Abstract
Key message
A novel MADS-box member SiMADS34 is essential for regulating inflorescence architecture and grain yield in Setaria italica.
Abstract
MADS-box transcription factors participate in regulating various developmental processes in plants. Inflorescence architecture is one of the most important agronomic traits and is closely associated with grain yield in most staple crops. Here, we isolated a panicle development mutant simads34 from a foxtail millet (Setaria italica (L.) P. Beauv.) EMS mutant library. The mutant showed significantly altered inflorescence architecture and decreased grain yield. Investigation of agronomic traits revealed increased panicle width by 16.8%, primary branch length by 10%, and number of primary branches by 30.9%, but reduced panicle length by 25.2%, and grain weight by 25.5% in simads34 compared with wild-type plants. Genetic analysis of a simads34 × SSR41 F2 population indicated that the simads34 phenotype was controlled by a recessive gene. Map-based cloning and bulked-segregant analysis sequencing demonstrated that a single G-to-A transition in the fifth intron of SiMADS34 in the mutant led to an alternative splicing event and caused an early termination codon in this causal gene. SiMADS34 mRNA was expressed in all of the tissues tested, with high expression levels at the heading and panicle development stages. Subcellular localization analysis showed that simads34 predominantly accumulated in the nucleus. Transcriptome sequencing identified 241 differentially expressed genes related to inflorescence development, cell expansion, cell division, meristem growth and peroxide stress in simads34. Notably, an SPL14–MADS34–RCN pathway was validated through both RNA-seq and qPCR tests, indicating the putative molecular mechanisms regulating inflorescence development by SiMADS34. Our study identified a novel MADS-box member in foxtail millet and provided a useful genetic resource for inflorescence architecture and grain yield research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Foxtail millet (Setaria italica (L) P. Beauv.) is an ancient C4 annual crop of dryland cultivation. It is a self-pollinating crop with chromosome number 2n = 18, and classified under the family Poaceae and subfamily Panicoideae (Fedorov 1974). Foxtail millet has been domesticated and has become an important crop grown worldwide; in particular, it has been grown in a wide area of northern China and East Asia for 10,000–11,000 years (Lu et al. 2009; Yang et al. 2012). It is the second most important millet (after pearl millet) and is distributed across warm and temperate regions of the world including Asia, Europe, America, Australia and Africa. Used as grain, forage or bird feed, foxtail millet is grown on approximately 2 million ha in China and produces nearly 6 Tg of grain per year (Diao 2011). China has been recognized as the center of origin and improvement of foxtail millet, the national gene bank of China conserving over 80% of the world’s Setaria accessions. Genetic characterization of DNA sequence polymorphisms from green foxtail germplasm is pivotal for analyzing domestication, evolution, and potential for breeding in wild grass species (Huang et al. 2014). To increase the yield and improve the quality of foxtail millet through traditional genetic improvement methods takes much time, because the yield and other agronomic traits are quantitative.
Genetic and phenotypic characterization of foxtail millet could provide valuable information to help explore the genetic variability and could be helpful in breeding programs and improving crops. Efforts in breeding research revealed that most inflorescence types are controlled by major genes, with complex inheritance patterns, while most agronomic traits are regulated by multiple genes or quantitative trait loci (QTLs), including plant height, inflorescence length, heading date, and nutrition-related characteristics (Diao and Jia 2017). Inflorescence development constitutes one of the most essential traits that determine the yield of many crops. Therefore, optimization of inflorescence size and architecture has become a priority target for high yield in breeding programs. According to previous reports, inflorescence size, inflorescence branching, pattern and number of spikelets are potentially determined by separate genetic mechanisms (Doust et al. 2005; Jia et al. 2013a). Inflorescence type in crops is a complex character that is affected by environment (Li et al. 1935). Earlier studies confirmed that heritability of inflorescence length is approximately 75% (Liu 1984; Diao and Jia 2017) and is closely associated with grain yield (Jia et al. 2013a).
MADS-box genes are key regulators of floral organ determination and inflorescence architecture in plants. The MADS-box family is characterized by a highly conserved N-terminal domain with a length of 58–60 amino acids known as the MADS domain (Passmore et al. 1988), which was named according to the first four members of this family, that is MINI-CHROMOSOME MAINTENANCE1 (MCM1) from yeast (Sommer et al. 1990), AGAMOUS (AG) from Arabidopsis thaliana (Yanofsky et al. 1990), DEFICIENS (DEF) from Antirrhinum majus (Norman et al. 1988), and SERUM RESPONSE FACTOR (SRF) from Homo sapiens (Honma and Goto 2001). Gramzow and Theißen (2010) reported that MADS-box genes have conserved functions in reproductive organ identity across different plant species. Previous studies indicated that MADS-box genes possess DNA-binding and dimerization functions (Hu and Liu 2012; Theißen et al 2016) and regulate flower development in Arabidopsis, rice and maize (Wang et al. 2008, 2012; Zhang et al. 2012). The MADS-box protein structure can be divided into four domains. The N-terminal end is the highly conserved DNA-binding domain; next to the MADS domain are the moderately conserved Intervening (I) and Keratin-like (K) domains, which are involved in specific protein–protein interactions (Jack 2004). The carboxyl-terminal (C) domain is highly variable and is involved in transcriptional activation and assembly of heterodimers and multimeric protein complexes (Riechmann and Meyerowitz 1997). The MADS domain proteins can bind to the DNA sequence CC[A/T]6GG which is also termed as the CArG-box (West et al 1997).
So far, the MADS-box gene family has been widely investigated in angiosperms, particularly in the model plant Arabidopsis thaliana (Ma 1991). The floral organ identity MADS-box genes have been divided into A, B, C, D, and E classes (Theissen 2001), among which the E-class genes include SEPALLATA1 (SEP1), SEP2, SEP3, SEP4 and OsMADS34 (Ditta et al. 2004). Among these genes, SEP4 in Arabidopsis and OsMADS34 in rice were reported to be associated with inflorescence architecture and floral organ development. OsMADS34 encodes a MADS-box protein and is considered a key regulator of rice inflorescence development. The number of primary branches, as well as spikelet number and spikelet morphology, were changed in the osmads34 mutant. OsMADS34 is also essential for sterile lemma identity and is required to prevent the formation of lemma/leaf-like organs (Gao et al. 2010). In Arabidopsis, SEP1, SEP2, SEP3, and SEP4 contribute to the development of stamens, petals, and carpels, as well as sepals, and play essential roles in meristem identity (Ditta et al. 2004). SEP4, together with SOC1, AGL24, and SVP, also plays important roles in regulating inflorescence branching in Arabidopsis (Liu et al. 2013).
Many previous studies indicate the importance of MADS genes; however, little research has been carried out in foxtail millet. In this study, we isolated a panicle morphology mutant from an ethyl methanesulfonate (EMS) mutant library of S. italica. Phenotypic characterization of the mutant simads34 has demonstrated significant differences in panicle size and primary branch lengths. The candidate gene was identified using map-based cloning and bulked-segregant analysis (BSA-) sequencing. A novel MADS-box member SiMADS34 was identified which is predicted to be responsible for the mutant phenotypes. Gene function characterization and transcriptome analysis indicate that SiMADS34 regulates panicle development in multiple regulatory pathways. In addition, this study provides useful information regarding inflorescence architecture and grain yield in foxtail millet.
Materials and methods
Plant materials and construction of the mapping population
For map-based cloning, the simads34 mutant was crossed with a foxtail millet cultivar SSR41, and the hybrids were self-pollinated to generate the F2 mapping population. For phenotype measurement and RNA-seq analysis, the simads34 mutant was backcrossed with wild-type Yugu1 three times and progeny of the recessive individuals from BC3F2 were used in subsequent experiments.
Plant growth and characterization of the agronomic traits
The plants were grown in an experimental field of the Chinese Academy of Agricultural Science, Beijing, China (40°N, 116°E) during the growth period from June to October in two consecutive seasons (2017 and 2018). Appropriate soil moisture content of 20% was achieved through field irrigation before cultivation. Seedlings were thinned manually at the five-leaf stage as recommended density which is 500,000 individuals per hectare. Five individuals of each experimental plot were used for agronomic traits measurements.
Agronomic traits of mutant simads34 and wild-type Yugu1 plants were measured (plant height, stem diameter, panicle length, flag leaf width, flag leaf length, panicle diameter, panicle weight, number of primary branches, primary branch length, number of seeds per primary branch, primary branch diameter, peduncle length, number of internodes, thousand seed weight, number of seeds per panicle, and grain weight per plant). Primary branch density was calculated as the number of primary branches per cm of mature panicle. Field management during the growing season (irrigation, weed management and fertilization) was done by trained farmers. Ten uniformly developed individuals were collected at the corresponding stages, and specific methodology applied to meet the standards of phenotype scoring (Jia et al. 2013b).
Statistical analysis
Agronomic data analysis was carried out using the SAS statistics program (SAS 9.2). The analyses included analysis of variance (ANOVA), means and standard error for both 2017 and 2018 seasons and combined analysis (Der and Everitt 2008).
Molecular mapping and identification of the candidate gene
In total, 395 recessive individuals resulting from the mutant simads34 × SSR41 F2 population were used to determine the candidate gene. DNA was extracted from young leaves following a standard CTAB protocol (Murray and Thompson 1980). Thermo Cycler (PCR-machine) and PCR Mix (TSE006, Tsingke) were used for DNA amplification. PCR products were separated by 8% polyacrylamide gel electrophoresis followed by silver staining, drying and photography (Bassam et al.1991). Forty-five polymorphic SSR markers uniformly representing the nine foxtail millet chromosomes were selected for primary mapping. Fine mapping was carried out with 240 F2 recessive individuals. Primers were designed using Primer 5.0 software and synthesized by Sangon Company (Shanghai, China). Primers used for gene mapping are listed in Supplementary Table S1.
To identify the candidate gene, whole-genome resequencing and MutMap analysis were used. DNA samples were extracted from individuals of the simads34 × Yugu1 BC1F2 population. Four DNA pools were constructed, comprising a recessive mutant individuals mixed pool, wild-type individuals mixed pool, female parental pool, and male parental pool. Whole-genome resequencing was carried out on the Illumina HiSeq 2500 platform. Using MutMap, the candidate gene was located in the 4.490–7.867 Mb region of chromosome 9. The sequencing reads generated from a DNA pool of 30 BC1F2 recessive individuals were aligned to the S. italica reference genome (phytozome.jgi.doe.gov). We calculated the SNP index value and all of the SNPs located in the candidate region with index value = 1 were collected for phenotype-relevant SNP verification. We selected the 4.490–7.867 Mb region of chromosome 9 as the candidate interval, and extracted a total of 226 SNPs and Indels. The SNP index represents the allele frequency of SNPs in different gene pools. A value of the SNP index > 0.9 means the SNP is homozygous; a SNP index value from 0.3 to 0.9 indicates the SNP is heterozygous. The SNP index of the candidate mutation site was 1.0, indicating a pure recessive mutation in the mutant pool, and it was heterozygous in the dominant pool.
Sequencing and phylogenetic analysis of the candidate gene
Reference sequences of the candidate gene set within the mapping region were retrieved from the S. italica genome project v2.2. Genes within the mapped region were PCR amplified and the PCR products were sequenced using an Applied Biosystems 3730 sequencer (Applied Biosystems, Foster City, CA, USA) and analyzed by DNAMAN8 software (Lynnon Biosoft, Quebec, Canada). A phylogenetic tree was constructed using MEGA 7.0 software (Tamura et al. 2011).
Vector construction and subcellular localization
The coding sequence of simads34 was fused at the C terminus with GFP in the pAN580 vector, and then plasmids were co-transformed into Setaria leaf protoplasts (Zhang et al. 2011) to explore the subcellular localization. The primers used are shown in Supplementary Table S2. Staining with 4′,6-diamidino-2-phenylindole (DAPI) indicated the nucleus. The GFP and DAPI signals were detected using an inverted fluorescence microscope (Zeiss LSM880).
RNA extraction and alternative splicing analysis
Wild-type Yugu1 and simads34 seedlings were used for total RNA extraction using the Purelink RNA kit (cat no. 12183018, Invitrogen, UK). cDNAs were generated by reverse transcription using a PrimeScript first-strand cDNA synthesis kit (cat. no. 6210A, TakaRa, Otsu Shiga, Japan). The cDNAs were used for identifying the simads34 transcript sequence in Yugu1 and simads34 mutants. The primers used for sequencing are listed in Supplementary Table S2. For tissue-specific expression analysis, 13 tissues from different developmental stages of wild-type Yugu1 were collected for total RNA sequencing. The transcriptome data can be accessed from the website www.setariamodel.cn.
Transcriptome sequencing of simads34 and quantitative real-time PCR (qRT-PCR) validation
Mutant simads34 and wild-type Yugu1 plants were grown in a growth chamber for 5 weeks with 10 h light at 28 °C and 14 h dark at 25 °C. The young panicles were harvested at the growing point differentiation stage, which was 1.5–2 cm panicle length (Fig. 3a below). Total RNA was extracted for transcriptome sequencing. RNA quality and purity were examined using an Agilent Bioanalyzer 2100 (Agilent Technologies, Waldbronn, Germany). The RNA-seq results were analyzed according to our previous report (Tang et al. 2017). The same plant samples were used for quantitative real-time PCR (qRT-PCR) validation. The experiment was performed on an Applied Biosystems 7300 Analyzer (Applied Biosystems, Foster City, USA) with Fast Start Universal SYBR Green Master mix (ROX) (Roche, Mannheim, Germany). Relative gene expression levels were calculated using the 2−ΔΔCt method.
Results
Isolation of a foxtail millet panicle development mutant with decreased grain yield
The simads34 mutant, which was isolated from an EMS-induced mutant library, had an abnormal panicle morphology compared with the wild-type Yugu1. The whole plant stature of simads34 was almost the same as that of the wild-type, except for the panicle (Fig. 1a; Table 1). Substantial differences were observed throughout the panicle formation process (Fig. 1b). The mutant panicle showed abnormal structure from day 1–12 after formation of the panicle (Fig. 1b). Throughout panicle development the mutant phenotype became progressively more severe (Fig. 1b). Moreover, simads34 had significantly wider panicles compared with those of Yugu1. However, the average panicle length, panicle weight, and grain weight per panicle of simads34 were lower than those of Yugu1 (Fig. 1c–d; Table 1).
Morphological and statistical comparisons between wild-type Yugu1 and mutant simads34. a General stature of Yugu1 (left) and simads34 (right) at the maturity stage. Bar = 15 cm. b Panicle development stage of Yugu1 (left) and simads34 (right). Bar = 1.5 cm. c Panicle morphology of Yugu1 (left) and simads34 (right). Bar = 8 cm. d Statistical comparisons of panicle length, panicle diameter, panicle weight and grain weight per panicle between Yugu1 and simads34. Values are means ± SD (n = 10 for each agronomic trait). Significant differences were determined using Student’s t-test (*P < 0.05, **P < 0.01)
These panicle development differences were seen as early as the inflorescence meristem (IM) differentiation stage (Fig. 2a–h). Some of the primary branches did not develop at the top of the IM (Fig. 2c, d). A wider and longer primary branch was observed at the same time as the growth of IM in simads34 compared with the wild-type (Fig. 2g, h). The abnormal panicle development ultimately affected the grain yield. The 1000-grain weight and number of seeds per panicle of the mutant were about 16% and 26% lower than those of the wild-type, respectively (Fig. 2i; Table 1). However, the seed length, seed width, and seed area of wild-type and simads34 were almost identical, indicating that the final grain yield reduction was attributable to panicle size, panicle shape, and grain filling.
Morphological observation of panicles at early differentiation stage and statistical comparisons of yield-related traits between wild-type Yugu1 and mutant simads34. a Morphology of panicle of Yugu1 at early differentiation stage. Bar = 0.5 mm. b Morphology of panicle of simads34 at early differentiation stage. Bar = 0.5 mm. c Apical panicle morphology of Yugu1. Bar = 0.5 mm. d Apical panicle morphology of simads34. Bar = 0.5 mm. e Basal panicle morphology of Yugu1. Bar = 0.5 mm. f Basal panicle morphology of simads34. Bar = 0.5 mm. g Morphology of young panicle of Yugu1. Bar = 2 mm. h Morphology of young panicle of simads34. Bar = 2 mm. i Statistical comparisons in 1000-grain weight, no. of seeds per panicle, seed length, seed width and seed area. Values are means ± SD (n = 10 for each agronomic trait). Significant differences were determined using Student’s t-test
Mutant simads34 exhibited abnormal inflorescence architecture
Detailed phenotype investigation was carried out to further understand the effect on panicle morphology in the mutant. Morphological and statistical comparisons of young and developed primary branches in apical, middle and basal panicles of simads34 and wild-type Yugu1 showed highly significant differences (Fig. 3a, b). The lengths of primary branches at the apical, middle and basal panicles in simads34 were longer by 59%, 58.8% and 7.3%, respectively, compared with the wild-type (Fig. 3b, c). In addition, the widths of primary branches at the apical, middle and basal panicles in simads34 were larger by 33.3%, 20% and 6.5%, respectively, compared with the wild-type. Overall, these comparisons showed less variation between the mutant and wild-type in primary branches at the basal panicle, while the variation was more notable at apical and middle panicles (Fig. 3c–g).
Microscopic observation of developing panicles, and morphological and statistical comparisons between wild-type Yugu1 and mutant simads34. a Morphology of young panicle of Yugu1 (left) and simads34 (right). Bar = 5 mm. b Morphology of young primary branches of Yugu1 (left) and simads34 (right). Bar = 1 mm. c Morphology of developed primary branches of Yugu1 (upper panel) and simads34 (lower panel). Bar = 1 cm. d Statistical comparison of young primary branch length of Yugu1 and simads34. e Statistical comparison of basal primary branch length of Yugu1 and simads34. f Statistical comparison of middle primary branch length of Yugu1 and simads34. g Statistical comparison of apical primary branch length of Yugu1 and simads34. Significant differences in D–G were determined using Student’s t-test (*P < 0.05, **P < 0.01)
These results confirmed that altered morphology of primary branches had a major and direct impact on the panicle architecture of simads34. In addition, we made a detailed analysis of the number of branches. As shown in Fig. 4, the number of primary branches was slightly higher in the simads34 mutant but not to a significant extent (Fig. 4a), while primary branch density was significantly higher (P = 5.91 × 10−9) in the mutant (Fig. 4b). We also investigated secondary branches. The results showed that the simads34 mutant had markedly more secondary branches than the wild-type (Fig. 4c, d). These results are consistent with those of previous studies on sep4/mads34 mutants in other species (Liu et al. 2013; Soyk et al. 2017).
Morphological variations in panicle branches. a Statistical comparison of no. of primary branches of Yugu1 and simads34. b Statistical comparison of density of primary branches of Yugu1 and simads34. Density value was calculated as number of primary branches per cm. c Morphology of panicle secondary branches of Yugu1 and simads34. Bar = 5 mm. d Statistical comparison of no. of secondary branches between Yugu1 and simads34. Significant differences were determined using Student’s t-test (n = 5)
Combined BSA-seq and map-based cloning identified the causal gene SiMADS34, which encodes a MADS-box transcription factor
To identify the candidate gene, simads34 was hybridized with SSR41: the resultant F1 generation showed normal panicle structure. Among the F2 population, we identified 395 individuals with mutant panicle structure, and 1,324 individuals with normal panicle structure. Genetic analysis indicated the segregation ratio of wild-type to mutant phenotype of the F2 population was 3:1 (χ2 = 3.64 < χ20.05,1 = 3.84), suggesting that simads34 was controlled by a single recessive nuclear gene (Supplementary Table S3).
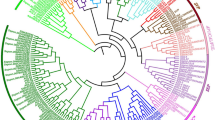
To map the locus responsible for the simads34 phenotype, we performed bulked-segregant analysis sequencing (BSA-seq) using a simads34 × wild-type Yugu1 BC1F2 population. DNA of the mutant plants and the wild-type individuals were pooled, and according to the resequencing results, the candidate gene was located in the 4.49–7.867 Mb interval of chromosome 9 (Fig. 5a). For further identification of the candidate gene, molecular markers were designed, based on the genome sequence of the candidate region. Using these molecular markers for screening of 240 F2 recessive individuals, we located the mutant gene in the 831-Kb region between the markers In9-4.829 and In9-5.66 (Chr. 9: 5,335,699–5,348,777 bp). The molecular marker In9-5.66 was closely linked to the candidate gene (Supplementary Table S4). We then screened for genomic variations where the SNP index of the recessive individual pool ≥ 0.9 and the SNP index was heterozygous in the dominant individual pool. Crucially, a causal SNP at 5,338,601 bp on chromosome 9 was identified. It was a G-to-A transition located in the genic region of Seita.9G088700 (Fig. 5b), indicating this gene might be responsible for the mutant phenotype. A BLAST search of Seita.9G088700 suggested that the causal gene encodes a MADS-box protein. MADS-box genes have many functions in plant growth and floral organ determination, and are essential for inflorescence development. A phylogenetic tree was constructed to examine the relationship of SiMADS34 protein with that of Arabidopsis and O. sativa (Goodstein et al. 2012) homologs. The phylogenetic tree grouped these MADS-box proteins into eight clades (Fig. 5C; Supplementary Table S5). The closest members of the MADS-box protein SEP-like (E) subfamily are OsMADS34 (O. sativa) and SEP4/AGL3 (Arabidopsis). Previous results identified OsMADS34 as a key regulator in the control of spikelet meristem and flower identity in rice (Agrawal et al. 2005; Gao et al. 2010; Kobayashi et al. 2009). In view of this similarity, we named Seita.9G088700 as SiMADS34. Previous research in Arabidopsis demonstrated that the SEP4 gene, together with three flower-related genes, regulate meristem specification and flower formation (Liu et al. 2013).
Identification of the SiMADS34 locus. a SNP index analysis: the X-axis shows physical positions on the nine foxtail millet chromosomes. The Y-axis shows the △SNP index, calculated according to Nakata et al. 2018. One candidate region was identified, located on chromosome 9. b Linked marker results for simads34 × SSR41 population. ♂: SSR41; ♀: simads34; F1: F1 population; P: DNA pool of recessive individuals of F2 population. Also fine mapping of mutant gene using map-based approaches; the vertical lines represent chromosome and marker names. Numbers below the horizontal lines indicate the physical distances between adjacent markers. c Phylogenetic relationship between simads34 and its homologous proteins in Arabidopsis and rice. The phylogenetic tree was constructed using the deduced full-length protein sequences of simads34 and other MADS proteins selected from previous publications. MEGA7 software was used with the maximum likelihood method. All protein accession IDs and sequences are listed in Supplementary Table S5
The simads34 single mutant displays normal floral organ identity
We named the mutant simads34 because map-based gene cloning and BSA-seq indicated that its defects are likely to be caused by mutation of a MADS-box gene that is homologous to OsMADS34 (see above). The MADS34 gene forms an integral part of the ABCDE model of MADS-box transcription factors that outlines the molecular basis of floral organ determination. To establish whether floret (spikelet) development was affected in simads34, we compared single floret structure between simads34 and wild-type plants. As shown in Fig. 6, all floral organs, including empty glume, lemma, palea, stamen, and pistil, displayed similar phenotype between simads34 and Yugu1, which indicated that the simads34 single mutation does not affect floral organ identity in Setaria (Fig. 6a–h).
Comparison of single floret structure between wild-type and simads34 mutant. a Morphology of floral organs of Yugu1. Bar = 1 mm. b Floret structure of Yugu1. Bar = 0.5 mm. c Morphology of floral organs of simads34. eg, empty glume; le, lemma; pa, palea; st, stamen pi, pistil. Bar = 1 mm. d Floret structure of simads34. Bar = 0.5 mm. e Morphology of necrotic floret of simads34. Bar = 1 mm. f Morphology of well-developed floret of Yugu1. Bar = 1 mm. g Empty glume morphology of Yugu1. Bar = 1 mm. h Empty glume morphology of simads34. Bar = 1 mm
However, both young and developed primary branches of simads34 panicles showed an apical abortion phenotype (Figs. 3a, b, 6e, f). A previous study indicated that apical abortion can cause a reduction in fertile spikelets and grain yield, and degeneration of spikelets at the inflorescence apex during late stage development might result from programmed cell death (Heng et al. 2018).
A single nucleotide change resulted in the alternative splicing of SiMADS34, which disturbs the K-box domain in the mutant
Sequencing of the full-length genomic DNA of SiMADS34 and Yugu1 confirmed a single nucleotide G-to-A transition existed at the join site of the fifth intron and exon of SiMADS34. Using RT-PCR and gel electrophoresis analyses of SiMADS34, a clear difference was detected at the RNA level. The splice site in the mutant was changed, which led to an intron retention (Fig. 7a, b). Domain analysis revealed that the K-box domain structure in the mutant has been severely disturbed compared with the wild-type (Fig. 7c, d).
Characterization of SiMADS34 in wild-type (Yugu1) and simads34 mutant. a PCR analysis of simads34 at the RNA level. Different bands were detected by RT-PCR in the simads34 mutant compared with the wild-type (Yugu1). b Comparison of transcripts and conserved domains between Yugu1 and simads34. c Comparison of aligned protein sequences of simads34 and its homologs from other species. d Comparison of domain structure in Yugu1 and simads34; the structural modeling of proteins was performed using SWISS-MODEL (swissmodel.expasy.org). In the simads34 mutant the MADS domain was not affected, while the K-box domain structure was disrupted
Expression patterns of SiMADS34
To investigate the tissue-specific expression pattern of SiMADS34 in foxtail millet, we evaluated its gene expression levels in different tissues and at different developmental stages. SiMADS34 was detected at the heading stage and all different stages of panicle development (Fig. 8a). We investigated the subcellular localization of SiMADS34 and determined that SiMADS34–GFP fusion protein predominantly accumulated in the nuclei of foxtail millet protoplasts (Fig. 8b).
Expression pattern of SiMADS34. a Characterization of SiMADS34 expression levels in different foxtail millet organs using RNA-seq. Mean expression levels and standard deviations were calculated by the TPM (Transcript per million) method. b Subcellular localization of SiMADS34–GFP fusion protein. Bar = 5 μm
Transcriptome sequencing and qPCR identified putative key biological process and candidate genes related to SiMADS34
To explore the molecular function and candidate pathways regulated by SiMADS34 in foxtail millet, we analyzed the transcriptome of young panicles of both mutant and wild-type plants. In total, 27,959 genes were detected in young panicles, of which 5,405 were differentially expressed (Log2FC ≥ 1 or ≤ − 1, and FDR ≤ 0.001) in the simads34 mutant compared with Yugu1. GO enrichment analysis indicated that numerous biological processes were significantly disturbed by the loss function of SiMADS34, including flower development, cell cycle regulation, abiotic stimulus responses, DNA transcription, chromatin assembly, and organic nitrogen metabolism (Fig. 9a). Pathway enrichment analysis showed that 19 molecular pathways were enriched for differentially expressed genes, most of them related to amino acid and carbohydrate metabolism. Photosynthesis and plant hormone signal transduction related pathways were also significantly enriched (Fig. 9b).
Enrichment analysis and candidate differentially expressed genes in the simads34 mutant. a Enriched GO terms for differentially expressed genes in simads34. Each circle represents a biological process GO term, and the lines show the relationship among different terms. Intensity of red color denotes extent of enrichment. b KEGG pathway enrichment bubble diagram of differentially expressed genes. Each circle represented a KEGG pathway, the names of which are shown in the legend on the left. The size of the circle represents the number of genes and the color gradient represents the extent of enrichment. Y-axis shows the gene ratio a to b, where a is the number of differentially expressed genes in the pathway, b is the total number of genes in related pathways. c Expression patterns of candidate genes in significantly enriched biological function categories. Each box represents a gene, and the color gradient represents the gene expression value. Gene ID and its expression value corresponding to each box are listed in Supplementary Table S6. d Transcript level analysis of candidate genes with high potential for acting as the downstream genes of SiMADS34. qRT-PCR was used to test the gene expression level in simads34 and wild-type plants. Young panicles of 1.5–2 cm in length were sampled for the experiment. The foxtail millet cullin gene (Seita.3G037700) was used as the internal control. Data are means ± SE (n = 3). Primers used for qRT-PCR are listed in Supplementary Table S7
By comparison of the results for gene expression profiles, gene functions, GO classification and KEGG enrichment, we identified four key biological processes and their related genes that might be regulated by SiMADS34 or severely affected by loss of function of this gene. These were: meristem growth, inflorescence structure, and flower development process and its candidate genes (e.g., RCN1, RCN2, and SPL14); cell elongation and cell division process and its candidate genes (e.g., ARR-B, AHK2, and PIN1), reactive oxygen species (ROS) scavenging process and its candidate genes (e.g., CSDs and APXs); and photosynthesis and its candidate genes (Fig. 9c; Supplementary Table S6).
Combining our RNA-seq results with those of previous publications (Nakagawa et al. 2002; Liu et al. 2013; Wang et al. 2015), 12 key differentially expressed genes with high potential for acting as the downstream genes of SiMADS34 were selected and validated by qRT-PCR. As we expected, 10 of the 12 genes showed consistent expression changes in both qRT-PCR and RNA-seq (Fig. 9d), which confirmed the gene expression patterns detected by high-throughput sequencing. Of particular significance is that a previously reported rice MADS34–RCN pathway was confirmed in Setaria, suggesting that our results were reliable.
Discussion
SiMADS34 encodes a MADS-box transcription factor that is important for inflorescence development in S. italica
Inflorescence development is one of the most important agronomic traits that are closely associated with the ultimate grain yield of various crop species. MADS-box transcription factors play important roles in almost every developmental process in plants. To date, a set of MADS-box genes have been studied in Arabidopsis thaliana, Glycine max, Cucumis sativus, Oryza sativa, Populus trichocarpa, Selaginella moellendorffii, and Physcomitrella patens (Parenicová et al. 2003; Leseberg et al. 2006; Arora et al. 2007; Hu and Liu 2012; Barker and Ashton 2013; Shu et al. 2013). Several MADS-box genes have conserved functions in inflorescence development (Wang et al. 2008, 2012; Zhang et al. 2012). In Arabidopsis and rice, the SEP4 gene determines the inflorescence architecture and the related mutant shows increased numbers of branches in the reproductive organs (Ditta et al. 2004; Gao et al. 2010). In the present study, we reported that SiMADS34 is a single-copy gene with a coding sequence length of 741 bp. The SiMADS34 peptide sequence contains 246 amino acid residues, with expected molecular mass of 27.8 kDa and an isoelectric point of 6.9. The phylogenetic analysis indicated that SiMADS34 showed high homology with the Os03g54170.1 (OsMADS34) (Fig. 5c). The foxtail millet simads34 mutant showed longer panicle primary branches, and the number of branches in the panicles was significantly higher than in wild-type Yugu1; these observations were consistent with the results for the mutant osmads34 in rice, as reported by Gao et al. (2010).
Setaria mads34 single mutant displays normal floral organ identity but a severe panicle apical abortion phenotype
We made a detailed investigation of the effect of the simads34 mutation on flower development. We found most flowers developed normally in the simads34 mutant (Fig. 6a–d). However, about 1–2% of florets, especially in the panicle apical region, showed necrosis and were unable to develop into fertile florets (Fig. 6e, f). Our results suggested that the simads34 single mutant did not have an obvious effect on floral organ determination. This result is comparable to previous research on Arabidopsis SEP4 (the orthologous gene of SiMADS34), which showed that the sep4 single mutant exhibited similar flower phenotype to the wild-type in Arabidopsis (Ditta et al. 2004). In rice, the osmads34 mutant showed elongated leaf-like empty glumes (eg) (Gao et al. 2010), while in Setaria we found very little difference in the development of eg between mutant and wild-type plants (Fig. 6). Why does MADS34 work differently in regulating eg development between rice and foxtail millet mutants? The difference might be attributable to the different floret structure between rice and foxtail millet. A rice floret has two egs, both of them very short. In contrast, a foxtail millet floret has four egs, two of them very long (second and third egs in Fig. 6c), while the other two are short (first eg in Fig. 6c, g; the fourth eg is rudimentary). These eg initiation and developmental differences imply that foxtail millet might have different molecular mechanisms controlling the development of egs, compared with those in rice.
Inflorescence apical abortion is a common phenomenon in most cereal crops. In our mutant, apical abortion might result from a burst of reactive oxygen species (ROS), since programmed cell death in apical inflorescences is reported to be one of the main factors resulting in inflorescence abortion (Heng et al. 2018). Moreover, in some other cereal crops, such as rice, the percentage of aborted spikelets in an inflorescence reached 22% on average and the number of grains per mature inflorescence dropped by 20%, compared with spikelets that did not suffer apical abortion. The growth of apical primary branches in our mutant was not greatly affected; in contrast, plant height in rice was affected because of apical abortion (Heng et al. 2018). A previous study by Yamagishi et al. (2004) indicated that inflorescence abortion is a physiological defect that reduces grain yield in rice and other cereal crops. Taken together, our results identified SiMADS34 protein as a key regulator of inflorescence architecture in S. italica.
Comparative analysis and differences between SiMADS34 and its homologs
A comparative analysis between SiMADS34 and its homologs helps us understand gene functions and how these genes regulate inflorescence development across different crops. Phenotypic analysis in our present study indicated that SiMADS34, a single recessive gene regulates inflorescence development via primary branch length and the number of primary branches per inflorescence, inflorescence width and length, and the number of seeds per inflorescence. Analysis of the homologous genes of simads34, such as those in rice, maize, Arabidopsis and cucumber, showed they have similar functions. In rice, a biological role of OsMADS34 was detected in controlling the development of spikelets; OsMADS34 encodes a MADS-box protein containing a short carboxyl terminus that lacked transcriptional activation activity when tested in yeast cells (Gao et al. 2010). OsMADS34 was previously identified as a key regulator of rice inflorescence architecture, and we identified a similar function for simads34 in foxtail millet. Gao et al. (2010) and Kobayashi et al. (2009) reported that the OsMADS34 has an important role in control of spikelet meristem development in rice; the number of primary branches increased, compared with the wild-type. Rice osmads34 mutants had fewer spikelets and shorter primary branches compared with the wild-type, while in our present study the primary branch length of simads34 mutants was higher, and the number of primary branches was lower compared with the wild-type; this inconsistency might be attributable to the different genetic backgrounds. Agrawal et al. (2005) reported similar results for the OsMADS5 gene in rice, which regulates inflorescence architecture via spikelet development.
In Arabidopsis, MADS-box genes participate in many developmental processes, such as meristem specification and inflorescence development (Smaczniak et al. 2012). The function of MADS-box genes in flower development has been clearly demonstrated (Bloomer and Dean 2017; Smaczniak et al. 2012; Yan et al. 2016). Several studies reported that MADS-box genes play essential roles in the flowering processes of maize, such as ZmMADS14 (GRMZM2G099522) and ZmMADS27 (GRMZM2G129034), which both show homology with our gene (Zhao et al. 2010). Further MADS-box genes, ZmMADS1 and ZmMADS3, have roles in regulating spikelet organ primordia during flower development (Heuer 2001).
Putative molecular mechanisms of SiMADS34 in regulating inflorescence development
Combining RNA-seq and qPCR analysis, ten differentially expressed genes were identified as showing high potential for acting as the up- or downstream genes of SiMADS34, including RCNs, PIN1 (auxin pathway), ARR-B (cytokinin pathway), SPLs, CSDs and APX. Both RCN1 and RCN2 were upregulated in the simads34 mutant. Rice RCN is the homolog of A. thaliana CENTRORADIALIS (ATC), which encodes a similar protein to TERMINAL FLOWER1 (TFL1). Overexpression of RCN1 and RCN2 led to a significant increase in secondary branches, which altered panicle morphology in rice (Nakagawa et al. 2002). Liu et al (2013) demonstrated that SEP4/MADS34 can bind to TFL1/RCN and directly suppress its expression. Combining previous reports with our results, we can conclude that loss of function of SiMADS34 led to overaccumulation of RCN1 (Seita.1G180300) and RCN2 (Seita.7G324800) in the Setaria mutant simads34, which thereby caused its highly branching phenotype. Interestingly, we also identified an upstream transcription factor Seita.6G205500 that was upregulated in simads34. Seita.6G205500 is homologous to rice SPL14, which was reported to bind directly to the promoter of MADS34 and positively regulate its expression in rice (Wang et al. 2015). In Setaria, the upregulation of Seita.6G205500 in the mads34 mutant implied there might be feedback regulation between SPL14 and MADS34. In summary, SiMADS34 might control inflorescence architecture mainly through an SPL14–MADS34–RCN regulatory module in Setaria italica.
Panicle apical abortion might result from a burst of reactive oxygen species (ROS) (Heng et al. 2018). In our present study, clear organ death was observed in apical spikelets of the mutant plants (Figs. 3a, b; 6e, f). According to previous research, ROS-triggered programmed cell death is one of the major causes of apical abortion (Heng et al. 2018). Our transcriptome sequencing demonstrated that two biological processes (‘response to ROS’ and ‘cell redox homeostasis’) and one KEGG pathway (‘peroxisome’) related to ROS scavenging were enriched in the simads34 mutant. Seventy-five ROS-inducible genes were differentially expressed, including some key ROS-scavenging enzymes such as copper/zinc superoxide dismutase (CSD1 and CSD3) and ascorbate peroxidase (APX1) (Fig. 9). These results implied that excessive ROS might arise in the simads34 mutant and lead to oxidative damage to the apical spikelets. More experiments are needed to elucidate how SiMADS34 regulates these potential molecular pathways.
Conclusion
We have identified a novel MADS-box transcription factor, SiMADS34, responsible for regulating inflorescence architecture in foxtail millet, Setaria italica. The potential of MADS-box genes in regulating yield component traits and floral organ development in this millet crop is well documented. Setaria is useful as an excellent genetic model system for studying grass functional genomics and comparative mapping. Our results help to fill a large knowledge gap regarding the biological processes that determine inflorescence architecture in foxtail millet. Furthermore, our results shed light on a crucial process, and consequently should enable further functional characterization. This novel information on the phenotypic effects and the underlying molecular mechanism of the SiMADS34 transcription factor should prove valuable in breeding for enhanced crop yield in foxtail millet.
References
Agrawal G, Abe K, Yamazaki M, Miyao A, Hirochika H (2005) Conservation of the E-function for floral organ identity in rice revealed by the analysis of tissue culture-induced loss-of-function mutants of the OsMADS1 gene. Plant Mol Biol 59(1):125–135
Arora R, Agarwal P, Ray S, Singh A, Singh VP, Tyagi A, Kapoor S (2007) MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress. BMC Genomics 8(1):242
Barker E, Ashton N (2013) A parsimonious model of lineage-specific expansion of MADS-box genes in Physcomitrella patens. Plant Cell Rep 32(8):1161–1177
Bassam B, Caetano-Anollés G, Gresshoff P (1991) Fast and sensitive silver staining of DNA in polyacrylamide gels. Anal Biochem 196(1):80–83
Bloomer R, Dean C (2017) Fine-tuning timing: natural variation informs the mechanistic basis of the switch to flowering in Arabidopsis thaliana. J Exp Bot 68(20):5439–5452
Der G, Everitt B (2008) A handbook of statistical analyses using SAS. Chapman and Hall, New York
Diao X, Jia G (2017) Foxtail millet germplasm and inheritance of morphological characteristics. In: Doust A, Diao X (eds) Genetics and genomics of setaria, plant genetics and genomics: crops and models, vol 19. Springer, Cham, pp 73–92
Diao X (2011) Current status of foxtail millet production in China and future development directions. The industrial production and development system of foxtail millet in China. pp 20–30
Ditta G, Pinyopich A, Robles P, Pelaz S, Yanofsky M (2004) The SEP4 gene of Arabidopsis thaliana functions in floral organ and meristem identity. Curr Biol 14(21):1935–1940
Doust A, Devos K, Gadberry M, Gale M, Kellogg E (2005) The genetic basis for inflorescence variation between foxtail and green millet (Poaceae). Genetics 169(3):1659–1672
Fedorov A (1974) Chromosome numbers of flowering plants. Otto Koeltz Science Publishers, Koenigstein
Gao X, Liang W, Yin C, Ji S, Wang H, Su X, Zhang D (2010) The SEPALLATA-like gene OsMADS34 is required for rice inflorescence and spikelet development. Plant Physiol 153(2):728–740
Goodstein D, Shu S, Howson R, Neupane R, Hayes R, Fazo J et al (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40(D1):D1178–D1186
Gramzow L, Ritz M, Theißen G (2010) On the origin of MADS-domain transcription factors. Trends Genet 26(4):149–153
Heng Y, Wu C, Long Y, Luo S, Ma J, Chen J et al (2018) OsALMT7 maintains panicle size and grain yield in rice by mediating malate transport. Plant Cell 30(4):889–906
Heuer S (2001) The Maize MADS Box Gene ZmMADS3 Affects node number and spikelet development and is co-expressed with ZmMADS1 during flower development, in egg cells, and early embryogenesis. Plant Physiol 127(1):33–45
Honma T, Goto K (2001) Complexes of MADS-box proteins are sufficient to convert leaves into floral organs. Natur 409(6819):525
Hu L, Liu S (2012) Genome-wide analysis of the MADS-box gene family in cucumber. Genome 55(3):245–256
Huang P, Feldman M, Schroder S, Bahri B, Diao X, Zhi H, Kellogg E et al (2014) Population genetics of Setaria viridis, a new model system. Mol Ecol 23(20):4912–4925
Jack T (2004) Molecular and genetic mechanisms of floral control. Plant Cell 16(suppl 1):S1–S17
Jia G, Shi S, Wang C, Niu Z, Chai Y, Zhi H, Diao X (2013) Molecular diversity and population structure of Chinese green foxtail [Setaria viridi1s (L.) Beauv.] revealed by microsatellite analysis. J Exp Bot 64(12):3645–3656
Jia G, Huang X, Zhi H, Zhao Y, Zhao Q, Li W et al (2013) A haplotype map of genomic variations and genome-wide association studies of agronomic traits in foxtail millet (Setaria italica). Nat Genet 45(8):957
Kobayashi K, Maekawa M, Miyao A, Hirochika H, Kyozuka J (2009) PANICLE PHYTOMER2 (PAP2), encoding a SEPALLATA subfamily MADS-box protein, positively controls spikelet meristem identity in rice. Plant Cell Physiol 51(1):47–57
Leseberg C, Li A, Kang H, Duvall M, Mao L (2006) Genome-wide analysis of the MADS-box gene family in Populus trichocarpa. Gene 378:84–94
Li W, Meng C, Liu T (1935) Problems in the breeding of millet (Setaria italica (L.) Beuav.). J Am Soc Agron 27:963–970
Liu C, Teo Z, Bi Y, Song S, Xi W, Yang X et al (2013) A conserved genetic pathway determines inflorescence architecture in Arabidopsis and rice. Dev Cell 24(6):612–622
Liu W (1984) Estimation of the genetic parameters for the main characters of millet and their application in breeding. J Shanxi Agric Univ 4:173–181
Lu H, Zhang J, Liu K, Wu N, Li Y, Zhou K et al (2009) Earliest domestication of common millet (Panicum miliaceum) in East Asia extended to 10,000 years ago. Proc Natl Acad Sci USA 106(18):7367–7372
Ma H, Yanofsky M, Meyerowitz E (1991) AGL1-AGL6, an Arabidopsis gene family with similarity to floral homeotic and transcription factor genes. Genes Dev 5:484–495
Murray M, Thompson W (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8(19):4321–4326
Nakata M, Miyashita T, Kimura R, Nakata Y, Yamakawa H (2018) Mutmapplus identified novel mutant alleles of a rice starch branching enzyme iib gene for fine-tuning of cooked rice texture. Plant Biotechnol J 16(1):111–123
Nakagawa M, Shimamoto K, Kyozuka J (2002) Overexpression of RCN1 and RCN2, rice TERMINAL FLOWER 1/CENTRORADIALIS homologs, confers delay of phase transition and altered panicle morphology in rice. Plant J 29(6):743–750
Norman C, Runswick M, Pollock R, Treisman R (1988) Isolation and properties of cDNA clones encoding SRF, a transcription factor that binds to the c-fos serum response element. Cell 55(6):989–1003
Parenicová L, De Folter S, Kieffer M, Horner D, Favalli C, Busscher J et al (2003) Molecular and phylogenetic analyses of the complete MADS-box transcription factor family in Arabidopsis: new openings to the MADS world. Plant Cell 15(7):1538–1551
Passmore S, Maine G, Elble R, Christ C, Tye B (1988) Saccharomyces cerevisiae protein involved in plasmid maintenance is necessary for mating of MAT$α$ cells. J Mol Biol 204(3):593–606
Riechmann J, Meyerowitz E (1997) MADS domain proteins in plant development. Biol Chem 378(10):1079–1102
Soyk S, Lemmon Z, Oved M, Fisher J, Liberatore K, Park S et al (2017) Bypassing negative epistasis on yield in tomato imposed by a domestication gene. Cell 169(6):1142-1155.e12
Shu Y, Yu D, Wang D, Guo D, Guo C (2013) Genome-wide survey and expression analysis of the MADS-box gene family in soybean. Mol Biol Rep 40(6):3901–3911
Smaczniak C, Immink R, Angenent G, Kaufmann K (2012) Developmental and evolutionary diversity of plant MADS-domain factors: insights from recent studies. Development 139(17):3081–3098
Sommer H, Beltran J, Huijser P, Pape H, Lönnig W, Saedler H, Schwarz-Sommer Z (1990) Deficiens, a homeotic gene involved in the control of flower morphogenesis in Antirrhinum majus: the protein shows homology to transcription factors. EMBO J 9(3):605–613
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Theissen G (2001) Development of floral organ identity: stories from the MADS house. Curr Opin Plant Biol 4:75–85
Tang S, Li L, Wang Y, Chen Q, Zhang W, Jia G et al (2017) Genotype-specific physiological and transcriptomic responses to drought stress in Setaria italica (an emerging model for Panicoideae grasses). Sci Rep 7(1):1–15
Theißen G, Melzer R, Rümpler F (2016) MADS-domain transcription factors and the floral quartet model of flower development: linking plant development and evolution. Development 143(18):3259–3271
Wang E, Wang J, Zhu X, Hao W, Wang L, Li Q et al (2008) Control of rice grain-filling and yield by a gene with a potential signature of domestication. Nat Genet 40(11):1370
Wang K, Tang D, Hong L, Xu W, Huang J, Li M et al (2010) DEP and AFO regulate reproductive habit in rice. PLoS Genet 6(1):e1000818
Wang L, Sun S, Jin J, Fu D, Yang X, Weng X, Zhang Q (2015) Coordinated regulation of vegetative and reproductive branching in rice. Proc Natl Acad Sci USA 112(50):15504–15509
Wang S, Wu K, Yuan Q, Liu X, Liu Z, Lin X et al (2012) Control of grain size, shape and quality by OsSPL16 in rice. Nat Genet 44(8):950
West A, Shore P, Sharrocks A (1997) DNA binding by MADS-box transcription factors: a molecular mechanism for differential DNA bending. Mol Cell Biol 17(5):2876–2887
Yamagishi J, Miyamoto N, Hirotsu S, Laza R, Nemoto K (2004) QTLs for branching, floret formation, and pre-flowering floret abortion of rice panicle in a temperate japonica × tropical japonica cross. Theoret Appl Genet 109(8):1555–1561
Yan W, Chen D, Kaufmann K (2016) Molecular mechanisms of floral organ specification by MADS domain proteins. Curr Opin Plant Biol 29:154–162
Yang X, Wan Z, Perry L, Lu H, Wang Q, Zhao C et al (2012) Early millet use in northern China. Proc Natl Acad Sci USA 109(10):3726–3730
Yanofsky M, Ma H, Bowman J, Drews G, Feldmann K, Meyerowitz E (1990) The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors. Nature 346(6279):35
Zhang X, Wang J, Huang J, Lan H, Wang C, Yin C et al (2012) Rare allele of OsPPKL1 associated with grain length causes extra-large grain and a significant yield increase in rice. Proc Natl Acad Sci USA 109(52):21534–21539
Zhang Y, Zhang B, Yan D, Dong W, Yang W, Li Q et al (2011) Two Arabidopsis cytochrome P450 monooxygenases, CYP714A1 and CYP714A2, function redundantly in plant development through gibberellin deactivation. Plant J 67(2):342–353
Zhao Y, Li X, Chen W, Peng X, Cheng X, Zhu S, Cheng B (2010) Whole-genome survey and characterization of MADS-box gene family in maize and sorghum. Plant Cell Tissue Organ Cult 105(2):159–173
Acknowledgements
This work was supported by National Key R&D Program of China (Grant Nos. 2019YFD1000700 and 2019YFD1000704), the National Natural Science Foundation of China (31871692), Fundamental Research Funds of CAAS (S2018PY03 to Sha Tang), the China Agricultural Research System (CARS06-13.5-A04), and the Agricultural Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences. We thank Huw Tyson, PhD in Plant Biochemistry, graduated from University of Cambridge, for editing the English text of a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
SHH, HW, HZ performed the molecular and field experiments; SHH, ST, CT, WZ extracted the data; SHH, ST analyzed the data; SHH, ST wrote the manuscript; SHH, ST, GJ were responsible for research methodology; ST, XD, HZ for review and editing; XD, ST for conceptualization, funding acquisition, supervision, project administration.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hussin, S.H., Wang, H., Tang, S. et al. SiMADS34, an E-class MADS-box transcription factor, regulates inflorescence architecture and grain yield in Setaria italica. Plant Mol Biol 105, 419–434 (2021). https://doi.org/10.1007/s11103-020-01097-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-020-01097-6