Abstract
Cyanobacteria acclimate and adapt to changing light conditions by controlling the energy transfer between photosystem I (PSI) and II (PSII) and pigment composition. Photosynthesis is driven by balancing the excitation between PSI and PSII. To predict the detailed electron transfer flux of cyanobacteria, we refined the photosynthesis-related reactions in our previously reconstructed genome-scale model. Two photosynthetic bacteria, Arthrospira and Synechocystis, were used as models. They were grown under various spectral light conditions and flux balance analysis (FBA) was performed using photon uptake fluxes into PSI and PSII, which were converted from each light spectrum by considering the photoacclimation of pigments and the distribution ratio of phycobilisome to PSI and PSII. In Arthrospira, the FBA was verified with experimental data using six types of light-emitting diodes (White, Blue, Green, Yellow, Red1, and Red2). FBA predicted the cell growth of Synechocystis for the LEDs, excepting Red2. In an FBA simulation, cells used respiratory terminal oxidases and two NADH dehydrogenases (NDH-1 and NDH-2) to balance the PSI and PSII excitations depending on the light conditions. FBA simulation with our refined model functionally implicated NDH-1 and NDH-2 as a component of cyclic electron transport in the varied light environments.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Photosynthetic organisms transform sunlight to chemical energy (ATP) and reductant (NADPH) for carbon fixation. These organisms contain specific pigments that include chlorophyll (Chl) and carotenoids (Car). Cyanobacteria have specific antenna pigment–protein complexes called phycobilisomes (PBS), which contain phycoerythrin, phycocyanin, and allophycocyanin. PBSs increase the light-harvesting ability of cyanobacteria. The light energy is transferred from each PBS to Chl in photosystem I (PSI) and photosystem II (PSII) (Gantt 1981; Glazer 1984; Mimuro and Kikuchi 2003). These light-harvesting pigments have different absorbance wavelengths and allow survival under various light environments (Grossman et al. 1995; Hohmann-Marriott and Blankenship 2011; Rockwell et al. 2014; Duanmu et al. 2017).
Photosynthesis is driven by two electron transport pathways, linear electron transport (LET) and cyclic electron transport (CET). Regulation of the ratio between LET and CET is particularly important for controlling the redox and energy balance of the cell by regulating the synthesis ratio of ATP and NADPH (Mullineaux 2014). The LET and CET activities are modified by maintaining a suitable balance of excitation between PSI and PSII. Energy transfer between both PSI and PSII, the energy distribution from each PBS to PSI/PSII, and the pigment composition are key factors for controlling the activities of electron transfer pathways to adapt to light environmental conditions (Akimoto et al. 2012, 2013; Dall’Osto et al. 2015; Yokono et al. 2015; Ho et al. 2017; Duanmu et al. 2017). However, the mechanism by which cyanobacteria selectively use these complicated electron transfer pathways depending on a light environment remains unclear.
Model-based approaches are increasingly used in photosynthetic organisms to gain functional insights and to predict the metabolic processes (Boyle and Morgan 2009; Chang et al. 2011; Yoshikawa et al. 2011, 2017; Imam et al. 2015; Broddrick et al. 2016; Yuan et al. 2016; Zakhartsev et al. 2016). One of the modeling approaches, flux balance analysis (FBA), is a constraint-based modeling approach that is widely used in predicting metabolic fluxes based on the stoichiometry of reactions by assuming a steady state of metabolic reactions (Schilling et al. 1999; Feist et al. 2009; Orth et al. 2010). Briefly, in FBA, all reactions of the metabolic model are described by a stoichiometric formula. Assuming a pseudo-steady state of the metabolic reactions, the maximum and minimum ranges of the flux for each reaction are defined. The FBA simulates a steady-state flux distribution that optimizes an objective function, e.g., maximizing biomass production (Price et al. 2004). FBA does not treat dynamic behavior of the intracellular metabolite concentrations or the kinetic representation of enzyme reactions. The details of FBA were previously described (Feist et al. 2009; Schilling et al. 1999).
Once this set of steady-state fluxes is defined, optimization technique is used to evaluate the performance of the biological system under various environmental conditions. In this study, electron transfer fluxes in photosynthetic pathways were predicted under differential spectral conditions. Based on the input rate of photons and the ratios provided, all other fluxes are adjusted to maximize the growth rate. The predicted fluxes are able to be compared with each other and with experimental data. These in silico metabolic simulations predict the metabolic flux changes according to culture conditions, such as nutrient limitation, and different trophic conditions. In this study, the specific growth rate was used as the objective function to be maximized, with the assumption that cellular metabolism is self-organized to maximize specific growth rate. The cell growth rates obtained by the experiments were compared with simulation. As the quantitative measurement of the electron flow through photosystems including complicated cyclic pathways remains challenging, FBA is a promising way of revealing the flux distribution of the photosystem based on the mass balance constraints. Previously, analysis of the adaptation mechanism of photosynthetic pathways and electron transfer in LET and CET (Nogales et al. 2012; Vu et al. 2012; Qian et al. 2017) were reported; however, the models lack the details of photosynthetic and respiratory electron transports in photosynthetic pathways under different spectral light information.
In the present study, we considered two photosynthetic bacteria, Synechocystis sp. PCC 6803 and Arthrospira platensis, as models. Synechocystis is a unicellular cyanobacterium, which is widely used as a model organism for studying photosynthesis (Burnap and Sherman 1991; Chitnis and Chitnis 1993; Ikeuchi and Tabata 2001). Its genome has been sequenced (Fujisawa et al. 2017) and its electron transport pathways and photosynthesis components have been determined. Nevertheless, for accurate simulation of the electron fluxes under various lightning conditions, the determination of the energy transfer ratio from PBS to PSI and PSII is essential. The ratio measured by time-resolved fluorescence spectroscopy has been reported in the filamentous cyanobacterium Arthrospira, which has been widely cultured for commercial applications (Akimoto et al. 2013). Assuming that both cyanobacteria utilize common metabolic and photosynthetic pathways, information of the photosynthetic characteristic of Arthrospira was used to understand the selective use of electron flow in Synechocystis.
This study sought to predict the fluxes of photosynthetic and respiratory electron transports under various light environments. For this goal, we developed an updated model that included detailed information of photosynthesis and respiratory chain electron transport pathways. The reconstruction was based on the previous genome-scale metabolic model of Synechocystis sp. PCC 6803 (Yoshikawa et al. 2011). Initially, the reconstructed model was verified with experimental data of cell growth and photon absorption of Arthrospira. It was confirmed that the model predicted the specific growth rate of Arthrospira under different spectral light conditions. Moreover, it also predicted the cell growth of Synechocystis sp. PCC 6803 with the specific parameters of this strain for absorbing different spectral lights. By using this model, it was clarified that the selective role of photosynthetic electron transport components and regulation of LET and CET under different spectral light were controlled by the change in energy transfer ratio from PBS to PSI and PSII. The simulation results provide new insight into the understanding of the photosynthetic system of cyanobacteria in the acclimation and adaptation to various light environments.
Materials and methods
Development of the genome-scale metabolic model
A genome-scale metabolic model of Synechocystis sp. PCC 6803 was developed according to our previous model (Yoshikawa et al. 2011). Photosynthesis reaction pathway in the previous model was refined based on the information from various sources, such as public databases, literature, and genome sequences (Mitchell 1975; Rich 1988; Sacksteder et al. 2000; Pils and Schmetterer 2001; Mullineaux 2008; Lea-Smith et al. 2013; Peltier et al. 2016). We collected the annotation and pathway data from CyanoBase (Nakamura et al. 1998) and Kyoto Encyclopedia of Genes and Genomes (KEGG) (Kanehisa and Goto 2000). The updated stoichiometric formulas from R0001 to R0043 in Table S1 were added to our previous model as photosynthetic reactions.
FBA
The metabolic flux distribution of the genome-scale metabolic model of Synechocystis sp. PCC 6803 was calculated using FBA, as described previously (Yoshikawa et al. 2011). The specific growth rate was used as the objective function to be maximized in all simulations presented in this study. All simulations were performed using the MATLAB R2018a (MathWorks Inc., Natick, MA, USA) with COBRA Toolbox v3.0 (Becker et al. 2007; Heirendt et al. 2019) and the open-source GLPK software (http://glpkmex.sourceforge.net/), which is an application that solves linear programming. Since linear programming problems can present multiple solutions with an identical value for the objective function, the flux variability of each reaction was calculated (Mahadevan and Schilling 2003). In brief, after obtaining a maximum specific growth rate by FBA, each flux was maximized and minimized under the condition of maximized specific growth rate, to check the variability of fluxes.
In FBA simulation, photon incident rates of Arthrospira and Synechocystis cells were normalized to 42.6 mmol gDW−1 h−1 and 78 mmol gDW−1 h−1, respectively. The values are adjusted so that the simulated cell growth rate under yellow light condition is coincident with the experimental data of the growth rate in each of Arthrospira and Synechocystis culture. We estimated that these values are much fewer than 230 µmol m−2 s−1 irradiation light intensity because large amount of irradiation photon is transmitted and not absorbed by the cells.
Growth of Synechocystis
Before the growth experiments, cells of Synechocystis sp. PCC 6803 glucose-tolerant strain (GT) (Williams 1988) were grown in BG-11 medium (Rippka et al. 1979) at 32 °C under continuous white LED light (20 µmol m−2 s−1) in flasks by constant shaking at 120 rpm. In the growth experiment, the culture was initially inoculated with an inoculum from the preculture in stationary phase. The cells were grown in 200-mL flasks containing 50 mL BG-11 under continuous different spectral LED lights at 230 µmol m−2 s−1. Six different light sources were used to cultivate the cells: white LED (hereafter referred as to White) and five different single-color LEDs with following spectral profiles: a single peak at 470 nm (blue light, hereafter referred as to Blue), 530 nm (green light, Green), 590 nm (yellow light, Yellow), 630 nm (red light 1, Red1), and 680 nm (red light 2, Red2).
Absorption spectrum of cell suspension
Absorption spectra of the cell suspensions were measured according to the so-called opal glass method described previously (Toyoshima et al. 2016), with a translucent cuvette placed in front of the detector in order to minimize the light scattering effect.
Measurements of photosynthetic oxygen evolution and chlorophyll content
All the analytical methods were essentially identical to those described previously (Toyoshima et al. 2016). Cells were maintained at 30 °C for 5 min and the oxygen consumption rate (A) was recorded. Next, the cells were irradiated by the six different LEDs at 230 µmol m−2 s−1 for 3 min, and the oxygen evolution rate (B) was recorded. Photosynthetic oxygen evolution rate was calculated as (B + A). To determine the Chl content, a 200-µL culture was mixed with 800 µL acetone. After a brief centrifugation in a microfuge at top speed, the supernatant was used for spectrophotometry at 710 and 630 nm using a model DU800 spectrophotometer (Beckman Coulter, Brea, CA, USA). Notably, Synechocystis sp. PCC 6803 only has Chl a. Thus, the amount of chlorophyll is the amount of Chl a, which was determined as previously described (Porra et al. 1989).
Results and discussion
Refinement of photosynthesis in the genome-scale metabolic model
We refined the photosynthesis-related reactions in the genome-scale model of Synechocystis sp. PCC 6803, developed in our previous studies (Yoshikawa et al. 2011, 2017). Initially, we updated a draft model of photosynthesis and electron transfer pathways based on the information provided by genome annotation from databases such as CyanoBase (Nakamura et al. 1998) and KEGG (Kanehisa and Goto 2000) (Fig. 1, Table S1). The metabolic reactions, excluding photosynthetic reactions, were the same as the constructed model in our previous study (Yoshikawa et al. 2011).
Scheme of photosynthetic reaction flow in a genome-scale model of Synechocystis sp. PCC 6803. Red lettering indicates the reaction name in our model (see Table S1). Abbreviations: PC, phycocyanin; APC, allophycocyanin; ApcDEF, phycobilisome anchor protein DEF; hv550, photon 550 nm; hv620, photon 620 nm; hv650, photon 650 nm; hv670, photon 670 nm; hv680, photon 680 nm; hv700, photon 700 nm; Fd, ferredoxin; Fdrd, reduced ferredoxin; Fdox, oxidized ferredoxin; Trdrd, reduced thioredoxin; Trdox, oxidized thioredoxin; OCP, orange carotenoid protein; Mn, manganese cluster; Yz, tyrosine Z; Phe, pheophytin; QA, first quinone electron acceptor; QB, second quinone electron acceptor; PQH2, plastoquinol; SDH, succinate dehydrogenase; NDH-1, type I NADH dehydrogenase; NDH-2, type II NADH dehydrogenase; succ, succinate; fum, fumarate; CCM, CO2-concentrating mechanism; CytQi, cytochrome b6f Qi site; CytQo, cytochrome b6f Qo site; Cyt b, cytochrome b; FeS, Rieske iron-sulfur cluster; Cyt f, cytochrome f; Pc, plastocyanin; A1, phylloquinone; A0, chlorophyll a0; A1, phylloquinone A1; FeSx, iron-sulfur cluster X; FeSA, iron-sulfur cluster A; FeSB, iron-sulfur cluster B; FTR, ferredoxin:thioredoxin reductase; FNR, ferredoxin:NADP+ reductase; TH, NADPH: NAD+ transhydrogenase; Cox, cytochrome c oxidase; Flv1/3, flavodiiron protein 1/3; H, hydrogen iron in cytosol; H[t], hydrogen iron in thylakoid lumen
We added type I NADH dehydrogenase (NDH-1; R0033 in Table S1) (Bernát et al. 2011; Peltier et al. 2016), type II NADH dehydrogenase (NDH-2; R0035 in Table S1) (Howitt et al. 1999; Peltier et al. 2016; Huokko et al. 2017), and succinate dehydrogenase (SDH; R031 and R0032 in Table S1) as CET in our model. In the KEGG database, NDH-1 complex accepts the electrons from NADH. However, previous studies elucidated that ferredoxin (Fd) is the electron donor for cyanobacterial NDH-1 (Battchikova et al. 2011; Peltier et al. 2016, Schuller et al. 2019). Thus, we used Fd as the electron donor of the NDH-1 reaction.
Furthermore, the Q cycle in cytochrome b6f complex (cyt b6f) was previously described in detail (R0017, R0018, and R0019 in Table S1) (Mitchell 1975; Rich 1988; Sacksteder et al. 2000). Synechocystis sp. PCC 6803 contains three respiratory terminal oxidases (RTOs): aa3-type cytochrome c oxidase complex (Cox) (Schmetterer et al. 1994), cytochrome bd-quinol oxidase complex (Cyd) (Howitt and Vermaas 1998), and the alternative respiratory oxidase (ARTO) (Pils et al. 1997; Howitt and Vermaas 1998). In our model, the electron donors for Cox and Cyd were designated plastocyanin (Pc; R0031 in Table S1) and plastoquinone (PQ; R0037 in Table S1), respectively, based on a previous study (Pils and Schmetterer 2001). As Cyd and ARTO have similar function (Pils et al. 1997; Pils and Schmetterer 2001; Lea-Smith et al. 2013), these oxidases are integrated into Cyd in our model. HCO3– uptake reaction was described as an ATP-dependent reaction (R0050 in Table S1). All CO2 that was taken us was temporarily converted to HCO3– (R063 and R0034 in Table S1). The HCO3− was converted to CO2 again by carbonic anhydrase for utilization by metabolic enzymes, which was termed the “CO2 concentrating mechanism” (R213 in Table S1).
We described PBS (light-harvesting pigment is phycocyanin), PSII (light-harvesting pigment is chlorophyll P680), and PSI (light-harvesting pigment is chlorophyll P700) as the incident photon sites (R0001, R0010, and R0022 in Table S1, respectively). The change in energy transfers from PBS to PSII and PSI (previously described as “state transitions”; Mullineaux 2008) was represented as R0006 and R0007, respectively. Experimental data of the energy transfer ratio of PBS to PSI and PSII for six wavelength lights (PSI/PSII* in Table 1) were used for the determination of the flux ratio of R0006 (from PBS to PSII energy transfer reaction) and R0007 (from PBS to PSI energy transfer reaction) to simulate under each color light condition (PSI/PSII* = R0007 flux/R0006 flux).
The most important stoichiometric constraint in the model is that the flux of electrons from PSII through cyt b6f to PSI and the flux of incident photons to PSI must be balanced to R0023 (Plcred + P700oxd → Plcoxd + P700red) flux. This constraint makes the PSI/PSII excitation ratio important in determining the flux distribution in FBA simulation. By this constraint, the fluxes of CET and respiration are required to adjust the flux balance of PSII and PSI. Additionally, by defining the energy distribution from PBS to PSI/PSII, the model suggested the effects of state transition on cell proliferation. In the previous study involving the Chlamydomonas model (Chang et al. 2011), a light-modeling approach was implemented to allow the quantitative growth prediction for a given light source, resolving wavelength and photon flux. We also constructed a novel light-modeling approach that enables quantitative growth prediction for a given light source, considering the absorption spectra of cell and photon flux. Furthermore, to perform FBA simulation more practically, we constructed the model that allows representation of the state of the photosynthetic apparatus, such as the different energy distribution state from PBS to PSII and PSI, in addition to providing an accurate description of the stoichiometric formula of the components of photosynthetic electron transport, such as NDH-1 and NDH-2.
Verification of updated model and FBA simulation using experimental data
Using the refined model, FBA simulations under various spectral light conditions were done. To calculate the photon uptake fluxes into PSI and PSII from each light spectrum, the experimental data of the photoacclimation of pigments and the distribution ratio of PBS to PSI and PSII are required. As these experimental data are available in Arthrospira, initially, to verify the results of the FBA simulations of Arthrospira using the refined genome-scale model, we compared the specific growth rate obtained by the FBA simulations with the growth data obtained by the experiments in previous studies (Akimoto et al. 2012, 2013). In these studies, the growth and absorption spectra of Arthrospira cultured under the six input LED lights [white LED (White) and five different single-color LEDs with following spectral profiles: a single peak at 470 nm (Blue), 530 nm (Green), 590 nm (Yellow), 630 nm (Red1), and 680 nm (Red2)], were measured. For FBA simulation of Arthrospira, the incident photon ratio of PBS, PSII, PSI, and Car (Table 1) was set by combining the data of the spectral profile of LED lights (Akimoto et al. 2012) and the cellular pigments (chlorophyll, phycocyanin, and carotenoid) ratio (Fig. S1) (Lichtenthaler 1987; Shen et al. 1993; Chen and Blankenship 2011; Akimoto et al. 2012; Collins et al. 2012; Ghosh et al. 2016; Ho et al. 2017). We defined the photons of any wavelength being as the same photons. Excited states resulting from absorption of blue photons are degraded within subpicoseconds to the level of red photons due to the heat dissipation intramolecularly before they are used for photosynthesis, although the blue photons contain more energy than red photons due to the Planck–Einstein relation (Björn et al. 2009).
The spectrum of each input light and absorption spectrum of the cells were calculated for Arthrospira. Relative spectral overlaps for Blue, Green, Yellow, Red1, and White lights to that for Red2 light were determined as 1.00, 0.616, 0.560, 0.981, and 0.952, respectively. Light absorption by Arthrospira at different wave length is summarized in Table 1. The energy of photons absorbed by Car was not used as an incident photon flux value in our FBA simulation (R0044–R0048 in Table S1) because Car is a highly effective light-activated energy quencher, which diverts energy away from both photosystems and switches the photosynthetic outputs to heat in cyanobacteria (Mullineaux 2014; Kirilovsky 2015; Kirilovsky and Kerfeld 2016; Sonani et al. 2018).
The flux distribution of the photosynthesis and metabolic pathways of Arthrospira was simulated by FBA using an objective function to maximize cell growth (Table S1) under the different spectral lights. The photon incident rate of the FBA simulation was normalized to 42.6 mmol gDW−1 h−1. The values are adjusted so that the simulated cell growth rate under yellow light condition is coincident with the experimental data of the growth rate of Arthrospira culture, as shown in Fig. 2b. A value obtained by multiplying the photon incident rate (42.6 mmol gDW−1 h−1) and the incident photon ratio in Table 1 was set as the flux of each photon incident site [PBS, PSII, and PSI (R0039, R0041, and R0043 in Table S1, respectively)]. To evaluate the simulation performance, we plotted the correlation between the estimated and experimental specific growth rates (Fig. 2). In Fig. 2a, the cell absorption for each wavelength of light was assumed to be uniform. The energy transfer ratio from PBS to PSI and PSII was free. The simulation results of specific growth rates under the Yellow and Red1 light conditions, which were calculated as the same value as under Red2 light, were far from the experimental data. Figure 2b shows the simulation results of the specific growth rate with the data of the absorption spectra of cell suspensions and the energy transfer ratio from PBS to PSI and PSII. The specific growth rates predicted by our refined model revealed a good agreement with the experimental data under the Blue, Green, Yellow, Red1, and Red2 light conditions. Consequently, the refined genome-scale metabolic model was demonstrated to accurately simulate the specific growth rate under different spectral light conditions by considering the detailed information of photosynthesis pathways, absorbed photon energy at different wavelength, and the PBS to PSI/PSII ratios.
Comparison of the specific growth rate of Arthrospira between FBA simulation and experimental results under different spectral lights. The specific growth rates experimentally obtained in Akimoto et al. (2012) were compared with the simulation result of our refined model. Photon uptake rate was set to a specific value, in which the specific growth rate is equivalent in the simulation and experimental results under Yellow. (a) The simulation with the flat shape of absorption spectra of cell. The energy transfer ratio from PBS to PSI and PSII is not considered. (b) The simulation with the data of incident photon ratio obtained from our experimental cellular absorption spectra (Table 1) and the energy transfer ratio from PBS to PSI and PSII (PSI/PSII* in Table 1) were set as the parameter
Prediction of cell density of Synechocystis sp. PCC 6803 grown under different spectral light conditions
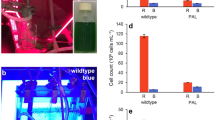
To apply the updated model to predict the growth of Synechocystis sp. PCC 6803 under the different spectral light conditions, cells were cultured using each LED (Fig. S2). Data of the growth and absorption spectra of the cells grown for 24 h under each spectral LED are presented in Fig. 3. As the photosynthetic apparatus and metabolic systems of Synechocystis and Arthrospira are not significantly different, the same model structure and stoichiometric equations were used for simulation of both organisms (Yoshikawa et al. 2015). Absorption of different spectral lights of Synechocystis was measured (Table 2). It was assumed that the energy transfer ratio for the six wavelength lights was the same as that obtained for Arthrospira and these values were used for simulation under each color light condition. Absorption peak of PBS (at 630 nm) under Yellow (a single peak at 590 nm) and Red2 (a single peak at 680 nm) lights was higher than those of other color lights (Fig. 3b). An increase in the Chl contents of the cells grown under Blue (a single peak at 470 nm) and Green (a single peak at 530 nm) lights (Fig. S3) was observed, which may have led to efficient photon absorption. Synechocystis sp. PCC 6803 was photoacclimatized (Ho et al. 2017) for each of the six kinds of LED lights. The cell growth at each wavelength was different, with slower growth under the Blue and Green light conditions and faster growth under the Yellow, Red1 (a single peak at 630 nm), Red2, and White light conditions. The dry cell weights of Synechocystis under the six kinds of LED light were not significantly different (Fig. S4). Biomass composition might be different under different spectral light conditions, but prediction performance was not seriously influenced.
Growth and cellular absorption spectra of Synechocystis sp. PCC 6803 under different spectral lights. (a) Growth of cells under six color LED lights (presented as Fig. S2) at a light intensity of 230 µmol m−2 s−1. (b) Absorption spectra of cell suspensions at 24 h. The spectra are normalized to absorption at 678 nm. Each value represents the mean ± S.D. of three independent experiments
Since the cell growth rates decreased late during culture under all the color condition, the shielding effect of the light was presumed to be important to represent the time courses of cell growth. To evaluate the shielding effect on the cells, we plotted the correlation between the specific growth rates and the optical density at 730 nm (OD730) (Fig. S5). It was considered that the shielding effects using light with different spectra were similar, and they were represented as the same negative straight line. The negative effect of photon irradiation shielding was determined as the slope of the straight line (Fig. S5) and used for simulation of OD730. The OD730 values of the culture grown under the six LED light types at 24, 48, 72, and 96 h were calculated using the simulated specific growth rates by FBA. The photon incident rate of FBA simulation was set to 78 mmol gDW−1 h−1. The values are adjusted so that the simulated cell growth rate under yellow light condition is coincident with the experimental data of the growth rate of Synechocystis culture, as shown in Fig. 4a. A value obtained by multiplying the photon incident rate (78 mmol gDW−1 h−1) and the incident photon ratio of Table 2 was set as the flux of each photon incident site [PBS, PSII, and PSI (R0039, R0041, and R0043 in Table S1, respectively)]. We plotted the correlation between the estimated and experimental OD730 (Figs. 4a, S6). The OD730 estimated by our refined model had good agreement with the experimental data under Blue, Green, Yellow, Red1, and White lights. The correlation between the estimated and experimental photosynthetic oxygen evolution rate at 24 h (Fig. S7) is presented in Fig. 4b. Experimental data of oxygen evolution were also efficiently predicted by simulation in case of Blue, Green, Yellow, Red1, and White lights. The model successfully predicted the flux distribution of photosynthesis of Synechocystis under Blue, Green, Yellow, Red1, and White light conditions. It was confirmed that the PBS to PSI/PSII ratios of Arthrospira were the same as those for simulating Synechocystis, except for Red2. The latter simulation results did not agree with the experimental data. Since the model for Arthrospira can predict cell growth under the Red2 condition, presumably the PBS to PSI/PSII ratios of Synechocystis under Red2 light are different from those of Arthrospira. In fact, even closely related species, such as Synechococcus elongates PCC 7942 and UTEX 2973 (Ungerer et al. 2018), have significantly different PSI/PSII stoichiometric ratios. During simulation of Synechocystis under Red2 light, the flux of proton transport into the thylakoid lumen increases by NDH-1; however, the actual activity of NDH-1 may not be dominant in Synechocystis. On the other hand, in the actual cells grown under the Red2 light condition, the use of ATP in central metabolism may not increase due to limitations, such as the activity of RuBisCo.
Comparison of the growth and photosynthetic activity of Synechocystis between FBA simulation and experimental results under different spectral lights. The OD730 experimentally obtained in Fig. 3a was compared with the simulation result of our refined model. Photon uptake rate was set to a specific value, in which the specific growth rate is equivalent in the simulation and experimental results under Yellow in simulation. (a) We plotted the correlation between the estimated and experimental OD730 at 24 h. (b) The photosynthetic oxygen evolution rates were experimentally obtained at 24 h (Fig. S7). Each experimental value represents the mean ± S.D. of three independent experiments. Color in each circle indicates the type of light source (see Fig. 2)
Prediction of selective use of photosynthetic electron transport system depending on PSI and PSII excitation ratio
Simulations by the refined model were done to provide visual information of photosynthetic systems. A schematic image of photon and electron fluxes in the photosynthetic apparatus of Synechocystis under the five different LED light conditions is presented in Fig. 5. As the data for cell growth simulation under Red2 light did not agree with the experimental data, the simulation results during photosynthesis under Red2 are not presented in the figure. Yellow, blue, and pink arrows indicate the incident photon, electron transfer, and proton pump fluxes, respectively. The line width of the arrows indicates their flux rate. Summation of proton pump fluxes under Red1 provides the maximum value among the investigated conditions, and consequently, highest ATP generation and cell growth are obtained.
Scheme of flux of photon and photosynthetic electron flow obtained by FBA simulation under different spectral lights. The fluxes of electron flow were simulated with the PBS to PSI/PSII ratios. a Blue, b Green, c Yellow, d Red1, and e White. The line width reflects the flux rate. Abbreviation: OEC, oxygen-evolving complex
Under Blue light, 85% of the photons were absorbed in Car (Table 2), and the photon energy was discarded as heat (Fig. 5a). Therefore, minimum ATP synthesis and cell growth was obtained. Under Green, Yellow, Red1 and White lights, both NDH-1 and NDH-2 were used as CET (Fig. 5b–e). Energy of the absorbed photon by PBS was transferred to the chlorophyll of PSII and PSI (P680 and P700, respectively) with the ratio shown in Table 2. More photons were absorbed by the Chl in PSII and PSI under the Red1 condition that under the Green, Yellow, and White light conditions. The total electron flux under the Red1 condition was larger than that under the Green, Yellow and White conditions. The total proton pump flux and the cell growth under the Red1 light condition were greater than those under the Green and Yellow light conditions. Under the White light conditions, the balance of ATP synthesis and NADPH synthesis was more suitable than Red1 light condition, which caused that cell growth was equivalent to that of the Red1 light condition.
To understand the role of the PSI/PSII excitation ratio in photosynthesis, the photosynthetic electron transport flux was simulated. The relationship between the PSI/PSII excitation ratio and fluxes in the photosynthetic pathways is depicted in Fig. 6. In the case of PSII, the incident flux was larger than PSI (PSI/PSII excitation ratio < 0.88, Phase 1 in Fig. 6). Cyd maintained the balance of PSII and PSI fluxes by uptake of the electron flux from PSII from PQ (R0037 in Table S1), because the Cyd does not transport the proton into thylakoid lumen. In this phase, Cox also maintains the electron balance of PSII and PSI fluxes as the PSI ratio increases (R0031 in Table S1). Since the Cox flux is enhanced with increasing PSI/PSII excitation ratio, ATP synthesis (R048 in Table S1) is elevated with the transfer of protons (pump) from the cytosol to the thylakoid lumen. In addition, as the photosynthetic NADPH synthesis flux by ferredoxin: NADP+ reductase (FNR) (photosynthetic NADPH synthesis) is proportional to PSI incident flux, the specific growth rate increases in proportion to the flux of NADPH synthesis (R543 in Table S1).
In Phase 2 in Fig. 6 (0.88 < PSI/PSII excitation ratio < 1.35), the specific growth rate attained the maximum value by taking the optimal ATP/NADPH synthesis ratio via photosynthesis. The optimal ATP/NADPH synthesis ratio of photosynthesis was calculated as 2.51. Proportional to the increase in the PSI/PSII excitation ratio and electron transfer from Pc to PSI (R0023 in Table S1), the flux from Pc to Cox (R0031 in Table S1) decreased to satisfy the balance of electron at Pc. Stoichiometric constraint in the linear electron transfer in PSI requires equivalent electron flux from Pc to the flux through PSI (R0024–R0028 in Table S1). Maximum ATP synthesis is maintained by increasing the NDH-1 flux until Cox flux becomes zero. The photosynthetic NADPH synthesis flux (R543 in Table S1) was remained constant in this phase due to the increased PSI flux, while the flux of NDH-1 increased (R0033 in Table S1).
When the incident flux of PSI became higher than that of PSII (1.35 < PSI/PSII excitation ratio, Phase 3 in Fig. 6), the Cox flux became zero, and NDH-1 (R0033 in Table S1) and NDH-2 (R0035 in Table S1) transferred excess electrons from PSI to the PQ pool. In this phase, the specific growth rate decreased because photosynthetic NADPH or ATP synthesis decreased. NDH-1 and NDH-2 do not have a unique flux solution; thus, the possible range of these fluxes is presented as the shaded area in the figure. In the case of increased photosynthetic ATP synthesis flux using NDH-1, the photosynthetic NADPH synthesis decreased correspondingly (Case 1). However, previous results of 13C metabolic flux analysis (13C-MFA) indicated small fluxes of the TCA cycle and that Case 1 would not occur (Nakajima et al. 2017). In contrast, NDH-2 does not transport the proton from the cytosol into the thylakoid lumen and ATP is not synthesized in photosynthesis. In this case, the photosynthetic NADPH synthesis increased dramatically. However, all of the increased flux was used for NDH-2 (R0035 in Table S1) through transhydrogenase (R052 in Table S1), and electron transfer was not effectively used for cell growth (Case 2). The Green, Yellow, Red1, and White light conditions are in Phase 3. The schematic image in Fig. 5 was drawn using the solutions with the largest values of NDH-2 because a large TCA cycle flux was not observed experimentally under these spectral conditions.
These simulation results indicate that the specific growth rate was maintained at the maximum value when the PSI/PSII excitation ratio ranged from 0.88 to 1.35 by adaptive selective use of electron transfer pathways. These results are consistent with the experimental results in previous studies, in which PSI/PSII stoichiometric ratio was maintained at an approximate value of one under light spectrum preferentially absorbed by Chl. This ratio was higher under the light absorbed by PBS (Kawamura et al. 1979; Myers et al. 1980), and the excessive PSI excitation prevented cell growth (Hihara et al. 1998; Fujimori et al. 2005). The simulated optimal value of the ATP/NADPH synthesis ratio of photosynthesis was 2.51 in this phase. This value is similar to the experimentally estimated ratio of the ATP/NADPH demand (ATP/NADPH = 2.94) based on the 13C-MFA (Nakajima et al. 2017).
Additionally, simulation indicated that respiration by Cyd and Cox, and CET by NDH-1 and NDH-2 can act to maintain the flux balance of PSII and PSI, depending on light conditions. FBA simulations of Cyd, Cox, NDH-1, and NDH-2 deletion revealed that Cyd and Cox can functionally complement each other, as well as NDH-1 and NDH-2 (Fig. S8). This is consistent with previous findings indicating little if any effect on growth or viability as a result of the Cox or Cyd single deletion (Howitt and Vermaas 1998; Pils and Schmetterer 2001; Nomura et al. 2006). The stoichiometric simulation results also suggested that the functions of NDH-1 and NDH-2 depend on the PSI/PSII excitation ratio. In support of this simulation result, previous experimental results indicated that NDH-1 acts functionally as the major CET route into the PQ pool and that NDH-2 hardly contributes to CET under White light condition (Howitt et al. 1999; Bernát et al. 2011; Peltier et al. 2016; Huokko et al. 2017). This switching from use of NDH-1 to NDH-2 might be controlled by a molecular regulatory mechanism.
FBA simulation using our refined model predicted the PSI/PSII excitation and the flux state of photosynthetic electron transport under various light conditions and indicated the optimal PSI/PSII excitation ratio for cell growth. The effect of ratio of PSI/PSII electron transfer on the ATP/NADPH synthesis under different light intensity conditions has been analyzed using FBA (Qian et al. 2017). The authors reported that the ATP/NADPH ratio and PSI/PSII electron transport was almost constant in the wide range of light intensity condition with red light. In our experimental and simulation results, the PSI/PSII excitation ratio depended on the spectrum of light.
Using this simulation, we obtained data that were valuable for optimizing the growth of cyanobacteria by developing light environment and genetic manipulation, leading to biomass production. For example, cell growth under the Red1 light condition was best of the investigated light condition. However, the Phase 3 state under Red1 light was not optimal for cell growth (Fig. 6). This shows the state of photosynthetic apparatus in which photoprotection is occurred. The value of energy transfer from PBS to PSI was high (5.68) under Red1 light (Table 2). It has been reported that higher plants, Chlamydomonas, and cyanobacteria acclimate and adapt to the changing light conditions by controlling the energy transfer between both PSs (spill-over), the energy distribution from PBS to PSI/PSII (state transition), and the pigments composition (Akimoto et al. 2012, 2013; Dall’Osto et al. 2015; Yokono et al. 2015; Ho et al. 2017; Duanmu et al. 2017). One possible genetic improvement strategy to increase the ATP generation and cell growth is to reduce the state change of the pigment composition by modification of signal transduction system of the cells by carefully considering the robustness against photoinhibition. Optimization of both increased cell growth and robustness against photoinhibition should be considered to create useful strains as biomass and valuable producing strains. Another operational strategy is to control the color or intensity of lights to irradiate the cells in the photobioreactor system. It is possible to optimize the spectrum and intensity using simulation data. These rational design methods of photosynthetic organisms and photobioreactors will be realized in the future.
Conclusions
We established the Synechocystis sp. PCC 6803 genome-scale model that refined the photosynthetic reaction and allowed an accurate prediction of the specific growth rate of the cells grown under various spectral lights using PBS to PSI/PSII ratios of Arthrospira and Synechocystis. FBA simulation with our refined model revealed the selective flow of photosynthetic electron transport, the NDH-1 and NDH-2 functions, depending on PSI/PSII excitation ratio, and the most efficient condition of photosynthesis under different spectral lights. The obtained results provide new insight into cyanobacterial photosynthesis and useful information to create appropriate photosynthetic systems under various light environmental conditions.
Abbreviations
- ARTO:
-
Alternative respiratory oxidase
- Car:
-
Carotenoid
- CET:
-
Cyclic electron transport
- Chl:
-
Chlorophyll
- Cox:
-
aa3-type cytochrome c oxidase complex
- Cyd:
-
Cytochrome bd-quinol oxidase complex
- cyt:
-
b6f Cytochrome b6f complex
- FBA:
-
Flux balance analysis
- Fd:
-
Ferredoxin
- FNR ferredoxin:
-
NADP+ reductase
- LED:
-
Light-emitting diodes
- LET:
-
Linear electron transport
- NDH-1:
-
Type I NADH dehydrogenase
- NDH-2:
-
Type II NADH dehydrogenase
- PBS:
-
Phycobilisome
- Pc:
-
Plastocyanin
- PQ:
-
Plastoquinone
- PSI:
-
Photosystem I
- PSII:
-
Photosystem II
- RTO:
-
Respiratory terminal oxidase
- SDH:
-
Succinate dehydrogenase
References
Akimoto S, Yokono M, Hamada F et al (2012) Adaptation of light-harvesting systems of Arthrospira platensis to light conditions, probed by time-resolved fluorescence spectroscopy. Biochim Biophys Acta 1817:1483–1489
Akimoto S, Yokono M, Aikawa S, Kondo A (2013) Modification of energy-transfer processes in the cyanobacterium, Arthrospira platensis, to adapt to light conditions, probed by time-resolved fluorescence spectroscopy. Photosynth Res 117:235–243
Battchikova N, Wei L, Du L et al (2011) Identification of novel Ssl0352 protein (NdhS), essential for efficient operation of cyclic electron transport around photosystem I, in NADPH:plastoquinone oxidoreductase (NDH-1) complexes of Synechocystis sp. PCC 6803. J Biol Chem 286:36992–37001
Becker SA, Feist AM, Mo ML et al (2007) Quantitative prediction of cellular metabolism with constraint-based models: the COBRA Toolbox. Nat Protoc 2:727–738
Bernát G, Appel J, Ogawa T, Rögner M (2011) Distinct roles of multiple NDH-1 complexes in the cyanobacterial electron transport network as revealed by kinetic analysis of P700+ reduction in various ndh-deficient mutants of Synechocystis sp. strain PCC6803. J Bacteriol 193:292–295
Björn LO, Papageorgiou GC, Blankenship RE, Govindjee (2009) A viewpoint: why chlorophyll a? Photosynth Res 99:85–98
Boyle NR, Morgan JA (2009) Flux balance analysis of primary metabolism in Chlamydomonas reinhardtii. BMC Syst Biol 3:4
Broddrick JT, Rubin BE, Welkie DG et al (2016) Unique attributes of cyanobacterial metabolism revealed by improved genome-scale metabolic modeling and essential gene analysis. Proc Natl Acad Sci USA 113:E8344–E8353
Burnap RL, Sherman LA (1991) Deletion mutagenesis in Synechocystis sp. PCC6803 indicates that the Mn-stabilizing protein of photosystem II is not essential for O2 evolution. Biochemistry 30:440–446
Chang RL, Ghamsari L, Manichaikul A et al (2011) Metabolic network reconstruction of Chlamydomonas offers insight into light-driven algal metabolism. Mol Syst Biol 7:518
Chen M, Blankenship RE (2011) Expanding the solar spectrum used by photosynthesis. Trends Plant Sci 16:427–431
Chitnis VP, Chitnis PR (1993) PsaL subunit is required for the formation of photosystem I trimers in the cyanobacterium Synechocystis sp. PCC 6803. FEBS Lett 336:330–334
Collins AM, Liberton M, Jones HDT et al (2012) Photosynthetic pigment localization and thylakoid membrane morphology are altered in Synechocystis 6803 phycobilisome mutants. Plant Physiol 158:1600–1609
Dall’Osto L, Bressan M, Bassi R (2015) Biogenesis of light harvesting proteins. Biochim Biophys Acta 1847:861–871
Duanmu D, Rockwell NC, Lagarias JC (2017) Algal light sensing and photoacclimation in aquatic environments. Plant Cell Environ 40:2558–2570
Feist AM, Herrgård MJ, Thiele I et al (2009) Reconstruction of biochemical networks in microorganisms. Nat Rev Microbiol 7:129–143
Fujimori T, Higuchi M, Sato H et al (2005) The mutant of sll1961, which encodes a putative transcriptional regulator, has a defect in regulation of photosystem stoichiometry in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol 139:408–416
Fujisawa T, Narikawa R, Maeda SI et al (2017) Cyanobase: a large-scale update on its 20th anniversary. Nucleic Acids Res 45:D551–D554
Gantt E (1981) Phycobilisomes. Annu Rev Plant Physiol 32:327–347
Ghosh T, Bhayani K, Paliwal C et al (2016) Cyanobacterial pigments as natural anti-hyperglycemic agents: an in vitro study. Front Mar Sci 3:146
Glazer AN (1984) Phycobilisome a macromolecular complex optimized for light energy transfer. Biochim Biophys Acta 768:29–51
Grossman AR, Bhaya D, Apt KE, Kehoe DM (1995) Light-harvesting complexes in oxygenic photosynthesis: diversity, control, and evolution. Annu Rev Genet 29:231–288
Heirendt L, Arreckx S, Pfau T et al (2019) Creation and analysis of biochemical constraint-based models: the COBRA Toolbox v3.0. Nat Protoc 14:639–702
Hihara Y, Sonoike K, Ikeuchi M (1998) A novel gene, pmgA, specifically regulates photosystem stoichiometry in the cyanobacterium Synechocystis sp. PCC 6803 in response to high light. Plant Physiol 117:1205–1216
Ho M-Y, Soulier NT, Canniffe DP et al (2017) Light regulation of pigment and photosystem biosynthesis in cyanobacteria. Curr Opin Plant Biol 37:24–33
Hohmann-Marriott MF, Blankenship RE (2011) Evolution of photosynthesis. Annu Rev Plant Biol 62:515–548
Howitt CA, Vermaas WFJ (1998) Quinol and cytochrome oxidases in the cyanobacterium Synechocystis sp. PCC 6803. Biochemistry 37:17944–17951
Howitt CA, Udall PK, Vermaas WFJ (1999) Type 2 NADH dehydrogenases in the cyanobacterium Synechocystis sp. Strain PCC 6803 are involved in regulation rather than respiration. J Bacteriol 181:3994–4003
Huokko T, Muth-Pawlak D, Battchikova N et al (2017) Role of type 2 NAD(P)H dehydrogenase NdbC in redox regulation of carbon allocation in Synechocystis. Plant Physiol 174:1863–1880
Ikeuchi M, Tabata S (2001) Synechocystis sp. PCC 6803—a useful tool in the study of the genetics of cyanobacteria. Photosynth Res 70:73–83
Imam S, Schäuble S, Valenzuela J et al (2015) A refined genome-scale reconstruction of Chlamydomonas metabolism provides a platform for systems-level analyses. Plant J 84:1239–1256
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kawamura M, Mimuro M, Fujita Y (1979) Quantitative relationship between two reaction centers in the photosynthetic system of blue-green algae. Plant Cell Physiol 20:697–705
Kirilovsky D (2015) Modulating energy arriving at photochemical reaction centers: orange carotenoid protein-related photoprotection and state transitions. Photosynth Res 126:3–17
Kirilovsky D, Kerfeld CA (2016) Cyanobacterial photoprotection by the orange carotenoid protein. Nat Plants 2:16180
Lea-Smith DJ, Ross N, Zori M et al (2013) Thylakoid terminal oxidases are essential for the cyanobacterium Synechocystis sp. PCC 6803 to survive rapidly changing light intensities. Plant Physiol 162:484–495
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzymol 148:350–382
Mahadevan R, Schilling CH (2003) The effects of alternate optimal solutions in constrain-based genome-scale metabolic models. Metab Eng 5:264–276
Mimuro M, Kikuchi H (2003) Antenna systems and energy transfer in cyanophyta and rhodophyta. In: Green BR, Parson WW (eds) Light harvesting antennas in photosynthesis. Kluwer Academic Publishers, Dordrecht, pp 281–306
Mitchell P (1975) The protonmotive Q cycle: a general formulation. FEBS Lett 59:137–139
Mullineaux CW (2008) Phycobilisome-reaction centre interaction in cyanobacteria. Photosynth Res 95:175–182
Mullineaux CW (2014) Electron transport and light-harvesting switches in cyanobacteria. Front Plant Sci 5:7
Myers J, Graham J-R, Wang RT (1980) Light-harvesting in Anacystis nidulans studies in pigmant mutants. Plant Physiol 66:1144–1149
Nakajima T, Yoshikawa K, Toya Y et al (2017) Metabolic flux analysis of the Synechocystis sp. PCC 6803 ΔnrtABCD mutant reveals a mechanism for metabolic adaptation to nitrogen-limited conditions. Plant Cell Physiol 58:537–545
Nakamura Y, Kaneko T, Hirosawa M et al (1998) CyanoBase, a www database containing the complete nucleotide sequence of the genome of Synechocystis sp. strain PCC6803. Nucleic Acids Res 26:63–67
Nogales J, Gudmundsson S, Knight EM et al (2012) Detailing the optimality of photosynthesis in cyanobacteria through systems biology analysis. Proc Natl Acad Sci USA 109:2678–2683
Nomura CT, Persson S, Shen G et al (2006) Characterization of two cytochrome oxidase operons in the marine cyanobacterium Synechococcus sp. PCC 7002: inactivation of ctaDI affects the PS I:PS II ratio. Photosynth Res 87:215–228
Orth J, Thiele I, Palsson B (2010) What is flux balance analysis? Nat Biotechnol 28:245–248
Peltier G, Aro E-M, Shikanai T (2016) NDH-1 and NDH-2 plastoquinone reductases in oxygenic photosynthesis. Annu Rev Plant Biol 67:55–80
Pils D, Schmetterer G (2001) Characterization of three bioenergetically active respiratory terminal oxidases in the cyanobacterium Synechocystis sp. strain PCC 6803. FEMS Microbiol Lett 203:217–222
Pils D, Gregor W, Schmetterer G (1997) Evidence for in vivo activity of three distinct respiratory terminal oxidases in the cyanobacterium Synechocystis sp. strain PCC6803. FEMS Microbiol Lett 152:83–88
Porra RJ, Thompson WA, Kriedemann PE (1989) Determination of accurate extinction coefficients and simultaneous equations for assaying chlorophylls a and b extracted with four different solvents: verification of the concentration of chlorophyll standards by atomic absorption spectroscopy. Biochim Biophys Acta 975:384–394
Price ND, Reed JL, Palsson BØ (2004) Genome-scale models of microbial cells: evaluating the consequences of constraints. Nat Rev Microbiol 2:886–897
Qian X, Kim MK, Kumaraswamy GK et al (2017) Flux balance analysis of photoautotrophic metabolism: uncovering new biological details of subsystems involved in cyanobacterial photosynthesis. Biochim Biophys Acta Bioeng 1858:276–287
Rich PR (1988) A critical examination of the supposed variable proton stoichiometry of the chloroplast cytochrome bf complex. Biochim Biophys Acta 932:33–42
Rippka R, Deruelles J, Waterbury JB et al (1979) Generic assignments, strain histories and properties of pure cultures of cyanobacteria. Microbiology 111:1–61
Rockwell NC, Duanmu D, Martin SS et al (2014) Eukaryotic algal phytochromes span the visible spectrum. Proc Natl Acad Sci USA 111:3871–3876
Sacksteder CA, Kanazawa A, Jacoby ME, Kramer DM (2000) The proton to electron stoichiometry of steady-state photosynthesis in living plants: a proton-pumping Q cycle is continuously engaged. Proc Natl Acad Sci USA 97:14283–14288
Schilling C, Schuster S, Palsson BØ et al (1999) Metabolic pathway analysis: basic concepts and scientific applications in the post-genomic era. Biotechnol Prog 15:296–303
Schmetterer G, Alge D, Gregor W (1994) Deletion of cytochrome c oxidase genes from the cyanobacterium Synechocystis sp. PCC6803: evidence for alternative respiratory pathways. Photosynth Res 42:43–50
Schuller JM, Birrell JA, Tanaka H et al (2019) Structure adaptations of photosynthetic complex I enable ferredoxin-dependent electron transfer. Science 363:257–260
Shen G, Boussiba S, Vermaas WFJ (1993) Synechocystis sp. PCC 6803 strains lacking photosystem I and phycobilisome function. Plant Cell 5:1853–1863
Sonani RR, Gardiner A, Rastogi RP et al (2018) Site, trigger, quenching mechanism and recovery of non-photochemical quenching in cyanobacteria: recent updates. Photosynth Res 137:171–180
Toyoshima M, Mori N, Moriyama T et al (2016) Analysis of triacylglycerol accumulation under nitrogen deprivation in the red alga Cyanidioschyzon merolae. Microbiology 162:803–812
Ungerer J, Lin PC, Chen HY, Pakrasi HB (2018) Adjustments to photosystem stoichiometry and electron transfer proteins are key to the remarkably fast growth of the cyanobacterium Synechococcus elongatus UTEX 2973. mBio 9:e002327–17
Vu TT, Stolyar SM, Pinchuk GE et al (2012) Genome-scale modeling of light-driven reductant partitioning and carbon fluxes in diazotrophic unicellular cyanobacterium Cyanothece sp. ATCC 51142. PLoS Comput Biol 8:e1002460
Williams JGK (1988) Construction of specific mutations in photosystem II photosynthetic reaction center by genetic engineering methods in Synechocystis 6803. Methods Enzymol 167:766–778
Yokono M, Takabayashi A, Akimoto S, Tanaka A (2015) A megacomplex composed of both photosystem reaction centres in higher plants. Nat Commun 6:6675
Yoshikawa K, Kojima Y, Nakajima T et al (2011) Reconstruction and verification of a genome-scale metabolic model for Synechocystis sp. PCC6803. Appl Microbiol Biotechnol 92:347–358
Yoshikawa K, Aikawa S, Kojima Y et al (2015) Construction of a genome-scale metabolic of Arthrospira platensis NIES-39 and metabolic design for cyanobacterial bioproduction. PLoS ONE 10:e0144430
Yoshikawa K, Toya Y, Shimizu H (2017) Metabolic engineering of Synechocystis sp. PCC 6803 for enhanced ethanol production based on flux balance analysis. Bioprocess Biosyst Eng 40:791–796
Yuan H, Cheung CYM, Hilbers PAJ, van Riel NAW (2016) Flux balance analysis of plant metabolism: the effect of biomass composition and model structure on model predictions. Front Plant Sci 7:537
Zakhartsev M, Medvedeva I, Orlov Y et al (2016) Metabolic model of central carbon and energy metabolisms of growing Arabidopsis thaliana in relation to sucrose translocation. BMC Plant Biol 16:262
Acknowledgements
This work was supported in part by Grants-in-Aid for Scientific Research (Grant Nos. 16H06552 and 16H06559).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Toyoshima, M., Toya, Y. & Shimizu, H. Flux balance analysis of cyanobacteria reveals selective use of photosynthetic electron transport components under different spectral light conditions. Photosynth Res 143, 31–43 (2020). https://doi.org/10.1007/s11120-019-00678-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-019-00678-x