Abstract
The chlorophyllide a oxygenase (CAO) plays a crucial role in the biosynthesis of chlorophyll b (Chl b). In the moss Physcomitrium patens (P. patens), two distinct gene copies, PpCAO1 and PpCAO2, are present. In this study, we investigate the differential expression of these CAOs following light exposure after a period of darkness (24 h) and demonstrate that the accumulation of Chl b is only abolished when both genes are knocked out. In the ppcao1cao2 mutant, most of the antenna proteins associated with both photosystems (PS) I and II are absent. Despite of the existence of LHCSR proteins and zeaxanthin, the mutant exhibits minimal non-photochemical quenching (NPQ) capacity. Nevertheless, the ppcao1cao2 mutant retains a certain level of pseudo-cyclic electron transport to provide photoprotection for PSI. These findings shed light on the dual dependency of Chl b synthesis on two CAOs and highlight the distinct effects of Chl b deprival on PSI and PSII core complexes in P. patens, a model species for bryophytes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oxygenic photosynthesis sustains most life on earth by converting light to chemical energy. Plants evolved various light harvesting complexes (LHCs) to capture sunlight under different conditions. However, excess light can cause photodamage to the photosynthetic apparatus (Perrine et al. 2012; Friedland et al. 2019; Wu et al. 2020). To prevent this, plants employ a rapid photoprotective mechanism called nonphotochemical quenching (NPQ) that dissipates excess energy as heat mostly in LHCs (Müller et al. 2001; Derks et al. 2015; Giovagnetti and Ruban 2018). The moss Physcomitrium patens (P. patens) has a diverse LHC system that includes both algal- and angiosperm-type NPQ pathways, unlike the green alga Chlamydomonas reinhardtii (Chlamydomonas) and the flowering plant Arabidopsis thaliana (Arabidopsis) that have one type each. (Gerotto et al. 2012; Ilik et al. 2017).
Chlorophyll (Chl) b only exists in the LHCs and play critical roles in stabilizing them (Green and Durnford 1996; Horn et al. 2007; Tanaka and Tanaka 2011). Chl b is synthesized from Chl a by the action of chlorophyllide a oxygenase (CAO), a key enzyme with three domains: A (regulatory), B (linker), and C (catalytic) (Oster et al. 2000; Eggink et al. 2004; Nagata et al. 2004; Kunugi et al. 2013). CAO is usually encoded by a single gene in green plants, except for rice which also has a pseudogene (Tanaka et al. 1998; Nagata et al. 2004; Lee et al. 2005; Kunugi et al. 2013; Bujaldon et al. 2017; Jung et al. 2021). In the prasinophyte Micromonas, the Rieske center and mononuclear iron-binding motif of the conserved site in C domain of CAO is encoded by different genes (Kunugi et al. 2013). The CAO knockout in Chlamydomonas and Arabidopsis leads to the absence of Chl b and the reduction of LHCs, especially LHCII. However, some PSII dimer and PSI-LHCI complex remain on the thylakoid membranes of these mutants (Tanaka and Tanaka 2005; Havaux et al. 2007; Kim et al. 2009; Dall’Osto et al. 2010; Takabayashi et al. 2011; Bujaldon et al. 2017). In the genome of P. patens, two complete CAO genes (PpCAO1 and PpCAO2) were detected in two chromosomes (Rensing et al. 2008), with unidentified physiological significance.
In this study, we employed CRISPR-Cas9 technology to knock out both CAO genes, thereby generating a Chl b-less mutant of P. patens, namely ppcao1cao2. We showed that both CAOs were essential for Chl b synthesis in the moss. Notably, the loss of Chl b had a more pronounced impact on the PSII core complexes compared to the PSI core complexes. This discrepancy may be attributed to the diminished NPQ observed in ppcao1cao2, which serves as the primary photoprotective mechanism for PSII. Nonetheless, the mutant displayed evident pseudo-cyclic electron transport, which acts as a safe valve for the acceptor side of PSI.
Experimental procedures
Plant culture, transformation and treatments
Plants were cultivated in a growth chamber under controlled conditions, including a 16 h light: 8 h dark photoperiod, 60% humidity, 23 °C and 50 μmol photons m−2 s−1 light intensity. P. patens Gransden wild-type strain was grown on solidified minimum PpNO3 medium supplemented with 0.5% glucose (Ashton et al. 1979; Pinnola et al. 2013, 2015, 2018; Yan et al. 2021). To disrupt the PpCAO1 and PpCAO2 genes, three target sites (Table S1) were selected for designing sgRNA (single-guide RNA) using the webtool CRISPR V1 against P. patens genome Phytozome V9 (http://crispor.tefor.net/crispor.py). CRISPR/Cas9 constructs were generated following a previously established protocol (Lopez-Obando et al. 2016). Protoplast isolation and transformation were performed using two rounds of 5-day-old moss protonema tissues (Ashton et al. 1979; Schaefer and Zrÿd 1997; Lopez-Obando et al. 2016). Subsequently, genome sequencing confirmed the generation of ppcao1, ppcao2, and ppcao1cao2 mutants.
Thylakoid extraction and gel electrophoresis
Thylakoid membranes were extracted from 10-day-old protonema tissues as described (Gerotto et al. 2012, 2019). Two different techniques, denaturing SDS-PAGE and lpBN-PAGE followed by 2D SDS-PAGE, were employed. For denaturing SDS-PAGE, thylakoids were solubilized with Laemmli buffer and separated by SDS-PAGE. Proteins were stained with Coomassie brilliant blue G250 or transferred to polyvinyl difluoride membranes and immunoblotted with the corresponding antibodies (the D1 (AS05084), D2 (AS06146), CP47 (AS04038), CP43 (AS06110), PSBS (AS09533), PSAB (AS10695), LHCA2 (AS01006), ATP β subunit (AS08370), Cyt b6 (AS03034) and Cyt f (AS06119) were purchased from Agrisera and the LHCBM, LHCSR, LHCB5 and LHCB6 were synthesized by Thermo-Fisher Co. For lpBN-PAGE followed by 2D SDS-PAGE, thylakoid membranes were first solubilized with 1% (w/v) β-DM at a final Chl concentration of 0.5 μg/μL and separated by lpBN gel system (Järvi et al. 2011). Then lanes were cut and solubilized with Laemmli buffer and separated by SDS-PAGE. Finally, the proteins were visualized by either CBB staining or western blotting.
Low-temperature fluorescence emission spectra
The thylakoid membranes were extracted and then rapidly frozen in liquid nitrogen at 20 µg/mL chlorophyll concentration (Casazza et al. 2001). The low temperature fluorescence emission spectra measurements were performed by exciting at 475 nm and recording in the 600–800 nm range. There were three major peaks: 682, 693 and 717 nm, which were attributed to LHCII, PSII core and PSI-LHCI respectively (Pinnola et al. 2015).
In vivo measurements of chlorophyll fluorescence
In vivo chlorophyll fluorescence was monitored with a Dual-PAM-100 (Walz, Germany). Plants were dark-acclimated for 30 min before measurements. The NPQ kinetics using 830 μmol photons m−2 s−1 of actinic light was measured for 10 min with saturating light of 6000 μmol photons m−2 s−1 and then for 10 min in dark.
Estimation of the functional antenna size of PSII and PSI
The relative antenna size of PSI was estimated according to the method described previously (Iwai et al. 2015; Pinnola et al. 2015) by analyzing time courses of P700 photo-oxidation upon illumination of the protonema with far-red light (710 nm, 200 μmol photons m−2 s−1). The kinetics were measured using a LED excitation-detection spectrometer JTS-10 (Bio-logic, France), and the protonema incubated with 30 µM DCMU and 100 µM MV in the dark for 40 min.
The relative antenna size of PSII was estimated by measuring the induction of Chl fluorescence emitted by P. patens protonema infiltrated with 20 µM DCMU for 30 min as described previously (Malkin et al. 1981; Havaux et al. 2007). The Chl fluorescence kinetics were measured using a Plant Efficiency Analyser (Hansatech, UK).
High-performance liquid chromatography
The chlorophyll concentrations were determined following extraction of the pigments in acetone as described previously (Yang et al. 1999) and the pigment compositions were analyzed by HPLC as described (Thayer and Björkman 1990; Qin et al. 2015). Briefly, 0.1 g fresh green tissues were used for extraction in a certain amount of acetone and the final volume and the pigment concentration after each extraction were measured. Each sample of 0.12 µg Chl amount was injected in the HPLC instrument to fit in the quantitative range. The pigment compositions were calculated based on the fresh weight of green tissues.
qRT-PCR
The transcripts were quantified by qRT-PCR using specific primers listed in Table S3. For qRT-PCR analysis, the first strand of cDNA was synthesized from total RNA prepared from 10 day-old protonemata, and qRT-PCR was performed using 2 × HQ SYBR qPCR Mix (High ROX) (ZOMANBIO) with the following thermal cycling program: 95 °C for 30 s, followed by 40 cycles of 95 °C for 10 s, 60 °C for 30 s. The data were analyzed using LightCycler480 Software release 1.5.0 (Roche). The relative gene expression levels were normalized by the Actin gene. qRT-PCR was carried out in duplicate for each sample in three independent experiments.
Statistical analysis
All experiments were repeated at least three times (n ≥ 3). Values are expressed as mean ± SD. One-way ANOVA and Tukey’s multiple comparison test were used to determine the significant difference between different treatments (α = 0.05).
Results
The transcriptional level of two CAO genes
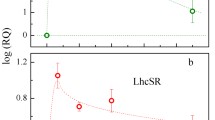
Two CAO homologous genes, PpCAO1 and PpCAO2 (Pp3c19_22390 and Pp3c22_20970 in P. patens genome v3.3: https://phytozome-next.jgi.doe.gov/), in P. patens are detected on chromosome 19 and 22 respectively with 80% homology. Motif analysis shows that both CAO genes possess the characteristic domains found in CAO genes of other species (Figs. S1–2). Phylogenetic analysis shows that the CAO genes of P. patens a occupy an intermediate position between those of green algae and vascular plants, but exhibit closer evolutionary affinity to the latter group (Fig. S3). Expression analysis revealed that both PpCAO1 and PpCAO2 are transcribed under both dark and light conditions (Fig. S4), which differs from the expression pattern observed in rice CAOs (Lee et al. 2005). To investigate the temporal expression profiles, quantitative real-time PCR (qRT-PCR) was performed at different time points (0.5, 1, 2, 4, 6, and 240 h) after 24 h of darkness followed by illumination. The results indicated that the expression level of PpCAO1 remains relatively constant regardless of the duration of light exposure. In contrast, the expression of PpCAO2 closely resembles that of PpCAO1 up to 4 h of light treatment, but shows significant upregulation thereafter (Fig. 1).
The ppcao1cao2 mutant possesses very low PS activities
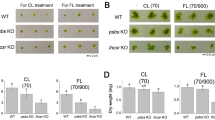
We employed the CRISPR-Cas9 technology to simultaneously knock out PpCAO1 and PpCAO2 genes in order to investigate the consequences of Chl b deprivation. The ppcao1, ppcao2, ppcao1cao2 mutants were identified by sequencing (Fig. S5). Analysis of pigment composition revealed a significant reduction in Chl b level in ppcao1 and ppcao2 than in WT (Fig. S6) Intriguingly, Chl b was completely absent in ppcao1cao2, which also exhibited approximately half the Chl a concentration of WT. Meanwhile, ppcao1cao2 displayed a distinct paler phenotype, accompanied by a substantial decrease in xanthophyll pigments. Importantly, the mutant exhibited a very low Fv/Fm, the maximum quantum yield of PSII, measuring only approximately about two-third of observed in the WT, indicating that strong PSII photoinhibition existed in ppcao1cao2 (Table 1, Fig. S7).
The induction of Chl a fluorescence with the presence of DCMU and P700 photoreduction with the presence of DCMU and MV were measured to reflect the estimated the functional antenna size for PSII and PSI (Malkin et al. 1981; Greene et al. 1988; Iwai et al. 2015; Pinnola et al. 2015; Bujaldon et al. 2017). The functional antenna size of both PSs was severely reduced in ppcao1cao2 compared with WT (Fig. 2a, b).
Analysis of the functional antenna size of PS and the low temperature 77 K fluorescence emission spectra in P. patens WT and the cao double mutant. a Induction curves of Chl fluorescence in tissue of moss infiltrated with 20 μmol DCMU in dark for 40 min. b The kinetics of P700+ formation in the presence of 30 µM DCMU and 100 µM methyl viologen (MV) in dark for 40 min. c, d The 77 K fluorescence emission spectra of P. patens WT and ppcao1cao2 thylakoid membrane that were extracted in the growth light condition before rapidly freezing in liquid nitrogen. The spectra with excitation at 475 nm. All spectra were normalized at 693 nm. Data are expressed as mean ± SD (n ≥ 3)
The low-temperature (77 K) fluorescence emission spectra of the P. patens WT thylakoid membranes had three characteristic peaks, located at 682, 693, and 717 nm, which were attributed to LHCII, PSII core, and PSI-LHCI, respectively, as previously reported (Pinnola et al. 2015). In ppcao1cao2, the peaks corresponding to LHCII and PSII core were significantly diminished, leaving only a prominent peak representing PSI (Fig. 2c, d).
Chl fluorescence and P700 redox analysis showed that ppcao1cao2 had much lower PSII and PSI activities than WT (Fig. S8a, b). To investgated the kinetic of NPQ, we measured the time courses of NPQ of WT and ppcao1cao2 under actinic light of 50 and 830 μmol photons m−2 s−1 (Fig. S8d, Fig. 3). Under both circumstrances, NPQ was clearly induced in WT, but ppcao1cao2 presented very low and plain NPQ kinetics. The ppcao1cao2 presented a much higher Y(ND) and a similar level of Y(NA) compared with WT (Fig. S8c, e), suggesting that the reduction in Y(I) of ppcao1cao2 can be attributed to limitations on the donor side.
The fluctuating light treatment (FL) simulates the unstable light conditions in the natural environment. All genotypes were exposed to FL consisting of a low-light phase (50 μmol photons m−2 s−1 for 5 min) and a high-light phase (500 μmol photons m−2 s−1 for 1 min) using the DUAL-PAM system. ppcao1cao2 possessed quite lower Y(I) and Y(II) than WT since the first cycle (Fig. S9a, b). NPQ increased gradually with FL cycles in WT, but remained very low and insensitive to FL in ppcao1cao2 (Fig. S9c). The reduction in Y(I) of ppcao1cao2 was due to donor side limitation as indicated by high Y(ND) (Fig. S9d). The unchanged Fv/Fm and Pm of WT and ppcao1cao2 before and after fluctuating light respectively indicated that the PSI and PSII were not photodamaged by the whole FL treatment in both genotypes (Fig. S9g, h).
The transcriptional level of genes related to photosynthesis
The expression levels of genes related to photosynthesis in WT and ppcao1cao2 were determined by qRT-PCR. The results revealed that the transcription levels of PsbB, PsbC, PsbD, PsaA, PsaO1, PsbS, LHCSR, FNR1, NdhA, PETE1, AtpA, PetA, PGR5 and Flv genes were upregulated in ppcao1cao2, whereas PsbA, rbcL, Lhca1, and Lhcbm were apparently downregulated in ppcao1cao2 compared with those of the WT (Fig. 4).
The accumulation of thylakoid membrane proteins in ppcao1cao2
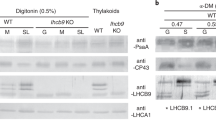
We analyzed the thylakoid proteins composition by SDS-PAGE and immunodetection. We used antibodies for LHCB1, LHCB5 (LHCB6) and LHCA2 representing the major, the minor antenna of PSII and PSI antenna respectively for westernblotting and found that these antenna subunits were almost diminished in ppcao1cao2 (Fig. 5). We also detected the subunits of other complexes on the thylakoid membranes. When thylakoid membrane samples with same amount of Chl a were loaded, more signals for bands representing PSII core subunits, PSI core subunits were observed in ppcao1cao2. LHCSR, an important NPQ player in P. patens, also accumulated more in ppcao1cao2 than in WT (Fig. 5). There are no significant changes in the signals of Cty b6f, ATPase and PsbS compared to WT (Fig. 5).
The organization and composition of the photosystems in ppcao1cao2
After solubilization in β-DM, the thylakoid membranes were separated by lpBN-PAGE to investigate composition of pigment protein complexes and the subunits comprising each protein complex were separated by 2D SDS-PAGE (Fig. 6). In ppcao1cao2, only three bands can be observed in lpBN-PAGE, representing PSI core complex, PSII core complex, and CP43-less PSII core complex, respectively (Fig. 6b). No bands representing PSII supercomplexes, PSII-LHC complex, LHCII trimer and LHCII monomers, which were present in WT, can be seen in ppcao1cao2. To investigate the distribution of PSII core proteins CP43 and CP47, western blotting after 2D SDS-PAGE were employed (Fig. 6c, d). The result showed that most CP47 were detected in the PSII core and CP43-less PSII core but only a small part of CP43 was found in the PSII core and a large amount of CP43 were detected as monomeric proteins in ppcao1cao2 (Fig. 6d).
2D-lpBN-SDS-PAGE of the thylakoid protein complexes in WT and ppcao1cao2. The thylakoid extracts of WT and ppcao1cao2 were firstly separated by lpBN-PAGE. A total of 2.9 μg of Chl a was loaded in each lane. The strips of WT and ppcao1cao2 were furtherly separated by 2D SDS-PAGE and stained with Coomassie brilliant blue G-250 (a, b) or immunoblotted with the antibodies for CP47 and CP43 (c, d)
The P700 re-oxidation capacity of ppcao1cao2
The P700 re-oxidation phenomenon has been observed exclusively in non-flowering plants (Ilik et al. 2017). The P700 re-oxidation capacity was measured by exposure of dark-adapted samples to actinic light (2000 μmol photons m−2 s−1). The initial oxidation of P700 (forming P700+) reflects the intrinsic activity of PSI and the subsequent P700+ reduction (forming P700) reflects the intake of electrons originating in water splitting at PSII, and the final re-oxidation of P700 can be attributed to the outflow of electrons from PSI (Schansker et al. 2003, 2005; Ilik et al. 2017). The result showed that after the initial oxidation of P700, WT presented a complete P700+ reduction before subsequent re-oxidation. In ppcao1cao2, after the initial oxidation of P700, the extent of P700+ reduction was much lower than in WT. notably, a faster re-oxidation rate was observed in ppcao1cao2 than in WT when a 0-1normalization was employed (Fig. S10).
Discussion
The moss P. patens, with detailed genome information and easy transformation methods, has been developed as a model organism in bryophytes (Schaefer 2002). In the wild, Mosses might grow on both sunlight energy and organic chemicals from decomposed biomass under tree canopies where they frequently experience much higher irradiation than aquatic environments (Cove 2005; Way and Pearcy 2012; Iwai and Yokono 2017). Studies have shown that P. patens possesses more family members of genes encoding antenna proteins compared to its green algae and angiosperm counterparts, Chlamydomonas and Arabidopsis (Iwai and Yokono 2017). Furthermore, P. patens persists algal- and angiosperm-type qE pathway to quench excessive energy (Iwai and Yokono 2017) and besides cyclic electron transport (CEF), it possesses very active pseudo-cyclic electron transport (PCEF) to drive active electron to FLV proteins (Yamamoto et al. 2016; Ilik et al. 2017), which is lost in angiosperms.
In green plants, Chl b molecules are only found in LHCs and plays important roles in the folding and function of the LHC complexes (Green and Durnford 1996; Hoober et al. 2007). CAO is the key enzyme that catalyzes the conversion of Chl a to Chl b. The precise localization of CAO and its role in chloroplast protein import remain elusive, because CAO is present at extremely low levels in chloroplasts and is not detectable by immunological or mass spectrometry methods (Tanaka and Tanaka 2019). Our attempt to make antibodies for CAOs in P. patens also failed. All studied species of green plants have only one copy of CAO gene in their genomes except for rice, which has two tandem copies on one chromosome; however, only one copy is active and the other seems to be a pseudogene (Lee et al. 2005; Kunugi et al. 2013; Jung et al. 2021). In P. patens, two copies of CAO genes were found on different chromosomes. We found the transcription of two CAOs was differently regulated upon light exposure (Fig. 1). Using mutants deficient either or both PpCAOs achieved by CRISPR-Cas9 technology (Lopez-Obando et al. 2016), it was revealed that only when both PpCAOs were knock out did Chl b synthesis stop completely in P. patens (Table 1, Fig. S6). Therefore, unlike green algae or vascular plants, P. patens is equipped with two functional CAO genes to guarantee the Chl b synthesis.
Previous studies have shown that Chl b deficiency in green plants specifically impacts the accumulation of LHC apoproteins, while their mRNA levels remain unaffected (Espineda et al. 1999; Nick et al. 2013). Appropriate reduction of Chl b in Chlamydomonas and higher plants can improve the photosynthetic activity and biomass, but excessive reduction of Chl b will lead to decreased photosynthetic efficiency and light tolerance of plants (Mussgnug et al. 2007; Friedland et al. 2019; Wu et al. 2020). In our study, ppcao1cao2, due to Chl b deficiency, has a lower Fv/Fm (Fig. S7). The majority of chlorophyll fluorescence is emitted by PSII, excited by energy captured mostly by LHCII. When LHCII was eliminated in the mutant, the level of fluorescence was very low and the kinetics of fluorescence was steady. We found a lot of PSII core subunits such as CP43 exist in the monomeric form in vivo (Fig. 6), which would enhance the Fo level. We observed a significant decrease in the expression of LHCBM and Lhca1. However almost no antenna proteins accumulated in the thylakoid membranes which means the transcription of antenna protein genes were downregulated when Chl b synthesis was blocked and even there were some mRNAs of antenna proteins, very few antenna proteins were detected because the synthesis of antenna proteins and Chl b might be tightly correlated or the newly-synthesized antenna proteins were rapidly degraded when Chl b was unavailable. As for core complex subunits, we found that the expression of PsbA was decreased, but PsbB, PsbC, PsbD were increased (Fig. 4b), suggesting that the expression of D1 is regulated under a distinct pattern from that of the other PSII core subunits. The expression of PsaA, encoding one major PSI subunit was markedly increased (Fig. 4a), which is consistent with findings in rice cao mutants (Jung et al. 2021). The expression of PsaO and FNR1, encoding two subunits of PSI, were also significantly increased (Fig. 4a). The final protein levels of both core complexes were similar in WT and ppcao1cao2 on an equal Chl a amount basis (Fig. 5). This suggest that unlike antenna proteins, the expression of all detected core complexes subunit genes except PsbA in ppcao1cao2 were enhanced (Fig. 4) and these subunit proteins can be synthesized and inserted into thylakoid membranes (Fig. 5), but a significant percentage of these proteins cannot be assembled into the whole complex and exist as monomeric form (Fig. 6). Moreover, genes encoding key players involved in NPQ, such as PsbS and LHCSR, as well as components related to CEF component PGR5, NdhA and PCEF component Flv were also enhanced in ppcao1cao2 (Fig. 4c, d). Similarly, the expression of PETE1 (encoding the major isoform of plastocyanin), PetA (encoding one cytochrome b6f subunit), AtpA (encoding ATP synthase alpha subunit) were also increased (Fig. 4c). However, the protein levels of subunits from cytochrome b6f and ATP synthase were not increased (Fig. 5), which means that post-transcriptional regulation might exist to repress the protein level of cytochrome b6f and ATP synthase.
Previous studies on Chl b-less mutants of vascular plants or algae have shown that varying abilities of different LHCs to withstand Chl b deprivation. Generally speaking, LHCI is less sensitive to Chl b deprivation than LHCII. In vascular plants, the absence of Chl b reduces the levels of not only LHCII but also some other antenna proteins in chloroplasts to varying degrees (Tanaka and Tanaka 2005; Takabayashi et al. 2011). In Chlamydomonas, both LHCII and LHCI could accumulate to wild-type levels in a CAO mutant if Chl a synthesis remains unaltered (Bujaldon et al. 2017). However, in ppcao1cao2, the protein accumulation of antenna proteins for both photosystems was significantly reduced, as evidenced by SDS-PAGE and western blotting analysis (Figs. 5 and 6) and only PSI and PSII core complexes were observed in lpBN-PAGE (Fig. 6). LHCSR, an essential component in NPQ, does not bind Chl b for its correct folding and LHCSR was also found in diatoms, which lack Chl b (Koziol et al. 2007). The transcriptional expression and protein levels of LHCSR were enhanced in ppcao1cao2 (Figs. 2d and 5). However, NPQ in ppcao1cao2 still cannot develop (Figs. 3, S8d, S9c), likely due to the very low accumulation of LHCBM (Fig. 5). A similar result has been reported in the Chl b-less Cbs3 strain of Chlamydomonas, where LHCSR3 accumulation failed to induce NPQ (Bonente et al. 2011). It had been shown that FLV-dependent PCEF, another fate of electrons past PSI, is especially active in fluctuating light conditions and essential for P. patens as safe valve (Gerotto et al. 2016; Ilik et al. 2017). In our results, although the P700+ reduction was not complete in ppcao1cao2 because of very low PSII activity (Fig. S10a), the kinetics of P700 re-oxidation was even faster than WT (Fig. S10b), partly resulting from higher expression of Flv (Fig. 4d). This pathway might consume active electrons at the acceptor side of PSI, decreasing Y(NA) (Figs. S8e, S9f), keeping PSI from damage by excessive electrons. This could explain why PSI core complexes were more stable than PSII complexes in ppcao1cao2 (Fig. 6).
Conclusions
Both CAOs, PpCAO1 and PpCAO2, are needed to maintain the amount of Chl b in P. patens, in which the existence of Chl b and antenna proteins are tightly correlated. With almost no antenna complexes and NPQ, the P. patens Chl b-less mutant ppcao1cao2 possesses much more functional PSI core complexes than PSII core complexes. The FLV-dependent PCEF activity in ppcao1cao2 might be the major photoprotective process for PSI in the mutant.
Data availability
It is not applicable.
References
Ashton NW, Grimsley NH, Cove DJ (1979) Analysis of gametophytic development in the moss, Physcomitrella patens, using auxin and cytokinin resistant mutants. Planta 144:427–435. https://doi.org/10.1007/bf00380118
Bonente G, Ballottari M, Truong TB et al (2011) Analysis of LhcSR3, a protein essential for feedback de-excitation in the green alga Chlamydomonas reinhardtii. PLoS Biol 9:e1000577. https://doi.org/10.1371/journal.pbio.1000577
Bujaldon S, Kodama N, Rappaport F et al (2017) Functional accumulation of antenna proteins in chlorophyll b-Less Mutants of Chlamydomonas reinhardtii. Mol Plant 10:115–130. https://doi.org/10.1016/j.molp.2016.10.001
Casazza AP, Tarantino D, Soave C (2001) Preparation and functional characterization of thylakoids from Arabidopsis thaliana. Photosynth Res 68:175–180. https://doi.org/10.1023/a:1011818021875
Cove D (2005) The Moss Physcomitrella patens. Annu Rev Genet 39:339–358. https://doi.org/10.1146/annurev.genet.39.073003.110214
Dall’Osto L, Cazzaniga S, Havaux M et al (2010) Enhanced photoprotection by protein-bound vs free xanthophyll pools: a comparative analysis of chlorophyll b and xanthophyll biosynthesis mutants. Mol Plant 3:576–593. https://doi.org/10.1093/mp/ssp117
Derks A, Schaven K, Bruce D (2015) Diverse mechanisms for photoprotection in photosynthesis: dynamic regulation of photosystem II excitation in response to rapid environmental change. Biochem Biophys Acta 1847:468–485. https://doi.org/10.1016/j.bbabio.2015.02.008
Eggink LL, LoBrutto R, Brune DC et al (2004) Synthesis of chlorophyll b: localization of chlorophyllide a oxygenase and discovery of a stable radical in the catalytic subunit. BMC Plant Biol 4:5. https://doi.org/10.1186/1471-2229-4-5
Espineda CE, Linford AS, Devine D et al (1999) The AtCAO gene, encoding chlorophyll a oxygenase, is required for chlorophyll b synthesis in Arabidopsis thaliana. PNAS 96:10507–10511. https://doi.org/10.1073/pnas.96.18.10507
Friedland N, Negi S, Vinogradova-Shah T et al (2019) Fine-tuning the photosynthetic light harvesting apparatus for improved photosynthetic efficiency and biomass yield. Sci Rep 9:13028. https://doi.org/10.1038/s41598-019-49545-8
Gerotto C, Alboresi A, Giacometti GM et al (2012) Coexistence of plant and algal energy dissipation mechanisms in the moss Physcomitrella patens. New Phytol 196:763–773. https://doi.org/10.1111/j.1469-8137.2012.04345.x
Gerotto C, Alboresi A, Meneghesso A et al (2016) Flavodiiron proteins act as safety valve for electrons in Physcomitrella patens. PNAS 113:12322–12327. https://doi.org/10.1073/pnas.1606685113
Gerotto C, Trotta A, Bajwa AA et al (2019) Thylakoid protein phosphorylation dynamics in amoss mutant lacking serine/threonine protein kinase STN8. Plant Physiol 180:1582–1597. https://doi.org/10.1104/pp.19.00117
Giovagnetti V, Ruban AV (2018) The evolution of the photoprotective antenna proteins in oxygenic photosynthetic eukaryotes. Biochem Soc Trans 46:1263–1277. https://doi.org/10.1042/bst20170304
Green BR, Durnford DG (1996) The chlorophyll-carotenoid proteins of oxygenic photosynthesis. Annu Rev Plant Physiol Plant Mol Biol 47:685–714. https://doi.org/10.1146/annurev.arplant.47.1.685
Greene BA, Staehelin LA, Melis A (1988) Compensatory alterations in the photochemical apparatus of a photoregulatory, chlorophyll b-deficient mutant of Maize. Plant Physiol 87:365–370. https://doi.org/10.1104/pp.87.2.365
Havaux M, Dall’osto L, Bassi R (2007) Zeaxanthin has enhanced antioxidant capacity with respect to all other xanthophylls in Arabidopsis leaves and functions independent of binding to PSII antennae. Plant Physiol 145:1506–1520. https://doi.org/10.1104/pp.107.108480
Hoober JK, Eggink LL, Chen M (2007) Chlorophylls, ligands and assembly of light-harvesting complexes in chloroplasts. Photosynth Res 94:387–400. https://doi.org/10.1007/s11120-007-9181-1
Horn R, Grundmann G, Paulsen H (2007) Consecutive binding of chlorophylls a and b during the assembly in vitro of light-harvesting chlorophyll-a/b protein (LHCIIb). J Mol Biol 366:1045–1054. https://doi.org/10.1016/j.jmb.2006.11.069
Ilik P, Pavlovic A, Kouril R et al (2017) Alternative electron transport mediated by flavodiiron proteins is operational in organisms from cyanobacteria up to gymnosperms. New Phytol 214:967–972. https://doi.org/10.1111/nph.14536
Iwai M, Yokono M (2017) Light-harvesting antenna complexes in the moss Physcomitrella patens: implications for the evolutionary transition from green algae to land plants. Curr Opin Plant Biol 37:94–101. https://doi.org/10.1016/j.pbi.2017.04.002
Iwai M, Yokono M, Kono M et al (2015) Light-harvesting complex Lhcb9 confers a green alga-type photosystem I supercomplex to the moss Physcomitrella patens. Nature Plants 1:14008. https://doi.org/10.1038/nplants.2014.8
Järvi S, Suorsa M, Paakkarinen V et al (2011) Optimized native gel systems for separation of thylakoid protein complexes: novel super- and mega-complexes. Biochemical Journal 439:207–214. https://doi.org/10.1042/BJ20102155
Jung YJ, Lee HJ, Yu J et al (2021) Transcriptomic and physiological analysis of OsCAO1 knockout lines using the CRISPR/Cas9 system in rice. Plant Cell Rep 40:1013–1024. https://doi.org/10.1007/s00299-020-02607-y
Kim EH, Li XP, Razeghifard R et al (2009) The multiple roles of light-harvesting chlorophyll a/b-protein complexes define structure and optimize function of Arabidopsis chloroplasts: a study using two chlorophyll b-less mutants. Biochem Biophys Acta 1787:973–984. https://doi.org/10.1016/j.bbabio.2009.04.009
Koziol AG, Borza T, Ishida K et al (2007) Tracing the evolution of the light-harvesting antennae in chlorophyll a/b-containing organisms. Plant Physiol 143:1802–1816. https://doi.org/10.1104/pp.106.092536
Kunugi M, Takabayashi A, Tanaka A (2013) Evolutionary changes in chlorophyllide a oxygenase (CAO) structure contribute to the acquisition of a new light-harvesting complex in micromonas. J Biol Chem 288:19330–19341. https://doi.org/10.1074/jbc.M113.462663
Lee S, Kim JH, Yoo ES et al (2005) Differential regulation of chlorophyll a oxygenase genes in rice. Plant Mol Biol 57:805–818. https://doi.org/10.1007/s11103-005-2066-9
Lopez-Obando M, Hoffmann B, Géry C et al (2016) Simple and efficient targeting of multiple genes through CRISPR-Cas9 in Physcomitrella patens. G3-Genes Genom Genet 6:3647–3653. https://doi.org/10.1534/g3.116.033266
Malkin S, Armond PA, Mooney HA et al (1981) Photosystem II photosynthetic unit sizes from fluorescenceinduction in leaves. Plant Physiol 67:570–579. https://doi.org/10.1104/pp.67.3.570
Müller P, Li XP, Niyogi KK (2001) Non-photochemical quenching: a response to excess light energy. Plant Physiol 125:1558–1566. https://doi.org/10.1104/pp.125.4.1558
Mussgnug JH, Thomas-Hall S, Rupprecht J et al (2007) Engineering photosynthetic light capture: impacts on improved solar energy to biomass conversion. Plant Biotechnol J 5:802–814. https://doi.org/10.1111/j.1467-7652.2007.00285.x
Nagata N, Satoh S, Tanaka R et al (2004) Domain structures of chlorophyllide a oxygenase of green plants and Prochlorothrix hollandica in relation to catalytic functions. Planta 218:1019–1025. https://doi.org/10.1007/s00425-003-1181-6
Nick S, Meurer J, Soll J et al (2013) Nucleus-encoded light-harvesting chlorophyll a/b proteins are imported normally into chlorophyll b-free chloroplasts of Arabidopsis. Mol Plant 6:860–871. https://doi.org/10.1093/mp/sss113
Oster U, Tanaka R, Tanaka A et al (2000) Cloning and functional expression of the gene encoding the key enzyme for chlorophyll b biosynthesis (CAO) from Arabidopsis thaliana. Plant J 21:305–310. https://doi.org/10.1046/j.1365-313x.2000.00672.x
Perrine Z, Negi S, Sayre RT (2012) Optimization of photosynthetic light energy utilization by microalgae. Algal Res 1:134–142. https://doi.org/10.1016/j.algal.2012.07.002
Pinnola A, Dall’Osto L, Gerotto C et al (2013) Zeaxanthin binds to light-harvesting complex stress-related protein to enhance nonphotochemical quenching in Physcomitrella patens. Plant Cell 25:3519–3534. https://doi.org/10.1105/tpc.113.114538
Pinnola A, Cazzaniga S, Alboresi A et al (2015) Light-harvesting complex stress-related proteins catalyze excess energy dissipation in both photosystems of Physcomitrella patens. Plant Cell 27:3213–3227. https://doi.org/10.1105/tpc.15.00443
Pinnola A, Alboresi A, Nosek L et al (2018) A LHCB9-dependent photosystem I megacomplex induced under low light in Physcomitrella patens. Nat Plants 4:910–919. https://doi.org/10.1038/s41477-018-0270-2
Qin XC, Suga M, Kuang T et al (2015) Structural basis for energy transfer pathways in the plant PSI-LHCI supercomplex. Science 348:989–995. https://doi.org/10.1126/science.aab0214
Rensing SA, Lang D, Zimmer AD et al (2008) The Physcomitrella genome reveals evolutionary insights into the conquest of land by plants. Science 319:64–69. https://doi.org/10.1126/science.1150646
Schaefer DG (2002) A new moss genetics: targeted mutagenesis in Physcomitrella patens. Annu Rev Plant Biol 53:477–501. https://doi.org/10.1146/annurev.arplant.53.100301.135202
Schaefer DG, Zrÿd JP (1997) Efficient gene targeting in the moss Physcomitrella patens. Plant J 11:1195–1206. https://doi.org/10.1046/j.1365-313x.1997.11061195.x
Schansker G, Srivastava A, Strasser RJ (2003) Characterization of the 820-nm transmission signal paralleling the chlorophyll a fluorescence rise (OJIP) in pea leaves. Funct Plant Biol 30:785–796. https://doi.org/10.1071/fp03032
Schansker G, Tóth SZ, Strasser RJ (2005) Methylviologen and dibromothymoquinone treatments of pea leaves reveal the role of photosystem I in the Chl a fluorescence rise OJIP. Biochim Biophys Acta 1706:250–261. https://doi.org/10.1016/j.bbabio.2004.11.006
Takabayashi A, Kurihara K, Kuwano M et al (2011) The oligomeric states of the photosystems and the light-harvesting complexes in the Chl b-less mutant. Plant Cell Physiol 52:2103–2114. https://doi.org/10.1093/pcp/pcr138
Tanaka R, Tanaka A (2005) Effects of chlorophyllide a oxygenase overexpression on light acclimation in Arabidopsis thaliana. Photosynth Res 85:327–340. https://doi.org/10.1007/s11120-005-6807-z
Tanaka R, Tanaka A (2011) Chlorophyll cycle regulates the construction and destruction of the light-harvesting complexes. Biochem Biophys Acta 1807:968–976. https://doi.org/10.1016/j.bbabio.2011.01.002
Tanaka A, Tanaka R (2019) The biochemistry, physiology, and evolution of the chlorophyll cycle—ScienceDirect. Adv Bot Res 90:183–212. https://doi.org/10.1016/bs.abr.2019.03.005
Tanaka A, Ito H, Tanaka R et al (1998) Chlorophyll a oxygenase (CAO) is involved in chlorophyll b formation from chlorophyll a. PNAS 95:12719–12723. https://doi.org/10.1073/pnas.95.21.12719
Thayer SS, Björkman O (1990) Leaf xanthophyll content and composition in sun and shade determined by HPLC. Photosynth Res 23:331–343. https://doi.org/10.1007/BF00034864
Way DA, Pearcy RW (2012) Sunflecks in trees and forests: from photosynthetic physiology to global change biology. Tree Physiol 32:1066–1081. https://doi.org/10.1093/treephys/tps064
Wu G, Ma L, Sayre RT et al (2020) Identification of the optimal light harvesting antenna size for high-light stress mitigation in plants. Front Plant Sci 11:505. https://doi.org/10.3389/fpls.2020.00505
Yamamoto H, Takahashi S, Badger MR et al (2016) Artificial remodelling of alternative electron flow by flavodiiron proteins in Arabidopsis. Nat Plants 2:16012. https://doi.org/10.1038/nplants.2016.12
Yan Q, Zhao L, Wang W et al (2021) Antenna arrangement and energy-transfer pathways of PSI-LHCI from the moss Physcomitrella patens. Cell Discov 7:10. https://doi.org/10.1038/s41421-021-00242-9
Yang C, Kosemund K, Cornet C et al (1999) Exchange of pigment-binding amino acids in light-harvesting chlorophyll a/b protein. Biochemistry 38:16205–16213. https://doi.org/10.1021/bi990738x
Funding
This work was supported by the Academician Workstation of Agricultural High-tech Industrial Area of the Yellow River Delta, National Center of Technology Innovation for Comprehensive Utilization of Saline-Alkali Land, Dongying, Shandong, China and Science & Technology Specific Projects in Agricultural High-tech Industrial Demonstration Area of the Yellow River Delta (Grant No: 2022SZX12) and the National Key R&D Program of China (Grant No: 2017YFA0503701).
Author information
Authors and Affiliations
Contributions
CL and CY planned and designed the research. LZ and CL carried out the experiments and took part in the data analysis. LZ and CL wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, L., Yang, C. & Liu, C. Revealing the significance of chlorophyll b in the moss Physcomitrium patens by knocking out two functional chlorophyllide a oxygenase. Photosynth Res 158, 171–180 (2023). https://doi.org/10.1007/s11120-023-01044-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-023-01044-8