Abstract
Purpose
A large fraction of a plant’s biomass is belowground, especially in shrublands that typically occur in episodically water-limited climates. Nonetheless, we have no standardized method to map the distribution of the root density (i.e., biomass per soil volumetric unit) of plant individuals (hereafter, Individual-level Root Density Distribution, IRDD). This type of information is difficult to collect, especially in woody plant communities in natural conditions where roots of different individuals can be highly intermingled.
Methods
We assess three methods to map IRDD of field shrubs: soil drilling to extract roots, plant injection with dyes, and microsatellite analysis for individual-level root identification. Using the resulting data, we fitted IRDD models obtaining comparable predictions of the root density of shrubs for each method.
Results
The proportion of identified roots was higher using plan injection, but the cost per linked roots was two orders of magnitude higher using microsatellite. Model results show that microsatellite markers had a similar success as compared to plant injection for those plant individuals for which it worked well, but it failed completely for several genotypes or individuals.
Conclusions
Core drilling machines and plant injection with dyes of different colors to link root fragments from the sample pool to plant individuals represent an affordable, reliable way to study the foraging behavior of woody plants which roots are highly intermingled.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Shrublands cover about 7–8% of Earth’s land surface (Lal 2004; Maestre et al. 2021), and are expanding because of anthropogenic desertification (Eldridge et al. 2011; Huang et al. 2020). Shrubs are an important functional type of vegetation amid largely undervalued by humankind (Tomaselli 1977; Kemper et al. 1999) and understudied by ecologists, as compared to forest and grassland systems (Wullschleger et al. 2014; Fusco et al. 2019; Schrader-Patton and Underwood 2021). Many shrubs grow in resource-poor soils, and allocate much of their biomass to their roots to acquire water and soluble nutrients (Schenk and Jackson 2002; Bardgett et al. 2014). As most plants, they absorb through their roots and compete for water, nitrogen, phosphorus, and many other soil resources that are subsequently transported to the leaves to photosynthesize (Lambers et al. 2008; Kirkham 2014). Compared to aboveground plant organs, we know very little about roots, since roots cannot be directly observed and are very difficult to measure (Jones et al. 2011; Lux and Rost 2012). Additionally, most of what we know about plant roots come from experiments conducted in controlled conditions, yet rooting behavior of plants may be very different in natural conditions (Poorter et al. 2016). Developing, testing, and standardizing methodologies to measure root traits in the field is a salient pending task for ecologists to address (Addo-Danso et al. 2016).
While many root variables can be measured (Freschet et al. 2021a, b), a map of the distribution of root density in soil (i.e., biomass per soil volumetric unit) of plant individuals (hereafter, Individual-level Root Density Distribution, IRDD) would provide the most comprehensive ecological information about plant foraging strategies (Cabal et al. 2021). IRDD maps allow researchers to study the root density distribution of neighboring plants in the soil and, by integrating such densities in three-dimensional space, the plant allocation of biomass to belowground tissues (Cabal et al. 2020). Estimating an IRDD would require samples of roots from known spatial coordinates and depths, and the assignment of roots in the sample to the surrounding individual plants.
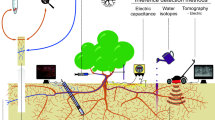
While several techniques exist to study roots in the field, such as the use of stable isotopes (Stahl et al. 2013), anatomical or chemical phenotype markers (Roumet et al. 2006; Lei and Bauhus 2010), tomography (Zenone et al. 2008; Weigand and Kemna 2017), or rhizotron systems (Arnaud et al. 2019) to name a few, only a few may allow researchers to map IRDD of plants from root mixture samples (Cabal et al. 2021). Core sampling allows obtaining root density information, as researchers can weight the extracted roots and know the volume of the sample. As for the identification of the roots at the individual-level (not the species-level), a few studies have used microsatellite analyses to link root samples to individual plants (Brunner et al. 2001; Saari et al. 2005; Lang et al. 2010). Dying plant roots with different colors by injecting dye into their aboveground stems might be a cheaper and easier alternative to DNA analysis. This technique has been used successfully in several experiments in controlled conditions, in plants grown in pots in the greenhouse (Murakami et al. 2006; Cahill et al. 2010; Cabal et al. 2020) and in tomato plants grown in outdoor containers (Murakami et al. 2011), but has never been used in woody plants in the field.
In this study, we evaluate three field methods to map IRDD in three mediterranean shrub species growing in central Spain. Firstly, we adapted a diamond core-drilling engine designed for the construction industry as a method for root extraction. Secondly, we compared two different root identification methods to link root fragments from soil samples to aboveground plants: microsatellite analyses and plant injection with dyes. Based in model predictions, we compare the root identification methods with each other to determine which is the most advisable to map the IRDD of plants in the field.
Materials and methods
Study site description
This study was carried out in a mediterranean shrubland in ‘Las Tejoneras’ (40°06′42.69″ N, 5°16′32.46″ W, 329 m.a.s.l.), Candeleda (Ávila, Spain), a small isolated granitic mountain that rises about 50 m above the surrounding plains. Leptosols and superficial bedrock are the dominant soils. The area presents a meso-mediterranean, sub-humid ombroclimate (Rivas-Matínez and Armaiz 1984) with characteristic arid summers, an annual precipitation of 797.9 mm, and a mean temperature of 16.17 °C during the last decade (from January 2010 to December 2019 data from the AEMET meteorological station in Candeleda, 40°08′21″ N, 5°18′41″ W, 350 m.a.s.l.). The vegetation growing in this terrain is a biodiverse mediterranean closed-canopy shrubland with over a dozen shrub species. Three dominant shrub species were selected: gum rockrose (Cistus ladanifer L.), rosemary (Salvia rosmarinus Schleid.), and hairy-fruited broom (Cytisus striatus [Hill] Rothm).
Field data collection
We selected seven 2 × 2 m plots based in the occurrence of individuals of the targeted species and in order to represent a range of plant sizes within each species from small to the largest in the region. We sampled one monospecific rosemary plot in the summer of 2018, accounting for all the 14 plants in and around the plot. Roots from this plot that could be linked to the species using DNA barcoding were identified using microsatellite markers (Segarra-Moragues and Gleiser 2009). We sampled six plots in the summer of 2019, with seven rosemary, eleven rockrose, and six broom plants of different sizes, whose roots were identified by root injection with dyes. In the latter six plots, a maximum of five focal individuals per plot were selected given that we used five different dye colors, but more individuals of the same and other species were found in and around the plot. We measured the total dry weight of photosynthetic and structural aboveground biomass of the selected focal plants from each plot.
Subsequently, we extracted soil cores of a maximum depth of 800 mm (minimum depth could be as low as 100 mm in the event of finding a large rock underground), and diameter of 104 mm, with a core drilling machine designed for construction, and adapted to the field (Fig. SM 1–3). We extracted a variable number of cores from each plot following a regular spatial pattern. We chose to sample following a regular pattern to represent all the parts of the plot and representing equally the different sides and distances from the focal plants. We obtained several soil samples from each core at different depths. We sifted soil samples using a 2 mm sieve, recovering mineral material (gravel and stones) whose volume was measured, and large root fragments whose diameter and dry weight was measured. We recovered the organic matter from the fraction of sample that passed through the sieve by flotation. Such organic matter was oven dried, we separated fine roots—all with mean diameter < 0.5 mm—from other materials, and weighed these roots in bulk.
We located the relative position of the insertion points of the stem of each focal plant to the soil surface (plant insertion) and the centroid of each cylinder-shaped soil sample (sample centroid) in the plot in three-dimensional space. To that end, we used a combination of drone photography, laser level measurements of the slope in the plot, and information about the minimum and maximum depth of each soil sample (Fig. SM 4–5).
Individual-level identification of roots
In the first plot, we analyzed 14 leaf samples representing the plant individuals and 904 root fragments extracted from 42 soil samples from 23 soil cores, linking roots to aboveground tissues using DNA analysis. Given the large number of collected root fragments, and the high cost of DNA analyses, we only retained root fragments with a diameter > 1 mm for the DNA analysis (415 root fragments). We analyzed the selected roots individually, following a dual approach. First, root DNA was isolated, and the region of the chloroplast ribulose-1,5-bisphosphate carboxylase/oxygenase large subunit (rbcL) gene was amplified by polymerase chain reaction (PCR) and sequenced. We compared the resulting sequences with barcode records available in two public reference databases to verify that the samples belonged to rosemary plants. Then, the roots with detectable rosemary barcodes (217 root fragments), together with the 14 leaf samples, were genotyped using six different microsatellite loci in four multiplex reactions.
In the six remaining plots, we injected 24 plants with dyes of different colors before soil coring. We extracted 2,840 root fragments from 216 soil samples in 81 soil cores and linked them to the plants based on the dye colors (Fig. SM 6). Because woody plants have never before been injected with dyes in order to link root fragments to plants, we developed a hydraulic model to estimate the velocity of dye in the stem vessels (\(\overrightarrow{{v}_{10}(f)}\)). This velocity is a function of the mean plant vessel diameter, the root length that the researcher wants to stain, the soil water potential, and the osmotic potential of the dyes used (Fig. SM 7). We obtained measures of the vessel diameter of the three focal species of our study and measured the osmotic potential of the dyes used with a psycrometer. The model assessment suggested a staining approach exerting 1.5 bar over a period of 30 h in our particular system. Other researchers aiming to use the plant injection method can use our model to estimate the pressure and time required in their own system. We identified root fragments in the lab using two methods (Fig. SM 8). First, we visually identified the color inside the roots under the bark. All roots for which color was not visible with the naked eye were introduced in a small plastic zip bag with water and kept at a constant temperature of 50 °C in a water bath. Bags were checked after five minutes, two hours, and one day, to see if any colorant dissolved in the water. When we detected a dye inside a root, visually or in dissolution in water, the root was linked to the plant stained with the same color in the plot.
Data processing
We calculated the Euclidian distance between every focal plant insertion and all soil sample centroids in the same plot, which allowed us to link data from the soil samples and plants. We sorted roots into diameter classes, differentiating fine (< 2 mm) and coarse roots (≥ 2 mm), and, within fine roots, diameters of: S (1.0 to 1.99 mm), XS (0.50 to 0.99 mm), and XXS (< 0.50 mm). For each soil sample, we calculated rooting volume (small mineral and organic particles and pores within) by subtracting the measured volume of gravel and stone from the total volume of the cylindrical sample, and root density as the root dry biomass (mg) per rooting volume (cm3) of the soil sample. Root densities (RD) could thus be calculated for the various diameter classes and for each individual plant that was dyed or genetically analyzed, or for all roots collectively.
Data analysis
We fitted generalized linear mixed (GLMM) models predicting plants’ individual root density in every point of three-dimensional soil; we called our models individual root density distribution models for fine-roots (fIRDD) and coarse-roots (cIRDD). We fitted these models only for rosemary plants, as this was the only species for which we identified roots using both methods. Only the rosemary individuals for which at least one root fragment was linked to a reference individual were considered, removing all plants for which no positive result was obtained (we removed seven out of fourteen individuals, clustering six out of eight genotypes in the microsatellite plot and none in the plant injection plots). Additionally, in the fine roots model we only accounted for the diameter class S, as we discarded all roots thinner than 1 mm using microsatellites.
We designed the fIRDD model to account for the mechanisms of the plant foraging behavior of the exploitative segregation theory (Cabal et al. 2020). Hence, we based this model on the euclidian distance from the plant stem to any point in soil (E), the non-self root density in that point of soil (NSR), the interaction beween both parameters, the depth in soil (D), the aboveground weight of the individual (AW), and the identification method used (m). The cIRDD model predicted the coarse root density based in the fine root density yielded by the fIRDD in the same sample (RDf) and the identification method used (m) (Table 1).
Notes
For a more detailed description and information of all our field methods, data processing, and statistical analyses, electronic supplementary materials are available:
-
(i)
Text and SM figures in “Detailed Materials and Methods” is an extended and detailed version of the Materials and Methods of this paper.
-
(ii)
The video “A Method for Identifying Shrub individual Root Density Distribution” describes the study site, and the method combining core extraction using a construction diamond drill and root staining by plant injection.
-
(iii)
The “Data and Code” folder includes all the collected data and several R scripts (R Core Team 2017) with all the code necessary to process the raw data, produce several graphical outputs, and perform statistical analyses.
The data collected as described in this paper covers much more ecological information (i.e., root density distribution in three-dimensional space, total root allocation, root ranges … of the three species studied). We will analyze and discuss the database produced from this work and fully available as a supplementary material in future publications focusing on plant ecology, foraging strategies, and inter-plant competition. In this paper, we focus exclusively on results that pertain to methodology, logistics, or that allow us to compare both identification methods tested.
Results
Hydraulic model results
The mean and range of values for water potential of each dye color measured are as shown in Fig. 1a. With these values, we plotted \(\overrightarrow{{v}_{10}(f)}\) for the different stem vessel diameters and dye water potentials (Fig. 1b).
Model results used to estimate pressure exertion time for plant injection at P = 1.5 bar and \({\Psi }_{w}^{soil}\) = -1 MPa. a- Means and range of values of the osmotic potential for the different color dyes injected to plants. b- Contour plot for \(\overrightarrow{{v}_{10}(f)}\), indicating the value for all the combinations of dye colors and focal species’ vessel diameters
Method comparison
We linked proportionally more root fragments to a plant using dye injection (~ 25%) than microsatellite analysis (~ 16%). However, the DNA and dying methods yielded consistent RD’s, because there was no significant effect of method on the estimated distributions for either fIRDD model (m, t = 0.770, p = 0.441) or cIRDD model (m, M t = -0.144, p = 0.889; PI t = -0.201, p = 0.845). This can be explained because, in the case of plant injection, most unidentified roots (75%) may correspond to individuals present in the plot and its surroundings that we did not stain. Contrastingly, in the case of the microsatellite plot we accounted for all the plants, hence in this case the 84% of unidentified or unspecific root samples must represent methodological errors. Given the total cost of each method applied, in our case the relative costs of linking roots was two orders of magnitude larger using microsatellite analysis (~ $400 per linked root) than dye injection (~ $4 per linked root) (Fig. 2, central).
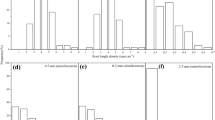
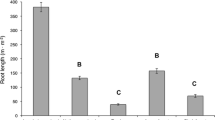
Summary of results from linking root fragments to plant individuals using two identification methods: Microsatellite (left) and plant injection with dyes (right). Central bar plots represent the proportion of analyzed roots successfully linked or not linked to plants. “Unspecific” results account for roots for which PCR/barcoding results were negative, non-specific, or roots that were associated with a different species (not rosemary). The left panel shows how different roots and plants analyzed with microsatellite analyses were linked to several genotypes (bottom), and how plants were spatially grouped by genotypes (top). The right panel shows the proportional performance of the visual and the dissolution identification methods per root diameter class, and the number of roots per color used successfully linked using the different identification techniques (visual and different dissolution times)
We assigned seven of the 14 plants we analyzed with the DNA method to two genotypes based in the analysis of leaves (Fig. 2, left panel). This illustrates that microsatellite markers cannot distinguish physically separate individuals of the same or similar genotype. These could be ramets, or alternatively separate individuals that the selected loci were not able to discern. Plant injection allows uncovering unambiguous connections between individuals that appear separate above ground but are connected belowground, because the dye is transported to all the living tissues of the physiological individual, including the leaves in the non-cut branches, where it can be seen with the naked eye.
Plant injection-based identification
The proportion of successful identifications of root fragments using dye injection varies between visual inspection and dissolution in root diameters and dye colors. Visual inspection was conducted first, and proved especially successful for thicker roots and for roots stained in cool colors (blue, green), which are difficult to confuse with natural wood colors. Of the remaining roots, we linked roots that were thinner and more often stained in warm colors (red, purple, yellow) using the water bath technique. Our results also show that the optimal time for dye dissolution is two hours, at which time we observe a peak in the fraction detected. Five minutes is not enough time for the dye to dissolve in the warm water, while 24 h proved to be too long (Fig. 2, right panel). After 24 h, most new identifications appeared to be yellow, and we suspect that roots may have stained the water in yellowish-brownish colors from the infusion of the natural plant materials after so much time soaking. This yellow color could have been misidentified as yellow or warm dyes, resulting in a potential source of error in positive results after 24 h of dissolution.
Discussion
Diverse coring techniques based on various mechanical systems already exist, but most ecological root studies still rely on hand-operated corers. Rotary drilling is a very effective coring technique, yet, like most mechanical coring systems, it involves equipment that is usually expensive, large and heavy (i.e. truck-mounted systems) (Abzalov 2016), which limits its use in many wild locations. However, more affordable and transportable diamond core drilling machines are widely available because of their use in construction industry to drill concrete. These machines can fit in most standard-sized cars, and researchers can transport them locally by hand. By adapting the use of a drilling machine of this type, we were able to extract large soil cores with woody roots from one of the hardest soils, a granitic leptosol. When encountering a rock, the machine was able to cut through it and, after extracting a granite column, we were able to continue sampling the soil underneath the rock. Generally, we extracted the cores over a period ranging from five minutes when we encountered no rocks, to around 30 min for the cores containing large rocks.
The comparison between the two identification methods demonstrates that the use of plant injection with dyes is less restrictive, more cost-effective, and had a lower degree of uncertainty than microsatellite markers for estimating IRDD. The GLMM results show that both methods’ results supported one another: there was no significant effect of the method on estimated RDs in the models. Even so, this result does not deny that microsatellite was overall less effective, as only the plants with at least one root identified were included in the model. This result indicates that microsatellite analysis yielded results similar to plant injection for the plant individuals for which it worked well, but it failed totally for many other individuals. Additionally, we found the microsatellite analysis restricted because of the limited availability of species-specific microsatellite markers (in our case, we could only use it for rosemary plants unless developing new markers) and its high financial costs. While both methods are subject to false negative errors (some of the roots that we could not link to any plant might actually belong to one of the focal plants), errors in positive results are more likely to occur using the microsatellite method because a root containing dye must belong to the corresponding injected plant. In addition, plant injection has other strengths such as allowing the measurement of sapwood area (Fig. SM 9), and the capacity to uncover unambiguous connections between individuals that appear separate aboveground. The dye diffuses to all the living tissues of the physiological individual including the all the standing branches and leaves, where it can be seen with the naked eye.
The different methods used to identify roots based in plant injection showed a different performance, but complemented each other well. While we could identify most of the linked roots visually, the dissolution in water baths allowed us to link additional roots to individuals that could not be linked visually, especially usefull for roots with thin diameters or stained in warm colors.
Conclusions
In this paper, we present and test an efficient and affordable protocol to map the IRDD of woody plants in the field. Tractable construction drills can be adapted to extract large soil samples of referenced spatial position relative to nearby plants and known volume in dry, stony soils. We must then link the root fragments from each core to the plants. We can do this successfully and at a reasonable cost by injecting dyes of different colors to the plant individuals before coring the soil, and identifying the colors inside the extracted root fragments. One important avenue for future research would be to experiment with different kinds of dyes and other chemical tags. For example, a dye that fluoresces at a specific wavelength could allow researchers to detect the presence of dye by an electronic sensor at concentrations that would have been invisible to the naked eye.
References
Abzalov M (2016) Drilling techniques and drill holes logging. In Applied mining geology. Modern approaches in solid earth sciences (12th ed.). Springer, Cham, pp. 39–77. https://doi.org/10.1007/978-3-319-39264-6_4
Addo-Danso SD, Prescott CE, Smith AR (2016) Methods for estimating root biomass and production in forest and woodland ecosystem carbon studies: A review. For Ecol Manage 359:332–351. https://doi.org/10.1016/j.foreco.2015.08.015
Arnaud M, Baird AJ, Morris PJ, Harris A, Huck JJ (2019) EnRoot: A narrow-diameter, inexpensive and partially 3D-printable minirhizotron for imaging fine root production. Plant Methods 15(1):1–9. https://doi.org/10.1186/s13007-019-0489-6
Bardgett RD, Mommer L, De Vries FT (2014) Going underground: Root traits as drivers of ecosystem processes. Trends Ecol Evol 29(12):692–699. https://doi.org/10.1016/j.tree.2014.10.006
Brunner I, Brodbeck S, Büchler U, Sperisen C (2001) Molecular identification of fine roots of trees from the Alps: Reliable and fast DNA extraction and PCR-RFLP analyses of plastid DNA. Mol Ecol 10(8):2079–2087. https://doi.org/10.1046/j.1365-294X.2001.01325.x
Cabal C, Deurwaerder HD, Matesanz S (2021) Field methods to study the spatial root density distribution of individual plants. Plant Soil 462:25–46. https://doi.org/10.1007/s11104-021-04841-z
Cabal C, Martínez-García R, De Castro Aguilar A, Valladares F, Pacala SW (2020) The Exploitative Segregation of Plant Roots. Science 370(6521):1197–1199. https://doi.org/10.1126/science.aba9877
Cahill JF, McNickle GG, Haag JJ, Lamb EG, Nyanumba SM, Clair CCS (2010) Plants integrate information about nutrients and neighbors. Science 328(5986):1657. https://doi.org/10.1126/science.1189736
Eldridge DJ, Bowker MA, Maestre FT, Roger E, Reynolds JF, Whitford WG (2011) Impacts of shrub encroachment on ecosystem structure and functioning: Towards a global synthesis. Ecol Lett 14(7):709–722. https://doi.org/10.1111/j.1461-0248.2011.01630.x
Freschet GT, Pagès L, Iversen CM, Comas LH, Rewald B, Roumet C, … McCormack ML (2021a) A starting guide to root ecology: strengthening ecological concepts and standardising root classification, sampling, processing and trait measurements. New Phytol 232(3):973–1122. https://doi.org/10.1111/nph.17572
Freschet GT, Roumet C, Comas LH, Weemstra M, Bengough AG, Rewald B, … Stokes A (2021b) Root traits as drivers of plant and ecosystem functioning: current understanding, pitfalls and future research needs. New Phytol 232(3):1123–1158. https://doi.org/10.1111/nph.17072
Fusco EJ, Rau BM, Falkowski M, Filippelli S, Bradley BA (2019) Accounting for aboveground carbon storage in shrubland and woodland ecosystems in the Great Basin. Ecosphere 10(8). https://doi.org/10.1002/ecs2.2821
Huang J, Zhang G, Zhang Y, Guan X, Wei Y, Guo R (2020) Global desertification vulnerability to climate change and human activities. Land Degrad Dev 31(11):1380–1391. https://doi.org/10.1002/ldr.3556
Jones FA, Erickson DL, Bernal MA, Bermingham E, Kress WJ, Herre EA, … Turner BL (2011) The roots of diversity: Below ground species richness and rooting distributions in a tropical forest revealed by DNA barcodes and inverse modeling. PLoS One 6(9):1-10 e24506. https://doi.org/10.1371/journal.pone.0024506
Kemper J, Cowling RM, Richardson DM (1999) Fragmentation of South African renosterveld shrublands: Effects on plant community structure and conservation implications. Biol Cons 90(2):103–111. https://doi.org/10.1016/S0006-3207(99)00021-X
Kirkham MB (2014) Principles of soil and plant water relations (2nd ed.). Elsevier, Oxford, UK
Lal R (2004) Carbon sequestration in dryland ecosystems. Environ Manage 33(4):528–544. https://doi.org/10.1007/s00267-003-9110-9
Lambers HFS, Chapin FS III, Pons TL (2008) Plant physiological ecology (Second). Springer, New York, US
Lang C, Dolynska A, Finkeldey R, Polle A (2010) Are beech (Fagus sylvatica) roots territorial? For Ecol Manage 260(7):1212–1217. https://doi.org/10.1016/j.foreco.2010.07.014
Lei P, Bauhus J (2010) Use of near-infrared reflectance spectroscopy to predict species composition in tree fine-root mixtures. Plant Soil 333(1):93–103. https://doi.org/10.1007/s11104-010-0325-2
Lux A, Rost TL (2012) Plant root research: the past, the present and the future. Ann Bot 110(2):201–204. https://doi.org/10.1093/aob/mcs156
Maestre FT, Benito BM, Berdugo M, Concostrina-Zubiri L, Delgado-Baquerizo M, Eldridge DJ, … Soliveres S (2021) Biogeography of global drylands. New Phytol. https://doi.org/10.1111/nph.17395
Murakami T, Shimano S, Kaneda S, Nakajima M, Urashima Y, Miyoshi N (2006) Multicolor staining of root systems in pot culture. Soil Sci Plant Nutr 52(5):618–622. https://doi.org/10.1111/j.1747-0765.2006.00078.x
Murakami T, Shimano S, Kaneda S, Nakajima M, Urashima Y, Miyoshi N (2011) Improvement of root staining method for field applications. Soil Sci Plant Nutr 57(4):541–548. https://doi.org/10.1080/00380768.2011.590945
Poorter H, Fiorani F, Pieruschka R, Wojciechowski T, van der Putten WH, Kleyer M, … Postma J (2016) Pampered inside, pestered outside? Differences and similarities between plants growing in controlled conditions and in the field. New Phytol 212(4):838–855. https://doi.org/10.1111/nph.14243
R Core Team (2017) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. Retrieved from https://www.r-project.org/. Accessed 03-06-2019
Rivas-Matínez S, Armaiz C (1984) Bioclimatologia Y Vegetacion en la Península Ibérica. Bull Soc Bot France Actualités Bot 5:110–120. https://doi.org/10.1080/01811789.1984.10826653
Roumet C, Picon-cochard C, Dawson LA, Joffre R, Mayes R, Blanchard A, Brewer MJ (2006) Quantifying species composition in root mixtures using two methods : near-infrared reflectance spectroscopy and plant wax markers. New Phytol 170:631–638. https://doi.org/10.1111/j.1469-8137.2006.01698.x
Saari SK, Campbell CD, Russell J, Alexander IJ, Anderson IC (2005) Pine microsatellite markers allow roots and ectomycorrhizas to be linked to individual trees. New Phytol 165(1):295–304. https://doi.org/10.1111/j.1469-8137.2004.01213.x
Schenk HJ, Jackson RB (2002) Rooting depths, lateral root spreads and below-ground/ above-ground allometries of plants in water-limited ecosystems. J Ecol 90:480–494. https://doi.org/10.1046/j.1365-2745.2002.00682.x
Schrader-Patton CC, Underwood EC (2021) New biomass estimates for chaparral-dominated southern California landscapes. Remote Sensing 13(8):1–24. https://doi.org/10.3390/rs13081581
Segarra-Moragues JG, Gleiser G (2009) Isolation and characterisation of di and tri nucleotide microsatellite loci in Rosmarinus officinalis (Lamiaceae), using enriched genomic libraries. Conserv Genet 10(3):571–575. https://doi.org/10.1007/s10592-008-9572-7
Stahl C, Herault B, Rossi V, Burban B, Brechet C, Bonal D (2013) Depth of soil water uptake by tropical rainforest trees during dry periods: does tree dimension matter? Oecologia 173(4):1191–1201. https://doi.org/10.1007/s00442-013-2724-6
Tomaselli R (1977) The Degradation of the Mediterranean Maquis. Ambio 6(6):356–362. Retrieved from http://www.jstor.org/stable/4312322. Accessed 07-02-2017
Weigand M, Kemna A (2017) Multi-frequency electrical impedance tomography as a non-invasive tool to characterize and monitor crop root systems. Biogeosciences 14(4). https://doi.org/10.5194/bg-14-921-2017
Wullschleger SD, Epstein HE, Box EO, Euskirchen ES, Goswami S, Iversen CM, … Xu X (2014) Plant functional types in Earth system models: Past experiences and future directions for application of dynamic vegetation models in high-latitude ecosystems. Ann Bot 114(1):1–16. https://doi.org/10.1093/aob/mcu077
Zenone T, Morelli G, Teobaldelli M, Fischanger F, Matteucci M, Sordini M, … Seufert G (2008) Preliminary use of ground-penetrating radar and electrical resistivity tomography to study tree roots in pine forests and poplar plantations. Funct Plant Biol 35(10):1047–1058. https://doi.org/10.1071/FP08062
Acknowledgements
We are very grateful to the Ayuntamiento de Candeleda for allowing us to work in their municipal properties in Las Tejoneras. Many thanks to Hannes De Deurwaeder for critical reading of the manuscript. Thanks to Ismael Aranda for sharing with us research materials and equipment. Thanks to Kristina K. Corvin for reviewing the English.
Funding
This work was funded by Princeton University’s High Meadows Environmental Institute-Carbon Mitigation Initiative (HMEI-CMI). CC acknowledges funding from The May Fellowship in the Department of Ecology and Evolutionary Biology, Princeton University. The authors have no relevant financial or non-financial interests to disclose. CC and SWP conceived the ideas; CC, JV, FV, and SWP designed methodology; CC, LRT, NMM, AV and AMB collected the data; CC analyzed the data; CC and NMM led the writing of the manuscript; FV and SWP funded this study; all authors contributed critically to the drafts and gave final approval for publication. The datasets generated during and analyzed during the current study are available as an electronic supplementary material.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Peter J. Gregory.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (MP4 1774593 kb)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cabal, C., Rodríguez-Torres, L., Marí-Mena, N. et al. Comparing two field protocols to measure individual shrubs’ root density distribution. Plant Soil 481, 691–699 (2022). https://doi.org/10.1007/s11104-022-05657-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05657-1