Abstract
Purpose
Farmers need alternative approaches to manage nitrogen (N) that meet crop needs while reducing loss to the environment. Identifying crop genotypes that promote decomposition of organic materials and understanding the potential mechanisms responsible could help address this challenge. Consequently, we aimed to 1) determine whether carrot genotypes differ in their potential to facilitate organic matter decomposition in soil, and 2) identify bacterial taxa that are stimulated by carrot roots, and thus could play a role in these processes.
Methods
We grew five genotypes expected to differ in N use efficiency in a nutrient-poor soil amended with 15 N-enriched corn residue, tracked changes in carbon (C) and N pools, quantified microbial activity and bacterial community composition, and predicted the potential expression of microbial genes involved in soil C and N cycles.
Results
Experimental genotype 8503 had the greatest capacity to promote decomposition of corn residues. This genotype had the highest % of N from the corn residue in its taproots and on average, promoted higher b-glucosidase activity in soils. Distinct bacterial communities from the families Micromonosporaceae, Chromatiaceae, and Rhodospirillaceae were also enriched in the soils of genotypes like 8503 that were most effective in obtaining N from the corn residue, and this was correlated with greater potential expression of genes responsible for β-glucosidase and nitrification activity.
Conclusions
Carrot genotypes do differ in their potential to alter soil bacterial communities and stimulate microbially-mediated decomposition of organic materials indicating that it may be possible to begin selecting for this important trait.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nitrogen (N) is an essential macronutrient required by all plants in large quantities to fulfill critical processes such as photosynthesis. Farmers apply substantial amounts of N fertilizers to meet crop plant needs, however, only 50% is generally captured by plants (Fageria and Baligar 2005). The remaining fertilizer is subject to loss via leaching and emissions of nitrous oxide (N2O), which negatively impacts environmental health (Erisman et al. 2013). One way to address this challenge is by improving crop nitrogen-use efficiency (NUE), which is broadly defined as the yield that a plant produces per unit of N fertilizer applied (Fageria and Baligar 2005). NUE can be further separated into components, such as a plants ability to recover N in soil, and its capacity to utilize N efficiently to support physiological functions. The former process is likely to be particularly critical for reducing N loss to the environment. Plant root traits most commonly known for their potential to influence N acquisition in soil include growth rate, architecture, amount and duration of root hairs, and ability to host symbiotic microbes such as arbuscular mycorrhizal fungi (AMF) (Bardgett et al. 2014; Blagodatskaya et al. 2014; Poirier et al. 2018). However, the ideal set of traits may vary given the type of fertilizer applied. For example, in agricultural systems that rely exclusively on inorganic forms of N fertililizers such as ammonium (NH4+) and nitrate (NO3−), the ideal root ideotype has been referred to as “steep, deep and cheap” (Lynch 2013). In this case, roots that grow quickly and deeply into the soil profile are useful in acquiring inorganic N compounds subject to leaching. In contrast, in systems where plants are more reliant on decomposing organic matter pools to obtain N, producing roots closer to the soil surface, and releasing compounds from roots that can stimulate microbially-mediated decomposition and N cycling processes in soils could be more important.
Farmers have long relied on organic nutrient sources such as animal manures, animal byproducts and residues from cover crops to help meet N fertility needs (Carr et al. 2020). While the use of these organic fertility sources fell out of favor during the twentieth century due to the widespread availability of inexpensive inorganic N fertilizers derived using the Haber–Bosch process, rising costs and environmental concerns have led to a resurgence in their use (Carr et al. 2020; Hirel et al. 2011). Plants can take up dissolved low molecular weight organic N compounds like urea, amino acids and oligopeptides to help fulfill their N needs (Moreau et al. 2019), however, this is energetically expensive so plants generally prefer NH4+ and NO3- since these compounds require less processing internally (Laberge et al. 2006). Decomposition and cycling of organic materials into these inorganic N compounds is carried out by a diverse set of soil macro and microfauna and these processes are dependent on environmental conditions such as temperature and moisture (Frouz 2018). This makes synchronizing plant available N with critical periods of plant uptake when relying on organic fertility sources challenging (Baresel et al. 2008; Hirel et al. 2011; Hoagland et al. 2008; Rudisill et al. 2016). Evidence is accumulating that plants can also influence decomposition of organic materials in soil to help meet their nutrient needs (Heijboer et al. 2016; Moreau et al. 2019; Yin et al. 2018), and this could aid in N acquisition from organic nutrient sources. Plants release up to 40% of their photosynthetically derived C from their roots through both active and passive processes (Bais et al. 2006). These rhizodeposits can influence decomposition of organic matter within soils in a phenomenon known as priming. This phenomenon has been extensively studied for its effects on C fluxes from soil and recent studies indicate that it could also be important for N cycling (Blagodatsky et al. 2010; Finzi et al. 2015; Meier et al. 2017; Murphy et al. 2015; Mwafulirwa et al. 2017; Paterson et al. 2016). As plants grow and N demand increases, plants that are able to exude more C from their roots to stimulate N cycling under situations where is limited, would have greater productivity.
While the health of soils and availability of organic materials is likely to be critical in mediating decomposition of organic materials in soil, plant genetics could also play an important role in this process (Dijkstra et al. 2013; Sasse et al. 2018; Terrazas et al. 2019). For example, wheat (Triticum aestivum L.) genotypes with high C investment in roots and root growth rates had greater capacity to stimulate decomposition and obtain N from compost to promote plant growth (Junaidi et al. 2018). Other researchers have demonstrated that the total amount of C released to soil through root exudates can vary among plant genotypes (Kuzyakov 2002; Schenk 2006; Yin et al. 2018), providing evidence that it may be possible to identify genotypes that can preferentially stimulate decomposition of organic residues in soil, thereby increasing NUE. Carrot (Daucus carota L.) is an ideal model crop to investigate how plant genetics can interact with microbial communities to facilitate N acquisition from organic sources in soil. Carrot is an important component of the rapidly growing organic vegetable industry (Kim et al. 2019), and this crop has one of the greatest market shares of organic vegetables with 14.3% of the total U.S. carrot crop grown on organic farms (Lucier and Lin 2011). Carrot is well known for its ability to scavenge N in soil and this crop is commonly included in crop rotation and intercropping strategies to help prevent N loss (Thorup-Kristensen 2006; Westerveld et al. 2006). Standard recommendations for N in carrots range from 50 to 220 kg/ha (Boskovic-Rakocevic et al. 2012; Veitch et al. 2014), though N uptake from inorganic fertilizers is notoriously low with only 8 to 33% of the N actually making it into carrot crops (Reid et al. 2017). Moreover, carrots do not respond to increased rates of inorganic N fertilizers to enhance yield suggesting that they can obtain sufficient N from other sources such as decomposing organic matter (Veitch et al. 2014).
In our organic carrot breeding program, we have observed potential differences in NUE among entries grown in our field trials (Simon et al. 2017). We have also determined that carrot genotypes host distinct microbial assemblages in their taproots which can play a role in helping carrot plants resist pathogens (Abdelrazek et al. 2020a, b), and we suspect that these microbes could also influence decomposition of plant residues and therefore NUE. To test these hypotheses, we grew five carrot genotypes suspected to differ in NUE from our field trials in a nutrient poor soil amended with a 15 N labeled corn residue and tracked changes in C and N cycling, soil bacterial composition and potential functional activity, and N uptake into carrot plants.
Materials and Methods
Greenhouse experiment
The experiment was conducted in the greenhouse using an artificial soil constructed from a 50:50 mix of agricultural field soil and sand. Initial characteristics of this soil were determined using standard procedures at the Midwest Soil Testing Laboratory in Omaha, NE (midwestlabs.com). Briefly, total soil organic matter was determined based on loss on ignition; pH was determined using a 1:1 soil water mixture; available phosphorous (P) was extracted as weak Bray and analyzed colorimetrically; exchangeable potassium (K), calcium (Ca) and magnesium (Mg) were extracted with neutral ammonium acetate (1 N) and quantified using inductively coupled argon plasma-mass spectrometry (ICP-MS); and base saturation and cation exchange capacity (CEC) were estimated from the results of exchangeable cations (Moore 2001). The soil mixture contained 1% total organic matter (OM), a pH of 7.2, and 110, 359, 327, and 1598 mg/kg available P and exchangeable K, Mg and Ca, respectively. Cation exchange capacity was 11.6 mmol ( +) /kg soil and the % base saturation on CEC sites was 8, 24 and 61 for K, Mg, and Ca respectively. Initial NO3-N and NH4-N values were 6.5 and 1.2 mg N/kg, respectively, which were determined using techniques described in Section "Quantification of plant available nitrogen in soils". Total elemental N, organic C and δ15N values of the soil and organic amendments (Table 1.) were determined using techniques described in Section "Soil and plant carbon and nitrogen elemental composition".
The artificial soil mixture was distributed into 15.2 × 40.6 cm pots, each with a total volume of 7.7 L and a soil bulk density of 1.03 g cm–3. Each pot was amended with 1.0 g of 15 N enriched corn material that was previously generated in a field trial by growing corn plants with 15 N enriched N fertilizer. The corn biomass was harvested from the field, dried and ground, and the total element N, organic C and δ15N values δ15N values determined using techniques described in Section "Soil and plant carbon and nitrogen elemental composition". Next, 1.2 g of organic feather meal fertilizer (Down to Earth, Eugene, Oregon) was added to each both and both substrates were mixed evenly in the soil prior to planting 7 seeds in each pot from one of five carrot genotypes (Table 2). The control experiment was only amended with 1.2 g of organic feather meal fertilizer. Soil, enriched corn material and feather meal fertilizer C: N ratios were 7:1, 62:1 and 3:1, respectively. These carrot genotypes were chosen because they have similar taproot morphological characteristics but are expected to vary in NUE based on our field trials. This included two experimental breeding lines and three commercial varieties. The pots were placed in the Purdue Greenhouse Horticulture Facility in West Lafayette, IN during the first week of June. The average day and nighttime temperatures were set at 22 °C and 18° C ± 1° C, respectively, and relative humidity was maintained at between 50 and 70%. Auto-drippers were placed in each pot, and all pots were irrigated daily as needed with the same amount of water throughout the experiment.
After germination, carrot seedlings were thinned to one per pot. The pots were arranged in a complete randomized block design with the five carrot genotypes and one unplanted control treatment, each with six replicates for a total of 36 pots. At 6 and 10 weeks after seeding and at harvest, three soil cores were collected to a depth of 0–10 cm in each pot with a 1.27 cm probe to minimize any potential damage to carrot roots. The three soil cores from each pot were mixed and homogenized. Every time the soil samples were recovered from the pot, the holes were covered to minimize the effects of soil disturbance. A subsample of the pooled soil cores collected from each pot was stored in a plastic bag at 4° C until microbial analysis and another was air-dried for two days and sieved (< 2 mm) in preparation for elemental analyses described below. Carrots were harvested in September, 90 days after seeding, when they reached maturity.
The total wet weight of above and below ground biomass was determined at harvest and the plant materials were oven dried at 70 °C until they no longer changed in moisture content (4 days) to quantify the dry weight of each fraction. The dry plant samples were ground using a UDY cyclone sample mill (UDY Corp., Boulder, Col) to obtain fragments of approximately 1 mm for analyses described below.
Quantification of plant available nitrogen in soils
A modified protocol by Bottomley et al. (1994) was used to quantify NH4+-N and NO3-N in each soil sample. Briefly, 12.5 mL of KCl (1 M) was added to 5 g soil samples and the slurries were incubated at room temperature for 30 min while shaking at 250 rpm. Samples were then filtered through a #42 Whatman filter paper for 10–15 min and the filtrate was stored at -20° C until analysis using an AQ2 Discrete SEAL Analyzer (Seal Analytical, Southampton, Hampshire, United Kingdom) to quantify nitrate (NO3-N) and ammonium (NH4-N) according to EPA methods AGR-231-A-Rev. 0 and AGR-210-A Rev. 0, respectively. All analyses were conducted in duplicate.
To estimate the corn residue decomposition, we calculated the mg of inorganic N per kg of soil made available from organic sources, or N made available (NRO), during the experiment as follows:
where \(IN\left(harvest\right)\) is the soil inorganic N (NH4-N + NO3-N) at harvest, \(IN(initial)\) is the initial soil inorganic N (NH4-N + NO3-N) in each pot, kg soil is the total soil in each pot, and \(total\;plant\;N\;uptake\) (total % N x total plant dry weight).
Soil ß -glucosidase enzyme assay
Soil ß-glucosidase activity was quantified at three time points (6 and 10 weeks after seeding and at harvest) in triplicate for each soil sample. The assay was based on the determination of p-nitrophenol released after incubating soil with p-nitrophenyl (Eivazi and Tabatabai 1988). Briefly, 1 g of moist soil per sample was incubated for 1 h with a solution containing 0.25 mL of toluene, 4 mL of modified universal buffer (MUB) and 1 mL of p-nitrophenyl ß-d-glucoside (PNG). After the incubation, a solution containing 1 mL of CaCl2 and 4 mL of tris buffer at pH 12 were added to stop the enzymatic reaction. The soil suspensions were then filtered immediately through #42 Whatman filter paper. Three technical replicates of each sample were then diluted by half and the color intensity of the filtrate at 400 nm was quantified. During each measurement an additional blank suspension with no soil was included as a control.
Soil and plant carbon and nitrogen elemental composition
Approximately 30 mg of the initial soil mixture was analyzed for total C and N with a Thermo Scientific FlashEA 1112 Nitrogen and Carbon Analyzer for Soils, Sediments, and Filters (CE Elantech, Lakewood, NJ). In addition, 0.5 mg of the organic feather meal fertilizer and 3.8 mg of the 15 N labeled corn residue were analyzed using the same procedure to quantify C and N. The total elemental (C and N) isotope composition of carrot root and shoot biomass, soil, organic feather meal fertilizer as well as values of delta 15 N (δ15N ‰), were quantified using the PDZ Europa Elemental Analyzer interfaced to a Sercon Ltd. 20–22 isotope ratio mass spectrometer (EA-IRMS) at the Purdue Stable Isotope Facility using established protocols.
Nitrogen isotope ratio calculations
The δ 15 N ‰ value is calculated as the proportional amount of 15 N (‰) in the sample relative to an international isotope standard of air (0.00366 atom ‰15 N) according to the following equation:
where R sample is the isotope ratio15N/14 N of the sample and the R standard is the atmospheric nitrogen with a 15 N/14 N ratio of 0.00368 (Van Cleemput et al. 2008).
Using the δ15N value obtained for carrot roots and shoots, we determined the %N in each plant that was obtained from the 15 N enriched corn residue (\(\%N FR)\) using the following calculation:
Equation (3) calculates %N obtained from 15 N labeled corn residue based on the δ 15 N values from carrot roots and shoots and δ 15 N values of enriched corn residue and the unlabeled corn residue and carrot roots and shoots controls.
Statistical analyses of soil and plant assays
Differences in soil ß-glucosidase activity, NH4-N, NO3-N, %N and %C collected at the three time points were evaluated using a linear mixed model with time and genotype as fixed factors, and pot as a random factor. NH4-N and NO3-N values were ln-transformed to meet normality assumptions and outliers were removed using the R function boxplot.stats with a 1.8 coefficient. A two-way ANOVA analysis was then conducted to determine if the effect of time, genotype and the genotype x time interaction were significantly different. Inorganic N made available and %N from residue during the experiment was evaluated using a linear model and a one-way ANOVA to determine if genotype was significant. Data from root and shoot dry weights that did not meet the normality assumptions were fitted to a linear model and a non-parametric Kruskal–Wallis test was conducted to determine if there were differences among the carrot genotypes. Then the root:shoot ratio (R:S), %C, %N, C:N ratio and total δ 15 N abundance values were each fitted to a linear regression model and subject to a one-way ANOVA. A post-hoc Tukey pairwise comparison was performed for each of these values to identify differences among the carrot genotypes. Finally, data was fitted to a linear regression and an ANOVA analysis was performed to determine if there were differences in \(\%N FR\) uptake among the genotypes.
Bacterial soil microbiome analysis
DNA was extracted from 0.25 g soil samples collected at harvest using DNeasy PowerSoil Kits (Qiagen, Venlo, Netherlands) in triplicate for each pot for a total of 108 extractions. The quantity and quality of each extraction was estimated using a Nanodrop 2000 microvolume spectrophotometer (Thermo Fischer Scientific, Waltman, MA, U.S.A), before pooling the three technical replicates from each pot. The concentration of DNA in each sample was then quantified using a Qubit Fluorometer (Thermofisher Scientific) to obtain a more accurate concentration and samples were diluted to 1 ng/μl in preparation for use as a template in PCR.
PCR reactions were conducted using primers targeting the V3-V4 16 s rRNA bacterial region: V3_341F: CCTACGGGAGGCAGCAG and V4_806R: GGACTACHVGGGTWTCTAAT. In addition, each primer contained a so-called “CS” linker sequence which is necessary to run a 2nd PCR with Illumina adapters and barcodes. 1 μl of DNA was used as a template in each 25 μl PCR reaction, which also included 10.5 μl H2O, 12.5 μl Q5Ⓡ High-fidelity Master Mix (NEB), 0.5 μl primer V3 (10 μM), and 0.5 μl primer V4 (10 μM). All PCR reactions were conducted in triplicate for each sample. PCR thermocycler conditions were as follows: initial denaturation at 95 °C for 5 min followed by 28 cycles of: 95 °C for 30 s, 50 °C for 30 s, 72 °C for 1 min and a final elongation at 72 °C for 10 min. After PCR, 4 μl of each amplification product (~ 460 pb) was visualized on a 1% agarose gel to verify that the PCR reaction was successful and the three replicate PCR reactions were pooled. Finally, a total of 36 samples (25 μl each) of final PCR products were sent to the Genome Research Core Facility at the University of Illinois in Chicago, U.S.A. for sequencing on the Illumina MiSeq platform.
Bioinformatic analysis of the soil microbiome data
Sequences obtained from the Illumina platform were demultiplexed based on individual barcodes assigned to each sample and trimmed according to a 25 phred quality value using the TrimGalore tool (Krueger 2015). Reads from forward and reverse primers were then merged using the join_paired_ends.py script found in QIIME v2. (Bolyen et al. 2019) and quality checked using the FastQC tool (Andrews 2010). QIIME v2. was used for picking, identifying chimeric sequences, assigning bacterial taxonomy, and aligning final operational taxonomic units (OTUs) with the SILVA 119 version bacterial data base (Pruesse et al. 2007), and then OTUs were filtered for chloroplasts and mitochondria sequences. Statistical analyses were conducted to calculate bacterial α-diversity (Shannon indexes) using the alpha_rarefaction.py script that uses a nonparametric index to measure the total number of OTUs and evenness (relative abundance of OTUs) using QIIME v2. A post-hoc Tukey pairwise comparison was performed to test for differences in α-diversity for each pair of means, and differences in individual families between genotypes. To analyze bacterial ß-diversity, a Bray–Curtis dissimilarity distance matrix was created and visualized using a PcoA ordination plot, and an ADONIS test was conducted to determine if there were statistical differences between the treatments using the R package “vegan” (Oksanen 2007), in R version 3.6.2 (R Development Core Team 2019).
The PICRUSt2 program was used to predict the abundance of genes from protein families involved in cellulose degradation and N metabolism using normalized OTU counts (Langille et al. 2013) (Table 3). Differences in the abundance of genes predicted between treatments were evaluated using a permutational analysis of variance (PERMANOVA) in the software PAST, based on Bray–Curtis distance and with 9999 bootstrap replications. Potential enrichment of specific genes among treatments were determined using both a correlation-based indicator species analysis and a differential abundance analysis. The correlation-based indicator species analysis was conducted using the “indicspecies” package implemented in R (De Cáceres and Legendre 2009), and was computed under a point-biserial correlation coefficient with 10,000 replications and a confidence of p < 0.05 (Hartman et al. 2018). The differential gene expression analysis was calculated using the edgeR package implemented in R (Robinson et al. 2009), computed using likelihood ration tests, and results were considered significantly different if they had a false discovery rate of p < 0.05 (Hartman et al. 2018).
Results
Soil ß-glucosidase enzyme assay
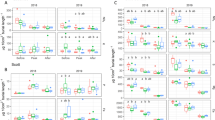
Soil ß-glucosidase activity differed based on the interaction between carrot genotype and time (p < 0.01) (Table S1). At 6 weeks after planting, there were no differences among the genotypes, however, at 10 weeks after seeding there were (Fig. 1a; Table S2). Specifically, carrot genotype 8503 had greater soil ß-glucosidase activity than Karotan and the unplanted control, and 3999 had greater levels than the unplanted control. At harvest there were no differences among genotypes (Fig. 1a; Table S2).
Plant available N (NH4-N and NO3-N) in soil
Soil NH4-N differed based on the interaction between carrot genotype and time (p < 0.001) (Table S1). Soil NH4-N levels were similar among carrot genotypes at 6 weeks after seeding, but differed at 10 weeks after seeding and harvest (Fig. 1b; Table S2). Specifically, at 10 weeks after seeding, there was greater concentrations of soil NH4-N in Karotan than Uppercut (Fig. 1b; Table S2). At harvest, 3999 had greater soil NH4-N values than 8503, Uppercut and the unplanted control, and Karotan had greater values than Uppercut or the unplanted control (Fig. 1b; Table S2).
Soil NO3-N values differed based on genotype (p < 0.01) and time (p < 0.01), but there was no interaction between these two factors (Fig. 1c; Table S1). With the exception of genotype Karotan, soil NO3-N values decreased across time. Soil NO3-N values were greater across time in the unplanted control relative to all pots planted with carrots, but there were no differences among genotypes within the individual time points (Fig. 1c; Table S2).
Soil total C and N, C:N ratio and elemental composition
Soil C concentration differed based on the interaction between carrot genotype and time (p < 0.001) (Table S1). There were no differences in soil %C levels among the genotypes at 6 weeks after seeding or at harvest, however, there were differences at 10 weeks after seeding (Fig. 1d; Table S2). Specifically, Sun255 and the unplanted control had higher soil %C levels than 3999, 8503 and Karotan.
Soil %N also differed based on an interaction between carrot genotype and time (p < 0.01) (Table S1). At 6 weeks after seeding, the unplanted control had higher %N levels than 8503 and Karotan (Fig. 1e; Table S2). At 10 weeks after seeding, Sun255 and the unplanted control had greater %N values than 8503, 3999, Uppercut and Karotan (Fig. 1e; Table S2). Finally, at harvest Sun255 and the unplanted control had greater soil %N values than Karotan (Fig. 1e; Table S2).
Soil C:N ratio was significantly different based on genotype only (p < 0.001) (Table S1). At 6 weeks, genotype 8503 had significantly greater soil C:N than 3999, Sun255 and Uppercut, while at 10 weeks genotype 8503 had significantly greater soil C:N than Sun255. There were no significant differences in the soil C:N ratio at harvest (Fig. 1f; Table S2).
Root and shoot dry weight, root:shoot ratio and carbon and nitrogen composition of plant tissues
There were no significant differences in total root, shoot or combined root and shoot dry weight (Table 4; Table S4). There were also no differences among genotypes in %C in roots or shoots, %N in roots, or C:N ratio in roots, however, there were significant differences in %N and the C:N ratio in shoots (Table 4). Specifically, there was greater %N in the shoots of 3999 than Karotan, Sun255 and Uppercut, and 8503 had greater %N than Uppercut, and Karotan, Sun255 and Uppercut had greater C:N ratios than 3999 and 8503 in carrot shoots.
There were no significant differences in the total amount of N in carrot roots, shoots or total plant biomass (Table 4; Table S4). Between 171–323 total mg N/kg was made available from the corn residue and taken up by carrot plants during the course of the experiment, and carrots obtained between 33–44% of the total N from this material (Table 4; Table S4). There were differences in the % of N in roots from the corn residue based on genotype, with greater % NFR in 8503 than Uppercut, but there were no differences in carrot shoots (Table 4).
Soil bacterial microbiome composition and diversity at harvest
Twenty-two unique bacterial families were identified in the soils collected at harvest and there were significant differences in families between carrot genotypes (Fig. 2; Table S5). For example, Oxalobacteraceae, Opitutaceae, Micrococcaceae families were detected in Karotan, Sun255 and Uppercut but not the other genotypes, whereas the Chromatiacea family was detected only in 3999, 8503 and Sun255. The Intrasporangiaceae family was found in Karotan and Sun255 and the Familyl family was detected only in 3999, Sun255 and Uppercut. 3999 and the unplanted control were the only treatments in which the Rhodospirillaceae family was found. Finally, Nitrosomonadaceae was greater in 3999 and 8503 than Karotan, Sun255 and Uppercut.
When comparing alpha diversity among the bacterial communities there were significant differences (p < 0.01) between 3999, 8503, Sun255 and the unplanted control treatment (Fig. 3a). Results of the principal components analysis (PCoA) and ADONIS test indicated that there were also significant differences in β diversity among the soil bacteria communities in the carrot genotypes (p < 0.001). Based on the R2 value in the ADONIS test, 50% of variation in distances was explained by carrot genotypes, and two of PCoA axes explained 21% of the total variability (Fig. 3b). Specifically, 3999 and 8503 clustered together in group A, while Karotan, Uppercut and Sun255 clustered in group B, and the unplanted control treatment clustered in group C (Fig. 3b).
Differences in α diversity (a) and β diversity (b) of soil bacterial communities in pots planted with five carrot genotypes and an unplanted control. Points in (a) indicate outliers for each group. Different letters in (a) indicate significant differences between genotypes in the Tukey test pairwise comparison (α value = 0.05). Principal components analysis (PcoA) in (b) explained 21% of the variability based on Bray–Curtis distances
3.6 PICRUSt’s prediction analysis at harvest
Results of the PICRUSt’s gene prediction indicated there were significant differences in the potential for expression of cellulose degradation and N metabolism genes in soil bacterial communities present at harvest after cultivation of the five carrot genotypes and unplanted control (Fig. 4; Table 5; Table S6). Specifically, the genes celA (p < 0.001), celF (p < 0.01), β-glucosidase (p < 0.01), and bglA (p < 0.001) involved in cellulose degradation were significantly different, as were genes norB (p < 0.01), pmoA-amoA (p < 0.01), hao (p < 0.01), narG-narZ-nxrA (p < 0.05), nirK (p < 0.001) and nifA (p < 0.01), which are involved in N metabolism.
Results of the indicator species analysis indicated that several cellulose degradation and N metabolism genes could be considered to be unique, or had potential for greater expression, in soil bacterial communities associated with the individual carrot genotypes (Table 5). Among the cellulose degradation genes, β -glucosidase, celF, bglA, and celA genes were associated with Sun255; β -glucosidase and celF were associated with 3999 and 8503; and, celF, bglA, and celA with Uppercut. Among the N metabolism genes, nirK, norB, narG-narZ-nxrA, nifA, p-amoA-amoA, and hao genes were associated with 3999 and 8503; nirK, norB, narG-narZ-nxrA and nifA were associated with Sun255; and, p-amoA-amoA was the only gene associated with the untreated control.
Results of the differential gene expression analysis were similar to those of the indicator species analysis, also indicating that there were potential differences in gene expression among soil bacterial communities following cultivation with the five carrot genotypes (Fig. 4; Table 5). Among the cellulose degradation genes, celF expression was expected to have greater expression in 3999, 8503, Sun255 and Uppercut relative to the other genotypes; and, blglA and celA expression was likely to have greater expression in Sun255 and Uppercut. Among the N metabolism genes, p-amoA-amoA and nifA expression were likely to have greater expression in 3999, 8503 and the untreated control; and, nifA was greater in Sun255.
Discussion
Determining how plants interact with soil microbial communities to facilitate C and N cycling has potential to increase the health and yield of crops while reducing N losses to the environment (de Vries and Wallenstein 2017; Dijkstra et al. 2013; Meier et al. 2017). One way that plants can influence C and N cycling processes is through priming. To quantify the potential for carrot plants to promote decomposition of organic materials in soil, we amended soils with an 15 N -isotopically labeled corn residue. The vast majority of N in the environment is present in the form 14 N, so introducing materials enriched in 15 N form allows researchers to track the movement of specific N compounds in the environment (de Oliveira Silva et al. 2017). The corn residue used in this study had an initial δ15N value of 603‰. This residue also had a high C:N ratio (62:1), so it should have been slow to decompose relative to the other N source added in this study, and thus serve as a good material to quantify priming activity by the carrot plants. The δ15N. values of the carrots from the control experiment ranged from 10.1 to 12, similar to other published values for carrots (Bateman et al. 2005). The δ 15 N values from the enriched experiments exhibited higher values ranging 13.2 to 15.1, confirming that the carrot plants did obtain N from the corn residue (Table 4). Specifically, carrots obtained around 4% of their total N from the corn residue and took up from 33–44% of the N that was added in the residue (Table 4; Table S4).
Determining whether genotypes differ in their potential to enhance decomposition of organic materials to obtain N is critical to being able to leverage this capacity to improve NUE in crop breeding programs. In this study, the experimental genotype 8503 appeared to have the greatest potential to obtain N from organic materials given the higher concentrations of N obtained from the corn residue in its roots at harvest (Table 4). Other evidence that experimental genotype 8503 was better in stimulating organic matter priming and obtaining N is provided by differences in soil β-glucosidase activity and C and N pools over the course of this study (Fig. 1; Tables S1 & S2). Soil β-glucosidase is an extracellular enzyme produced by many soil microbes to break down cellulose, the primary component of most plant materials, making it a valuable indicator for tracking rhizosphere induced priming activity (Chang et al. 2007). For example, Zhu et al. (2014) observed 19–56% increases in β-glucosidase activity in the presence of living sunflower and soybean roots that were positively correlated with gross N mineralization rates in soil. While there were no differences in ß-glucosidase activity at 6 weeks after seeding in this study when the carrots were still very small, 8503 had lower total soil N and a higher soil C:N ratio, indicating that it was better at obtaining N from more easily degraded organic matter pools. As the experiment progressed, differences in soil ß-glucosidase activity and C and N pools became much more dramatic providing stronger evidence of varietal differences in N mineralization processes (Fig. 1; Tables S1 & S2). For example, at 10 weeks after planting, carrot genotype 8503 had the highest ß-glucosidase activity and soil C:N ratio, supporting our assertion that this genotype was most effective at stimulating organic matter decomposition and obtaining N from organic residues. Genotypes 3999 and Uppercut also had higher ß-glucosidase activity and Uppercut had lower NH4-N at this time point, indicating that these genotypes were also becoming more effective than genotypes like Karotan, which had the highest levels of NH4-N, and thus lowest uptake. Finally, since Sun255 had higher soil C and N than the other treatments, we suspect that this genotype had the lowest capacity to promote soil priming processes.
One of the reasons that experimental genotype 8503 may have had greater potential to promote decomposition and obtain N from organic materials in this study was that it had a moderately higher root:shoot ratio (although it was non-significant, p < 0.10) than the other genotypes (Table S3). This phenomenon has been demonstrated in wheat, where genotypes with a greater investment in belowground structures responded more positively to compost additions and had greater N uptake from these materials (Junaidi et al. 2018). A higher investment in root biomass may allow for greater exudation rates of low molecular weight substrates from plants roots that are critical for promoting positive rhizosphere priming processes (Pausch et al. 2016). Differences in root architecture and root plasticity have also been reported to enhance the potential for crop varieties to obtain N from decaying organic materials in soil (Andresen et al. 2016; Junaidi et al. 2018). This was not likely to be the case in this study since carrots were grown in pots where root spread was limited, however, future studies should consider exploring this mechanisms in larger pots and/or field studies, since it can also be an important factor influencing differences in NUE among crop genotypes.
While many studies have focused on quantifying changes in broad soil microbial processes such as ß-glucosidase activity to evaluate priming processes, it is also important to consider changes in specific microbial communities. For example, Fontaine et al. (2003) pointed out that many soil microbes are r-strategists that are adapted to decomposing labile and nutrient-rich substrates, and these microbes proliferate rapidly when labile C compounds such as root exudates are introduced into soil. In contrast, other soil microbes are k-strategists that are more efficient at depolymerizing and degrading more recalcitrant or specific organic substrates, and it is the competition between microbes with these two diverse strategies that actually contributes to priming processes in soil. In this study, we observed distinct differences in β diversity among soil bacteria associated with carrot plants in comparison with the unplanted control, as well as lower α diversity, providing evidence that carrot plants do recruit and support specific microbial taxa from soil (Fig. 3b). This was not surprising, given that the composition of rhizosphere microbial communities often represents a subset of microbes in bulk soil since only a subset of the soil microbial community can respond to and effectively compete for the labile carbon compounds released by plant roots. For example, Hargreaves et al. (2015) observed distinct differences in microbial communities inhabiting bulk soil and the rhizosphere of switchgrass (Panicum virgatum L.). Moreover, they observed greater abundances of taxa known to be active in degrading cellulose and chitin in the switchgrass rhizosphere, indicating that switchgrass plants can promote positive priming of organic materials by recruiting and supporting the growth of these specific taxa. Similarly, Turner et al. (2013) identified distinct differences in rhizosphere community structure between pea (Pisum sativum L.) and wheat plants along with greater concentrations of bacteria with cellulolytic activity in the rhizospheres of wheat plants, which do not host symbiotic microbes that can fix atmospheric N in nodules like pea. Consequently, we suspect that the unique microbial taxa recruited by carrot plants in this study could play a role in helping carrot plants promote soil priming processes to obtain N.
Previous studies have demonstrated that the composition of plant root exudates can vary among plant genotypes (Micallef et al. 2009), and this can in turn alter soil microbial community structure and influence organic matter decomposition rates (Zwetsloot et al. 2020). While explicitly quantifying differences in root exudates was beyond the scope and resources available for this study, we suspect that differences in the composition of root exudates among the carrot genotypes evaluated likely contributed to differences in the soil bacterial families observed (Fig. 2; Table S5), and this could have influenced decomposition rates. For example, soil bacterial communities associated with 8503 and 3999, which both had greater soil ß-glucosidase activity (Fig. 1; Tables S1 & S2) and greater %N in their shoot tissues at harvest (Table 4), clustered together and were clearly separate from Uppercut, Sun255 and Karotan (Fig. 3b). Moreover, 8503 was the only genotype to contain abundant populations of Micromonosporaceae. Many species of these bacteria have been noted for their ability to degrade chitin, cellulose, lignin and pectin, and play important roles in the turnover of organic materials in soil (Trujillo et al. 2014). Thus, we suspect that these taxa could have played a role in the greater soil β-glucosidase activity observed with this genotype. Populations of Rhodospirillaceae, which have potential to fix atmospheric N (Madigan et al. 1984) were enriched in 3999, indicating that these bacteria could have aided in the potential for this genotype to obtain greater concentrations of N in its tissues at harvest. Microbes belonging to the Chromatiaceae were enriched in 8503, 3999 and Sun255. These bacteria are well known for their role in the sulfur cycle and some species such as Nitrosococcus, are nitrifiers. While most Chromatiaceae are phototrophic, there is evidence that they can colonize plant rhizospheres by switching their growth strategies to chemo-, organo-, and/or mixo-trophy (Cúcio et al. 2016). Consequently, it is possible that microbes belonging to this family also aided in the potential for these carrot genotypes to obtain critical plant nutrients in this experiment.
Interestingly, other bacterial taxa were enriched in the soils of carrot genotypes that did not appear to be as effective at promoting C and N cycling, indicating that these microbes could have been better at immobilizing and preventing these genotypes from obtaining N. For example, Opitutaceae were enriched in Karotan, Sun255 and Uppercut. These bacteria belong to the phylum Verrucomicrobia. The ecological role of bacteria belonging to this phylum is still not well understood, though they have been negatively correlated with soil fertility (Navarrete et al. 2015). Other studies investigating relationships between rhizosphere bacterial community structure and N availability have noted that these microbes are a subset of microbial taxa that can tolerate low soil nutrients and persist in the rhizosphere until harvest (Bell et al. 2015). Consequently, they may be k-strategists that are better at decomposing more specific substrates. Interestingly, a recent study using 13C labeling of chitin residues demonstrated that while Opitutaceae did not appear to degrade chitin directly, they did become enriched in 13C at later time points indicating that they may have been utilizing products of the chitin degraders, such as extracellular polysaccharides (Wieczorek et al. 2019). The genotypes Karotan, Sun255 and Uppercut also had a lower relative abundance of microbes from the family Nitrosomonadaceae. These are ammonia oxidizers that can rapidly transform ammonia to nitrate (Thion et al. 2016).
Consequently, having a lower relative abundance of these microbes could reduce the capacity for these genotypes to obtain nitrate relative to 8503 and 3999.
To better estimate whether there were differences in the potential for genotype-induced differences in the soil bacterial communities to cycle C and N in this study, we used PICRUSt2 to quantify differences in the potential abundance of genes involved in cellulose degradation and N metabolism (Fig. 4; Table S6). Interestingly, genes involved in β -glucosidase were enriched in 8503, 3999 and Uppercut, supporting our assertions that these genotypes can better stimulate decomposition of organic materials in soil. Moreover, none of the genes involved in cellulose degradation or N metabolism were enriched in Karotan, which appeared to have the lowest priming activity in this study (Hartman et al. 2018). In addition to facilitating decomposition, plants can also recruit and support microbes that mineralize and release plant available N (Rudisill et al. 2016). In this study, many genes associated with N metabolism were more associated with 3999 and 8503, providing evidence that these genotypes might be better able to recruit and support bacteria that can contribute to NUE. Finally, it is interesting to note that genes associated with cellobiose degradation were enriched in Sun255 and Uppercut, which were also enriched with microbial taxa from the family Opitutaceae. Since these microbes are suspected to play a role in degrading substrates produced by other microbes rather than directly degrading complex substrates on their own (Wieczorek et al. 2019), this may help explain why these genotypes were less effective in stimulating positive priming processes in this study. However, it is important to evaluate these results with caution, since the potential presence of the gene does not always correlate with enhanced gene expression (Zheng et al. 2018).
Finally, it is important to point out that differences in soil microbial communities induced by the carrot genotypes in this study (Fig. 2; Table S5), could play other important roles in carrot production systems such as resistance to root-knot nematodes. Several studies have identified distinct microbiomes associated with resistant and susceptible cucumber and tomato genotypes (Kwak et al. 2018; Upreti and Thomas 2015; Yao and Wu 2010), and we recently discovered this phenomenon in carrot (Abdelrazek et al. 2020a, 2020b). The exact cause of the genetic resistance to root-knot nematodes in genotypes like 3999 and 8503 is not yet clear (Williamson et al. 2009), though it is possible that these genotypes are able to sense the presence of these nematodes at the root surface and strengthen their cell walls to resist invasion in class hypersensitive response. We have theorized that these genotypes may also be able to host endophytic microbes within carrot taproots that help prevent root-knot nematodes from infecting carrot plants (Abdelrazek et al. 2020a). In contrast, Sun255, Karotan and Uppercut may need to mount alternative strategies to resist attack by pathogenic nematodes, such as recruiting microbes with antagonistic activity against pathogens in their rhizosphere. Interestingly, taxa belonging to the Oxalobacteriaceae and Micrococcaceae, which were enriched in the soil of these three genotypes, are present in elevated concentrations in so-called ‘disease suppressive soils’ (Cretoiu et al. 2013; van der Voort et al. 2016). Consequently, it is possible that these microbes could help these genotypes fight pathogens, and this should be explored in future studies.
Conclusions
Results of this study confirm previous reports that carrots are efficient at obtaining N in soil (Thorup-Kristensen 2006; Veitch et al. 2014; Westerveld et al. 2006), and at least part of this activity could be due to bacterial-induced priming processes. The results also confirm previous reports that at least part of the composition and functional potential of soil microbial communities associated with carrot roots are under genetic control (Abdelrazek et al. 2020a; 2020b). In addition to serving as a first line of defense against invading pathogens as previously reported, results of this study indicate that changes in bacterial community structure induced by carrot genotypes also have potential to influence nutrient use efficiency in carrot crops. Finally, carrot genotypes appear to differ in their capacity to promote decomposition of plant residues to obtain at least part of their nitrogen needs. By learning more about the genetic mechanisms regulating these relationships, it may someday be possible to leverage these beneficial plant-soil-microbial interactions in crop breeding programs (Hoagland 2015). Currently most plant breeding programs are conducted under conventional farming systems where plants are selected for high yield in the presence of inorganic fertility programs. As a result, researchers have theorized that varieties developed in conventional systems may not have the traits needed to interact with soil microbes to obtain nutrients from organic sources (Lammerts van Bueren and Struik 2017; Mwafulirwa et al. 2016). For example, it is energetically expensive for plants to provide C resources to support soil microbes, and thus high rates of inorganic N fertilizers could disrupt relationships between plants and microbes that play a role in organic matter decomposition (Heijboer et al. 2016). In contrast, selecting varieties in organic farming systems where plants are more reliant on organic materials to obtain nutrients could lead to the development of varieties that promote organic matter priming. Consequently, it is tempting to speculate that our organic breeding program has selected for genotypes that are better at obtaining nutrients from organic materials, since experimental genotypes 8503 and 3999 have been performing well in our organic field trials (Simon et al. 2017). However, future studies conducted under a diverse set of environmental conditions will be needed to confirm whether this is the case. Since fungi also play a critical role in the decomposition of organic materials in soils, future studies should also investigate genotype-induced changes in these communities as well. Regardless of the approach, researchers need to improve understanding of how plants interact with their microbiomes to address environmental challenges and improve productivity and NUE in agricultural systems (Vandenkoornhuyse et al. 2015).
Data Availability
The sequencing data have been deposited with the BioProject accession number PRJNA714923 in the NCBI BioProject database (https://www.ncbi.nlm.nih.gov/bioproject/). The datasets analysed during the current study are available from the corresponding author upon request.
References
Abdelrazek S, Choudhari S, Thimmapuram J et al (2020a) Changes in the core endophytic mycobiome of carrot taproots in response to crop management and genotype. Sci Rep 10:13685. https://doi.org/10.1038/s41598-020-70683-x
Abdelrazek S, Simon P, Colley M et al (2020b) Crop management system and carrot genotype affect endophyte composition and Alternaria dauci suppression. PLoS One 15:e0233783. https://doi.org/10.1371/journal.pone.0233783
Andresen M, Dresbøll DB, Jensen LS et al (2016) Cultivar differences in spatial root distribution during early growth in soil, and its relation to nutrient uptake - a study of wheat, onion and lettuce. Plant Soil 408:255–270. https://doi.org/10.1007/s11104-016-2932-z
Andrews S (2010) FastQC: A quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/. Accessed 5 Jan 2021
Bais HP, Weir TL, Perry LG et al (2006) The role of root exudates in rhizosphere interactions with plants and other organisms. Annu Rev Plant Biol 57:233–266. https://doi.org/10.1146/annurev.arplant.57.032905.105159
Bardgett RD, Mommer L, De Vries FT (2014) Going underground: Root traits as drivers of ecosystem processes. Trends Ecol Evol 29:692–699. https://doi.org/10.1016/j.tree.2014.10.006
Baresel JP, Zimmermann G, Reents HJ (2008) Effects of genotype and environment on N uptake and N partition in organically grown winter wheat (Triticum aestivum L.) in Germany. Euphytica 163:347–354. https://doi.org/10.1007/s10681-008-9718-1
Bateman AS, Kelly SD, Jickells TD (2005) Nitrogen isotope relationships between crops and fertilizer: Implications for using nitrogen isotope analysis as an indicator of agricultural regime. J Agric Food Chem 53:5760–5765. https://doi.org/10.1021/jf050374h
Bell CW, Asao S, Calderon F et al (2015) Plant nitrogen uptake drives rhizosphere bacterial community assembly during plant growth. Soil Biol Biochem 85:170–182. https://doi.org/10.1016/j.soilbio.2015.03.006
Blagodatskaya E, Littschwager J, Lauerer M, Kuzyakov Y (2014) Plant traits regulating N capture define microbial competition in the rhizosphere. Eur J Soil Biol 61:41–48. https://doi.org/10.1016/j.ejsobi.2014.01.002
Blagodatsky S, Blagodatskaya E, Yuyukina T, Kuzyakov Y (2010) Model of apparent and real priming effects: Linking microbial activity with soil organic matter decomposition. Soil Biol Biochem 42:1275–1283. https://doi.org/10.1016/j.soilbio.2010.04.005
Bolyen E, Rideout JR, Dillon MR et al (2019) QIIME 2: Reproducible, interactive, scalable, and extensible microbiome data science. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Braker G, Tiedje JM (2003) Nitric oxide reductase (norB) genes from pure cultures and environmental samples. Appl Environ Microbiol 69(6):3476–3483
Boskovic-Rakocevic L, Pavlovic R, Zdravkovic J et al (2012) Effect of nitrogen fertilization on carrot quality. Afr J Agric Res 7:2884–2900. https://doi.org/10.5897/ajar11.1652
Bottomley PS, Angle S, Weaver RW (1994) Methods of soil analysis, Part 2: Microbiological and Biochemical Properties. American Society of Agronomy. https://doi.org/10.2136/sssabookser5.2
Carr PM, Cavigelli MA, Darby H et al (2020) Green and animal manure use in organic field crop systems. Agron J 112:648–674. https://doi.org/10.1002/agj2.20082
Chang E-H, Chung R-S, Tsai Y-H (2007) Effect of different application rates of organic fertilizer on soil enzyme activity and microbial population. Soil Sci Plant Nutr 53:132–140. https://doi.org/10.1111/j.1747-0765.2007.00122.x
Cretoiu MS, Korthals GW, Visser JHM, Van Elsas JD (2013) Chitin amendment increases soil suppressiveness toward plant pathogens and modulates the actinobacterial and oxalobacteraceal communities in an experimental agricultural field. Appl Environ Microbiol 79:5291–5301. https://doi.org/10.1128/AEM.01361-13
Cúcio C, Engelen AH, Costa R, Muyzer G (2016) Rhizosphere microbiomes of European + seagrasses are selected by the plant, but are not species specific. Front Microbiol 7:440. https://doi.org/10.3389/fmicb.2016.00440
De Cáceres M, Legendre P (2009) Associations between species and groups of sites: Indices and statistical inference. Ecology 90:3566–3574. https://doi.org/10.1890/08-1823.1
de Vries FT, Wallenstein MD (2017) Below-ground connections underlying above-ground food production: a framework for optimising ecological connections in the rhizosphere. J Ecol 105:913–920. https://doi.org/10.1111/1365-2745.12783
de Oliveira SA, Camberato JJ, Coram T, Filley T, Vyn TJ (2017) Applicability of a “multi-stage pulse labeling” 15N approach to phenotype n dynamics in maize plant components during the growing season. Front Plant Sci 8:1360. https://doi.org/10.3389/fpls.2017.01360
Dijkstra FA, Carrillo Y, Pendall E, Morgan JA (2013) Rhizosphere priming: A nutrient perspective. Front Microbiol 4:1–8. https://doi.org/10.3389/fmicb.2013.00216
Eivazi F, Tabatabai MA (1988) Glucosidases and galactosidases in soils. Soil Biol Biochem 20:601–606. https://doi.org/10.1016/0038-0717(88)90141-1
Erisman JW, Galloway JN, Seitzinger S et al (2013) Consequences of human modification of the global nitrogen cycle. Philos Trans R Soc B Biol Sci 368:20130116. https://doi.org/10.1098/rstb.2013.0116
Fageria NK, Baligar VC (2005) Enhancing nitrogen use efficiency in crop plants. Adv Agron 88:97–185. https://doi.org/10.1016/S0065-2113(05)88004-6
Fan TW, Conn EE (1985) Isolation and characterization of two cyanogenic beta-glucosidases from flax seeds. Arch Biochem Biophys 243(2):361–373. https://doi.org/10.1016/0003-9861(85)90513-2
Fauré J, Dagher MC (2001) Interactions between Rho GTPases and Rho GDP dissociation inhibitor (Rho-GDI). Biochimie 83(5):409–414. https://doi.org/10.1016/S0300-9084(01)01263-9
Finzi AC, Abramoff RZ, Spiller KS et al (2015) Rhizosphere processes are quantitatively important components of terrestrial carbon and nutrient cycles. Glob Chang Biol 21:2082–2094. https://doi.org/10.1111/gcb.12816
Fontaine S, Mariotti A, Abbadie L (2003) The priming effect of organic matter: A question of microbial competition? Soil Biol. Biochem 35:837–843. https://doi.org/10.1016/S0038-0717(03)00123-8
Frouz J (2018) Effects of soil macro- and mesofauna on litter decomposition and soil organic matter stabilization. Geoderma 332:161–172. https://doi.org/10.1016/j.geoderma.2017.08.039
Gussin GN, Ronson CW, Ausubel FM (1986) Regulation of nitrogen fixation genes. Ann Rev Genet 20:567–591
Hargreaves SK, Williams RJ, Hofmockel KS (2015) Environmental filtering of microbial communities in agricultural soil shifts with crop growth. PLoS One 10:e0134345. https://doi.org/10.1371/journal.pone.0134345
Hartman K, van der Heijden MGA, Wittwer RA et al (2018) Cropping practices manipulate abundance patterns of root and soil microbiome members paving the way to smart farming. Microbiome 6:1–14. https://doi.org/10.1186/s40168-017-0389-9
Hasper AA, Dekkers E, van Mil M, van de Vondervoort PJ, de Graaff LH (2002) EglC, a new endoglucanase from Aspergillus niger with major activity towards xyloglucan. Appl Environ Microbiol 68(4):1556–1560. https://doi.org/10.1128/AEM.68.4.1556-1560
Heijboer A, ten Berge HFM, de Ruiter PC et al (2016) Plant biomass, soil microbial community structure and nitrogen cycling under different organic amendment regimes; a 15N tracer-based approach. Appl Soil Ecol 107:251–260. https://doi.org/10.1016/j.apsoil.2016.06.009
Hirel B, Tétu T, Lea PJ, Dubois F (2011) Improving nitrogen use efficiency in crops for sustainable agriculture. Sustainability 3:1452–1485. https://doi.org/10.3390/su3091452
Hoagland L, Carpenter-Boggs L, Granatstein D et al (2008) Orchard floor management effects on nitrogen fertility and soil biological activity in a newly established organic apple orchard. Biol Fertil Soils 45:11–18. https://doi.org/10.1007/s00374-008-0304-4
Hoagland L (2015) Breeding for beneficial plant-microbial relationships: how do we get there? Proceedings of the Eucarpia Workshop on Implementing Plant-Microbe Interaction in Plant Breeding, Weihenstephan
Junaidi J, Kallenbach CM, Byrne PF, Fonte SJ (2018) Root traits and root biomass allocation impact how wheat genotypes respond to organic amendments and earthworms. PLoS ONE 13:1–24. https://doi.org/10.1371/journal.pone.0200646
Kim GS, Seok JH, Mark TB, Reed MR (2019) The price relationship between organic and non-organic vegetables in the U.S.: Evidence from Nielsen scanner data. Appl Econ 51:1025–1039. https://doi.org/10.1080/00036846.2018.1524570
Krueger F (2015) Wrapper tool around Cutadapt and FastQC to consistently apply quality and adapter trimming to FastQ files. The Babraham Institute, Bioinformatics Group. https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/. Accessed 5 Jan 2021
Kuzyakov Y (2002) Review: Factors affecting rhizosphere priming effects. J. Plant. Nutr Soil Sci-Zeitschrift Fur Pflanzenernahrung Und Bodenkunde 165:382–396. https://doi.org/10.1002/1522-2624(200208)165:4%3c382::AID-JPLN382%3e3.0.CO;2-#
Kwak M-J, Kong HG, Choi K et al (2018) Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat. Biotechnol 36. https://doi.org/10.1038/nbt.4232
Laberge G, Ambus P, Hauggaard-Nielsen H, Jensen ES (2006) Stabilization and plant uptake of N from 15N-labelled pea residue 16.5 years after incorporation in soil. Soil Biol Biochem 38:1998–2000. https://doi.org/10.1016/j.soilbio.2005.11.023
Lammerts van Bueren ET, Struik PC (2017) Diverse concepts of breeding for nitrogen use efficiency. A review. Agron Sustain Dev 37. https://doi.org/10.1007/s13593-017-0457-3
Langille MGI, Zaneveld J, Caporaso JG et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Lucier G, Lin BH (2007) Factors affecting carrot consumption in the United States. Eating right: The consumption of fruits and vegetables 199–220. https://www.ers.usda.gov/webdocs/outlooks/39511/11719_vgs31901_1_.pdf?v=9138
Lynch JP (2013) Steep, cheap and deep: An ideotype to optimize water and N acquisition by maize root systems. Ann Bot 112:347–357. https://doi.org/10.1093/aob/mcs293
Madigan M, Cox SS, Stegeman RA (1984) Nitrogen fixation and nitrogenase activities in members of the family Rhodospirillaceae. J Bacteriol 157:73–78. https://doi.org/10.1128/jb.157.1.73-78.1984
Meier IC, Finzi AC, Phillips RP (2017) Root exudates increase N availability by stimulating microbial turnover of fast-cycling N pools. Soil Biol Biochem 106:119–128. https://doi.org/10.1016/j.soilbio.2016.12.004
Micallef SA, Shiaris MP, Colón-Carmona A (2009) Influence of Arabidopsis thaliana accessions on rhizobacterial communities and natural variation in root exudates. J Exp Bot 60:1729–1742. https://doi.org/10.1093/jxb/erp053
Moore, GA (2001) Soilguide (Soil guide) : a handbook for understanding and managing agricultural soils. Department of Primary Industries and Regional Development. Western Australia, Perth. Bulletin 4343.
Moreau D, Bardgett RD, Finlay RD et al (2019) A plant perspective on nitrogen cycling in the rhizosphere. Funct Ecol 33:540–552. https://doi.org/10.1111/1365-2435.13303
Murphy CJ, Baggs EM, Morley N et al (2015) Rhizosphere priming can promote mobilisation of N-rich compounds from soil organic matter. Soil Biol Biochem 81:236–243. https://doi.org/10.1016/j.soilbio.2014.11.027
Mwafulirwa L, Baggs EM, Russell J et al (2016) Barley genotype influences stabilization of rhizodeposition-derived C and soil organic matter mineralization. Soil Biol Biochem 95:60–69. https://doi.org/10.1016/j.soilbio.2015.12.011
Mwafulirwa LD, Baggs EM, Russell J et al (2017) Combined effects of rhizodeposit C and crop residues on SOM priming, residue mineralization and N supply in soil. Soil Biol Biochem 113:35–44. https://doi.org/10.1016/j.soilbio.2017.05.026
Navarrete AA, Soares T, Rossetto R et al (2015) Verrucomicrobial community structure and abundance as indicators for changes in chemical factors linked to soil fertility. Antonie Van Leeuwenhoek 108:741–752. https://doi.org/10.1007/s10482-015-0530-3
Oakley BB, Francis CA, Roberts KJ, Fuchsman CA, Srinivasan S, Staley JT (2007) Analysis of nitrite reductase (nirK and nirS) genes and cultivation reveal depauperate community of denitrifying bacteria in the Black Sea suboxic zone. Environ Microbiol 9(1):118–130
Oksanen J (2007) vegan : Community Ecology Package. R package version 1.8–5
Paterson E, Baggs EM, de la Fuente CC et al (2016) Barley genotype influences stabilization of rhizodeposition-derived C and soil organic matter mineralization. Soil Biol Biochem 95:60–69. https://doi.org/10.1016/j.soilbio.2015.12.011
Pausch J, Loeppmann S, Kühnel A et al (2016) Rhizosphere priming of barley with and without root hairs. Soil Biol Biochem 100:74–82. https://doi.org/10.1016/j.soilbio.2016.05.009
Poirier V, Roumet C, Munson AD (2018) The root of the matter: Linking root traits and soil organic matter stabilization processes. Soil Biol Biochem 120:246–259. https://doi.org/10.1016/j.soilbio.2018.02.016
Pruesse E, Quast C, Knittel K et al (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196. https://doi.org/10.1093/nar/gkm864
R Development Core Team (2019) A language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria
Reid JB, Hunt AG, Johnstone PR et al (2017) On the responses of carrots ( Daucus carota L.) to nitrogen supply. N. Z. J. Crop Hortic. Sci 0:1–21. https://doi.org/10.1080/01140671.2017.1402790
Robinson MD, McCarthy DJ, Smyth GK (2009) edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26:139–140. https://doi.org/10.1093/bioinformatics/btp616
Rudisill MA, Turco RF, Hoagland LA (2016) Fertility practices and rhizosphere effects alter ammonia oxidizer community structure and potential nitrification activity in pepper production soils. Appl Soil Ecol 99:70–77. https://doi.org/10.1016/j.apsoil.2015.10.011
Sasse J, Martinoia E, Northen T (2018) Feed your Friends: Do plant exudates shape the root microbiome? Trends Plant Sci 23:25–41. https://doi.org/10.1016/j.tplants.2017.09.003
Sayavedra-Soto LA, Hommes NG, Arp DJ (1994) Characterization of the gene encoding hydroxylamine oxioreducase in Nitrosomonas europaea. J Bacteriol 176(2):504–510. https://doi.org/10.1128/jb.176.2.504-510
Schenk HJ (2006) Root competition: Beyond resource depletion. J Ecol 94:725–739. https://doi.org/10.1111/j.1365-2745.2006.01124.x
Smith CJ, Nedwell DB, Dong LF, Osborn AM (2007) Diversity and abundance of nitrate reductase genes (narG and napA), nitrite reductase genes (nirS and nrfA), and their transcripts in estuarine sediments. Appl Environ Microbiol 73(11):3612–l
Simon PW, Navazio JP, Colley M et al (2017) The CIOA (Carrot Improvement for Organic Agriculture) project: location, cropping system and genetic background influence carrot performance including top height and flavour. Acta Hortic 1153:1–8. https://doi.org/10.17660/ActaHortic.2017.1153.1
Sinigalliano CD, Kuhn DN, Jones RD (1995) Amplfication of the amoA gene from diverse species of ammonium-oxidizing bacteria and from an indigenous bacterial population from seawater. Appl Environ Microbiol 61(7):2702–2706
Terrazas RA, Pietrangelo L, Corral AM et al (2019) Nitrogen availability modulates the host control of the barley rhizosphere microbiota. bioRxiv 605204. https://doi.org/10.1101/605204
Thion CE, Poirel JD, Cornulier T et al (2016) Plant nitrogen-use strategy as a driver of rhizosphere archaeal and bacterial ammonia oxidiser abundance. FEMS Microbiol Ecol 92:fiw091. https://doi.org/10.1093/femsec/fiw091
Thompson J, Ruvinov SB, Freedberg DI, Hall BG (1999) Cellobiose-6-phosphate hydrolase (CelF) of Escherichia coli: characterization and assignment to the unusual family 4 of glycosylhydrolase. J Bacteriol 181:7339–7345. https://doi.org/10.1128/JB.181.23.7339-7345.1999
Thorup-Kristensen K (2006) Root growth and nitrogen uptake of carrot, early cabbage, onion and lettuce following a range of green manures. Soil Use Manag 22:29–38. https://doi.org/10.1111/j.1475-2743.2005.00012.x
Trujillo ME, Hong K, Genilloud O (2014) The family Micromonosporaceae. In: The Prokaryotes: Actinobacteria. Springer-Verlag Berlin Heidelberg, pp 499–569 https://doi.org/10.1007/978-3-642-30138-4_196
Turner TR, Ramakrishnan K, Walshaw J et al (2013) Comparative metatranscriptomics reveals kingdom level changes in the rhizosphere microbiome of plants. ISME J 7:2248–2258. https://doi.org/10.1038/ismej.2013.119
Upreti R, Thomas P (2015) Root-associated bacterial endophytes from Ralstonia solanacearum resistant and susceptible tomato cultivars and their pathogen antagonistic effects. Front Microbiol 6. https://doi.org/10.3389/fmicb.2015.00255
Van Cleemput O, Zapata F, Vanlauwe B (2008) Use of tracer technology in mineral fertilizer management. In: Guidelines on Nitrogen Management in Agricultural Systems 19–125. International Atomic Energy Agency, Vienna, Austria.
van der Voort M, Kempenaar M, van Driel M et al (2016) Impact of soil heat on reassembly of bacterial communities in the rhizosphere microbiome and plant disease suppression. Ecol Lett 19:375–382. https://doi.org/10.1111/ele.12567
Vandenkoornhuyse P, Quaiser A, Duhamel M et al (2015) The importance of the microbiome of the plant holobiont. New Phytol 206:1196–1206. https://doi.org/10.1111/nph.13312
Vázquez-Garcidueñas S, Leal-Morales CA, Herrera-Estrella A (1998) Analysis of the beta-1,3-Glucanolytic System of the Biocontrol Agent Trichoderma harzianum. Appl Environ Microbiol 64(4):1442–1446. https://doi.org/10.1128/AEM.64.4.1442-1446
Veitch RS, Lada RR, Adams A, MacDonald MT (2014) Carrot yield and quality as influenced by nitrogen application in cut-and-peel carrots. Commun Soil Sci Plant Anal 45:887–895. https://doi.org/10.1080/00103624.2014.880713
Westerveld SM, McKeown AW, McDonald MR (2006) Relationship between nitrogen application and alternaria leaf blight [Alternaria dauci] and cercospora leaf blight [Cercospora carotae] of carrots. HortScience 46:1522–1527. https://doi.org/10.21273/hortsci.43.5.1522
Welsh A, Chee-Sanford JC, Connor LM, Löffler FE, Sanford RA (2014) Refined NrfA phylogeny improves PCR based NrfA gene detection. Appl Environ Microbiol 80(7):2110–2119
Wieczorek AS, Schmidt O, Chatzinotas A et al (2019) Ecological functions of agricultural soil bacteria and microeukaryotes in chitin degradation: A case study. Front Microbiol 10:1293. https://doi.org/10.3389/fmicb.2019.01293
Williamson VM, Roberts PA, Perry RN (2009) Root-knot nematodes. In: Mechanisms and genetics of resistance. Root-knot nematodes. p 301
Yao H, Wu F (2010) Soil microbial community structure in cucumber rhizosphere of different resistance cultivars to fusarium wilt. FEMS Microbiol Ecol 72:456–463. https://doi.org/10.1111/j.1574-6941.2010.00859.x
Yeoman CJ, Han Y, Dodd D, Schroeder CM, Mackie RI, Cann IK (2010) Thermostable enzymes as biocatalysts in the biofuel industry. Adv Appl microbiol 70:1–55
Yin L, Dijkstra FA, Wang P et al (2018) Rhizosphere priming effects on soil carbon and nitrogen dynamics among tree species with and without intraspecific competition. New Phytol 218:1036–1048. https://doi.org/10.1111/nph.15074
Yu WL, Jiang YL, Pikis A, Cheng W, Bai XH, Ren YM, Thompson J, Zhou CZ, Chen Y (2013) Structural insights into the substrate specificity of a 6-phospho-β-glucosidase BglA-2 from Streptococcus pneumoniae TIGR4. J Biol Chem 288(21):14949–14958. https://doi.org/10.1074/jbc.M113.454751
Zheng W, Zhao Z, Gong Q et al (2018) Effects of cover crop in an apple orchard on microbial community composition, networks, and potential genes involved with degradation of crop residues in soil. Biol Fertil 54:743–759. https://doi.org/10.1007/s00374-018-1298-1
Zhu B, Gutknecht JLM, Herman DJ et al (2014) Rhizosphere priming effects on soil carbon and nitrogen mineralization. Soil Biol Biochem 76:183–192. https://doi.org/10.1016/j.soilbio.2014.04.033
Zumft WG, Dreusch A, Löchelt S, Cuypers H, Friedrich B, Schneider B (1992) Derived amino acid sequences of the nosZ gene (respiratory N2O reductase) from Alcaligenes eutrophus, Pseudomonas aeruginosa and Pseudomonas stutzeri reveal potential copper-binding residues: Implications for the CuA site of N2O reductase and cytochrome-c oxidase. Eur J Biochemi 208(1):31–40
Zwetsloot MJ, Ucros JM, Wickings K et al (2020) Prevalent root-derived phenolics drive shifts in microbial community composition and prime decomposition in forest soil. Soil Biol. Biochem 145:107797. https://doi.org/10.1016/j.soilbio.2020.107797
Acknowledgements
We would like to express our appreciation to Mr. Nathan Deppe for his assistance in setting up and management of the greenhouse trials, as well as Ms. Leidy Mejia, who assisted with harvest and processed soil samples. We also thank Ms. Nicole S. De Armond in the Purdue Agronomy Department and Ms. Janine Sparks in the Purdue Stable Isotope Facility for their assistance with laboratory assays.
Funding
This project was funded by the NIFA-OREI program through grant #2016–51300-25721, as well as NIFA-Hatch project #’s 1007553 and 1015999.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Lori Hoagland, Alejandro Rodriguez-Sanchez and Narda J Triviño. The first draft of the manuscript was written by Narda J Triviño and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible Editor: Johannes Postma.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Triviño, N.J., Rodriguez-Sanchez, A., Filley, T. et al. Carrot genotypes differentially alter soil bacterial communities and decomposition of plant residue in soil. Plant Soil 486, 587–606 (2023). https://doi.org/10.1007/s11104-023-05892-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-05892-0