Abstract
Flavodoxins are small proteins with a non-covalently bound FMN that can accept two electrons and accordingly adopt three redox states: oxidized (quinone), one-electron reduced (semiquinone), and two-electron reduced (quinol). In iron-deficient cyanobacteria and algae, flavodoxin can substitute for ferredoxin as the electron carrier in the photosynthetic electron transport chain. Here, we demonstrate a similar function for flavodoxin from the green sulfur bacterium Chlorobium phaeovibrioides (cp-Fld). The expression of the cp-Fld gene, found in a close proximity with the genes for other proteins associated with iron transport and storage, increased in a low-iron medium. cp-Fld produced in Escherichia coli exhibited the optical, ERP, and electron-nuclear double resonance spectra that were similar to those of known flavodoxins. However, unlike all other flavodoxins, cp-Fld exhibited unprecedented stability of FMN semiquinone to oxidation by air and difference in midpoint redox potentials for the quinone–semiquinone and semiquinone–quinol couples (− 110 and − 530 mV, respectively). cp-Fld could be reduced by pyruvate:ferredoxin oxidoreductase found in the membrane-free extract of Chl. phaeovibrioides cells and photo-reduced by the photosynthetic reaction center found in membrane vesicles from these cells. The green sulfur bacterium Chl. phaeovibrioides appears thus to be a new type of the photosynthetic organisms that can use flavodoxin as an alternative electron carrier to cope with iron deficiency.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Living organisms require iron (Fe)-containing prosthetic groups, such as hemes, iron–sulfur (FeS) clusters, and non-heme Fe centers, to catalyze redox and other reactions. Fe is the most abundant metal with a variable valency in the Earth’s crust and biosphere. However, its soluble Fe(II) forms are not easily available to living organisms in most aerobic ecological niches because of being oxidized by molecular oxygen to low-soluble Fe(III) forms. One way to overcome Fe limitation is to increase its acquisition from the Fe(III) compounds, for instance, by using various siderophores (De Serrano et al. 2016). Alternatively, Fe-containing enzymes are replaced by their functional non-Fe protein analogs. Flavin-containing flavodoxin is a typical example of such substitution of Fe-containing ferredoxin in many bacteria and algae that cope with Fe deficiency (LaRoche et al. 1996; Sancho 2006).
Flavodoxins are small (140–180 amino acid residues in length), strongly acidic proteins that contain a non-covalently bound FMN molecule as the prosthetic group (Simondsen and Tollin 1980). In structural terms, flavodoxins are three-layered α/β proteins, with a central five-stranded β-sheet surrounded by two helical layers (Burnett et al. 1974; Watt et al. 1991) and are classified into long-chain and short-chain types, depending on the presence/absence of an ~ 20-residue loop in strand 5 of the β-sheet (Peelen et al. 1996). Flavodoxins have been found in many but not all prokaryotes of different taxonomic groups, including most cyanobacteria. In the eukaryotic world, flavodoxins occur in some algae, but not in plants, presumably, because plants evolved from the flavodoxin-lacking algae (Pierella Karlusich et al. 2014).
The key property of flavodoxins is their ability to substantially stabilize a one-electron-reduced (semiquinone) form of FMN because of a large difference in the midpoint redox potentials (Em) for the quinone/semiquinone and semiquinone/quinol transitions (− 90 to − 245 and − 370 to − 480 mV, respectively) (Dubourdieu et al. 1975; Watt 1979; Sykes and Rogers 1984; Biel et al. 1996; Lawson et al. 2004; Segal et al. 2017). This property allows flavodoxins to function as one-electron carriers of reducing equivalents between various partner proteins, for instance, the type I photosynthetic reaction center complex (RC) and ferredoxin-NAD(P)+ reductase (FNR) in the photosynthetic electron transport chain of cyanobacteria and algae (Sétif 2001; Pierella Karlusich et al. 2014). Other important physiological reactions in which flavodoxins can substitute for ferredoxins include N2 fixation (Segal et al. 2017), dissimilatory sulfate reduction (Simondsen and Tollin 1980), and anaerobic fermentation (Chowdhury et al. 2016). The in vivo function of flavodoxin is associated with the semiquinone/quinol transition; however, a physiological significance of the quinone/semiquinone transition cannot be excluded.
Green sulfur bacteria (GSB) are similar to cyanobacteria in that both are photo-autotrophs. Furthermore, GSB have to cope with Fe deficiency because their habitat is enriched with reduced sulfur compounds, including H2S (Sakurai et al. 2010), which forms virtually insoluble Fe sulfides. A search through available prokaryotic genomes indicates the presence of flavodoxin homologs in many GSB, such as Chlorobium phaeovibrioides, Chlorobium limicola, Chlorobaculum tepidum, Chloroherpeton thalassium, and different Prosthecochloris species. The products of the putative flavodoxin genes found in GSB belong to a flav_long family (TIGR01752) that comprises long-chain flavodoxins. Noteworthy, the genes for flavodoxin and three other proteins presumably associated with Fe storage, transport, and regulation (ferritin, the ferrous ion transporter FeoABC, and the Fe(Mn)-dependent repressor DtxR) are typically located close to each other in the chromosomes of different GSB (Fig. 1), suggesting their common role in maintaining viability under conditions of Fe limitation. Nevertheless, there is no literature report describing flavodoxin functioning in GSB to date (Fromme 1999). The results reported below provide a first experimental demonstration of a possible role of flavodoxin as an electron carrier in the photosynthetic electron transport chain of a green sulfur bacterium, a new type of the photosynthetic organism, which uses flavodoxin in this capacity.
Typical arrangements of flavodoxin-associated genes in genomes of green sulfur bacteria. FeoA–C subunits of the ferrous ion transporter FeoABC, Fld flavodoxin, Fd ferredoxin, DtxR iron-dependent repressor, HP1, HP2, and HP3 proteins with unknown functions. aChl. phaeovibrioides (GenBank IDs: ABP36541 to ABP36547 from left to right, respectively); bChl. limicola (ACD89833–ACD89838); cChlorobaculum tepidum (AAM72966–AAM72958); dChloroherpeton thalassium (ACF13400–ACF13403); eProsthecochloris sp. CIB 2401 (ANT64387–ANT64376)
Materials and methods
Bacterial strain and growth conditions
Chlorobium phaeovibrioides DSM 265 cells were obtained from the Leibniz Institute Collection of Microorganisms and Cell Cultures (DSMZ). The cells were grown anaerobically at 28 °C and a light illuminance of 1000 lx, as described by Malik (1983). The growth medium contained 10 g/L NaCl, 0.5 g/L each of MgSO4·7H2O, NH4Cl, KH2PO4, and ammonium acetate, 0.05 g/L CaCl2·2H2O, 2 g/L NaHCO3, 2 g/L Na2S2O3, 1 g/L Na2S, 0.15 or 3.00 μM ferric citrate, trace element solution SL-6 (1:500), 20 μg/L cyanocobalamin, 5 mg/L resazurin, and 20 mM MOPS (final pH of the medium 6.9).
Fractionation of Chlorobium phaeovibrioides cells
The cells were harvested by centrifugation (10,000×g, 10 min) and washed with anaerobic buffer A (170 mM KCl, 5 mM MgSO4, 10 mM β-mercaptoethanol, and 30 mM MOPS–Tris, pH 7.0). The cell pellet was suspended in buffer A, and the suspension was passed twice through a French press (16,000 psi). Undamaged cells and cell debris were removed by centrifugation at 22,500×g (10 min), and the supernatant was further centrifuged at 180,000×g for 75 min. The pellet containing membrane vesicles was suspended in buffer A. The supernatant is referred to below as the cytoplasmic fraction of Chl. phaeovibrioides.
Construction of the plasmid encoding flavodoxin from Chlorobium phaeovibrioides
The gene encoding Chl. phaeovibrioides DSM 265 flavodoxin (GenBank ID: ABP36546.1) was amplified from genomic DNA of this bacterium by PCR using high-fidelity Tersus polymerase (Evrogen, Russia) and the forward/reverse primers 5´-CATATGAAAAGAACAGGCATT/5´-CTCGAGGGAAAGCAGCGGGCT (restriction sites for NdeI and XhoI are underlined). The amplified 513-bp fragment was cloned into the pSCodon vector (Delphi Genetics) using the NdeI/XhoI sites, resulting in the plasmid pSC_fld. The cp-Fld-encoding region of pSC_fld was verified using DNA sequencing, and the plasmid was transformed into the Escherichia coli BL21 (DE3) strain.
Isolation of recombinant 6 × His-tagged cp-Fld
Escherichia coli/pSC_fld cells were grown at 32 °C to mid-exponential phase (A600 = 0.3–0.4), and cp-Fld synthesis was induced with 0.2% (w/v) lactose. The cells additionally grown for 3 h were harvested by centrifugation (10,000×g, 10 min) and washed twice with the medium containing 300 mM KCl, 5 mM MgSO4, and 10 mM Tris–HCl, pH 8.0. The cell pellet was suspended in the medium containing 300 mM KCl, 5 mM MgSO4, 1 mM phenylmethylsulfonyl fluoride, 20 mM Tris–HCl, and 5 mM imidazole–HCl, pH 8.0, and the suspension was passed twice through a French press (16,000 psi). Cell debris and membranes were removed by centrifugation at 180,000×g for 75 min. The supernatant was loaded onto a Ni–NTA column equilibrated with buffer B (10 mM Tris–HCl, 5 mM imidazole–HCl, and 300 mM KCl, pH 8.0). The column was washed first with buffer B and then with buffer B containing 10 mM imidazole–HCl. The 6 × His-tagged cp-Fld was eluted with buffer B containing 100 mM imidazole–HCl. The protein fraction was concentrated and kept frozen at − 80 °C until use. Typical yield of cp-Fld was ~ 1.2 μmol per 1 L of cell culture.
X-band EPR spectroscopy
EPR measurements were accomplished using a Bruker ESP-300 spectrometer. The temperature of the sample (80 K) was controlled by an ESR 900 cryostat with an ITC4 temperature controller (Oxford Instruments). Magnetic field modulation frequency in continuous-wave EPR measurements was 100 kHz.
For pulse proton electron-nuclear double resonance (ENDOR) experiments, a dielectric ENDOR resonator (Bruker EN 4118X-MD4) was used. The Davies ENDOR microwave pulse sequence π–T–π/2–τ–π–τ– echo was used, with a radio frequency π pulse applied during the T-interval. For the low-frequency range (7.5–22.5 MHz), the microwave π pulse duration of 200 ns, radio frequency π pulse of 16 μs, τ interval of 400 ns, and shot repetition time of 3 ms were used. These selected experimental parameters ensured good resolution of the ENDOR spectrum of protons with weak hyperfine coupling. For the high-frequency range (22.5–37.5 MHz), the microwave π pulse duration of 40 ns, radio frequency π pulse of 8 μs, τ interval of 160 ns, and shot repetition time of 1 ms were used. This parameter set allowed to increase the signal-to-noise ratio for the ENDOR signal of strongly coupled H(5) protons.
Redox titration
The redox potentials of cp-Fld were determined at 25 °C, by the method described by Efimov et al. (2014). In titrations of the cp-Fldox/cp-Fldsq transition, the anaerobic cuvette contained 50 μM cp-Fldox, 10–30 μM redox dye (resorufin, Nile blue, or phenosafranine), 100 mM potassium phosphate/20 mM EDTA buffer (pH 7.0), and 1 μM riboflavin as a source of electrons. The stepwise cp-Fld reduction was achieved by short illuminations of the mixture through a 380–480-nm bandpass filter with a 100-W halogen lamp equipped with an infrared cut-off filter. In the titration of the cp-Fldsq/cp-Fldred transition, the anaerobic cuvette contained 70 μM cp-Fldsq, 30 μM methyl viologen, and 100 mM potassium phosphate/20 mM EDTA buffer (pH 7.0). The mixture was first reduced by 1 mM dithionite and then oxidized by (bi)sulfite additions, as described previously (Mayhew 1978). Complete reduction of cp-Fld and methyl viologen was achieved by shifting the pH to 9.0, by KOH addition at the end of the titration. The reduction reactions were monitored spectrophotometrically. The spectrum of the equilibrated mixture at each titration point was deconvoluted using the program GIM, and the concentrations of the oxidized and reduced forms of cp-Fld and the dye were calculated.
Quantitative reverse transcription polymerase chain reaction (RT-qPCR)
RNA was extracted from Chl. phaeovibrioides cells using the ExtractRNA and CleanRNA Standard kits (Evrogen) and digested with the RNase-free DNase I (Thermo Scientific) at 37 °C for 1 h. cDNA was synthesized using the MMLV RT kit (Evrogen) using random decanucleotide primers. A control reaction without reverse transcriptase was included for each sample. RT-qPCR assays were performed with the qPCRmix-HS SYBR kit (Evrogen), using the cDNA preparations as templates and 5′-GACTGGGCGGTGTTCCTT/5′-GGCCGACGAACTTTCCAT primer pair. 16S rRNA was used for normalization (primer pair 5′-CAGCCACATTGGAACTGAGA/5′-GCTTATTCGCAGAGTACCGT). Serial dilutions of Chl. phaeovibrioides genomic DNA, containing genes for cp-Fld and 16S rRNA in a 1:1 ratio, were used for calibration.
Determination of enzymatic activities
cp-Fld reduction by pyruvate:ferredoxin oxidoreductase (PFOR) or photoreduction by membrane vesicles from Chl. phaeovibrioides was determined at 25 °C in a 3.2-mL anaerobic cuvette. For the PFOR-catalyzed reaction, the assay mixture contained 37 or 75 μM cp-Fldox or 1 mM methyl viologen, 0.1 mM thiamine pyrophosphate, 5 mM pyruvate, 0.2 mM CoA, 15 U glucose oxidase, 15 U catalase, 10 mM glucose, 2 mM MgSO4, and 100 mM MOPS–Tris (pH 7.0). The reaction was initiated by the addition of the cytoplasmic fraction of Chl. phaeovibrioides (50 or 125 μg protein). In cp-Fld photoreduction, the assay mixture contained 50 μM cp-Fld (cp-Fldox or cp-Fldsq), membrane vesicles from Chl. phaeovibrioides (3–5 μg/mL bacteriochlorophyll d), 15 U glucose oxidase, 15 U catalase, 10 mM glucose, 10 mM β-mercaptoethanol, and 100 mM MOPS–Tris (pH 7.0). The reaction was initiated by light (730 ± 20 nm) from a 3-W light-emitting diode and monitored using a Hitachi 557 spectrophotometer. One unit of the enzyme activity was defined as the amount of the enzyme required to catalyze a one-electron reduction of 1 μmol of cp-Fld (ε577 = 4.4 mM−1 cm−1) or methyl viologen (ε606 = 13.7 mM−1 cm−1 (Watanabe and Honda 1982)) per 1 min.
Flavin extraction and separation
Non-covalently bound flavins were extracted from cp-Fld, as described previously (Bertsova et al. 2014), and separated using thin-layer chromatography on silica gel plates (Sorbfil, 15 × 15 cm); a 5% (w/v) Na2HPO4·12 H2O solution was used as the mobile phase (Fazekas and Kokai 1971). The Rf values for riboflavin, FMN, and FAD were 0.37, 0.49, and 0.6, respectively.
Protein and chlorophyll concentrations
Protein concentrations were determined by the bicinchoninic acid method (Smith et al. 1985) using bovine serum albumin as the standard. Concentrations of bacteriochlorophyll d (the main chlorophyll of Chl. phaeovibrioides chlorosomes) were determined spectrophotometrically, as described by Savvichev et al. (2018).
Results
Isolation and initial characterization of Chlorobium phaeovibrioides flavodoxin produced in Escherichia coli
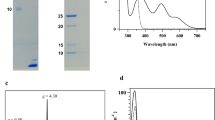
The gene for putative flavodoxin (ABP36546) was amplified from Chl. phaeovibrioides genomic DNA and cloned into the expression vector pSCodon, containing a T7 promoter and an insert for a 6 × His-tag at the C-terminus of the encoded protein, which was heterologously produced in E. coli BL21 (DE3) cells. Purification of flavodoxin (cp-Fld) from these cells using metal-chelating chromatography yielded a protein that was detected as a single ~ 20-kDa band on SDS-polyacrylamide gel electrophoresis (data not shown; the calculated mass of the 6 × His-tagged cp-Fld is 18.6 kDa).
Flavin cofactors were extracted from cp-Fld by treatment with trichloroacetic acid and separated by thin-layer chromatography. Only FMN was detected in the extract, in a ratio of 0.8 mol/mol protein (data not shown), like in other flavodoxins characterized to date (Sancho 2006). However, cp-Fld differed from known flavodoxins, all of which are yellow in the “as isolated” state (flavos is yellow in Latin), by having an unusual purple appearance (Fig. 2a).
Spectral characteristics of Chl. phaeovibrioides flavodoxin produced in Escherichia coli cells. acp-Fld preparation obtained from 1 L of cell culture. b Electronic absorption spectra of cp-Fld in “as isolated,” completely oxidized (ox), and semiquinone (sq) forms. The two last forms were obtained by oxidation with 1 mM hexaammineruthenium (III) and photoreduction in the presence of catalytic amounts (0.5 µM) of riboflavin, respectively. The inset shows the differential cp-Fldsqminus cp-Fldox spectrum
The electronic absorption spectrum of “as isolated” cp-Fld (Fig. 2b) exhibited two maxima at 352 and 577 nm, respectively. The spectrum resembles those for one-electron reduced flavodoxins (Mayhew 1978; Biel et al. 1996), namely, the “blue” neutral flavin radical (Massey and Palmer 1966). The contribution of the oxidized form with a maximum at 455 nm was quite small (~ 8%, Fig. 2b). These findings indicated that although cp-Fld was isolated under aerobic conditions, it was mainly represented by its semiquinone form.
Aerobic incubation of cp-Fld solution for 1 day at room temperature in the dark did not cause significant oxidation of the FMN semiquinone. Complete oxidation of FMN could be accomplished by incubation with 1 mM ferricyanide or hexaammineruthenium (III) and proceeded in a minute or second time-domain, respectively. The electronic absorption spectrum of the oxidized cp-Fld was similar to the spectra of other isolated flavodoxins (Biel et al. 1996; Lawson et al. 2004; Chowdhury et al. 2016) and exhibited maxima at 394 and 455 nm, with a shoulder at 485 nm (Fig. 2b). Hexaammineruthenium (III)-oxidized cp-Fld (cp-Fldox) was purified from the reaction mixture by metal affinity chromatography and used along with its “as isolated” semiquinone form (cp-Fldsq) in further characterizations.
The denaturation of cp-Fldox and cp-Fldsq, which was achieved by boiling in the presence of 1% SDS, resulted in FMN release into solution. Its amount was estimated using the known extinction coefficient (Cerletti 1959), which allowed calculation of the extinction coefficients for cp-Fldox and cp-Fldsq, ε455 and ε577, of 10.8 and 4.4 mM−1 cm−1, respectively (Fig. 2b).
EPR characterization of cp-Fldsq
The continuous-wave EPR spectrum of cp-Fldsq (Fig. 3a) demonstrated a radical signal at g = 2.00 with the peak-to-peak linewidth of about 20 G, characteristic of the neutral flavosemiquinones (Palmer et al. 1971), in agreement with the data in Fig. 2b. To further characterize this radical signal, its pulsed proton X-band ENDOR spectra were determined in radio frequency ranges of 7.5–22.5 (Fig. 3b) and 22.5–37.5 MHz (Fig. 3c). Resonances from the H(5) proton at 28 (peak) and 33 MHz (edge of the spectrum) in Fig. 3c provided direct support for the neutral character of the flavin radical in cp-Fldsq (Schleicher et al. 2010). Noteworthy, resonances from three protons of the methyl group attached at the C(8α) position were determined at 10.9 and 19.0 MHz (Fig. 3b), that is, the principal values A1 and A2 of the hyperfine coupling tensor for these protons were ≈ 8 MHz. This value is typical of the neutral flavin radicals in flavodoxins (Schleicher et al. 2010; Martínez et al. 2014), wherein the xylene ring of the isoalloxazine moiety of FMN is significantly exposed to the solvent (Schleicher et al. 2010).
EPR spectroscopy of cp-Fld. a X-band continuous-wave EPR spectrum of cp-Fldsq. Sample composition: ~ 3 mM cp-Fldsq, 100 mM KCl, 10 mM Tris–HCl (pH 8.0). EPR settings: microwave frequency, 9.6936 GHz; microwave power, 2 μW; temperature, 80 K; modulation amplitude, 1 G. b and c X-band Davies ENDOR spectra of cp-Fldsq in radio frequency regions of 7.5–22.5 and 22.5–37.5 MHz, respectively. Magnetic field corresponded to the maximum of the echo-detected EPR spectrum of cp-Fldsq. Sample composition was the same as for panel (a). Labels mark resonances from the hyperfine couplings arising from protons H(8α), H(6), and H(5), as well as the Larmor frequency of free proton (νH). The structure of the flavin isoalloxazine ring with labeled relevant atomic positions is shown as the inset in (c). Note that spectra b and c were measured under different instrumental settings (see Materials and Methods) and their amplitudes cannot, therefore, be directly compared
Redox titration of cp-Fld
Em values for quinone/semiquinone and semiquinone/quinol transitions were determined by redox titrations of cp-Fldox and cp-Fldsq in the presence of various redox indicators at pH 7.0, as described by Efimov et al. (2014). Neutral or positively charged indicators, such as resorufin (\(E^{\prime}_{0}\) = − 51 mV; all potentials cited in this paper are given vs Standard Hydrogen Electrode), Nile blue (\(E^{\prime}_{0}\) = − 116 mV), phenosafranine (\(E^{\prime}_{0}\) = − 252 mV) (Clark 1960), or methyl viologen (\(E^{\prime}_{0}\) = − 446 mV) (Michaelis and Hill 1933) were found to be most suitable. With negatively charged indicators, such as various indigo sulfonates, the redox equilibration was too slow for this purpose.
Reductive titration of cp-Fldox demonstrated that the quinone is reduced to its semiquinone approximately in parallel with Nile blue, after resorufin and before phenosafranine (Fig. 4). The titration curve could be satisfactorily described by the Nernst equation for a one-electron transition, with Em of –110 ± 5 mV. Further titration of cp-Fldsq by such a strong reducing agent as dithionite yielded only sub-stoichiometric amounts of quinol; namely, the titration curve was incomplete (Fig. 4). However, the known amplitude of the signal for the cp-Fldsq→ cp-Fldred transition, measured by shifting pH to 9.0 at the end of the titration, allowed estimation of the Em value for this transition of approximately − 530 mV from the left part of the titration curve.
Redox titration of cp-Fldox (filled symbols) and cp-Fldsq (open symbols). A typical titration curve is shown, which was reproducible in replicate runs. The redox dyes were as follows: open circles, methyl viologen; filled circles, phenosafranine; triangles, Nile blue; squares, resorufin. The lines show the best fit of the Nernst equation for one-electron transitions, yielding Em values of − 110 (solid curve) and − 530 mV (dotted curve)
Flavodoxin as an electron carrier in Chlorobium phaeovibrioides
The cp-Fld mRNA level in Chl. phaeovibrioides cells grown in the presence of 0.15 or 3 µM ferric citrate was determined by RT-qPCR using 16S rRNA as a reference. The ratio cp-Fld mRNA/rRNA ± standard deviation found in triplicate measurements was 0.47 ± 0.12 (× 10−3) and 0.13 ± 0.035 (× 10−3), respectively. The amount of cp-Fld mRNA is, therefore, measurable in the cells grown in the presence of excess Fe but is 3.5-fold greater in the cells grown in the low-Fe medium. This finding is consistent with the role of cp-Fld as a substituent for a Fe-containing protein in GSB.
This hypothesis was further corroborated and detailed using a PFOR reaction:
This reaction is involved in electron transfer from the RC, generally carried out by ferredoxin (Hauska et al. 2001), to an Arnon–Buchanan cycle enzyme (Buchanan and Arnon 1990). The ability of cp-Fld to react with PFOR was estimated by measuring cp-Fldox reduction by PFOR found in the membrane-depleted extract of Chl. phaeovibrioides cells. Its addition to the mixture of PFOR substrates and cp-Fldox resulted in a nearly linear increase in absorbance at 577 nm (phase I) (Fig. 5a). The differential spectrum of the reaction mixture after completion of this phase of the reaction (Fig. 5b) corresponded to the one-electron cp-Fldox → cp-Fldsq transition (Fig. 2b, inset), indicating the ability of PFOR to catalyze it. The rate of cp-Fldox reduction could be calculated from these data to be 22–25 nmol/min per 1 mg cell extract protein, which was very similar to PFOR activity (22 nmol/min per 1 mg cell extract protein) measured with 1 mM artificial electron acceptor methyl viologen. These findings are consistent with the data of Biel et al. (1996) for a different system.
PFOR-catalyzed reduction of cp-Fldox. a The time-course of absorbance change at 577 nm. Curve labels refer to the concentrations of cytoplasmic protein/cp-Fldox (milligram and micromole per liter, respectively). Phases I and II are indicated for only the black curve. b, c The difference spectra of the reaction mixture for phases I and II, respectively
The linear phase of the rise in absorbance seen in Fig. 5a was followed by its decline (phase II). The time before the onset of phase II was proportional to the amounts of the cell extract and cp-Fldox added, whereas the height of the peak depended only on the latter parameter (Fig. 5a). The resulting differential spectrum of phase II (Fig. 5c) corresponded to the cp-Fldsq → cp-Fldred transition (Bogachev et al. 2009). The initial rate of phase II was 8–12 nmol/min per 1 mg of cell extract protein. However, this transition did not proceed to completion, and only around one-sixth of cp-Fldsq was further reduced before the reaction stopped. The same yield was obtained when the “as isolated” cp-Fldsq was used in the PFOR-catalyzed reaction instead of cp-Fldox (data not shown). This finding indicated that the incompleteness of the reduction was caused by the extremely low Em value for the cp-Fldsq/cp-Fldred transition, rather than by substrate depletion.
The ability of cp-Fld to capture reducing equivalents initially produced by the RC of Chl. phaeovibrioides was deduced from experiments measuring the light-induced reduction of cp-Fldox in the presence of the high-potential electron donor β-mercaptoethanol, which demonstrated higher photoreduction rate by comparison with ascorbate plus 2,6-dichlorophenolindophenol. The rate of cp-Fldox reduction to cp-Fldsq by inverted membrane vesicles under illumination at 730 nm was 17 nmol/min per 1 mg of protein or 450 nmol/min per 1 mg of bacteriochlorophyll d (data not shown), which exceeded, considerably, the reported rate of ferredoxin reduction by the RC of GSB (Buchanan and Evans 1969). Although the cp-Fldox to cp-Fldsq transition can hardly have a physiological role, these data indicated the ability of cp-Fld to interact with RC. Upon completion of the cp-Fldox → cp-Fldsq transition, the reaction stopped, presumably because the photosynthetic electron transport chain switched to its cyclic functioning mode.
Discussion
Flavodoxin genes, found in many GSB, encode ≈ 17.5 kDa proteins that belong to the flav_long family (TIGR01752). Many properties of cp-Fld characterized in this study are typical of flavodoxins (Sancho 2006). Thus, cp-Fld contains a non-covalently bound FMN as the only prosthetic group, which, like in all known flavodoxins (Sykes and Rogers 1984), can undergo two consecutive one-electron reduction reactions, with profoundly different Em values to form its semiquinone and quinol forms.
The principal difference between cp-Fld and known flavodoxins is the unusual stability of the radical (semiquinone) form of FMN to oxidation by air. All currently described flavodoxins are isolated in a fully oxidized (quinone) form of FMN (Dubourdieu et al. 1975; Sykes and Rogers 1984; Biel et al. 1996; Lawson et al. 2004; Chowdhury et al. 2016; Segal et al. 2017), whereas FMN of the isolated cp-Fld is in the neutral semiquinone form. Based on spectral characteristics, the neutral form of the flavin radical is generally referred to as a “blue flavin radical,” to distinguish it from the anionic “red flavin radical” (Massey and Palmer 1966), and the isolated cp-Fld provides rare visual support for this nomenclature (Fig. 2a). Flavin, in a completely or partially radical state, was earlier observed in an “as isolated” multi-subunit Na+-translocating NADH:quinone oxidoreductase (Bogachev et al. 2002) and nitric oxide synthase (Perry et al. 1998), wherein the FMN group may be sterically shielded from air oxygen. In contrast, flavodoxins are small monomeric proteins with a surface-located FMN that is partially exposed to the solution (Watt et al. 1991), which should allow its direct contact with oxygen.
The Em value of the quinone/semiquinone transition in cp-Fld (− 110 mV) is among the highest for flavodoxins, whose Em values lie between − 90 and − 245 mV (Dubourdieu et al. 1975; Watt 1979; Sykes and Rogers 1984; Biel et al. 1996; Lawson et al. 2004; Segal et al. 2017). The high redox potential hampers electron transfer from FMN to oxygen, both thermodynamically and kinetically. The kinetic effect is manifested in the slow reaction with redox mediators, negatively charged, in particular. Conversely, the Em value of the semiquinone/quinol transition in cp-Fld is unusually low (− 530 mV) and lies outside the range generally found for other flavodoxins (between − 370 and − 480 mV) (Dubourdieu et al. 1975; Sykes and Rogers 1984; Biel et al. 1996; Lawson et al. 2004; Segal et al. 2017), the lowest reported value being − 515 mV (Azotobacter vinelandii flavodoxin I) (Watt 1979). Accordingly, the difference between the Em values for the quinone/semiquinone and semiquinone/quinol transitions in cp-Fld is the largest among all known flavodoxins, as is the stability constant for the semiquinone formation,
Consequently, the reduction of cp-Fldox to cp-Fldsq should be virtually complete before the onset of the cp-Fldsq → cp-Fldred reaction, explaining the sharp reversal of the slope in the time-course in Fig. 5a.
The structural basis for the unusual redox characteristics of cp-Fld remains to be determined. Noteworthy, the proton ENDOR spectrum of cp-Fldsq (Fig. 3b, c) is very similar to those for other flavodoxins (Schleicher et al. 2010; Martínez et al. 2014), indicating similar positioning of FMN and the exposure of the xylene ring of the isoalloxazine moiety to the solvent (Schleicher et al. 2010). Solving the 3D structure of cp-Fld will eventually solve this conundrum.
The extremely low Em value for the cp-Fldsq/cp-Fldred transition may be associated with a specific role of flavodoxin in GSB. As Fig. 5a highlights, only one-sixth of the cp-Fldsq pool can be reduced by PFOR, even under optimal conditions (high concentrations of pyruvate and CoA, low concentration of acetyl-CoA), presumably because of the lower Em value for the cp-Fldsq/cp-Fldred couple in comparison with the (pyruvate + CoA)/(acetyl-CoA + CO2) couple. As a contrasting example, Wolinella succinogenes flavodoxin, with the Em value of − 450 mV for the Fldsq/Fldred transition, is completely reduced in the PFOR-catalyzed reaction (Biel et al. 1996). The redox characteristics of cp-Fld are, thus, optimal for the reverse pyruvate synthase reaction of PFOR, consistent with the physiological requirement of the Arnon–Buchanan cycle in GSB (Buchanan and Arnon 1990). Noteworthy, the Em value of the semiquinone/quinol transition in cp-Fld (− 530 mV) is quite close to those for ferredoxins I and II (− 514 and − 584 mV, respectively) in the green sulfur bacterium Chlorobaculum tepidum (Yoon et al. 2001). This observation provides support for the proposed functional equivalence of flavodoxin and ferredoxin in GSB, that is, their common capacity to serve as the source of reducing equivalents in the PFOR-catalyzed pyruvate synthase reaction. In the reverse reaction, the carrier function belongs to rubredoxin, with a more positive redox potential (Yoon et al. 1999).
Beside the redox properties of flavodoxin, the hypothesis that it may substitute for ferredoxin in GSB under conditions of Fe limitation is supported by the results of the genome context analysis (Fig. 1) and the induction of cp-Fld gene expression at low Fe content in the growth medium. Noteworthy, the expression level was relatively high even in the Chl. phaeovibrioides cells grown in the Fe-rich medium (3 μM). This result is apparently explained by the presence of soluble sulfides in the used growth medium, which convert the added Fe into its low-soluble sulfides, like in the natural habitat of this and other GSB (Savvichev et al. 2018).
To summarize, Chl. phaeovibrioides cp-Fld, a first characterized flavodoxin of GSB, can functionally interact with the RC and PFOR of Chl. phaeovibrioides and substitute for ferredoxin in a Fe-deficient medium. The ease of cp-Fld production and the high stability of the protein itself and the FMN semiquinone make the novel flavodoxin an auspicious object for studies of the flavin radical by various EPR techniques, which generally require high amounts of the protein in the paramagnetic form.
Abbreviations
- cp-Fld, cp-Fldox, cp-Fldsq, cp-Fldred :
-
Flavodoxin from Chl. phaeovibrioides and its oxidized, semiquinone, and completely reduced forms, respectively
- E m :
-
Midpoint redox potential
- ENDOR:
-
Electron-nuclear double resonance
- GSB:
-
Green sulfur bacteria
- PFOR:
-
Pyruvate:ferredoxin oxidoreductase
- RC:
-
Photosynthetic reaction center
- RT-qPCR:
-
Quantitative reverse transcription polymerase chain reaction
References
Bertsova YV, Kostyrko VA, Baykov AA, Bogachev AV (2014) Localization-controlled specificity of FAD:threonine flavin transferases in Klebsiella pneumoniae and its implications for the mechanism of Na+-translocating NADH:quinone oxidoreductase. Biochim Biophys Acta 1837:1122–1129
Biel S, Klimmek O, Gross R, Kröger A (1996) Flavodoxin from Wolinella succinogenes. Arch Microbiol 166:122–127
Bogachev AV, Bertsova YV, Ruuge EK, Wikström M, Verkhovsky MI (2002) Kinetics of the spectral changes during reduction of the Na+-motive NADH:quinone oxidoreductase from Vibrio harveyi. Biochim Biophys Acta 1556:113–120
Bogachev AV, Bloch DA, Bertsova YV, Verkhovsky MI (2009) Redox properties of the prosthetic groups of Na+-translocating NADH:quinone oxidoreductase. 2. Study of the enzyme by optical spectroscopy. Biochemistry 48:6299–6304
Buchanan BB, Arnon DI (1990) A reverse KREBS cycle in photosynthesis: consensus at last. Photosynth Res 24:47–53
Buchanan BB, Evans MC (1969) Photoreduction of ferredoxin and its use in NAD(P)+ reduction by a subcellular preparation from the photosynthetic bacterium, Chlorobium thiosulfatophilum. Biochim Biophys Acta 180:123–129
Burnett RM, Darling GD, Kendall DS, LeQuesne ME, Mayhew SG, Smith WW, Ludwig ML (1974) The structure of the oxidized form of clostridial flavodoxin at 1.9-Ǻ resolution. J Biol Chem 249:4383–4392
Cerletti P (1959) Properties of riboflavin phosphates. Anal Chim Acta 20:243–250
Chowdhury NP, Klomann K, Seubert A, Buckel W (2016) Reduction of flavodoxin by electron bifurcation and sodium ion-dependent reoxidation by NAD+ catalyzed by ferredoxin-NAD+ reductase (Rnf). J Biol Chem 291:11993–12002
Clark WM (1960) Oxidation-reduction potentials of organic systems. Williams and Wilkin Co., Baltimore
De Serrano LO, Camper AK, Richards AM (2016) An overview of siderophores for iron acquisition in microorganisms living in the extreme. Biometals 29:551–571
Dubourdieu M, le Gall J, Favaudon V (1975) Physicochemical properties of flavodoxin from Desulfovibrio vulgaris. Biochim Biophys Acta 376:519–532
Efimov I, Parkin G, Millett ES, Glenday J, Chan CK, Weedon H, Randhawa H, Basran J, Raven EL (2014) A simple method for the determination of reduction potentials in heme proteins. FEBS Lett 588:701–704
Fazekas AG, Kokai K (1971) Extraction, purification, and separation of tissue flavins for spectrophotometric determination. Methods Enzymol 18:385–398
Fromme P (1999) Biology of photosystem I: structural aspects. In: Singhal GS, Renger G, Sopory SK, Irrgang KD, Govindjee (eds) Concepts in photobiology. Narosa Publishing House, New Dehli, pp 181–220
Hauska G, Schoedl T, Remigy H, Tsiotis G (2001) The reaction center of green sulfur bacteria. Biochim Biophys Acta 1507:260–277
LaRoche J, Boyd PW, McKay RML, Geider RJ (1996) Flavodoxin as an in situ marker for iron stress in phytoplankton. Nature 382:802–805
Lawson RJ, von Wachenfeldt C, Haq I, Perkins J, Munro AW (2004) Expression and characterization of the two flavodoxin proteins of Bacillus subtilis, YkuN and YkuP: biophysical properties and interactions with cytochrome p450 BioI. Biochemistry 43:12390–12409
Malik KA (1983) A modified method for the cultivation of phototrophic bacteria. J Microbiol Methods 1:343–352
Martínez JI, Alonso PJ, García-Rubio I, Medina M (2014) Methyl rotors in flavoproteins. Phys Chem Chem Phys 16:26203–26212
Massey V, Palmer G (1966) On the existence of spectrally distinct classes of flavoprotein semiquinones. A new method for the quantitative production of flavoprotein semiquinones. Biochemistry 5:3181–3189
Mayhew SG (1978) The redox potential of dithionite and SO−2 from equilibrium reactions with flavodoxins, methyl viologen and hydrogen plus hydrogenase. Eur J Biochem 85:535–547
Michaelis L, Hill ES (1933) The viologen indicators. J Gen Physiol 16:859–873
Palmer G, Muller F, Massey V (1971) Electron paramagnetic resonance studies on flavoprotein radicals. In: Kamin H (ed) Flavins and flavoproteins. University Park Press & Butterworths, Baltimore, pp 123–139
Peelen S, Wijmenga S, Erbel PJ, Robson RL, Eady RR, Vervoort J (1996) Possible role of a short extra loop of the long-chain flavodoxin from Azotobacter chroococcum in electron transfer to nitrogenase: complete 1H, 15N and 13C backbone assignments and secondary solution structure of the flavodoxin. J Biomol NMR 7:315–330
Perry JM, Moon N, Zhao Y, Dunham WR, Marletta MA (1998) The high-potential flavin and heme of nitric oxide synthase are not magnetically linked: implications for electron transfer. Chem Biol 5:355–364
Pierella Karlusich JJ, Lodeyro AF, Carrillo N (2014) The long goodbye: the rise and fall of flavodoxin during plant evolution. J Exp Bot 65:5161–5178
Sakurai H, Ogawa T, Shiga M, Inoue K (2010) Inorganic sulfur oxidizing system in green sulfur bacteria. Photosynth Res 104:163–176
Sancho J (2006) Flavodoxins: sequence, folding, binding, function and beyond. Cell Mol Life Sci 63:855–864
Savvichev AS, Babenko VV, Lunina ON, Letarova MA, Boldyreva DI, Veslopolova EF, Demidenko NA, Kokryatskaya NM, Krasnova ED, Gaisin VA, Kostryukova ES, Gorlenko VM, Letarov AV (2018) Sharp water column stratification with an extremely dense microbial population in a small meromictic lake, Trekhtzvetnoe. Environ Microbiol 20:3784–3797
Schleicher E, Wenzel R, Ahmad M, Batschauer A, Essen LO, Hitomi K, Getzoff ED, Bittl R, Weber S, Okafuji A (2010) The electronic state of flavoproteins: investigations with proton electron-nuclear double resonance. Appl Magn Reson 37:339–352
Segal HM, Spatzal T, Hill MG, Udit AK, Rees DC (2017) Electrochemical and structural characterization of Azotobacter vinelandii flavodoxin II. Protein Sci 26:1984–1993
Sétif P (2001) Ferredoxin and flavodoxin reduction by photosystem I. Biochim Biophys Acta 1507:161–179
Simondsen RP, Tollin G (1980) Structure-function relations in flavodoxins. Mol Cell Biochem 33:13–24
Smith PK, Krohn RI, Hermanson GT, Mallia AK, Gartner FH, Provenzano MD, Fujimoto EK, Goeke NM, Olson BJ, Klenk DC (1985) Measurement of protein using bicinchoninic acid. Anal Biochem 150:76–85
Sykes GA, Rogers LJ (1984) Redox potentials of algal and cyanobacterial flavodoxins. Biochem J 217:845–850
Watanabe T, Honda K (1982) Measurement of the extinction coefficient of the methyl viologen cation radical and the efficiency of its formation by semiconductor photocatalysis. J Phys Chem 86:2617–2619
Watt GD (1979) An electrochemical method for measuring redox potentials of low potential proteins by microcoulometry at controlled potentials. Anal Biochem 99:399–407
Watt W, Tulinsky A, Swenson RP, Watenpaugh KD (1991) Comparison of the crystal structures of a flavodoxin in its three oxidation states at cryogenic temperatures. J Mol Biol 218:195–208
Yoon KS, Hille R, Hemann C, Tabita FR (1999) Rubredoxin from the green sulfur bacterium Chlorobium tepidum functions as an electron acceptor for pyruvate ferredoxin oxidoreductase. J Biol Chem 274:29772–29778
Yoon KS, Bobst C, Hemann CF, Hille R, Tabita FR (2001) Spectroscopic and functional properties of novel 2[4Fe-4S] cluster-containing ferredoxins from the green sulfur bacterium Chlorobium tepidum. J Biol Chem 276:44027–44036
Acknowledgements
This work was supported by the Russian Science Foundation research project 19-14-00063. We are indebted to Prof. R.N. Ivanovsky for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bertsova, Y.V., Kulik, L.V., Mamedov, M.D. et al. Flavodoxin with an air-stable flavin semiquinone in a green sulfur bacterium. Photosynth Res 142, 127–136 (2019). https://doi.org/10.1007/s11120-019-00658-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-019-00658-1