Abstract
Plants are constantly exposed to a variety of biotic and abiotic stresses, which include pathogens and conditions of high salinity, low temperature, and drought. Abscisic acid (ABA) is a major plant hormone involved in signal transduction pathways that mediate the defense response of plants to abiotic stress. Previously, we isolated Ring finger protein gene (CaRING1) from pepper (Capsicum annuum), which is associated with resistance to bacterial pathogens, accompanied by hypersensitive cell death. Here, we report a new function of the CaRING1 gene product in the ABA-mediated defense responses of plants to dehydration stress. The expression of the CaRING1 gene was induced in pepper leaves treated with ABA or exposed to dehydration or NaCl. Virus-induced gene silencing of CaRING1 in pepper plants exhibited low degree of ABA-induced stomatal closure and high levels of transpirational water loss in dehydrated leaves. These led to be more vulnerable to dehydration stress in CaRING1-silenced pepper than in the control pepper, accompanied by reduction of ABA-regulated gene expression and low accumulation of ABA and H2O2. In contrast, CaRING1-overexpressing transgenic plants showed enhanced sensitivity to ABA during the seedling growth and establishment. These plants were also more tolerant to dehydration stress than the wild-type plants because of high ABA accumulation, enhanced stomatal closure and increased expression of stress-responsive genes. Together, these results suggest that the CaRING1 acts as positive factor for dehydration tolerance in Arabidopsis by modulating ABA biosynthesis and ABA-mediated stomatal closing and gene expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Abiotic and biotic stresses, such as pathogenic infections and drought, adversely influence plant growth and crop production. Under these stress conditions, plants activate several defense responses by altering physiological and molecular systems via the synthesis, remodeling, and degradation of proteins (Stone and Callis 2007; Huang et al. 2014; Lim et al. 2014). The ubiquitin-26S proteasome system is important for post-translational modification and regulates crucial eukaryotic cellular processes, including DNA repair, cell signaling, and defense responses (Lee et al. 2011; Sadanandom et al. 2012; Callis 2014; Seo et al. 2014). This mechanism has a diverse range of substrates, including transcription factors, hormone receptors, cell cycle regulators, light regulators, and misfolded proteins (Hershko and Ciechanover 1998; Gagne et al. 2004; Zhang et al. 2007; Irigoyen et al. 2014; Seo et al. 2014). Ubiquitin is bound to target proteins by a series of well-characterized reactions that result in the target proteins being marked for degradation (Pickart 2001; Glickman and Adir 2004; Smalle and Vierstra 2004).
Ubiquitin is activated by an E1 ubiquitin-activating enzyme in an ATP-dependent manner and is then transferred to an E2 ubiquitin-conjugating enzyme. Finally, the E2-ubiquitin complex interacts with an E3 ubiquitin-ligase that catalyzes the formation of a bond between the target protein and ubiquitin, leading to autoubiquitination of E3 ligase and ubiquitination of the target protein (Ciechanover 1998; Ciechanover and Schwartz 1998; Jacobson et al. 2009). In vitro ubiquitination analysis have suggested that the combined activities of these three enzymes are necessary and sufficient for autoubiquitination and ubiquitination (Callis 2014). In this system, E3 ubiquitin-ligase has multiple isoforms, indicating that this enzyme selects target proteins for ubiquitination (Callis 2014). The E3 ubiquitin ligases are divided into two groups based on their structure (Pickart 2001). One group functions as a single subunit, including RING (Really Interesting New Gene), U-box, and HECT (Homology to E6-AP Carboxyl Terminus) E3 ligases (Hatakeyama et al. 2001; Miao and Zentgraf 2010; Kim and Kim 2013a, b; Marin 2013). The other group functions as a multisubunit, including APC (Anaphase Promoting Complex), CUL4-DDB1 (CULLIN4-Damaged-specific DNA binding protein 1), and SCF (Skp1, Cullin, F-box) (Zheng et al. 2002; Pazhouhandeh et al. 2011; Chang et al. 2014). Among these, RING type E3 ubiquitin ligases constitute the third largest gene family in Arabidopsis with more than 477 genes (Stone et al. 2005); however, the function of only a few RING type E3 ligases has been elucidated. In plants, several RING type ligases are involved in cellular processes, including development, hormonal signaling, and defense responses to biotic and abiotic stresses (Xie et al. 2002; Stone et al. 2006; Bu et al. 2009; Lee et al. 2011; Kim and Kim 2013a, b).
In particular, many studies have shown that RING type E3 ligases play a crucial role in defense mechanisms and that they are specifically induced by pathogen infection as well as environmental stresses (Zeng et al. 2006; Ryu et al. 2010; Lee et al. 2011). For instance, RING type E3 ligases, such as ACRE132 and ACRE189, act as positive regulators of disease resistance. On the contrary, CaRFP1 is associated with disease susceptibility and abiotic stress tolerance (Hong et al. 2007). Several RING type E3 ligases, which are associated with stress tolerance/susceptibility, are involved in the degradation of ABA signal transduction components. ABA is one of the major plant hormones that play an important role in seed dormancy, plant development, and adaptation to abiotic stress (Bartels and Sunkar 2005; Finkelstein et al. 2002; Leung and Giraudat 1998; Rock 2000). The expression of a number of genes involved in plant defense responses to abiotic stress is regulated by ABA (Jakab et al. 2005). The ABA-induced C3H2C3-type RING E3 ligase AtAIRP1 functions as a positive regulator of drought stress in an ABA dependent manner (Ryu et al. 2010). Moreover, ABI5 binding protein (AFP) acts as a negative regulator of ABA by ubiquitinating and degrading the ABI5 transcription factor (Lopez-Molina et al. 2003). These studies suggest that various RING type ligases are both negative and positive regulators of ABA signaling, which affect tolerance to abiotic stress.
Previously, we reported that the CaRING1 (Ca psicum a nnuum RING finger protein 1) gene is differentially expressed in pepper leaves that are infected by virulent or avirulent strains of Xanthomonas campestris pv. vesicatoria (Xcv) (Lee et al. 2011). CaRING1 contains an amino-terminal transmembrane domain and a c-terminal RING domain, which is essential for E3 ubiquitin ligase activity. CaRING1 is involved in cell death, which is accompanied by changes in reactive oxygen species (ROS) and salicylic acid (SA) accumulation, and PR gene expression. CaRING1-silenced pepper plants display enhanced susceptibility to an avirulent strain of Xcv, while CaRING1-overexpressing (OX) Arabidopsis plants are resistant to Pseudomonas syringae pv. tomato (Pst) DC3000 and Hyaloperonospora arabidopsidis infections. In this study, based on the expression profiles of the CaRING1 gene in pepper treated with ABA, dehydration, NaCl, and H2O2, we evaluated their responses to ABA and dehydration by using CaRING1-silenced pepper and overexpressing transgenic Arabidopsis plants. Our data suggest that CaRING1 protein is involved in regulation of ABA biosynthesis and ABA signaling, leading enhanced dehydration tolerance.
Materials and methods
Plant materials
Pepper (Capsicum annuum L., cv. Hanbyul) seeds were sown in a steam-sterilized compost soil mix (peat moss, perlite, and vermiculite, 5:3:2, v/v/v), sand, and loam soil (1:1:1, v/v/v). The pepper plants were raised in a growth room at 27 ± 1 °C with 80 μmol photons m−2 s−1 (white fluorescent light) for 16 h per day as described previously (Lee et al. 2008). Seeds of 35S:CaRING1 Arabidopsis plants were obtained from individual stable transformants (line #13 and #16) that were used in a previous study (Lee et al. 2011). 35S:CaRING1 transgenic mutants and wild-type plants (ecotype Col-0) were routinely grown in a 9:1:1 ratio of peat moss, perlite, and vermiculite under fluorescent light (130 μmol photons m−2 s−1) at 24 °C with 60 % humidity and a 16-h light/8-h dark cycle. Arabidopsis seeds were surface sterilized with 70 % ethanol for 1 min for in vitro culture. After treatment with 2 % sodium hydroxide for 10 min, seeds were washed 10 times in sterile distilled water and were finally sown on Murashige and Skoog (1962) (MS) agar (Sigma, St. Louis, MO, USA) supplemented with 1 % sucrose. The plates were sealed and incubated at 24 °C in a chamber exposed to a 16-h light/8-h dark cycle.
Virus-induced gene silencing (VIGS) of the CaRING1 gene in pepper
To knockdown the CaRING1 gene in pepper plants, VIGS was performed as described by Lee et al. (2011). Briefly, Agrobacterium tumefaciens strain GV3101 carrying pTRV1 and pTRV2:CaRING1 or pTRV2:00 as a negative control was co-infiltrated into the fully expanded cotyledons of pepper plants (OD600 = 0.2 for each construct). Plants were placed in a growth room at 24 °C with a 16 h light and 8 h dark photoperiod for growth and spread of the virus.
ABA, H2O2, NaCl and dehydration treatment
At the six-leaf-stage, pepper plants were used to examine expression pattern of CaRING1 gene. Pepper plants were sprayed with 100 μM ABA and 100 μM H2O2. For treatment with salt stress, plants were irrigated with a salt solution (200 mM). For dehydration stress, plants were carefully removed from the soil to prevent injury and then dried on 3 MM paper (Whatman, Clifton, UK) or only aerial parts of pepper plants were dried after removing their roots.
For the seedling growth test of Arabidopsis plant, 100 seeds of wild-type and 35S:CaRING1 transgenic Arabidopsis lines were sown on plates containing MS agar medium supplemented with various concentrations of ABA, and seedlings with green cotyledons were counted 7 days later. In parallel, seedlings from each line were vertically grown on the MS plates for 7 days and the root lengths of seedlings were measured. For qRT-PCR analysis, 4-week-old 35S:CaRING1 mutants and wild-type plants were treated with 50 μM ABA, or were carefully removed from the soil to be subjected to dehydration stress, and harvested at the given time points after treatment.
Dehydration tolerance assays
Dehydration tolerance assays were carried out as described by Lim and Lee (2014). One-week-old seedlings from the wild-type and 35S:CaRING1 lines were randomly planted in a pot containing soil mix (peat moss, perlite, and vermiculite, 9:1:1) and were grown under normal watering conditions for 2 weeks. To impose dehydration stress, watering was withheld for 2 weeks and the survival rate of the plants with rehydrated leaves was calculated after rewatering for 3 days. Rates of water loss were measured to determine the dehydration tolerance of 35S:CaRING1 mutant plants in a quantitative manner. Ten leaves were detached from 4-week-old plants of each line and placed in petri dishes. The dishes were kept in a growth chamber with 40 % relative humidity, and loss of fresh weight was determined at the indicated times. The experiments were repeated three times.
Stomatal aperture bioassay
A stomatal aperture bioassay was carried out as described previously with the following modifications (Lee et al. 2013). Briefly, leaf peels were harvested from the rosette leaves of 4-week-old plants and floated in stomatal opening solution (SOS: 50 mM KCl and 10 mM MES-KOH, pH 6.15, 10 mM CaCl2) in the light for 2.5 h. To induce stomatal closing, the buffer was replaced with fresh SOS containing various concentrations of ABA and the leaf peels were further incubated for 2.5 h. In each sample, 100 stomata were randomly observed under a Nikon eclipse 80i microscope, and the width and length of individual stomata were recorded using Image J 1.46r (http://imagej.nih.gov/ij). Each experiment was performed three times independently.
RNA isolation and quantitative reverse transcription-polymerase chain reaction (qRT-PCR)
Total RNA was isolated from leaves of pepper and Arabidopsis treated with ABA or subjected to dehydration stress using an RNeasy Mini kit (Qiagen, Valencia, CA, USA), following the manufacturer’s instructions. All RNA samples were digested with RNA-free DNase to remove genomic DNA. After quantification using a spectrophotometer, 1 μg of total RNA was used as a template to synthesize cDNA using a Transcript First Strand cDNA Synthesis kit (Roche, Indianapolis, IN, USA), according to the manufacturer’s instructions. In parallel, PCR was performed without reverse transcriptase and the products were subjected to qRT-PCR to confirm the absence of genomic DNA contamination in the cDNA samples. The synthesized cDNA was amplified in a CFX96 Touch™ Real-Time PCR detection system (Bio-Rad, Hercules, CA, USA) with iQ™ SYBR Green Supermix and specific primers (Supplemental table S1). Every reaction was performed in triplicate. The PCR was programmed as follows: 95 °C for 5 min, 45 cycles at 95 °C for 20 s, 60 °C for 20 s, and 72 °C for 20 s. The relative expression of each gene was calculated using the ∆∆Ct method, as previously described (Livak and Schmittgen 2001). The Arabidopsis Actin8 gene (AtACT8) and pepper Actin1 (CaACT1) was used for normalization.
DAB staining
Staining with 3,3′-diaminobenzidine (DAB) was carried out to visualize H2O2 in pepper leaves treated with dehydration stress, according to the method described by Lee et al. (2011). Briefly, leaf samples were collected and submerged into 1 mg ml−1 DAB (Sigma) solution (pH 3.8). After incubation for 16 h, the leaf samples were boiled in 100 % ethanol for 10 min to remove the chlorophyll and were then photographed.
Measurement of ABA content
For determination of ABA content, leaves were harvested from pepper and Arabidopsis plants treated with dehydration for 2 h and immediately frozen in liquid nitrogen. Approximately 50 mg of ground tissue were extracted overnight in 1 ml of ABA extraction buffer (methanol, containing 100 mg l−1 butylated hydroxyl toluene, 0.5 g l−1 citric acid monohydrate) at 4 °C on a rotary shaker. After centrifuged at 1500g, the supernatant was transferred to new tube and dried using a speed vac. ABA content of each sample was quantified by using the Phytodetek-ABA kit (Agdia Inc., Elkhart, IN, USA) according to manufacturer’s instruction. ABA contents were expressed as pmol mg−1 fresh weight of the tissue.
Results and discussion
Induction of the CaRING1 in pepper leaves by ABA, H2O2, dehydration, and high salinity
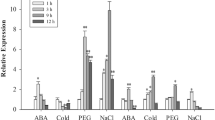
Many of RING type E3 ligases exhibits specific induction in response to both pathogen attacks and environmental stresses (Zeng et al. 2006; Ryu et al. 2010; Lee et al. 2011). In this study, we investigated involvement of CaRING1 in abiotic stress. Expression pattern of CaRING1 was examined in pepper leaves treated with ABA, a phytohormone that acts as a core regulator in abiotic stress responses (Fig. 1a). CaRING1 transcripts were first detected 2 h after ABA treatment, with maximal levels detected at 12 h. ABA induces H2O2 production in guard cells of Arabidopsis (Pei et al. 2000) and Vicia faba (Zhang et al. 2007). H2O2 levels are also enhanced by ABA in maize embryos and seedlings (Jiang and Zhang 2001). Based on these, we examined whether H2O2 promotes expression of CaRING1 gene. Following treatment with 100 μM of H2O2, expression level of CaRING1 gene gradually increased up to three times in pepper leaves (Fig. 1b). The accumulation of ABA is indispensable for plant defense responses to abiotic stresses, it is synthesized in various tissues, and its levels are increased in plant tissues, particularly the leaves, in response to osmotic stress (Cutler et al. 2010; Hubbard et al. 2010). To ascertain the effect of osmotic stress on the expression of the CaRING1 gene, pepper plants were subjected to dehydration and high salinity treatments (Fig. 1c, d). As shown in Fig. 1c, dehydration stress rapidly induced CaRING1 transcription after 2-h exposure and rapidly decreased expression between 6 and 24 h after treatment. Moreover, in pepper leaves treated with 200 mM NaCl, CaRING1 transcripts started to accumulate 2 h after treatment and reached peak levels at 12 h (Fig. 1d). These findings indicate that CaRING1 is induced in plants in response to ABA, dehydration, high salinity and H2O2. In Arabidopsis genome, there are four genes, At1g20823, At1g76410, At2g17450, and At4g35480, sharing high sequence similarity with CaRING1 (Lee et al. 2011). These genes also exhibit specific expression under treatment with ABA, drought, osmotic, cold, and high salinity (Supplemental figs. S1 and S2). In particular, ABA treatment leads to high induction of those genes except At1g76410 in leaves and guard cell. Similarly, the three genes At1g20823, At2g17450, and At4g35480 are significantly induced at the early time point under drought condition. Of the four genes, At1g20823 show specific induction in response to all treatments and was chosen for further study. These data provide the possibility that they have similar function in response to those treatments.
Expression pattern of the CaRING1 genes in pepper leaves treated with abscisic acid (100 μM; a), H2O2 (100 μM; b), dehydration (c), or NaCl (200 mM; d). Relative expression (∆∆CT) of the CaRING1 gene was normalized to that of that of CaACT1 as internal control gene and compared with the value for non-treated leaves. Data are the mean ± standard error from three independent experiments
Involvement of CaRING1 in regulation of ABA- and dehydration-induced stomatal closure.
To analyze the biological role of the CaRING1 gene in response to ABA treatment and dehydration stress, we applied the VIGS technique to induce the knockdown gene expression of CaRING1 in pepper. qRT-PCR analysis was performed to test the efficiency of VIGS using empty vector control (TRV:00) and CaRING1-silenced (TRV:CaRING1) pepper leaves harvested before and after dehydration treatment for 2 h (Fig. 2a). Under non-treatment, expression of CaRING1 gene in TRV:CaRING1 leaves was hardly detected (>7.4-fold decrease) compared with TRV:00 leaves. Dehydration stress triggered strong induction of CaRING1 gene in both plant leaves: 3.6 to 6.1-fold in TRV:00 leaves and 3.0 to 6.1-fold in TRV:CaRING1 leaves compared with that of the non-treated plant leaves. However, the expression level was still lower (>6.6-fold) in TRV:CaRING1 leaves than that of TRV:00 leaves. Next, 4-week-old plants of TRV:00 and TRV:CaRING1 pepper grown under normal condition were subjected to dehydration stress for 10 days. We measured leaf surface temperature using an Infrared Thermal Imaging Camera (T420, FLIR system, USA). Leaf temperature can be an indirect indicator for stomatal aperture and transpirational rate, which are controlled by ABA, because stomatal closing induces decreased evaporative cooling and finally increase leaf temperature (Park et al. 2015). As shown in Fig. 2b, CaRING1-silenced pepper plants exhibited low leaf temperature relative to that of the control plants. Previous studies have used measurements of stomatal movement to establish that high sensitivity to ABA in leaves leads to increased dehydration tolerance (Cheong et al. 2007; Lim and Lee 2014). In the absence of ABA, TRV:CaRING1 plants had larger stomatal apertures compared with TRV:00 plants (Fig. 2c, d). This significant difference was still observed after treatment with ABA, indicating silencing of CaRING1 allowed partial inhibition of ABA-induced stomatal closing. Consistently, rare of transpirational water loss was higher in TRV:CaRING1 plants than TRV:00 (Fig. 2e). These data suggests that CaRING1 acts as a positive regulator in ABA and dehydration-induced stomatal closing.
Reduced stomatal closing in leaves of CaRING1-silenced pepper plants. a RT-PCR analysis of CaRING1 expression in leaves of pepper plants transfected with the empty vector (TRV:00) as control or the silencing constructs (TRV:CaRING1) 2 h after dehydration. Relative expression level (∆∆CT) of CaRING1 gene was normalized to that of CaACT1 as internal control gene. Four-week-old plants of TRV:00 and TRV:CaRING1 peppers were subjected to dehydration stress by removing their roots. b Representative thermographic image of CaRING1-silenced pepper plants 10 days after dehydration treatment and mean leaf temperature was measured from the four upper leaves. c, d Stomatal apertures in control and CaRING1-silenced pepper plants treated with ABA. Stomatal apertures were measured under the microscope after leaf peels harvested from the 4-week-old plants of each line were incubated for 2 h in SOS buffer containing 20 μM ABA (d) and representative images were taken. Data are the mean ± standard error from three independent experiments. e Water loss from leaves of TRV:00 and TRV:CaRING1 pepper plants at various times after detachment of leaves. Data are the mean ± standard error from three independent experiments. Asterisks indicate significant differences in three independent experiments (Student’s t test; P < 0.05)
Reduced tolerance of CaRING1-silenced pepper plants to dehydration stress
Based on the data shown in Fig. 2, we postulated that CaRING1-silenced pepper plants are more vulnerable to dehydration stress than the control plants. To prove this hypothesis, 4-week-old plants of TRV:CaRING1 and TRV:00 were subjected to dehydration stress by withholding watering for 14 days (Fig. 3a). There were no phenotypic differences between the two plant lines under well-watered conditions. However, dehydration stress treatment made TRV:CaRING1 plants rapidly dried relative to TRV:00 plants. Upon re-watering, over 73 % of the TRV:CaRING1 plants did not resume growth, while TRV:00 plants exhibited more survival rates approximately 83 %. These data indicated that silencing of CaRING1 gene confer reduced dehydration tolerance.
Reduced tolerance of CaRING1-silenced pepper plants to dehydration stress. a Dehydration sensitivity of CaRING1-silenced pepper plants. Four-week-old CaRING1-silenced pepper plants (TRV:CaRING1) and control plants (TRV:00) were subjected to dehydration stress by withholding water for 14 days. Representative images were taken before (right) and after (middle) dehydration stress and at 3 days after rewatering (right). Survival rate was measured by counting plants that have green and rehydrated leaves 3 days after rewatering. b RT-PCR analysis of ABA-responsive gene and ABA biosynthesis-related gene expression in leaves of TRV:00 and TRV:CaRING1 pepper plants after treatment with dehydration for 2 h. Relative expression level (∆∆CT) of each gene was normalized to that of CaACT1 as internal control gene. Four-week-old pepper plants of TRV:00 and TRV:CaRING1 were subjected to dehydration stress by drying aerial part of the plants after removing their roots. c ABA content in leaves of TRV:00 and TRV:CaRING1 pepper plants after dehydration treatment for 2 h. Data are the means ± standard error from three independent experiments. d Hydrogen peroxide production in the leaves of TRV:00 and TRV:CaRING1 pepper plants in response to dehydration. Four-week-old pepper plants were treated with dehydration as mentioned above. Leaves harvested 3 h after treatment were stained with DAB solution and representative images were taken
Drought sensitivity is correlated with level of ABA and drought-related gene expression (Gonzalez-Guzman et al. 2012; Li et al. 2011; Ryu et al. 2010). The primary function of ABA is the regulation of tolerance to abiotic stress, and ABA signal transduction is associated with defense response to abiotic stress (Zhu 2002). In particular, ABA contributes to plant adaptation to water-deprivation through regulatory circuits that induce gene expression (Wasilewska et al. 2008; Lee and Luan 2012). Based on these, we performed qRT-PCR analysis to examine expression levels of ABA- and drought-responsive gene CaRD29B and ABA biosynthesis-related gene CaNCED3 in the leaves of TRV:00 and TRV:CaRING1 plants treated with dehydration for 2 h (Fig. 3b; Lim et al. 2015). Before treatment, TRV:CaRING1 plants exhibited low expression of CaRD29B (>2.5-fold decrease) and CaNCED3 (>2.1-fold decrease) genes, compared with TRV:00 plants. Dehydration stress treatment triggered significant induction of the two genes in the two plant lines: CaRD29B, 11.7 to 14.5-fold in TRV:CaRING1 and 13.2 to 15.4-fold in TRV:00; CaNCED3, 1.9 to 3.4-fold in TRV:CaRING1 and 1.5 to 4.2-fold in TRV:00. However, their expression levels were still lower in TRV:CaRING1 than in TRV:00, consistent with reduced tolerance of TRV:CaRING1 plants to dehydration. Especially, CaNCED3 gene induction by ABA was not surprised because some of ABA biosynthesis-related genes have been induced by endogenous and exogenous ABA (Barrero et al. 2006; Cheng et al. 2002; Xiong et al. 2002). The differential expression of CaNCED3 gene between TRV:CaRING1 and TRV:00 plants raised the possibility that CaRING1 functions in regulation of ABA biosynthesis. To test this possibility, we measured ABA level in the leaves of the two plant lines harvested at the same time as qRT-PCR analysis (Fig. 3c). Consistent with the expression level of CaNCED3 gene, TRV:CaRING1 plants exhibited low ABA level, compared with TRV:00 plants, and even this pattern maintained after dehydration stress which elevated ABA biosynthesis.
Water stress induces production of H2O2 as well as ABA in plant cell and ABA is also essential for H2O2 production (Hu et al. 2006). The association of ABA signaling and H2O2 production is observed during ABA-induced stomatal closure in Arabidopsis guard cell (Pei et al. 2000; Kwak et al. 2003). As the second messenger in ABA signaling pathway, H2O2 is downstream of ABA-INSENSITIVE1 (ABI1), one of the protein phosphates acting as negative regulator in ABA signaling, and upstream of ABI2 (Umezawa et al. 2009; Murata et al. 2001). As shown in Fig. 1b, H2O2 treatment triggered gradual accumulation of CaRING1 transcripts. Based on these, we measured H2O2 production in the leaves of TRV:CaRING1 and TRV:00 plants treated with dehydration stress through DAB staining (Fig. 3d). Dehydration stress led to significant accumulation of H2O2 in the both plant line leaves, but its amount were greater in TRV:CaRING1 plant leaves than in TRV:00 plant leaves. Taken together, these data suggest that silencing of CaRING1 reduced dehydration tolerance via downregulation of ABA biosynthesis, ABA-response gene expression, and H2O2 production.
Enhanced ABA sensitivity of 35S:CaRING1 Arabidopsis transgenic plants
We previously generated Arabidopsis transgenic plants constitutively expressing CaRING1 under the control of the strong constitutive 35S promoter (35S: CaRING1; Lee et al. 2011). Of the 35S: CaRING1 plant lines used in the previous study, two lines (#13 and #16) expressing CaRING1 at high level were selected for use in the present study (Supplemental fig. S3). Under our laboratory conditions, we examined the phenotype of 35S:CaRING1 plants and found them to be indistinguishable from the wild-type plants in terms of the shape of leaf, flower, silique, leaf number, and flowering time (data not shown). To further investigate functional involvement of CaRING1 in ABA response using 35S:CaRING1 Arabidopsis plants, we initially analyzed the effect of ABA on seed germination and seedling establishment of 35S:CaRING1 plants. In the germination assay, there were no significant differences in seed germination between 35S:CaRING1 and wild-type plants with or without ABA treatment (data not shown). However, application of norflurazon, an inhibitor of endogenous ABA synthesis (Piskurewicz et al. 2008), allowed observing effect of CaRING1 constitutive expression during germination. Compared with wild-type plants, germination rate was low in 35S:CaRING1 plants only at 2 days after sowing (Fig. 4a). Although norflurazon prevented root elongation and seedling establishement, the lengths of the radicles emerged from 35S:CaRING1 seeds were quite shorter than those of wild-type seeds (Supplemental fig. S4). Consistently, application of 0.75 μM ABA into the media led to strong inhibition of 35S: CaRING1 seedling growth, relative to that of wild-type plants (Fig. 4b). The development of an expanded green cotyledon in the 35S:CaRING1 plants was also less than 11–21 % that of the wild-type plants (Fig. 4c). We then analyzed the root lengths of wild-type and 35S:CaRING1 plants treated with various concentrations of ABA (Fig. 4d, e). Seven days after treatment, the inhibitory effects of ABA on root growth were observed in wild-type and 35S:CaRING1 plants (Fig. 4d). In the presence of 0.75 μM ABA, growth of 35S:CaRING1 roots was retarded in contrast to that observed in wild-type roots. In plants treated with 0.75 or 1.0 μM ABA, the root length of 35S:CaRING1 plants was less than 40–60 % of that in the wild-type plants (Fig. 4e). One of the Arabidopsis genes has high sequence homology with CaRING1, loss-of-function mutants of RFH2 gene (At1g20823) exhibited the opposite pattern in root elongation in the presence of 0.75 μM ABA (Supplemental fig. S5a and b). Root lengths of rfh2 mutants were at least threefold longer than those of WT. These results indicate that constitutive expression of the CaRING1 gene confers enhanced ABA sensitivity in Arabidopsis during seedling establishment.
Phenotypic analysis of the 35S:CaRING1 transgenic Arabidopsis mutants under ABA treatment. a Germination rates of wild-type and transgenic plants exposed to norflurazon and ABA. The percentage of seeds with radicle emergence was scored 2 and 4 days after plating on 0.5X MS containing 50 μM norflurazon and various concentration of ABA. b, c Seedling establishment of wild-type and 35S:CaRING1 plants in 0.5X MS containing 0.75 μM ABA. The number of seedlings with green cotyledons was counted 7 days after sowing and representative images were simultaneously taken. d, e Seedling growth of wild-type and 35S:CaRING1 plants exposed to ABA. The seedlings were grown vertically in 0.5X MS containing 0.75 and 1.0 μM ABA for 7 days. The representative images were taken (d) and root length of each line was measured (e). Data are the mean ± standard deviation from three independent experiments each evaluating 50 seeds. Different letters indicate significant differences in three independent experiments (ANOVA; P < 0.05)
Enhanced tolerance of 35S:CaRING1 Arabidopsis transgenic plants to dehydration stress
Since the CaRING1 gene was induced in dehydrated pepper leaves (Fig. 1) and silencing of CaRING1 gene in pepper plants reduced dehydration tolerance (Fig. 3), we investigated the dehydration tolerance of 35S:CaRING1 plants (Fig. 5). Wild-type and 35S:CaRING1 plants were grown for 4 weeks under normal growth conditions and were then subjected to dehydration stress when grown under well-watered conditions (Fig. 5a, left panel), and after dehydration treatment in which water was withheld for 12 days (Fig. 5a, middle panel), there was no phenotypic difference between the two plant lines. However, after re-watering for 3 days (Fig. 5a, right panel), the transgenic lines exhibited a phenotype with less wilting compared with that observed in wild-type plants. Only 15 % of wild-type plants survived, whereas 90 and 85 % of 35S:CaRING1 lines #13 and #16, respectively, were able to resume their growth and survived (Fig. 5a). Interestingly, loss-of-function mutants of RFH2 gene showed no difference in dehydration tolerance compared with wild-type plants (Supplemental fig. S5c), indicating that CaRING1 has multiple functions relative to Arabidopsis RFH2 gene.
Enhanced dehydration tolerance of the 35S:CaRING1 transgenic Arabidopsis plants. a Dehydration sensitivity of the 35S:CaRING1 plants. Three-week-old wild-type (WT) and transgenic plants were treated with dehydration stress by withholding water for 12 days, followed by rehydration for 3 days. The representative images were taken and percentages of plants that survived were measured. b Transpirational water loss of leaves of wild-type and transgenic plants. Rate of water loss from leaves of each line was measured every hour after detachment of leaves for 8 h. Data are the mean ± standard error from three independent experiments each evaluating 12 plants. c ABA content in leaves of 35S:CaRING1 and wild-type plants after dehydration treatment for 2 h. Data are the mean ± standard error from three independent experiments. Different letters indicate significant differences in three independent experiments (ANOVA; P < 0.05)
To further evaluate the response to dehydration stress, we measured the fresh weight of detached rosette leaves to monitor transpirational water loss and to thus determine whether the dehydration tolerant phenotype of the 35S:CaRING1 plants resulted from a low transpiration rate (Fig. 5b). The weights of rosette leaves detached from 4-week old plants of both wild-type and 35S:CaRING1 lines were measured over time (0–8 h) and approximately 47 and 42–43 % of their fresh weights were decreased, respectively (Fig. 5b). Since silencing of CaRING1 partially suppressed ABA biosynthesis (Fig. 3c), we measured ABA levels in leaves of 35S:CaRING1 plants treated with dehydration stress for 2 h (Fig. 5c). Under normal condition, ABA levels were not significantly differences between 35S:CaRING1 and wild-type plants. However, upon dehydration stress, 35S:CaRING1 plants exhibited high ABA accumulation compared with wild-type plants. These data suggest that the level of CaRING1 expression is positively correlated with stress tolerance. The molecular mechanisms responsible for the positive effects of ABA under dehydration stress are well established (Zhu 2002; Yamaguchi-Shinozaki and Shinozaki 2006). In addition, many studies have reported that ABA sensitivity is associated with dehydration tolerance (Cheong et al. 2007; Lim and Lee 2014; Ryu et al. 2010; Santiago et al. 2009). Based on these, constitutive expression of CaRING1 gene may lead to increase of ABA-mediated dehydration tolerance. However, we cannot rule out the possibility that ABA-independent responses were also altered, because induction of CaRING1 gene expression in pepper plant was faster by dehydration stress than by ABA treatment (Fig. 1a, b).
Participation of CaRING1 in ABA-induced stomatal aperture and induction of ABA- or dehydration-related genes in 35S:CaRING1 Arabidopsis transgenic plants
To determine whether the enhanced dehydration tolerance of 35S:CaRING1 Arabidopsis transgenic plants is associated with the ABA response, we measured the size of stomatal pores with or without ABA treatment (Fig. 6). There were no significant differences in the size of the stomatal aperture between the wild-type and 35S:CaRING1 leaves in the absence of ABA. However, the size of stomatal pores decreased more dramatically in leaves of 35S:CaRING1 plants than in those of wild-type plants after treatment with 10 or 20 μM ABA. These data indicate that ABA hypersensitivity in the guard cells of 35S:CaRING1 plants may enhance water retention, leading to a dehydration tolerance phenotype.
ABA-induced stomatal closing in leaves of 35S:CaRING1 mutants. Stomatal apertures were measured from leaf peels of wild-type and 35S:CaRING1 mutants plants treated with 10 and 20 μM ABA for 2.5 h and the pictures show representative stomata from each sample (a). Average stomatal aperture was calculated as pore width/length (b). Data are the means ± standard errors (n = 100). Values are the mean ± SE from three independent experiments. Statistical analysis was performed using ANOVA test (P < 0.05) and significant differences between wild-type and mutant plants are indicated by different letters
Since the ectopic expression of CaRING1 is positively correlated with ABA sensitivity, we examined whether CaRING1 affects the expression levels of ABA biosynthesis-related or ABA-induced genes including NCED3, NCED5, ABF3, ABI5, RAB18, and RD29B (Fig. 7a). Under normal growth conditions, the expression levels of these genes were not significantly different between the wild-type and 35S:CaRING1 plants except for NCED3. Compared with the wild-type plant, low expression of NCED3 in 35S:CaRING1 did not support the possibility that CaRING1 promotes NCED3 gene expression as shown in TRV:CaRING1 (Fig. 3c). However, in contrast to pepper NCED3 gene, alteration of Arabidopsis NCED3 gene did not made difference in ABA content between 35S:CaRING1 plants and wild-type plants under non-treatment condition (Fig. 5c). This discrepancy may be explained by functional diversity of CaRING1 and different function of NCED3 between two plant species. To examine effect of ABA on transcriptional alteration of those ABA signaling-related genes, 4-week-old wild-type and 35S:CaRING1 plants were treated with 50 μM ABA. The level of NCED3 and NCED5 transcription, which are associated with ABA synthesis (Frey et al. 2012), was significantly lower in 35S:CaRING1 than in wild-type plants and not different, respectively, (Fig. 7a). However, there was higher accumulation of ABF3 and ABI5, which encode basic leucine zipper (bZIP) transcription factors (Chen et al. 2013), in 35S:CaRING1 compared with wild-type plants, indicating that CaRING1 may act upstream of these transcription factors in the ABA signal transduction pathway. Moreover, induction of the ABA-responsive marker genes RAB18 and RD29B was also significantly higher in 35S:CaRING1 than in the wild-type plants (Fig. 7a). This increased expression of several ABA-induced marker genes may reflect ABA hypersensitivity of 35S:CaRING1 plants.
Expression analysis of ABA biosynthesis-related and ABA-responsive genes in leaves of the 35S:CaRING1 transgenic plants and wild-type plants treated with ABA (50 μM; a) and subjected to dehydration (b). Relative expression level (∆∆CT) of each gene was normalized to that of Actin8 as an internal control gene and compared with the value for mock-treated leaves. Data are the means ± standard error from three independent experiments. Different letters indicate significant differences in three independent experiments (ANOVA; P < 0.05)
In line with these results, to determine that the tolerance phenotype of 35S:CaRING1 plants to dehydration stress was also influenced by the level of ABA- or dehydration-induced marker gene expression, we performed qRT-PCR analysis in wild-type and 35S:CaRING1 plants subjected to dehydration conditions (Fig. 5b). There was higher accumulation of NCED3 transcripts in 35S:CaRING1 plants compared with wild-type plants after treatment with dehydration stress, whereas the level of NCED5 transcription was lower in 35S:CaRING1 plants. This may contribute to high accumulation of ABA in 35S:CaRING1 plants treated with dehydration stress (Fig. 5c). As shown in Fig. 7b, the expression levels of transcription factors and dehydration -induced marker genes, including ABF3, ABI5, RAB18, and RD29B, were significantly higher in the 35S:CaRING1 plants than in the wild-type plants in response to dehydration-stress conditions, indicating that changes in the expression of stress genes may underlie the altered dehydration tolerance in 35S:CaRING1 plants. In addition, although stress-related gene expression showed some degree of variation in the two transgenic lines, ectopic expression of CaRING1 may positively regulate the expression of stress marker genes in plants treated with ABA and exposed to dehydration stress, thereby presumably enhancing dehydration stress tolerance in the 35S:CaRING1 plants.
In conclusion, these findings suggest that CaRING1 acts as a positive regulator to provide tolerance to dehydration stress through regulation of ABA biosynthesis, ABA-induced stomatal closure, and ABA-mediated stress-responsive gene expression. Although we elucidated the in vivo functions of CaRING1 in plant defense responses to pathogen infection and dehydration, it is still unclear how CaRING1 serves as a positive regulator of biotic and abiotic stress responses. Subsequent molecular and physiological analysis of the downstream target(s) of CaRING1 will improve our understanding of the function of CaRING1 under biotic and abiotic stress conditions.
Abbreviations
- ABA:
-
Abscisic acid
- Pst :
-
Pseudomonas syringae pv. tomato
- PR:
-
Pathogenesis related
- qRT-PCR:
-
Quantitative reverse transcription-polymerase chain reaction
- ROS:
-
Reactive oxygen species
- SA:
-
Salicylic acid
- SOS:
-
Stomatal opening solution
- VIGS:
-
Virus-induced gene silencing
- Xcv :
-
Xanthomonas campestris pv. vesicatoria
References
Barrero JM, Rodriguez PL, Quesada V, Piqueras P, Ponce MR, Micol J (2006) Both abscisic acid (ABA)-dependent and ABA-independent pathways govern the induction of NCED3, AAO3 and ABA1 in response to salt stress. Plant Cell Environ 29:2000–2008
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24:23–58
Bu Q, Li H, Zhao Q, Jiang H, Zhai Q, Zhang J, Wu X, Sun J, Xie Q, Wang D, Li C (2009) The Arabidopsis RING finger E3 ligase RHA2a is a novel positive regulator of abscisic acid signaling during seed germination and early seedling development. Plant Physiol 150:463–481
Callis J (2014) The ubiquitination machinery of the ubiquitin system. Arabidopsis Book 12:e0174
Chang L, Zhang Z, Yang J, McLaughlin SH, Barford D (2014) Molecular architecture and mechanism of the anaphase-promoting complex. Nature 513:388–393
Chen YT, Liu H, Stone S, Callis J (2013) ABA and the ubiquitin E3 ligase KEEP ON GOING affect proteolysis of the Arabidopsis thaliana transcription factors ABF1 and ABF3. Plant J 75:965–976
Cheng WH, Endo A, Zhou L, Penney J, Chen HC, Arroyo A, Leon P, Nambara E, Asami T, Seo M, Koshiba T, Sheen J (2002) A unique short-chain dehydrogenase/reductase in Arabidopsis glucose signaling and abscisic acid biosynthesis and functions. Plant Cell 14:2723–2743
Cheong YH, Pandey GK, Grant JJ, Batistic O, Li L, Kim BG, Lee SC, Kudla J, Luan S (2007) Two calcineurin B-like calcium sensors, interacting with protein kinase CIPK23, regulate leaf transpiration and root potassium uptake in Arabidopsis. Plant J 52:223–239
Ciechanover A (1998) The ubiquitin-proteasome pathway: on protein death and cell life. EMBO J 17:7151–7160
Ciechanover A, Schwartz AL (1998) The ubiquitin-proteasome pathway: the complexity and myriad functions of proteins death. Proc Natl Acad Sci USA 95:2727–2730
Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679
Finkelstein RR, Gampala SSL, Rock CD (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell Suppl 14:15–45
Frey A, Effroy D, Lefebvre V, Seo M, Perreau F, Berger A, Sechet J, To A, North HM, Marion-Poll A (2012) Epoxycarotenoid cleavage by NCED5 fine-tunes ABA accumulation and affects seed dormancy and drought tolerance with other NCED family members. Plant J 70:501–512
Gagne JM, Smalle J, Gingerich DJ, Walker JM, Yoo SD, Yanagisawa S, Vierstra RD (2004) Arabidopsis EIN3-binding F-box 1 and 2 form ubiquitin-protein ligases that repress ethylene action and promote growth by directing EIN3 degradation. Proc Natl Acad Sci USA 101:6803–6808
Glickman MH, Adir N (2004) The proteasome and the delicate balance between destruction and rescue. PLoS Biol 2:E13
Gonzalez-Guzman M, Pizzio GA, Antoni R, Vera-Sirera F, Merilo E, Bassel GW, Fernández MA, Holdsworth MJ, Perez-Amador MA, Kollist H, Rodriguez PL (2012) Arabidopsis PYR/PYL/RCAR receptors play a major role in quantitative regulation of stomatal aperture and transcriptional response to abscisic acid. Plant Cell 24:2483–2496
Hatakeyama S, Yada M, Matsumoto M, Ishida N, Nakayama KI (2001) U box proteins as a new family of ubiquitin-protein ligases. J Biol Chem 276:33111–33120
Hershko A, Ciechanover A (1998) The ubiquitin system. Annu Rev Biochem 67:425–479
Hong JK, Choi HW, Hwang IS, Hwang BK (2007) Role of a novel pathogen-induced pepper C3–H–C4 type RING-finger protein gene, CaRFPI, in disease susceptibility and osmotic stress tolerance. Plant Mol Biol 63:571–588
Hu X, Zhang A, Zhang J, Jiang M (2006) Abscisic acid is a key inducer of hydrogen peroxide production in leaves of maize plants exposed to water stress. Plant Cell Physiol 47:1484–1495
Huang Y, Minaker S, Roth C, Huang S, Hieter P, Lipka V, Wiermer M, Li X (2014) An E4 ligase facilitates polyubiquitination of plant immune receptor resistance proteins in Arabidopsis. Plant Cell 26:485–496
Hubbard KE, Nishimura N, Hitomi K, Getzoff ED, Schroeder JI (2010) Early abscisic acid signal transduction mechanisms: newly discovered components and newly emerging questions. Genes Dev 24:1695–1708
Irigoyen ML, Iniesto E, Rodriguez L, Puga MI, Yanagawa Y, Pick E, Strickland E, Paz-Ares J, Wei N, De Jaeger G, Rodriguez PL, Deng XW, Rubio V (2014) Targeted degradation of abscisic acid receptors is mediated by the ubiquitin ligase substrate adaptor DDA1 in Arabidopsis. Plant Cell 26:712–728
Jacobson AD, Zhang NY, Xu P, Han KJ, Noone S, Peng J, Liu CW (2009) The lysine 48 and lysine 63 ubiquitin conjugates are processed differently by the 26s proteasome. J Biol Chem 284:35485–35494
Jakab G, Ton J, Flors V, Zimmerli L, Metraux JP, Mauch-Mani B (2005) Enhancing Arabidopsis salt and drought stress tolerance by chemical priming for its abscisic acid responses. Plant Physiol 139:267–274
Jiang M, Zhang J (2001) Effect of abscisic acid on active oxygen species, antioxidative defence system and oxidative damage in leaves of maize seedlings. Plant Cell Physiol 42:1265–1273
Kim JH, Kim WT (2013a) The Arabidopsis RING E3 ubiquitin ligase AtAIRP3/LOG2 participates in positive regulation of high-salt and drought stress responses. Plant Physiol 162:1733–1749
Kim SJ, Kim WT (2013b) Suppression of Arabidopsis RING E3 ubiquitin ligase AtATL78 increases tolerance to cold stress and decreases tolerance to drought stress. FEBS Lett 587:2584–2590
Kwak JM, Mori IC, Pei ZM, Leonhardt N, Torres MA, Dangl JL, Bloom RE, Bodde S, Jones JD, Schroeder JI (2003) NADPH oxidase AtrbohD and AtrbohF genes function in ROS-dependent ABA signaling in Arabidopsis. EMBO J 22:2623–2633
Lee SC, Luan S (2012) ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ 35:53–60
Lee SC, Hwang IS, Choi HW, Hwang BK (2008) Identification and functional expression of the novel pepper antimicrobial protein, CaAMP1 enhances broad-spectrum disease resistance in transgenic Arabidopsis. Plant Physiol 148:1004–1020
Lee DH, Choi HW, Hwang BK (2011) The pepper E3 ubiquitin ligase RING1 gene, CaRING1, is required for cell death and the salicylic acid-dependent defense response. Plant Physiol 156:2011–2025
Lee SC, Lim CW, Lan W, He K, Luan S (2013) ABA signaling in guard cells entails a dynamic protein-protein interaction relay from the PYL-RCAR family receptors to ion channels. Mol Plant 6:528–538
Leung J, Giraudat J (1998) Abscisic acid signal transduction. Annu Rev Plant Physiol Plant Mol Biol 49:199–222
Li H, Jiang H, Bu Q, Zhao Q, Sun J, Xie Q, Li C (2011) The Arabidopsis RING finger E3 ligase RHA2b acts additively with RHA2a in regulating abscisic acid signaling and drought response. Plant Physiol 156:550–563
Lim CW, Lee SC (2014) Functional roles of the pepper MLO protein gene, CaMLO2, in abscisic acid signaling and drought sensitivity. Plant Mol Biol 85:1–10
Lim CW, Luan S, Lee SC (2014) A prominent role for RCAR3-mediated ABA signaling in response to Pseudomonas syringae pv. tomato DC3000 infection in arabidopsis. Plant Cell Physiol 55:1691–1703
Lim CW, Lim S, Baek W, Lee SC (2015) The pepper late embryogenesis abundant protein CaLEA1 acts in regulating abscisic acid signaling, drought and salt stress response. Physiol Planta 154:526–542
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Lopez-Molina L, Mongrand S, Kinoshita N, Chua NH (2003) AFP is a novel negative regulator of ABA signaling that promotes ABI5 protein degradation. Genes Dev 17:410–418
Marin I (2013) Evolution of plant HECT ubiquitin ligases. PLoS ONE 8:e68536
Miao Y, Zentgraf U (2010) A HECT E3 ubiquitin ligase negatively regulates Arabidopsis leaf senescence through degradation of the transcription factor WRKY53. Plant J 63:179–188
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio-assays with tobacco tissue cultures. Physiol Plant 15:473–497
Murata Y, Pei ZM, Mori IC, Schroeder J (2001) Abscisic acid activation of plasma membrane Ca2+ channels in guard cells requires cytosolic NAD(P)H and is differentially disrupted upstream and downstream of reactive oxygen species production in abi1-1 and abi2-1 protein phosphatase 2C mutants. Plant Cell 13:2513–2523
Park SY, Peterson FC, Mosquna A, Yao J, Volkman BF, Cutler SR (2015) Agrochemical control of plant water use using engineered abscisic acid receptors. Nature 520:545–548
Pazhouhandeh M, Molinier J, Berr A, Genschik P (2011) MSI4/FVE interacts with CUL4-DDB1 and a PRC2-like complex to control epigenetic regulation of flowering time in Arabidopsis. Proc Natl Acad Sci USA 108:3430–3435
Pei ZM, Murata Y, Benning G, Thomine S, Klüsener B, Allen GJ, Grill E, Schroeder JI (2000) Calcium channels activated by hydrogen peroxide mediate abscisic acid signalling in guard cells. Nature 406:731–734
Pickart CM (2001) Mechanisms underlying ubiquitination. Annu Rev Biochem 70:503–533
Piskurewicz U, Jikumaru Y, Kinoshita N, Nambara E, Kamiya Y, Lopez-Molina L (2008) The gibberellic acid signaling repressor RGL2 inhibits Arabidopsis seed germination by stimulating abscisic acid synthesis and ABI5 activity. Plant cell 20:2729–2745
Rock C (2000) Pathways to abscisic acid-regulated gene expression. New Phytol 148:357–396
Ryu MY, Cho SK, Kim WT (2010) The Arabidopsis C3H2C3-type RING E3 ubiquitin ligase AtAIRP1 is a positive regulator of an abscisic acid-dependent response to drought stress. Plant Physiol 154:1983–1997
Sadanandom A, Bailey M, Ewan R, Lee J, Nelis S (2012) The ubiquitin-proteasome system: central modifier of plant signalling. New Phytol 196:13–28
Santiago J, Rodrigues A, Saez A, Rubio S, Antoni R, Dupeux F, Park SY, Márquez JA, Cutler SR, Rodriguez PL (2009) Modulation of drought resistance by the abscisic acid receptor PYL5 through inhibition of clade A PP2Cs. Plant J 60:575–588
Seo KI, Lee JH, Nezames CD, Zhong S, Song E, Byun MO, Deng XW (2014) ABD1 is an Arabidopsis DCAF substrate receptor for CUL4-DDB1-based E3 ligases that acts as a negative regulator of abscisic acid signaling. Plant Cell 26:695–711
Smalle J, Vierstra RD (2004) The ubiquitin 26S proteasome proteolytic pathway. Annu Rev Plant Biol 55:555–590
Stone SL, Callis J (2007) Ubiquitin ligases mediate growth and development by promoting protein death. Curr Opin Plant Biol 10:624–632
Stone SL, Hauksdottir H, Troy A, Herschleb J, Kraft E, Callis J (2005) Functional analysis of the RING-type ubiquitin ligase family of Arabidopsis. Plant Physiol 137:13–30
Stone SL, Williams LA, Farmer LM, Vierstra RD, Callis J (2006) KEEP ON GOING, a RING E3 ligase essential for Arabidopsis growth and development, is involved in abscisic acid signaling. Plant Cell 18:3415–3428
Umezawa T, Sugiyama N, Mizoguchi M, Hayashi S, Myouga F, Yamaguchi-Shinozaki K, Ishihama Y, Hirayama T, Shinozaki K (2009) Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. Proc Natl Acad Sci USA 106:17588–17593
Wasilewska A, Vlad F, Sirichandra C, Redko Y, Jammes F, Valon C, Frey NFD, Leung J (2008) An update on abscisic acid signaling in plants and more. Mol Plant 1:198–217
Xie Q, Guo HS, Dallman G, Fang S, Weissman AM, Chua NH (2002) SINAT5 promotes ubiquitin-related degradation of NAC1 to attenuate auxin signals. Nature 419:167–170
Xiong LM, Lee HJ, Ishitani M, Zhu JK (2002) Regulation of osmotic stress-responsive gene expression by the LOS6/ABA1 locus in Arabidopsis. J Biol Chem 277:8588–8596
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Zeng LR, Vega-Sanchez ME, Zhu T, Wang GL (2006) Ubiquitination-mediated protein degradation and modification: an emerging theme in plant-microbe interactions. Cell Res 16:413–426
Zhang Y, Yang C, Li Y, Zheng N, Chen H, Zhao Q, Gao T, Guo H, Xie Q (2007) SDIR1 is a RING finger E3 ligase that positively regulates stress-responsive abscisic acid signaling in Arabidopsis. Plant Cell 19:1912–1929
Zheng N, Schulman BA, Song L, Miller JJ, Jeffrey PD, Wang P, Chu C, Koepp DM, Elledge SJ, Pagano M, Conaway RC, Conaway JW, Harper JW, Pavletich NP (2002) Structure of the Cul1-Rbx1-Skp1-F boxSkp2 SCF ubiquitin ligase complex. Nature 416:703–709
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgments
This work was supported by a Grant from “The Next-Generation BioGreen 21 Program for Agriculture & Technology Development (Project No. PJ011010501)” Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights
Human participants and/or animals have not involved in this research.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Expression patterns of CaRING1-homologous genes from Arabidopsis in response to ABA and drought stress. (A and B) Relative expression levels of Arabidopsis RING-H2 genes in guard cells and leaves of 5-week-old plants 3 h after treatment with 50 μM ABA or ethanol (control). (C and D) Relative expression levels of Arabidopsis RING-H2 genes in 7-day-old seedling treated with 10 μM ABA. (E and F) Relative expression levels of Arabidopsis RING-H2 genes in 18-day-old seedling treated with drought stress. For drought stress treatment, plants grown on rafts in Magenta boxes were exposed to the air stream for 15 min with loss of approximately 10 % fresh weight. Data were obtained from Arabidopsis eFP Browser in the Bio-Analytic Resource for Plant Biology (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi). (PPTX 624 kb)
Supplementary Fig. 2
Expression patterns of CaRING1-homologous genes from Arabidopsis in response to cold, osmotic, and salt stress. Relative expression levels of Arabidopsis RING-H2 genes in shoot (right) and root (left) of 18-day-old seedling treated with cold (A; continuous 4 °C on crushed ice in cold chamber), osmotic (B; 300 mM mannitol), and salt (C; 150 mM NaCl). Data were obtained from Arabidopsis eFP Browser in the Bio-Analytic Resource for Plant Biology (http://bar.utoronto.ca/efp/cgi-bin/efpWeb.cgi). (PPTX 81 kb)
Supplemental Fig. S3
Constitutive expression of CaRING1 in the transgenic lines. RT-PCR analysis of CaRING1 gene expression in wild-type and transgenic lines. Actin 8 was used as an internal control. (PPTX 53 kb)
Supplemental Fig. S4
Effect of norflurazon (NF) and ABA during seed germination and seedling establishment of 35S:CaRING1 mutants. The seeds of 35S:CaRING1 mutants and wild-type plants were germinated on 0.5X MS containing 50 μM NF alone or 50 μM NF and 0.75 μM ABA. After 10 days, the representative images were taken. (PPTX 125 kb)
Supplemental Fig. S5
ABA and dehydration sensitivity of rfh2 mutants. (A) Scheme of the Arabidopsis RFH2 gene structure. White and black boxes indicate exons and introns, respectively. The positions of the T-DNA insertions in rhf2 mutants are indicated by triangles. (B) Seedling growth of wild-type and rhf2 mutants exposed to ABA. The seedlings were grown vertically in 0.5X MS containing 0.75 μM ABA for 7 days. The representative images were taken. (C) Dehydration sensitivity of the rfh2 mutants. Three-week-old wild-type (WT) and rfh2 mutants were treated with dehydration stress by withholding water for 14 days, followed by rehydration for 3 days. The representative images were taken and percentages of plants that survived were measured. (PPTX 1015 kb)
Rights and permissions
About this article
Cite this article
Lim, C.W., Hwang, B.K. & Lee, S.C. Functional roles of the pepper RING finger protein gene, CaRING1, in abscisic acid signaling and dehydration tolerance. Plant Mol Biol 89, 143–156 (2015). https://doi.org/10.1007/s11103-015-0359-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-015-0359-1