856
Lagoja IM (2007) Pyrimidine as constituent of natural biologically active compounds. Chem Biodivers 2:1–50
Nguyen HP, Seto NOL, Cai Y, Leinala EK, Borisova SN, Palcic MM, Evans SV (2003) The influence of an intramolecular hydrogen bond in differential recognition of inhibitory acceptor analogs by human ABO(H) blood group A and B glycosyltransferases. J Biol Chem 278:49191–49195
Song Y, Zhang W, Ji H, Zhou Y, Zhu J, Lu J (2001). Zhongguo Yaowu Huaxue Za Zhi 11:311–316
Raissi H, Khanmohammadi A, Yoosefian M, Mollania F (2013) Ab initio and DFT studies on 1-(thionitrosomethylene) hydrazine: conformers, energies, and intramolecular hydrogen-bond strength. Struct Chem 24:1121–1133
Raissi H, Yoosefian M, Mollania F, Khoshkhou S (2013) Electronic structures, intramolecular interactions, and aromaticity of substituted 1-(2-iminoethylidene) silan amine: a density functional study. Struct Chem 24:123–137
Raissi H, Khoshbin Z, Mollania F (2014) The analysis of structural and electronic properties for assessment of intramolecular hydrogen bond (IMHB) interaction: a comprehensive study into the effect of substitution on intramolecular hydrogen bond of 4-nitropyridine-3-thiol in ground and electronic excited state. Struct Chem 25:515–538
Westphal M, Hilt DC, Bortey E, Delavault P, Olivares R, Warnke PC, Whittle IR, Jääskeläinen J, Ram Z (2003) A phase 3 trial of local chemotherapy with biodegradable carmustine (BCNU) wafers (Gliadel wafers) in patients with primary malignant glioma. Neuro-Oncology 5:79–88
Reithmeier T, Graf E, Piroth T, Trippel M, Pinsker MO, Nikkhah G (2010) BCNU for recurrent glioblastomamultiforme: efficacy, toxicity and prognostic factors. BMC Cancer:10–30
Qian L, Zheng J, Wang K, Tang Y, Zhang X, Zhang H, Huang F, Pei Y, Jiang Y (2013) Cationic coree shell nanoparticles with carmustine contained within O6-benzylguanine shell for glioma therapy. Biomaterials 34:8968–8978
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Montgomery JA, Vreven Jr T, Kudin KN, Burant JC, Millam JM, Iyengar SS, Tomasi J, Barone V, Mennucci B, Cossi M, Scalmani G, Rega N, Petersson GA, Nakatsuji H, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Klene M, Li X, Knox JE, Hratchian HP, Cross JB, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Ayala PY, Morokuma K, Voth GA, Salvador P, Dannenberg JJ, Zakrzewski VG, Dapprich S, Daniels AD, Strain MC, Farkas O, Malick DK, Rabuck AD, Raghavachari K, Foresman JB, Ortiz JV, Cui Q, Baboul AG, Clifford S, Cioslowski J, Liu G, Stefanov BB, Liashenko A, Piskorz P, Komaromi I, Martin RL, Fox DJ, Keith T, Al-Laham MA, Peng CY, Nana-yakkara A, Challacombe M, Gill PMW, Johnson B, Chen W, Wong MW, Gonzalez C, Pople JA (2003) Gaussian 03, revision C.02 (or D.01). Gaussian Inc., Pittsburgh
Becke AD (1993) A new mixing of Hartree-Fock and local density functional theories. J Chem Phys 98:1372–1377
Lee C, Yang W, Parr RG (1988) Development of the Colle-Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B 37:785–789
VMiertus S, Scrocco EC, Tomasi J (1981) Electrostatic interaction of asolute with a continuum. A direct utilization of ab initio molecular potentials for the prevision of solvent effects. J Chem Phys 55:117–129
Mennucci B (2012) Polarizable continuum model. Wiley Interdiscip. Rev Comput Mol Sci 2:386–404
Boys SF, Bernardi F (1970) The calculation of small molecular interactions by the differences of separate total energies. Some procedures with reduced errors. Mol Phys 19:553–566
RFW B (1990) Atoms in molecules—a quantum theory. Clarendon Press, Oxford A. E Reed, L A Curtiss and F Weinhold, Chem Rev, 88: 899
Espinosa E, Molins E (2000) Retrieving interaction potentials from the topology of the electron density distribution: the case of hydrogen bonds. J Chem Phys 113:5686–5694
Espionsa E, Souhassou M, Lachekar H, Lecomte C (1999) Topological analysis of the electron density in hydrogen bonds. Acta Crystallogr B 55:563–572
Safdari F, Raissi H, Shahabi M, Zaboli M (2017) DFT calculations and molecular dynamics simulation study on the adsorption of 5-fluorouracil anticancer drug on Graphene oxide nanosheet as a drug delivery vehicle. J Incl Phenom Macrocycl Chem 27:805–817
Shahabi M, Raissi H, Mollania F (2014) Electronic structures, intramolecular hydrogen bond interaction, and aromaticity of substituted 4-amino-3-penten-2-one in ground and electronic excited state. Struct Chem 26:491–506
Kamel M, Raissi H, Morsali A, Shahabi M (2017) Assessment of the adsorption mechanism of flutamide anticancer drug on the functionalized single-walled carbon nanotube surface as a drug delivery vehicle: an alternative theoretical approach based on DFT and MD. Appl Surf Sci, In Press
Shahabi M, Raissi H (2017) Screening of the structural, topological, and electronic properties of the functionalized Graphene nanosheets as potential Tegafur anticancer drug carriers using DFT method, J Biomol Struct Dyn, In Press
Murray JS, Sen K (1996) Molecular electrostatic potentials concepts and applications. Elsevier, Amsterdam
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(DOCX 37 kb)
Rights and permissions
About this article
Cite this article
Khorram, R., Raissi, H. & Shahabi, M. Analysis of the structures, energetics, and vibrational frequencies for the hydrogen-bonded interaction of nucleic acid bases with Carmustine pharmaceutical agent: a detailed computational approach. Struct Chem 29, 1165–1174 (2018). https://doi.org/10.1007/s11224-018-1102-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-018-1102-8