Abstract
The development of oxygenic photosynthesis by primordial cyanobacteria ~2.7 billion years ago led to major changes in the components and organization of photosynthetic electron transport to cope with the challenges of an oxygen-enriched atmosphere. We review herein, following the seminal contributions as reported by Jaganathan et al. (Functional genomics and evolution of photosynthetic systems, vol 33, advances in photosynthesis and respiration, Springer, Dordrecht, 2012), how these changes affected carriers and enzymes at the acceptor side of photosystem I (PSI): the electron shuttle ferredoxin (Fd), its isofunctional counterpart flavodoxin (Fld), their redox partner ferredoxin-NADP+ reductase (FNR), and the primary PSI acceptors F x and F A/F B. Protection of the [4Fe–4S] centers of these proteins from oxidative damage was achieved by strengthening binding between the F A/F B polypeptide and the reaction center core containing F x, therefore impairing O2 access to the clusters. Immobilization of F A/F B in the PSI complex led in turn to the recruitment of new soluble electron shuttles. This function was fulfilled by oxygen-insensitive [2Fe–2S] Fd, in which the reactive sulfide atoms of the cluster are shielded from solvent by the polypeptide backbone, and in some algae and cyanobacteria by Fld, which employs a flavin as prosthetic group and is tolerant to oxidants and iron limitation. Tight membrane binding of FNR allowed solid-state electron transfer from PSI bridged by Fd/Fld. Fine tuning of FNR catalytic mechanism led to formidable increases in turnover rates compared with FNRs acting in heterotrophic pathways, favoring Fd/Fld reduction instead of oxygen reduction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oxygenic photosynthesis involves the combined function of two multi-subunit membrane-bound complexes, photosystem (PS) I and II, which provide the energy transducing machinery to drive electrons against a thermodynamic gradient at the expense of light (Jagannathan et al. 2012). Coupling of these two photosystems in the photosynthetic electron transport chain (PETC) of primordial cyanobacteria was the seminal event in the development of oxygenic photosynthesis. On one end of the PETC, the evolution of a water-splitting system led to oxygen production, an event of catastrophic proportions that changed the redox conditions on Earth and allowed the appearance of life forms based on oxygen respiration. On the opposite end, reduced intermediates were produced in the form of NADPH and low-potential electron transfer shuttles to deliver hydride groups and electrons to a plethora of metabolic, dissipative and regulatory pathways (Hohmann-Marriott and Blankenship 2011).
Electron shuttling is the currency for energy flow throughout cellular metabolism. Reducing equivalents are mobilized across networks of redox enzymes from the sources of reducing power, including light-driven reactions, to electron-consuming pathways and processes via a small suite of diffusible electron carriers. The most extensively used among them are ferredoxins (Fds), small mobile metalloproteins containing one or more iron-sulfur clusters of different stoichiometry ([4Fe–4S], [2Fe–2S]) as prosthetic groups. Ferredoxins, whose name, according to San Pietro (2006) were chosen after a contest at the Dupont lab where the first Fd (from Clostridium pasteurianum) was discovered, are very ancient proteins that were extensively employed as electron shuttles by anaerobes long before the advent of oxygenic photosynthesis (Caetano-Anollés et al. 2007). Those present in oxygen-evolving phototrophs harbor a single [2Fe–2S] center, and constitute a large protein family that also includes Fds from mitochondria and proteobacteria (Hase et al. 2006; Meyer 2008).
The acceptor end of the PETC in plants is made up by a group of soluble and thylakoid-bound components, including the primary acceptors of PSI, Fd and the flavoprotein ferredoxin-NADP+ reductase (FNR), which catalyzes an electron-hydride exchange between reduced Fd and NADP+ to yield NADPH (Carrillo and Ceccarelli 2003; Aliverti et al. 2008). Oxygen build-up forced major changes on these components to adapt to the novel, potentially harmful environmental conditions. In spite of these adaptations, the acceptor side of PSI remains the main site of oxygen reduction in photosynthetic cells and tissues (Kozuleva and Ivanov 2016). It is worth noting that this reduction leads to the synthesis of various reactive oxygen species (ROS) such as the radical anion superoxide, hydrogen peroxide, and the hydroxyl radical, all of which are more damaging to biomolecules than oxygen itself.
We review in this article the selective forces and evolutionary changes that shaped the composition and function of the acceptor end of the PETC in modern-day phototrophic organisms, including the incorporation of new players and the modification of existing ones to meet the challenges of an oxygen-rich atmosphere.
Phototrophy before oxygen
Current proposals on the living conditions prevailing in the young Earth agree that primordial cells thrived in an anaerobic world by exploiting every available redox pair to obtain energy from purely chemotrophic metabolic pathways (Zahnle et al. 2007). In this oxygen-free environment, photosynthetic reaction centers likely originated from pre-existing pigment-carrying proteins involved in protection against ultraviolet irradiation, through sequential acquisition of redox cofactors that helped to stabilize charge separation states (Mulkidjanian and Junge 1997). If so, original photosynthesis must have been anoxygenic and presumably based on a single photosystem.
A survey of extant phototrophic microorganisms indicates that, besides cyanobacteria, there are six bacterial phyla with members able to perform chlorophyll-based photosynthesis: Firmicutes (heliobacteria), Chlorobi (green sulfur bacteria), Chloroflexi (green non-sulfur bacteria) and Proteobacteria (purple sulfur and non-sulfur bacteria), as well as recently added species from Acidobacteria and Gemmatimonadetes (Zeng et al. 2014). Heliobacteria, Chlorobi, and Acidobacteria contain Type I reaction centers with an iron-sulfur cluster as the terminal acceptor, therefore resembling PSI. The remaining three phyla have Type II centers that use a mobile quinone as terminal acceptor, similar to PSII. Of them, only Chlorobi are strict photoautotrophs. The two types of photosystems are proposed to derive from a common ancestor by gene duplication and divergence, with basal sequences lying near the split between heliobacteria and green sulfur bacteria (Raymond and Blankenship 2006). Therefore, the original phototrophs may have contained a Type I reaction center (Blankenship 2001), similar to that present in heliobacteria, which has the most rudimentary photosystem described to date (Oh-Oka 2007).
While there is ample agreement among researchers that oxygenic photosynthesis arose after anoxygenic photosynthesis, the question of whether the two photosystems of cyanobacteria became divergent in the same or different genomes is still a matter of debate (Hohmann-Marriott and Blankenship 2011). Initial models on the evolution of the photosynthetic apparatus assumed that the current PETC architecture with photosystems operating in tandem evolved from the union of pre-existing Type I and II reaction centers within a primordial cyanobacterium (Raymond and Blankenship 2006, and references therein). However, Mulkidjanian et al. (2006) have advanced an alternative proposal, suggesting that cyanobacteria were actually the earliest phototrophs and contained the two types of reaction centers, which were delivered to other phyla by independent lateral gene transfer events well before the development of the oxygen-evolving machinery associated to PSII. This hypothesis gains support from the lack of a deep split in chlorophyll biosynthesis that would mirror the split of the photosystems (Sousa et al. 2013), suggesting that the two reaction centers arose and diverged within the same cell. Allen and Martin (2007) have postulated that in these primitive cyanobacteria, the co-existing photosystems were not coupled and played different roles, with the Type I center engaged in linear electron transfer using H2, H2S or ferrous iron as electron donors, and the Type II center carrying out cyclic electron transport. The proposed functions are exactly the opposite of those displayed by their descendants, PSI and PSII, in modern-day oxygenic phototrophs. They resemble, instead, the activities of type I reaction centers in current green sulfur bacteria, which mediate linear electron transport between H2S and Fd, and of type II reaction centers in purple non-sulfur bacteria, which generate ATP via cyclic electron flow (Jagannathan et al. 2012). The opposing views in this active field of research are outside the scope of this article. The subject has been extensively reviewed by Cardona (2015).
As the availability of suitable electron donors became progressively limiting, the acquisition of the Mn4Ca cluster capable of water oxidation would have had a formidable selective advantage, and the presence of both reaction centers in a single organism allowed them to operate concertedly to synthesize oxygen and provide reducing power for metabolic routes, giving birth to oxygenic photosynthesis.
Oxygen build-up as driving force for the evolution of the reducing side of PSI
The fossil record, especially in the form of stromatolites, times the onset of oxygenic photosynthesis at about 2.7 billion years ago (Kerr 2005). Reaction with ferrous and sulfide ions, as well as a number of geological processes, sequestered enough atmospheric oxygen for the Earth to remain anaerobic during the following 2 billion years or so, but the continuous production of intracellular oxygen started a number of adaptations which affected all aspects of metabolism and regulation, including photosynthesis (Kirschvink and Kopp 2008). Components of the reducing side of PSI were among the most affected by these environmental changes.
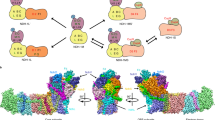
As already indicated, Type I photosystems rely heavily on iron-sulfur clusters. The core complex, which is a heterodimer (PsaA/PsaB) in PSI and a homodimer in bacterial photosystems, contains the reaction center chlorophylls (P700) and three other electron transfer components operating sequentially: chlorophyll A0, phylloquinone A1 and the [4Fe–4S] cluster F x, associated to the PsaC protein. Electrons are then delivered to a di-cluster protein containing two [4Fe–4S] centers (F A and F B), in the sequence F x → F A → F B, and eventually to Fd and FNR (Jagannathan et al. 2012). The reader is referred to the article by Pierre Setif in this issue for a comprehensive description of PSI structure and electron transfer properties.
Oxygen build-up negatively affected the functions of these metalloproteins in a number of ways. First, spin-pairing rules dictate that molecular oxygen accepts electrons one at a time rather than in pairs, discouraging reaction with most organic biomolecules but facilitating oxidation of transition metals, which are good univalent electron donors. As a consequence, oxygen oxidized ferrous iron in the environment to its ferric form, which rapidly precipitated as ferric polyhydroxides or formed insoluble salts. The outcome was that as oxygen accumulated, iron decreased its bioavailability and became a limiting nutrient in most aerobic habitats (Anbar 2008).
Second, partial reduction of oxygen generates superoxide, H2O2 and hydroxyl radicals, oxidants which display still higher reactivity than oxygen. Even under normal growth conditions, up to 10% of all electrons moving through the photosynthetic or respiratory chains can be adventitiously delivered to oxygen with concomitant ROS generation (Badger et al. 2000). This fraction increases dramatically under adverse environmental conditions, which also promote the generation of highly reactive singlet oxygen molecules by energy transfer from excited chlorophylls of PSII (Pierella Karlusich et al. 2014; Kozuleva and Ivanov 2016). These compounds can react with and inactivate many different biomolecules including proteins, lipids, and nucleic acids (Møller et al. 2007).
Iron-sulfur centers are vulnerable to ROS attack to various extents, depending on solvent exposure and the polypeptide environment surrounding the cluster. Oxidation yields unstable intermediates that quickly decompose, resulting in protein inactivation and iron release. Elevated concentrations of free iron can wreak cellular havoc and lead to oxidative damage by engaging in Fenton-type reactions with hydrogen peroxide to generate the extremely toxic hydroxyl radical (Imlay 2006). This process is generally self-propagating. In photosynthetic organisms, for instance, stress-dependent Fd down-regulation leads to over-reduction of the PETC due to shortage of electron acceptors, and under such circumstances the electron surplus can be passed straight to oxygen resulting in runaway ROS generation (Voss et al. 2008; Blanco et al. 2011).
The type of modifications undergone by photosynthetic organisms in response to the threat of an increasingly oxidative environment can be visualized by comparing components of the PETC in cyanobacteria and chloroplasts with their equivalent counterparts of anaerobic phototrophs. Changes can be tracked in F x and F A/F B, in the mobile low-potential shuttles and in FNR.
Of the various iron-sulfur clusters operating at the acceptor side of Type I reaction centers of anaerobes, the equivalents of F x, F A, and F B are particularly oxygen-labile. In the anaerobic world in which photosynthesis initially evolved, these terminal electron carriers had no need to be protected against oxidative destruction. They were just placed at the shortest possible distance to optimize the rate of forward electron transfer. The possibility of side-reaction with increasing oxygen levels required significant modifications to protect the F A/F B cluster from denaturation. The polypeptide holding these iron-sulfur centers displayed a loose and dynamic interaction with the reaction center core in anaerobic phototrophs, which contrasts with the tight binding of PsaC to PSI in chloroplasts and cyanobacteria. Indeed, contemporary heliobacteria and green sulfur bacteria still harbor mobile iron-sulfur proteins displaying high oxygen sensitivity (Romberger et al. 2010). Jagannathan et al. (2012) have proposed that the increase in binding interaction, achieved through the development of a network of oppositely charged amino acids in the F x and F A/F B proteins, was critical to the protection of the clusters by the formation of an oxygen-impenetrable interface between them. Fastening of PsaC to the PsaA/PsaB heterodimer was further ensured by acquisition of PsaD and PsaE subunits. These proteins flank PsaC on either side of the stromal ridge of PSI and provide additional solvent shielding to F A/F B (Antonkine et al. 2003).
As important as this protection could be, this was not the only modification undergone by the primary acceptors of PSI. In the reaction centers of chloroplasts and cyanobacteria there is an inversion of the redox potentials between F A and F B that makes the F A → F B electron transfer thermodynamically unfavorable. The penalty introduced by this change in the global rates of electron transfer (and in the quantum yield) is marginal, due to the high difference in redox potential between F x and the mobile acceptor of F B (Jagannathan et al. 2012). However, the inversion changed dramatically the half-life of reduced F B, and therefore its availability to oxidation by oxygen. The difference in midpoint potential with F A (~60 mV) implies that only ~10% of reduced F B would be present at any given time for reaction with oxygen (Jagannathan et al. 2012).
Weak binding of the di-cluster F A/F B protein to Type I reaction centers in anaerobes allowed these components to act as shuttles between F x and soluble redox targets (Jagannathan and Golbeck 2008; Romberger and Golbeck 2010, 2012). A consequence of F A/F B immobilization in oxygenic phototrophs was the requirement of an alternative mobile electron carrier to fulfill this role. This led to the evolution of oxygen-insensitive Fds and the recruitment of flavodoxin (Fld) at the reducing side of PSI.
Beyond PSI: the mobile electron shuttles
Development of an oxygen-insensitive ferredoxin
Several lines of evidence support the notion that iron-sulfur clusters were among the earliest, if not the first, biocatalysts on Earth (Wächtershäuser 2007). They are chemically versatile and can form spontaneously in the presence of Fe2+, sulfide, and an external thiolate (Martin and Russel 2003), all of which were abundant in the anaerobic Archaean ocean. Moreover, Fe–S cubanes can be readily assembled into cysteine-containing polypeptides through a thiol-ligand exchange reaction (Que et al. 1974). As a consequence, iron-sulfur proteins were widely used by anaerobes, both photosynthetic and heterotrophic, prior to the appearance of oxygen evolution, and Fds were among the oldest of them. Inferences on the physiology and habitat of the last universal common ancestor (LUCA) suggest that LUCA’s biochemistry was replete with Fe–S centers and radical-based reactions, its lifestyle relying heavily on transition metals, flavins and Fds among other basic components (Weiss et al. 2016).
The Fd apoprotein is made up of a conserved fold of ~60 amino acids that binds the Fe–S cluster and presumably evolved from an early gene duplication event of a 30-residue sequence, a “protoferredoxin” composed of a primeval subset of amino acids (Eck and Dayhoff 1966; Trifonov 2000). Phylogenomic analysis of protein architecture indicates that this fold is very ancient, actually the fifth oldest domain on Earth, and second among redox-related structures after the NAD(P+)(H)-binding Rossmann domain (Caetano-Anollés et al. 2007).
Based on fold mapping, high network centrality, and abundance in anaerobic, thermophilic environments, it has been concluded that [4Fe–4S] clusters were the first to be formed and assembled into proteins in the O2-free primordial oceans. They are more flexible and chemically versatile than their [2Fe–2S] counterparts, being able to act as Lewis acids in dehydration reactions catalyzed by hydro-lyases such as fumarase and aconitase. Using profile-vs-profile and position-specific iterative-BLAST alignments, Harel et al. (2014) concluded that Fds with [4Fe–4S] clusters were the ancestral representatives of the family, from which [2Fe–2S] Fds followed by duplication and divergence. They also found connections of iron-sulfur proteins with 4-Cys iron (FeS4) and heme domains, all of them sharing a common origin, whereas bimetallic domains such as the MoFe and VFe centers of nitrogenase were more distantly related (Harel et al. 2014).
Anaerobes thriving prior to oxygen evolution were then supposed to use mostly [4Fe–4S] Fds as low-potential electron carriers, and some contemporary phototrophs still make ample use of these proteins for electron transfer. For instance, heliobacteria and green sulfur bacteria have small, mobile Fds with two [4Fe–4S] clusters that accept electrons from the Type I reaction center (Jagannathan and Golbeck 2008; Romberger and Golbeck 2010, 2012; García Costas et al. 2012).These carriers are typically much more susceptible to oxygen inactivation than the photosynthetic [2Fe–2S] Fds present in contemporary cyanobacteria and chloroplasts. Oxygen insensitivity was gained by solvent extrusion from the [2Fe–2S] cluster, which is located in a shielded environment in these proteins, whereas the µ-sulfido atoms of typical anaerobic bacterial Fds are solvent exposed (Jagannathan et al. 2012). Fds with [2Fe–2S] clusters replaced existing oxygen-sensitive counterparts, and evolved to get even greater levels of oxygen tolerance, without substantially altering their capacity for rapid electron exchange with the F A/F B protein of the reaction center (PsaC in chloroplasts and cyanobacteria). Fd docking was optimized through interactions with the PsaD and PsaE subunits. Analysis of the crystal structure of cyanobacterial PSI indicates that the main forces involved in Fd binding are electrostatic, between acidic residues of the electron shuttle and positive charges on PsaC, PsaD, and PsaE (Fromme et al. 2001).
Fds containing [2Fe–2S] clusters are found in both photosynthetic and non-photosynthetic organisms. They group together in a general phylogenetic tree that includes Fd-like domains of multi-domain enzymes (Fig. 1a), thus confirming a monophyletic origin for all these Fds. It is possible to distinguish two major subfamilies: the Fds from cyanobacteria and plastids, which constitute the “plant-type” cluster (Fig. 1b), and those present in mitochondria and most heterotrophic microorganisms (Fig. 1a), that have been termed “proteo-type” Fds and include adrenodoxin and putidaredoxin (Ewen et al. 2011). Yeh et al. (2000) have identified still another family of [2Fe–2S] Fds based on a thioredoxin-like fold, which are unrelated to those depicted in Fig. 1.
Phylogenetic relationships among [2Fe–2S] Fds and Fd-like domains. a Phylogenetic tree of Fd and Fd-like domain sequences. The tree topology indicates that Fds from cyanobacteria and plastids have a monophyletic origin. The same holds true for the Fds from proteobacteria and mitochondria (“proteo-type”). b Phylogeny of Fds from cyanobacteria and plastids, amplified from the corresponding region of panel a. Sequences are color-coded according to their taxonomic assignment. The scale bars indicate the number of expected amino acid substitutions per site per unit of branch length. Sequences corresponding to PFAM domain PF00111 were aligned with MAFFT version 6 (Katoh and Toh 2008) and the trees were calculated using PhyML 3.0 (Guindon et al. 2010), using the LG model of amino acid substitution and four substitution rate categories
Canonical Fds from photosynthetic organisms form a highly supported cluster which includes those present in non-photosynthetic plastids from plant roots and alveolates, and the Fd isoforms responsible for electron donation to nitrogenase in diazotrophic β-cyanobacteria (FdxH, Fig. 1b). The FdC1 and FdC2 proteins from plants, green algae, and cyanobacteria are more divergent [2Fe–2S] proteins of unknown function (Voss et al. 2011; Zhao et al. 2015; Li et al. 2015), but they also share a common origin with canonical plant-type Fds (Fig. 1b). The Fd tree is plagued with putative interdomain HGT events involving gene transfers between oxygenic phototrophs, the Halobacteria archaeal class, and even a few proteobacteria/actinobacteria/Aquificae (Armengaud et al. 1994; Frolow et al. 1996; Meyer et al. 2002; Sjöblom et al. 2008; Grinter et al. 2012; Pierella Karlusich et al. 2015). These sequences appear scattered throughout the tree (Fig. 1b). In addition, several studies have demonstrated the acquisition of photosynthetic genes, including those encoding Fd, by different marine cyanophages. This incorporation is thought to help maintain host photosynthetic activity during infection (Lindell et al. 2004).
The paralogy of Fd-coding genes varies among species. Most cyanobacteria contain three or more isoforms, including divergent Fds such as FdC1 and FdC2 (Poncelet et al. 1998; Voss et al. 2011). Among eukaryotes, glaucophytes and algae from the red-plastid lineage usually have a single Fd encoded in the plastid genome, whereas green algae and land plants contain numerous isoforms expressed from nuclear genes. Six different Fd variants have been described in Chlamydomonas reinhardtii, a freshwater chlorophyte (Peden et al. 2013; see also Dubini, this issue), in the C3 plant Arabidopsis thaliana, with isoforms located in leaves and roots (Hanke et al. 2004; Voss et al. 2011), and in the C4 plant maize, present in root plastids and in mesophyll and bundle sheath chloroplasts (Matsumura et al. 1997; Sakakibara 2003; Cheng et al. 2008). Photosynthetic isoforms usually display higher level of sequence identity within the same species than among different species, indicating that they are the result of relatively recent gene duplication events (Hanke and Hase 2008). In general, the main Fd isoform engaged in photosynthetic electron transport accounts for 80–90% of the total Fd pool in leaves (Hanke et al. 2004; Terauchi et al. 2009).
Fd expression is affected by many environmental factors. All isoforms are down-regulated by iron starvation and adverse conditions leading to oxidative stress (Thimm et al. 2001; Singh et al. 2003; Mazouni et al. 2003; Tognetti et al. 2006; Hruz et al. 2008; Terauchi et al. 2009; Thompson et al. 2011; Ceccoli et al. 2011). Tissue specificity broadly correlates with labor division and expression patterns. Isoforms involved in photosynthesis are usually light-responsive (Vorst et al. 1993; Lemaire et al. 1999; Whitney et al. 2011). This regulation is complex and involves post-transcriptional mechanisms (Dickey et al. 1992). Root Fds that participate in nitrogen assimilation as electron donors for nitrite reductase are induced by nitrate and other oxidized nitrogen sources, and repressed by ammonia (Matsumura et al. 1997; Patterson et al. 2010). Two Fd isoforms are present in maize root plastids; one of them is expressed constitutively (FdIII), while the other (FdVI) responds to nitrate (Matsumura et al. 1997). FdVI is also nitrate induced in mesophyll chloroplasts (Sakakibara 2003). It is unclear whether this Fd is co-expressed with the photosynthetic isoform in the same cells and chloroplasts, or if it is present in heterotrophic cells of the leaf (Hanke and Mulo 2013). Some of the non-photosynthetic isoforms have less negative midpoint redox potentials compared to photosynthetic Fds (Hanke et al. 2004; Gou et al. 2006), indicating that they may have a narrower range of suitable electron partners (Peden et al. 2013).
Dinitrogen fixation in microorganisms is an anaerobic process because nitrogenase is very sensitive to oxygen inactivation (Schrautemeier et al. 1995). Nitrogen-fixing cyanobacteria usually express one or more FdxH isoforms as dedicated electron donors for the nitrogenase, in addition to the major Fd variant that mediates photosynthesis. Expression of these “diazotrophic” Fds is regulated by different mechanisms depending on the morphological and/or physiological strategy for dinitrogen fixation adopted by the microorganism. In those cyanobacteria in which heterocysts provide the anaerobic environment, such as Anabaena sp. PCC 7120, a Fd isoform specific for these differentiated cells is expressed under dinitrogen-fixing conditions (Masepohl et al. 1997). In contrast, in the non-heterocystous, filamentous cyanobacterium Plectonema boryanum, which fixes nitrogen only in low oxygen environments, the single FdxH accumulates under microaerobic conditions (Schrautemeier et al. 1994). The two types of FdxH isoforms and regulatory mechanisms can be present in the same organism, as it occurs in heterocyst-forming Anabaena variabilis (Schrautemeier et al. 1995).
Then, as the atmosphere became progressively oxidant, recruitment, expansion, and diversification of [2Fe–2S] Fds increased exponentially from anaerobes to facultatives to aerobes, including phototrophs (Harel et al. 2014). While the modifications introduced on Fd sequence, structure, and interactions significantly increased the capacity of these proteins to tolerate the oxygen threat, the other unwanted consequence of an oxidative atmosphere, namely, limitation of iron bioavailability, could not be solved. Fld was the ultimate solution, since it is both oxygen-insensitive and does not require iron.
Recruitment of flavodoxin at the acceptor side of photosystem I
Flds are electron carriers that contain flavin mononucleotide as prosthetic group instead of an iron-sulfur center (Sancho 2006). Fld redox properties largely match those of Fd, being able to replace the metalloprotein in most reactions (Pierella Karlusich et al. 2014). They are present in several bacterial phyla (including cyanobacteria), and in the chloroplasts of some algae, but not in plants, fungi, or animals (Pierella Karlusich et al. 2015). Both the [2Fe–2S] cluster of Fd and the flavin group of Fld can in principle exchange one or two electrons. However, experimental measurements showed that they behave as obligatory one-electron carriers, switching between the Fe3+, Fe3+/Fe3+, Fe2+ (Bott 1999) and the semiquinone/hydroquinone states, respectively (Nogués et al. 2005). These transitions have similar redox potentials (−400 to −430 mV for the photosynthetic isoforms), which allow the two proteins to fulfill their roles as low-potential electron shuttles.
Photosynthetic Fd has been shown to be more efficient than Fld in all reactions assayed in vitro, including NADP+ photoreduction by isolated thylakoids and thioredoxin reduction by Fd-thioredoxin reductase in reconstituted systems (Tognetti et al. 2006). FNR-mediated reactions are the best characterized at the kinetic level, revealing some interesting features. The k cat/K M value of Anabaena Fd was reported to be ~25-fold higher than that of Fld (Medina et al. 1998). The strength of the interactions with Anabaena FNR, as reflected by the Michaelis constants, was similar for both carriers, indicating that the gain in efficiency was at the expense of electron transfer rates (Medina et al. 1998). It is likely that the complex electronic configuration of the flavin is not flexible enough to attain the high electron transfer rates typical of transition metals. Iron ions contain incompletely filled d orbitals which can readily accept electrons from different partners with various geometries, making them particularly versatile in oxido-reductive processes. It is actually remarkable that some of these electron transfer reactions can be mimicked by the particular arrangement of π-orbitals found in the isoalloxazine ring system.
The subcellular location of Fld in phototrophs corresponds to that of Fd, namely, the cyanobacterial cytosol and algal chloroplasts (La Roche et al. 1996). When both carriers are present in the same species, the Fld gene is typically induced as an adaptive resource under environmental or nutritional hardships that compromise Fd expression or activity such as iron limitation and oxidative stress. The relevance of this replacement for survival and reproduction varies among species. In the cyanobacterium Synechocystis sp. PCC 6803, for instance, disruption of the Fld-coding gene has no fatal consequences, even under iron limitation (Kutzki et al. 1998), whereas the gene encoding the main Fd isoform was found to be essential in spite of Fld induction (Poncelet et al. 1998). However, several exceptions to this rule are documented, and a few Fld-specific pathways have been described in non-photosynthetic prokaryotes. Indeed, Fld is an essential gene in Escherichia coli and Helycobacter pylori, while Fd is not (Zheng et al. 1999; Freigang et al. 2002; Puan et al. 2005).
Fld is induced under various environmental hardships to take over Fd functions, but iron limitation appears to be the key factor determining its adaptive value. Evaluation of genomic and metagenomic data showed that microorganisms lacking this flavoprotein are usually confined to iron-rich coastal areas while Fld-containing marine microorganisms are preferably located in oceanic environments (Toulza et al. 2012; Pierella Karlusich et al. 2015). The importance of Fld in the dynamics of sea ecology can be gauged by its use as a proxy for iron deficiency in the oceans (La Roche et al. 1996; Erdner and Anderson 1999). Since Fld role as a backup of Fd appears to be a response to aerobiosis, it is natural to assume that Fld evolved well after Fd. Surprisingly, this is not the case.
Phylogenomic evaluation of protein fold architecture actually placed the Fld structure among the nine most ancestral and widely shared folds, appearing immediately after Fd, and long before diversification of prokaryotes (Caetano-Anollés et al. 2007). Then, these early Flds either played roles independent from Fd substitution or responded to environmental cues not related to aerobiosis and/or iron starvation. Most likely, they were recruited to replace iron- and oxygen-sensitive Fd at a later stage, after oxygen build-up. The widespread presence of Fld in anaerobes and the existence of Fld-specific metabolic routes (Freigang et al. 2002; Puan et al. 2005) provide circumstantial evidence to this hypothesis.
Flds are classified as short-chain or long-chain depending on the presence of a 20-amino acid loop of unknown function, with the photosynthetic Flds belonging to the latter class (Sancho 2006). A complete Fld tree has been recently reported (Pierella Karlusich et al. 2015). It shows that long-chain Flds form a monophyletic group distributed between photosynthetic eukaryotes and various bacterial phyla besides cyanobacteria (Pierella Karlusich et al. 2015). The flavoproteins present in phototrophs are themselves divided in three different clades. The algal and α-cyanobacterial sequences assemble into a single clade, whereas the Flds from β-cyanobacteria are distributed in the two other clades (Pierella Karlusich et al. 2015).
The most remarkable observation derived from the phylogenetic analysis was that the eukaryotic Flds are basal with respect to their α-cyanobacterial counterparts in the algal/α-cyanobacterial clade (Pierella Karlusich et al. 2015). This topology differs from those inferred from other genetic markers such as 16S rRNA, PsbO or PsbA, which show all cyanobacterial sequences grouping in a basal position with respect to the eukaryotic clades, in agreement with the existence of the primary endosymbiotic event (Keeling 2013). The conflicting topology involving the α-cyanobacterial position in the Fld tree with respect to the cyanobacterial-plastid tree might indicate an HGT event from an eukaryotic donor to an α-cyanobacterium (Pierella Karlusich et al. 2015).
The phylogenetic distribution analysis indicates that retention of the Fld-coding gene along the path of evolution has been disparate, showing a clear bias towards iron-deficient habitats such as open oceans (Erdner and Anderson 1999; see also Fig. 2). This situation occurs even among species of the same genera, such as the closely related chlorophytes Bathycoccus and Ostreococcus (Pierella Karlusich et al. 2015). In this context, HGT from an eukaryote to an α-cyanobacterium seems to be related to ecological adaptations to iron bioavailability. The α-cyanobacterial lineage has a freshwater origin, and early diverging α-cyanobacteria, which are from high-iron environments, do not contain Fld (Pierella Karlusich et al. 2015). When they colonized marine environments, the reacquisition of Fld from an eukaryote might have helped this lineage to tolerate iron deficiencies typical of the open oceans (Pierella Karlusich et al. 2015).

(adapted from Pierella Karlusich et al. 2015)
Phylogenetic distribution of Fd and Fld in cyanobacteria and photosynthetic eukaryotes and its relationship with the iron bioavailability in their habitats. The phylogenetic trees of cyanobacteria and photosynthetic eukaryotes are based on the 16S and 18S rRNA sequences, respectively. For each species, the presence (green circles) or absence of Fd- and Fld-coding genes, as well as the iron levels at the site of isolation (red bars), are shown. Lengths of the red bars provide an estimation of iron level as high, intermediate, or low
The loss of the Fld gene from the plant genome might also be related to ecological adaptations to iron bioavailability and the successive stages of land colonization (Pierella Karlusich et al. 2014). The phylogenetic distribution of the Fld-coding gene implies that it has been most likely lost long before the origin of plants, as it has not been found in charophytes, a group of freshwater and terrestrial algae that includes the sister lineage of land plants (Leliaert et al. 2011). It is likely that the Fld-coding gene was no longer required in an environment in which iron was both abundant and readily accessible (Pierella Karlusich et al. 2015). Under such conditions, selection pressures for Fld retention as an adaptive trait to replace iron-demanding Fd might have been relaxed.
The collected results indicate that the Fld-coding gene has a complex evolutionary dynamics strongly associated with the geochemistry of iron. This behavior contrasts with that of Fd-coding genes, which have been found in all described algae and cyanobacteria, even in those thriving in iron-limited environments (Fig. 2).
Evolution of the transducer, ferredoxin-NADP+ reductase
FNRs are FAD-containing enzymes that mediate the reversible electron transfer between two molecules of Fd or Fld and one molecule of NADP(H) (Aliverti et al. 2008). While Fd/Fld can exchange reducing equivalents with many different enzymes (Hase et al. 2006; Hanke and Mulo 2013), FNR is certainly the main partner in photosynthetic tissues. This reductase behaves as a general electronic switch that connects the universe of obligatory two-electron carriers (i.e., pyridine nucleotides) with that of obligatory one-electron shuttles such as transition metals, taking advantage of the relatively small differences existing in the redox potentials for the one- and two-electron transfer processes of its FAD cofactor (Carrillo and Ceccarelli 2003). Considering their strategic position as electronic hubs, it is not surprising that FNRs are found in all kinds of organisms displaying very different lifestyles. Noteworthy, not all flavoenzymes exhibiting FNR activity derive from a common ancestor. Indeed, at least three different classes have been defined based on their basic 3D-conformation. The enzymes present in Gram-positive bacteria belong to the NADPH-dependent thioredoxin reductase family of flavoproteins (Seo et al. 2016a, b), and are termed the TR-class FNRs. They are also found in archaea and some Gram-negative prokaryotes including green sulfur bacteria and proteobacteria. The two other classes are structurally related to the disulfide reductases, whose prototype is glutathione reductase (Ziegler and Schultz 2000), and to the flavin reductases (Bruns and Karplus 1995), constituting the DR and FR classes, respectively. Then, in a remarkable case of convergent evolution, nature made use of different flavoproteins as bauplan to develop the desired function, underscoring the relevance of this reductase for electron transfer under a wide range of physiological conditions. Most organisms contain FNRs of a single class (FR in cyanobacteria), but plants also have a reductase from the DR class in mitochondria, presumably derived from the α-proteobacterial endosymbiont.
The FR-class FNRs of chloroplasts and cyanobacteria are made up of two structural domains, with the N-terminal region binding FAD and the C-terminal region NADP(H) in a typical Rossman-fold conformation (Bruns and Karplus 1995; Aliverti et al. 2008). Docking of Fd and Fld takes place at the cleavage between the two domains (Martínez-Júlvez et al. 1999). Despite their disparate origin, FR- and DR- class FNRs can be functionally exchanged in vitro (Faro et al. 2003; Zöllner et al. 2004). The isoforms present in non-photosynthetic plastids are most likely the product of a duplication event with minimal functional changes relative to their chloroplast counterparts (Ceccarelli et al. 2004).
Interestingly, some cyanobacterial FNRs harbor an extra domain of ~10 kDa at their N-termini, which displays significant sequence similarity with the 9-kDa CpcD rod-capping polypeptide (Schluchter and Bryant 1992) and the C-terminal region of CpcC (Six et al. 2005), both components of the phycobilisomes (PBS), the light-harvesting antenna complexes in most cyanobacteria (Watanabe and Ikeuchi 2013). Indeed, this longer 3-domain version of FNR has only been found in cyanobacteria containing PBS. Conversely, not all PBS-containing species have 3-domain FNRs (Fig. 3). The CpcD-like domain is responsible for attachment of this FNR to the peripheral rods of PBS (Morsy et al. 2008; Arteni et al. 2009; Korn et al. 2009). Phylogenetic analysis shows that basal cyanobacteria [Gloeobacter, Synechococcus sp. JA-3-3Ab, Synechococcus sp. JA-2-3B’a (2–13), Pseudanabaena] only have two-domain FNRs although they do contain PBS (Fig. 3), suggesting that the CpcD-like domain was acquired at a later stage, probably after transfer to the first photosynthetic eukaryote through the primary endosymbiotic event, since both glaucophytes and red algae have PBS and two-domain FNRs (Fig. 3). In green algae and plants, as well as in secondary-plastid-bearing algae, PBS were replaced by light-harvesting complexes (LHC, Fig. 3). Cryptophytes have phycobiliproteins but they are not organized as PBS (Cheregi et al. 2015).
Phylogenetic distribution of PBS and two- or three-domain FNRs in cyanobacteria and photosynthetic eukaryotes. The data suggest that the cyanobacterium that gave origin to modern plastids already lacked the CpcD-domain in its FNR. The fact that the identity of the cyanobacterial lineage that gave rise to primary plastids is still controversial is indicated by a dashed line
The results therefore suggest that the gene coding for the two-domain FNR was fused to CpcD-like sequences in a cyanobacterial lineage with PBS, giving rise to the three-domain FNR gene, which then spread to other cyanobacteria by vertical transfer. The CpcD-like domain was independently lost in four cyanobacterial lineages that also lost PBS (β-cyanobacteria Acaryochloris sp. CCMEE 5410, Prochloron and Prochlorothrix, and the terminal α-cyanobacteria Prochlorococcus), but was maintained in all other lineages keeping PBS (Fig. 3). In Prochlorococcus, the N-terminal domain of FNR corresponds to a transmembrane α-helix, while the basal α-cyanobacteria (Cyanobium and Synechococcus) have three-domain FNRs and PBS.
Some cyanobacteria have genes encoding for a three-domain FNR but they can produce the two-domain version by using an alternative promoter and an internal Met codon (Thomas et al. 2006; Omairi-Nasser et al. 2011). Interestingly, PBS-containing cyanobacteria that are able to produce both FNRs (Synechocystis sp. PCC6803, Synechococcus sp. PCC7002, Anabaena sp. PCC7120) are facultative heterotrophs, whereas obligate phototrophs (Synechococcus elongates, Thermosynechococcus elongatus) lack the internal Met and only express the three-domain FNR (Thomas et al. 2006). Morsy et al. (2008) have studied the subcellular localization of FNR in the cyanobacterium Spirulina platensis and the red alga Cyanidium caldarium. Although both microorganisms are photoautotrophs bearing PBS as light-harvesting complexes, FNR localization and membrane-binding characteristics were different. Spirulina expressed a three-domain reductase which was distributed between PBS and thylakoid membranes, while the FNR from Cyanidium was a two-domain protein tightly bound to thylakoids, without any detectable interaction with PBS (Morsy et al. 2008).
Removal of the CpcD-like domain results in solubilization of an active two-domain FNR from PBS, and there is ample evidence that soluble FNR favors electron exchange in the direction of NADPH oxidation in cyanobacteria (Korn et al. 2009). This might represent a labor division in those cases in which there exists the possibility of a dynamic exchange between membrane and solution. It has been argued that the three-domain PBS-associated FNR could participate in NADP+ photoreduction, and the soluble one in cyclic electron transport and heterotrophic redox pathways (Korn et al. 2009). Indeed, election between the “long” and “short” versions of the flavoprotein might respond to these demands. In Synechocystis sp. PCC6803, a fresh-water non-diazotrophic cyanobacterium, expression of the two-domain FNR is induced by heterotrophic growth conditions, and by nitrogen and iron deficiency, whereas accumulation of the three-domain isoform was not affected (Thomas et al. 2006). In diazotrophic Anabaena sp. PCC7120, expression of the two- and three-domain FNRs was perfectly split between heterocysts and vegetative cells, respectively (Omairi-Nasser et al. 2014).
Then, FNR evolved from a soluble enzyme in heterotrophic bacteria to a membrane-bound reductase in phototrophs, using different mechanisms of membrane association. In plants, two-domain FNR molecules are dynamically exchanged among three different chloroplast compartments: the thylakoid membrane, the stroma, and the inner envelope membrane (Hanke et al. 2005; Benz et al. 2009). Two Arabidopsis chloroplast proteins, At-TROL and At-TIC62, which possess conserved proline-rich FNR-binding motif(s) in their C-termini, have been shown to mediate anchoring of FNR to the thylakoid membrane (Benz et al. 2009; Juric et al. 2009; Alte et al. 2010; Lintala et al. 2014). FNR assembles into thylakoid protein complexes of approximately 500 and 190 kDa with At-TIC62 and At-TROL, respectively (Benz et al. 2009; Juric et al. 2009). The physiological roles of the soluble and membrane-bound pools of FNR have not been clarified. Several observations suggested that the rate of NADP+ photoreduction was higher in the thylakoid-bound complex than in the soluble form, as in cyanobacteria (Forti and Bracale 1984; Rodríguez et al. 2007). Nevertheless, as plant performance was not markedly affected in Arabidopsis fnr1 and tic62 trol mutants lacking FNR associated to thylakoid membranes, the soluble reductase pool also appears to be photosynthetically competent (Lintala et al. 2007, 2014), so that the importance of membrane attachment remains an open question. Yang et al. (2016) identified a rice protein, LIR1, as an FNR-interacting partner that modulates binding of the flavoenzyme to the membrane anchor. Rice lir1 knock-out plants had slightly impaired photosynthetic capacity, whereas no such effect was observed in the analogous Arabidopsis mutants (Yang et al. 2016). Distribution of FNR in thylakoids, envelope, and stroma also depends on the nature of the isoform involved (Twachtmann et al. 2012).
Further adaptations to oxygenic photosynthesis: reversion of electron transfer direction and increased catalytic efficiency of FNR
As already indicated, enzymes with FNR activity are widespread, and were certainly present in anaerobes prior to the evolution of oxygenic photosynthesis. In heterotrophic microorganisms, mitochondria, and non-photosynthetic plastids, the direction of electron transfer is from NADPH to oxidized Fd or Fld, and the main function of the reductase was to provide these low-potential promiscuous carriers to electron-consuming reactions (Carrillo and Ceccarelli 2003). In chloroplasts and cyanobacteria, instead, electron flow proceeds backwards. While the overall electron exchange process remains fully reversible, the change in direction was accompanied by modifications in the midpoint redox potential of FNR and/or Fd/Fld that favored NADP+ reduction.
Besides reversing the direction of preferred electron transport, photosynthetic FNRs experienced a formidable increase in catalytic competence relative to their heterotrophic counterparts, with turnover numbers of 200–500 s−1 for the reductases of cyanobacteria and plants, and less than 10 s−1 for the FNRs present in most bacterial species (Carrillo and Ceccarelli 2003; Ceccarelli et al. 2004; see however; Catalano Dupuy et al. 2011). This improvement is entirely accounted for by a drastic increase in the k cat values, while the K M for NADP(H), Fd, and Fld remain in the low micromolar range for all reductases (Carrillo and Ceccarelli 2003). Optimization of FNR catalytic efficiency might be related to the demands of the photosynthetic process that requires a very fast electron flow to sustain CO2 fixation rates. In organisms growing on heterotrophic metabolisms or anoxygenic photosynthesis, FNR is involved in pathways that proceed at a much slower pace (McLean et al. 2003; Bortolotti et al. 2009; Yeom et al. 2009; Seo et al. 2016a, b). Then, the maximal rates that each reductase can attain correlate fairly well with the demands of the pertinent metabolic pathway of which it is a committed member. Exceptions are the FNR isoforms present in plastids from non-photosynthetic plant tissues (such as root amyloplasts) that display high specific activities in the context of a heterotrophic metabolism whose partner enzymes (i.e., nitrite reductase) turn over at 10–20 s−1 (Hirasawa et al. 1993). It should be kept in mind, however, that these flavoenzymes evolved from photosynthetic ancestors present in preexisting chloroplasts (the primary descendants of the cyanobacterial endosymbiont) in relatively recent times. The advent of terrestrial plants with diversified tissues and organs gave origin to the non-photosynthetic plastids of roots, fruits, petals, etc., all derived from chloroplasts and harboring the same reduced genome (Waters and Pyke 2005). Most plastid-targeted photosynthetic components ceased to be expressed in these tissues but a few, including FNR, were recruited to play new functions and acquired novel regulatory mechanisms of expression (Hanke et al. 2005; Hachiya et al. 2016). Then, the high activity profile of amyloplast FNR, for instance, is likely a remnant of its recent chloroplastic past.
An additional consequence of the increased activity of photosynthetic FNR was faster reoxidation of Fd for reaction with reduced F A/F B. Together with the redox potential inversion described before, this FNR modification improved Fd competition with oxygen for the reducing equivalents generated at PSI, further contributing to the protection of the [4Fe–4S] di-cluster from oxidative denaturation.
Concluding remarks
The reducing side of PSI is made up of a group of electron transfer proteins containing iron-sulfur clusters and flavins as prosthetic groups. All these carriers and enzymes underwent profound and concerted evolutionary changes along the path that led from anoxygenic to oxygenic photosynthesis. Structural comparisons and phylogenetic analyses reveal that modifications were directed to protect themselves against inactivation by oxygen and its reactive derivatives generated by partial reduction at PSI and energy transfer at PSII. Changes introduced included shielding from solvent of the [4S–4Fe] clusters of F A/F B and the [2S–2Fe] center of Fd, transient recruitment of iron-free Fld, and significant increases in the catalytic turnover of FNR. The resulting electron transfer system was able to operate at high rates, compatible with the demands of CO2 fixation, and minimal electron leakage to oxygen, even in the super-saturated environment of oxygenic photosynthesis.
References
Aliverti A, Pandini V, Pennati A, de Rosa M, Zanetti G (2008) Structural and functional diversity of ferredoxin-NADP+ reductases. Arch Biochem Biophys 474:283–291
Allen JF, Martin W (2007) Evolutionary biology: out of thin air. Nature 445:610–612. doi:10.1038/445610a
Alte F, Stengel A, Benz JP, Petersen E, Soll J, Groll M, Bolter B (2010) Ferredoxin:NADPH oxidoreductase is recruited to thylakoids by binding to a polyproline type II helix in a pH-dependent manner. Proc Natl Acad Sci USA 107:19260–19265. doi:10.1073/pnas.1009124107
Anbar AD (2008) Oceans. Elements and evolution. Science 322:1481–1483. doi:10.1126/science.1163100
Antonkine ML, Jordan P, Fromme P, Krauss N, Golbeck JH, Stehlik D (2003) Assembly of protein subunits within the stromal ridge of photosystem I. Structural changes between unbound and sequentially PS I-bound polypeptides and correlated changes of the magnetic properties of the terminal iron sulfur clusters. J Mol Biol 327:671–697
Armengaud J, Meyer C, Jouanneau Y (1994) Recombinant expression of the fdxD gene of Rhodobacter capsulatus and characterization of its product, a [2Fe–2S] ferredoxin. Biochem J 300:413–418
Arteni AA, Ajlani G, Boekema EJ (2009) Structural organisation of phycobilisomes from Synechocystis sp. strain PCC6803 and their interaction with the membrane. Biochim Biophys Acta 1787:272–279. doi:10.1016/j.bbabio.2009.01.009
Badger MR, von Caemmerer S, Ruuska S, Nakano H (2000) Electron flow to oxygen in higher plants and algae: rates and control of direct photoreduction (Mehler reaction) and rubisco oxygenase. Philos Trans R Soc Lond B Biol Sci 355:1433–1446. doi:10.1098/rstb.2000.0704
Benz JP, Stengel A, Lintala M, Lee YH, Weber A, Philippar K, Gugel IL, Kaieda S, Ikegami T, Mulo P, Soll J, Bolter B (2009) Arabidopsis Tic62 and ferredoxin-NADP(H) oxidoreductase form light-regulated complexes that are integrated into the chloroplast redox poise. Plant Cell 21:3965–3983. doi:10.1105/tpc.109.069815
Blanco NE, Ceccoli RD, Segretin ME, Poli HO, Voss I, Melzer M, Bravo-Almonacid FF, Scheibe R, Hajirezaei MR, Carrillo N (2011) Cyanobacterial flavodoxin complements ferredoxin deficiency in knocked-down transgenic tobacco plants. Plant J 65:922–935. doi:10.1111/j.1365-313X.2010.04479.x
Blankenship RE (2001) Molecular evidence for the evolution of photosynthesis. Trends Plant Sci 6:4–6
Bortolotti A, Pérez-Dorado I, Goñi G, Medina M, Hermoso JA, Carrillo N, Cortez N (2009) Coenzyme binding and hydride transfer in Rhodobacter capsulatus ferredoxin/flavodoxin NADP(H) oxidoreductase. Biochim Biophys Acta 1794:199–210. doi:10.1016/j.bbapap.2008.09.013
Bott AW (1999) Redox properties of electron transfer metalloproteins. Curr Sep 18:47–54
Bruns CM, Karplus AP (1995) Refined crystal structure of spinach ferredoxin reductase at 1.7 Å resolution: oxidized, reduced and 2′-phospho-5′-AMP bound states. J Mol Biol 247:125–145. doi:10.1006/jmbi.1994.0127
Caetano-Anollés G, Kim HS, Mittenthal JE (2007) The origin of modern metabolic networks inferred from phylogenomic analysis of protein architecture. Proc Natl Acad Sci USA 104:9358–9363. doi:10.1073/pnas.0701214104
Cardona T (2015) A fresh look at the evolution and diversification of photochemical reaction centers. Photosynth Res 126:111–134. doi:10.1007/s11120-014-0065-x
Carrillo N, Ceccarelli EA (2003) Open questions in ferredoxin-NADP+ reductase catalytic mechanism. Eur J Biochem 270:1900–1915. doi:10.1046/j.1432-1033.2003.03566.x
Catalano Dupuy DL, Musumeci MA, López-Rivero A, Ceccarelli EA (2011) A highly stable plastidic-type ferredoxin-NADP(H) reductase in the pathogenic bacterium Leptospira interrogans. PLoS One 6:e26736. doi:10.1371/journal.pone.0026736
Ceccarelli EA, Arakaki AK, Cortez N, Carrillo N (2004) Functional plasticity and catalytic efficiency in plant and bacterial ferredoxin-NADP(H) reductases. Biochim Biophys Acta 1698:155–165. doi:10.1016/j.bbapap.2003.12.005
Ceccoli RD, Blanco NE, Medina M, Carrillo N (2011) Stress response of transgenic tobacco plants expressing a cyanobacterial ferredoxin in chloroplasts. Plant Mol Biol 76:535–544. doi:10.1007/s11103-011-9786-9
Cheng Y-Q, Liu Z-M, Xu J, Zhou T, Wang M, Chen Y-T, Li H-F, Fan Z-F (2008) HC-Pro protein of sugar cane mosaic virus interacts specifically with maize ferredoxin-5 in vitro and in planta. J Gen Virol 89:2046–2054. doi:10.1099/vir.0.2008/001271-0
Cheregi O, Kotabova E, Prasil O, Schroder WP, Kana R, Funk C (2015) Presence of state transitions in the cryptophyte alga Guillardia theta. J Exp Bot 66:6461–6470. doi:10.1093/jxb/erv362
Dickey LF, Gallo-Meagher M, Thompson W (1992) Light regulatory sequences are located within the 5′ portion of the Fed-1 message sequence. EMBO J 11:2311–2317
Eck RV, Dayhoff MO (1966) Evolution of the structure of ferredoxin based on living relics of primitive amino acid sequences. Science 152:363–366. doi:10.1126/science.152.3720.363
Erdner DL, Anderson DM (1999) Ferredoxin and flavodoxin as biochemical indicators of iron limitation during open-ocean iron enrichment. Limnol Oceanogr 44:1609–1615. doi:10.4319/lo.1999.44.7.1609
Ewen KM, Hannemann F, Iametti S, Morleo A, Bernhardt R (2011) Functional characterization of Fdx1: evidence for an evolutionary relationship between P450-type and ISC-type ferredoxins. J Mol Biol 413:940–951. doi:10.1016/j.jmb.2011.09.010
Faro M, Schiffler B, Heinz A, Nogués I, Medina M, Bernhardt R, Gómez-Moreno C (2003) Insights into the design of a hybrid system between Anabaena ferredoxin-NADP+ reductase and bovine adrenodoxin. Eur J Biochem 270:726–735
Forti G, Bracale M (1984) Ferredoxin-ferredoxin NADP+ reductase interaction: catalytic differences between the soluble and thylakoid-bound complex. FEBS Lett 166:81–84. doi:10.1016/0014-5793(84)80049-6
Freigang J, Diederichs K, Schäfer KP, Welte W, Paul R (2002) Crystal structure of oxidized flavodoxin, an essential protein in Helicobacter pylori. Protein Sci 11:253–261. doi:10.1110/ps.28602
Frolow F, Harel M, Sussman JL, Mevarech M, Shoham M (1996) Insights into protein adaptation to a saturated salt environment from the crystal structure of a halophilic 2Fe–2S ferredoxin. Nat Struct Biol 3:452–458
Fromme P, Jordan P, Krauss N (2001) Structure of photosystem I. Biochim Biophys Acta 1507:5–31
García Costas AM, Liu Z, Tomsho LP, Schuster SC, Ward DM, Bryant DA (2012) Complete genome of Candidatus Chloracidobacterium thermophilum, a chlorophyll-based photoheterotroph belonging to the phylum acidobacteria. Environ Microbiol 14:177–190. doi:10.1111/j.1462-2920.2011.02592.x
Gou P, Hanke GT, Kimata-Ariga Y, Standley DM, Kubo A, Taniguchi I, Nakamura H, Hase T (2006) Higher order structure contributes to specific differences in redox potential and electron transfer efficiency of root and leaf ferredoxins. BioChemistry 45:14389–14396. doi:10.1021/bi061779d
Grinter R, Milner J, Walker D (2012) Ferredoxin-containing bacteriocins suggest a novel mechanism of iron uptake in Pectobacterium spp. PLoS One 7:e33033. doi:10.1371/journal.pone.0033033
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. doi:10.1093/sysbio/syq010
Hachiya T, Ueda N, Kitagawa M, Hanke GT, Suzuki A, Hase T, Sakakibara H (2016) Arabidopsis root-type ferredoxin:NADP(H) oxidoreductase 2 is involved in detoxification of nitrite in roots. Plant Cell Physiol 57:2440–2450. doi:10.1093/pcp/pcw158
Hanke GT, Hase T (2008) Variable photosynthetic roles of two leaf-type ferredoxins in Arabidopsis, as revealed by RNA interference. Photochem Photobiol 84:1302–1309. doi:10.1111/j.1751-1097.2008.00411.x
Hanke G, Mulo P (2013) Plant type ferredoxins and ferredoxin-dependent metabolism. Plant Cell Environ 36:1071–1084. doi:10.1111/pce.12046
Hanke GT, Kimata-Ariga Y, Taniguchi I, Hase T (2004) A post genomic characterization of Arabidopsis ferredoxins. Plant Physiol 134:255–264. doi:10.1104/pp.103.032755
Hanke GT, Okutani S, Satomi Y, Takao T, Suzuki A, Hase T (2005) Multiple iso-proteins of FNR in Arabidopsis: evidence for different contributions to chloroplast function and nitrogen assimilation. Plant Cell Environ 28:1146–1157
Harel A, Bromberg Y, Falkowski PG, Bhattacharya D (2014) Evolutionary history of redox metal-binding domains across the tree of life. Proc Natl Acad Sci USA 111:7042–7047. doi:10.1073/pnas.1403676111
Hase T, Schürmann P, Knaff DB (2006) The interaction of ferredoxin with ferredoxin-dependent enzymes. In: Golbeck JH (ed) Photosystem I: the light-driven plastocyanin:ferredoxin oxidoreductase. Springer, Dordrecht, pp 477–498
Hirasawa M, Jappe H, Knaff DB (1993) The effect of lysine- and arginine-modifying reagents on spinach ferredoxin:nitrite oxidoreductase. Biochim Biophys Acta 1140:304–312
Hohmann-Marriott MF, Blankenship RE (2011) Evolution of photosynthesis. Annu Rev Plant Biol 62:515–548. doi:10.1146/annurev-arplant-042110-103811
Hruz T, Laule O, Szabo G, Wessendorp F, Bleuler S, Oertle L, Widmayer P, Gruissem W, Zimmermann P (2008) Genevestigator V3: a reference expression database for the meta-analysis of transcriptomes. Adv Bioinform 420747. doi:10.1155/2008/420747
Imlay JA (2006) Iron-sulphur clusters and the problem with oxygen. Mol Microbiol 59:1073–1082. doi:10.1111/j.1365-2958.2006.05028.x
Jagannathan B, Golbeck JH (2008) Unifying principles in homodimeric type I photosynthetic reaction centers: properties of PscB and the F A , F B and F X iron-sulfur clusters in green sulfur bacteria. Biochim Biophys Acta 1777:1535–1544. doi:10.1016/j.bbabio.2008.09.001
Jagannathan B, Shen GZ, Golbeck JH (2012) The evolution of Type I reaction centers: the response to oxygenic photosynthesis. In: Burnap RL, Vermaas W (eds) Functional genomics and evolution of photosynthetic systems, vol 33, advances in photosynthesis and respiration. Springer, Dordrecht, pp 285–316
Juric S, Hazler-Pilepic K, Tomasic A, Lepedus H, Jelicic B, Puthiyaveetil S, Bionda T, Vojta L, Allen JF, Schleiff E, Fulgosi H (2009) Tethering of ferredoxin:NADP+ oxidoreductase to thylakoid membranes is mediated by novel chloroplast protein TROL. Plant J 60:783–794. doi:10.1111/j.1365-313X.2009.03999.x
Katoh K, Toh H (2008) Improved accuracy of multiple ncRNA alignment by incorporating structural information into a MAFFT-based framework. BMC Bioinform 9:212. doi:10.1186/1471-2105-9-212
Keeling PJ (2013) The number, speed, and impact of plastid endosymbioses in eukaryotic evolution. Annu Rev Plant Biol 64:583–607. doi:10.1146/annurev-arplant-050312-120144
Kerr RA (2005) Earth science. The story of O2. Science 308:1730–1732. doi:10.1126/science.308.5729.1730
Kirschvink JL, Kopp RE (2008) Palaeoproterozoic ice houses and the evolution of oxygen-mediating enzymes: the case for a late origin of photosystem II. Philos Trans R Soc Lond B Biol Sci 363:2755–2765. doi:10.1098/rstb.2008.0024
Korn A, Ajlani G, Lagoutte B, Gall A, Setif P (2009) Ferredoxin:NADP+ oxidoreductase association with phycocyanin modulates its properties. J Biol Chem 284:31789–31797. doi:10.1074/jbc.M109.024638
Kozuleva MA, Ivanov BN (2016) The mechanisms of oxygen reduction in the terminal reducing segment of the chloroplast photosynthetic electron transport chain. Plant Cell Physiol 57:1397–1404. doi:10.1093/pcp/pcw035
Kutzki C, Masepohl B, Böhme H (1998) The isiB gene encoding flavodoxin is not essential for photoautotrophic iron limited growth of the cyanobacterium Synechocystis sp. strain PCC 6803. FEMS Microbiol Lett 160:231–235. doi:10.1111/j.1574-6968.1998.tb12916.x
La Roche J, Boyd PW, McKay RML, Geider RJ (1996) Flavodoxin as an in situ marker for iron stress in phytoplankton. Nature 382:802–805. doi:10.1038/382802a0
Leliaert F, Verbruggen H, Zechman FW (2011) Into the deep: new discoveries at the base of the green plant phylogeny. Bioessays 33:683–692. doi:10.1002/bies.201100035
Lemaire SD, Stein M, Issakidis-Bourguet E, Keryer E, Benoit V, Pineau B, Gérard-Hirne C, Miginiac-Maslow M, Jacquot J-P (1999) The complex regulation of ferredoxin/thioredoxin-related genes by light and the circadian clock. Planta 209:221–229. doi:10.1007/s004250050626
Li C, Hu Y, Huang R, Ma X, Wang Y, Liao T, Zhong P, Xiao F, Sun C, Xu Z, Deng X, Wang P (2015) Mutation of FdC2 gene encoding a ferredoxin-like protein with C-terminal extension causes yellow-green leaf phenotype in rice. Plant Sci 238:127–134. doi:10.1016/j.plantsci.2015.06.010
Lindell D, Sullivan MB, Johnson ZI, Tolonen AC, Rohwer F, Chisholm SW (2004) Transfer of photosynthesis genes to and from Prochlorococcus viruses. Proc Natl Acad Sci USA 101:11013–11018. doi:10.1073/pnas.0401526101
Lintala M, Allahverdiyeva Y, Kidron H, Piippo M, Battchikova N, Suorsa M, Rintamaki E, Salminen TA, Aro EM, Mulo P (2007) Structural and functional characterization of ferredoxin-NADP+ oxidoreductase using knock-out mutants of Arabidopsis. Plant J 49:1041–1052. doi:10.1111/j.1365-313X.2006.03014.x
Lintala M, Schuck N, Thormahlen I, Jungfer A, Weber KL, Weber AP, Geigenberger P, Soll J, Bolter B, Mulo P (2014) Arabidopsis tic62 trol mutant lacking thylakoid-bound ferredoxin-NADP+ oxidoreductase shows distinct metabolic phenotype. Mol Plant 7:45–57. doi:10.1093/mp/sst129
Martin W, Russell MJ (2003) On the origins of cells: a hypothesis for the evolutionary transitions from abiotic geochemistry to chemoautotrophic prokaryotes, and from prokaryotes to nucleated cells. Philos Trans R Soc Lond B Biol Sci 358:59–83. doi:10.1098/rstb.2002.1183 (discussion 83–55)
Martínez-Júlvez M, Medina M, Gómez-Moreno C (1999) Ferredoxin-NADP+ reductase uses the same site for the interaction with ferredoxin and flavodoxin. J Biol Inorg Chem 4:568–578
Masepohl B, Scholisch K, Gorlitz K, Kutzki C, Böhme H (1997) The heterocyst-specific fdxH gene product of the cyanobacterium Anabaena sp. PCC 7120 is important but not essential for nitrogen fixation. Mol Gen Genet 253:770–776
Matsumura T, Sakakibara H, Nakano R, Kimata Y, Sugiyama T, Hase T (1997) A nitrate-inducible ferredoxin in maize roots. Genomic organization and differential expression of two nonphotosynthetic ferredoxin isoproteins. Plant Physiol 114:653–660. doi:10.1104/pp.114.2.653
Mazouni K, Domain F, Chauvat F, Cassier-Chauvat C (2003) Expression and regulation of the crucial plant-like ferredoxin of cyanobacteria. Mol Microbiol 49:1019–1029. doi:10.1046/j.1365-2958.2003.03609.x
McLean KJ, Scrutton NS, Munro AW (2003) Kinetic, spectroscopic and thermodynamic characterization of the Mycobacterium tuberculosis adrenodoxin reductase homologue FprA. Biochem J 372:317–327. doi:10.1042/BJ20021692
Medina M, Martínez-Júlvez M, Hurley JK, Tollin G, Gómez-Moreno C (1998) Involvement of glutamic acid 301 in the catalytic mechanism of ferredoxin-NADP+ reductase from Anabaena PCC 7119. BioChemistry 37:2715–2728. doi:10.1021/bi971795y
Meyer J (2008) Iron-sulfur protein folds, iron-sulfur chemistry, and evolution. J Biol Inorg Chem 13:157–170. doi:10.1007/s00775-007-0318-7
Meyer J, Clay MD, Johnson MK, Stubna A, Munck E, Higgins C, Wittung-Stafshede P (2002) A hyperthermophilic plant-type [2Fe–2S] ferredoxin from Aquifex aeolicus is stabilized by a disulfide bond. BioChemistry 41:3096–3108
Møller IM, Jensen PE, Hansson A (2007) Oxidative modifications to cellular components in plants. Annu Rev Plant Biol 58:459–481. doi:10.1146/annurev.arplant.58.032806.103946
Morsy FM, Nakajima M, Yoshida T, Fujiwara T, Sakamoto T, Wada K (2008) Subcellular localization of ferredoxin-NADP+ oxidoreductase in phycobilisome retaining oxygenic photosysnthetic organisms. Photosynth Res 95:73–85. doi:10.1007/s11120-007-9235-4
Mulkidjanian AY, Junge W (1997) On the origin of photosynthesis as inferred from sequence analysis. Photosynth Res 51:27–42
Mulkidjanian AY, Koonin EV, Makarova KS, Mekhedov SL, Sorokin A, Wolf YI, Dufresne A, Partensky F, Burd H, Kaznadzey D, Haselkorn R, Galperin MY (2006) The cyanobacterial genome core and the origin of photosynthesis. Proc Natl Acad Sci USA 103:13126–13131. doi:10.1073/pnas.0605709103
Nogués I, Pérez-Dorado I, Frago S, Bittel C, Mayhew SG, Gómez-Moreno C, Hermoso JA, Medina M, Cortez N, Carrillo N (2005) The ferredoxin–NADP(H) reductase from Rhodobacter capsulatus: molecular structure and catalytic mechanism. BioChemistry 44:11730–11740. doi:10.1021/bi0508183
Oh-Oka H (2007) Type 1 reaction center of photosynthetic heliobacteria. Photochem Photobiol 83:177–186. doi:10.1562/2006-03-29-IR-860
Omairi-Nasser A, de Gracia AG, Ajlani G (2011) A larger transcript is required for the synthesis of the smaller isoform of ferredoxin:NADP oxidoreductase. Mol Microbiol 81:1178–1189. doi:10.1111/j.1365-2958.2011.07739.x
Omairi-Nasser A, Galmozzi CV, Latifi A, Muro-Pastor MI, Ajlani G (2014) NtcA is responsible for accumulation of the small isoform of ferredoxin:NADP+ oxidoreductase. Microbiology 160:789–794. doi:10.1099/mic.0.076042-0
Patterson K, Cakmak T, Cooper A, Lager I, Rasmusson AG, Escobar MA (2010) Distinct signalling pathways and transcriptome response signatures differentiate ammonium-and nitrate-supplied plants. Plant Cell Environ 33:1486–1501. doi:10.1111/j.1365-3040.2010.02158.x
Peden EA, Boehm M, Mulder DW, Davis R, Old WM, King PW, Ghirardi ML, Dubini A (2013) Identification of global ferredoxin interaction networks in Chlamydomonas reinhardtii. J Biol Chem 288:35192–35209. doi:10.1074/jbc.M113.483727
Pierella Karlusich JJ, Lodeyro AF, Carrillo N (2014) The long goodbye: the rise and fall of flavodoxin during plant evolution. J Exp Bot 65:5161–5178. doi:10.1093/jxb/eru273
Pierella Karlusich JJ, Ceccoli RD, Graña M, Romero H, Carrillo N (2015) Environmental selection pressures related to iron utilization are involved in the loss of the flavodoxin gene from the plant genome. Genome Biol Evol 7:750–767. doi:10.1093/gbe/evv031
Poncelet M, Cassier-Chauvat C, Leschelle X, Bottin H, Chauvat F (1998) Targeted deletion and mutational analysis of the essential (2Fe–2S) plant-like ferredoxin in Synechocystis PCC6803 by plasmid shuffling. Mol Microbiol 28:813–821. doi:10.1046/j.1365-2958.1998.00844.x
Puan KJ, Wang H, Dairi T, Kuzuyama T, Morita CT (2005) fldA is an essential gene required in the 2-C-methyl-d-erythritol 4-phosphate pathway for isoprenoid biosynthesis. FEBS Lett 579:3802–3806. doi:10.1016/j.febslet.2005.05.047
Que L Jr, Anglin JR, Bobrik MA, Davison A, Holm RH (1974) Synthetic analogs of the active sites of iron-sulfur proteins. IX. Formation and some electronic and reactivity properties of Fe4S4 Glycyl-l-cysteinylglycyl oligopeptide complexes obtained by ligand substitution reactions. J Am Chem Soc 96:6042–6048
Raymond J, Blankenship RE (2006) Evolutionary relationships among type I photosynthetic reaction centers. In: Golbeck JH (ed) Photosystem I: the light-driven plastocyanin:ferredoxin oxidoreductase. Springer, Dordrecht, pp 669–681
Rodríguez RE, Lodeyro A, Poli HO, Zurbriggen M, Peisker M, Palatnik JF, Tognetti VB, Tschiersch H, Hajirezaei MR, Valle EM, Carrillo N (2007) Transgenic tobacco plants overexpressing chloroplastic ferredoxin-NADP(H) reductase display normal rates of photosynthesis and increased tolerance to oxidative stress. Plant Physiol 143:639–649. doi:10.1104/pp.106.090449
Romberger SP, Golbeck JH (2010) The bound iron-sulfur clusters of type-I homodimeric reaction centers. Photosynth Res 104:333–346. doi:10.1007/s11120-010-9543-y
Romberger SP, Golbeck JH (2012) The FX iron-sulfur cluster serves as the terminal bound electron acceptor in heliobacterial reaction centers. Photosynth Res 111:285–290. doi:10.1007/s11120-012-9723-z
Romberger SP, Castro C, Sun Y, Golbeck JH (2010) Identification and characterization of PshBII, a second FA/FB-containing polypeptide in the photosynthetic reaction center of Heliobacterium modesticaldum. Photosynth Res 104:293–303. doi:10.1007/s11120-010-9558-4
Sakakibara H (2003) Differential response of genes for ferredoxin and ferredoxin:NADP+ oxidoreductase to nitrate and light in maize leaves. J Plant Physiol 160:65–70. doi:10.1078/0176-1617-00919
San Pietro A (2006) A personal historical introduction to photosystem I: ferredoxin + FNR, the key to NADP+ reduction. In: Golbeck JH (ed) Photosystem I: the light-driven plastocyanin:ferredoxin oxidoreductase. Springer, Dordrecht, pp 1–8
Sancho J (2006) Flavodoxins: sequence, folding, binding, function and beyond. Cell Mol Life Sci 63:855–864. doi:10.1007/s00018-005-5514-4
Schluchter WM, Bryant DA (1992) Molecular characterization of ferredoxin-NADP+ oxidoreductase in cyanobacteria: cloning and sequence of the petH gene of Synechococcus sp. PCC 7002 and studies on the gene product. BioChemistry 31:3092–3102
Schrautemeier B, Cassing A, Böhme H (1994) Characterization of the genome region encoding an fdxH-type ferredoxin and a new 2[4Fe–4S] ferredoxin from the nonheterocystous, nitrogen-fixing cyanobacterium Plectonema boryanum PCC 73110. J Bacteriol 176:1037–1046
Schrautemeier B, Neveling U, Schmitz S (1995) Distinct and differently regulated Mo-dependent nitrogen-fixing systems evolved for heterocysts and vegetative cells of Anabaena variabilis ATCC 29413: characterization of the fdxH1/2 gene regions as part of the nif1/2 gene clusters. Mol Microbiol 18:357–369. doi:10.1111/j.1365-2958.1995.mmi_18020357.x
Seo D, Kitashima M, Sakurai T, Inoue K (2016a) Kinetics of NADP+/NADPH reduction-oxidation catalyzed by the ferredoxin-NAD(P)+ reductase from the green sulfur bacterium Chlorobaculum tepidum. Photosynth Res 130:479–489. doi:10.1007/s11120-016-0285-3
Seo D, Soeta T, Sakurai H, Setif P, Sakurai T (2016b) Pre-steady-state kinetic studies of redox reactions catalysed by Bacillus subtilis ferredoxin-NADP+ oxidoreductase with NADP+/NADPH and ferredoxin. Biochim Biophys Acta 1857:678–687. doi:10.1016/j.bbabio.2016.03.005
Singh AK, McIntyre LM, Sherman LA (2003) Microarray analysis of the genome-wide response to iron deficiency and iron reconstitution in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol 132:1825–1839. doi:10.1104/pp.103.024018
Six C, Thomas JC, Thion L, Lemoine Y, Zal F, Partensky F (2005) Two novel phycoerythrin-associated linker proteins in the marine cyanobacterium Synechococcus sp. strain WH8102. J Bacteriol 187:1685–1694. doi:10.1128/JB.187.5.1685-1694.2005
Sjöblom S, Harjunpaa H, Brader G, Palva ET (2008) A novel plant ferredoxin-like protein and the regulator Hor are quorum-sensing targets in the plant pathogen Erwinia carotovora. Mol Plant Microbe Interact 21:967–978. doi:10.1094/MPMI-21-7-0967
Sousa FL, Shavit-Grievink L, Allen JF, Martin WF (2013) Chlorophyll biosynthesis gene evolution indicates photosystem gene duplication, not photosystem merger, at the origin of oxygenic photosynthesis. Genome Biol Evol 5:200–216. doi:10.1093/gbe/evs127
Terauchi AM, Lu S-F, Zaffagnini M, Tappa S, Hirasawa M, Tripathy JN, Knaff DB, Farmer PJ, Lemaire SD, Hase T (2009) Pattern of expression and substrate specificity of chloroplast ferredoxins from Chlamydomonas reinhardtii. J Biol Chem 284:25867–25878. doi:10.1074/jbc.M109.023622
Thimm O, Essigmann B, Kloska S, Altmann T, Buckhout TJ (2001) Response of Arabidopsis to iron deficiency stress as revealed by microarray analysis. Plant Physiol 127:1030–1043. doi:10.1104/pp.010191
Thomas JC, Ughy B, Lagoutte B, Ajlani G (2006) A second isoform of the ferredoxin:NADP+ oxidoreductase generated by an in-frame initiation of translation. Proc Natl Acad Sci USA 103:18368–18373. doi:10.1073/pnas.0607718103
Thompson AW, Huang K, Saito MA, Chisholm SW (2011) Transcriptome response of high- and low-light-adapted Prochlorococcus strains to changing iron availability. ISME J 5:1580–1594. doi:10.1038/ismej.2011.49
Tognetti VB, Palatnik JF, Fillat MF, Melzer M, Hajirezaei M-R, Valle EM, Carrillo N (2006) Functional replacement of ferredoxin by a cyanobacterial flavodoxin in tobacco confers broad-range stress tolerance. Plant Cell 18:2035–2050. doi:10.1105/tpc.106.042424
Toulza E, Tagliabue A, Blain S, Piganeau G (2012) Analysis of the global ocean sampling (GOS) project for trends in iron uptake by surface ocean microbes. PLoS One 7:e30931. doi:10.1371/journal.pone.0030931
Trifonov EN (2000) Consensus temporal order of amino acids and evolution of the triplet code. Gene 261:139–151
Twachtmann M, Altmann B, Muraki N, Voss I, Okutani S, Kurisu G, Hase T, Hanke GT (2012) N-Terminal structure of maize ferredoxin:NADP+ reductase determines recruitment into different thylakoid membrane complexes. Plant Cell 24:2979–2991. doi:10.1105/tpc.111.094532
Vorst O, Dam F, Weisbeek P, Smeekens S (1993) Light-regulated expression of the Arabidopsis thaliana ferredoxin A gene involves both transcriptional and post-transcriptional processes. Plant J 3:793–803. doi:10.1111/j.1365-313X.1993.00793.x
Voss I, Koelmann M, Wojtera J, Holtgrefe S, Kitzmann C, Backhausen JE, Scheibe R (2008) Knockout of major leaf ferredoxin reveals new redox-regulatory adaptations in Arabidopsis thaliana. Physiol Plant 133:584–598. doi:10.1111/j.1399-3054.2008.01112.x
Voss I, Goss T, Murozuka E, Altmann B, McLean KJ, Rigby SE, Munro AW, Scheibe R, Hase T, Hanke GT (2011) FdC1, a novel ferredoxin protein capable of alternative electron partitioning, increases in conditions of acceptor limitation at photosystem I. J Biol Chem 286:50–59. doi:10.1074/jbc.M110.161562
Wächtershäuser G (2007) On the chemistry and evolution of the pioneer organism. Chem Biodivers 4:584–602. doi:10.1002/cbdv.200790052
Watanabe M, Ikeuchi M (2013) Phycobilisome: architecture of a light-harvesting supercomplex. Photosynth Res 116:265–276. doi:10.1007/s11120-013-9905-3
Waters M, Pyke K (2005) Plastid development and differentiation. Blackwell, Oxford
Weiss MC, Sousa FL, Mrnjavac N, Neukirchen S, Roettger M, Nelson-Sathi S, Martin WF (2016) The physiology and habitat of the last universal common ancestor. Nat Microbiol 1:16116. doi:10.1038/nmicrobiol.2016.116
Whitney LP, Lins JJ, Hughes MP, Wells ML, Chappell PD, Jenkins BD (2011) Characterization of putative iron responsive genes as species-specific indicators of iron stress in thalassiosiroid diatoms. Front Microbiol 2:234. doi:10.3389/fmicb.2011.00234
Yang C, Hu H, Ren H, Kong Y, Lin H, Guo J, Wang L, He Y, Ding X, Grabsztunowicz M, Mulo P, Chen T, Liu Y, Wu Z, Wu Y, Mao C, Wu P, Mo X (2016) LIGHT-INDUCED RICE1 regulates light-dependent attachment of LEAF-TYPE FERREDOXIN-NADP+ OXIDOREDUCTASE to the thylakoid membrane in rice and Arabidopsis. Plant Cell 28:712–728. doi:10.1105/tpc.15.01027
Yeh AP, Chatelet C, Soltis SM, Kuhn P, Meyer J, Rees DC (2000) Structure of a thioredoxin-like [2Fe–2S] ferredoxin from Aquifex aeolicus. J Mol Biol 300:587–595. doi:10.1006/jmbi.2000.3871
Yeom J, Jeon CO, Madsen EL, Park W (2009) Ferredoxin-NADP+ reductase from Pseudomonas putida functions as a ferric reductase. J Bacteriol 191:1472–1479. doi:10.1128/JB.01473-08
Zahnle K, Arndt N, Cockell C, Halliday A, Nisbet E, Selsis F, Sleep NH (2007) Emergence of a habitable planet. Space Sci Rev 129:35–78. doi:10.1007/s11214-007-9225-z
Zeng Y, Feng F, Medová H, Dean J, Koblížek M (2014) Functional type 2 photosynthetic reaction centers found in the rare bacterial phylum Gemmatimonadetes. Proc Natl Acad Sci USA 111:7795–7800
Zhao J, Qiu Z, Ruan B, Kang S, He L, Zhang S, Dong G, Hu J, Zeng D, Zhang G, Gao Z, Ren D, Hu X, Chen G, Guo L, Qian Q, Zhu L (2015) Functional inactivation of putative photosynthetic electron acceptor ferredoxin C2 (FdC2) induces delayed heading date and decreased photosynthetic rate in rice. PLoS One 10:e0143361. doi:10.1371/journal.pone.0143361
Zheng M, Doan B, Schneider TD, Storz G (1999) OxyR and SoxRS regulation of fur. J Bacteriol 181:4639–4643
Ziegler GA, Schulz GE (2000) Crystal structures of adrenodoxin reductase in complex with NADP+ and NADPH suggesting a mechanism for the electron transfer of an enzyme family. BioChemistry 39:10986–10995. doi:10.1021/bi000079k
Zöllner A, Nogués I, Heinz A, Medina M, Gómez-Moreno C, Bernhardt R (2004) Analysis of the interaction of a hybrid system consisting of bovine adrenodoxin reductase and flavodoxin from the cyanobacterium Anabaena PCC 7119. Bioelectrochemistry 63:61–65. doi:10.1016/j.bioelechem.2003.10.014
Acknowledgements
This work was supported by Grant PICT 2012–2851 from the National Agency for the Promotion of Science and Technology (ANPCyT, Argentina). NC is a staff member of CONICET, Argentina, and JJPK is a fellow of Bunge & Born Foundation (Fundación Bunge y Born, Argentina). NC and JJPK are Faculty members of the Molecular Biology Unit, School of Biochemical and Pharmaceutical Sciences, University of Rosario (Facultad de Ciencias Bioquímicas y Farmacéuticas, Universidad Nacional de Rosario, Argentina).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pierella Karlusich, J.J., Carrillo, N. Evolution of the acceptor side of photosystem I: ferredoxin, flavodoxin, and ferredoxin-NADP+ oxidoreductase. Photosynth Res 134, 235–250 (2017). https://doi.org/10.1007/s11120-017-0338-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-017-0338-2