Abstract
Ubiquitination is one of the most common posttranslational modifications. A series of E3 ligases are implicated in plant abiotic stress signaling, regulating the degradation of multiple specific target proteins. Here, we showed that a novel gene ABA-RESPONSE KELCH PROTEIN 1 (AtARKP1), which encodes an F-box subunit of Skp-cullin-F-box (SCF) ubiquitin ligase complex, was localized in the nucleus and could be induced by phytohormone abscisic acid (ABA) in Arabidopsis. ARKP1 interacted with ASK1 and ASK2, which tethered the rest of the complex to an F-box protein, suggesting that they might form an SCF ubiquitin ligase complex. Further analysis revealed that ARKP1 was exclusively expressed in the seed, rosette leaf, and root. arkp1 T-DNA insertion mutant plants were insensitive to ABA, displaying reduced ABA-mediated inhibition of seed germination, root elongation, and water loss rate of detached leaves. In contrast, transgenic plants showed enhanced sensitivity to ABA and tolerance to water deficit. Accordingly, the expressions of ABA and drought responsive marker genes were markedly upregulated in ARKP1 overexpressing plants than the wild-type and arkp1 mutant plants. Taken together, our findings suggest that AtARKP1 plays a positive role in ABA signaling network.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Abscisic acid (ABA) plays an important role during the life cycle of plants, such as seed dormancy, germination, and early seedling growth. More importantly, ABA enables the plants to tolerate the water-related stresses like drought, salinity, and wounding (Koornneef et al. 2002; Xiong et al. 2002; Zhu 2002). Much of E3 ligases have been reported to be involved in ABA signaling pathway. For example, abscisic acid insensitive 5 (ABI5) acts as a positive transcription factor that functions downstream of central protein kinases sucrose nonfermenting 1-related protein kinases (SnRK2s) and binds to the ABRE element in the promoter region of multiple ABA responsive genes (Lopez-Molina and Chua 2000; Lopez-Molina et al. 2001). ABI5 degradation via the ubiquitin/26S proteasome proteolytic pathway is regulated by keep on going (KEG) and ABI five binding protein 1 (AFP1) in nucleus (Lopez-Molina et al. 2003; Stone et al. 2006). ABI3 interacting protein 2 (AIP2) controls the transcription factor abscisic acid insensitive 3 (ABI3) stability (Zhang et al. 2005). F-box protein drought tolerance repressor (DOR) acts independent of phospholipase Da1 in an ABA signaling pathway to inhibit the ABA-induced stomatal closure under drought stress (Zhang Ye et al. 2008). Recently, two studies have firstly reported that ABA receptor RCAR3 is regulated by two different type multiple subunit E3 ligase complex. DET1-, DDB1-associated 1 (DDA1), as part of the CDD complex, directly binds to RCAR3 in vivo and facilitates its proteasomal degradation in the nucleus (Irigoyen et al. 2014). RSL1, a plasma membrane-located single RING-type protein, interacts with another two ABA receptors PYL4 and PYR1 to mediate their half-life (Bueso et al. 2014). Although some receptors and downstream transcription factors have been identified to be modified by E3 ligases, much of the other important components of ubiquitination remain largely unknown.
Ubiquitination mediated by 26S proteasome system, which is one of the most common posttranslational modifications, regulates multiple life cycles from cell division to cell death in higher plants (Moon et al. 2004; Smalle and Vierstra 2004; Dreher and Callis 2007). As so far, over 1400 genes encoding E3 ligases have been found in Arabidopsis thaliana genome (Vierstra 2009). Briefly, they can be divided into two groups based on their structure: single-subunit type and multiple-subunit type. Single-subunit E3 ligases are further categorized into distinct families based on the presence of specific domains, such as RING type (for really interesting new gene), U-Box type, and HECT (for homology to E6-AP carboxyl terminus) (Kraft et al. 2005; Stone et al. 2005). The other group is cullin-based complex E3 ligases, such as Skp1-cullin-F-box (SCF) complex, BCR complex, and DCX complex. Among them, the SCF E3 ligase complex is the best-characterized member of the family that consists of four subunits. The adaptor protein S-phase kinase-associated proteins (SKPs they mean ASKs in Arabidopsis) can bind to N-terminal of CULLIN1 and connect the F-box proteins with S-phase kinase-interacting protein (SKIPs) together (Craig and Tyers 1999; Lechner et al. 2006).
In this study, we identified a novel gene that encodes a kelch repeat F-box protein ABA-response kelch protein 1 (ARKP1). The expression of ARKP1 was induced by ABA, suggesting that it might be involved in ABA signaling pathway. The ABA-related phenotypes of T-DNA mutant and transgenic overexpression plants also indicated that ARKP1 played a positive role in the ABA signaling pathway.
Materials and Methods
Plant Materials and Growth Conditions
Arabidopsis thaliana plants, including the wild-type, T-DNA insertion mutant, and transgenic plants, were all the Columbia-0 ecotype. Plants were grown under greenhouse conditions in pots containing soil and vermiculite. Seeds were sterilized by treatment with bleach for 10 min and, finally, four times washing with sterile distilled water. Stratification of the seeds was conducted in the dark at 4 °C for 3 days. Then, seeds were grown on half strength Murashige and Skoog (MS) (Murashige and Skoog 1962) medium supplemented with 1 % (w/v) sucrose, 1 % (w/v) agar (pH 5.8). Plates were sealed and incubated in greenhouse at 22 °C under a 16-h light/8-h dark photoperiod.
Yeast Two-Hybrid Assay
The ARKP1 coding regions were fused to the GAL4 activation domain in pGADT7 as the prey construct. ASK1 and ASK2 were fused to the GAL4 DNA binding domain in pGBKT7 as the bait constructs (Clontech). Bait and prey constructs were cotransformed into yeast AH109 cells using PEG/LiAC method. All of the clones grew well on SD medium minus leucine and tryptophan (−LW). The positive clones were identified by the ability to grow on SD medium minus leucine, tryptophan, His, and adenine (−LWHA). The photographs were taken after 2 or 3 days. The primers used in this assay are listed in Supplemental Table S1.
Subcellular Localization
For subcellular localization, the ARKP1 open reading frame was cloned into the pSK-EGFP vector (Invitrogen) and fused with the N-terminal of enhanced GFP gene. About 4-week-old Arabidopsis plants were used for protoplast transfection as described previously (Yoo et al. 2007). The ARKP1-GFP plasmid was transfected into the digested protoplasts, and GFP signal was investigated after 16 h. 4′,6-Diamidino-2-phenylindole (DAPI) staining indicated the nucleus of protoplast. Photos were captured by a confocal fluorescence microscope (TCS SP5 II system, Leica, Germany).
Confirmation of T-DNA Mutant Plants
The T-DNA insertion mutant arkp1 (SALK_078824) was ordered from Arabidopsis Biological Resource Center (ABRC). The genomic DNA of mutant plants was extracted from the 3-week-old young leaves by Edward Buffer containing 100 mM Tris-HCl, 250 mM NaCl, 25 mM EDTA, and 0.5 % SDS, and followed the method performed as described previously (Edwards et al. 1991). The polymerase chain reaction experiment was performed using gene-specific primers arkp1-F (LP), arkp1-R (RP), and T-DNA-specific primer LBb. The primers used in this assay are listed in Supplemental Table S1.
Generation of ARKP1 Overexpressing Transgenic Plants
The ARKP1 coding region was ligated to the C-terminal of synthetic 6*HA (Influenza Hemagglutinin) peptide of pUC19 vector, then the coding region of ARKP1 with 6*HA was subcloned into the binary vector pBI121 (Jefferson et al. 1987) by replacing the GUS (β-glucuronidase) gene. The floral dip method used in this study was described previously (Clough and Bent 1998). The single coping transgenic plants which grow well (with green leaves and longer primary roots) with Mendelian segregation ratio (3:1) on Kanamycin resistance medium were transferred to soil in order to obtain their seeds (T2 seeds). Then, the T2 seeds were sown on Kanamycin resistance medium to identify the homozygous transgenic lines. After this, the homozygous seeds were used for phenotypic analysis. The primers used in this assay are listed in Supplemental Table S1.
Phenotypic Analysis
For germination assay in Arabidopsis, about 200 stratified seeds were sown on each medium with different ABA concentrations. Then, plants were cultivated in growth chambers under long-day conditions. The seed germination was indicated when the radicle extension reached 0.1 cm.
For root growth assay, seedlings were grown on vertically oriented MS plates for 4 to 5 days. Afterward, plants were transferred to new MS plates supplemented with 20 μM ABA. The root lengths were measured after 11 days.
For water loss assay, growing in the normal growth condition was used for the water loss assay. Four-week-old seedling rosette leaves were detached from the filter paper, and then the water loss rate was measured at the indicated intervals.
For drought tolerance assay, 4-week-old seedlings were stopped watering for 15 days and re-watering. The photographs were taken 5 days after re-watering. All of the phenotypic assays were performed by three technical replicates and three independent biological repetitions with the same results.
qRT-PCR Analysis of Gene Expression
For ABA and NaCl treatment assay, 9-day-old Arabidopsis seedlings (Col-0) grow in medium were transferred to medium with or without 50 μM ABA or 200 mM NaCl. Total RNA was extracted using liquid nitrogen from plants harvested at the indicated times (1, 3, 6, and 12 h) after each treatment using the TRIzol Reagent (Life Technologies, USA), and reverse transcription was performed with the PrimeScript™ RT reagent Kit with gDNA Eraser (TAKARA, Japan). For tissue specificity analysis of ARKP1, 5-week-old Col-0 plants were cultivated in growth chambers under long-day conditions. Roots, rosette leaves, open flowers, stems, and dry seeds were harvested for RNA isolation. All the qRT-PCR experiments for the analysis of different genes expression were performed using the instrument of CFX96 and the reagent iQ™ Multiplex Powermix (Bio-Rad). ACTIN2 was used as an internal control. All the qRT-PCR assays were performed by three technical replicates and three independent biological repetitions with the same results. The primers used in this assay are listed in Supplemental Table S1.
Results
Analysis of AtARKP1 Expression and Tissue Specificity
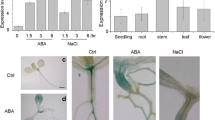
To identify novel components of E3 ligase involved in ABA signaling, we selected several F-box kelch proteins (FBKs) of previously published FBKs (Risseeuw et al. 2003) to search which of them were induced by the phytohormone ABA using the online eFP microarray database (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi). It showed that a candidate with the accession number At2g02870, encoding kelch repeat F-box E3 ligase, was induced by several phytohormone, such as ABA (10 μM, 3 h, Fig. 1a). To further confirm the microarray data, quantitative reverse transcript PCR (qRT-PCR) was performed to determine the expression of At2g02870 in response to ABA and NaCl. Our analysis demonstrated that the expression of At2g02870 was markedly induced by ABA treatment compared with normal condition. The expression level of At2g02870 reached a peak at 6 h with 5.2-fold increase and subsequently dropped down to 3.5-fold at 12 h (Fig. 1b). Moreover, the transcript abundance of At2g02870 was at a high level, about 3-fold under the treatment with exogenous NaCl compared with no stress treatment (Fig. 1b). Therefore, we named it ABA-response kelch protein 1 (AtARKP1).
Expression pattern of ARKP1 in Arabidopsis. a Microarray data showed ABA mediated upregulation of ARKP1 expression treatment for 0.5, 1, and 3 h in whole 7-day-old seedlings. Data were available at http://www.Arabidopsis.org/info/expression/ATGenExpress.jsp. b Analysis of the expression of ARKP1 by hormone and abiotic stress (50 μM ABA or 200 mM NaCl for 1 to 6 h) treatment in 9-day-old wild-type Arabidopsis using real-time quantitative PCR. The expression level under non-stress conditions was set to 1. ACTIN2 was used as an internal control. c Analysis of ARKP1 gene expression among different tissues in Arabidopsis using real-time quantitative PCR. The expression level in flower was set to 1. All the data in b, c represent the means ± SD of triplicate biological results
Furthermore, expression pattern analysis indicated that a higher expressional level of AtARKP1 transcripts was detected in 5-week-old rosette leaves, stems, roots, siliques, and dry seed (3.4-, 2-, 2.6-, 2.4-, and 5.3-fold, respectively) compared with the transcripts in open flowers (Fig. 1c). These data suggested that AtARKP1 is induced by ABA and expressed in different tissues of Arabidopsis, especially in leaves and seeds.
ARKP1 Is an F-box E3 Ubiquitin Ligase and Localizes in the Nucleus
AtARKP1 is constituted with 2318 base pairs and encodes 467 amino acid residues. The alignment of protein sequences from NCBI (http://www.ncbi.nlm.nih.gov/) showed that ARKP1 shares 65 to 95 % similar sequences to the deduced protein in Morus notabilis, Citrus sinensis, Nicotiana sylvestris, Arabidopsis alpine, Brassica rapa, Eutrema salsugineum, Capsella rubella, and Arabidopsis lyrata (Fig. 2a). Motif Scan (http://myhits.isb-sib.ch/cgi-bin/motif_scan) analysis (Fig. 2b) showed that ARKP1 contained four conserved domains: an F-box domain (116-163) and three kelch domains (236-343). F-box proteins in Arabidopsis have been classified into three groups (Gagne et al. 2002), and ARKP1 belongs to Clade A, suggesting that ARKP1 plays an important role in posttranslational during long life stage in plants (Schumann et al. 2011).
ARKP1 is an F-box kelch repeat protein in Arabidopsis. a Phylogenetic tree of plant ARKP1 proteins. The tree was built and visualized using MEGA 5 software based on the multiple sequence alignment from ClustalX V2.0 software. b A schematic showing the positions of the F-box E3 ligase activity domain and three conserved kelch repeat domains in the C-terminal of ARKP1 protein. Data were obtained from Motif Scan (http://myhits.isb-sib.ch/cgi-bin/motif_scan) analysis. c Yeast two-hybrid analysis of ARKP1 interaction with two members of ASKs. Interaction was determined by growth assay on selective SD medium lacking Leucine and Tryptophan (−LW) or Leucine, Tryptophan, His, and adenine (-LWHA). Dilutions (10−1, 10−2, and 10−3) of saturated cultures were spotted onto the plates. Photographs were taken 2 and 3 days after spotting the cultures onto the −LW and −LWHA medium respectively. The empty vector AD and BD was used as a negative control. d The subcellular localization of ARKP1-GFP fusion in Arabidopsis mesophyll protoplasts. 35S::ARKP1-GFP construct was transfected into the protoplasts, and the GFP signal was detected 16 h after transfection using a fluorescence microscope. DAPI staining indicated the nuclei. Scale bars 10 μm
Because the SCF complex is composed of four major subunits: the scaffolding protein CULLIN1, the adaptor ASKs, the E2 binding protein RBX1, and the F-box SKIP proteins which recognize specific substrate, ASKs, bind F-box proteins and scaffolding proteins CULLINs together as bridge (Lechner et al. 2006). Therefore, the interaction between ASKs and ARKP1 was investigated in this study. In yeast two-hybrid assay, the plasmids truncated with AD-ARKP1, BD-ASK1, and BD-ASK2 were cotransfected into the yeast AH109 cells. All the clones grew well on the SD medium lacking leucine and tryptophan (−LW), while on the medium lacking leucine, tryptophan, histidine, and adenine (−LWHA), only the truncations followed with both ARKP1 and ASK1/2 could grow (Fig. 2c). The results suggested that ARKP1 interacted with both ASK1 and ASK2 in yeast cells.
Furthermore, GFP fluorescence signal was detected after 35S::ARKP1-GFP constructs were transformed in Arabidopsis protoplasts and incubated for 16 h. With the same position indicated by DAPI staining, the GFP fluorescence signal was observed only in the nucleus. No GFP signal was found in the cytoplasm or cell membrane (Fig. 2d). These data suggest that ARKP1 is a nucleus-localized protein.
ARKP1 Is a Positive Regulator in ABA Responsive Phenotype
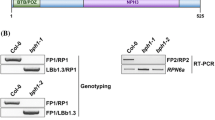
Because the messenger RNA (mRNA) level of ARKP1 is induced by ABA, so it prompted us to analyze its function in the ABA signaling in plants. The T-DNA insertion mutant arkp1 (SALK_078824) seeds were obtained from ABRC. PCR assay confirmed that the T-DNA was inserted into the 300 base pair promoter region of ARKP1 genomic DNA (Fig. 3a, c). The ARKP1 transcript level in the mutant plants decreased about 50 % compared with the wild-type plants (Fig. 3c, d). On the other hand, the ARKP1-transgenic plants were constructed by creating ARKP1 under the control of Cauliflower Mosaic Virus 35S promoter. Twenty individual ARKP1-overexpressing plants were isolated from T0 generation seeds. Among the 20 putative T1 transgenic lines, only eight lines segregated for in 3:1 kanamycin resistance separation ratio in the T1 generation. Three T2 homozygous transgenic lines (OE-1-7, OE-6-11, and OE-7-3) that expressed relatively higher ARKP1 levels were used for further phenotypic analysis (Fig. 3b, d).
Identification of arkp1 mutant plants and generation of ARKP1-overexpressing transgenic plants. a The genomic structure of ARKP1. The white boxes indicate 5′ UTR and 3′ UTR, and the black boxes indicate the exons and the black line indicates the intron. b Analysis of the genomic DNA level of ARKP1 transgenic plants. c PCR analysis of T-DNA insertion mutant arkp1 and RT-PCR analysis of ARKP1 transcript in arkp1 mutant. TUBULIN was used as an internal control. d Analysis of ARKP1 transcript expression in wild-type, mutant, and transgenic plants using real-time quantitative PCR. ACTIN2 was used as an internal control. The expression of ARKP1 in wild-type plants was set to 1. All the data represent the means ± SD of triplicate biological results
A seed germination assay on media with 0.5, 1, and 3 μM ABA was performed to investigate whether ARKP1 was involved in the ABA-dependent seed germination response. After 2 days, the germination rates of wild-type, ARKP1 overexpression, and T-DNA insertion mutant seeds were almost 100 % of media without ABA. However, in the presence of different concentration ABA, the seeds of ARKP1-overexpressing plants showed much lower germination rate than those of the wild-type and arkp1 mutant plants. In the medium containing 0.5 μM ABA, almost all the arkp1 mutant seeds were germinated at the fourth day, but the seeds of wild-type and ARKP1 overexpressing plants only reached 76 and 54 %, respectively (Fig. 4a). In the medium containing 1 μM ABA, about 20 % of arkp1 mutant seeds were germinated, but the wild-type seeds only reached 6 % germination rate. Moreover, the transgenic seeds did not germinate at all (Fig. 4b). After 6 days on medium containing 3 μM ABA, the germination rates of the wild-type, ARKP1-OE-1-7, ARKP1-OE-6-11, ARKP1-OE-7-3, and T-DNA mutant seeds were around 17.8, 9.4, 12.4, 8.6, and 51.5 %, respectively (Fig. 4c). In the presence of 1 μM ABA at the fourth day, 22 % seeds of wild-type and 8 % seeds of ARKP1 overexpressing plants germinated compared to 74 % in arkp1 mutant plants (Fig. 4d). These results demonstrated that ARKP1 positively regulated the physiological responses in Arabidopsis.
ARKP1 increased the sensitivity of Arabidopsis to ABA. a–c The germination rates on plates with different ABA concentration (0.5, 1, and 3 μM). Radicle reaching 0.1 millimeter indicates the seeds germination. d Wild-type (Col-0), arkp1 mutant, and ARKP1-overexpressing plants were grown for 5 days on an increasing concentration of ABA. After that, the germination rate was determined. All the data represent the means ± SD of triplicate biological results (n = 200). e, f Root growth of Col-0 wild-type, arkp1 mutant, and ARKP1-overexpressing plants that were vertically grown on medium in the absence or presence of 30 μM ABA for 11 days after sowing. Values are mean ± SD (n = 15). Significant difference was determined by a Student’s t test; a star indicates a P value of *P < 0.05; **P < 0.01. All the data represent the means ± SD of triplicate biological results
To identify whether ARKP1 functioned during the postgermination stage, germinated seedlings were then transferred to medium supplemented with or without ABA. In the root growth assay, no significant difference was observed in root lengths among all the plants without ABA (Fig. 4e). On the contrary, three ARKP1 overexpression transgenic lines displayed shorter primary roots compared with mutant plants supplemented with 20 μM ABA. After sowing for 11 days, the primary root length of wild type, ARKP1-OE-1-7, ARKP1-OE-6-11, and ARKP1-OE-7-3 only reached 29.6, 21.5, 26.1, and 20.5 mm, respectively, but that of the arkp1 mutant plants displayed about 38.3 mm (Fig. 4f).
ABA is one of the key phytohormone to regulate drought resistance in higher plants (Koornneef et al. 2002). So, water loss rate assay was employed to determine the function of ARKP1 in response to drought stress. Our analysis demonstrated that the water loss rate of arkp1 mutant plants was faster than the wild-type and overexpression plants, and ARKP1 overexpression plants showed the lowest water loss rate. After the leaves had been detached for 3 h, the arkp1 mutant plants contained only 31.8 % of the initial fresh weight, whereas the water loss rate was about 44.5, 63.8, 62.7, and 65.7 % in the wild-type and plants overexpressing ARKP1-OE-1-7, ARKP-OE-6-11, and ARKP-OE-7-3, respectively (Fig. 5a).
ARKP1 reduces drought tolerance in Arabidopsis. a Water loss rates of detached rosette leaves of the wild-type, arkp1 mutant, and ARKP1-overexpressing plants. Leaf weights of 4-week-old plants were determined at the indicated times. Data represent means ± SD of six leaves from each of three independent experiments. b Drought tolerance assay of 4-week-old Col-0 wild-type, arkp1 mutant, and ARKP1-overexpressing plants was performed by withholding water for 15 days and subsequently re-watering and examining after 5 days. Results are from three replicates, and values represent means ± SD (n = 20)
Since overexpression of ARKP1 conferred hypersensitivity to ABA during germination and postgermination stages, it was of particular interest to determine if loss of ARKP1 induces other ABA-associated phenotypes, such as increasing the drought tolerance. As expected, the arkp1 mutant plants showed less tolerance in drought condition (without watering for 15 days) compared with wild-type and overexpressing plants (Fig. 5b). After re-watering for 5 days, almost all arkp1-mutant plants were dead, whereas the survival rates were about 60 and 76 % in the wild-type and overexpression plants, respectively. Taken together, our results suggested that ARKP1 played a positive role in ABA signaling pathway during germination and postgermination stage.
ARKP1 Affects Expression of ABA Responsive Genes
The exogenous ABA induces the expression of ABA responsive genes, such as RELATED TO ABSCISIC ACID 18 (RAB18) and RESPONSIVE TO DESSICATION 29A (RD29A) (Lång and Palva 1992; Yamaguchi-Shinozaki and Shinozaki 1993; Lång et al. 1994). To further investigate whether the transcription levels of ABA and drought responsive genes might be affected by ARKP1, the mRNA levels of RAB18 and RD29A were analyzed in the wild-type, arkp1 mutant, and ARKP1-overexpression plants. In the presence of 50 μM ABA for 2 h, the transcripts of ABA responsive genes RAB18 and RD29A were about 3-fold higher in ARKP1-overexpressing plants compared with the wild type, while those in the arkp1 mutant plants reduced 40 % compared with wild-type plants (Fig. 6). For abscisic acid insensitive 2 (ABI2), the dominant negative regulator in ABA signaling (Leung et al. 1997), its transcript level was 1.5-fold higher in the arkp1 mutant plants compared with the wild-type plants, while the overexpression plants displayed decreased expression level. On the other hand, the expression of abscisic acid responsive elements-binding factor 3 (ABF3), a positive transcription factor in ABA signaling (Choi et al. 2000) expression, was elevated in ARKP1-overexpressing plants compared with the wild-type plants. Collectively, the results indicate that ARKP1 regulated ABA signaling pathway by altering multiple important components in ABA signaling pathway (Fig. 6).
The transcript abundance of ABA responsive genes. Nine-day-old seedlings were treated with liquid half strength MS culture with or without 50 μM ABA. The expression of different genes in wild-type plants was set to 1. ACTIN2 was used as an internal control. All the data represent the means ± SD of triplicate biological results
Discussion
ABA signaling pathway, which is regulated by E3 ligases, has been demonstrated to play vital roles in many cellular events, from biosynthesis to signaling cascade (Raab et al. 2009). The degradation of ABI5, an important positive transcription factor, was controlled by a number of E3 ligases, such as KEG, AFP1, DWA1, DWA2, and ABD1 (Lee et al. 2010; Seo et al. 2014). Not only the positive regulators mediated by the 26S proteasome E3 ligases in ABA signaling but also the dominant negative component, homeodomain transcription factor AtHB6, are substrates of CULLIN3-based E3 ligase MATH-BTB proteins (BPMs) (Himmelbach et al. 2002; Lechner et al. 2011). The BPM-ATHB6 model could be used to set up a similar model in this study. ARKP1 was shown to interact with ASK1 and ASK2 in yeast, indicating that they might form CULLIN1-based SCF type multiple-subunit E3 ligase to perform its posttranslational regulatory function. Subcellular localization analysis revealed that ARKP1 predominantly localized in the nucleus (Fig. 2d). Thus, ARKP1 may be involved in ABA signaling through regulation of certain ABA signaling factors in the nucleus.
It is noteworthy that the ARKP1 mRNA level was rapidly induced by the ABA treatment (Fig. 1a). Interestingly, the expression of ATHB7 and ATHB12 is increased after ABA treatment, but ATHB5 and ATHB6 are downregulated by external ABA (Soderman et al. 1994, 1996, 1999; Lee et al. 2001). So, the change of ATHBs protein level could also affect the transcription of these genes to construct a feedback in the signaling.
Furthermore, ABA-related phenotypic analysis indicated that downregulation of ARKP1 in Arabidopsis reduced ABA sensitivity in seed germination, postgermination, and drought tolerant behavior, while overexpression of ARKP1 increased the sensitivity of plants to ABA (Figs. 4 and 5). The appearance of three aspects of classic ABA phenotypes in ARKP1 transgenic or mutant plants may result from the regulation of some important upstream factors in the early stage of ABA signaling pathway. Recently, an important breakthrough was the identification of new ABA receptors named PYR/PYL/RCAR family proteins (Ma et al. 2009; Park et al. 2009). Thus, the major components of early ABA signaling pathway have been identified. ABA receptors (PYR/PYL/RCARs) bind with PP2Cs phosphatases to form coreceptor complex, and the receptor-phosphatase complex directly regulate the central SnRK2 protein kinase activity (Fujii and Zhu 2009). Thus, ARKP1 may interact with some negative regulators which localize in the nucleus, such as AHG1 and AHG3 (Yoshida et al. 2006; Nishimura et al. 2007).
In addition, the gene expression analysis (Fig. 6) revealed that the expression of ABA responsive genes, such as RAB18 and RD29A, was dramatically decreased in ARKP1 overexpression plants compared with the wild-type and T-DNA insertion arkp1 mutant plants (Lång and Palva 1992; Yamaguchi-Shinozaki and Shinozaki 1993; Lång et al. 1994). The RD29A promoter contains several DREs and one ABRE that could bind many abiotic responsive transcription factors to induce or reduce their expression (Uno et al. 2000; Sakuma et al. 2006a, b). Localized in the nucleus, ARKP1 may link some promoter regions of ABA responsive genes to regulate ABA signaling positively. Thus, ARKP1 is likely to play different roles in ABA signaling by acting as an F-box E3 ligase or a transcription activator. The expression of the positive transcription factor ABF3 was increased in ARKP1-overexpressing transgenic plants compared with the wild-type and arkp1 mutant plants, suggesting that ARKP1 plays a positive role in the upstream of ABF3. In conclusion, the F-box E3 ligase ARKP1 plays a positive role in ABA-mediated responses in Arabidopsis.
References
Bueso E, Rodriguez L, Lorenzo‐Orts L, Gonzalez‐Guzman M, Sayas E, Muñoz‐Bertomeu J, Ibañez C, Serrano R, Rodriguez PL (2014) The single‐subunit RING‐type E3 ubiquitin ligase RSL1 targets PYL4 and PYR1 ABA receptors in plasma membrane to modulate abscisic acid signaling. Plant J 80(6):1057–1071
Choi H-i, J-h H, J-o H, J-y K, Kim SY (2000) ABFs, a family of ABA-responsive element binding factors. J Biol Chem 275(3):1723–1730
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium‐mediated transformation ofArabidopsis thaliana. Plant J 16(6):735–743
Craig KL, Tyers M (1999) The F-box: a new motif for ubiquitin dependent proteolysis in cell cycle regulation and signal transduction. Prog Biophys Mol Biol 72(3):299–328
Dreher K, Callis J (2007) Ubiquitin, hormones and biotic stress in plants. Ann Bot 99(5):787–822
Edwards K, Johnstone C, Thompson C (1991) A simple and rapid method for the preparation of plant genomic DNA for PCR analysis. Nucleic Acids Res 19(6):1349
Fujii H, Zhu J-K (2009) Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc Natl Acad Sci 106(20):8380–8385
Gagne JM, Downes BP, Shiu S-H, Durski AM, Vierstra RD (2002) The F-box subunit of the SCF E3 complex is encoded by a diverse superfamily of genes in Arabidopsis. Proc Natl Acad Sci 99(17):11519–11524
Himmelbach A, Hoffmann T, Leube M, Hohener B, Grill E (2002) Homeodomain protein ATHB6 is a target of the protein phosphatase ABI1 and regulates hormone responses in Arabidopsis. EMBO J 21(12):3029–3038
Irigoyen ML, Iniesto E, Rodriguez L, Puga MI, Yanagawa Y, Pick E, Strickland E, Paz-Ares J, Wei N, De Jaeger G (2014) Targeted degradation of abscisic acid receptors is mediated by the ubiquitin ligase substrate adaptor DDA1 in Arabidopsis. Plant Cell Online 26(2):712–728
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6(13):3901
Koornneef M, Bentsink L, Hilhorst H (2002) Seed dormancy and germination. Curr Opin Plant Biol 5(1):33–36
Kraft E, Stone SL, Ma L, Su N, Gao Y, Lau O-S, Deng X-W, Callis J (2005) Genome analysis and functional characterization of the E2 and RING-type E3 ligase ubiquitination enzymes of Arabidopsis. Plant Physiol 139(4):1597–1611
Lång V, Palva ET (1992) The expression of a rab-related gene, rab18, is induced by abscisic acid during the cold acclimation process of Arabidopsis thaliana (L.) Heynh. Plant Mol Biol 20(5):951–962
Lång V, Mantyla E, Welin B, Sundberg B, Palva ET (1994) Alterations in Water Status, Endogenous Abscisic Acid Content, and Expression of rab18 Gene during the Development of Freezing Tolerance in Arabidopsis thaliana. Plant Physiol 104(4):1341–1349
Lechner E, Achard P, Vansiri A, Potuschak T, Genschik P (2006) F-box proteins everywhere. Curr Opin Plant Biol 9(6):631–638
Lechner E, Leonhardt N, Eisler H, Parmentier Y, Alioua M, Jacquet H, Leung J, Genschik P (2011) MATH/BTB CRL3 receptors target the homeodomain-leucine zipper ATHB6 to modulate abscisic acid signaling. Dev Cell 21(6):1116–1128
Lee YH, Oh HS, Cheon CI, Hwang IT, Kim YJ, Chun JY (2001) Structure and expression of the Arabidopsis thaliana homeobox gene Athb-12. Biochem Biophys Res Commun 284(1):133–141
Lee J-H, Yoon H-J, Terzaghi W, Martinez C, Dai M, Li J, Byun M-O, Deng XW (2010) DWA1 and DWA2, two Arabidopsis DWD protein components of CUL4-based E3 ligases, act together as negative regulators in ABA signal transduction. Plant Cell Online 22(6):1716–1732
Leung J, Merlot S, Giraudat J (1997) The Arabidopsis ABSCISIC ACID-INSENSITIVE2 (ABI2) and ABI1 genes encode homologous protein phosphatases 2C involved in abscisic acid signal transduction. Plant Cell Online 9(5):759–771
Lopez-Molina L, Chua N-H (2000) A null mutation in a bZIP factor confers ABA-insensitivity in Arabidopsis thaliana. Plant Cell Physiol 41(5):541–547
Lopez-Molina L, Mongrand S, Chua N-H (2001) A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc Natl Acad Sci 98(8):4782–4787
Lopez-Molina L, Mongrand S, Kinoshita N, Chua N-H (2003) AFP is a novel negative regulator of ABA signaling that promotes ABI5 protein degradation. Genes Dev 17(3):410–418
Ma Y, Szostkiewicz I, Korte A, Moes D, Yang Y, Christmann A, Grill E (2009) Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 324(5930):1064–1068
Moon J, Parry G, Estelle M (2004) The ubiquitin-proteasome pathway and plant development. Plant Cell 16(12):3181–3195
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15(3):473–497
Nishimura N, Yoshida T, Kitahata N, Asami T, Shinozaki K, Hirayama T (2007) ABA-Hypersensitive Germination1 encodes a protein phosphatase 2C, an essential component of abscisic acid signaling in Arabidopsis seed. Plant J 50(6):935–949
Park S-Y, Fung P, Nishimura N, Jensen DR, Fujii H, Zhao Y, Lumba S, Santiago J, Rodrigues A, Tsz-fung FC (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324(5930):1068–1071
Raab S, Drechsel G, Zarepour M, Hartung W, Koshiba T, Bittner F, Hoth S (2009) Identification of a novel E3 ubiquitin ligase that is required for suppression of premature senescence in Arabidopsis. Plant J 59(1):39–51
Risseeuw EP, Daskalchuk TE, Banks TW, Liu E, Cotelesage J, Hellmann H, Estelle M, Somers DE, Crosby WL (2003) Protein interaction analysis of SCF ubiquitin E3 ligase subunits from Arabidopsis. Plant J 34(6):753–767
Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006a) Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18(5):1292–1309
Sakuma Y, Maruyama K, Qin F, Osakabe Y, Shinozaki K, Yamaguchi-Shinozaki K (2006b) Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc Natl Acad Sci U S A 103(49):18822–18827
Schumann N, Navarro-Quezada A, Ullrich K, Kuhl C, Quint M (2011) Molecular evolution and selection patterns of plant F-box proteins with C-terminal kelch repeats. Plant Physiol 155(2):835–850
Seo K-I, Lee J-H, Nezames CD, Zhong S, Song E, Byun M-O, Deng XW (2014) ABD1 is an Arabidopsis DCAF substrate receptor for CUL4-DDB1–based E3 ligases that acts as a negative regulator of abscisic acid signaling. Plant Cell Online 26(2):695–711
Smalle J, Vierstra RD (2004) The ubiquitin 26S proteasome proteolytic pathway. Annu Rev Plant Biol 55:555–590
Soderman E, Mattsson J, Svenson M, Borkird C, Engstrom P (1994) Expression patterns of novel genes encoding homeodomain leucine-zipper proteins in Arabidopsis thaliana. Plant Mol Biol 26(1):145–154
Soderman E, Mattsson J, Engstrom P (1996) The Arabidopsis homeobox gene ATHB-7 is induced by water deficit and by abscisic acid. Plant J 10(2):375–381
Soderman E, Hjellstrom M, Fahleson J, Engstrom P (1999) The HD-Zip gene ATHB6 in Arabidopsis is expressed in developing leaves, roots and carpels and up-regulated by water deficit conditions. Plant Mol Biol 40(6):1073–1083
Stone SL, Hauksdóttir H, Troy A, Herschleb J, Kraft E, Callis J (2005) Functional analysis of the RING-type ubiquitin ligase family of Arabidopsis. Plant Physiol 137(1):13–30
Stone SL, Williams LA, Farmer LM, Vierstra RD, Callis J (2006) KEEP ON GOING, a RING E3 ligase essential for Arabidopsis growth and development, is involved in abscisic acid signaling. Plant Cell Online 18(12):3415–3428
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci U S A 97(21):11632–11637
Vierstra RD (2009) The ubiquitin–26S proteasome system at the nexus of plant biology. Nat Rev Mol Cell Biol 10(6):385–397
Xiong L, Schumaker KS, Zhu JK (2002) Cell signaling during cold, drought, and salt stress. Plant Cell 14(suppl 1):S165–183
Yamaguchi-Shinozaki K, Shinozaki K (1993) Characterization of the expression of a desiccation-responsive rd29 gene of Arabidopsis thaliana and analysis of its promoter in transgenic plants. Mol Gen Genet MGG 236(2-3):331–340
Yoo S-D, Cho Y-H, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2(7):1565–1572
Yoshida T, Nishimura N, Kitahata N, Kuromori T, Ito T, Asami T, Shinozaki K, Hirayama T (2006) ABA-hypersensitive germination3 encodes a protein phosphatase 2C (AtPP2CA) that strongly regulates abscisic acid signaling during germination among Arabidopsis protein phosphatase 2Cs. Plant Physiol 140(1):115–126
Zhang Ye XW, Li Z, Deng XW, Wu W, Xue Y (2008) F-box protein DOR functions as a novel inhibitory factor for abscisic acid-induced stomatal closure under drought stress in Arabidopsis. Plant Physiol 148(4):2121–2133
Zhang X, Garreton V, Chua N-H (2005) The AIP2 E3 ligase acts as a novel negative regulator of ABA signaling by promoting ABI3 degradation. Genes Dev 19(13):1532–1543
Zhu J-K (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247
Acknowledgments
This work was supported by grants from the National 973 Project (2015CB755700) and National Natural Science Foundation of China (NSFC 31171586, 31271758, and 31300996).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplementary Table S1
(DOCX 121 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Liu, Z., Wang, J. et al. The Arabidopsis Kelch Repeat F-box E3 Ligase ARKP1 Plays a Positive Role for the Regulation of Abscisic Acid Signaling. Plant Mol Biol Rep 34, 582–591 (2016). https://doi.org/10.1007/s11105-015-0942-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-015-0942-2