Abstract
Purpose
The effects of trees on soil nematode communities are related to nutrient cycles in forest ecosystems. We conducted greenhouse pot experiments to determine the effects of a single tree species for each of coniferous and broad-leaved tree on soil nematodes.
Methods
Soils were collected from a coniferous plantation and broad-leaved forests. Seedlings of a coniferous tree (Cryptomeria japonica) and a broad-leaved tree (Quercus serrata) were planted in soils derived from each species. After 11 months, seedling biomass, soil properties, and ectomycorrhizal fungal colonization of Q. serrata were measured. Soil nematodes were morphologically identified to the genus/family level and differentiated by community and trophic composition.
Results
C. japonica root biomass was significantly higher than that of Q. serrata regardless of the soil and nematode community structures were significantly different between the species. The fungal: bacterial ratio and density of fungivorous nematodes were significantly higher in broad-leaved soils. Herbivorous nematodes increased significantly in C. japonica seedlings grown in broad-leaved soils. Structural equation modeling indicated that soil origin and tree species directly regulated nematode trophic compositions.
Conclusion
Our findings suggest that tree species modify soil micro-food webs by affecting microbial abundance and nematode trophic composition. Specifically, C. japonica, with a larger root biomass, increased the number of herbivorous nematodes, whereas Q. serrata, with ectomycorrhizal fungal symbiosis, increased the number of fungivorous nematodes. Thus, tree species are tightly involved in shaping nematode communities in forest ecosystems through root traits and mycorrhizal types.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
In the rhizosphere, root exudates and sloughing root cells provide resources that support hotspots of microbes and soil fauna (Bonkowski et al. 2009). Lower trophic level animals such as herbivores and bacterivores are more responsive to changes in plant species diversity and composition than organisms at higher trophic levels (De Deyn et al. 2004). Consumers at higher trophic levels may have access to a larger variety of prey than those at lower trophic levels (Scherber et al. 2010). Extramatrical fungal hyphae are vulnerable to grazing by soil fauna (Klironomos and Hart 2001). Thus, the bottom-up control of plant diversity may be weaker at higher trophic levels. Bacteria and fungi, together with the soil fauna that feed on them, are organized into micro-food webs, and their diverse interactions significantly affect plant growth (Berg et al. 2014; Fierer 2017).
Nematodes are a major group of soil microfauna and the most abundant animals on Earth (van den Hoogen et al. 2019). They play crucial roles in ecosystem processes, such as improving soil physical properties, participating in carbon and nitrogen cycling by feeding on bacteria and fungi (Ingham et al. 1985), and maintaining ecosystem health by occupying key positions in the soil food cycle (Bongers and Bongers 1998; Ferris 2010; Zhang et al. 2017; Thakur and Geisen 2019). In contrast, soil nematode communities are affected by subtle changes in both abiotic and biotic factors (Neher et al. 2005). For example, soil pH and organic carbon (C) are factors that determine the spatial distribution of soil nematodes in forest soils (Kitagami et al. 2020; Liu et al. 2019). In terms of biotic factors, root traits influence soil nematode abundance and community structure (De Deyn et al. 2008; Wilschut et al. 2019; Zhang et al. 2020, 2022). However, most observations have been based on herbaceous plants in agricultural soils, and there is a lack of knowledge regarding how abiotic and biotic factors affect the nematode community structure in woody plants in forest soils. Tree species influence soil biological, physical, and chemical properties through different root qualities (De Schrijver et al. 2012; Keith et al. 2009); yet our understanding of root-mediated effects on the composition and diversity of soil biota is still limited. In this study, we identified the factors that determine the structure of soil nematode communities by comparing the community patterns of these organisms in coniferous and broad-leaved forest soils.
Coniferous and broad-leaved trees exhibit significantly different effects on nematode trophic groups (Kondratow et al. 2019; Liu et al. 2023). Differences in litter quality among tree species have different effects on soil chemical properties (such as pH) and microbial communities (Sawada et al. 2021; Tanikawa et al. 2023), which, in turn, alter the trophic composition of nematodes (Cesarz et al. 2013). Moreover, root-associated mycorrhizal types, either arbuscular or ectomycorrhizal, are related to nematode communities in forest soils (Kudrin et al. 2021; Liu et al. 2023). Mycorrhizal associations shape activity and biomass of soil bacterial and fungal communities which are food resources for nematodes, i.e., arbuscular mycorrhizal trees showed higher bacterial growth whereas ectomycorrhizal trees showed higher fungal growth (Heděnec et al. 2020). Thus, tree species identities, through root traits and mycorrhizal types, can shape specific nematode communities; therefore, they are tightly involved in shaping nematode communities in forest ecosystems.
Japanese cedar, Cryptomeria japonica (Cupressaceae), is afforested in a large area as much as 18% of Japan (Forest Agency of Japan 2022). Recently, monocultural forests have been converted to coniferous-broadleaf mixed forests to maintain ecosystem functioning and services (Cavard et al. 2011). Alternatively, Quercus serrata (Fagaceae) is one of the candidate representative broadleaf trees for planting, especially in warm temperate climatic zones (Forest Agency of Japan 2022). In ecological relevance, the fine roots of C. japonica and Q serrata are associated with arbuscular mycorrhizal and ectomycorrhizal fungi, respectively (Matsuda et al. 2013, 2021). Such a different mycorrhizal type affects soil ecological processes and functions (Tedersoo et al. 2020).
This study aimed to determine the effects of a single tree species for each of conifer and broad-leaved tree on micro-food webs of soil nematodes communities by a pot experiment in a greenhouse. Seedlings of coniferous (C. japonica) and broad-leaved (Q. serrata) trees, both of which are the predominant species in Japan (Forest Agency of Japan 2022), were grown in monospecific pots with soils derived from either tree species. This monoculture of the pot experiment in a greenhouse could be useful for focusing on the effects of tree species and avoiding interference from other factors such as microclimate, topography, tree age, or the admixture of other plant species that generally exist in field experiments (Stefanowicz et al. 2021). We hypothesized that the abundance, community, and trophic composition of nematodes would be synergistically affected by the root traits and soil origin of tree species. Moreover, we hypothesized that nematodes were affected by soil biotic (i.e., microbes and root traits) and abiotic factors (i.e., pH and C/N), which are characterized by each tree species.
Materials and methods
Experimental set-up
In November 2019, we collected approximately 150 L each of soil layer from 0 to 20 cm in depth from a 50-year-old coniferous C. japonica plantation and broad-leaved Q. serrata forest in Mie Prefectural Forest Institute, Japan (34°65′N, 136°37′E). Soil from each site was classified as Brown Forest Soil (Forest Soil Division 1976) or Inceptisol (Soil Survey Staff 2014). Soil pH at the collecting points was slightly acidic, 5.62 and 4.17, at the coniferous plantation and broad-leaved forest, respectively. Annual precipitation and annual mean temperature in 2018, recorded at the nearest weather station (Tsu, 20 km from the forest), were 1630 mm and 16.9 °C.
Coarse roots, litter, and stones in the soil samples were discarded using a 2-cm mesh sieve. The sieved soil was gently homogenized before filling up Wagner pots (NF-5, top φ17 cm × bottom φ16 cm × height 19 cm) with 2 L of soils. After immersion in running water for 24 h, three seeds of either C. japonica or Q. serrata were sown in individual pots. The pots were placed in a greenhouse at 25 °C under a natural light condition and were supplied with distilled water for the first 3 months every day. For the next 8 months, 450 ml of distilled water was added to the pots every 3 days to maintain the water content at more than 10%. In total, 96 pots were established by combining two soil origins (C. japonica and Q. serrata) × two tree species (C. japonica and Q. serrata) × 24 replicates. Thus, four soil and tree species treatment combinations were established (Fig. S1): C. japonica soil with C. japonica seedlings (CC), Q. serrata soil with C. japonica seedlings (QC), C. japonica soil with Q. serrata seedlings (CQ), Q. serrata soil with Q. serrata seedlings (QQ).
Plant biomass, root traits, and mycorrhizal colonization
After 11 months, we carefully harvested the C. japonica and Q. serrata seedlings from each pot. The seedlings were used to observe root morphology and to measure shoot and root biomass. The soil remaining in the pots was used to measure environmental conditions and nematode extraction. The harvested seedlings were divided into shoot and root systems using pruning shears. Root systems in the soil blocks were washed in a 2-mm-mesh sieve under running tap water to remove adhering soil particles. The root systems were divided into two categories based on diameter: coarse roots ≥2 mm and fine roots ≤2 mm. The shoots and coarse and fine roots were dried at 70 °C for 2 weeks and then weighed. Images of all individual fine roots were acquired using a scanner (500 dpi; GT-9800, EPSON, Nagano, Japan). The average diameter and length of all individual fine roots were obtained using the image analysis software IJ Rhizo (Pierret et al. 2013). From these data, the specific root length (SRL), that is, the root length per unit dry weight (expressed as m g−1), was calculated. The SRL represents the cost-benefit relationship of fine root production (Eissenstat and Yanai 1997). A higher SRL corresponds to thinner roots and a more frequent branching pattern, indicating efficient nutrient uptake (Ostonen et al. 2007).
Quercus serrata seedlings per pot were transferred onto 9-cm Petri dishes to count the number of root tips under a stereomicroscope (SZX16, Olympus, Tokyo, Japan). Root tips were defined as ectomycorrhizal (ECM) roots when fungal sheaths covered them entirely, with no cracks or shrinkage (Matsuda et al. 2013). We defined root tips with root hairs without ECM fungal colonization as non-mycorrhizal (non-ECM).
Soil environmental conditions
Soil pH (H2O) was determined using air-dried fine soil (soil-to-distilled water ratio, 1:2.5) using a pH meter (MP220; Mettler-Toledo, Greifensee, Switzerland). A subset of the pot soil was oven-dried at 105 °C for 24 h and ground for the analysis of total C and N concentrations measured with an elemental analyzer (vario EL cube; Elementar Japan Ltd., Yokohama, Japan).
Bacterial and fungal abundances quantified by real-time PCR
Total DNA was extracted from 250 mg of moist soil using a Fast DNA SPIN Kit for Soil (Q-Biogene, Ohio, USA) following the manufacturer’s instructions. Subsequently, all DNAs were quantified using QuantiFluor™ dsDNA System (Promega, Madison, USA) and DNA concentration for each extraction was standardized to 20 ng μL−1. The relative abundances of bacterial and fungal communities of each pot were assessed for each DNA extract by qPCR using a StepOne™ Real-Time PCR system (Applied Biosystems, Waltham, USA). For bacteria, a fragment of the 16S ribosomal ribonucleic acid (rRNA) gene was amplified using the Eub338/Eub518 primer pair (Fierer et al. 2005). For fungi, a fragment of the 18S rRNA gene was amplified using FR1/FF390 primers (Chemidlin Prévost-Bouré et al. 2011). Each 20 μL reaction contained 10 μL THUNDERBIRD™ Next SYBR® qPCR Mix (TOYOBO, Osaka, Japan), 0.4 μL per primer (10 μM), 2 μL template DNA (2 ng), and 7.2 μL H2O. All samples were analyzed in duplicate using PCR strip tubes (Axygen, Thermo Fisher Scientific Inc., Waltham, USA). PCR amplification condition was set as follows: for bacteria; 95 °C for 2 min followed by 40 cycles of 95 °C for 15 s, 60 °C for 60 s (two-step), and for fungi; 95 °C for 15 min followed by 40 cycles of 95 °C for 60 s, 58 °C for 30 s, and 72 °C for 60 s. After amplification, melting curve and gel electrophoresis were performed to confirm that the amplified products were of the appropriate size. The qPCR efficiencies were 156.923% (R2 = 0.947) for bacteria, and 100.648% (R2 = 0.977) for fungi.
Standards for the qPCR assays were generated by PCR, amplifying each gene of interest from the genomic DNA of Escherichia coli clones containing plasmids that carried the respective PCR target fragment as an insert of a soil bacterial (unknown taxon) and fungal (Botrytis cinerea 20,201,009) strain using the primers previously described for each kingdom. The PCR products were confirmed on an agarose gel and cloned using a TA-Enhancer Cloning Kit (Nippon Gene, Toyama, Japan) following the manufacturer’s instructions. Positive clones were further amplified using KOD One PCR Master Mix (TOYOBO, Osaka, Japan) and primers previously described for each kingdom. The thermal cycle program comprised 25 cycles of 98 °C for 10 sec, 55 °C for 5 sec, and 72 °C for 10 sec. Serial dilutions of each plasmid DNA were prepared, and bacterial and fungal gene copy numbers were calculated using a regression equation for each assay, relating the cycle threshold (Ct) value to the known number of copies in the standards, as described (Qian et al. 2007). The copy numbers of bacterial 16S rRNA and fungal 18S rRNA genes were expressed as copy number log 10 transformed per gram of dry weight soil. The fungi/bacteria (F/B) ratio was calculated for each pot based on the number of copies of the 16S rRNA or 18S rRNA genes log-transformed.
Nematode extraction and observation
Nematodes were extracted from 100 g of fresh soil samples from each pot by decanting and sieving, following Cobb’s method (van Bezooijen 2006). Each soil sample was placed in a 2 L beaker and mixed with tap water. The water suspensions were stirred with a plastic spatula and decanted immediately into another 2 L beaker covered with a 1 mm mesh sieve (15 cm in diameter) to screen out stones and roots. The water suspensions were further mixed and decanted through successive 75 μm and 25 μm mesh sieves (15 cm in diameter). Finally, the remnants on each mesh sieve were carefully collected and pooled into a 50 mL beaker. The remaining sample matter was transferred into individual funnels (9 cm in diameter) covered with a paper-cleaning cloth (Pro-wipe S200; Elleair, Tokyo, Japan) and further processed to extract nematodes using the Baermann funnel technique for 24 h at 25 °C (Baermann 1917). The extracted nematodes were transferred into a Petri dish filled with distilled water and counted under a stereomicroscope to estimate their density (per gram of dried soil). Nematodes were then transferred to glass slides with vertical lines, and up to the first 100 individuals encountered in each sample (some samples contained as few as 80 nematodes) were identified at 100–400× magnification under a compound light microscope (BX53, Olympus, Tokyo, Japan). Nematodes were identified to the genus/family level following Bongers (1988) and divided into four trophic groups (bacterivorous, fungivorous, herbivorous, and predatory-omnivorous) based on the structural characteristics of their oral parts (Yeates et al. 1993).
Data and statistical analysis
To compare nematode densities among the treatments, the Steel–Dwass test was performed using the NSM3 package (Schneider et al. 2021) in R software version 4.3.1 (R Development Core Team 2023). We generated a non-metric multidimensional scaling (NMDS) scatter plot based on the Chao dissimilarity index using nematode taxonomic data, by generating 20 permutations using the meataMDS function in the vegan package (Oksanen et al. 2022) in R. In addition, permutational multivariate analysis of variance (PERMANOVA) was performed to determine whether the nematode community structures differed among the treatments; 9999 permutations were conducted using the adonis function in vegan. The effects of soil, plant, and microbial properties on nematode community structure were determined by conducting 9999 permutations using the envfit function in vegan. Indicator species analysis (Dufrêne and Legendre 1997) was performed to determine whether any characteristic nematode genera were present among the treatments. This analysis was performed using indicator values for each nematode genus, and statistical significance was tested using a Monte Carlo test (9999 permutations). These analyses were performed using the indicspecies package (Cáceres and Legendre 2009) in R.
A structural equation model was constructed to provide a mechanistic understanding of how soil origin and tree species affect the trophic groups of nematodes. Structural equation modeling is a multivariate statistical method that allows for hypothesis testing of complex path-relation networks under any conditions (Grace et al. 2007). The full initial model (Fig. S2) was constructed a priori based on the hypotheses that (1) soil origins and tree species affected soil abiotic factors, root biomass, F/B ratio, and the abundance of nematode trophic groups; (2) soil abiotic factors affected root biomass, F/B ratio, and the abundance of nematode trophic groups; (3) F/B ratio affected bacterivorous and fungivorous nematodes; (4) root biomass affected herbivorous nematodes; (5) predatory and omnivorous nematodes feed on bacterivorous, fungivorous, and herbivorous ones. We added double headed arrows specifying residual covariance (Bacterivores < − > Fungivores, Bacterivores < − > Herbivores, Fungivores < − > Herbivores) because the prey-predator interaction was fundamentally unknown. The model was constructed using data from all pots (n = 96). The analysis was performed using the piecewise SEM package (Lefcheck 2016) in R. In this analysis, soil origin and tree species were indicated as categorical explanatory variables, that is, soil origin: C. japonica (1) and Q. serrata (2) and tree species: C. japonica (1) and Q. serrata (2). Principal component analysis (PCA) was performed to reduce the dimensions of the soil abiotic factors, i.e., soil pH and C/N. The first principal component of soil properties (Soil PC1) was negatively correlated with soil pH and the C/N ratio (Fig. S3). First, all non-significant paths (P > 0.05) were removed from the initial model. We then compared several models that incorporated all or part of the significant paths. The overall fit of the piecewise SEM was assessed using Shipley’s test of d-separation, which combines the P-values associated with each of the k independence claims in Fisher’s C statistics (Shipley 2009). For all analyses, the significance level was set at P < 0.05, unless otherwise stated.
Results
Plant biomass, root traits, and mycorrhizal colonization
Shoot biomass values were significantly greater with CC (24.2 ± 0.3 g) and QC than CQ and QQ (Table 1, Steel–Dwass, P < 0.05), but root biomass values were significantly greater with CC (5.7 ± 0.3 g), CQ and QC than QQ (Table 1, Steel–Dwass, P < 0.05). Fine root biomass values were significantly greater with CC (5.1 ± 0.2 g) and QC than CQ and QQ (Table 1, Steel–Dwass, P < 0.05). SRL values were significantly greater with CQ and QQ than CC and QC (Table 1, Steel–Dwass, P < 0.05). In the CQ and QQ treatments, ECM formation was confirmed in all the seedlings. The number of Q. serrata root tips was not significantly different between CQ (241 ± 5) and QQ (241 ± 6) (Wilcoxon rank sum test, W = 322, P = 0.48), and ECM root frequencies were not significantly different between CQ (68.9 ± 2.7%) and QQ (62.8 ± 2.8%) (W = 360, P = 0.14), respectively.
Soil environmental conditions
The soil pH of each pot treatment ranged from 4.18 ± 0.03 to 5.80 ± 0.03, and CC was significantly the highest among the four treatments (Table 1, Steel–Dwass, P < 0.05). The total C was significantly higher at CC and CQ than at other treatments (Table 1, Steel–Dwass, P < 0.05), and the total N was significantly higher at CC, CQ, and QC than at QQ (Table 1, Steel-Dwass, P < 0.05). The C/ N ratio ranged from 16.4 ± 0.7–20.5 ± 0.2 and CQ had the highest value (Table 1, Steel–Dwass, P < 0.05).
Bacterial and fungal DNA abundance
Bacterial gene copy numbers log 10 transformed per gram dry weight soil were significantly greater at CC, QC, and QQ than that at CQ. Fungal gene copy numbers were significantly the highest in QQ (Table 1, Steel–Dwass, P < 0.05). The F/B ratio was significantly the highest in QQ (Table 1, Steel–Dwass, P < 0.05).
Nematode abundance of each trophic group and community structure
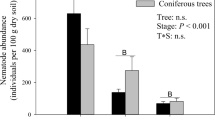
The abundance of total nematodes (number per 100 g dry soil) in pots ranged from 212 ± 28 to 620 ± 64 and the number was significantly highest in QC (Fig. 1, Steel–Dwass, P < 0.05). The abundance of herbivorous nematodes was significantly highest in QC (Fig. 1, Steel–Dwass, P < 0.05). The abundance of fungivorous nematodes were significantly higher in QQ and QC than the others (Fig. 1, Steel–Dwass, P < 0.05). Bacterivorous and predatory-omnivorous nematodes were not significantly different among pots (Fig. 1, Steel–Dwass, P > 0.05).
Number of total nematodes and four trophic groups per 100-g dry soil at four different treatments. Columns are means (n = 24) with vertical bars of standard errors, and different characters upper to the bars indicated significant differences in nematode numbers among the treatments (Steel–Dwass test, P < 0.05). Treatment codes show Cryptomeria japonica soil with C. japonica seedlings (CC); Quercus serrata soil with C. japonica seedlings (QC); C. japonica soil with Q. serrata seedlings (CQ); Q. serrata soil with Q. serrata seedlings (QQ)
The NMDS scatter plot shows that the nematode community structures were significantly clustered among the four pot treatments (Fig. 2; stress value = 0.223, PERMANOVA, permutation 9999, P < 0.001, R2 = 0.70). Among the environmental factors measured, soil origin, tree species, shoot biomass, root biomass, fine roots, SRL, soil pH, soil C, soil N, C/N ratio, bacterial and fungal gene copy numbers, and the F/B ratio significantly influenced the nematode community structure. Across all treatments, soil origin accounted for the most variation (R2 = 0.76) in the nematode communities. Moreover, tree species also accounted for the variation in nematode communities (R2 = 0.18, among the treatments).
Non-metric multidimensional scaling scatterplot of Chao dissimilarity based on the abundance of nematode taxa derived from four treatments. Stress value = 0.223. Nematode community structures were clustered significantly into C. japonica soil with Cryptomeria. japonica seedlings (CC); Quercus serrata soil with C. japonica seedlings (QC); C. japonica soil with Q. serrata seedlings (CQ); Q. serrata soil with Q. serrata seedlings (QQ) (PERMANOVA, P < 0.001, R2 = 0.70). Soil origins, Tree species, Shoot biomass (SB), Root biomass (RB), Fine root biomass (FR), Specific root length (SRL), soil pH, C, N, C/N ratio, Bacterial and fungal DNA copy number and Fungi/Bacteria (F/B) ratio showed significant influences on nematode communities (Envfit, permutation 9999 times, P < 0.05)
Indicator species analysis was performed for each pot treatment (Table 2). Eleven taxa (Acrobeles, Acrobeloides, Aphanolaimus, Criconematidae, Diphtherophora, Ecphyadophora, Heteroderidae, Helicotylenchus, Plectus, Teratocephalus and Wilsonema) were selected as indicator taxa for CC (P < 0.05), four taxa (Miconchus, Monhystera, Rhabditidae and Rhabdolaimus) for CQ, Trichodoridae and Tylencholaimellus for QC, and four taxa (Cephalobus, Ditylenchus, Filenchus/Tylenchus and Mylonchulus) for QQ.
Structural equation modeling (SEM) analysis
The final structural equation model adequately fitted the obtained dataset. The model explained the effects of soil origin and tree species on soil PC1, root biomass, F/B ratio, and nematode trophic group abundance (Fig. 3; Fisher’s C = 27.844, df = 32, P = 0.677, AICc = 73.844). Soil PC1 was significantly positively correlated with Q. serrata soil. The F/B ratio was significantly positively correlated with Q. serrata soil and trees, whereas root biomass was significantly positively correlated with C. japonica soil and trees. The abundance of fungivorous nematodes was significantly positively correlated with Q. serrata soils but not with the F/B ratio. There was a positive covariance between the nematode abundance of bacterivores and fungivores. The abundance of herbivorous nematodes was significantly and positively correlated with Q. serrata soils and C. japonica trees, but not with root biomass. Herbivorous nematode abundance had a significant positive relationship with predator-omnivore abundance.
Structural equation model of effects of soil origins and tree species on soil abiotic factors, root biomass, F/B ratio and the abundance of nematode trophic groups (Fisher’s C = 27.844, df = 32, P = 0.677, AICc = 73.844). R squared values for endogenous variables are noted at the bottom of each factor. Numbers on arrows are standardized path coefficients. Solid and dashed lines represent significant and non-significant pathways, respectively. Soil origins and tree species were indicated as a categorical explanatory variable, i.e. soil origins; Cryptomeria japonica (1) and Quercus serrata (2), tree species; C. japonica (1) and Q. serrata (2). Principal component analysis (PCA) was performed to reduce the dimensions of soil abiotic factors, i.e., soil pH and C/N. The first principal component of soil properties (Soil PC1) was negatively correlated with soil pH and C/N ratio (see Fig. S3). Bact, abundance of bacterivores; Fung, abundance of fungivores; Herb, abundance of herbivores; Pred&Omni, abundance of predators and omnivores
Discussion
Effects of tree species identity and soil origin on nematode community structures
Nematode community structures were significantly different between the tree species examined (Fig. 2), which agrees with the findings of previous studies (Cesarz et al. 2013; Gilarte et al. 2021). Moreover, the nematode community structure with trees is apparently shifted and altered into different ones without trees (Fig. S4). Tree species identity is a major factor affecting root-associated microbial communities (Sawada et al. 2021; Tanikawa et al. 2023) and nematode diversity (Li et al. 2023). Since nematodes are involved in nutrient cycling by feeding on bacteria, fungi (Ingham et al. 1985) and roots (Wilschut and Geisen 2021), nematode communities are possibly tightly connected with microbial and root trait responses driven by tree species. For example, finer root systems enhance the complexity of nematode communities by increasing the relative abundance of high-trophic-level omnivores and predators (Zhang et al. 2022). Tree species identity can affect soil fertility by regulating soil nematode communities in coniferous and broad-leaved forests (Liu et al. 2023). Moreover, the nematode community structures differed significantly between soil origins obtained from either coniferous C. japonica or broad-leaved Q. serrata forests (Fig. 2). Previous studies have reported that nematode community structures are affected by soil origin (e.g., Richter et al. 2023). In the present study, nematode community structures were significantly coordinated by several proximate factors, particularly soil pH and C/N. These are generally related to both abiotic and biotic factors, that is, resource quality and quantity, which may indirectly influence nematode community structures (Kitagami et al. 2020; Liu et al. 2019). From our findings, the more fungal biomass present in Q. serrata forest soils, which can influence nematode community structures by changing fungivorous nematode abundance. Thus, nematode community structures are shaped by cascading effects of tree species identities on soil properties (e.g., microbial biomass) through root traits.
Direct and indirect effects of tree species identity and soil origin on nematode trophic compositions
The abundance of nematodes was significantly the highest in QC among all treatments (Fig. 1). Moreover, nematode densities with trees tended to be higher than those without trees (Fig. S5). Root biomass is well documented to have positive effects on nematode densities, particularly on herbivorous ones (Keith et al. 2009; Salamon and Wolters 2009). The proportion of herbivorous nematodes in the trophic composition was higher than that in other treatments. Among these, Trichodoridae was the most dominant taxon (Table 2), which could be naturally maintained at a low density in Q. serrata forest soils. Members of this taxon may have possibly increased due to a bottom-up effect of the increase in C. japonica fine roots in the QC treatment. This result suggests that Trichodoridae fed on C. japonica roots, because some species Trichodoridae are frequently found in various tree rhizosphere species, including C. japonica (Mamiya 1967). Under low plant diversity conditions, specialist herbivorous nematodes can play an important role by contributing to negative plant–soil feedback toward a certain dominant plant species (Wilschut and Geisen 2021). In such a situation, the living fine root tissue is consumed by herbivorous nematodes, leading to the deterioration of the absorptive function of root systems (Maboreke et al. 2017; Saucet et al. 2016). There was no significant difference in the shoot and root biomass of C. japonica grown in the soils of either tree species. This result showed that damage by Trichodoridae had a negligible effect on host growth and there was no negative feedback to the host plant. Regarding root traits, a low root tissue density (i.e., cells are soft and easily penetrate the nematode stylet) resulted in increased herbivorous nematode density (de Deyn et al. 2008; Wilschut and Geisen 2021), and herbivorous nematode abundance also increased in C. japonica, a tree with low root tissue density (Table 2). The diameters of fine roots in coniferous trees, including C. japonica were greater than those in broad-leaved trees (Wada et al. 2019); therefore, feeding area for herbivorous nematodes was large, which may have led to significant increments. Feeding by herbivorous nematodes also indirectly benefit plant performance by inducing increased plant exudation that stimulates microbial activity, enhancing N and P availability (Gebremikael et al. 2016). Other trophic groups of nematodes are distributed across the entire soil and rhizosphere food web (Bonkowski et al. 2009), and structure the rhizobiome, with potential positive feedback to plant performance (Thakur and Geisen 2019).
Soil origin exhibited significant effects on fungivore abundance, which was higher under Q. serrata soils than under C. japonica soils (Fig. 1), which was possibly due to intimate mycorrhizal associations between Q. serrata and ECM fungi (Matsuda et al. 2013). In the SEM analysis (Fig. 3), Q. serrata soils and their tree species positively affected the F/B ratio, and more than half of the root tips of Q. serrata seedlings showed ECM formation in both soil origins, which may contribute to fungal biomass. Although a significant path from F/B to fungivorous nematodes was not selected, the Q. serrata soil may have contained many potential food resources for fungivorous nematodes. In support of this idea, girdling of Picea obovata stems decreased the abundance of fungivorous nematodes by decreasing the amount of extramatrical mycelia derived from ECM fungi, which are the food resource of fungivorous nematodes (Kudrin et al. 2021). The extramatrical hyphae of ECM fungi extending into soils may provide a food resource, thus improving the habitat quality of fungivorous nematodes (Kitagami and Matsuda 2020). The community structures of bacteria, fungi, and nematodes varied between the tree species and mycorrhizal symbionts, suggesting adaptation of microbial communities. Easily degradable substrates can be provided under trees associated with AM fungi, and recalcitrant substrates associated with ECM fungi (Eagar et al. 2022; Heděnec et al. 2020). Fungal community compositions differed between C. japonica plantations and natural forests in soils; for example, saprophytic, endophytic, and plant pathogenic fungi dominated in C. japonica plantations, whereas ECM fungi were predominant in natural forests (Sawada et al. 2021). The fungivorous nematode Aphelenchoididae was dominant in all treatments, suggesting that the nematodes fed on saprophytic or ECM fungi, regardless of the fungal ecological type. These nematode groups have a wide range of food resources because they can proliferate by feeding on saprophytic or ECM fungi (Kitagami and Matsuda 2022; Ruess et al. 2000). In the indicator species analysis, the fungivorous Ditylenchus and Filenchus/Tylenchus were selected as significant indicator species in QQ. ECM fungi may be suitable food sources for the reproduction of these nematodes. Selective grazing by fungivorous nematodes may facilitate the compensation of fungal growth, inducing to initial decomposition (Mamilov and Dilly 2002). Bacterivorous and fungivorous nematodes can influence microbe-mediated ecosystem processes by affecting microbial population size and traits (Thakur and Geisen 2019). Thus, fungivorous nematode abundance is influenced by mycorrhizal types as well as fungal ecological types in forest ecosystems.
An increase in the abundance of herbivorous, bacterivorous, and fungivorous nematodes, which are lower trophic groups, increases the abundance of predatory and omnivorous nematodes, which are higher trophic groups (Kitagami and Matsuda 2020; Zhang et al. 2020). In the SEM analysis, there was a tight connection in the abundance of nematode trophic groups between herbivorous and predatory-omnivorous nematodes. This is possibly explained by the feeding relationships between these two groups and suggests that predatory-omnivorous nematodes feed specifically on herbivorous nematodes. In fact, field experiments and molecular gut analyses have suggested that some predatory-omnivorous nematodes feed on herbivorous nematodes (Bilgrami et al. 2008; Cabos et al. 2013). Herbivorous nematodes are poorly motile and most of the ones recorded in this study are ectoparasitic, i.e., prey on epidermal and cortical root cells (Wyss 1997), which may have made it easier for predatory-omnivorous nematodes to approach and feed on herbivorous nematodes.
Conclusion
Our study revealed that tree species identity and soil origin synergistically influence nematode communities. Herbivorous nematodes significantly increased at C. japonica tree species in Q. serrata forest soils. Fungivorous nematodes were significantly greater in Q. serrata soils than in C. japonica soils because of the mycorrhizal associations between Q. serrata and ECM fungi. Moreover, there was a covariation between the abundance of herbivorous and predatory-omnivorous nematodes. Our findings suggest that tree species identity modifies soil micro-food webs by affecting microbial abundance and nematode trophic composition. However, our results do not necessarily reflect community patterns of soil organisms in field environments because the present experiments were performed in a controlled greenhouse using the early stages of tree seedlings. Nevertheless, our results will help us better understand the effects of tree species identity on belowground biodiversity and functioning. Further research integrating multi-dimensional root traits and manipulating tree symbiont partners is needed to generalize of multi-trophic interactions among trees, microbes, and soil animals. Besides, the monitoring soil communities over time after being converted to coniferous-broadleaf mixed forests would provide insights into the structure and stability of soil micro-food webs.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Baermann G (1917) Eine einfache Methode zur Auffindung von Ancylostomum (Nematoden) larven in Erdproben. Geneeskd Tijdschr Ned Indie 57:131–137
Berg G, Grube M, Schloter M, Smalla K (2014) Unraveling the plant microbiome: looking back and future perspectives. Front Microbiol 5:1–7. https://doi.org/10.3389/fmicb.2014.00148
Bilgrami AL, Brey C, Gaugler R (2008) First field release of a predatory nematode, Mononchoides gaugleri (Nematoda: Diplogastrida), to control plant-parasitic nematodes. Nematology 10:143–146. https://doi.org/10.1163/156854108783360177
Bonkowski M, Villenave C, Griffiths B (2009) Rhizosphere fauna: the functional and structural diversity of intimate interactions of soil fauna with plant roots. Plant Soil 321:213–233. https://doi.org/10.1007/s11104-009-0013-2
Bongers T (1988) The nematodes of the Netherlands. The nematodes of the Netherlands. KNNV Publishing, Zeist
Bongers T, Bongers M (1998) Functional diversity of nematodes. Appl Soil Ecol 10:239–251. https://doi.org/10.1016/S0929-1393(98)00123-1
Cabos RYM, Wang KH, Sipes BS, Heller WP, Matsumoto TK (2013) Detection of plant parasitic nematode DNA in the gut of predatory and omnivorous nematodes. Nematropica 43:44e48
Cáceres MD, Legendre P (2009) Associations between species and groups of sites: indices and statistical inference. Ecology 90:3566–3574. https://doi.org/10.1890/08-1823.1
Cesarz S, Ruess L, Jacob M, Jacob A, Schaefer M, Scheu S (2013) Tree species diversity versus tree species identity: driving forces in structuring forest food webs as indicated by soil nematodes. Soil Biol Biochem 62:36–45. https://doi.org/10.1016/j.soilbio.2013.02.020
Cavard X, Macdonald SE, Bergeron Y, Chen HY (2011) Importance of mixedwoods for biodiversity conservation: evidence for understory plants, songbirds, soil fauna, and ectomycorrhizae in northern forests. Environ Rev 19:142–161. https://doi.org/10.1139/a11-004
Chemidlin Prévost-Bouré N, Christen R, Dequiedt S, Mougel C, Leliévre M, Jolivet C, Shahbazkia HR, Guillou L, Arrouays D, Ranjard L (2011) Validation and application of a PCR primer set to quantify fungal communities in the soil environment by real-time quantitative PCR. PLoS One 6:e24166. https://doi.org/10.1371/journal.pone.0024166
De Deyn GB, Raaijmakers CE, Van Ruijven J, Berendse F, van der Putten WH (2004) Plant species identity and diversity effects on different trophic levels of nematodes in the soil food web. Oikos 106:576–586. https://doi.org/10.1111/j.0030-1299.2004.13265.x
De Deyn GB, Cornelissen JHC, Bardgett RD (2008) Plant functional traits and soil carbon sequestration in contrasting biomes. Ecol Lett 11:516–531. https://doi.org/10.1111/j.1461-0248.2008.01164.x
De Schrijver A, Vesterdal L, Hansen K, De Frenne P, Augusto L, Achat DL, Staelens J, Baeten L, De Keersmaeker L, De Neve S, Verheyen K (2012) Four decades of post-agricultural forest development have caused major redistributions of soil phosphorus fractions. Oecologia 169:221–234. https://doi.org/10.1007/s00442-011-2185-8
Dufrêne M, Legendre P (1997) Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol Monogr 67:345–366. https://doi.org/10.1890/0012-9615(1997)067[0345:SAAIST]2.0.CO;2
Eagar AC, Mushinski RM, Horning AL, Smemo KA, Phillips RP, Blackwood CB (2022) Arbuscular mycorrhizal tree communities have greater soil fungal diversity and relative abundances of saprotrophs and pathogens than ectomycorrhizal tree communities. Appl Environ Microbiol 88:e01782–e01721. https://doi.org/10.1128/AEM.01782-21
Eissenstat DM, Yanai RD (1997) The ecology of root lifespan. Adv Ecol Res 27:1–60
Ferris H (2010) Form and function: metabolic footprints of nematodes in the soil food web. Eur J Soil Biol 46:97–104. https://doi.org/10.1016/j.ejsobi.2010.01.003
Fierer N, Jackson JA, Vilgalys R, Jackson RB (2005) Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71:4117–4120. https://doi.org/10.1128/AEM.71.7.4117-4120.2005
Fierer N (2017) Embracing the unknown: disentangling the complexities of the soil microbiome. Nat Rev Microbiol 15:579–590. https://doi.org/10.1038/nrmicro.2017.87
Forest Agency of Japan (2022) Annual report on trends in forests and forestry 2022. Tokyo (in Japanese)
Forest Soil Division (1976) Classification of forest soils in Japan. Bull Gov For Exp Sta 280:1–28 (in Japanese with English summary)
Gebremikael MT, Steel H, Buchan D, Bert W, De Neve S (2016) Nematodes enhance plant growth and nutrient uptake under C and N-rich conditions. Sci Rep 6:32862. https://doi.org/10.1038/srep32862
Gilarte P, Pendall E, Carrillo Y, Nielsen UN (2021) Plant functional identity has predictable effects on nematode communities across successional stages. Soil Biol Biochem 162:108406. https://doi.org/10.1016/j.soilbio.2021.108406
Grace JB, Allain LK, Sankaran M, Knops J, Weiher E, Ritchie M, Anderson TM, Meche G, Andelman SJ, Smith MD, Jutila H, Seabloom E, Willig MR (2007) Does species diversity limit productivity in natural grassland communities? Ecol Lett 10:680–689. https://doi.org/10.1111/j.1461-0248.2007.01058.x
Heděnec P, Nilsson LO, Zheng H, Gundersen P, Schmidt IK, Rousk J, Vesterdal L (2020) Mycorrhizal association of common European tree species shapes biomass and metabolic activity of bacterial and fungal communities in soil. Soil Biol Biochem 149:107933. https://doi.org/10.1016/j.soilbio.2020.107933
Ingham RE, Trofymow JA, Ingham ER, Coleman DC (1985) Interactions of bacteria, fungi, and their nematode grazers: effects on nutrient cycling and plant growth. Ecol Monogr 55:119–140. https://doi.org/10.2307/1942528
Keith AM, Brooker RW, Osler GHR, Chapman SJ, Burslem DFRP, van der Wal R (2009) Strong impacts of belowground tree inputs on soil nematode trophic composition. Soil Biol Biochem 41:1060–1065. https://doi.org/10.1016/j.soilbio.2009.02.009
Kitagami Y, Matsuda Y (2020) Temperature changes affect multi-trophic interactions among pines, mycorrhizal fungi, and soil nematodes in a microcosm experiment. Pedobiologia 78:150595. https://doi.org/10.1016/j.pedobi.2019.150595
Kitagami Y, Tanikawa T, Matsuda Y (2020) Effects of microhabitats and soil conditions on structuring patterns of nematode communities in Japanese cedar (Cryptomeria japonica) plantation forests under temperate climate conditions. Soil Biol Biochem 151:108044. https://doi.org/10.1016/j.soilbio.2020.108044
Kitagami Y, Matsuda Y (2022) Effect of ectomycorrhizal fungal species on population growth and food preference of a fungivorous nematode. Mycorrhiza 32:95–104. https://doi.org/10.1007/s00572-021-01063-0
Klironomos JN, Hart MM (2001) Animal nitrogen swap for plant carbon. Nature 410:651–652. https://doi.org/10.1038/35070643
Kondratow F, Chauvin C, Villenave C, Andrieu E, Brin A (2019) Nematode communities after the reintroduction of silver fir in beech-dominated forests. Eur J For Res 138:957–965. https://doi.org/10.1007/s10342-019-01216-z
Kudrin AA, Zuev AG, Taskaeva AA, Konakova TN, Kolesnikova AA, Gruzdev IV, Gabov DN, Yakovleva EV, Tiunov AV (2021) Spruce girdling decreases abundance of fungivorous soil nematodes in a boreal forest. Soil Biol Biochem 155:108184. https://doi.org/10.1016/j.soilbio.2021.108184
Lefcheck JS (2016) piecewiseSEM: piecewise structural equation modelling in R for ecology, evolution, and systematics. Meth Evol Ecol 7:573–579. https://doi.org/10.1111/2041-210X.12512
Li Y, Du X, Su X, Han X, Liang W, Wang Z, Bruelheide H, Bezemer TM, Li Q (2023) Local-scale soil nematode diversity in a subtropical forest depends on the phylogenetic and functional diversity of neighbor trees. Plant Soil 486:441–454. https://doi.org/10.1007/s11104-023-05882-2
Liu T, Hu F, Li H (2019) Spatial ecology of soil nematodes: perspectives from global to micro scales. Soil Biol Biochem 137:107565. https://doi.org/10.1016/j.soilbio.2019.107565
Liu J, Wang X, Kou Y, Zhao W, Liu Q (2023) Differences in the effects of broadleaf and coniferous trees on soil nematode communities and soil fertility across successional stages. Plant Soil 485:197–212. https://doi.org/10.1007/s11104-022-05677-x
Maboreke HR, Graf M, Grams TEE, Herrmann S, Scheu S, Ruess L (2017) Multitrophic interactions in the rhizosphere of a temperate forest tree affect plant carbon flow into the belowground food web. Soil Biol Biochem 115:526–536. https://doi.org/10.1016/j.soilbio.2017.09.002
Mamilov AS, Dilly OM (2002) Soil microbial eco-physiology as affected by short-term variations in environmental conditions. Soil Biol Biochem 34:1283–1290. https://doi.org/10.1016/S0038-0717(02)00071-8
Mamiya Y (1967) Descriptive notes on three species of Trichodorus (Dorylaimide : Trichodoridae) from Forest nurseries in Japan. Appl Entomol Zool 2:61–68. https://doi.org/10.1303/aez.2.61
Matsuda Y, Takano Y, Shimada H, Yamanaka T, Ito SI (2013) Distribution of ectomycorrhizal fungi in a Chamaecyparis obtusa stand at different distances from a mature Quercus serrata tree. Mycoscience 54:260–264. https://doi.org/10.1016/j.myc.2012.09.019
Matsuda Y, Kita K, Kitagami Y, Tanikawa T (2021) Colonization status and community structure of arbuscular mycorrhizal fungi in the coniferous tree, Cryptomeria japonica, with special reference to root orders. Plant Soil 468:423–438. https://doi.org/10.1007/s11104-021-05147-w
Neher DA, Wu J, Barbercheck ME, Anas O (2005) Ecosystem type affects interpretation of soil nematode community measures. Appl Soil Ecol 30:47–64. https://doi.org/10.1016/j.apsoil.2005.01.002
Oksanen J, Simpson G, Blanchet F, Kindt R, Legendre P, Minchin P, O'Hara R, Solymos P, Stevens M, Szoecs E, Wagner H, Barbour M, Bedward M, Bolker B, Borcard D, Carvalho G, Chirico M, De Caceres M, Durand S, Evangelista H, FitzJohn R, Friendly M, Furneaux B, Hannigan G, Hill M, Lahti L, McGlinn D, Ouellette M, Ribeiro Cunha E, Smith T, Stier A, Ter Braak C, Weedon J (2022) Package “vegan.” Community Ecol. Package version 2.6–4. https://CRAN.R-project.org/package=vegan. Accessed 6 Dec 2023
Ostonen I, Püttsepp Ü, Biel C, Alberton O, Bakker M, Lõhmus K, Majdi H, Metcalfe D, Olsthoorn A, Pronk A (2007) Specific root length as an indicator of environmental change. Plant Biosyst 141:426–442. https://doi.org/10.1080/11263500701626069
Pierret A, Gonkhamdee S, Jourdan C, Maeght JL (2013) IJ_Rhizo: an open-source software to measure scanned images of root samples. Plant Soil 373:531–539. https://doi.org/10.1007/s11104-013-1795-9
Qian H, Hu B, Cao D, Chen W, Xu X, Lu Y (2007) Bio-safety assessment of validamycin formulation on bacterial and fungal biomass in soil monitored by real-time PCR. Bull Environ Contam Toxicol 78:239–244. https://doi.org/10.1007/s00128-007-9148-0
R Development Core Team (2023) R: a language and environment for statistical computing. R foundation for statistical computing, Austria. https://www.R-project.org/. Accessed 6 Dec 2023
Richter A, Ewald M, Hemmerling C, Schöning I, Bauhus J, Schall P, Ruess L (2023) Effects of management intensity, soil properties and region on the nematode communities in temperate forests in Germany. For Ecol Manag 529:120675. https://doi.org/10.1016/j.foreco.2022.120675
Ruess L, Zapata EJG, Dighton J (2000) Food preferences of a fungal-feeding Aphelenchoides species. Nematology 2:223–230. https://doi.org/10.1163/156854100508962
Salamon JA, Wolters V (2009) Nematoda response to forest conversion. Eur J Soil Biol 45:184–191. https://doi.org/10.1016/j.ejsobi.2008.09.014
Saucet SB, van Ghelder C, Abad P, Duval H, Esmenjaud D (2016) Resistance to root-knot nematodes Meloidogyne spp. in woody plants. New Phytol 211:41–56. https://doi.org/10.1111/nph.13933
Sawada K, Inagaki Y, Sugihara S, Funakawa S, Ritz K, Toyota K (2021) Impacts of conversion from natural forest to cedar plantation on the structure and diversity of root-associated and soil microbial communities. Appl Soil Ecol 167:104027. https://doi.org/10.1016/j.apsoil.2021.104027
Scherber C, Eisenhauer N, Weisser WW, Schmid B, Voigt W, Fischer M, Schulze ED, Roscher C, Weigelt A, Allan E, Beler H, Bonkowski M, Buchmann N, Buscot F, Clement LW, Ebeling A, Engels C, Halle S, Kertscher I et al (2010) Bottom-up effects of plant diversity on multitrophic interactions in a biodiversity experiment. Nature 468:553–556. https://doi.org/10.1038/nature09492
Schneider G, Chicken E, Becvarik R, Schneider MG (2021) Functions and datasets to accompany Hollander, Wolfe, and Chicken—nonparametric statistical methods. R Package Version 1:16
Shipley B (2009) Confirmatory path analysis in a generalized multilevel context. Ecology 90:363–368. https://doi.org/10.1890/08-1034.1
Soil Survey Staff (2014) Keys to soil taxonomy, 11th edn. USDA-Natural Resources Conservation Service, Washington, DC
Stefanowicz AM, Rożek K, Stanek M, Rola K, Zubek S (2021) Moderate effects of tree species identity on soil microbial communities and soil chemical properties in a common garden experiment. For Ecol Manag 482:118799. https://doi.org/10.1016/j.foreco.2020.118799
Tanikawa T, Maie N, Fujii S, Sun L, Hirano Y, Mizoguchi T, Matsuda Y (2023) Contrasting patterns of nitrogen release from fine roots and leaves driven by microbial communities during decomposition. Sci Total Environ 855:158809. https://doi.org/10.1016/j.scitotenv.2022.158809
Tedersoo L, Bahram M, Zobel M (2020) How mycorrhizal associations drive plant population and community biology. Science 367:1–9. https://doi.org/10.1126/science.aba1223
Thakur MP, Geisen S (2019) Trophic regulations of the soil microbiome. Trends Microbiol 27:771–780. https://doi.org/10.1016/j.tim.2019.04.008
van Bezooijen J (2006) Methods and techniques for nematology. Wageningen University, Wageningen, pp 30–33
van den Hoogen J, Geisen S, Routh D, Ferris H, Traunspurger W, Wardle DA, de Goede RGM, Adams BJ, Ahmad W, Andriuzzi WS, Bardgett RD, Bonkowski M, Campos-Herrera R, Cares JE, Caruso T, de Brito CL, Chen X, Costa SR, Creamer R et al (2019) Soil nematode abundance and functional group composition at a global scale. Nature 572:194–198. https://doi.org/10.1038/s41586-019-1418-6
Wada R, Tanikawa T, Doi R, Hirano Y (2019) Variation in the morphology of fine roots in Cryptomeria japonica determined by branch order-based classification. Plant Soil 444:139–151. https://doi.org/10.1007/s11104-019-04264-x
Wilschut RA, Geisen S, Martens H, Kostenko O, de Hollander M, ten Hooven FC, Weser C, Snoek LB, Bloem J, Caković D, Čelik T, Koorem K, Krigas N, Manrubia M, Ramirez KS, Tsiafouli MA, Vreš B, van der Putten WH (2019) Latitudinal variation in soil nematode communities under climate warming-related range-expanding and native plants. Glob Change Biol 25:2714–2726. https://doi.org/10.1111/gcb.14657
Wilschut RA, Geisen S (2021) Nematodes as drivers of plant performance in natural systems. Trends Plant Sci 26:237–247. https://doi.org/10.1016/j.tplants.2020.10.006
Wyss U (1997) Root parasitic nematodes: an overview. In: Fenoll C, Grundler FMW, Ohl SA (eds) Cellular and molecular aspects of plant-nematode interactions. Kluwer Academic Publishers, Dordrecht, pp 5–22
Yeates GW, Bongers T, De Goede RGM, Freckman DW, Georgieva SS (1993) Feeding habits in soil nematode families and genera—an outline for soil ecologists. J Nematol 25:315–331
Zhang X, Ferris H, Mitchell J, Liang W (2017) Ecosystem services of the soil food web after long-term application of agricultural management practices. Soil Biol Biochem 111:36–43. https://doi.org/10.1016/j.soilbio.2017.03.017
Zhang C, Wang J, Ren Z, Hu Z, Tian S, Fan W, Chen X, Griffiths BS, Hu F, Liu M (2020) Root traits mediate functional guilds of soil nematodes in an ex-arable field. Soil Biol Biochem 151:108038. https://doi.org/10.1016/j.soilbio.2020.108038
Zhang J, Hu Z, Zhang C, Tao Y, Chen X, Griffiths BS, Liu M (2022) Roots with larger specific root length and C: N ratio sustain more complex rhizosphere nematode community. Plant Soil 477:693–706. https://doi.org/10.1007/s11104-022-05465-7
Acknowledgments
We thank Mr. G. Yamanaka (The Mie Prefectural Forestry Research Center) for permission to access study sites. We also thank members of the laboratory of Forest Mycology at Mie University for their support with field sampling. We would like to thank Editage (www.editage.com) for English language editing. This study was supported in part by the Grants-in-Aid for Scientific Research (B) 18H02237, 21H02232 to YM and JSPS Research Fellow 18 J13285, Grant-in-Aid for Young Scientists 21 K14876 to YK from the Japan Society for the Promotion of Science.
Funding
This study was supported in part by the Grants-in-Aid for Scientific Research (B) 18H02237, 21H02232 to YM and JSPS Research Fellow 18 J13285, Grant-in-Aid for Young Scientists 21 K14876 to YK from the Japan Society for the Promotion of Science.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by YK, KS and YM. The first draft of the manuscript was written by YK and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have influenced the work reported in this paper.
Additional information
Responsible Editor: Sven Marhan.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(XLSX 41 kb)
ESM 2
Fig. S1. A conceptual figure of pot experiment. (a) 4 soil and tree species treatment combinations were established; Cryptomeria japonica soil with C. japonica seedlings (CC); Quercus serrata soil with C. japonica seedlings (QC); C. japonica soil with Q. serrata seedlings (CQ); Q. serrata soil with Q. serrata seedlings (QQ). (b) C. japonica and Q. serrata seedlings in pots grown for 11 months. (c) Procedures of experimental set-up. Fig. S2. Hypothesized initial full model. Soil origins and tree species were indicated as a categorical explanatory variable, i.e. soil origins; C. japonica (1) and Q. serrata (2), tree species; C. japonica (1) and Q. serrata (2). Principal component analysis (PCA) was performed to reduce the dimensions of soil abiotic factors, i.e., soil pH and C/N. The first principal component of soil properties (Soil PC1) was negatively correlated with soil pH and C/N ratio (see Fig. S3). Bact, abundance of bacterivores; Fung, abundance of fungivores; Herb, abundance of herbivores; Pred&Omni, abundance of predators and omnivores. Fig. S3. Principal component analysis of soil abiotic factors (soil pH and C/N ratio) from 96 pots. Cryptomeria japonica soil with C. japonica seedlings (CC); Quercus serrata soil with C. japonica seedlings (QC); C. japonica soil with Q. serrata seedlings (CQ); Q. serrata soil with Q. serrata seedlings (QQ). Fig. S4. Non-metric multidimensional scaling scatterplot of Chao dissimilarity based on the abundance of nematode taxa derived from four treatments and no seedling pots as control. Stress value = 0.227. Treatment codes show Cryptomeria japonica soil with C. japonica seedlings (CC); Quercus serrata soil with C. japonica seedlings (QC); C. japonica soil with Q. serrata seedlings (CQ); Q. serrata soil with Q. serrata seedlings (QQ). The “origin” showed the dataset at the time of sampling. The “control” showed the dataset of no tree treatments. Nematode community structures were clustered significantly into each treatment (PERMANOVA, P < 0.001, R2 = 0.73). Fig. S5. Number of total nematodes and four trophic groups per 100-g dry soil at four different treatments and no seedling pots as control. Columns are means (n = 24) with standard errors excepting for the number of control pots (n = 9). Treatment codes show Cryptomeria japonica soil with C. japonica seedlings (CC); Quercus serrata soil with C. japonica seedlings (QC); C. japonica soil with Q. serrata seedlings (CQ); Q. serrata soil with Q. serrata seedlings (QQ). C and Q indicate no tree treatments, i.e., only C. japonica and Q. serrata soils, respectively. (PPTX 452 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kitagami, Y., Suzuki, K. & Matsuda, Y. Effects of tree species identity and soil origin on soil nematode communities and trophic composition in coniferous and broad-leaved forests. Plant Soil (2024). https://doi.org/10.1007/s11104-024-06599-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11104-024-06599-6