Abstract
Depending upon their growth responses to high and low irradiance, respectively, thermophilic Synechococcus sp. isolates from microbial mats associated with the effluent channels of Mushroom Spring, an alkaline siliceous hot spring in Yellowstone National Park, can be described as either high-light (HL) or low-light (LL) ecotypes. Strains isolated from the bottom of the photic zone grow more rapidly at low irradiance compared to strains isolated from the uppermost layer of the mat, which conversely grow better at high irradiance. The LL-ecotypes develop far-red absorbance and fluorescence emission features after growth in LL. These isolates have a unique gene cluster that encodes a putative cyanobacteriochrome denoted LcyA, a putative sensor histidine kinase; an allophycocyanin (FRL-AP; ApcD4-ApcB3) that absorbs far-red light; and a putative chlorophyll a-binding protein, denoted IsiX, which is homologous to IsiA. The emergence of FRL absorbance in LL-adapted cells of Synechococcus sp. strain A1463 was analyzed in cultures responding to differences in light intensity. The far-red absorbance phenotype arises from expression of a novel antenna complex containing the FRL-AP, ApcD4-ApcB3, which is produced when cells were grown at very low irradiance. Additionally, the two GAF domains of LcyA were shown to bind phycocyanobilin and a [4Fe-4S] cluster, respectively. These ligands potentially enable this photoreceptor to respond to a variety of environmental factors including irradiance, redox potential, and/or oxygen concentration. The products of the gene clusters specific to LL-ecotypes likely facilitate growth in low-light environments through a process called Low-Light Photoacclimation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mushroom and Octopus Springs are alkaline, siliceous hot springs in the Lower Geyser Basin of Yellowstone National Park, WY, USA. The run-off channels of these thermal features harbor phototrophic microbial communities that have served as models for studies of microbial ecophysiology and speciation for more than 50 years (Brock 1978, 1985; Ward et al. 1998). Because light is the primary energy source available, these chlorophototrophic communities exhibit remarkable diversity. More than seventeen different chlorophototrophs have been identified in these mats, and they represent much of the known diversity of chlorophyll (Chl)-based phototrophs (Tank et al. 2017; Thiel et al. 2018; Saini et al., 2020). The steep chemical and physical gradients that occur in the mats, as well as metabolic interactions among community members, provide strong selective pressures for speciation (e.g., Becraft et al. 2015). Members of the phyla Cyanobacteria and Chloroflexota are the predominant phototrophic organisms in these mat communities. The cyanobacteria are mostly Synechococcus spp., and the members of the Chloroflexota mostly belong to the genera Roseiflexus, Chloroflexus, and Candidatus Roseilinea (Klatt et al. 2011, 2013; Liu et al. 2011, 2012; Thiel et al. 2016, 2017; Tank et al. 2017).
Chl a absorbs all wavelengths of visible light, and thus the light reaching the lowest levels of the photic layer of the mat is strongly filtered by cyanobacteria and the other chlorophotorophs in the upper layers of the mat. Due to the density of cells in the mats and the physical properties of light, an interesting ecophysiological question thus arises: how do the cyanobacteria living in the lowest regions of the euphotic layer of the mat acquire sufficient light to maintain energy production for growth? In some hot spring mat communities, for example in the mats of La Duke Hot Springs (Gardiner, MT, USA) (Gan et al., 2014, 2015) or the mats of Nakabusa hot springs in Japan (Ohkubo and Miyashita 2017), a process known as far-red light photoacclimation (FaRLiP) occurs that allows some cyanobacteria to acclimate to the FRL-enriched regions in the lower levels of such mats. However, the genomes of the cyanobacteria that inhabit the mats of Octopus and Mushroom Springs do not have the FaRLiP gene cluster, and thus they must employ a different mechanism to grow in the deeper layers of those mat communities.
Some Synechococcus spp. strains isolated from deep (~ 600 to 900 μm) within the euphotic layer of the Mushroom Spring mat community demonstrated improved growth at low irradiance (~ 25 μmol photons m−2 s−1) compared to isolates of “high-light” (HL) ecotypes (HLEs) isolated from nearer the surface (Nowack et al. 2015). Among other differences, these LL-ecotypes (LLEs) were found to have a unique cluster of four genes related to light harvesting that is absent in HLEs (suppl. Fig. S1). This cluster encodes a putative photoreceptor-histidine kinase (LcyA), two allophycocyanin (AP) subunits (ApcD4 and ApcB3), and a putative Chl a-binding protein, denoted IsiX, which is homologous to IsiA (Olsen et al. 2015). The LLEs also possessed a far-red fluorescence emission feature when cells grown in LL were preferentially excited with light absorbed by phycobiliproteins (PBP(s)) that was not observed in a HLE or when the LLEs were grown in HL (Nowack et al. 2015). It was subsequently discovered that two genes, apcD4 and apcB3, in the LLE-specific gene cluster, encode an FRL-AP that could be responsible for some or all of the observed FRL fluorescence (Soulier et al. 2020; Soulier and Bryant 2021). The process by which LL-adapted Synechococcus spp. acquire FRL absorbance after growth at low irradiance was termed low-light photoacclimation (LoLiP) (Zhao et al. 2015; Ho et al. 2017c; Soulier et al. 2020). In this study the influences of light intensity and some other factors on FRL absorbance and transcription of the LoLiP genes in Synechococcus sp. strain A1463 (hereafter Synechococcus A1463) were analyzed. Additionally, the two ligand-binding GAF (cGMP-specific phosphodiesterases, Adenylyl cyclases and FhlA) domains of the putative cyanobacteriochrome (CBCR) photoreceptor, LcyA, were heterologously expressed and characterized. The data suggest that LoLiP is a response that includes the production of a novel antenna complex.
Materials and methods
Strain information and growth conditions
Synechococcus A1463, also known as Synechococcus sp. 63AY4M1 (NCBI Taxonomy ID: 1,353,263), is an LLE enrichment culture from Mushroom Spring, Yellowstone National Park. A draft version of the genome sequence of Synechococcus A1463 is available in GenBank as accession GCA_002760395.1. The creation of enrichment cultures of various Synechococcus spp. ecotypes from this hot spring and Octopus Spring has been described previously (Allewalt et al. 2006; Becraft et al. 2015). B-HEPES medium (Rippka et al. 1979; Dubbs and Bryant 1991) supplemented with 1 mM biotin was used to grow Synechococcus A1463 at 58 °C. Cultures were slowly sparged with 5% CO2 and 95% N2 (v/v). Unless otherwise specified, cultures were not stirred and received approximately 40 μmol photons m−2 s−1 of continuous, white light provided by a soft white halogen bulb, although self-shading could lower this value for most cells in the flask. Cells grown this way or at lower light intensities are termed “LL-acclimated”. White light intensities were measured with a model QSL-100 light meter (Biospherical Instruments Inc., San Diego, CA) that is sensitive to visible light (PAR). FRL intensities were measured with a model MP-200 solar radiation meter (Apogee Instruments, Logan, UT). Synechococcus sp. PCC 7002 (hereafter Synechococcus 7002) was grown as previously described (Ludwig and Bryant 2011).
α-Select Silver Efficiency Escherichia coli competent cells (Bioline, Memphis, TN) were used for routine cloning and preparation of plasmids used in this study. E. coli strain BL21 (DE3) (New England Biolabs, Ipswich, MA) was used for heterologous protein production. E. coli cells were grown in Luria–Bertani lysogeny broth (LB) medium (pH = 7.0) supplemented with kanamycin (100 μg mL−1), chloramphenicol (25 μg mL−1), carbenicillin (200 μg mL−1), and spectinomycin (50 μg mL−1) as appropriate at 30 °C with shaking at 200 rpm. When cultures of engineered E. coli expression strains reached OD600 = 0.7, expression was induced by addition of 1 mM isopropyl β-D-1-thiogalactopyranoside. Cultures were grown an additional 4 h at 30 °C with shaking before reducing the temperature to 18 °C and shaking overnight. E. coli cells were pelleted by centrifugation at 9,184 × g and stored at − 20 °C until needed.
Isolation of photosynthetic complexes from Synechococcus A1463 and Synechococcus 7002
Thylakoid and PBP complexes from Synechococcus A1463 and Synechococcus 7002 were extracted and separated by sucrose density ultracentrifugation as previously described by Gan et al. (2014) with the following differences. Thylakoid membrane samples were solubilized for 45 min with 1% (v/v) Triton X-100 prior to loading a 2-mL sample on each gradient. Continuous gradients designed to separate thylakoid complexes were constructed with 5–25% sucrose (22 mL) with a 70% (w/v) sucrose “cushion” at the bottom of the gradient tube (2 mL). All gradient solutions contained 0.1% (v/v) Triton X-100. The discontinuous high-phosphate (0.75 M) sucrose gradients designed to separate PBP complexes were prepared as previously described (Soulier et al. 2020).
Clear-native polyacrylamide gel electrophoresis
Clear-native polyacrylamide gel electrophoresis (CN-PAGE) was used so that absorbance and fluorescence emission spectra could be acquired from excised gel bands prior to tryptic peptide analysis. The stacking and resolving gels were made with 4% and 5–15% (v/v) acrylamide-bisacrylamide solution, respectively. The acrylamide-bisacrylmide solution contained a total concentration of 49.5% (v/v) acrylamide and bisacrylamide monomers, and 3% bisacrylamide (v/v) in distilled water. Buffers and running conditions were as previously described for CN-PAGE with membrane proteins (Wittig and Schägger 2005). Sample loading buffer (3×) was identical to the gel buffer with the addition of 2% (w/v) n-dodecyl β-D-maltoside and 0.001% (w/v) bromophenol blue dye to mark the ion front during electrophoresis.
Identification of tryptic peptides by mass spectrometry
Peptides in sucrose gradient fractions and in excised gel bands were subjected to digestion with trypsin as described by (Ho et al. 2017b) and by following protocols recommended by the Penn State Proteomics and Mass Spectroscopy Core Facility. The LC–MS-MS analyses were subsequently performed on a Thermo LTQ Orbitrap Velos ETD mass spectrometer equipped with a Dionex UltiMate 3000 Nano-flow 2D LC system (Thermo Fisher Scientific, Waltham, MA). Proteins were identified by custom database searches against the predicted proteomes of Synechococcus A1463 or Synechococcus 7002 supplemented with heterologously produced proteins and appropriate control proteins.
DNA constructs for heterologous production of LoLiP proteins and GAF domains of LcyA
The apcD4 and apcB3 genes, isiX, and regions of the lycA gene encoding the GAF domains were amplified from the LoLiP gene cluster in Synechococcus A1463 using Q5 High-Fidelity polymerase Master Mix (NEB #M0492S; New England Biolabs, Ipswich, MA) and the oligonucleotide primers listed in suppl. Table S1. The genes apcD4, apcB3, and isiX from the LoLiP gene cluster were inserted as an operon into plasmid pAQ1Ex (Xu et al. 2011) by Gibson assembly (Gibson et al. 2009) to produce plasmid pAQ1Ex-LL, which was then transformed into E. coli α-Select Silver Efficiency competent cells (Bioline, Memphis, TN) for subsequent DNA amplification of the construct. Plasmid pAQ1Ex-LL was then transformed into the model cyanobacterium Synechococcus 7002, producing strain Synechococcus 7002-LL, for heterologous overexpression as described (Xu et al. 2011). The previously described plasmid pAQ1Ex-apcD4apcB3 (Soulier et al. 2020) was also transformed into Synechococcus 7002, producing Synechococcus 7002-apcD4B3, for heterologous overexpression.
The lcyA gene segment to produce GAF1 (amino acids 679–831) was inserted into plasmid pET-15b via Gibson assembly (Gibson et al. 2009) prior to transformation of the construct into α-Select Silver Efficiency E. coli competent cells (Bioline, Memphis, TN). After DNA extraction, construct pET-15b-GAF1 was transformed into E. coli BL21 (DE3) competent cells containing the pPcyA plasmid for production of phycocyanobilin (Biswas et al. 2010; Soulier et al. 2020; Soulier and Bryant 2021).
The lcyA gene segment to produce the GAF2 domain (amino acids 870–1012) was similarly inserted into pET-15b by Gibson assembly and transformed into α-Select Silver Efficiency E. coli competent cells (Bioline, Memphis, TN) prior to DNA extraction and transformation of construct pET-15b-GAF2 into BL21(DE3) ΔiscR competent cells for heterologous overexpression (Akhtar and Jones 2008; Walters and Golbeck 2018).
Purification of recombinant proteins and polypeptides
Recombinant proteins were individually expressed with an N-terminal [His]10-tag for genes inserted into pAQ1Ex and with an N-terminal [His]6-tag for genes inserted into pET-15b. When multiple genes were expressed from an operon, only the most upstream gene encoded the N-terminal polyhistidine tag. Cell lysates were produced by passing cell suspensions three times through a chilled French pressure cell at 124 MPa and were clarified by ultracentrifugation before purification of the recombinant protein by immobilized metal affinity chromatography (IMAC) as previously described (Soulier et al. 2020).
The GAF2 domain of LcyA of Synechococcus A1463 formed inclusion bodies when heterologously overexpressed in E. coli strain BL21(DE3) ΔiscR from plasmid pET-15b-GAF2, and thus it was purified as follows. E. coli cells engineered and induced to produce the GAF2 polypeptide were lysed in cold 50 mM Tris–HCl buffer, pH = 8.3, with 20 mM NaCl and 2 mM 2-mercaptoethanol. The lysate was centrifuged at 90,000 × g for 30 min at 4 °C to pellet debris and inclusion bodies. The supernatant was discarded, and the pellet was resuspended in 50 mM Tris–HCl, pH = 8.3, containing 20 mM NaCl and 4 mM 2-mercaptoethanol. The resulting suspension was again centrifuged at 90,000×g for 30 min at 4 °C, and this wash-centrifugation step was repeated once. The twice-washed pellet was resuspended in 50 mM Tris–HCl buffer, pH = 8.3, with 20 mM NaCl and 40 mM 2-mercaptoethanol with 9 M urea. The suspension was stirred for 1 h at room temperature prior to a final centrifugation (90,000×g for 30 min at 4 °C). The supernatant was loaded onto a column packed with with ~ 1.5 mL Ni2+-nitrilotriacetate affinity resin (Goldbio, St. Louis, MO). The column was washed with five column volumes of 50 mM Tris–HCl (pH = 8.3) containing 20 mM NaCl before addition of the same buffer containing 200 mM imidazole and elution of the light-brown-colored polypeptide. The solution containing the GAF2 polypeptide was placed inside an anoxic chamber and passed through a PD-10 column (Sephadex G-25; GE Healthcare, Marlborough, MA) pre-equilibrated with anoxic 50 mM Tris–HCl buffer, pH = 8.3, to remove imidazole and oxygen.
Chemical reconstitution of the GAF2 polypeptide iron-sulfur cluster
After IMAC purification of the soluble GAF2 polypeptide, the iron-sulfur cluster coordinated by GAF2 was chemically reconstituted under anoxic conditions by a procedure adapted from Heinnickel et al. (2005). A stir-bar and 4% (v/v) 2-mercaptoethanol were added to the anoxic solution of the GAF2 polypeptide before it was sealed in a glass bottle with a rubber stopper inside the anoxic chamber. Separate solutions of 60 mM ferrous ammonium sulfate and sodium sulfide were also prepared inside the chamber in anoxic 50 mM Tris–HCl buffer, pH = 8.3, and were moved out of the chamber with the sample to a cold room (~ 4 °C). Using a plastic syringe and needle to pierce the rubber stopper, 60 mM ferrous ammonium sulfate was slowly added to the stirring sample of GAF2 over the course of ten minutes, to a final concentration of ~ 800 μM. The stock solution of 60 mM sodium sulfide was added over a second ten-minute period to the stirred sample, also to a final concentration of ~ 800 μM. The sample was then sealed with Parafilm and stirred in the cold room overnight. Excess iron and sulfide were removed the next day from the reconstituted GAF2 polypeptide sample by chromatography on a PD-10 desalting column that had been equilibrated with anoxic 50 mM Tris–HCl buffer, pH = 8.3, inside the anoxic chamber.
Absorbance, fluorescence, and EPR spectroscopy
A Cary 14 spectrophotometer modified for on-line data acquisition (On-line Systems, Inc.; Bogart, GA) was used to measure absorbance spectra. Fluorescence emission spectra were measured at 77 K for each protein using an SLM 8000C spectrofluorometer modified for digital data acquisition by On-line Systems, Inc. (Bogart, GA). To excite Chls or PBPs preferentially, samples were excited at either 440 nm or 590 nm, respectively.
Low temperature CW X-band EPR was performed using a Bruker ELEXSYS E-500 spectrometer (Bruker BioSpin Corporation, Billerica, MA) as described by Walters and Golbeck (2018). Specific instrument settings are described in the corresponding figure legends.
RNA sequencing
Separate total RNA extractions were performed on Synechococcus A1463 cells that had been grown for 3 days at 10 μmol photons m−2 s−1 (LL), 3 days at 500 μmol photons m−2 s−1 (HL), two weeks at 20 μmol photons m−2 s−1 (LL), and two weeks at 400 μmol photons m−2 s−1 (HL). The bead-phenol–chloroform method of extraction described by Kim et al. (2006) was used in sequence with the High Pure RNA Isolation Kit (Roche Applied Science, Mannheim, Germany) to produce highly purified samples of total RNA. Approximately 10 µg of total RNA from HL and LL-grown cells were sent to the MiGS Microbial Genome Sequencing Center (Pittsburgh, PA) for rRNA depletion and RNA sequencing. Depletion of rRNA was performed using the RiboZero Plus rRNA Depletion kit (Illumina). Approximately 12 million reads for each sample were acquired with Illumina Stranded RNA library preparation. Adapter trimming and quality control was performed with bcl2fastq. Read mapping was performed with HISAT2, and the number of HISAT2 counts was 2.01E + 07 for the HL sample and 1.47E + 07 for the LL sample. The featureCounts function of Subread was used to quantify reads. Read counts were loaded into R and normalized using the Trimmed Mean of M values (TMM) algorithm of edgeR, with subsequent values converted to counts per million (cpm). Analysis of differential expression was performed using edgeR’s exact test for differences between two groups of negative-binomial counts with an estimated dispersion value of 1.0. A subset of the quantified genes with |logFC|> 1 and p < 0.05 were reported as differentially expressed.
Reverse-transcription PCR
Some of the RNA extracted from cells grown for 3 days at 10 μmol photons m−2 s−1 (LL) was used for reverse-transcription PCR (RT-PCR) with the OneTaq One-Step RT-PCR Kit (NEB #E5315S; New England Biolabs, Ipswich, MA). Supplemental Table S1 includes the primers that were used in amplifying across each intergenic region within the LoLiP cluster, namely lcyA-apcD4, apcD4-apcB3, apcB3-isiX, and lcyA-isiX to demonstrate co-expression. The identities of the amplified fragments were verified by DNA sequencing of the produced amplicons.
Results
Absorbance of Synechococcus A1463 cells from different growth conditions
Previous studies indicated that LLEs of Synechococcus acclimated to low irradiance acquire a new absorbance band centered at ~ 710 nm and a fluorescence emission band at ~ 715 nm (Nowack et al. 2015). Additionally, the genomes of LLEs encode a four-gene cluster encoding proteins that are related to light harvesting and that are not present in HLEs (Olsen et al. 2015). Thus, the presence (or absence) of an absorbance feature at ~ 710 nm was monitored in cells grown in various conditions. A culture of Synechococcus A1463 grown in HL (600 μmol photons m−2 s−1) that lacked FRL absorbance was transferred to 50 μmol photons m−2 s−1 for one day and then to 7 μmol photons m−2 s−1. Over the ensuing 3 days, an FRL absorbance band developed at ~ 715 nm (Fig. 1). When cells were continuously stirred with a magnetic stir-bar, the stirred cells lacked FRL absorbance after the irradiance was reduced from 50 to 10 μmol photons m−2 s−1 (suppl. Fig. S2). Two populations of cells, sedimented and dispersed, appeared in cultures when stirring ceased and the light intensity was maintained at 10 μmol photons m−2 s−1. The absorbance spectra of cells from both populations had prominent absorbance features at ~ 616, ~ 653, and 680 nm due to PBPs and Chl a, respectively, but only the sedimented cells developed a significant shoulder at ~ 717 nm (suppl. Figs. S2 and S3).
Changes in the whole-cell absorbance spectra of LL-adapted Synechococcus A1463 after growth in extremely low irradiance for 3 days. Cells that had lost all FRL absorbance after being subjected to ~ 600 μmol photons m−2 s−1 for 72 h were grown at 50 μmol photons m−2 s−1 for one day before reducing the light intensity to 7 μmol photons m−2 s−1. Beginning at 24 h after reducing the light intensity to the lower level, a steady increase in FRL absorbance (715 nm feature) was observed over time, peaking at 72 h. Additionally, the QY absorbance band associated with Chl a-binding proteins shifted 3-nm to the blue—from 680 to 677 nm. Cells were not stirred over the course of the experiment and were very slowly sparged with a gas mixture of 5% carbon dioxide balanced with 95% (v/v) nitrogen
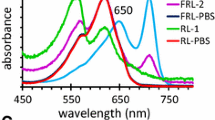
The absorbance of Synechococcus A1463 cells was also monitored after growth in FRL and red light (RL). A combination of overlapping red and green plastic filters, or a red plastic filter alone, was used to grow cells in FRL and RL, respectively (Gan et al. 2014; Shen et al. 2019). Compared to cells grown in white light, cells that were placed in FRL for 7 days exhibited the following: an increase in absorbance at ~ 712 nm; a reduction in amplitude as well as a blue-shift in the absorbance at 680 to ~ 673 nm; and an increase in the PBP-associated absorbance at ~ 617 nm (Fig. 2). However, cells incubated in FRL did not grow, and the amplitudes of both the FRL absorbance band and the absorbance associated with Chl-binding proteins (~ 673 nm) were greatly reduced after 21 days (Fig. 2). After growth in RL for 24 days, the absorbance due to phycocyanin at ~ 618 nm increased considerably compared to that of cells grown in white light (i.e., day 0), but no absorbance changes were observed in the 710–720 nm spectral region (suppl. Fig. S4).
Absorbance spectra of Synechococcus A1463 cells grown in white light and FRL. The spectrum of cells from the WL starter culture (Day 0, black line) is shown in both conditions. a The absorbance spectra of cells grown in white light (WL; ~ 40 μmol photons m−2 s−1) for 3, 7, and 21 days are shown with green, blue, and violet lines, respectively. The amplitude of the FRL absorbance feature remained relatively constant in WL cells. b The absorbance spectra of cells grown in FRL (~ 40 μmol photons m−2 s−1) for 3, 7, and 21 days are shown with goldenrod, orange, and red lines, respectively. The FRL culture flask was wrapped in overlapping red- and green-colored filter plastic (GamColor 250 and GamColor 650, respectively; Shen et al. 2019) and illuminated with a soft white halogen bulb. The amplitude of the FRL feature (~ 712 nm) initially increased in FRL cells over 7 days, but after 21 days in FRL cells had very little FRL absorbance. The amplitude of the absorbance associated with Chl-binding proteins (~ 680 nm) decreased and was slightly blue-shifted (~ 673 nm) before disappearing from FRL cells, which turned a pale blue color after 21 days
Identifying the source of FRL absorbance in Synechococcus A1463
Phycobilisomes (PBS), Photosystem I (PSI), and Photosystem II (PSII) complexes were isolated by sucrose density ultracentrifugation from Synechococcus A1463 cells exhibiting FRL absorbance. PBS isolated from such cells did not incorporate ApcD4-ApcB3, the previously characterized FRL-AP from LLEs (Fig. 3) (Soulier et al. 2020; Soulier and Bryant 2021). Low levels of peptides from ApcD4 alone were detected in fractions F1 and F2, and peptides from ApcB3 were detected in the PBS fraction (F3), but peptides from both subunits were not detected in any gradient fraction (suppl. Table S2). FRL absorbance and fluorescence emission were observed in the uppermost blue fraction of gradients (Fig. 3, F1) but not in PBS (Fig. 3, F3). It is possible that ApcD4 in F1 formed complexes with ApcB1 to form a FRL-absorbing AP complex, and the presence of ApcB3 in the PBS fraction could similarly be explained by the structural similarity of ApcB1 and ApcB3 leading to complexes other than the expected ApcD4-ApcB3 complex (see Soulier and Bryant 2021 for further details).
Absorbance and fluorescence emission spectra of PBPs and PBS extracted from Synechococcus A1463 cells acclimated to LL. The absorbance (a) and fluorescence emission (b) spectra of whole cells (WC) prior to lysis are shown with a green line. PBPs and PBS complexes were separated into three fractions by sucrose density ultracentrifugation in 0.75 M K-phosphate buffer, as pictured. The absorbance (a) and fluorescence emission spectra (b, λex = 590 nm) of the fractions are shown in cyan (F1), blue (F2), and black (F3). F3 corresponds to intact PBS, while F2 probably corresponds to phycocyanin-containing rod complexes. F1, the only fraction with FRL absorbance, consists of trimeric or hexameric PBPs
FRL absorbance was observed for several fractions in sucrose gradients used to separate complexes from solubilized membranes of Synechococcus A1463 (Fig. 4 and suppl. Fig. S5). Fractions F3 and F4 had a small FRL absorbance feature (~ 710 nm) after an initial round of sucrose density ultracentrifugation (suppl. Fig. S5, F3 and F4). The FR fluorescence emission was greater when Chl-binding proteins were preferentially excited at 440 nm than when PBPs were excited at 590 nm. Fractions F3, F4, and F5, the latter of which corresponds to trimeric PSI, were dialyzed to eliminate sucrose, and each was subjected to a second round of fractionation on sucrose gradients. These “Round 2” gradients (Fig. 4) improved the resolution of the various complexes from fractions F3, F4, and F5 of the “Round 1” gradients (suppl. Fig. S5) and revealed a FRL absorbance feature around 710 nm in two blue-green fractions (Fig. 4, F3-4 and F4-1) and in one green fraction (F4-2); a FRL shoulder was present in the absorbance spectrum of the PSI trimer fraction (F5-2). As with fractions 3 and 4 from the initial gradient, fractions F3-4, F4-1, and F4-2 mostly exhibited FRL fluorescence emission (λmax = ~ 716–719 nm) when Chl was preferentially excited at 440 nm. Tryptic peptide fingerprinting of the “Round 2” sucrose gradient fractions (Table 1) indicated that fractions F3-4, F4-1, and F4-2 were a mixture of PBPs, PSII, and some PSI proteins, but all included both ApcD4 and ApcB3 as well as IsiX. IsiX was not detected in fraction F5-2, possibly due to low concentration or the overall hydrophobicity of the sample, but both ApcD4 and ApcB3 were again present. Notably, peptides from LcyA, the putative CBCR of the LoLiP cluster, were also detected in fraction F3-1 (Table 1).
Absorbance and fluorescence emission spectra of proteins and protein complexes extracted from LL-acclimated Synechococcus A1463 thylakoid membranes and separated by two rounds of sucrose density ultracentrifugation. The “Round 1” gradient (continuous 5–25% sucrose with a 70% sucrose cushion) shown on the left was separated into six fractions (see suppl. Fig. S5). Fractions F3, F4 and F5 from Round 1 were dialyzed, concentrated and further separated by sucrose density ultracentrifugation (“Round 2”, 5–18% continuous sucrose gradients). a Fraction 3 was separated into four sub-fractions: F3-1, F3-2, F3-3 and F3-4, depicted with black, red, blue and green lines, respectively. b Fraction 4 was separated into five sub-fractions, though only two fractions had unique spectra: F4-1 and F4-2, depicted with black and red lines, respectively. c Fraction 5 was separated into two sub-fractions, F5-1 and F5-2, depicted with black and red lines, respectively. Fluorescence emission spectra were collected at low temperature (77 K) for all fractions, with the excitation wavelength set to either 440 nm (solid lines) or 590 nm (dotted lines) for preferential excitation of Chls or PBPs, respectively. After the second round of separation, FRL absorbance (λmax ~ 709 nm) was present in fractions F3-4, F4-1, and F4-2. FRL fluorescence emission (λmax = ~ 716–719 nm) was only observed for these samples when Chl-binding proteins were preferentially excited at 440 nm, as opposed to PBPs
Fraction F3-4 (Fig. 4) was further resolved into several bands by CN-PAGE (Fig. 5a). The absorbance spectra of the excised bands indicated the presence of FRL-absorbing AP in bands 1 and 3 (Fig. 5a). Band 1 had the most pronounced FRL absorbance and exhibited far-red fluorescence emission (λmax ~ 715 nm) upon excitation with light of wavelengths 440 nm, 590, and 690 nm (Fig. 5b). Tryptic peptide fingerprinting of the excised bands showed that band 1 contained IsiX and ApcB3 and that band 3 contained IsiX, ApcD4, and ApcB3 (Table 2). The failure to detect ApcD4 in band 1 may have been due to the low protein concentration of the analyzed samples, because the sharp absorbance observed at ~ 710 nm results from the excitonic coupling of the phycocyanobilin on ApcD4 with the chromophore on ApcB3 of an adjacent monomer in trimeric FRL-AP (Soulier et al. 2021). Peptides from ApcD4 and IsiX were not identified in bands 2, 4, and 5, which had very little or no FRL absorbance (Fig. 5a, Table 2). Collectively, these data from sucrose density gradients and the CN-PAGE suggest that ApcD4, ApcB3, and IsiX form an FRL-absorbing complex. The tailing FRL absorbance between 700 and 730 nm observed in the PSI complexes in fraction F5-2 (Fig. 4) can be explained by the presence of some FRL-absorbing Chl a molecules, which are often associated with PSI (Sener et al. 2002; Karpetyan et al. 2014; Tros et al. 2020; Toporik et al. 2020).
Absorbance spectra of protein bands separated by CN-PAGE from LL-acclimated Synechococcus A1463 and fluorescence emission spectra of a FRL-absorbing gel band (band #1) at three excitation wavelengths. a FRL-absorbing sucrose gradient fraction F3-4 (Fig. 4) was resolved by CN-PAGE (see inset). Five bands were excised, and the absorbance spectra of bands 1 through 5 are shown with black, red, blue, magenta, and green lines, respectively. Tryptic peptides identified in each band by mass spectrometry are shown in Table 2. b Low temperature (77 K) fluorescence emission spectra were collected from the blue-green band 1 at three excitation wavelengths: 440 nm (black line), 590 nm (red line), and 690 nm (blue line)
Heterologous expression of LL-ecotype genes in Synechococcus 7002
The apcD4, apcB3, and isiX genes from Synechococcus A1463 were concatenated to form an operon between the NdeI and BamHI restriction sites of the pAQ1Ex expression plasmid and transformed into the model strain Synechococcus 7002, which does not normally produce FRL-absorbing pigment-proteins. PBP and PBS fractions isolated from the control, wild-type Synechococcus 7002 cells did not exhibit FRL absorbance for any of the fractions obtained from sucrose gradients (suppl. Fig. S6). PBS (Fig. 6) were also isolated from strain Synechococcus 7002-LL cells that exhibited FRL absorbance. As in Synechococcus A1463, the PBS fraction (Fig. 6a, F3) from Synechococcus 7002-LL did not exhibit FRL absorbance, but the upper two blue-colored gradient fractions had strong absorbance at ~ 710 nm (Fig. 6a, F2 and especially F1). Fraction F1 also had intense FRL fluorescence emission at ~ 716 nm when this fraction was excited with 590 nm light (Fig. 6b, F1). Both ApcD4 and ApcB3 were detected in fractions F1 and F2, but only ApcD4 was detected in PBS (F3) by peptide analysis of the three PBP fractions from Synechococcus 7002-LL (suppl. Table S3). IsiX was not detected in any of these fractions. The number of uniquely detected peptides for ApcD4 and ApcB3 indicated that these proteins were present in highest concentration in fraction F1.
Absorbance and fluorescence emission spectra of PBPs and PBS complexes extracted from Synechococcus 7002-LL. Cells were lysed in 0.75 M K-phosphate buffer, pH 7.0, and the soluble fraction of the lysate was separated into fractions by sucrose density ultracentrifugation in 0.75 M K-phosphate (see inset gradient at left). Absorbance (a) and fluorescence emission (b, λex = 590 nm) spectra were collected for fractions F1, F2, and F3, which are colored with black, red, and blue lines, respectively. No FRL absorbance was detected in PBS. F2 had a small absorbance feature at 708 nm, perhaps due to cross-contamination with F1. F1 was likely composed of dissociated PBPs (suppl. Table S3) and had very intense absorbance with a maximum at 708 nm resulting from ApcD4-ApcB3
Thylakoid complexes were also isolated from strain Synechococcus 7002-LL (Fig. 7) and compared to similar fractions from cells of strain Synechococcus 7002-apcD4B3 (Fig. 8) and the wild type (suppl. Fig. S7). Unlike Synechococcus A1463, the only fraction isolated from the thylakoids of Synechococcus 7002-LL that had a distinct FRL absorbance feature was fraction F3, corresponding to trimeric PSI (Fig. 7, F3). On the other hand, no FRL absorbance was observed in thylakoid fractions from cells expressing only apcD4-apcB3 (Compare Figs. 7 and 8). The overlapping fluorescence emission from Synechococcus 7002 PSI at 715 nm and that from fractions containing ApcD4-ApcB3 at 716 nm obviously complicates the interpretation of the fluorescence data for these fractions. However, it was notable that the PSI fraction from Synechococcus 7002-LL also had a strong emission at 672 nm when excited at 440 nm that was not observed in PSI complexes from the wild-type strain or strain Synechococcus 7002-apcD4B3 (Figs. 8 and suppl. Fig. S7, F3). Tryptic peptide fingerprinting of fractions F2, F3, and F4 revealed the presence of ApcD4 and ApcB3 in every fraction with the highest abundance of detected peptides in fraction F3, but no peptides from IsiX were detected in any fraction (Table 3).
Absorbance and fluorescence emission spectra of complexes extracted from Synechococcus 7002-LL thylakoid membranes. The solubilized thylakoid membranes were separated into four fractions, F1, F2, F3, and F4, by sucrose density ultracentrifugation (see inset gradient). The absorbance spectra (a) and corresponding fluorescence emission spectra (b λex = 440 nm; c λex = 590 nm) are shown with black, red, blue, and magenta lines, respectively. FRL absorbance was only detected in F3, which corresponds to trimeric PSI potentially associated with ApcD4-ApcB3 (Table 3). F4 has two equally prominent fluorescence features at 672 nm and 717 nm when Chls are preferentially excited at 440 nm (b blue line). Trimeric PSI from Synechococcus A1463 had no FRL absorbance or 672 nm fluorescence emission feature, although the fractions immediately above it, F3-4, F4-1, and F4-2 (Fig. 4), were similar to F3 from this gradient in having FRL absorbance at 709 nm and fluorescence emission features at both ~ 672 nm and ~ 717 nm (λex = 440 nm)
Absorbance and fluorescence emission spectra of complexes extracted from the thylakoid membranes of Synechococcus 7002 expressing apcD4 and apcB3. The solubilized thylakoid membranes were separated into four fractions, F1, F2, F3, and F4, by sucrose density ultracentrifugation (see inset gradient). The absorbance spectra (a) and corresponding fluorescence emission spectra (b λex = 440 nm; c λex = 590 nm) are shown with black, red, blue, and magenta lines, respectively, while the absorbance of whole cells prior to lysis is shown in green. FRL absorbance was present in whole cells (WC, green line) but no pronounced FRL absorbance was observed for any thylakoid fraction. The absorbance and fluorescence emission spectra of the thylakoid complexes closely resemble those isolated from wild-type cells (suppl. Fig. S7)
RT-PCR analysis of transcription from the LoLiP gene cluster
The inability to inactivate genes in Synechococcus A1643 limits the ability to demonstrate functions for genes in the lcyA-apcD4-apcB3-isiX gene cluster. Other than colocalization with genes involved in light harvesting in the genomes of LLEs, there is no evidence that LcyA is functionally associated with the other genes/proteins in the LoLiP gene cluster. The gene arrangement suggests that these four genes could form an operon (suppl. Fig. S1), and thus the four genes might be cotranscribed. To test this possibility, reverse-transcription PCR analysis was performed using RNA isolated from cells grown in LL as the template and oligonucleotide primers (suppl. Table S1) designed to amplify the three intergenic regions as well as a region spanning lcyA-isiX. Demonstrating that there was no contaminating DNA in the RNA template, no amplicons were obtained when reverse transcriptase was omitted from the PCR reactions (suppl. Fig. S11). However, amplicons were obtained for each of the intergenic regions as well as the four-gene operon. These results show that lcyA, apcD4, apcB3, and isiX are cotranscribed, although transcripts could subsequently be processed to produce more stable segments. These findings strengthen the hypothesis that lcyA is the CBCR photoreceptor controlling the LoLiP response.
Heterologous expression of two GAF domains from the putative CBCR unique to LL-ecotypes
Repeated attempts to express full-length LcyA in both E. coli and Synechoocccus 7002 were unsuccessful. Thus, segments of the lcyA gene encoding its two GAF domains were separately expressed in E. coli with polyhistidine tags and added start and stop codons for individual characterization. Affinity purification of the N-terminal GAF domain, GAF1, from an E. coli strain encoding genes for the synthesis of phycocyanobilin resulted in a blue-colored polypeptide with broad absorbance band (λmax ~ 615 nm) and a narrow fluorescence emission band (λmax ~ 645 nm) upon excitation with 590 nm light (Fig. 9). Zinc-enhanced fluorescence emission of the denatured GAF1 polypeptide after SDS-PAGE confirmed that the phycocyanobilin chromophore was covalently attached to the GAF1 polypeptide, consistent with the hypothesis that LcyA is a photoreceptor. The GAF1 domain contains two cysteine residues (Fig. 9). Changing the first cysteine residue in GAF1 to alanine (C763A) resulted in a colorless polypeptide that did not bind phycocyanobilin. However, when the second cysteine was changed to alanine (C796A), the GAF1 protein retained the ability to bind phycocyanobilin covalently (data not shown).
Characterization of the first GAF domain (GAF1) from LcyA, the CBCR potentially controlling LoLiP gene expression in LL-adapted Synechococcus strains. a Domains of LcyA are depicted in the cartoon (top), with GAF1 highlighted in yellow, and the amino acid sequence of GAF1 from LcyA in Synechococcus A1463 is shown with Cys residues in red font. The N-terminal Cys residue in GAF1 (amino acid 763) is the site of chromophore attachement (see text). Recombinant GAF1 (expected size ~ 20 kDa) was heterologously co-expressed in E. coli with the genes for phycocyanobilin synthesis (pcyA and hox1) and a blue polypeptide was purified by IMAC before acquisition of absorbance (solid line) and fluorescence spectra (dashed line, λex = 590 nm). An SDS-PAGE gel stained first with 25 mM ZnSO4 (Zinc) and then Coomassie (Coo) is shown in the inset. A fluorescent band was visible in the zinc-stained gel under UV light, indicating covalent binding of phycocyanobilin (Berkelman and Lagarias 1986)
The GAF2 polypeptide did not bind phycocyanobilin after heterologous expression in an E. coli strain producing PCB. The protein sequence of the second GAF domain (GAF2) contained a CxxCxxCP motif similar to those found in proteins that bind a [4Fe-4S] cluster (Fig. 10), and so the plasmid encoding the GAF2 gene segment was transformed into a strain of E. coli engineered for improved expression of iron-sulfur cluster proteins, E. coli BL21(DE3) ΔiscR. After purification from inclusion bodies and subsequent chemical reconstitution of the iron-sulfur cluster under anoxic conditions, the brown-colored GAF2 polypeptide exhibited a broad absorbance band around 420 nm, which slowly disappeared after the addition of oxic buffer (suppl. Fig. S8). This behavior is consistent with the oxidation of a reduced iron-sulfur cluster. The EPR spectrum of the as-purified, reduced GAF2 polypeptide resembled that of proteins with an axial [4Fe-4S] cluster (Fig. 10). Redox titration of the purified and reconstituted GAF2 polypeptide corroborated the presence of a [4Fe-4S] cluster with a midpoint redox potential of − 554 ± 9 mV (suppl. Fig. S9).
Characterization of the second GAF domain (GAF2) from LcyA. a Domains of LcyA are depicted in the diagram, and the amino acid sequence of GAF2 from LcyA in Synechococcus A1463 is shown with Cys residues in red font. From its amino acid sequence, this domain was predicted to bind an iron-sulfur cluster. Recombinant GAF2 was heterologously produced in E. coli, solubilized from inclusion bodies, and purified by IMAC prior to chemical reconstitution of the iron-sulfur cluster (see Methods). b After anaerobic concentration and desalting, the brown-colored protein was analyzed by SDS-PAGE (see inset image; expected size ~ 18 kDa). EPR spectroscopy detected an axial signal with g values 2.04 and 1.93, characteristic of a single [4Fe-4S] cluster (Walters and Golbeck 2018). EPR parameters were as follows: microwave power, 2 mW; microwave frequency, 9.38 GHz; modulation amplitude, 1 mT; modulation frequency, 100 kHz; and temperature, 13 K
Comparison of transcript abundances in Synechococcus A1463 cells grown in HL and LL
A starter culture of Synechococcus A1463 was grown under 500 μmol photons m−2 s−1 for three weeks (OD680 ~ 1.0) and divided into two subcultures, each of which was diluted by half with fresh growth medium. One subculture was maintained at 500 μmol photons m−2 s−1 while the other was transferred to an irradiance of 10 μmol photons m−2 s−1 (suppl. Fig. S10). After 72 h in these HL and LL conditions, the cells from each culture were separately harvested, and total RNA was extracted. RNA sequencing of the LL and HL samples indicated that transcript abundances for several genes were higher in the LL-grown cells compared to the HL-grown cells. For example, transcripts for phycocyanin genes (cpcA2, cpcB2, cpcB1) were 3- to 4-fold higher in abundance in the LL cells. This finding agrees with the higher content of phycocyanin in LL cells (Fig. 1) Transcripts from the genes of the LoLiP gene cluster were also somewhat more abundant in cells grown in LL, although less obviously so (Table 4). In this experiment, only transcripts for apcD4 were more than two-fold more abundant in cells grown in LL. Another interesting observation was that transcripts for several genes associated with assembly of functional nitrogenase were more abundant in LL-grown cells (Table 4).
Discussion
LoLiP and FaRLiP share several properties, the most notable being the development of FRL absorbance when some cyanobacterial cells are grown in LL or FRL, respectively (Gan et al. 2014; Nowack et al. 2015; Olsen et al. 2015; Ho et al. 2016). Both types of photoacclimation involve distinctive gene clusters that encode FRL-absorbing APs, Chl-binding proteins, and CBCR or phytochrome photoreceptor-like proteins with one or two GAF domains (Gan et al. 2014; Olsen et al. 2015; Ho et al. 2017a; Soulier et al. 2020). Prior to this study, no specific linkage had been established between the products of the LoLiP gene cluster and the FRL absorbance that appears when LLE cells are grown under LL, although it had been shown that both processes likely involve FRL-absorbing forms of allophycocyanin (Soulier et al, 2020; Soulier and Bryant 2021). Little was known about the LcyA and IsiX proteins in the cluster beyond the predicted homology of the gene products to other proteins. Regulation of the LoLiP genes remains a mystery, in large part because no genetic system exists in Synechococcus A1463 to facilitate such studies. This study showed that the products of the LoLiP gene cluster are cotranscribed and are responsible for the FRL absorbance that is observed when LLEs are grown under LL conditions. It also showed that these products are found together with thylakoid membrane complexes and not with phycobilisomes. It also ruled out several other possible regulatory signals of lcyA, apcD4, apcB3, and isiX in Synechococcus A1463.
Growth conditions resulting in increased absorbance of FRL by Synechococcus A1463
The appearance of the FRL absorbance at ~ 712 nm in Synechococcus A1463 was monitored after cell growth under several conditions. Three growth conditions led to increased FRL absorbance: (1) growth in LL (Fig. 1); (2) growth in FRL (suppl. Fig. S3); and (3) allowing cells to settle and grow on the bottom of the flask or on its sides as biofilms in the absence of agitation (suppl. Fig. S2). Self-shading occurs in still cultures, and this can readily result in LL conditions for cells that are not directly in the light path. Moreover, the light gradient imposed by self-shading should mimic that experienced by cells in a microbial mat. FRL becomes enriched in situ as a function of mat depth because longer wavelengths of light are more weakly scattered and because FRL is not absorbed by most cyanobacteria that only produce Chl a, which absorbs very weakly beyond 700 nm (Gan et al. 2014; Gan and Bryant 2015). This applies to cells experiencing self-shading in cultures as well. Thus, unagitated cultures produce conditions that simulate both LL and FRL conditions for shaded cells. To differentiate between the effects of LL and FRL on the absorbance of unstirred cells, a culture of Synechococcus A1463 was placed in FRL. This initially resulted in an increase in the amplitude of the FRL absorbance feature as well as an increase in the absorbance associated with PBPs (~ 618 nm; Fig. 2b) and a decrease in amplitude of the absorbance of the photosystems (~ 680 nm). Unlike cells capable of FaRLiP, however, Synechococcus A1463 was unable to grow in FRL, and the cells slowly died. Interestingly, FRL absorbance did not increase in continuously agitated cells grown in FRL (suppl. Fig. S13). These results suggest that FRL alone is insufficient to activate transcription of the LoLiP genes and that other environmental factors, such as irradiance level, oxygen concentration, or other unknown factors are also required.
Isolation of proteins with FRL absorbance from Synechococcus A1463
Separation of complexes from cell lysates demonstrated that the FRL absorbance in Synechococcus A1463 cells grown in LL was mostly associated with the thylakoid membranes (suppl. Fig. S2). The chlF gene encoding Chl f synthase (Ho et al. 2016) does not occur in LLEs of thermophilic Synechococcus spp., and Synechococcus A1463 only synthesizes Chl a, so the presence of FRL absorbance in green fractions from sucrose density separation of thylakoids was unexpected. Because it was previously demonstrated that the LoLiP-associated AP, ApcD4-ApcB3, is a water-soluble FRL-AP (λmax ~ 710 nm) (Soulier et al. 2020; Soulier and Bryant 2021), PBS were expected to contribute to the FRL absorbance observed for whole cells. However, PBS in Synechococcus A1463 were completely devoid of FRL absorbance, which was only present in the upper fractions of sucrose density gradients prepared with 0.75 M K-phosphate buffer to stabilize PBS. Those fractions correspond to free PBPs or dissociated PBP complexes that are much smaller and less dense than PBS (Fig. 3a, F1).
These observations were consistent with results from heterologous expression of apcD4-apcB3-isiX in the model cyanobacterium, Synechococcus 7002. Although the FRL-AP proteins were clearly produced and accumulated in cells, the PBS of this strain did not include both ApcD4 and ApcB3 (Fig. 6). Tryptic peptide fingerprinting of PBP fractions isolated from both Synechococcus A1463 and Synechococcus 7002-LL demonstrated that the FRL-absorbing AP subunits accumulated predominantly in the uppermost fractions of sucrose gradients (suppl. Tables S2 and S4). The absorbance and low-temperature (77 K) fluorescence emission spectra of FRL-absorbing PBP fractions from high-phosphate sucrose gradients separating Synechococcus A1463 PBP complexes (Fig. 3) are distinct from the bicylindrical core complexes isolated from FaRLiP organisms such as Synechococcus sp. PCC 7335, which absorb maximally in the far-red (~ 710 nm) and have a single, prominent far-red fluorescence emission peak at about 730 nm (Ho et al. 2017b; Soulier et al. 2020). Thus, the FRL-absorbing AP from the LoLiP gene cluster does not assemble into complexes like those formed by the FRL-absorbing APs from the FaRLiP gene cluster. Those FRL-APs assemble into either bicylindrical core complexes or PBS cores depending on the organism (Gan et al. 2014; Ho et al. 2017b; Herrera-Salgado et al. 2018; Soulier et al. 2020).
Sucrose gradients prepared with low ionic strength buffer supplemented with detergent were used to separate complexes from solubilized thylakoid membranes isolated from Synechococcus A1463 cells grown in LL. The FRL absorbance in such gradients was associated with a fraction containing a mixture of monomeric and dimeric PSI and PSII (suppl. Fig. S5, F3). Due to the complex mixture of proteins present, fractions of interest were dialyzed to remove the sucrose prior to separation on a second sucrose density gradient, which resulted in improved resolution (Fig. 4). Tryptic peptide fingerprinting of FRL-absorbing fractions from the second round of sucrose density gradient separation consistently showed the co-occurrence of ApcD4, ApcB3, and IsiX (Table 1). Peptides from some of these proteins were identified in other fractions as well, but not together and not with equivalent abundance (as estimated from the % coverage and number of unique peptides detected).
The presence of FRL absorbance in greenish-blue fractions from sucrose gradients prepared with low ionic strength buffer suggests the association of ApcD4-ApcB3 with a Chl a-binding protein (Table 1). In some cyanobacterial strains, the linker polypeptide CpcL permits direct attachment of PBP rod complexes to PSI (Kondo et al. 2007; Watanabe et al. 2014; Liu et al. 2019), but the absence of a cpcL-like gene in LLEs necessitates another explanation. This explanation probably is found in the LoLiP operon itself, which encodes a homolog of iron-stress induced protein A (IsiA). The paralogous protein, denoted IsiX, shares ~ 56% protein sequence identity with IsiA from Synechococcus A1463 and is likewise predicted to be a Chl a-binding protein. Two complexes formed from ApcD4, ApcB3, and IsiX were observed after CN-PAGE analysis of the proteins in sucrose gradient fraction F3-4 (Fig. 4), providing strong evidence for the occurrence of such a complex. The synthesis of IsiX was not regulated by the iron status of cells (data not shown; see Soulier 2021). IsiX could potentially form a ring or partial-ring structure around one or both photosystems, as IsiA does around PSI (Boekema et al. 2001; Shen et al. 2016; Akita et al. 2020). Together with ApcD4-ApcB3, such a complex could enhance light harvesting under LL conditions. Further experimentation will be required to demonstrate if this is truly the case.
Heterologous production of LoLiP proteins in Synechococcus 7002
As observed in LL-grown cells of Synechococcus 1463, heterologous expression of the apcD4-apcB3-isiX operon in Synechococcus 7002-LL resulted in accumulation of FRL-absorbing proteins in the uppermost soluble PBP fraction from high-phosphate sucrose gradients but not in PBS (Fig. 6). However, the uppermost blue fraction in those gradients was significantly more enriched in FRL-absorbing ApcD4-ApcB3 than the equivalent fraction from Synechococcus A1463 (Fig. 3, F1). Localization of FRL-absorbing proteins in thylakoid complexes differed between the two strains: trimeric PSI had the most obvious FRL absorbance feature of the thylakoid complexes isolated from Synechococcus 7002-LL (Fig. 7, F3), but in Synechococcus A1463 FRL-absorbing proteins were mostly present in fractions less dense than trimeric PSI and containing both PSII dimers and PSI monomers (Fig. 4, F3-4, F4-1, F4-2; suppl. Fig. S5, F3). Unlike Synechococcus A1463, no IsiX was detected in any sucrose gradient fraction from Synechococcus 7002-LL (Table 3), even though the nucleotide sequence of the inserted operon was determined to be free of mutations, and the adjacent apcD4 and apcB3 genes were clearly expressed.
ApcD4 and ApcB3 were associated with an FRL-absorbing thylakoid fraction isolated from Synechococcus 7002-LL (Table 3, F3 containing PSI). However, when only apcD4 and apcB3 were heterologously expressed in Synechococcus 7002, very little FRL absorbance was observed in thylakoid complexes (Fig. 8a), and PSI was missing the pronounced fluorescence emission at 672 nm (440 nm; Fig. 8b) that was present in Synechococcus 7002-LL (Fig. 7b). The presence of IsiX with ApcD4-ApcB3 in FRL-absorbing fractions from Synechococcus A1463 (Tables 1 and suppl. Table S3) supports its involvement in a putative, light-harvesting complex incorporating ApcD4 and ApcB3 trimers and associated with one or both photosystems, but we were not successful in identifying such a complex after heterologous expression of apcD4, apcB3, and isiX in Synechococcus 7002. Further studies will be required to explain these observations.
GAF domains from LcyA, the putative photoreceptor of LoLiP
The putative LoLiP CBCR, LcyA, is homologous to photosensitive histidine kinases that are often involved in photoacclimation responses. LcyA has four PAS domains, two GAF domains, and a histidine kinase domain at its C-terminus (Figs. 9 and 10). Production of LcyA in Synechococcus A1463 cells was confirmed by detection of tryptic peptides (Table 1, F3-1). However, the absence of a genetic system and the inability to express the full-length protein (1,247 amino acids) heterologously made it difficult to determine its function. The demonstration that lcyA, apcD4, apcB3, and isiX are cotranscribed provides circumstantial evidence that the proteins encoded by the LoLiP gene cluster are functionally interconnected. The potential sensing function(s) of LcyA was evaluated by heterologous expression and characterization of its individual GAF domains.
The N-terminal GAF domain, GAF1, binds a phycocyanobilin chromophore covalently and absorbs light around 615 nm (Fig. 9), identifying LcyA as a potential photoreceptor that might respond to light wavelength or intensity. While exposure to FRL resulted in an increase in FRL absorbance of cells, FRL absorbance in whole cells did not increase after growth in red light (suppl. Fig. S4), which better complements the absorbance of the recombinant GAF1 polypeptide as isolated. Photoconversion of GAF1 was not observed upon illumination of a solution containing the GAF1 polypeptide with orange light (620 nm) or FRL (700 nm), as would be expected for the chromophore-binding GAF domain of a CBCR photoreceptor that responds to either condition (Ikeuchi and Ishizuka 2008). However, if LcyA acts as a sensor of power (i.e., light intensity), then it is possible that the GAF1 polypeptide reverted to its 615 nm-absorbing state too quickly to detect such a change under our experimental conditions (Rockwell et al. 2012; Fushimi et al. 2016, 2017; Hasegawa et al. 2018). By binding a phycocyanobilin chromophore, however, LcyA has the capacity to function as a sensor of light wavelength and/or intensity, and it is possible that the spectroscopic properties of the GAF1 domain alone differ from those of full-length LcyA.
After heterologous expression and chemical reconstitution, the GAF2 domain of LcyA was shown to coordinate a [4Fe-4S] cluster (Figs. 10 and suppl. Fig. S9). To our best knowledge, LcyA is the first CBCR photoreceptor that also binds an iron-sulfur cluster. Regulatory iron-sulfur proteins in bacteria sense changes in diverse conditions including iron concentration, oxygen concentration, and redox potential (Crack et al. 2012; Ludwig et al. 2014). Because iron and oxygen concentration serve as regulators of gene expression in the mat environment (Bhaya et al. 2007; Liu et al. 2011), it is possible that the [4Fe-4S] cluster of LcyA might sense changes in one of these environmental conditions. It is also possible that the [4Fe-4S] cluster in LcyA is sensitive to changes in redox potential, as described for numerous regulatory proteins binding iron-sulfur clusters including OrpR, a sigma-54-dependent transcriptional activator with a PAS domain with a [4Fe-4S] cluster, in Desulfovibrio vulgaris (Fiévet et al. 2021) and the Chloroplast Sensor Kinase of plants and cyanobacteria. The latter coordinates an iron-sulfur cluster with its GAF domain much like LcyA (Ibrahim et al. 2020). Although the precise function of LcyA is unknown, the coordination of an iron-sulfur cluster by a probable photoreceptor is a novel discovery that expands the already remarkable diversity of photoreceptors identified in cyanobacteria.
Regulation of genes and gene products associated with LoLiP
In the microbial mats of Mushroom Spring, transcripts for apcD4, apcB3, and isiX are most abundant in the early morning (8:00–10:00 AM) and afternoon-evening (3:00–5:30 PM), which correspond to periods of LL, anoxia, and nitrogen fixation for Synechococcus spp. (Olsen et al. 2015). Together with the observation that FRL absorbance and fluorescence emission increased after growth in low irradiance (Fig. 1; Nowack et al. 2015), it seems reasonable to assume that these genes are regulated by light intensity. However, some findings in this study are seemingly inconsistent with this hypothesis. Dispersed cells did not develop the FRL absorbance feature even after growth in LL (10 μmol photons m−2 s−1; suppl. Fig. S2). This observation, together with the result that transcript abundances of the four LoLiP genes (lcyA, apcD4, apcB3, and isiX) were only slightly higher in LL-grown cells (Table 4), suggests that transcription of the LoLiP genes is dependent upon more than one parameter and that the appropriate conditions are difficult to replicate so far under laboratory conditions. Although the GAF1 domain binds a chromophore and may be able to respond to light intensity and/or wavelength, the coordination of an iron-sulfur cluster by the second GAF domain of LcyA suggests that LcyA has an additional sensing function. What this could be remains a mystery for now, but it is possible that multiple conditions must be met to strongly induce the expression of LoLiP genes and/or the accumulation of their products.
The accumulation of FRL-absorbing proteins under growth conditions where LL and/or FRL occur makes physiological sense in the context of the microbial mat. Cells at the depths from which Synechococcus A1463 and other LLEs were isolated would receive mostly low-intensity light that is strongly enriched in wavelengths beyond 700 nm. A few trends emerge on the level of LoLiP protein accumulation in Synechococcus A1463 that corroborate this conclusion. Specifically, the FRL-absorbing proteins, known to include minimally ApcD4-ApcB3 from the cluster and perhaps IsiX as well, accumulate at low irradiance (Fig. 1). However, under the laboratory conditions tested, transcript levels for the genes of the LoLiP cluster were only modestly higher in LL-grown cells. It is possible that post-transcriptional regulation is more important in this case. Alternatively, FRL (Fig. 2) and some factor(s) sensed by the Fe-S cluster bound to the GAF2 domain might be most important for the transcription of the LoLiP gene cluster. Further studies will be required to improve the laboratory growth conditions to facilitate the study of this phenomenon.
Conclusions
FRL absorbance is the hallmark of the LoLiP photoacclimation response in LLEs of thermophilic Synechococcus spp. from Mushroom Spring in Yellowstone National Park. Through analysis of proteins and complexes from the LLE Synechococcus A1463, a source was defined for this FRL absorbance: ApcD4-ApcB3, a PBP encoded by a four-gene cluster that also contains genes for a putative photoreceptor and the putative Chl a-binding protein IsiX. LcyA, the CBCR photoreceptor and histidine kinase encoded by the LoLiP gene cluster, was shown to bind both a phycocyanobilin chromophore and a [4Fe-4S] cluster, potentially enabling it to respond to numerous environmental factors in addition to light wavelength and/or intensity. Several environmental conditions that could be sensed by LcyA, namely red light, FRL, LL, anoxia, and iron starvation, were monitored for accumulation of ApcD4-ApcB3 and the other products of the cluster. Of these, only LL and FRL conditions resulted in an increase in FRL absorbance and thus the accumulation of ApcD4-ApcB3 in Synechococcus A1463. RNA sequencing demonstrated only a modest increase in transcript abundance for genes of the LoLiP cluster in LL-grown cells (compared to HL-grown cells). It is thus possible that post-transcriptional effects ensure accumulation of LoLiP gene products in LL and FRL conditions. The properties and function of IsiX remain elusive due to an inability to express this protein heterologously or purify the native protein completely. However, repeated detection of IsiX together with ApcD4 and ApcB3 in complexes derived from FRL-absorbing thylakoids suggests that these three proteins form a novel complex when Synechococcus A1463 cells undergo LoLiP. Future studies will be directed towards gaining further structural and biochemical details for this complex.
References
Akhtar MK, Jones PR (2008) Deletion of iscR stimulates recombinant clostridial Fe-Fe hydrogenase activity and H2-accumulation in Escherichia coli BL21(DE3). Appl Microbiol Biotechnol 78:853–862. https://doi.org/10.1007/s00253-008-1377-6
Akita F, Nagao R, Kato K et al (2020) Structure of a cyanobacterial photosystem I surrounded by octadecameric IsiA antenna proteins. Commun Biol 3:232. https://doi.org/10.1038/s42003-020-0949-6
Allewalt JP, Bateson MM, Revsbech NP et al (2006) Effect of temperature and light on growth of and photosynthesis by Synechococcus isolates typical of those predominating in the Octopus Spring microbial mat community of Yellowstone National Park. Appl Environ Microbiol 72:544–550. https://doi.org/10.1128/AEM.72.1.544-550.2006
Becraft ED, Wood JM, Rusch DB et al (2015) The molecular dimension of microbial species: 1. Ecological distinctions among, and homogeneity within, putative ecotypes of Synechococcus inhabiting the cyanobacterial mat of Mushroom Spring, Yellowstone National Park. Front Microbiol 6:590. https://doi.org/10.3389/fmicb.2015.00590
Berkelman TR, Lagarias JC (1986) Visualization of bilin-linked peptides and proteins in polyacrylamide gels. Anal Biochem 156:194–201. https://doi.org/10.1016/0003-2697(86)90173-9
Bhaya D, Grossman AR, Steunou A-S et al (2007) Population level functional diversity in a microbial community revealed by comparative genomic and metagenomic analyses. ISME J 1:703–713. https://doi.org/10.1038/ismej.2007.46
Biswas A, Vasquez YM, Dragomani TM et al (2010) Biosynthesis of cyanobacterial phycobiliproteins in Escherichia coli: chromophorylation efficiency and specificity of all bilin lyases from Synechococcus sp. strain PCC 7002. Appl Environ Microbiol 76:2729–2739. https://doi.org/10.1128/AEM.03100-09
Boekema EJ, Hifney A, Yakushevska AE et al (2001) A giant chlorophyll-protein complex induced by iron deficiency in cyanobacteria. Nature 412:745–748. https://doi.org/10.1038/35089104
Brock TD (1978) Thermophilic microorganisms and life at high temperatures. Springer-Verlag, New York
Brock TD (1985) Life at high temperatures. Science 230:132–138. https://doi.org/10.1126/science.230.4722.132
Crack JC, Green J, Hutchings MI et al (2012) Bacterial iron–sulfur regulatory proteins as biological sensor-switches. Antioxid Redox Signal 17:1215–1231. https://doi.org/10.1089/ars.2012.4511
Dubbs JM, Bryant DA (1991) Molecular cloning and transcriptional analysis of the cpeBA operon of the cyanobacterium Pseudanabaena species PCC 7409. Mol Microbiol 5:3073–3085. https://doi.org/10.1111/j.1365-2958.1991.tb01867.x
Fiévet A, Merrouch M, Brasseur G et al (2021) OrpR is a σ54-dependent activator using an iron-sulfur cluster for redox sensing in Desulfovibrio vulgaris Hildenborough. Mol Microbiol 116:231–244
Fushimi K, Enomoto G, Ikeuchi M, Narikawa R (2017) Distinctive properties of dark reversion kinetics between two red/green-type cyanobacteriochromes and their application in the photoregulation of cAMP synthesis. Photochem Photobiol 93:681–691. https://doi.org/10.1111/php.12732
Fushimi K, Rockwell NC, Enomoto G et al (2016) Cyanobacteriochrome photoreceptors lacking the canonical Cys residue. Biochemistry 55:6981–6995. https://doi.org/10.1021/acs.biochem.6b00940
Gan F, Bryant DA (2015) Adaptive and acclimative responses of cyanobacteria to far-red light. Environ Microbiol 17:3450–3465. https://doi.org/10.1111/1462-2920.12992
Gan F, Shen G, Bryant DA (2015) Occurrence of far-red light photoacclimation (FaRLiP) in diverse cyanobacteria. Life 5:4–24. https://doi.org/10.3390/life5010004
Gan F, Zhang S, Rockwell NC et al (2014) Extensive remodeling of a cyanobacterial photosynthetic apparatus in far-red light. Science 345:1312–1317. https://doi.org/10.1126/science.1256963
Gibson DG, Young L, Chuang R-Y et al (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345. https://doi.org/10.1038/nmeth.1318
Hasegawa M, Fushimi K, Miyake K et al (2018) Molecular characterization of DXCF cyanobacteriochromes from the cyanobacterium Acaryochloris marina identifies a blue-light power sensor. J Biol Chem 293:1713–1727. https://doi.org/10.1074/jbc.M117.816553
Heinnickel M, Shen G, Agalarov R, Golbeck JH (2005) Resolution and reconstitution of a bound Fe−S protein from the photosynthetic reaction center of Heliobacterium modesticaldum. Biochemistry 44:9950–9960. https://doi.org/10.1021/bi050588s
Herrera-Salgado P, Leyva-Castillo LE, Ríos-Castro E, Gómez-Lojero C (2018) Complementary chromatic and far-red photoacclimations in Synechococcus ATCC 29403 (PCC 7335). I: the phycobilisomes, a proteomic approach. Photosynth Res 138:39–56. https://doi.org/10.1007/s11120-018-0536-6
Ho M-Y, Shen G, Canniffe DP et al (2016) Light-dependent chlorophyll f synthase is a highly divergent paralog of PsbA of photosystem II. Science 353:aaf9178. https://doi.org/10.1126/science.aaf9178
Ho M-Y, Gan F, Shen G et al (2017a) Far-red light photoacclimation (FaRLiP) in Synechococcus sp. PCC 7335: I. Regulation of FaRLiP gene expression. Photosynth Res 131:173–186. https://doi.org/10.1007/s11120-016-0309-z
Ho M-Y, Gan F, Shen G, Bryant DA (2017b) Far-red light photoacclimation (FaRLiP) in Synechococcus sp. PCC 7335: II. Characterization of phycobiliproteins produced during acclimation to far-red light. Photosynth Res 131:187–202. https://doi.org/10.1007/s11120-016-0303-5
Ho M-Y, Soulier NT, Canniffe DP et al (2017c) Light regulation of pigment and photosystem biosynthesis in cyanobacteria. Curr Opin Plant Biol 37:24–33. https://doi.org/10.1016/j.pbi.2017.03.006
Ibrahim IM, Wu H, Ezhov R et al (2020) An evolutionarily conserved iron-sulfur cluster underlies redox sensory function of the chloroplast sensor kinase. Commun Biol 3:1–11. https://doi.org/10.1038/s42003-019-0728-4
Ikeuchi M, Ishizuka T (2008) Cyanobacteriochromes: a new superfamily of tetrapyrrole-binding photoreceptors in cyanobacteria. Photochem Photobiol Sci 7:1159–1167. https://doi.org/10.1039/B802660M
Karapetyan NV, Bolychevtseva YV, Yurina NP et al (2014) Long-wavelength chlorophylls in photosystem I of cyanobacteria: origin, localization, and functions. Biochem 79:213–220. https://doi.org/10.1134/S0006297914030067
Kim B-H, Oh H-M, Lee Y-K et al (2006) Simple method for RNA preparation from cyanobacteria. J Phycol 42:1137–1141. https://doi.org/10.1111/j.1529-8817.2006.00263.x
Klatt CG, Liu Z, Ludwig M et al (2013) Temporal metatranscriptomic patterning in phototrophic Chloroflexi inhabiting a microbial mat in a geothermal spring. ISME J 7:1775–1789. https://doi.org/10.1038/ismej.2013.52
Klatt CG, Wood JM, Rusch DB et al (2011) Community ecology of hot spring cyanobacterial mats: predominant populations and their functional potential. ISME J 5:1262–1278. https://doi.org/10.1038/ismej.2011.73
Kondo K, Ochiai Y, Katayama M, Ikeuchi M (2007) The membrane-associated CpcG2-phycobilisome in Synechocystis: a new photosystem I antenna. Plant Physiol 144:1200–1210. https://doi.org/10.1104/pp.107.099267
Liu H, Weisz DA, Zhang MM et al (2019) Phycobilisomes harbor FNRL in cyanobacteria. Mbio. https://doi.org/10.1128/mBio.00669-19
Liu Z, Klatt CG, Wood JM et al (2011) Metatranscriptomic analyses of chlorophototrophs of a hot-spring microbial mat. ISME J 5:1279–1290. https://doi.org/10.1038/ismej.2011.37
Liu Z, Klatt CG, Ludwig M et al (2012) “Candidatus Thermochlorobacter aerophilum:” an aerobic chlorophotoheterotrophic member of the phylum Chlorobi defined by metagenomics and metatranscriptomics. ISME J 6:1869–1882. https://doi.org/10.1038/ismej.2012.24
Ludwig M, Bryant DA (2011) Transcription profiling of the model cyanobacterium Synechococcus sp. strain PCC 7002 by next-gen (SOLiDTM) sequencing of cDNA. Front Microbiol 2:41. https://doi.org/10.3389/fmicb.2011.00041
Ludwig M, Pandelia M-E, Chew CY et al (2014) ChlR protein of Synechococcus sp. PCC 7002 is a transcription activator that uses an oxygen-sensitive [4Fe-4S] cluster to control genes involved in pigment biosynthesis. J Biol Chem 289:16624–16639. https://doi.org/10.1074/jbc.M114.561233
Nowack S, Olsen MT, Schaible GA et al (2015) The molecular dimension of microbial species: 2. Synechococcus strains representative of putative ecotypes inhabiting different depths in the Mushroom Spring microbial mat exhibit different adaptive and acclimative responses to light. Front Microbiol 6:626. https://doi.org/10.3389/fmicb.2015.00626
Ohkubo S, Miyashita H (2017) A niche for cyanobacteria producing chlorophyll f within a microbial mat. ISME J 11:2368–2378. https://doi.org/10.1038/ismej.2017.98
Olsen MT, Nowack S, Wood JM et al (2015) The molecular dimension of microbial species: 3. Comparative genomics of Synechococcus strains with different light responses and in situ diel transcription patterns of associated putative ecotypes in the Mushroom Spring microbial mat. Front Microbiol 6:604. https://doi.org/10.3389/fmicb.2015.00604
Rippka R, Deruelles J, Waterbury JB et al (1979) Generic assignments, strain histories and properties of pure cultures of cyanobacteria. Microbiology 111:1–61. https://doi.org/10.1099/00221287-111-1-1
Rockwell NC, Martin SS, Lagarias JC (2012) Red/green cyanobacteriochromes: sensors of color and power. Biochemistry 51:9667–9677
Saini MK, ChihChe W, Soulier N et al (2020) Caldichromatium japonicum gen. nov., sp. nov., a novel thermophilic phototrophic purple sulphur bacterium of the Chromatiaceae isolated from Nakabusa hot springs, Japan. Int J Syst Evol Microbiol 70:5701–5710. https://doi.org/10.1099/ijsem.0.004465
Sener MK, Lu D, Ritz T et al (2002) Robustness and optimality of light harvesting in cyanobacterial photosystem I. J Phys Chem B 106:7948–7960. https://doi.org/10.1021/jp020708v
Shen G, Gan F, Bryant DA (2016) The siderophilic cyanobacterium Leptolyngbya sp. strain JSC-1 acclimates to iron starvation by expressing multiple isiA-family genes. Photosynth Res 128:325–340. https://doi.org/10.1007/s11120-016-0257-7
Shen G, Canniffe DP, Ho M-Y et al (2019) Characterization of chlorophyll f synthase heterologously produced in Synechococcus sp. PCC 7002. Photosynth Res 140:77–92. https://doi.org/10.1007/s11120-018-00610-9
Soulier NT (2021) Light-harvesting proteins and complexes in cyanobacteria acclimated to far-red and low-light conditions. Doctoral dissertation, The Pennsylvania State University
Soulier N, Bryant DA (2021) The structural basis of far-red light absorbance by allophycocyanins. Photosynth Res 147:11–26. https://doi.org/10.1007/s11120-020-00787-y
Soulier N, Laremore TN, Bryant DA (2020) Characterization of cyanobacterial allophycocyanins absorbing far-red light. Photosynth Res. https://doi.org/10.1007/s11120-020-00775-2
Tank M, Thiel V, Ward DM, Bryant DA (2017) A panoply of phototrophs: an overview of the thermophilic chlorophototrophs of the microbial mats of alkaline siliceous hot springs in Yellowstone National Park, WY, USA. In: Hallenbeck PC (ed) Modern Topics in the Phototrophic Prokaryotes: Environmental and Applied Aspects. Springer International Publishing, Cham, pp 87–137. https://doi.org/10.1007/978-3-319-46261-5_3
Thiel V, Hügler M, Ward DM, Bryant DA (2017) The dark side of the Mushroom Spring microbial mat: life in the shadow of chlorophototrophs. II. Metabolic functions of abundant community members predicted from metagenomic analyses. Front Microbiol 8:943
Thiel V, Tank M, Bryant DA (2018) Diversity of chlorophototrophic bacteria revealed in the -omics era. Annu Rev Plant Biol 69:21–49. https://doi.org/10.1146/annurev-arplant-042817-040500
Thiel V, Wood JM, Olsen MT et al (2016) The dark side of the Mushroom Spring microbial mat: life in the shadow of chlorophototrophs. I. Microbial diversity based on 16S rRNA gene amplicons and metagenomic sequencing. Front Microbiol 7:919. https://doi.org/10.3389/fmicb.2016.00919
Toporik H, Khmelnitskiy A, Dobson Z et al (2020) The structure of a red-shifted photosystem I reveals a red site in the core antenna. Nat Commun 11:5279. https://doi.org/10.1038/s41467-020-18884-w
Tros M, Bersanini L, Shen G et al (2020) Harvesting far-red light: functional integration of chlorophyll f into Photosystem I complexes of Synechococcus sp. PCC 7002. Biochim Biophys Acta–Bioenerg 1861:148206. https://doi.org/10.1016/j.bbabio.2020.148206
Walters KA, Golbeck JH (2018) Expression, purification and characterization of an active C491G variant of ferredoxin sulfite reductase from Synechococcus elongatus PCC 7942. Biochim Biophys Acta - Bioenerg 1859:1096–1107. https://doi.org/10.1016/j.bbabio.2018.06.014
Ward DM, Ferris MJ, Nold SC, Bateson MM (1998) A natural view of microbial biodiversity within hot spring cyanobacterial mat communities. Microbiol Mol Biol Rev 62:1353–1370. https://doi.org/10.1128/MMBR.62.4.1353-1370.1998
Watanabe M, Semchonok DA, Webber-Birungi MT et al (2014) Attachment of phycobilisomes in an antenna–photosystem I supercomplex of cyanobacteria. Proc Natl Acad Sci USA 111:2512–2517. https://doi.org/10.1073/pnas.1320599111
Wittig I, Schägger H (2005) Advantages and limitations of clear-native PAGE. Proteomics 5:4338–4346. https://doi.org/10.1002/pmic.200500081
Xu Y, Alvey RM, Byrne PO et al (2011) Expression of genes in cyanobacteria: adaptation of endogenous plasmids as platforms for high-level gene expression in Synechococcus sp. PCC 7002. Methods Mol Biol 684:273–293. https://doi.org/10.1007/978-1-60761-925-3_21
Zhao C, Gan F, Shen G, Bryant DA (2015) RfpA, RfpB, and RfpC are the master control elements of far-red light photoacclimation (FaRLiP). Front Microbiol 6:1303. https://doi.org/10.3389/fmicb.2015.01303
Acknowledgements
DAB and JHG gratefully acknowledge support from the U. S. National Science Foundation (MCB-1613022). This research was also conducted under the auspices of the Photosynthetic Antenna Research Center (PARC), an Energy Frontier Research Center funded by the DOE, Office of Science, Office of Basic Energy Sciences under Award Number DE-SC 0001035 (DAB). Molecular graphics and analyses performed with UCSF-Chimera, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, were supported by NIH award P410GM103311.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that this research was conducted in the absence of any commercial, financial, or non-financial relationships that could be construed as a potential conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Soulier, N., Walters, K., Laremore, T.N. et al. Acclimation of the photosynthetic apparatus to low light in a thermophilic Synechococcus sp. strain. Photosynth Res 153, 21–42 (2022). https://doi.org/10.1007/s11120-022-00918-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-022-00918-7