Abstract
We performed a BLAST search of the rice database and screened four group 3 late embryogenesis abundant genes (OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5) that shared characteristics of canonical G3LEAs such as multiple copies of consensus motif, hydrophilic, structural intrinsic disorder, thermostability, abscisic acid (ABA)-responsiveness, and high G and C content in gene sequence. Under nonstress conditions, OsG3LEA-20.5 and OsG3LEA-24.5 were constitutively expressed, whereas OsG3LEA-47.3 and OsG3LEA-41.9 were expressed in a stage- or tissue-specific manner. Transcripts of OsG3LEA-20.5 and OsG3LEA-24.5 accumulated under salt, ABA, and cold stress treatment. By contrast, OsG3LEA-47.3 and OsG3LEA-41.9 showed less responsiveness to stress in tillering and heading stages, respectively. All genes showed enhanced transcript levels after pollination and embryo development. To investigate the functions, we overexpressed OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 in Arabidopsis (47.3-ox, 41.9-ox, 20.5-ox, and 24.5-ox). Analysis of tolerance to drought stress revealed higher recovery after 14 days of dehydration treatment for 47.3-ox or 41.9-ox than the wild type (WT), 20.5-ox, or 24.5-ox. In addition, 47.3-ox and 24.5-ox plants showed higher survival than WT, 41.9-ox, and 20.5-ox plants under heat treatment, which induced similar expression patterns of heat shock protein genes in WT, 47.3-ox, 41.9-ox, 20.5-ox, and 24.5-ox plants. In vitro chaperone activity with the model substrate citrate synthase was comparable between OsG3LEA-47.3 and OsHSP16.9A, a rice molecular chaperone providing thermoprotection in vivo and in vitro. These results suggest that rice G3LEAs share many physical and biological features but function in contrasting ways.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Late embryogenesis abundant (LEA) proteins were first identified and characterized in cotton and wheat during the last stage of seed maturation (Galau and Hughes 1987). Up to 4 % of total cellular proteins during seed development are LEA proteins (Roberts et al. 1993). LEA proteins are grouped into various families by the presence of a particular sequence motif and distinct physico-chemical properties, with no significant similarity among family members (Bies-Etheve et al. 2008; Jaspard et al. 2012). LEA proteins have high hydrophilicity and a high proportion of glycine or small amino acids (alanine, serine, and threonine), but they lack or have low content of tryptophan and cysteine residues (Stacy and Aalen 1998). LEA proteins are also characterized by their thermal stability, nonglobular structure, and low complexity. They may fold into special structures in response to stress conditions (Lisse et al. 1996; Soulages et al. 2002; Tompa 2002; Goyal et al. 2003; Soulages et al. 2003; Wise and Tunnacliffe 2004). Some Arabidopsis LEA genes are activated in vegetative tissues under nonstress conditions (Bies-Etheve et al. 2008). However, the physiological functions of LEA proteins are still unclear.
The group 3 LEA proteins are principally composed of multiple tandem repeats of an 11-mer amino acid motif (TAQAAKEKAGE) that forms an amphiphilic helical structure (Dure 1993). Group 3 LEA proteins are involved in binding and replacement of water, ion sequestration, maintenance of protein and membrane structure, molecular chaperoning, and regulation of development (Bray 1993; Koag et al. 2003). Barley HVA1, a group 3 LEA gene, can be rapidly induced by abscisic acid (ABA) and accumulates at high levels in the aleurone layer of barley seeds (Hong et al. 1992). Overexpression of barley HVA1 can confer high tolerance to water deficit and hypersalinity in transgenic rice (Xu et al. 1996). Furthermore, these transgenic rice plants grew faster than controls after recovery under nonstress conditions (Xu et al. 1996). AavLEA1, a group 3 LEA protein from the nematode Aphelenchus avenae, behaves as a molecular shield to prevent protein aggregation (Tunnacliffe and Wise 2007).
Rice, a model plant in monocots, is one of the most important cereal crops in tropic and temperate regions of the world. The normal growth and yield may be decreased by environmental stress. LEA proteins are upregulated in vegetative tissues in response to water deficit and osmotic stress. OsLEA3, a rice group 3 LEA protein, shows ABA responsiveness and tissue-specific expression under salt stress. Overexpressing OsLEA3 in rice enhanced drought tolerance in the field (Machuka et al. 1999).
Here, we used the conserved tandem repeat of HVA1 as a template with a BLAST search to screen unidentified or uncharacterized rice group 3 LEA genes (OsG3LEAs) from the rice database. We found four putative genes of OsG3LEA in cultivar Tainung 67 (TNG67) (GenBank accession no. AP004018, AC073556, AC098833, and AP000836), designated as OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5, respectively (Table 1). In agreement with characteristics of group 3 LEA proteins, these four proteins contained multiple copies of an 11 amino acid consensus motif, apparent amphipathic α helices and random coils, and high G and C content (Table 1). Gain-of-function analyses revealed that overexpression of the OsG3LEA genes conferred drought tolerance or thermotolerance in transgenic Arabidopsis plants (Xu et al. 1996).
Materials and Methods
Plant Materials
Seeds of rice (Oryza sativa L. cv. TNG67) were surface-sterilized in bleach for 20 min and washed with water. The seeds were allowed to germinate for 5∼8 days at 28 °C; then, the seedlings were transplanted into soil and grown under 14-h light/10-h dark conditions. For full-length gene cloning, the 20-day-old rice seedlings with 20-μM ABA treatment for 12 h were used. For ABA, NaCl, and cold treatment, samples collected from four development stages (seedling, tillering, heading, and seed maturation) were incubated in shaking buffer (1 mM potassium phosphate, 1 % sucrose, 50 μg/ml chloramphenicol, pH 6.0) containing 20 μM ABA or 150 mM NaCl or treated with 4 °C as indicated. For drought treatment, samples collected from seedling and heading stage were allowed to dry (28 °C, 60 % relative humidity) for the indicated times. For heat shock (HS) treatment, samples collected from seedling and heading stage were incubated in shaking buffer at 41 °C for the indicated times. The samples were frozen in liquid nitrogen and stored at −80 °C.
The Arabidopsis thaliana ecotype Col-0 was used as the wild type (WT). Arabidopsis plants were grown at 23 °C in a 16-h light/8-h dark cycle in a growth chamber with 60 % relative humidity.
Bioinformatics Analysis and Data Mining
The MEME tool (http://meme.sdsc.edu/meme) was used for searching the conserved motif of rice OsG3LEA genes. Gene sequence similarity analyses involved use of LALIGN (http://www.ch.embnet.org/software/LALIGN_form.html) and the European Bioinformatics Institute tool box ClustalW (http://www.ebi.ac.uk/Tools/clustalw2/index.html). A phylogenetic tree was built with use of ClustalΩ. For secondary structure predictions, NPS@ (http://npsa-pbil.ibcp.fr/cgi-bin/npsa_automat.pl?page=npsa_nn.html) and the Kyte and Doolittle program (http://www.expasy.org/tools/protscale.html) were used (Kyte and Doolittle 1982). Protein signal sequences and cellular localization were predicted by use of Signal Peptide Prediction/SIG-Pred (http://bioinformatics.leeds.ac.uk/prot_analysis/Signal.html), SOSUI (http://bp.nuap.nagoya-u.ac.jp/sosui/sosuisignal/sosuisignal_submit.html#sample), Signal Peptide web server (http://www.cbs.dtu.dk/services/SignalP/), and MultiLoc/TargetLoc (http://www-bs.informatik.uni-tuebingen.de/Services/MultiLoc/). IUPred (http://iupred.enzim.hu/index.html) was used to characterize intrinsically unstructured protein (Dosztanyi et al. 2005).
Subcellular Localization Analysis
The coding regions of OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 complementary DNA (cDNA) were PCR-amplified with the primers in supplementary Table S1. The PCR products were then separately fused upstream of the green fluorescence protein (GFP) at the XbaI/BamHI sites in the p326GFP vector (35S::GFP) (Lee et al. 2001), which resulted in 47.3-, 41.9, 20.5, and 24.5 GFP constructs under the control of the CaMV35S promoter. The vector p326GFP was used as a free GFP control, and the vector pARR6-GFP (35S::ARR6-GFP) was used to show the site of nucleus (Hwang and Sheen 2001). The plasmids were transiently expressed in epidermal onion cells by particle bombardment as described (Guan et al. 2010). A helium biolistic particle delivery system (model PDS-1000, Bio-Rad) was used for particle bombardment. In total, 5 μg recombinant plasmid was used per transformation and all target materials were bombarded twice. The bombarded onion epidermal cells were recovered at 25 °C in the dark for at least 16 h and observed under an Olympus U-LH 100HG inverted fluorescence microscope.
Reverse Transcription Polymerase Chain Reaction
Transcript levels in different tissues and under salt, ABA, and cold treatments were analyzed by reverse transcription polymerase chain reaction (RT-PCR). Total RNA was extracted as described (Guan et al. 2010). The first-strand cDNA was synthesized from 2 μg total RNA in a 20-μl reaction volume with use of the SuperScriptIII First-Strand Synthesis system (Invitrogen). RNA and cDNA were quantified by optical density measurement with use of a spectrophotometer (NanoDrop 1000, Thermo Scientific, Waltham, MA, USA). PCR amplification conditions were 24 cycles of 30 s at 95 °C, 30 s at 55 °C, and 30 s at 72 °C, then 5 min at 72 °C. Primers used for analysis of gene expression were designed with use of Primer3 (http://frodo.wi.mit.edu/) and are in supplementary Table S1. DNA from 15 μl of each PCR reaction underwent 1.2 % (w/v) agarose gel electrophoresis. ImageJ for Windows (http://rsbweb.nih.gov/ij/) was used to quantify the intensity of the ethidium-bromide–stained DNA bands. The expression of 18S ribosomal RNA (rRNA) was an internal control. The relative expression level was visualized as heatmap using the Heatmap Illustrator, version 1.0 (Deng et al. 2014).
Quantitative Real-Time Reverse Transcription Polymerase Chain Reaction
Transcript levels of the OsG3LEAs under drought and heat stress were analyzed by quantitative real-time RT-PCR (qRT-PCR) by use of the iCycler-iQ5 Multicolor Real-time PCR Detection System and iQ SYBR Green Hot-start Supermix (Bio-Rad, Hercules, CA, USA) with the primer sequences in Supplementary Table S1. Real-time PCR reaction involved 60 ng cDNA with each set of primers and the iQ™ SYBR® Green Supermix (Bio-Rad). PCR cycling included an initial step at 95 °C for 5 min, then 40 cycles of 10 s at 95 °C, 30 s at 56 °C, and 20 s at 72 °C. The comparative CT method was used to determine the relative amount of each sample, with the expression of 18S rRNA as an internal control.
Preparation of DNA Constructs and Transformation
For analysis of physiological functions, a DNA fragment of CaMV35S-Nos bearing cloning sites for XbaI/BamHI/KpnI was cut from pBI121 shuttle vector and subcloned into pCAMBIA1301. Then, the XbaI/BamHI fragments of OsG3LEA-47.3, OsG3LEA-41.9, and OsG3LEA-20.5 coding regions and the XbaI/KpnI fragment of the OsG3LEA-24.5 coding region were separately cloned into the vector in the sense orientation downstream of the CaMV35S promoter to generate the overexpression vectors pOsG3LEA-47.3-ox, pOsG3LEA-41.9-ox, pOsG3LEA-20.5-ox, and pOsG3LEA-24.5-ox. Sequences for all primers are in supplementary Table S1. All constructs were verified by sequencing analysis. The recombinant vectors were transferred to Agrobacteria (GV3101) and used to transform A. thaliana by the floral-dip technique (Clough and Bent 1998). Transformed Arabidopsis seedlings were selected on 1/2 Murashige & Skoog medium (MS; Duchefa) that contained 50 μg/ml hygromycin, 1 % sucrose (w/v), and 0.8 % agar. The hygromycin-resistant T1 seedlings were transferred to soil and grown to maturity. Homozygous T2 plants were selected by growth of their T3 generations on 1/2 MS medium containing 50 μg/ml hygromycin, 1 % sucrose (w/v), and 0.8 % agar. T3 seeds derived from homozygous T2 plants were used for subsequent tests.
Stress Treatment of Transgenic Arabidopsis Plants
For HS treatment, T3 generation transgenic Arabidopsis was germinated on 1/2 MS medium containing 1 % sucrose (w/v) and 0.8 % agar for 7 days and then shifted to 44 °C for 27 min, then 23 °C for 7 days of recovery. For drought tolerance assay, 2-week-old plants grown in soil were subjected to progressive drought stress by withholding water for 14 days and then re-watered for the next 7 days of recovery. Survival (ratio of surviving to total plants) was calculated after stress treatments.
Expression and Purification of Rice G3LEA Fusion Proteins
To express rice G3LEA genes, DNA constructs encoding OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 were prepared by PCR and subcloned into the BamHI-XhoI site of the pET21a expression vector (Novagen) to produce fusion proteins as described (Yeh et al. 2002). Expressed in E. coli, the plasmids produced the rice OsG3LEA proteins fused with an N-terminal 1.7 kDa-T7 tag and a C-terminal 6XHis tag. Transformed E. coli cells were grown in Luria-Bertani (LB) broth containing 100 μg/ml ampicillin at 37 °C overnight. The overnight cultures were diluted 10-fold with fresh LB broth plus ampicillin, and incubation continued at 37 °C until mid-log phase (OD600 = ∼0.6). Isopropyl-β-d-thiogalactopyranoside (IPTG) was then added to a final concentration of 1 mM and incubation continued at 37 °C for 2 h. After IPTG induction, cultures were harvested and cell pellets were obtained by centrifugation. Pellets were resuspended in phosphate-buffered saline (150 mM NaCl, 16 mM Na2HPO4, 4 mM NaH2PO4·2H2O, pH 7.8), and cells were sonicated for 10 min on ice. After centrifugation (18,000 g, 15 min at 4 °C), the supernatant was applied to a nickel-agarose column (Novagen). Purified fusion proteins were eluted with 1 M imidazole according to the manufacturer’s instructions (Novagen).
Chaperone Activity Analysis
Thermal aggregation suppression analysis was performed as described (Yeh et al. 2002) with porcine heart citrate synthase (CS; Sigma) used as a substrate. An amount of 360 nM of CS (monomer concentration) was incubated with or without 3.6 μM of purified rice G3LEA recombinant proteins in 50 mM potassium buffer, pH 7.5 (800 μl total). Samples were monitored for light scattering at 320 nm with a Hitachi U3200 spectrophotometer with a thermostated cell compartment preheated at 43 °C.
Statistical Analysis
Data are shown as mean ± SE from three independent experiments. Statistical differences were analyzed by Student’s t test or Duncan’s multiple range test. P < 0.05 was considered statistically significant.
Results
Screen for Rice G3LEA Proteins
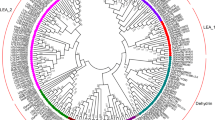
On the basis of G3LEA proteins with consensus 11-mer amino acid tandem repeats, we cloned and deduced four putative proteins containing the specific motif for further assay. The accession numbers of the proteins are BAD19162 (OsG3LEA-47.3), AAL84288 (OsG3LEA-41.9), AAU43988 (OsG3LEA-20.5), and BAD81113 (OsG3LEA-24.5), with 12, 13, 6, and 5 conserved motifs, respectively (Table 1) (Dure 1993). Multiple sequence alignment showed OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, or OsG3LEA-24.5 with 44, 57, 61, and 35 % identity, respectively, to HVA1 from barley (Hong et al. 1988); 38, 71, 98, and 28 % identity to OsLEA3 from rice (Moons et al. 1997); and 39, 64, 32, and 4 % identity to AavLEA1 from nematodes (Solomon et al. 2000) (Fig. 1a). Thus, we considered OsG3LEA-20.5 as an allele of OsLEA3 on the basis of their sequence similarity and location on chromosome 5 (Machuka et al. 1999). Phylogenetic tree analysis with other G3LEA genes from alfalfa, Arabidopsis, barley, brine shrimp, common bean, chickpea, cotton, grass, green algae, maize, nematode, rape, rice, sorghum, and wheat revealed the sequence association as three major clades in the evolution (Fig. 1b).
Amino acid sequence alignment of rice OsG3LEAs with other G3LEAs. a The sequences aligned were from rice OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, OsG3LEA-24.5, Barley HVA1, rice OsG3LEA, and Nematodes AavLEA1. The putative consensus tandem repeats are indicated by italics. Regions in black- and gray-shaded boxes correspond to fully conserved residues and partially conserved residues, respectively. The alignment involved ClustalΩ (http://www.ebi.ac.uk/Tools/msa/clustalo/). The N-terminal membrane anchor signal peptide of OsG3LEA-41.9 is underlined. b Phylogenic tree with 44 group 3 LEA proteins from Aphelenchus avenae, Triticum aestivum, Hordeum vulgare, Gossypium hirsutum, Medicago sativa, Oryza sativa, Zea mays, Brassica napus, Steinernema carpocapsae, Chlorella vulgaris, Arabidopsis thaliana, Phaseolus vulgaris, Agropyron mongolicum, Artemia franciscana, Sorghum bicolor, Sporobolus stapfianus, and Zizania latifolia. A phylogenetic tree was created from the aligned sequences by the neighbor-joining method with the MEGA program
Intrinsic disorder/unstructured proteins (IDPs/IUPs) may exert chaperone activity to prevent the aggregation of proteins caused in many stress-response processes and are found in most LEA proteins (Jaspard et al. 2012; Pazos et al. 2013). Protein structure analysis indicated that OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 were hydrophilic proteins (Supplementary Fig. S1) and included in the IDP/IUP family (Supplementary Fig. S2). In addition, transcription pattern analysis of the four rice G3LEA genes revealed that the mRNA levels in roots and shoots of rice seedlings were markedly higher than that without ABA (Fig. 3a), which implicates the genes in the rice seedling response to ABA. These characterizations were confirmed with the expression of typical G3LEA genes.
Subcellular Analysis of Selected Rice G3LEA Proteins
Bioinformatics analysis revealed that OsG3LEA-47.3, OsG3LEA-20.5, and OsG3LEA-24.5 were cytoplasmic proteins, and OsG3LEA-41.9 was an extracellular protein containing a putative membrane anchor signal peptide in the N-terminus. To acquire experimental data about the subcellular distribution of the four rice G3LEA proteins, the corresponding coding sequences were genetically fused with GFP in vectors designed for transient expression. Recombinant constructs were transformed into onion epidermal cells by particle bombardment. Fusion proteins were visualized by fluorescence microscopy. The fluorescence of OsG3LEA-47.3, OsG3LEA-20.5, and OsG3LEA-24.5 fusion proteins was predominantly in nuclei and cytosol fractions, similar to the localization pattern of GFP (Fig. 2). In contrast, OsG3LEA-41.9 fluorescence exhibited an exclusive cytosolic localization. 35S::ARR6-GFP construct was used as a nucleus specificity control.
Subcellular localization of OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5. Onion epidermal cells were particle-bombarded with the OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 fusion constructs, and localization of fluorescent signal was examined at 20 h after bombardment. As controls, GFP and NLS-GFP expression alone was analyzed
Expression of Rice G3LEA Genes
To study the role of rice G3LEA genes in rice, we examined expression profiles in vegetative and reproductive tissues (Fig. 3). The sampling points are shown in supplementary Fig. S3. We used the C-repeated binding factor 2 gene (RCBF2, GenBank accession no. AY345234), a rice transcription factor involved in dehydration- and cold-inducible gene expression, as a stress treatment indicator (Liu et al. 2007). The transcripts of OsG3LEA-20.5 and OsG3LEA-24.5 were detected in all plant organs at the stages tested, whereas those of OsG3LEA-41.9 were not detected in seedling roots and shoots, and OsG3LEA-47.3 was not expressed in roots and shoots during tillering. RCBF2 was expressed at most stages under normal conditions (Fig. 3).
Expression profiles of rice G3LEA genes in root (Rt), shoot (Sh), stem (St), and leaf (Lf). Transcript levels in different tissues and under salt, ABA, and cold treatments were analyzed by RT-PCR at a seedling, b tillering, c heading, and d reproductive stage. The transcript level was converted into heatmap. The five bar items with green (0.5-fold) and red (2.5-fold) gradient color change. Black color indicates no signal detected in the test (Color figure online)
Next, we examined the expression profiles of these four G3LEA genes in rice under stress treatment. In seedling roots, the transcripts of OsG3LEA-47.3 were significantly elevated with 1 h of NaCl treatment and then decreased, but OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 were activated after 1 h of salinity treatment and expressed up to 48 h (Fig. 3a). The four LEA genes showed ABA responsiveness under exogenous 20-μM ABA treatment and expression detected after 24 h of 4 °C treatment. At tillering stage, NaCl, ABA, and 4 °C treatment activated the expression of OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 in all organs. Cold stress did not activate the expression of OsG3LEA-47.3, and NaCl and ABA had less effect on the induction of OsG3LEA-47.3 (Fig. 3b). At heading stage, the expression of the four genes in stems peaked after 24 h of salinity or 4 °C treatment (Fig. 3c). OsG3LEA-41.9 and OsG3LEA-20.5 transcripts were less accumulated during NaCl treatment but showed sensitivity to ABA or 4 °C induction in root tissues. However, OsG3LEA-47.3 showed ABA responsiveness, and OsG3LEA-47.3 and OsG3LEA-24.5 peaked in level after 1 and 24 h of 4 °C treatment, respectively (Fig. 3c). The four rice G3LEA genes were highly expressed in the reproductive stage, especially in mature seeds (Fig. 3d). These results suggest that these four rice G3LEA genes function differentially in response to environmental stresses.
Under natural growth conditions of plants, heat-induced rapid and continuous evaporation or a decrease in water content of the leaf results in immediate desiccation. We thus studied the four G3LEA gene expression profilings of heat and drought responsiveness at rice seedling or heading stage. After drought treatment, elevated OsG3LEA-47.3, OsG3LEA-41.9, and OsG3LEA-20.5 expression was detected in the roots of rice seedlings, whereas no significant increase in OsG3LEA-24.5 transcripts was found (Fig. 4a). Regarding the effect of heat stress on induction of the four G3LEA genes, high temperature was shown to induce the accumulation of OsG3LEA-47.3, OsG3LEA-41.9, and OsG3LEA-20.5 transcripts in the root of rice seedling, but the expression of OsG3LEA-24.5 was decreased during heat stress (Fig. 4b).
Expression levels of the OsG3LEAs in rice. qRT-PCR analysis of the four G3LEAs in roots (R) and shoots (S) of seedling stage and leaves (L) from heading stage of rice plants treated with drought (a) and HS (b) for the indicated time. Bars are mean expression relative to that of 18S rRNA from three independent experiments. Nonstress control condition (Ctrl). *P < 0.05, **P < 0.01 compared with WT
Overexpression of OsG3LEA-47.3 or OsG3LEA-41.9 Enhances Drought Tolerance in Transgenic Arabidopsis Plants
To further investigate the roles of rice G3LEA proteins in drought and heat stress, we developed transgenic A. thaliana overexpressing OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 in ecotype Col-0 plants. The T3 homozygous transgenic lines 47.3-ox, 41.9-ox, 20.5-ox, and 24.5-ox, constitutively expressing OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, OsG3LEA-24.5, respectively, were used for further characterization. We compared plant height, leaf morphology, and flowering time between the WT and transgenic Arabidopsis plants grown under normal conditions. The 47.3-ox plants showed accelerated flowering, but the rosette leaf number was the same as for the WT (Supplementary Fig. S4). Plant height, leaf morphology, and flowering time did not differ among the WT, 41.9-ox, 20.5-ox, and 24.5-ox plants (Supplementary Fig. S5 and data not shown).
We tested the tolerance of 47.3-ox, 41.9-ox, 20.5-ox, and 24.5-ox plants to water deficiency. For drought stress treatment, water was withheld for 14 days from 14-day-old seedlings grown in pots at nonstress growth temperature. Under this dehydration condition, most WT and transgenic lines wilted (data not shown). As compared with the WT, 47.3-ox (Fig. 5a) and 41.9-ox (Fig. 5b) plants showed 3.5∼5.1-fold and 4.0∼8.4-fold higher recovery after 7 days of re-watering, respectively, with no effect on survival in 20.5-ox and 24.5-ox plants (data not shown). Thus, pre-accumulation of OsG3LEA-47.3 or OsG3LEA-41.9 protein in Arabidopsis transgenic plants could enhance the tolerance to a desiccation environment.
Drought tolerance of transgenic Arabidopsis overexpressing a OsG3LEA-47.3 and b OsG3LEA-41.9. Water was withheld from 14-old-day seedlings for 14 days. Survival (%) with 7 days of watering was calculated. RT-PCR analysis of mRNA expression of OsG3LEA-47.3 and OsG3LEA-41.9 in transgenic plants shown in the middle panel. Data are mean ± SD from three independent experiments (30 plants in each test). *P < 0.05 compared with Col-0 WT
Overexpression of OsG3LEA-47.3 Increases Thermotolerance in Arabidopsis Seedlings
We investigated the effect of overexpressing OsLEA3-47.3, OsLEA3-41.9, OsLEA3-20.5, or OsLEA3-24.5 on thermotolerance of transgenic Arabidopsis seedlings. After HS treatment, survival and recovery were better for OsG3LEA-47.3 and OsG3LEA-24.5 transgenic lines (57∼66 % and 43∼55 %, respectively) than the WT (21 %) (difference in survival rates >2-fold) (Fig. 6a, b). By contrast, survival of OsG3LEA-41.9 and OsG3LEA-20.5 transgenic plants was consistent with that of WT plants (11∼22and 22∼30 %, respectively) (Fig. 6b). The presence of OsG3LEA-47.3 and OsG3LEA-24.5 in Arabidopsis plants can protect cells against heat damage.
Viability of seedlings treated with HS (44 °C for 27 min). WT and OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5 Arabidopsis transgenic plants (47.3-ox, 41.9-ox, 20.5-ox, and 24.5-ox) were germinated on 1/2 MS medium for 7 days and then shifted to HS. a, b Survival (%) with 7 days of recovery at 23 °C. c RT-PCR of the mRNA expression of AtHSP genes (AtHSP17.7, AtHSP25.3, and AtHSP101) in heat-stressed Arabidopsis seedlings. The expression of genes is relative to that of the Col-0 WT control, and β-tubulin expression was an internal control. Data are mean ± SD from three independent experiments (30 plants in each test). *P < 0.05, **P < 0.01 compared with WT
Furthermore, we compared the transcript accumulation of some AtHSPs in OsG3LEA-47.3 and OsG3LEA-24.5 transgenic and WT plants. The expression of AtHSP17.7, AtHSP25.3, and AtHSP101 did not differ from the WT in OsG3LEA-47.3 transgenic lines under control or HS treatment (Fig. 6c). Similar expression levels of AtHSP17.7, AtHSP25.3, and AtHSP101 were also found in OsG3LEA-24.5 transgenic lines (data not shown). Overexpression of OsG3LEA-47.3 or OsG3LEA-24.5 did not upregulate the expression of AtHSP genes in Arabidopsis plants.
OsG3LEA-47.3 and OsG3LEA-24.5 Fusion Proteins Can Prevent Thermal Aggregation of Citrate Synthase
To assess the stability of OsG3LEA-47.3, OsG3LEA-24.5, and OsG3LEA-20.5 fusion proteins, we examined the change in light scattering of recombinant proteins at 44 °C. We used the OsHSP16.9A fusion protein, identified as a molecular chaperone to increase the thermotolerance of transgenic cells (Yeh et al. 2002), as a control. Incubation at high temperature did not increase the light scattering of OsG3LEA-47.3, OsG3LEA-24.5, and OsG3LEA-20.5 fusion proteins (Fig. 7a), so they were stable under thermal stress. As well, OsLEA3-41.9 fusion protein was stable at 44 °C (data not shown). The OsG3LEA-47.3 and OsG3LEA-24.5 proteins may be able to confer thermotolerance in transgenic Arabidopsis plants because of their ability to prevent thermal aggregation of cellular proteins. Thus, we compared the anti-aggregation property of the four G3LEA fusion proteins using the model substrate CS. Light scattering with CS aggregation was detected at 43 °C but was suppressed with 3.6 μM of OsG3LEA-24.5, OsG3LEA-47.3, or OsHSP16.9A fusion protein (58, 76, or 81 % reduction, respectively) (Fig. 7b). However, 3.6 μM of OsG3LEA-20.5 fusion protein could not efficiently suppress CS thermal aggregation (21 % reduction). OsLEA3-41.9 fusion protein, like OsG3LEA-20.5 fusion protein, did not prevent CS thermal aggregation (data not shown). Therefore, OsG3LEA-47.3 and OsG3LEA-24.5 fusion proteins could provide substantial thermostabilization of CS against heat aggregation in vitro.
In vitro chaperone activity of OsG3LEA recombinant proteins. a Light scattering analysis (320 nm) at 44 °C of stability of 3.6 μM purified OsG3LEA-47.3, OsG3LEA-20.5, OsG3LEA-24.5, and OsHSP16.9A and b with and without citrate synthase (CS; 360 nM) incubation at 43 °C for up to 30 min. Curves are CS alone and with OsG3LEA-47.3, OsG3LEA-20.5, OsG3LEA-24.5, or OsHSP16.9A. Data are mean ± SD from three independent experiments
Discussion
With knowledge of the well-documented 11-mer repeat of G3LEA proteins, we screened OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5, which showed features typical of the OsG3LEA proteins: a predicted protein sequence that is highly disordered, with significant hits for sets of motifs and strong 11-mer periodicity (Fig. 1a). The structures and physiological functions of LEA proteins may be strongly related to the types of amino acids, especially those located in the conserved repeat motifs (Dure 1993). These proteins also formed a good cluster with other members of the OsG3LEA proteins on phylogenetic analysis (Fig. 1b).
In Arabidopsis, some cytosolic LEA proteins can diffuse into the nucleus during stress conditions, which suggests that LEA proteins are associated with the cell membrane, cytoskeleton, and within the nucleus for their protective functions (Candat et al. 2014). Our results showed OsG3LEA-47.3, OsG3LEA-20.5, and OsG3LEA-24.5 with a dual cytosolic–nuclear localization (Fig. 2), whereas predictions denoted the three proteins as cytosolic proteins. Dual cytosolic–nuclear localization was hypothesized to result from passive diffusion of the cytosolic LEA–GFP fusion protein into the nucleus (Candat et al. 2014). Of note, GFP fusion protein analysis revealed that OsG3LEA-41.9, with an N-terminal membrane anchor signal peptide (Fig. 1a), localized exclusively in the cytosol, even though it was predicted as an extracellular protein. Bai et al. (2012) reported that accumulation of MsLEA3-1, an alfalfa LEA3 protein with a putative signal sequence in the N-terminal region, was able to mitigate membrane damage induced by salt stress. Thus, OsG3LEA-41.9 may be a membrane stabilizer when facing stress-induced membrane disturbance. In this study, we found that OsG3LEA-41.9 showed significant sensitivity to ABA and drought condition in rice root (Figs. 3 and 4a), and OsG3LEA-41.9-overexpressing Arabidopsis plants had higher recovery after drought treatment (Fig. 5b). Drought tolerance is known to be correlated with maintenance of membrane integrity (Premachandra et al. 1990). Our results agree with that OsG3LEA-41.9 may function in stabilizing the membrane system of rice plants exposed to detrimental conditions.
Water uptake in the roots is important for drought tolerance and transpiration efficiency. Based on the results of gene expression analyses, OsG3LEA-47.3 and OsG3LEA-41.9 transcripts significantly accumulated in rice roots after ABA, cold, drought, or heat treatment, whereas OsG3LEA-24.5 expression was repressed in seedling root during heat stress. Ectopic expression of OsG3LEA-47.3 and OsG3LEA-41.9 in Arabidopsis conferred resistance to drought stress (Figs. 4a and 5a, b), which coincided with LEA function in stabilization of the cell structure and water molecular binding and replacement (Bray 1993; Koag et al. 2003). Of note, after HS treatment, the survival was significantly higher in OsG3LEA-47.3 and OsG3LEA-24.5 transgenic Arabidopsis plants than the WT (Fig. 6b), which suggests that the function of OsG3LEA-24.5 is closely related with its accumulation level during heat stress. These results also suggest that OsG3LEA-47.3 may function in tolerance against heat and drought stresses through a pathway different from that for OsG3LEA-41.9 or OsG3LEA-24.5. In contrast, OsG3LEA-20.5 was constitutively expressed in all plant organs at the stages tested and showed sensitivity to stress induction. However, we found the survival of OsG3LEA-20.5 transgenic Arabidopsis plants comparable to that of WT plants after drought or HS treatment (Fig. 6b and data not shown), which suggests that OsG3LEA-20.5 may only function in rice.
Under HS, heat shock proteins (HSPs) are the most predominant proteins in plants, and upregulation of HSPs may protect metabolic activities and enhance heat tolerance (Yeh et al. 1997; Chang et al. 2007; Jagadish et al. 2010). However, we found similar accumulation levels of AtHSP transcripts among the WT, 47.3-ox, and 24.5-ox plants after HS treatment (Fig. 6c and data not shown), which suggests that overexpression of OsG3LEA-47.3 or OsG3LEA-24.5 can confer thermotolerance in Arabidopsis transgenic plants without mediating the expression of endogenous AtHSP genes. Furthermore, in the in vitro assay for preventing CS thermal aggregation, as compared with 3.6 μM OsHSP16.9A, OsG3LEA-47.3 or OsG3LEA-24.5 could efficiently prevent CS thermal aggregation (Fig. 7b). The chaperone functions of some molecular chaperones depend on ATP or other carbohydrates. LEA proteins could independently exhibit chaperone activity or cooperate with other chaperones to prevent the formation of damaging protein aggregates during stress (Goyal et al. 2005). However, in this study, OsG3LEA-47.3 may function as a chaperone depending on the concentration of the substrate (Supplementary Fig S6).
In conclusion, we identified four rice G3LEA proteins, OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5, that expressed differently under abiotic stresses. OsG3LEAs showed different expression patterns in tissues and developmental progress. OsG3LEA-47.3 plays an important role in conferring cross-tolerance for alleviating the detrimental effects of drought and heat treatment; OsG3LEA-41.9 and OsG3LEA-24.5 can confer resistance to drought and heat stress, respectively. OsG3LEA-20.5 is constitutively expressed under nonstress, ABA, NaCl, or low-temperature conditions but is unable to confer tolerance in Arabidopsis plants under heat or drought treatment. OsG3LEA-47.3 or OsG3LEA-24.5 can dose-dependently function in thermoprotection of proteins in vitro.
Abbreviations
- ABA:
-
Abscisic acid
- CS:
-
Citrate synthase
- GFP:
-
Green fluorescence protein
- HS:
-
Heat shock
- HSPs:
-
Heat shock proteins
- IPTG:
-
Isopropyl-β-d-thiogalactopyranoside
- LEAs:
-
Late embryogenesis abundant proteins
- RCBF2 :
-
C-repeated binding factor 2 gene
- WT:
-
Wild type
References
Bai Y, Yang Q, Kang J, Sun Y, Gruber M, Chao Y (2012) Isolation and functional characterization of a Medicago sativa L. gene, MsLEA3-1. Mol Biol Rep 39:2883–2892
Bies-Etheve N, Gaubier-Comella P, Debures A, Lasserre E, Jobet E, Raynal M, Cooke R, Delseny M (2008) Inventory, evolution and expression profiling diversity of the LEA (late embryogenesis abundant) protein gene family in Arabidopsis thaliana. Plant Mol Biol 67:107–124
Bray EA (1993) Molecular responses to water deficit. Plant Physiol 103:1035–1040
Candat A, Paszkiewicz G, Neveu M, Gautier R, Logan DC, Avelange-Macherel MH, Macherel D (2014) The ubiquitous distribution of late embryogenesis abundant proteins across cell compartments in Arabidopsis offers tailored protection against abiotic stress. Plant Cell 26:3148–3166
Chang CC, Huang PS, Lin HR, Lu CH (2007) Transactivation of protein expression by rice HSP101 in planta and using Hsp101 as a selection marker for transformation. Plant Cell Physiol 48:1098–1107
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Deng W, Wang Y, Liu Z, Cheng H, Xue Y (2014) HemI: a toolkit for illustrating heatmaps. PLoS One 7, e111988
Dosztanyi Z, Csizmok V, Tompa P, Simon I (2005) IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics 21:3433–3434
Dure L 3rd (1993) A repeating 11-mer amino acid motif and plant desiccation. Plant J 3:363–369
Galau GA, Hughes DW (1987) Coordinate accumulation of homeologous transcripts of seven cotton Lea gene families during embryogenesis and germination. Dev Biol 123:213–221
Goyal K, Tisi L, Basran A, Browne J, Burnell A, Zurdo J, Tunnacliffe A (2003) Transition from natively unfolded to folded state induced by desiccation in an anhydrobiotic nematode protein. J Biol Chem 278:12977–12984
Goyal K, Walton LJ, Tunnacliffe A (2005) LEA proteins prevent protein aggregation due to water stress. Biochem J 388:151–157
Guan JC, Yeh CH, Lin YP, Ke YT, Chen MT, You JW, Liu YH, Lu CA, Wu SJ, Lin CY (2010) A 9 bp cis-element in the promoters of class I small heat shock protein genes on chromosome 3 in rice mediates L-azetidine-2-carboxylic acid and heat shock responses. J Exp Bot 61:4249–4261
Hong B, Uknes SJ, Ho TH (1988) Cloning and characterization of a cDNA encoding a mRNA rapidly-induced by ABA in barley aleurone layers. Plant Mol Biol 11:495–506
Hong B, Barg R, Ho TH (1992) Developmental and organ-specific expression of an ABA- and stress-induced protein in barley. Plant Mol Biol 18:663–674
Hwang I, Sheen J (2001) Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 413:383–389
Jagadish SV, Muthurajan R, Oane R, Wheeler TR, Heuer S, Bennett J, Craufurd PQ (2010) Physiological and proteomic approaches to address heat tolerance during anthesis in rice (Oryza sativa L.). J Exp Bot 61:143–156
Jaspard E, Macherel D, Hunault G (2012) Computational and statistical analyses of amino acid usage and physico-chemical properties of the twelve late embryogenesis abundant protein classes. PLoS One 7, e36968
Koag MC, Fenton RD, Wilkens S, Close TJ (2003) The binding of maize DHN1 to lipid vesicles. Gain of structure and lipid specificity. Plant Physiol 131:309–316
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132
Lee YJ, Kim DH, Kim YW, Hwang I (2001) Identification of a signal that distinguishes between the chloroplast outer envelope membrane and the endomembrane system in vivo. Plant Cell 13:2175–2190
Lisse T, Bartels D, Kalbitzer HR, Jaenicke R (1996) The recombinant dehydrin-like desiccation stress protein from the resurrection plant Craterostigma plantagineum displays no defined three-dimensional structure in its native state. Biol Chem 377:555–561
Liu JG, Zhang Z, Qin QL, Peng RH, Xiong AS, Chen JM, Xu F, Zhu H, Yao QH (2007) Isolated and characterization of a cDNA encoding ethylene-responsive element binding protein (EREBP)/AP2-type protein, RCBF2, in Oryza sativa L. Biotechnol Lett 29:165–173
Machuka J, Bashiardes S, Ruben E, Spooner K, Cuming A, Knight C, Cove D (1999) Sequence analysis of expressed sequence tags from an ABA-treated cDNA library identifies stress response genes in the moss Physcomitrella patens. Plant Cell Physiol 40:378–387
Moons A, De Keyser A, Van Montagu M (1997) A group 3 LEA cDNA of rice, responsive to abscisic acid, but not to jasmonic acid, shows variety-specific differences in salt stress response. Gene 191:197–204
Pazos F, Pietrosemoli N, Garcia-Martin JA, Solano R (2013) Protein intrinsic disorder in plants. Front Plant Sci 4:363
Premachandra GS, Saneoka H, Ogata S (1990) Cell membrane stability an indicator of drought tolerance as affected by applied nitrogen in soybean. J Agri Sci 115:63–66
Roberts JK, DeSimone NA, Lingle WL, Dure L 3rd (1993) Cellular concentrations and uniformity of cell-type accumulation of two lea proteins in cotton embryos. Plant Cell 5:769–780
Solomon A, Salomon R, Paperna I, Glazer I (2000) Desiccation stress of entomopathogenic nematodes induces the accumulation of a novel heat-stable protein. Parasitology 121:409–416
Soulages JL, Kim K, Walters C, Cushman JC (2002) Temperature-induced extended helix/random coil transitions in a group 1 late embryogenesis-abundant protein from soybean. Plant Physiol 128:822–832
Soulages JL, Kim K, Arrese EL, Walters C, Cushman JC (2003) Conformation of a group 2 late embryogenesis abundant protein from soybean. Evidence of poly (L-proline)-type II structure. Plant Physiol 131:963–975
Stacy RA, Aalen RB (1998) Identification of sequence homology between the internal hydrophilic repeated motifs of group 1 late-embryogenesis-abundant proteins in plants and hydrophilic repeats of the general stress protein GsiB of Bacillus subtilis. Planta 206:476–478
Tompa P (2002) Intrinsically unstructured proteins. Trends Biochem Sci 27:527–533
Tunnacliffe A, Wise MJ (2007) The continuing conundrum of the LEA proteins. Naturwissenschaften 94:791–812
Wise MJ, Tunnacliffe A (2004) POPP the question: what do LEA proteins do? Trends Plant Sci 9:13–17
Xu D, Duan X, Wang B, Hong B, Ho T, Wu R (1996) Expression of a late embryogenesis abundant protein gene, HVA1, from barley confers tolerance to water deficit and salt stress in transgenic rice. Plant Physiol 110:249–257
Yeh CH, Chang PF, Yeh KW, Lin WC, Chen YM, Lin CY (1997) Expression of a gene encoding a 16.9-kDa heat-shock protein, Oshsp16.9, in Escherichia coli enhances thermotolerance. Proc Natl Acad Sci U S A 94:10967–10972
Yeh CH, Chen YM, Lin CY (2002) Functional regions of rice heat shock protein, Oshsp16.9, required for conferring thermotolerance in Escherichia coli. Plant Physiol 128:661–668
Acknowledgments
This work was supported by the National Science Council, Taiwan, ROC (no. NSC97-2313-B-008-001-MY3) to CH Yeh.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table S1
Oligonucleotides used in RT-PCR and coding region (CD) cloning. (DOCX 15 kb)
Supplementary Fig. S1
Hydropathy profiles for the deduced rice group 3 LEA proteins OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5. (PPTX 296 kb)
Supplementary Fig. S2
Schematic representation of the structural features of the deduced rice group 3 LEA proteins OsG3LEA-47.3, OsG3LEA-41.9, OsG3LEA-20.5, and OsG3LEA-24.5. The disorder prediction agrees with the standard 0.5 threshold. (PPTX 120 kb)
Supplementary Fig. S3
The growth stage and sampling points in TNG67. (PPTX 151 kb)
Supplementary Fig. S4
Overall appearance of Col-0 WT and OsG3LEA-47.3 Arabidopsis transgenic plants grown on soil for 24 days at 23oC. (PPTX 557 kb)
Supplementary Fig. S5
Leaf number at flowering of OsG3LEA-41.9 transgenic Arabidopsis plants. (PPTX 53 kb)
Supplementary Fig. S6
Concentrations of OsG3LEA-47.3 with citrate synthase (CS)under 43oC. (PPTX 118 kb)
Rights and permissions
About this article
Cite this article
Ke, YT., Lu, CA., Wu, SJ. et al. Characterization of Rice Group 3 LEA Genes in Developmental Stages and Under Abiotic Stress. Plant Mol Biol Rep 34, 1003–1015 (2016). https://doi.org/10.1007/s11105-016-0983-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-016-0983-1