Abstract
Purpose
Soil acidification influences competitive N uptake between plants and microorganisms. The mechanisms by which soil acidification affects competition between maize and microorganisms for organic N must be determined to understand N cycling and adjust the forms and levels of N fertilisation.
Methods
The uptake of glycine, mineral N after glycine decomposition, and NH4+ by maize and microorganisms was investigated using 13C and 15N labelling. Microbial community composition biomarkers were analysed using phospholipid fatty acid (PLFA) analysis. Mineralisation of organic N was monitored via CO2 production, and gross NH4+/NO3− production and consumption was assessed using 15N pool dilution.
Results
Soil acidification (pH from 7.6 to 5.1) increased the intact glycine uptake by maize roots (from 0.7 to 2.4% of added 15N) but decreased its uptake by microorganisms (from 32 to 2.4% of added 15N). Soil acidification altered the microbial community composition: the PLFA of arbuscular mycorrhizal fungi and anaerobes decreased by 6- and 1.5-fold, respectively. Soil acidification reduced the decomposition rates of proteins, peptides, and amino acids as indicated by the CO2 release. This corresponded to a gross NH4+ production increase by 1.3-fold and a gross NO3− production decrease by 97%, compared with soil at pH 7.1.
Conclusions
Acidification led to decreased microbial biomass, shift in the microbial community, and the strong decrease (10–15-fold) in amino acid uptake by microorganisms, and was beneficial to maize plants, which assimilated 2.4% of the N added as glycine. However, these quantities of N are insufficient for a substantial increase in the N nutrition of the plants.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
In soil, organic N accounts for 70–95% of the total N (Hill and Jones 2019). It comprises mainly amino acids, which are the most abundant source of bioavailable N and contain organic monomers derived from protein and peptide hydrolysis (Farrell et al. 2011a, b). NH4+ and NO3− are the main N forms taken up by plants, uptake of organic N has been largely ignored (Fokin et al. 1993). Organic N is also an important N source for plants in cold ecosystems, such as the polar, northern tundra, and northern boreal forest regions (Schmidt et al. 2008; Schmidt et al. 2016). Some agricultural plants, such as maize (Xu et al. 2008), rye (Jämtgård et al. 2008), wheat (Ma et al. 2018), and soybeans (Ma et al. 2021a) can take up amino acid-N directly, but this uptake is marginal compared to their mineral N uptake (Biernath et al. 2008; Rasmussen et al. 2010; Moran-Zuloaga et al. 2015). Despite the minor role of amino acid uptake in plant nutrition, some amino acids may act as signalling molecules for sensing plant N status and are therefore crucial to N metabolism (Guo et al. 2021). Amino acids also regulate plant growth by changing the shapes of roots and shoots, and regulate defence responses against biotic and abiotic stressors (Zeier 2013; Guo et al. 2021). Many factors such as pH, microbial community, and the content and composition of soil organic N affect amino acid uptake by plants (Ma et al. 2016). However, the mechanisms by which these factors affect soil amino acid bioavailability remain uncertain.
Typically, proteins and peptides are hydrolysed by exoenzyme proteases to form amino acids (Greenfield et al. 2020), and their decomposition rate depends on protein characteristics and soil properties (Nannipieri and Eldor 2009; Greenfield et al. 2020). Peptides and amino acids can be rapidly decomposed within minutes to hours (Jones et al. 2005a, b; Apostel et al. 2013), while protein decomposition is slower and may take days (Jan et al. 2009). Organic N decomposition is regulated by various environmental factors including soil moisture, temperature, and pH (Zhang et al. 2019). Changes in the soil pH alter microbial activity and the adsorption of proteins and peptides in the soil (Zhang et al. 2019; Greenfield et al. 2020). This may further affect amino acid production, and ultimately, the competition between plants and microorganisms for amino acids.
Nitrogen availability is one of the key factors limiting plant growth and productivity in natural ecosystems (Mckane et al. 2002; Jiang et al. 2018). Plant amino acid uptake is an energy-driven process, wherein outward-facing membrane H+ ATPases generate a dynamic proton gradient to drive inward-facing amino acid–H+ cotransport (Owen and Jones 2001; Nsholm et al. 2009). Plants can take up 0.2–21% of intact amino acids within hours (Ma et al. 2018; Hill and Jones 2019), but microbial uptake of amino acids is considerably faster, ranging from minutes to hours (Kuzyakov and Xu 2013); hence, microorganisms are the main competitors for amino acid uptake (Hill and Jones 2019; Moran-Zuloaga et al. 2015). Understanding the mechanisms underlying microbial amino acid uptake and determining the microbial communities involved in these processes can aid in elucidating the constraints faced by plants in amino acids uptake.
Soil pH is a primary factor that affects nutrient availability, microbial community composition, and plant growth. Excess NH4+-based fertilisation induces severe soil acidification (Guo et al. 2010; Zamanian et al. 2018). Soil acidification affects root cell plasma membranes, root membrane transporter activity, and the Km associated with the transport system, thereby decreasing the amino acid uptake of plants (Reid and Hayes 2003). Soil pH also affects microbial N use by altering the soil microbial biomass (Pietri and Brookes 2009) or community composition (Pietri and Brookes 2009; Rousk et al. 2010). The dynamic equilibrium of the amino acid pool is regulated by amino acid production and mineralisation (Jones and Kielland 2012), which are both influenced by several factors including microbial activity and soil pH. Soil pH modifies 1) microbial biomass and community structure (Zhang et al. 2019); 2) mineralisation of amino acids (Rousk et al. 2010; Malik et al. 2018); 3) activities of proteases, peptidases (Jérémy et al. 2019), and other enzymes; and 4) binding of proteins and peptides by soil particles (Greenfield et al. 2020), which affects their mineralisation and ultimately amino acid production. Thus, determining N cycling processes is key to explain how environmental changes affect amino acid utilisation by microbes, and how roots compete with microbes for amino acids under soil acidification.
This study investigated plant–microorganism competition for N depending on soil pH, using a staple crop, maize, cultivated with a mixture of 13C,15N dual-labelled glycine and 15NH4+. We tested the following hypotheses: 1) Soil acidification decreases glycine uptake but increases NH4+ uptake by both maize and microorganisms; 2) soil acidification decreases the microbial biomass and alters the microbial community, causing Gram-negative bacteria (G−) numbers to increase and those of arbuscular mycorrhizal fungi to decrease; 3) soil acidification reduces the decomposition rates of proteins, peptides, and amino acids, thereby decreasing the gross mineralisation rate and limiting soil N cycling.
Materials and methods
Soil
The soil samples were collected from Jinhua City, Zhejiang Province, China (29°19′09″ N, 119°43′43″ E, 72.8 m a.s.l.) and classified as Haplic Acrisol soil (Food and Agriculture Organisation; FAO). The initial physico-chemical characteristics of soil are listed in Table S1. Soil pH was measured using 1:2.5 (w/v) soil:distilled water extracts. Soil organic C was determined using K2Cr2O7 redox titration (Yeomans and Bremner 1988). Total N content was determined following semi-automatic Kjeldahl digestion with 5 mL concentrated H2SO4 (Bao 2000). Alkali-hydrolysed N content was determined using the alkaline diffusion method (Mulvaney and Khan 2001). Soil available P was detected in ammonium fluoride hydrochloride (pH < 7) and sodium bicarbonate (pH > 7) extracts using the molybdenum blue method (Hylander et al. 1996). The soil available K (K+), Mg (Mg2+), and Ca (Ca2+) contents were measured in 1:10 (w/v) soil:CH3COONH4 (ammonium acetate) extracts by performing inductively coupled plasma atomic emission spectroscopy (ICP-AES) (Thermo Fisher Scientific, Waltham, MA, USA). Soil available Al III (Al3+) was detected in 1:10 (v/v) soil:1 M KCl extracts using graphite furnace atomic absorption spectrometry (GFAAS) (AA 800; PerkinElmer, Waltham, MA, USA). To test the contents of NH4+ and NO3−, 5 g of soil was first extracted with 25 mL of 1 M KCl and shaken at 220 rpm for 1 h. Then, the contents of NH4+ and NO3− in the soil extracts were colorimetrically detected using the salicylic acid and vanadate methods, respectively, with a PowerWave HT microplate spectrophotometer (BioTek Instruments Inc., Winooski, VT, USA) (Mariano et al. 2016).
Soil pH adjustment
Soil samples with four different pH values were produced using the electrokinetic method (Xiao et al. 2013). The electrokinetic device consisted of a direct current (DC) power supply, a plastic basin (450 × 250 × 100 mm soil reaction chamber), and anode and cathode chambers (Fig. S1) (Xiao et al. 2013). The DC power supply had an adjustable voltage (0–220 V) and each electrode chamber contained two graphite electrodes (100 × 120 × 5 mm). Two electrodes were inserted into the soil on either side of the reaction chamber to induce a DC electrical field and allow the electrode surfaces to initiate water electrolysis. The anode was placed into the leftmost soil sample in the reaction chamber, and the cathode was placed into the rightmost soil sample (Fig. S1). Oxidation occurred at the anode and reduction occurred at the cathode (Saichek and Reddy 2003). The OH− and H+ ions moved to the cathode and anode, respectively, via electromigration, electroosmosis, and diffusion. This caused the pH of the soil near the cathode and anode to rise and fall, respectively (Saichek and Reddy 2003). This technique resembles natural acidification because it alters the contents of soil Ca2+, Mg2+, Al3+, and other cations (Saichek and Reddy 2003; Xiao et al. 2013). The soil pH varied depending on the distance from the anode. The soil closer to the anode contained more H+ ions, indicating a lower pH, while the soil closer to the cathode harboured more OH− ions, indicating a higher pH.
The air-dried soil sample was passed through a 5-mm sieve before being placed in the reaction chamber at a depth of 8 cm. Subsequently, purified water was added until the soil was saturated, and the two electrodes were inserted on each side of the reaction chamber. The DC power supply was set to 220 V and run for 15 h. Purified water was added periodically to maintain soil saturation. After electrolysis, the soil was removed from the reaction chamber and divided into four parts according to its distance from the anode (0–10, 10–20, 20–30, and 30–40 cm). For each electrolysed soil sample, four replicates were prepared. The soil portions were air-dried and passed through a 2-mm sieve before subsequent incubation experiments (Xiao et al. 2013). To restore the microbial diversity and abundance, 100 g of the original un-electrolysed soil (10%) was added to 1000 g of electrolysed soil and the mixture was incubated at 25 °C with a 60% water content for 10 d. The soil pH in the four groups was successfully adjusted to 5.1, 6.0, 6.5, and 7.6.
Plant cultivation
Maize seeds were germinated in Petri dishes in the dark for 2 d until the roots grew to 1–2 cm in length. Electrolysed soil (50 g) was placed in 50 mL centrifuge tubes; a single germinated maize seed was placed in each tube and covered with 2 g soil. Then, 2 mL pure water was added, and the tube was placed in an artificial climate chamber. The day/night temperatures and relative humidity were 25/20 °C and 60/40%, respectively. Water was added 2–3 times daily. After 20 d of cultivation, the tubes with maize seedlings were divided into two groups. Group A was used in the subsequent experiments on plant/microbial competition for amino acids and the 13C-PLFA analysis (Sections 2.4 and 2.5). Group B was used in the subsequent experiments on organic N mineralisation and the gross NH4+/NO3− production/consumption analysis (Sections 2.6 and 2.7).
Amino acid uptake by maize roots and soil microorganisms
A 5 mL tracer solution was added to the samples from Group A. The soil water content was maintained at 30% to ensure that the tracers rapidly diffused throughout the culture tube. The following solution combinations were used in various treatments: 100 μM 99.0% 13C,15N-glycine +100 μM NH4+; 100 μM 99% 15NH4+ + 100 μM glycine; and 100 μM NH4+ + 100 μM glycine (unlabelled control). The glycine concentration used here reflects the typical free amino acid content associated with root and microbial cell lysis in agricultural soils (Jones et al. 2005a, b).
After 4 h of tracer uptake, the maize seedlings were removed from the tubes. The roots and soil were separated by gentle shaking. The soil samples were subjected to microbial amino acid uptake and PLFA analyses. The roots were washed thrice with 50 mM CaCl2 and purified water to remove any residual 13C/15N from the root surfaces. The seedlings were oven-dried at 90 °C and pulverised in a ball mill (Shanghai Jingxin Co. Ltd., Shanghai China). Two maize seedlings were combined in each replicate, with four replicates per treatment. The 13C/15N content in the maize seedlings was measured using element analyser-stable isotope mass spectrometry (EA-IRMS) (IsoPrime100; Isoprime Ltd., Cheadle Hulme, UK).
To detect the 13C and 15N in the microbial biomass, 20 g soil sample from Group A (4 h after tracer addition) was fumigated with 3 mL alcohol-free chloroform for 24 h. The chloroform was then removed using a vacuum desiccator, evacuating the vessel thrice (20 min each time). The soil was extracted by shaking in 0.01 M KCl at 180 rpm for 30 min. Another 20 g of soil from Group A remained unfumigated and was directly extracted under similar conditions. There were four replicates; the extracted samples were oven-dried at 45 °C and placed in tin capsules. The 13C/15N abundance in the soil microorganisms was detected by EA-IRMS.
13C-PLFA analysis
Freeze-dried soil sample from Group A (1.5 g) that had received 13C,15N-glycine was extracted twice with 7.6 mL chloroform/methanol/citrate (1:2:0.8 (v/v/v), 0.15 M, pH 4.0) single-phase buffer mixture. The neutral lipids and glycolipids were removed using a silica column, while the phospholipids were retained. Thereafter, the collected phospholipids were dried with N2, and 100 μL methyl nonadecanoate fatty acid (19:00) was added as an internal standard (Bobbie and White 1980). After phospholipid methylation, the PLFA methyl ester was separated with 60 μL hexane and identified using gas chromatography (GC 7890A; Agilent Technologies, Santa Clara, CA, USA) and a MIDI Sherlock Microbial Identification System 6.2B (MIDI; Newark, DE, USA). 13C content in the PLFA compounds was detected using gas chromatography combustion isotope ratio mass spectrometry in a Trace GC Isolink II gas chromatograph; it was fitted with a combustion column attached via a GC Combustion III to a Delta V Advantage isotope ratio mass spectrometer (Thermo Fisher Scientific, Waltham, MA, USA) (Thornton et al. 2011). Cyclopropyl and monounsaturated fatty acids are biomarkers of G− while ante-iso- and iso-branched fatty acids are indicators of Gram-positive bacteria (G+). The 10Me branched PLFA is an indicator of actinobacteria (Ge et al. 2017). Saturated straight-chain fatty acids are considered nonspecific PLFAs (general FAMEs) present in various microorganisms. The 18:1ω9 is regarded as a G− indicator in agricultural soil (Åsa et al. 2011).
Protein, peptide, and amino acid decomposition
To estimate organic compound decomposition rates, a 5 g sample of soil sown with maize (Group B, without glycine or NH4+ addition) was placed in a 50 mL gas culture bottle, and 0.5 mL protein (albumin from bovine serum albumin), peptide (L-triglycine), or L-amino acid (glycine) was added to the soil at a concentration equivalent to 1 mM N, respectively. Each treatment had four replicates, and 0.5 mL purified water was used as the control. The bottles were incubated in the dark at 20 °C and CO2 was collected from the bottles after 0.5, 1, 3, 9, 24, and 48 h (amino acids and peptides) and after 1, 3, 9, 24, 96, and 192 h (proteins), which were then detected using gas chromatography (Varian, Palo Alto, CA, USA). The CO2 sampling time point series was longer for the proteins as they decompose more slowly than amino acids and peptides.
Gross NH4 + and NO3 − production and consumption
Using Group B samples, the gross NH4+ and NO3− production and consumption in soils after maize cultivation were detected following the 15N isotope pool dilution method with slight modifications (Wanek et al. 2010). In brief, the NH4+ and NO3− pools were labelled with enriched 15N tracers and the gross target pool influx and efflux rates were calculated according to the target pool size and the differences in the target pool 15N composition between two time points. Considering the stimulating effect of adding an 15N tracer on the immobilisation rate, inorganic 15N was added at <20% of the initial pool size. Soil (20 g) was combined with 2 mL of 0.1 mM 20% atm 15NH4+ or 15NO3−. After 15 min and 24 h, the soils were extracted using 80 mL of 0.5 M K2SO4 solution and centrifuged at 1000 rpm for 1 min. The soil NH4+ or NO3− content was measured by microdetection colorimetry (Mariano et al. 2016) and the 15N isotope ratio of NH4+ or NO3− was detected using a modified diffusion method (Corre et al. 2007). To induce the conversion of 15NH4+ to 15NH3, 0.2 g MgO was added to a 100 mL Duran bottle containing 60 mL 0.5 M K2SO4 extract. A 7 mm filter disc acidified with 2.5 M KHSO4 was suspended below the cap before sealing the vessel. The bottles were shaken carefully twice daily, and the filter discs were removed after 6 d. For 15NO3− detection, 0.2 g MgO was added to the extract, and the bottles were left open for 3 d to remove any residual 15NH4+ and 15NH3. Then, 0.2 g MgO and 0.4 g Devarda’s alloy were added to each bottle. The KHSO4-impregnated filter discs were set inside the bottles as stated above, and the vessels were shaken twice daily. The filter discs were removed after 6 d. All filter discs were freeze-dried and their 15N content was analysed using the EA-IRMS.
Data processing and statistical analysis
The amino acid, peptide, and protein decomposition rates were calculated based on the amounts of CO2 produced at different time points (Orchard and Cook 1983):
where CT is the amount of CO2 produced per gram soil, Ct is the measured amount of CO2, VH is the headspace volume, and Vt is the measuring volume.
The peptide and amino acid mineralisation was assessed using the two-process double-first-order kinetic decay model (Glanville et al. 2016). Relevant details are shown in the supplementary information.
The amount of 13C and 15N taken up by the maize from the labelled glycine was obtained by subtracting the amount of 13C or 15N detected in the control from the corresponding amount detected in the labelled sample. This calculation was first presented by Sauheitl et al. (2009):
where Cu is the uptake of 13C or 15N from the labelled glycine (μg), CT-c is the total C or N in the maize, and AS and AC are the 13C or 15N atom% in the test and control samples, respectively.
The complete glycine uptake of the maize was calculated from the percentage of 13C uptake (Ge et al. 2017). The post-mineralisation 15N absorption was calculated from the total 15N absorption minus the intact 15N absorption. The 13C and 15N uptake by the microbial biomass was calculated from the difference in 13C or 15N between fumigated and unfumigated soil extracts, divided by 0.45 (Ganeteg et al. 2016):
Where CM is the amount of 13C or 15N absorbed by the soil microorganisms, A is the 13C or 15N atom% in the soil extract, C is the total C or N content in the soil extract, and W is the soil sample weight. F, UF, L, and UL represent fumigated, unfumigated, labelled, and unlabelled samples, respectively.
The gross NH4+/NO3− production (GP) and consumption (GC) were calculated using the isotope dilution equation (Braun et al. 2018). The atom percent excess (A) in the equation is the 15N atom% of the sample minus the natural 15N abundance in the unlabelled control samples.
where Q1 and Q2 are the NH4+/NO3− concentrations and A1 and A2 are the 15N atom% in NH4+/NO3− at times t1 (3 h) and t2 (24 h), respectively, after the NH4+/NO3− pool was labelled.
Data were analysed using SAS v.8.2 (SAS Institute Inc., Cary, NC, USA). We performed analysis of variance (ANOVA) and a post hoc Tukey’s test (p < 0.05) to compare the differences between the treatments. The data are presented as the means ± standard error (SE). Principal component analysis (PCA) was applied to the data pertaining to the 13C ratios in the PLFA indicators at four different pH values to determine the effects of pH on soil microbial communities. A redundancy analysis (RDA) was conducted using CANOCO v.5.0 (Microcomputer Power, Ithaca, NY, USA) to identify the effect of environmental factors (pH, total N, organic matter, NH4+, NO3−, Mg2+, Ca2+, and Al3+) on N uptake by maize and microbes, including the 15N intact glycine, mineral 15N after glycine deamination, and 15NH4+ uptake. Figures were plotted in Origin v.8.1 (OriginLab, Northampton, MA, USA).
Results
Effects of soil acidity on maize uptake of 13C and 15N derived from glycine or 15N-ammonium
After 20 d cultivation, the maize biomass was approximately 30% larger in the 6.5 and 7.6 pH soil samples than in the soil samples with lower pH (Fig. S2). The amount of 13C reflects the proportion of glycine that bypassed microbial degradation and was directly taken up by the maize. Soil acidification increased the uptake of intact amino acid by maize (Fig. 1A). In contrast, 15N uptake by maize after glycine mineralisation was the highest at pH 6.5 (2.4% of 15N) (Fig. 1A).
Effects of soil pH on maize uptake of 13C,15N-labelled intact glycine, 15N after the mineralisation of glycine, and 15NH4+ (A), and microbial uptake of glycine and 15NH4+ (B). Whiskers represent the means ± SEM of four replicates. Lowercase letters indicate significant differences in glycine uptake among soil with different pH (p < 0.05). Soil pH did not significantly alter maize uptake of 15NH4+ and 15N after mineralisation. Soil pH did not significantly alter microbial 15NH4+ uptake
Effects of soil pH on microbial uptake of 13C and 15N derived from glycine
Soil acidification from pH 7.6 to 5.1 caused a 5- and 2.2-fold decrease in microbial biomass C and N, respectively (Table. S1). Microorganisms assimilated maximum glycine within 4 h at pH 6.5 (32% of the total) compared to other pH values (Fig. 1B). In contrast, microbial uptake of NH4+ was constant among pH values (Fig. 1B).
15N distribution in maize, microorganisms, and soil
After the addition of 13C,15N-glycine to the soil, the 15N uptake by the microbial community was 10-fold higher than that of the maize (including glycine and 15N after mineralisation) at pH 6.5 and 7.6. In contrast, the 15N-glycine uptake by the microbial community at pH 5.1 and 6.0 was similar to that of maize, both less than 5% (Fig. 2).
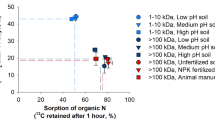
Intact glycine uptake by maize increased with increasing SOM and NH4+, whereas it decreased with increasing pH, total N, and NO3−, as well as microbial biomass. Glycine uptake by microorganisms (MB-15N) increased by 10–15 folds with soil pH (Fig. 3A) and increased with increasing microbial biomass (Fig. S3).
Redundancy analysis (RDA) of the effects of environmental factors on glycine uptake by maize and soil microorganisms (A), and principal component analysis (PCA) of 13C PLFA depending on soil pH (B). TN represents the total N; OM represents the organic matter; M-B represents the maize biomass; M-15N represents the percentage of 15N uptake by maize; M-15NH4+ represents the percentage of 15NH4+ uptake by maize; M-13C represents the percentage of 13C uptake by maize; MB-N represents the microbial biomass N; MB-C represents the microbial biomass C; MB-15N represents the percentage of 15N uptake by microorganisms; MB-15NH4+ represents the percentage of 15NH4+ uptake by microorganisms; and MB-13C represents the percentage of 13C uptake by microorganisms
The incorporation of 13C from glycine into PLFAs depends on soil acidity
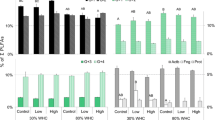
The 13C-PLFA was used to identify the microbial groups competing with maize roots for glycine. The total 13C-PLFA was highest at pH 5.1. G− bacteria predominated at four pH and absorb the highest amount of amino acids in the entire microbial community at pH 6.5 (Fig. 4). The 16:1ω5c ratio, representing arbuscular mycorrhizal (AM) fungi, increased 9-fold with increasing pH. Anaerobes decreased from 7.0% to 2.9% with decreasing pH, while actinomycetes increased from 8.7% to 17.9%. Neither G− nor G+ bacteria were affected by soil acidification (Fig. 4).
Ratios of 13C in PLFA based on soil pH (% of total 13C-PLFA). PLFAs are classified as G+ bacteria, G− bacteria, general FAME, Anaerobes, Actinomycetes, and AM fungi. Lowercase letters indicate significant differences among soils of different pH values (p < 0.05). The lines represent the means ± SEM of four replicates
The PCA of the 13C amount incorporated into PLFAs revealed differences in microbial community responses to acidification. The 13C-PLFA composition at pH 6.5 differed from that at pH 6.0 and 5.1, and the composition at pH 7.6 differed from that at pH 5.1 and 6.0 (Fig. 3B).
Effects of soil acidification on organic N decomposition
The decomposition of amino acids, peptides, and proteins over 48 h was the highest at pH 6.5. The peptide decomposition rates at pH 5.1 and 6.0 were 72% and 51% lower, respectively, than that at pH 6.5. The protein decomposition rates decreased with decreasing pH (Fig. 5).
The gross NH4+ production increased by 58% in soil at pH 7.6 compared to that at pH 5.1. In contrast, the gross NH4+ consumption remained unaffected by soil acidity (p > 0.05; one-way ANOVA). The gross NO3− production and consumption increased by 97% and 113%, respectively, when the pH increased from 5.1 to 7.6, with the lowest production and consumption rates observed at pH 5.1. The production and consumption rates of NO3− were higher than those of NH4+ (Fig. 6).
Discussion
Competition for glycine between maize and soil microorganisms
Owing to the high mineral N fertiliser input in agricultural soils and intense competition from microorganisms, organic N has a limited role in plant nutrition (Kuzyakov and Xu 2013; Ganeteg et al. 2016; Moran-Zuloaga et al. 2015). Ma et al. (2018) investigated in situ uptake by adding 100 μM 13C,15N-labelled glycine to the wheat rhizosphere for 4 h. They only recovered 6–21% of it from the roots, suggesting that most of the glycine was incorporated by microorganisms. Our short-term (4 h) uptake test showed that 0.7–2.4% of the intact glycine and 0.8–2.2% of the inorganic N after glycine mineralisation were taken up by maize. However, some of the glycine added to the soil might be rapidly converted to other organic substances, and then incorporated by the plants (Vranova et al. 2012; Moran-Zuloaga et al. 2015). Moreover, the ability of maize roots to absorb and utilise organic N may have been underestimated, as maize respiration releases 13CO2 (Jones et al. 2005a, b; Rasmussen et al. 2010). Concurrently, part of the mineralised 13C remained as H13CO3− that could be taken up by maize under a high pH (Wu and Xing 2012). Thus, in the future, H13CO3− uptake concentration needs to be measured to better understand the uptake of intact amino acids by maize.
The uptake of organic N by maize was primarily restricted by competition with microorganisms. Soil acidification decreases plant N uptake and alters the microbial community structure (Reid and Hayes 2003; Malik et al. 2018). However, maize can take up more amino acids with soil acidification, refuting our first hypothesis. Soil acidification increased the intact glycine uptake of maize from 1.2 to 3.8% of the total added amount (Fig. 1A). Maize incorporates more amino acids at low pH, which may depend on H+ cotransporters (Hawkins and Robbins 2010; Ma et al. 2016). The pH affects the H+ concentration in the plasma membrane of root surface cells, which in turn affects the activity and efficiency of amino acid transporters, thereby modifying intact glycine uptake (Guo et al. 2021). Consequently, soil acidification may increase the activity of H+ cotransporters on the surface of plant root cells, facilitating the uptake of more intact glycine by the plants (Fig. 7).
Mechanisms underlying the effects of soil pH on the competition between maize and soil microorganisms for amino acid uptake, including the abilities of maize and microorganisms to take up amino acids, and soil N cycling rates. The black lines represent soil N cycling processes. The red line indicates the processes affected by soil pH. The black dashed line represents peptides that may be utilised by microorganisms directly. The red lines indicate that the microbial biomass, microbial community, plant uptake capacity, and gross NH4+/NO3− production and consumption are affected by soil acidification. The blue numbers represent the percentage of added 15N recovered from maize plants or microorganisms. The green numbers refer to gross NH4+/NO3− production or consumption rates. The purple numbers indicate organic N mineralisation rates
The intact glycine uptake by plants faces fierce competition from soil microorganisms (Hill et al. 2012; Moran-Zuloaga et al. 2015). The incorporation of glycine into microorganisms increased from 2.4 to 32% with increasing pH, i.e., the glycine uptake by microorganisms was proportional to the pH (Fig. 3A). A lower pH decreased microbial biomass and respiration, which reduced microbial glycine uptake and mineralisation, and decreased microbial competition, allowing the maize to take up more glycine. Consequently, soil acidification increased amino acid uptake by maize but decreased that by microorganisms (Fig. 7).
Effects of pH on microbial glycine uptake
The N uptake of glycine by microorganisms increased 10-fold as the pH increased from 5.1 to 7.6, while the microbial biomass N increased 3-fold (Fig. S3). High pH condition may be a suitable growth environment for microbes because their activity and enzyme secretions are maintained. Consequently, a higher pH would facilitate the decomposition of proteins, peptides, and amino acids. In our study, higher soil pH decreased the concentration of Al3+ (Table S1), thereby reducing its toxic effects on microorganisms and ultimately increasing microbial biomass and amino acid absorption (Pietri and Brookes 2008). The plant biomass also increased at higher pH (Fig. S2), indicating that the roots could release more exudates (Zhao et al. 2021). As a result, more C was taken up by microorganisms, which may have increased their biomass and activity. Thus, the competition for glycine between plants and microbes may have intensified at higher pH.
By determining how acidity affects soil microbial communities, it is possible to identify the microorganisms that compete with plants for organic N (Singh et al. 2009). G− bacteria constituted approximately 40% of the amino acid-absorbing microbial community and remained stable regardless of the pH (Fig. 4). Thus, G− bacteria dominated the microbial community and were presumed to be the main microbes competing with maize for organic N. The PCA showed significant differences between the microbial communities identified at pH 6.5 and 7.6 and those at pH 5.1 and 6.0 (Fig. 3B). Certain PLFAs might be directly associated with pH changes. Soil pH strongly affected the PLFA components, especially for actinomycetes and anaerobes (Fig. 4). The 16:1ω5 refers to AM fungi and decreased from 1.3% to 0.23% with soil acidification (Hgberg et al. 2007; Rousk et al. 2010). Owing to the decrease in microbial biomass and change of community composition with soil acidification, organic N (proteins, peptides, and amino acids) mineralisation and the ability of microorganisms to take up glycine decreased, allowing its uptake by maize. Consequently, the changes in soil microbial biomass and community composition caused by changes in the pH directly influence plant–microbial competition for the amino acids (Fig. 7).
Effects of soil acidification on organic N mineralisation
Most organic N must be cleaved prior to being incorporated by plants (Greenfield et al. 2020; Ma et al. 2020a; Moran-Zuloaga et al. 2015). We found that the decomposition of amino acids, peptides, and proteins was affected by soil pH (Fig. 5). Microbial biomass C and microbial biomass N increased with increasing pH (Table S1). The rapid mineralisation of amino acid (a1) increased 1.2-fold from pH 5.1 to 7.6 (Table S2). Increasing pH shifted the bacterial community towards more G− bacteria (Fig. 4), which may have accelerated microbial growth and turnover (Pietri and Brookes 2009). Thus, more C and N uptake were needed to support microbial growth at high pH, and amino acids, as a low-molecular form of organic N, were rapidly utilised (Kuzyakov 2010; Hagerty et al. 2014).
Peptides have more complex structures than amino acids (Farrell et al. 2011a, b; Hill et al. 2011). Warren (2021) found that large amounts of small peptides and amino acids are produced when proteins depolymerise, and the production rate of peptides in soils is 11–68 times faster than that of amino acids, indicating that small peptides can constitute an important source of organic N for microorganisms (Farrell et al. 2011a, b). However, the rate at which peptides are depolymerised and utilised by microorganisms may be regulated by pH, which may indirectly affect the ability of microorganisms to utilise amino acids. Consistent with previous findings (Ma et al. 2020b), peptide and amino acid mineralisation (t1/2) was similar at pH 6.5. At pH 5.1, however, the peptide decomposition rate reduced 2-fold compared with that at pH 6.5 (Fig. 5; Table S2). We suggest that peptides may be directly utilised by microorganisms without first being decomposed by peptidases into amino acids (Hill et al. 2012; Hill and Jones 2019). This may, in turn, influence the competition for amino acids.
Protein hydrolysis is the rate-limiting step in amino acid production (Jan et al. 2009; Yuntao et al. 2018), and protein decomposition rates rapidly decreased with decreasing pH (Fig. 5). A decrease in the protein decomposition rate may reduce amino acid production. Soil acidity is a major factor controlling microbial community composition, metabolism, and protease activity (Jones et al. 2019). We considered that increases in the protein mineralisation rate primarily depend on three mechanisms: (i) increased protection offered by soil particles, which involves the relationship between the soil pH and the protein isoelectric point, to make the proteins inaccessible to exoenzymes secreted by microorganisms (Ma et al. 2016; Greenfield et al. 2020); (ii) increased ex vivo modification of original compounds, as some proteins are more difficult for exoenzymes to decompose after modification (Liang et al. 2017); and (iii) decreased microbial biomass and changes in microbial community composition owing to acidification (Farrell et al. 2011a, b; Giagnoni et al. 2011). The protein mineralisation rate was considerably slower than those of amino acids and peptides; the t1/2 of proteins was less than 7 h at pH 6.5, while those of amino acids and peptides were less than 5.8 and 4.5 h, respectively (Table S2). This suggests that protein decomposition occurred rapidly at high pH; hence, we predicted that the degradation of soluble low-molecular-weight proteins by microorganisms would be much faster than that of insoluble high-molecular-weight proteins. In addition, the priming effect of pH on organic N mineralisation was not considered.
The rate of gross NH4+ production increased 1.5-fold when pH decreased from 6.5 to 5.1 (Fig. 6), refuting our third hypothesis. Generally, the amount of organic N utilised by microorganisms depends primarily on their growth and energy metabolism (Geisseler et al. 2010; Zhang et al. 2019; Ma et al. 2021b). If organic N decomposition decreases, then gross NH4+ production should also decrease. Soil surface charge may be one of the factors affecting gross NH4+ production. At a soil pH of 5.1, H+ ions directly participate in NO3− reduction and affect the adsorption of NO3− on reactive sites (soil corrosion products) (Huang and Zhang 2004). This may increase the gross NH4+ production. Furthermore, the increase in the gross NO3− production with increasing pH may be explained by pH-dependent alterations in the activity of acid-sensitive nitrifying bacteria (Cheng et al. 2013; Tang et al. 2021).
The decomposition rates of the proteins, peptides, and amino acids decreased with soil acidification, while the gross NH4+ content increased. We hypothesised that the electrostatic adsorption of differently charged proteins, peptides, and amino acids on the soil stationary phase was affected by the solute charge induced by soil acidification. Microorganisms can utilise only specific organic substrates, and the high utilisation of these substrates may increase the gross NH4+ production. Future studies are required to identify the specific organic substrates that are utilised by microorganisms under soil acidification.
Conclusions
Soil acidification reduced glycine uptake by microorganisms, while it increased uptake by maize. Decrease in the microbial biomass and changes in the community composition are the major factors affecting the competition between maize roots and microbes for amino acids. Soil acidification decreased organic N (amino acids, peptides, and proteins) mineralisation, but increased gross NH4+ production. Microbial activity declined because soil acidification not only reduced amino acid production (protein and peptide hydrolysis), but also affected processes associated with amino acid consumption, such as ammonification and subsequent nitrification (NH4+ and NO3− production). Enhanced amino acid uptake was observed in maize following soil acidification, owing to the decreases in the microbial biomass and changes in the community composition. However, this amino acid uptake remained insufficient to induce a corresponding increase of N nutrition in the plants.
References
Apostel C, Dippold M, Glaser B, Kuzyakov Y (2013) Biochemical pathways of amino acids in soil: assessment by position-specific labeling and 13C-PLFA analysis. Soil Biol Biochem 67:31–40
Åsa F, Tunlid A, Bååth E (2011) Use and misuse of PLFA measurements in soils. Soil Biol Biochem 43:1621–1625
Bao SD (2000) Soil and agricultural chemical analysis. China Agriculture Press, Beijing. 10
Biernath C, Fischer H, Kuzyakov Y (2008) Root uptake of N-containing and N-free low molecular weight organic substances by maize: a 14C/15N tracer study. Soil Biol Biochem 40:2237–2245
Bobbie RJ, White DC (1980) Characterization of benthic microbial community structure by high-resolution gas chromatography of fatty acid methyl esters. Appl Environ Microbiol 39:1212–1222
Braun J, Mooshammer M, Wanek W, Prommer J, Walker TWN, Rütting T, Richter A (2018) Full 15N tracer accounting to revisit major assumptions of 15N isotope pool dilution approaches for gross nitrogen mineralization. Soil Biol Biochem 117:16–26
Cheng Y, Wang J, Mary B, Zhang JB, Cai ZC, Chang SX (2013) Soil pH has contrasting effects on gross and net nitrogen mineralizations in adjacent forest and grassland soils in Central Alberta, Canada. Soil Biol Biochem 57:848–857
Corre MD, Brumme R, Veldkamp E, Beese FO (2007) Changes in nitrogen cycling and retention processes in soils under spruce forests along a nitrogen enrichment gradient in Germany. Glob Chang Biol 13:1509–1527
Farrell M, Hill PW, Farrar J, Bardgett RD, Jones DL (2011a) Seasonal variation in soluble soil carbon and nitrogen across a grassland productivity gradient. Soil Biol Biochem 43:835–844
Farrell M, Hill PW, Wanniarachchi SD, Farrar J, Bardgett RD, Jones DL (2011b) Rapid peptide metabolism: a major component of soil nitrogen cycling? Glob Biogeochem Cycles 25:GB3014
Fokin AD, Knyazev DA, Kuzyakov YV (1993) Destruction of C-14-labeled and N-15-labeled amino-acids and nucleic bases in soil and the supply of their transformation products to plants. Eurasian Soil Sci 25:109–122
Ganeteg U, Ahmad I, Jmtgrd S, Aguetoni-Cambui C, Nsholm T (2016) Amino acid transporter mutants of Arabidopsis provides evidence that a non-mycorrhizal plant acquires organic nitrogen from agricultural soil. Plant Cell Environ 40:413–423
Ge T, Li B, Zhu Z, Hu Y, Yuan H, Dorodnikov M, Jones DL, Wu J, Kuzyakov Y (2017) Rice rhizodeposition and its utilization by microbial groups depends on N fertilization. Biol Fertil Soils 53:37–48
Geisseler D, Horwath WR, Joergensen RG, Ludwig B (2010) Pathways of nitrogen utilization by soil microorganisms–a review. Soil Biol Biochem 42:2058–2067
Giagnoni L, Magherini F, Landi L, Taghavi S, Modesti A, Bini L, Nannipieri P, Van der Lelie D, Renella G (2011) Extraction of microbial proteome from soil: potential and limitations assessed through a model study. Eur J Soil Sci 62:74–81
Glanville HC, Hill PW, Schnepf A, Oburger E, Jones DL (2016) Combined use of empirical data and mathematical modelling to better estimate the microbial turnover of isotopically labelled carbon substrates in soil. Soil Biol Biochem 94:154–168
Greenfield LM, Hill PW, Seaton FM, Paterson E, Jones DL (2020) Is soluble protein mineralisation and protease activity in soil regulated by supply or demand? Soil Biol Biochem 150:108007
Guo JH, Liu XJ, Zhang Y, Shen JL, Han WX, Zhang WF, Christie P, Goulding K, Vitousek PM, Zhang FS (2010) Significant acidification in major Chinese croplands. Science 327:1008–1010
Guo N, Zhang S, Gu M, Xu G (2021) Function, transport, and regulation of amino acids: what is missing in rice? Crop J 9:530–542
Hagerty SB, Van Groenigen KJ, Allison SD, Hungate BA, Schwartz E, Koch GW, Kolka RK, Dijkstra P (2014) Accelerated microbial turnover but constant growth efficiency with warming in soil. Nat Clim Chang 4:903–906
Hawkins BJ, Robbins S (2010) pH affects ammonium, nitrate and proton fluxes in the apical region of conifer and soybean roots. Physiol Plant 138:238–247
Hgberg MN, Hgberg P, Myrold DD (2007) Is microbial community composition in boreal forest soils determined by pH, C-to-N ratio, the trees, or all three? Oecologia 150:590–601
Hill PW, Jones DL (2019) Plant–microbe competition: does injection of isotopes of C and N into the rhizosphere effectively characterise plant use of soil N? New Phytol 221:796–806
Hill PW, Farrell M, Roberts P, Farrar J, Grant H, Newsham KK, Hopkins DW, Bardgett RD, Jones DL (2011) Soil- and enantiomer-specific metabolism of amino acids and their peptides by Antarctic soil microorganisms. Soil Biol Biochem 43:2410–2416
Hill PW, Farrell M, Jones DL (2012) Bigger may be better in soil N cycling: does rapid acquisition of small l-peptides by soil microbes dominate fluxes of protein-derived N in soil? Soil Biol Biochem 48:106–112
Huang YH, Zhang TC (2004) Effects of low pH on nitrate reduction by iron powder. Water Res 38:2631–2642
Hylander LD, Svensson HI, Simán G (1996) Different methods for determination of plant available soil phosphorus. Commun Soil Sci Plant Anal 27:1501–1512
Jämtgård S, Näsholm T, Huss-Danell K (2008) Characteristics of amino acid uptake in barley. Plant Soil 302:221–231
Jan MT, Roberts P, Tonheim SK, Jones DL (2009) Protein breakdown represents a major bottleneck in nitrogen cycling in grassland soils. Soil Biol Biochem 41:2272–2282
Jérémy P, Jones B, Goodall T, Mang D, Griffiths R (2019) The pH optimum of soil exoenzymes adapt to long term changes in soil pH. Soil Biol Biochem 138:107601
Jiang L, Wang S, Zhe P, Xu X, Kardol P, Li Y, Zhang L, Wang Y, Lei Z, Lan Z (2018) Plant organic N uptake maintains species dominance under long-term warming. Plant Soil 433:243–255
Jones DL, Kielland K (2012) Amino acid, peptide and protein mineralization dynamics in a taiga forest soil. Soil Biol Biochem 55:60–69
Jones DL, Healey JR, Willett VB, Farrar JF, Hodge A (2005a) Dissolved organic nitrogen uptake by plants—an important N uptake pathway? Soil Biol Biochem 37:413–423
Jones DL, Kemmitt SJ, Wright D, Cuttle SP, Bol R, Edwards AC (2005b) Rapid intrinsic rates of amino acid biodegradation in soils are unaffected by agricultural management strategy. Soil Biol Biochem 37:1267–1275
Jones DL, Cooledge EC, Hoyle FC, Griffiths RI, Murphy DV (2019) pH and exchangeable aluminum are major regulators of microbial energy flow and carbon use efficiency in soil microbial communities. Soil Biol Biochem 138:107584
Kuzyakov Y (2010) Priming effects: interactions between living and dead organic matter. Soil Biol Biochem 42:1363–1371
Kuzyakov Y, Xu X (2013) Competition between roots and microorganisms for nitrogen: mechanisms and ecological relevance. New Phytol 198:656–669
Liang C, Schimel JP, Jastrow JD (2017) The importance of anabolism in microbial control over soil carbon storage. Nat Microbiol 2:1–6
Ma Q, Cao X, Xie Y, Yan G, Wu L (2016) Effect of pH on the uptake and metabolism of glycine in pak choi (Brassica chinensis L.). Environ Exp Bot 133:139–150
Ma Q, Wu L, Wang J, Ma J, Zheng N, Hill PW, Chadwick DR, Jones DL (2018) Fertilizer regime changes the competitive uptake of organic nitrogen by wheat and soil microorganisms: an in-situ uptake test using 13C, 15N labelling, and 13C-PLFA analysis. Soil Biol Biochem 125:319–327
Ma Q, Wen Y, Pan W, Macdonald A, Jones DL (2020a) Soil carbon, nitrogen, and Sulphur status affects the metabolism of organic S but not its uptake by microorganisms. Soil Biol Biochem 149:107943
Ma Q, Wen Y, Wang D, Sun X, Jones DL (2020b) Farmyard manure applications stimulate soil carbon and nitrogen cycling by boosting microbial biomass rather than changing its community composition. Soil Biol Biochem 144:107760
Ma Q, Kuzyakov Y, Pan W, Tang S, Chadwick DR, Wen Y, Hill PW, Macdonld A, Ge T, Si L, Wu L, Jones DL (2021a) Substrate control of organic Sulphur utilisation and microbial stoichiometry in soil: results of 13C, 15N, 14C, and 35S quad labelling. ISME J 15:3148–3158
Ma Q, Pan W, Tang S, Sun X, Xie Y, Chadwick DR, Hill PW, Si L, Wu L, Jones DL (2021b) Maize and soybean experience fierce competition from soil microorganisms for the uptake of organic and inorganic nitrogen and Sulphur: a pot test using 13C, 15N, 14C, and 35S labelling. Soil Biol Biochem 157:108260
Malik AA, Jeremy P, Buckeridge KM, Tim G, Nico J, Somak C, Soon GH, Peyton JM, Mason KE, Maaike VA (2018) Land use driven change in soil pH affects microbial carbon cycling processes. Nat Commun 9:3591
Mariano E, Jones DL, Hill PW, Trivelin PCO (2016) Mineralisation and sorption of dissolved organic nitrogen compounds in litter and soil from sugarcane fields. Soil Biol Biochem 103:522–532
Mckane RB, Johnson LC, Shaver GR, Nadelhoffer KJ, Rastetter EB, Fry B, Giblin AE, Kielland K, Kwiatkowski BL, Laundre JA (2002) Resource-based niches provide a basis for plant species diversity and dominance in arctic tundra. Nature 415:68–71
Moran-Zuloaga D, Dippold M, Glaser B, Kuzyakov Y (2015) Organic nitrogen uptake by plants: reevaluation by position-specific labeling of amino acids. Biogeochemistry 125:359–374
Mulvaney RL, Khan SA (2001) Diffusion methods to determine different forms of nitrogen in soil hydrolysates. Soil Sci Soc Am J 65:1284–1292
Nannipieri P, Eldor P (2009) The chemical and functional characterization of soil N and its biotic components. Soil Biol Biochem 41:2357–2369
Nsholm T, Kielland K, Ganeteg U (2009) Uptake of organic nitrogen by plants. New Phytol 182:31–48
Orchard VA, Cook FJ (1983) Relationship between soil respiration and soil moisture. Soil Biol Biochem 15:447–453
Owen AG, Jones DL (2001) Competition for amino acids between wheat roots and rhizosphere microorganisms and the role of amino acids in plant N acquisition. Soil Biol Biochem 33:651–657
Pietri JA, Brookes PC (2008) Relationships between soil pH and microbial properties in a UK arable soil. Soil Biol Biochem 40:1856–1861
Pietri JCA, Brookes PC (2009) Substrate inputs and pH as factors controlling microbial biomass, activity, and community structure in an arable soil. Soil Biol Biochem 41:1396–1405
Rasmussen J, Sauheitl L, Eriksen J, Kuzyakov Y (2010) Plant uptake of dual-labeled organic N biased by inorganic C uptake: results of a triple labeling study. Soil Biol Biochem 42:524–527
Reid R, Hayes J (2003) Mechanisms and control of nutrient uptake in plants. Int Rev Cytol 229:73–115
Rousk J, Brookes PC, Bååth E (2010) The microbial PLFA composition as affected by pH in an arable soil. Soil Biol Biochem 42:516–520
Saichek RE, Reddy KR (2003) Effect of pH control at the anode for the electrokinetic removal of phenanthrene from kaolin soil. Chemosphere 51:273–287
Sauheitl L, Glaser B, Weigelt A (2009) Uptake of intact amino acids by plants depends on soil amino acid concentrations. Environ Exp Bot 66:145–152
Schmidt PS, Zhu CT, Das J, Batavia M, Yang L, Eanes WF (2008) An amino acid polymorphism in the couch potato gene forms the basis for climatic adaptation in Drosophila melanogaster. Proc Natl Acad Sci 105:16207–16211
Schmidt SM, Lukasiewicz J, Farrer R, Van Dam P, Bertoldo C, Rep M (2016) Comparative genomics of fusarium oxysporum f. sp. melonis reveals the secreted protein recognized by the Fom-2 resistance gene in melon. New Phytol 209:307–318
Singh BK, Campbell CD, Sorenson SJ, Zhou J (2009) Soil genomics. Nat Rev Microbiol 7:758–759
Tang S, Ma Q, Luo J, Xie Y, Pan W, Zheng N, Liu M, Wu L (2021) The inhibition effect of tea polyphenols on soil nitrification is greater than denitrification in tea garden soil. Sci Total Environ 778:146328
Thornton B, Zhang Z, Mayes RW, Högberg MN, Midwood AJ (2011) Can gas chromatography combustion isotope ratio mass spectrometry be used to quantify organic compound abundance? Rapid Commun Mass Spectrom 25:2433–2438
Vranova V, Zahradnickova H, Janous D, Skene KR, Matharu AS, Rejsek K, Formanek P (2012) The significance of D-amino acids in soil, fate and utilization by microbes and plants: review and identification of knowledge gaps. Plant Soil 354:21–39
Wanek W, Mooshammer M, Blöchl A, Hanreich A, Richter A (2010) Determination of gross rates of amino acid production and immobilization in decomposing leaf litter by a novel 15N isotope pool dilution technique. Soil Biol Biochem 42:1293–1302
Warren C (2021) What are the products of enzymatic cleavage of organic N? Soil Biol Biochem 154:108152
Wu YY, Xing DK (2012) Effect of bicarbonate treatment on photosynthetic assimilation of inorganic carbon in two plant species of Moraceae. Photosynthetica 50(4):587–594
Xiao K, Xu J, Tang C, Zhang J, Brookes PC (2013) Differences in carbon and nitrogen mineralization in soils of differing initial pH induced by electrokinesis and receiving crop residue amendments. Soil Biol Biochem 67:70–84
Xu X, Stange CF, Richter A, Wanek W, Kuzyakov Y (2008) Light affected the competition for inorganic and organic nitrogen between maize and soil microorganisms. Plant Soil 304:59–72
Yeomans JC, Bremner JM (1988) A rapid and precise method for routine determination of organic carbon in soil. Commun Soil Sci Plant Anal 19:1467–1476
Yuntao H, Qing Z, Shasha Z, Lisa N, Wolfgang W (2018) Significant release and microbial utilization of amino sugars and d-amino acid enantiomers from microbial cell wall decomposition in soils. Soil Biol Biochem 123:115–125
Zamanian K, Zarebanadkouki M, Kuzyakov Y (2018) Nitrogen fertilization raises CO2 efflux from inorganic carbon: a global assessment. Glob Chang Biol 24:2810–2817
Zeier J (2013) New insights into the regulation of plant immunity by amino acid metabolic pathways. Plant Cell Environ 36:2085–2103
Zhang S, Zheng Q, Noll L, Hu Y, Wanek W (2019) Environmental effects on soil microbial nitrogen use efficiency are controlled by allocation of organic nitrogen to microbial growth and regulate gross N mineralization. Soil Biol Biochem 135:304–315
Zhao M, Zhao J, Yuan J, Hale L, Wen T, Huang Q, Vivanco JM, Zhou J, Kowalchuk GA, Shen Q (2021) Root exudates drive soil-microbe-nutrient feedbacks in response to plant growth. Plant Cell Environ 44:613–628
Acknowledgements
This research was financed by the National Natural Science Foundation of China (31872180; 32172674; 32102488) and the National Key Research and Development Program of China (2020YFD1100402). YK offers thanks for the support of the Government Program of Competitive Growth of Kazan Federal University, RUDN University Strategic Academic Leadership Program, Government Program of Competitive Growth of Kazan Federal University, and West-Siberian Interregional Science and Education Center’s project “TerrArctic” (89-DON 1).
Author information
Authors and Affiliations
Contributions
Wankun Pan, Qingxu Ma, Sheng Tang, and Lianghuan Wu conceived and designed the study, analysed data, and drafted the manuscript; Jingjie Zhou, Mengjiao Liu, and Meng Xu collected the data; Yakov Kuzyakov interpreted the results, and revised the manuscript and data presentations.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Ad C. Borstlap.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pan, W., Tang, S., Zhou, J. et al. Plant–microbial competition for amino acids depends on soil acidity and the microbial community. Plant Soil 475, 457–471 (2022). https://doi.org/10.1007/s11104-022-05381-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-022-05381-w