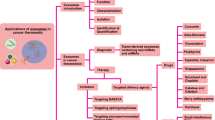
Abstract
Exosomes are extracellular vesicles secreted by cells with a particle size of 30–150 nm in diameter. Exosomes can be used as natural drug carriers. The treatment of cancer with drug-loaded exosomes is an area of high interest. This review introduces the composition, function, isolation and characterization of exosomes, and briefly describes the selection of exosome donor cells and methods for drug loading. Through studies on therapies with drug-loaded exosomes in gastric cancer, lung cancer, brain cancer and other cancers, the advantages and disadvantages of drug-loaded exosomes have been analyzed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer has become a research hotspot because of its high mortality and incurability. Common treatments for cancer therapy include surgery, chemotherapy, radiotherapy, immunotherapy, small chemical molecules and small biological molecules (including RNA and exosomes). Exosomes are a subpopulation of extracellular vesicles secreted by cells with a particle size of 30–150 nm in diameter that exist in various body fluids[1, 2]. Exosomes was first discovered in reticulocytes in 1983[3] and named in 1987[4]. Exosome release was originally thought to be a way for cells to excrete waste. Until 1996, Graca Raposo et al. found that exosomes released from B lymphoblastoid cells had anti-tumor effect[5]. Exosomes contain lipids, proteins, nucleic acids and other substances. In 2007, Hadi Valadi et al. found that mRNA and microRNA in exosomes could be transferred between cells, resulting in genetic exchange[6]. In the last decade, researches on the exosome field has made advanced progress rapidly. A large number of studies on exosomes as liquid biopsy and drug delivery platforms have been carried out. Due to the overlap of size and shape between exosomes and small-sized macrovesicles (with size less than 200 nm in diameter), it is difficult to obtain relatively pure preparation and characterize properly, leading to controversy over the term of exosome. In 2018, the International Society for Extracellular Vesicles proposed using small extracellular vesicles (sEVs) instead of exosomes as an optional term for EVs with a size less than 200 nm in diameter[7]. However, in the last decade, the term of exosome still has been widely used in scientific publications. For the convenience of readers with different expertise, we will keep using the word of exosome in this review.
The formation of exosomes mainly includes the following steps: firstly, endocytic vesicles are formed through plasmalemma invagination; secondly, early endosomes form and then mature into late endosomes; thirdly, with the entry of "cargo", such as cytoplasmic nucleic acids and proteins, many intraluminal vesicles (ILVs) are produced in late endosomes, and endosomes are then transformed into multivesicular bodies (MVBs) through inward budding; and finally, MVBs can fuse with lysosomes, which can degrade ILVs. On the other hand, MVBs can be transported to and then fuse with the plasma membrane to secrete ILVs outside the cell[4, 8, 9]. In this scenario, the ILVs are considered exosomes.
Exosomes derived from different cells have different functions, such as affecting tumor invasion[10], tissue repair[11], and immune regulation[12] and using in drug delivery [13, 14]. At present, drug carriers consist of nanoparticles, liposomes, natural or synthetic macromolecular materials, etc. Drug carriers can control the rate of drug release and actively or passively deliver drugs to the target site, avoiding the limitations of general drug therapy and being beneficial to the treatment of tumors; however, these drug carriers also have some drawbacks. As foreign substances, nanoparticles can induce an immune response, which is characterized by poor biocompatibility. In addition, they can be cleared by the mononuclear phagocyte system, leading to reduced efficacy. In contrast, exosomes, vesicles secreted by the body’s own cells, have good biocompatibility and do not cause immune reactions[15]. Elimination by the mononuclear phagocyte system can be avoided, which leads to a long cyclic half-life. The plasma half-life of BAY55-9837 loaded in exosome-SPION was reported to be 27-fold longer than that of BAY55-9837 alone[16]. Therefore, exosomes are a promising carrier for drug delivery. Studies in this field are of great significance for cancer treatment. This review summarizes the basic characteristics of exosomes, the application of exosomes as carriers for cancer treatment (in gastric cancer, lung cancer, brain cancer and other cancers), the selection of cell sources of exosomes, drug-loading methods, and the advantages and disadvantages of exosomes as drug carriers.
Isolation and characterization of exosomes
To load drugs and better study the composition and functions of exosomes, it is necessary to separate exosomes from cell culture medium, biological fluids or tissues. At present, there are various methods for exosome isolation, but there is no gold standard. The commonly used methods for the isolation of exosomes include ultracentrifugation (UC), ultrafiltration, immunoaffinity capture methods, chromatography, etc.
One of the most common isolation techniques is UC. UC can be divided into differential UC and density gradient UC[17]. The procedure for differential ultracentrifugation is to first remove dead cells and cell fragments by low-speed centrifugation and then to precipitate exosomes by ultracentrifugation steps (with a centrifugal force of 100,000–120,000 × g) according to the difference in sizes among the substances. This results in a mixture of exosomes and some contaminating proteins, so the precipitate needs to be resuspended in PBS and centrifuged again. Finally, the supernatant is removed to obtain purer exosomes. It should be noted that centrifugation operations should always be conducted at 4 °C[18]. The theory of density gradient centrifugation is that when there is a difference in the sedimentation coefficient between different particles, under a certain centrifugal force, the particles with each coefficient settle at a certain speed, ultimately forming zones in different areas of the density gradient. The commonly used density gradient solvents are sucrose and iodixanol[19]. Both sucrose and iodixanol are inert solutions capable of forming density gradient solutions. Sucrose is readily soluble in water. Exosomes could float in a sucrose gradient solution with a density of 1.10–1.18 g/ml, which is close to the density of exosomes (1.13–1.19 g/ml)[20]. Iodixanol is a highly hydrophilic substance that can form a continuous or discontinuous density gradient solution. Iodixanol has some advantages over sucrose. Firstly, it can form iso-osmotic solutions at all densities, which helps preserve the size and shape of isolated exosomes[21]; Secondly, iodixanol density gradients could accurately separate different subtypes of EVs, while the sucrose density gradients could not[22]; and thirdly, iodixanol density gradients enables the separation of other substances which has similar densities to exosomes, such as viruses and microparticles. But sucrose density gradients cannot effectively separate them[23].
Ultrafiltration can use ultrafiltration membranes with a certain size pore or molecular weight cutoff (MWCO) to selectively separate exosomes under the action of centrifugal force. Zhang et al. [24] used a 0.22-μm filter to isolate exosomes derived from human umbilical cord mesenchymal stem cells (MSCs). The ultrafiltration method is simple to perform and has a low cost. Additionally, He et al. [25] optimized an ultrafiltration method by passing urine samples sequentially through a 0.22-μm filter and a dialysis membrane with an MWCO of 10,000 kDa. Their results demonstrated that this optimized method could obtain exosomes with higher purity.
The immunoaffinity capture approach is also called the immunomagnetic bead method. There are many specific markers in the exosome membrane, such as CD9, CD63, CD81, TSG101, and Alix[26, 27]. By incubating exosomes with magnetic beads coated with specific antibodies, antigens can bind to the antibodies on the magnetic beads through antigen–antibody interactions[28]. Thus, exosomes can be specifically captured. In addition to the exosomal markers mentioned above, exosomes from certain cells may also express specific markers. For example, exosomes derived from melanoma cells express chondroitin sulfate peptidoglycan 4(CSPG4), whereas normal cell-derived exosomes do not. Therefore, exosomes from melanoma cells can be captured using anti-CSPG4 monoclonal antibodies (mAbs) by the immunoaffinity capture method[29].
In addition to the commonly used methods described above, exosome isolation methods also include size exclusion chromatography (SEC)[30], a polyethylene glycol (PEG)-based precipitation method[31], an exosome isolation kit method[32], and microfluidics[33]. These techniques for isolating exosomes have their own advantages and limitations, and the purity, recovery rate, and stability of the isolated exosomes vary (Table I). When UC and SEC were compared for exosome isolations, it was found that fewer exosomes were recovered by the UC, but the purity was higher than that achieved with SEC[34]. Compared with SEC, combined use of UC and SEC can obtain better separation results[35]. In the separation of exosomes released from human colon cancer cells, Tauro et al. compared UC, density gradient centrifugation and immunoaffinity capture methods and found that the immunoaffinity capture method was the best[36]. The appropriate isolation method can be selected according to downstream analyses and applications. Sometimes two or more isolation methods can be used in combination to exploit their advantages and make up for their defects. A study showed that the number of exosomes separated by combined use of ultrafiltration and SEC could be up to 58 times than that isolated with UC[37].
After separation and purification, isolated substances need to be characterized to verify whether they are exosomes. The characterization of exosomes is based on the size, morphology, density, and protein content of exosomes. Commonly used exosomal characterization methods include electron microscopy (including scanning electron microscopy and transmission electron microscopy (TEM)), western blot analysis, enzyme-linked immunosorbent assay, FACS analysis of labeled exosomes bound to beads, dynamic light scattering (DLS) and nanoparticle tracking analysis (NTA)[38,39,40]. The size of exosomes can be determined by DLS and NTA, and many studies have reported that the size of exosomes is 30–150 nm[41, 42]. TEM can be used to observe the morphology of exosomes. Zhu et al. found that macrophage-derived exosomes and nicotine-induced macrophage-derived exosomes are cup or sphere shaped[43]. In addition, the proteins enriched in exosomes and proteins deficient in exosomes can be used together to characterize exosomes. Jan Lötvall et al. [44] found that the proteins enriched in exosomes include tetraspanins (CD9, CD63, and CD81), integrins, and endosome- or membrane-binding proteins (TSG101, annexins, and Rabs). The proteins lacking in exosomes are HSP90B1, GM1330, CYC1, etc.
Composition and functions of exosomes
The main components of exosomes include proteins (such as tetraspanins, enzymes, transcription factors, and heat shock proteins), lipids, and RNAs (such as mRNA, miRNA, lncRNA and cirRNA) (Fig. 1)[54,55,56,57]. There are several exosome-related databases, such as ExoCarta[58], Vesiclepedia[59], and exoRBase[60]. These databases can provide information about proteins, lipids and RNAs in exosomes. However, some contents of exosomes are present in only exosomes derived from special cells, and some contents are common among all exosomes. Obviously, the content of exosomes derived from different cells varies; for example, the proteins and lipids contained in exosomes isolated from ovarian surface epithelial cells and ovarian cancer cells are clearly different[61]. The physiological and pathological states of cells also have an impact on the composition of exosomes. One study found that the level of the lncRNA PTENP1 was lower in the plasma exosomes of bladder cancer patients than in those of healthy controls[62].
At present, there are some controversies about the inclusion of DNA in exosomes. Many studies have shown the presence of DNA in exosomes[63, 64]. Moreover, Takahashi et al. [65] stated the opinion that exosomes derived from some cells can maintain cellular homeostasis by releasing harmful cytoplasmic DNA outside the cell. In contrast, Jeppesen et al. [66] used high-resolution density gradient centrifugation combined with direct immunoaffinity capture to isolate exosomes, and found that double-stranded DNA and DNA-binding histones were not detected in exosomes by nucleic acid analysis. The difference may be because different studies use exosomes produced by different cells, or because of the different isolation methods, the existence of other extracellular vesicles or non-vesicular substances may ultimately affect the experimental results. The composition of exosomes has not been fully determined; thus, more in-depth research is needed.
The formation of exosomes was described in the introduction. In the process of exosome generation, there are some mechanisms that affect the sorting of endogenous cargoes into exosomes. Cargo sorting mechanisms can be divided into two types: the endosomal sorting complex required for transport (ESCRT)-dependent mechanism and ESCRT-independent mechanisms. The ESCRT-dependent mechanism plays a key role. The ESCRT-dependent mechanism involves the ESCRT-0, ESCRT-I, ESCRT-II and ESCRT-III pathways. They work together to recognize and capture ubiquitinated proteins, and these proteins can enter endosomes through invagination and rupture of the endosomal membrane[67]. ESCRT-independent mechanisms include lipid raft- and ceramide-mediated mechanisms, which also modulate the composition of exosomes[68].
The biological functions of exosomes have been gradually elucidated with increasing research on exosomes. Exosomes are known to be involved in intercellular communication, disease diagnosis and treatment, tissue repair, and drug delivery. However, different types of cells produce exosomes with different functions; for example, exosomes from cardiac progenitor cells can inhibit cardiomyocyte apoptosis, thereby protecting the heart[69], while gastric cancer cell-derived exosomes can promote the proliferation and metastasis of gastric cancer cells by inducing neutrophil tumor-promoting activity[70]. There are also some studies showing that exosomes can be used as carriers for drug delivery; for example, exosomes derived from HEK293T cells and loaded with the superrepressor IκB can reduce the mortality of septic mice[71]. In this review, the role of exosomes as drug carriers is the main topic discussed below.
Selection of donor cells for exosomes
Almost all cells release exosomes, and exosomes are present in all body fluids[72]. Exosomes derived from different types of cells are different. Their drug-loading efficiency, yield and antitumor capacity vary. Kanchanapally et al. found that pancreatic cancer cells released more exosomes than pancreatic stellate cells and macrophages and that the released exosomes had the highest loading efficiency for doxorubicin. However, doxorubicin-loaded and macrophage-derived exosomes had the strongest antitumor effects[73]. It was also found that the inhibitory effects of taxol-loaded exosomes from different cell sources on A549 lung cancer cells exhibited the following hierarchies: taxol-loaded exosomes secreted by HuVEC-4 cells < taxol-loaded exosomes secreted by HuVEC-6 cells < taxol-loaded exosomes secreted by MSCs[74]. These results may be because the contents of exosomes are partially derived from the parental cells[75], which also plays a role in cancer treatment to some extent. Therefore, the efficacy of drug-loaded exosomes secreted by different cell types will be different. In addition, a study found that the distribution of exosomes in vivo could be influenced by the cell source of exosomes and the route of administration. HEK293T cell-derived exosomes were mainly distributed in the liver, spleen and gastrointestinal tract after intravenous injection[76]. Exosomes from some cell types have natural targeting capabilities; for example, exosomes from endothelial cells are more likely to be distributed in the bone than those secreted by osteoblasts or bone marrow MSCs[77]. For cancer treatment, in addition to exploiting the natural targeting capabilities of exosomes, exosomes can be engineered to accumulate at the cancer site[78].
Methods of loading exosomes with drugs
As natural drug delivery vehicles, exosomes can be loaded with chemotherapeutic drugs, therapeutic nucleic acids, proteins, etc. It has been reported that there are many approaches for loading drugs into exosomes for the purpose of treating many diseases (Table II). We speculate that the selection of exosome donor cells, the types of drugs, and the methods of drug loading would all affect the efficacy of exosome-based drug delivery systems. Here, we divide the methods for loading drugs into exosomes into two categories: 1) treatment of cells with drugs before the isolation of exosomes from the cells, so that the cells can release drug-containing exosomes and 2) loading of drugs into exosomes after isolation. An overview of these methods is provided below.
Treating cells before exosome isolation
Drug loading can be performed before exosomes are released from cells by transfection or mixing cells with drugs. Transfection methods include liposomal transfection, electroporation transfection, and viral infection. Hepatocyte growth factor (HGF)-specific siRNA can be transferred into HEK293T cells by using a transfection reagent; thus, the transfected cells will produce exosomes encapsulating HGF-specific siRNA. These exosomes are then taken up by gastric cancer cells, in which they play a role in inhibiting tumor growth[79]. Kalimuthu et al. found that mixing MSCs with paclitaxel (PTX) and then extruding the cells using an extruder with a polycarbonate membrane could obtain PTX-loaded exosome mimics[84].
Although there are many studies on loading chemotherapeutic drugs and RNA, it is difficult to load proteins into exosomes by the commonly used methods described above [85]. To solve this problem, some new approaches for loading proteins into exosomes have emerged. These methods are linked to the biogenesis process of exosomes. Yim et al. [86] designed “exosomes for protein loading via optically reversible protein–protein interactions” (EXPLORs) for protein delivery; this technology forms protein–protein modules by using two proteins, CIBN and CRY2. Under blue light, CIBN and CRY2 combine together, so CIBN-conjugated CD9 (an exosomal membrane protein) and a CRY2-conjugated target protein would bind, allowing the target protein to enter exosomes. After turning off the light source, the target protein can be separated from CD9 on the membrane and enter the exosomal cavity so that the target protein can enter target cells. In addition, another study showed that the late-domain-containing protein Ndfip1 could recognize a WW-tagged protein, driving ubiquitination and exosomal entry of the tagged protein during the exosomal biological process[87].
Loading drugs after exosome isolation
Methods for loading drugs after exosome isolation include electroporation, coincubation of exosomes with drugs at room temperature, sonication, extrusion, dialysis, and freeze–thaw cycling. The method of co-incubating exosomes with drugs is simple. Incubating exosomes derived from raw bovine milk with curcumin at room temperature can achieve a drug loading rate of 18–24%[88]. Sometimes saponin is added to assist in loading, that is, exosomes are incubated with drugs and saponins at room temperature[89]. When electroporation is used to load drugs, the electric current breaks through the exosomal membrane reversely, and temporary pores form in the lipid bilayer of the exosomal membrane so that exogenous drugs can enter the exosomes. However, at the same time, molecules in the exosomes may flow out[90]. Therefore, after electroporation of a mixture of exosomes and drugs, it is necessary to incubate at 37 °C to restore the exosomal membrane[83]. Sonication is also one of the methods for loading exosomes with drugs. After 6 cycles of sonication of a mixture of PTX and M1 macrophage-derived exosomes (30 s per cycle, with a 2-min cooling time after each cycle), PTX was shown to be successfully loaded into exosomes. Similar to electroporation, it is also necessary for a sonicated mixture to be incubated at 37 °C after sonication to restore the exosomal membrane[91]. In addition, dialysis can also be used to load drugs. In a study, a mixture of doxorubicin hydrochloride and exosomes derived from bone marrow MSCs was desalted and then dialyzed against PBS overnight to obtain doxorubicin-encapsulated exosomes[92].
There are differences in the drug-loading capacity and the size and morphology of obtained drug-exosome formulations. In a study on different methods of loading catalase into exosomes to treat Parkinson’s disease, different amounts of catalase were found to be loaded into exosomes. Sonication and extrusion methods could load the most exosomes, followed by freeze–thaw cycles, but the coincubation method loaded the fewest exosomes[13]. Similarly, comparing the loading efficiency of incubation to that of electroporation and sonication, it was found that the efficiency of incubation was the lowest, while electroporation was higher than that of incubation but much lower than that of sonication[93].
Usually, the drug-loading efficiency of exosomes is less than 30%[13, 93], while that of other nanoparticles can reach more than 90%[94]. The methods for fabricating high drug-loading nanoparticles include pre-loading, co-loading and post-loading. In the pre-loading method, drug nanoparticles is first prepared, then coated with a layer of other materials. Drug-loaded nanoparticles can also be formed by the self-assembly of drug-conjugated macromolecules, which is called the co-loading method. In the post-loading method, usually porous nanoparticles is first formed, followed by mixing with drug[95].
Modification of exosomes with targeting ligands
The in-vivo biodistribution of drug-loaded exosomes is of great importance during cancer treatment. Specific delivery of anti-cancer drugs to cancer sites or cancer cells can significantly increase drug killing efficiency and prevent drug damage to normal cells. Exosomes which inherit the characteristics of donor cells, express ligands that naturally target certain recipient cells, and are eventually taken up by recipient cells[96]. Furthermore, the targeting ability of exosomes can also be improved by surface modification of exosome membrane.
The surface modification of exosome membrane could be achieved by direct modification and indirect modification. The indirect modification is referred to the engineering of cells that release exosomes. Cell engineering methods include genetic engineering, metabolic engineering and direct membrane engineering of the parent cells [97]. The most common method is genetic engineering. It includes viral and non-viral methods. Fusing the targeting peptide to the N-terminus of exosome-associated transmembrane protein Lamp2b, and followed by transfecting the encoded plasmid into cells would result in the display of targeting peptide on the membrane surface of isolated exosomes[98]. Through genetic engineering of HEK293 cells, the released exosomes expressing IL3-Lamp2 could target the interleukin 3 receptor (IL3 receptor) which is highly expressed in Chronic Myeloid Leukemia cells. When compared with exosomes which were not modified by IL3, exosomes expressing IL3 were more tumor-targeted[99].
Direct modification methods could be achieved for induced targeting by chemical modification. Bio-orthogonal copper-free azide alkyne cyclo-addition (called click chemistry) was found to be able to conjugate ligands to the surface of exosomes. The cyclo (Arg-Gly-Asp-D-Tyr-Lys) peptide (c(RGDyK)) was able to bind to integrin αvβ3 in ischemic cerebral vascular endothelial cells. Conjugating (c(RGDyK)) to the surface of exosomes by click chemistry enabled the effective targeting of ischemic brain[100]. In addition, exosomes could be delivered to target sites using a magnetic field. Co-incubation of iron-loaded transferrin-conjugated Fe3O4 superparamagnetic nanoparticles (SPMNs) with exosomes enabled to form SPMN-exosomes by targeting the transferrin receptor on the exosome membrane[101].
Drug-loaded exosomes in cancer therapy
Because exosomes secreted by one type of cell can be taken up by another type of cell, it is believed that exosomes can be used as natural carriers for drug delivery. Drug-loaded exosomes can be used for the treatment of cancer, central nervous system diseases, inflammation, etc.[100, 102, 103]. Here, we mainly discuss the application of drug-loaded exosomes in the field of cancer therapy.
Gastric cancer
Gastric cancer is the fifth most common cancer type in the world, and has the third highest mortality rate[104]. Radiotherapy, chemotherapy and surgery are commonly used in the treatment of gastric cancer[105]. However, when chemotherapy is chosen, patients may have adverse reactions because not all drugs exclusively target the tumor site. Drug resistance often occurs, resulting in a poor prognosis. Therefore, considering that exosomes can be used as carriers for drug delivery, many studies have focused on the use of exosomes to target gastric cancer for treatment.
As mentioned above, the types of drugs that can be delivered by exosomes include small-molecule drugs, miRNAs, siRNAs, and proteins. It has been shown that HGF can promote the growth of gastric cancer[106], while HGF-specific siRNA can reduce HGF levels, thus inhibiting the growth of tumors. Zhang et al. applied exosomes to deliver HGF-specific siRNA to the tumor site, inhibiting tumor growth by suppressing the proliferation, migration and angiogenesis of a gastric cell line (SGC-7901)[79]. Similar to HGF, c-Met, as the receptor of HGF, is also highly expressed in gastric cancer, which accounts for the development of resistance to cisplatin, leading to a poor prognosis. Encapsulating c-Met-specific siRNA into exosomes extracted from the cell culture medium of HEK293-T cells and then delivering the exosomes to gastric cancer cells can reduce c-Met levels, thereby reducing resistance to cisplatin[104]. In addition, exosomes combined with photodynamic therapy (PDT) can be used to treat gastric cancer. Ce6 used in PDT can produce singlet oxygen under laser irradiation to induce cell apoptosis. Considering the poor compatibility of Ce6, Pan et al. synthesized a nanocomposite material, PMA/Au-BSA@Ce6, and loaded it into exosomes from patient urine to obtain Exo-PMA/Au-BSA@Ce6. Compared with free CE6 and PMA/Au-BSA@CE6, Exo-PMA/Au-BSA@CE6 accumulated more specifically in tumor sites and could significantly reduce the survival rate of gastric cancer cells (MGC-803 cells)[107]. Engineered tumor-targeting exosomes can also be obtained by plasmid transfection. Exosomes are transfected with plasmids expressing a fusion protein of internalized RGD (iRGD), and the resulting iRGD-Exos can target tumors. Recombinant methioninase (rMETase) can clear methionine, thereby inhibiting the G2 phase and tumor growth. A study found that rMETase-loaded iRGD-Exos (iRGD-Exos-rMETase) had a stronger inhibitory effect on tumors than rMETase and blank-Exos-rMETase[108].
Lung cancer
Lung cancer is a common type of cancer with a high mortality rate, and it can also be treated with chemotherapy, radiotherapy and surgery[109]. Lung cancer can be divided into small cell lung carcinoma (SCLC) and non-small cell lung carcinoma (NSCLC) according to histopathology. NSCLC is often treated with a combination of platinum drugs and paclitaxel[110]. However, free drugs and traditional nanocarrier drugs lack specificity and can cause side effects. The emergence of exosomes as carriers with the characteristics of low immunogenicity and a tumor-homing capability can significantly improve therapeutic effects. Therefore, using exosomes to deliver drugs to treat lung cancer has become a new strategy.
The targeting of exosome-drug formulations to lung cancer can be improved, by exploiting the specific binding between lung cancer cells and exosomes. For example, integrin beta 4 on the surface of exosomes (231-Exos) derived from breast cancer cells (MDA-MB-231) can specifically bind to surfactant protein C on the surface of lung cancer cells; that is, 231-Exos are enriched in the lungs. Therefore, miRNA-126, which has an inhibitory effect on lung cancer cells, can be loaded into 231-Exos to target and suppress lung cancer[111]. Appropriate modification of exosomes can also achieve a desired therapeutic effect. For example, exosomes were modified with anisamide-polyethylene glycol (AA-PEG), as AA can bind to the sigma receptor overexpressed in lung cancer cells (3LL-M27 cells), so that paclitaxel encapsulated in the exosomes could be delivered to 3LL-M27 cells. PEG is beneficial for avoiding clearance and prolonging the circulation half-life of exosomes in vivo [112]. Folic acid (FA)-functionalized exosomes can also be used to target tumors. After intravenous injection of different formulations in a lung cancer model, the accumulation of the dye Alexa Fluor 750 (AF750) in the tumor site increased in the following order: AF750<ExoAF750<FA-ExoAF750. This suggests that FA functionalization can allow more exosomes to be taken up at the tumor site[113].
Brain cancer
Brain diseases, especially brain cancer, are seriously harmful to human health. The presence of the blood–brain barrier makes the treatment of brain cancer difficult. Many antitumor drugs are unable to cross the blood–brain barrier (BBB) and thus fail to exert therapeutic effects. Therefore, approaches to enable drugs to cross the blood–brain barrier has become a focus in brain cancer therapy. At present, nanodrug delivery systems, such as liposomes, can cross the blood–brain barrier and can be used to treat brain diseases[114]. However, they are biologically toxic as foreign substances and have poor stability[115]. Exosomes with the ability to cross the blood–brain barrier and low immunogenicity have become a new type of drug carrier and have attracted much attention.
It has been found that the cell source and administration route of exosomes have effects on the distribution of exosomes in the body. For example, after intravenous injection, exosomes mainly accumulate in the liver and spleen and exhibit little accumulation in the brain and tumors[76]. Therefore, in the treatment of malignant brain tumors, it is very important to enhance the targeting of exosomes to the brain so that drug-loaded exosomes can cross the blood–brain barrier to play a pharmacological role. The methods that have been found to enable drug-loaded exosomes to reach the brain include intranasal administration, the use of exosomes with natural brain-targeting capabilities, and exosome engineering (Fig. 2). Regarding the administration route, intranasal administration, as an administration route that effectively bypasses the blood–brain barrier, can enable drug-loaded exosomes to reach the brain. For example, curcumin-loaded exosomes administered intranasally can significantly inhibit the growth of brain tumors[116]. In addition, exosomes produced by certain cells have natural brain-targeting capabilities. Macrophage-derived exosomes can target brain endothelial cells through the interaction between the integrin lymphocyte function-associated antigen 1 (LFA-1) on exosomes and intercellular adhesion molecule 1 (ICAM-1) on brain endothelial cells[96, 117]. Exosomes derived from brain endothelial cells can also be loaded with antitumor drugs to treat brain cancer[118]. Reticulocyte-derived exosomes in the blood achieve brain-targeting through the interaction between transferrin and transferrin receptors[119]. Exosomes from brain-metastatic cancer cell lines (e.g., SK-Mel-28) can also enter the brain[120, 121]. Exosomes without a natural brain-targeting ability can be modified to distribute in the brain. One approach is to engineer cells to produce exosomes that target the brain. In a study, dendritic cells were engineered to release exosomes expressing lamp2b because lamp2b can fuse with the neuron-specific RVG peptid; thus, exosomes could reach the brain[98]. Another approach is to directly modify exosomes after isolation. Exosomes modified with the RGERPPR (RGE) peptide can target gliomas that overexpress Neuropilin-1 (NRP-1) through the interaction between RGE and NRP-1[122]. Exosomes modified with the Cyclo (Arg-Gly-Asp-D-Tyr-Lys) peptide can also target glioblastoma that overexpresses αvβ3 integrin receptors[123].
Methods for Exosomes to Cross the bbb to Reach the Brain. Four types of Exosomes could Target Brain: (1) Exosomes from Certain Cells could cross the bbb through Ligand-receptor Interaction (such as lfa-1 and icam-1, transferrin and transferrin receptor, rgd and rgd receptor); (2) exosomes from brain endothelial cells have natural brain targeting capability; (3) brain-metastatic cancer cells could breach the bbb through transcytosis; (4) exosomes modified with ligands that target brain ecs could interact with receptors on ecs to cross the bbb
Other cancers
In addition to gastric cancer, lung cancer, and brain cancer, which are described in detail above, other cancers can also be treated with drug-loaded exosomes. As reported previously, Anthos-loaded exosomes and paclitaxel-loaded exosomes can significantly inhibit the growth of ovarian cancer[124]. Compared with paclitaxel alone, paclitaxel-loaded M1 macrophage-derived exosomes have a stronger anti-breast cancer effect[91]. To target triple-negative breast cancer (TNBC), exosomes can be modified with a disintegrin, metalloproteinase 15 (binding to the integrin αvβ3)[125] and FA (binding to FA receptors overexpressed by TNBC cells)[126]. Similarly, exosomes can be used as carriers loaded with drugs to treat pancreatic cancer[127], prostate cancer[128], colorectal cancer[129], cervical cancer[88], chronic myelogenous leukemia[99] and so on and play an effective antitumor role.
In the process of applying drug-loaded exosomes as a cancer therapy, the accumulation of exosomes in the tumor site is very important for the loaded drug to exert its therapeutic effect. In addition to some exosomes having natural targeting capabilities, modification of exosomes to improve their targeting is often employed. This type of modification can allow exosomes to accumulate mainly in specific locations but may also cause an immune response. This is due to the presence of major histocompatibility complex molecules and costimulatory molecules on the exosomal membrane derived from dendritic cells, which can enhance the immune response[96, 130]. However, exosomes with a natural targeting ability also have the disadvantage of a low yield. Therefore, it is very important to choose a suitable cell source for exosomes and increase the production of exosomes to treat cancers.
Advantages and disadvantages of drug-loaded exosomes in cancer therapy
Due to the enhanced permeability and retention effect on tumor tissue, a nanodrug delivery system (NDDS) can pass through blood vessels to accumulate in tumors[131]; however, the NDDS is immunogenic and toxic[132]. As natural drug carriers, exosomes have many advantages in cancer therapy. First, exosomes, as vesicles produced by body cells, have superior biocompatibility and low immunogenicity[74]. Second, exosomes have natural or acquired targeting characteristics, allowing loaded drugs to accumulate in specific organs or tissues. For enabling targeting of tumors by exosomes, there are three approaches: exploiting a receptor-ligand interaction or antigen–antibody binding and targeting the microenvironment[133]. We also know that exosomes derived from tumor cells have tumor-homing properties. Cells tend to take up exosomes produced by the same cell type[134]. Therefore, tumor-derived exosomes usually accumulate at the tumor site[14]. In addition, the exosomal protein CD47 can bind to signal regulatory protein alpha, which triggers “don’t eat me” signaling to prevent clearance by the mononuclear macrophage system, thus prolonging the half-life of exosomes[127]. Preinjection of an exosomal blockade agent can also be used to block the phagocytosis of exosomes by macrophages, prevent massive accumulation in the liver and spleen, and further allow engineered exosomes to reach the target site[135]. Due to the above advantages of exosomes, drug-loaded exosomes can significantly inhibit the growth of tumors compared to free drugs and nanocarrier-encapsulated drugs, such as liposomes-encapsulated drugs[127].
However, the application of drug-loaded exosomes in cancer treatment also has the defects of low drug-loading efficiency and a low yield. Both this and the low production of exosomes limit the clinical application of exosomes. To overcome these problems, researchers have performed a substantial amount of research and developed many ways to increase the production of exosomes. For example, culturing cells on a cell nanoporation chip and separating and loading exosomes simultaneously. Compared with the electroporation drug-loading method, this method increases the output of exosomes by 50 times[136]. Extrusion of exosomes with synthetic lipids can increase the number of exosomes by 6–43 times while maintaining the targeting property[137]. Some exosome-mimetic nanovesicles have been designed to increase yield. Serial extrusion of cells using nanometer-sized membranes or filters can produce a large number of exosome-mimetic nanovesicles. The output can be increased by 100 times[138, 139]. Furthermore, the emergence of artificial chimeric exosomes can also solve the problem of low yield of exosomes[140]. The artificial chimeric exosomes are prepared by integrating the cell membrane proteins of various cells into the phospholipid bilayer, which can ultimately possess multiple targeting capabilities from all parental cells and be produced in large scale. For instance, artificial chimeric exosomes produced using membrane proteins of red blood cells and MCF-7 cancer cells can not only express CD47 protein, avoiding being cleared by macrophages, but also inherit the targeting effect of cancer cells[141]. Although the above-mentioned exosome-mimetic nanovesicles are different from natural exosomes in some aspects, they can be a future developmental direction to overcome the weaknesses of exosomes.
In addition to the drug-loading efficiency and yield, the cell source of exosomes, exosome isolation methods, drug-loading methods and route of administration may all affect the therapeutic effect of drug-loaded exosomes[76]. All these aspects need to be considered to make the most favorable choice for treatment.
Conclusion and perspective
On the basis of many years of research, the composition, function, isolation and characterization of exosomes are relatively clear. Many studies have been performed on exosomes, and exosomes can be used as carriers for drug delivery. Exosomes, as a new type of drug carrier, have attracted extensive attention in the field of cancer therapy. Different antitumor effects are produced after drug loading with exosomes from different cellular sources with different compositions, yields, drug loading efficiencies and targeting capabilities; thus, any specific study must select appropriate exosomes. To treat cancer, it is often necessary to engineer exosomes so that the exosomes can accumulate at the tumor site through antigen–antibody binding, a receptor-ligand interaction, or targeting of the microenvironment. Moreover, in research on drug-loaded exosomes in cancer therapy, drug-loaded exosomes have been shown to have the advantages of relatively strong antitumor efficacy, low immunogenicity, a good targeting capability, and a long half-life in vivo. This makes drug-loaded exosomes more advantageous than traditional nanocarrier drugs. The best cell sources and the best drug-loading methods for the treatment of various cancers have not yet been fully elucidated. These points require further research in the future.
References
Boyiadzis M, Whiteside TL. The emerging roles of tumor-derived exosomes in hematological malignancies. Leukemia. 2017;31(6):1259–68.
Li M, Zeringer E, Barta T, Schageman J, Cheng A, Vlassov AV. Analysis of the RNA content of the exosomes derived from blood serum and urine and its potential as biomarkers. Philos Trans R Soc Lond B Biol Sci. 2014;369(1652):20130502.
Pan BT, Johnstone RM. Fate of the transferrin receptor during maturation of sheep reticulocytes in vitro: selective externalization of the receptor. Cell. 1983;33(3):967–78.
Johnstone RM, Adam M, Hammond JR, Orr L, Turbide C. Vesicle formation during reticulocyte maturation Association of plasma membrane activities with released vesicles (exosomes). J Biol Chem. 1987;262(19):9412–20.
Raposo G, Nijman HW, Stoorvogel W, Liejendekker R, Harding CV, Melief CJ, Geuze HJ. B lymphocytes secrete antigen-presenting vesicles. J Exp Med. 1996;183(3):1161–72.
Valadi H, Ekström K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 2007;9(6):654–9.
Théry C, Witwer KW, Aikawa E, Alcaraz MJ, Anderson JD, Andriantsitohaina R, Antoniou A, Arab T, Archer F, Atkin-Smith GK, Ayre DC, Bach JM, Bachurski D, Baharvand H, Balaj L, Baldacchino S, Bauer NN, Baxter AA, Bebawy M, Beckham C, Bedina Zavec A, Benmoussa A, Berardi AC, Bergese P, Bielska E, Blenkiron C, Bobis-Wozowicz S, Boilard E, Boireau W, Bongiovanni A, Borràs FE, Bosch S, Boulanger CM, Breakefield X, Breglio AM, Brennan M, Brigstock DR, Brisson A, Broekman ML, Bromberg JF, Bryl-Górecka P, Buch S, Buck AH, Burger D, Busatto S, Buschmann D, Bussolati B, Buzás EI, Byrd JB, Camussi G, Carter DR, Caruso S, Chamley LW, Chang YT, Chen C, Chen S, Cheng L, Chin AR, Clayton A, Clerici SP, Cocks A, Cocucci E, Coffey RJ, Cordeiro-da-Silva A, Couch Y, Coumans FA, Coyle B, Crescitelli R, Criado MF, D’Souza-Schorey C, Das S, Datta Chaudhuri A, de Candia P, De Santana EF, De Wever O, Del Portillo HA, Demaret T, Deville S, Devitt A, Dhondt B, Di Vizio D, Dieterich LC, Dolo V, Dominguez Rubio AP, Dominici M, Dourado MR, Driedonks TA, Duarte FV, Duncan HM, Eichenberger RM, Ekström K, El Andaloussi S, Elie-Caille C, Erdbrügger U, Falcón-Pérez JM, Fatima F, Fish JE, Flores-Bellver M, Försönits A, Frelet-Barrand A, Fricke F, Fuhrmann G, Gabrielsson S, Gámez-Valero A, Gardiner C, Gärtner K, Gaudin R, Gho YS, Giebel B, Gilbert C, Gimona M, Giusti I, Goberdhan DC, Görgens A, Gorski SM, Greening DW, Gross JC, Gualerzi A, Gupta GN, Gustafson D, Handberg A, Haraszti RA, Harrison P, Hegyesi H, Hendrix A, Hill AF, Hochberg FH, Hoffmann KF, Holder B, Holthofer H, Hosseinkhani B, Hu G, Huang Y, Huber V, Hunt S, Ibrahim AG, Ikezu T, Inal JM, Isin M, Ivanova A, Jackson HK, Jacobsen S, Jay SM, Jayachandran M, Jenster G, Jiang L, Johnson SM, Jones JC, Jong A, Jovanovic-Talisman T, Jung S, Kalluri R, Kano SI, Kaur S, Kawamura Y, Keller ET, Khamari D, Khomyakova E, Khvorova A, Kierulf P, Kim KP, Kislinger T, Klingeborn M, Klinke DJ 2nd, Kornek M, Kosanović MM, Kovács ÁF, Krämer-Albers EM, Krasemann S, Krause M, Kurochkin IV, Kusuma GD, Kuypers S, Laitinen S, Langevin SM, Languino LR, Lannigan J, Lässer C, Laurent LC, Lavieu G, Lázaro-Ibáñez E, Le Lay S, Lee MS, Lee YXF, Lemos DS, Lenassi M, Leszczynska A, Li IT, Liao K, Libregts SF, Ligeti E, Lim R, Lim SK, Linē A, Linnemannstöns K, Llorente A, Lombard CA, Lorenowicz MJ, Lörincz ÁM, Lötvall J, Lovett J, Lowry MC, Loyer X, Lu Q, Lukomska B, Lunavat TR, Maas SL, Malhi H, Marcilla A, Mariani J, Mariscal J, Martens-Uzunova ES, Martin-Jaular L, Martinez MC, Martins VR, Mathieu M, Mathivanan S, Maugeri M, McGinnis LK, McVey MJ, Meckes DG Jr, Meehan KL, Mertens I, Minciacchi VR, Möller A, Møller Jørgensen M, Morales-Kastresana A, Morhayim J, Mullier F, Muraca M, Musante L, Mussack V, Muth DC, Myburgh KH, Najrana T, Nawaz M, Nazarenko I, Nejsum P, Neri C, Neri T, Nieuwland R, Nimrichter L, Nolan JP, Nolte-’t Hoen EN, Noren Hooten N, O’Driscoll L, O’Grady T, O’Loghlen A, Ochiya T, Olivier M, Ortiz A, Ortiz LA, Osteikoetxea X, Østergaard O, Ostrowski M, Park J, Pegtel DM, Peinado H, Perut F, Pfaffl MW, Phinney DG, Pieters BC, Pink RC, Pisetsky DS, Pogge von Strandmann E, Polakovicova I, Poon IK, Powell BH, Prada I, Pulliam L, Quesenberry P, Radeghieri A, Raffai RL, Raimondo S, Rak J, Ramirez MI, Raposo G, Rayyan MS, Regev-Rudzki N, Ricklefs FL, Robbins PD, Roberts DD, Rodrigues SC, Rohde E, Rome S, Rouschop KM, Rughetti A, Russell AE, Saá P, Sahoo S, Salas-Huenuleo E, Sánchez C, Saugstad JA, Saul MJ, Schiffelers RM, Schneider R, Schøyen TH, Scott A, Shahaj E, Sharma S, Shatnyeva O, Shekari F, Shelke GV, Shetty AK, Shiba K, Siljander PR, Silva AM, Skowronek A, Snyder OL 2nd, Soares RP, Sódar BW, Soekmadji C, Sotillo J, Stahl PD, Stoorvogel W, Stott SL, Strasser EF, Swift S, Tahara H, Tewari M, Timms K, Tiwari S, Tixeira R, Tkach M, Toh WS, Tomasini R, Torrecilhas AC, Tosar JP, Toxavidis V, Urbanelli L, Vader P, van Balkom BW, van der Grein SG, Van Deun J, van Herwijnen MJ, Van Keuren-Jensen K, van Niel G, van Royen ME, van Wijnen AJ, Vasconcelos MH, Vechetti IJ Jr, Veit TD, Vella LJ, Velot É, Verweij FJ, Vestad B, Viñas JL, Visnovitz T, Vukman KV, Wahlgren J, Watson DC, Wauben MH, Weaver A, Webber JP, Weber V, Wehman AM, Weiss DJ, Welsh JA, Wendt S, Wheelock AM, Wiener Z, Witte L, Wolfram J, Xagorari A, Xander P, Xu J, Yan X, Yáñez-Mó M, Yin H, Yuana Y, Zappulli V, Zarubova J, Žėkas V, Zhang JY, Zhao Z, Zheng L, Zheutlin AR, Zickler AM, Zimmermann P, Zivkovic AM, Zocco D, Zuba-Surma EK. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J Extracell Vesicles. 2018;7(1):1535750.
He C, Zheng S, Luo Y, Wang B. Exosome Theranostics: Biology and Translational Medicine. Theranostics. 2018;8(1):237–55.
Hessvik NP, Llorente A. Current knowledge on exosome biogenesis and release. Cell Mol Life Sci. 2018;75(2):193–208.
Xu J, Xiao Y, Liu B, Pan S, Liu Q, Shan Y, Li S, Qi Y, Huang Y, Jia L. Exosomal MALAT1 sponges miR-26a/26b to promote the invasion and metastasis of colorectal cancer via FUT4 enhanced fucosylation and PI3K/Akt pathway. J Exp Clin Cancer Res. 2020;39(1):54.
Xing X, Han S, Cheng G, Ni Y, Li Z, Li Z. Proteomic Analysis of Exosomes from Adipose-Derived Mesenchymal Stem Cells: A Novel Therapeutic Strategy for Tissue Injury. Biomed Res Int. 2020;2020:6094562.
Zhou X, Xie F, Wang L, Zhang L, Zhang S, Fang M, Zhou F. The function and clinical application of extracellular vesicles in innate immune regulation. Cell Mol Immunol. 2020;17(4):323–34.
Haney MJ, Klyachko NL, Zhao Y, Gupta R, Plotnikova EG, He Z, Patel T, Piroyan A, Sokolsky M, Kabanov AV, Batrakova EV. Exosomes as drug delivery vehicles for Parkinson’s disease therapy. J Control Release. 2015;207:18–30.
Qiao L, Hu S, Huang K, Su T, Li Z, Vandergriff A, Cores J, Dinh PU, Allen T, Shen D, Liang H, Li Y, Cheng K. Tumor cell-derived exosomes home to their cells of origin and can be used as Trojan horses to deliver cancer drugs. Theranostics. 2020;10(8):3474–87.
Allahverdiyev AM, Parlar E, Dinparvar S, Bagirova M, Abamor ES. Current aspects in treatment of breast cancer based of nanodrug delivery systems and future prospects. Artif Cells Nanomed Biotechnol. 2018;46(sup3):S755–62.
Zhuang M, Du D, Pu L, Song H, Deng M, Long Q, Yin X, Wang Y, Rao L. SPION-Decorated Exosome Delivered BAY55–9837 Targeting the Pancreas through Magnetism to Improve the Blood GLC Response. Small. 2019;15(52):e1903135.
Li P, Kaslan M, Lee SH, Yao J, Gao Z. Progress in Exosome Isolation Techniques. Theranostics. 2017;7(3):789–804.
Purushothaman A. Exosomes from cell culture-conditioned medium: isolation by ultracentrifugation and characterization. Methods Mol Biol. 2019;1952:233–44.
Momen-Heravi F. Isolation of Extracellular Vesicles by Ultracentrifugation. Methods Mol Biol. 2017;1660:25–32.
Admyre C, Johansson SM, Qazi KR, Filén JJ, Lahesmaa R, Norman M, Neve EP, Scheynius A, Gabrielsson S. Exosomes with immune modulatory features are present in human breast milk. J Immunol. 2007;179(3):1969–78.
Van Deun J, Mestdagh P, Sormunen R, Cocquyt V, Vermaelen K, Vandesompele J, Bracke M, De Wever O, Hendrix A. The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. J Extracell Vesicles. 2014;3(1):24858.
Kowal J, Arras G, Colombo M, Jouve M, Morath JP, Primdal-Bengtson B, Dingli F, Loew D, Tkach M, Théry C. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc Natl Acad Sci U S A. 2016;113(8):E968-977.
Cantin R, Diou J, Bélanger D, Tremblay AM, Gilbert C. Discrimination between exosomes and HIV-1: purification of both vesicles from cell-free supernatants. J Immunol Methods. 2008;338(1–2):21–30.
Zhang Y, Hao Z, Wang P, Xia Y, Wu J, Xia D, Fang S, Xu S. Exosomes from human umbilical cord mesenchymal stem cells enhance fracture healing through HIF-1alpha-mediated promotion of angiogenesis in a rat model of stabilized fracture. Cell Prolif. 2019;52(2):e12570.
He L, Zhu D, Wang J, Wu X. A highly efficient method for isolating urinary exosomes. Int J Mol Med. 2019;43(1):83–90.
Khushman M, Bhardwaj A, Patel GK, Laurini JA, Roveda K, Tan MC, Patton MC, Singh S, Taylor W, Singh AP. Exosomal Markers (CD63 and CD9) Expression Pattern Using Immunohistochemistry in Resected Malignant and Nonmalignant Pancreatic Specimens. Pancreas. 2017;46(6):782–8.
Sun Z, Wang L, Dong L, Wang X. Emerging role of exosome signalling in maintaining cancer stem cell dynamic equilibrium. J Cell Mol Med. 2018;22(8):3719–28.
Hannafon BN, Trigoso YD, Calloway CL, Zhao YD, Lum DH, Welm AL, Zhao ZJ, Blick KE, Dooley WC, Ding WQ. Plasma exosome microRNAs are indicative of breast cancer. Breast Cancer Res. 2016;18(1):90.
Sharma P, Ludwig S, Muller L, Hong CS, Kirkwood JM, Ferrone S, Whiteside TL. Immunoaffinity-based isolation of melanoma cell-derived exosomes from plasma of patients with melanoma. J Extracell Vesicles. 2018;7(1):1435138.
Ludwig N, Razzo BM, Yerneni SS, Whiteside TL. Optimization of cell culture conditions for exosome isolation using mini-size exclusion chromatography (mini-SEC). Exp Cell Res. 2019;378(2):149–57.
Weng Y, Sui Z, Shan Y, Hu Y, Chen Y, Zhang L, Zhang Y. Effective isolation of exosomes with polyethylene glycol from cell culture supernatant for in-depth proteome profiling. Analyst. 2016;141(15):4640–6.
Macias M, Rebmann V, Mateos B, Varo N, Perez-Gracia JL, Alegre E, Gonzalez A. Comparison of six commercial serum exosome isolation methods suitable for clinical laboratories. Effect in cytokine analysis. Clin Chem Lab Med. 2019;57(10):1539–45.
Ayala-Mar S, Perez-Gonzalez VH, Mata-Gomez MA, Gallo-Villanueva RC, Gonzalez-Valdez J. Electrokinetically Driven Exosome Separation and Concentration Using Dielectrophoretic-Enhanced PDMS-Based Microfluidics. Anal Chem. 2019;91(23):14975–82.
Takov K, Yellon DM, Davidson SM. Comparison of small extracellular vesicles isolated from plasma by ultracentrifugation or size-exclusion chromatography: yield, purity and functional potential. J Extracell Vesicles. 2019;8(1):1560809.
An M, Wu J, Zhu J, Lubman DM. Comparison of an Optimized Ultracentrifugation Method versus Size-Exclusion Chromatography for Isolation of Exosomes from Human Serum. J Proteome Res. 2018;17(10):3599–605.
Tauro BJ, Greening DW, Mathias RA, Ji H, Mathivanan S, Scott AM, Simpson RJ. Comparison of ultracentrifugation, density gradient separation, and immunoaffinity capture methods for isolating human colon cancer cell line LIM1863-derived exosomes. Methods. 2012;56(2):293–304.
Shu S, Yang Y, Allen CL, Hurley E, Tung KH, Minderman H, Wu Y, Ernstoff MS. Purity and yield of melanoma exosomes are dependent on isolation method. J Extracell Vesicles. 2020;9(1):1692401.
Théry C, Amigorena S, Raposo G, Clayton A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol. 2006;30(1):3–22.
Que ZJ, Luo B, Wang CT, Qian FF, Jiang Y, Li Y, Han XH, Li HG, Liu JX, Tian JH. Proteomics analysis of tumor exosomes reveals vital pathways of Jinfukang inhibiting circulating tumor cells metastasis in lung cancer. J Ethnopharmacol. 2020;256:112802.
Abbastabar M, Sarfi M, Golestani A, Karimi A, Pourmand G, Khalili E. Tumor-derived urinary exosomal long non-coding RNAs as diagnostic biomarkers for bladder cancer. EXCLI J. 2020;19:301–10.
Ramanathan S, Shenoda BB, Lin Z, Alexander GM, Huppert A, Sacan A, Ajit SK. Inflammation potentiates miR-939 expression and packaging into small extracellular vesicles. J Extracell Vesicles. 2019;8(1):1650595.
Zhao Z, McGill J, Gamero-Kubota P, He M. Microfluidic on-demand engineering of exosomes towards cancer immunotherapy. Lab Chip. 2019;19(10):1877–86.
Zhu J, Liu B, Wang Z, Wang D, Ni H, Zhang L, Wang Y. Exosomes from nicotine-stimulated macrophages accelerate atherosclerosis through miR-21-3p/PTEN-mediated VSMC migration and proliferation. Theranostics. 2019;9(23):6901–19.
Lotvall J, Hill AF, Hochberg F, Buzas EI, Di Vizio D, Gardiner C, Gho YS, Kurochkin IV, Mathivanan S, Quesenberry P, Sahoo S, Tahara H, Wauben MH, Witwer KW, Thery C. Minimal experimental requirements for definition of extracellular vesicles and their functions: a position statement from the International Society for Extracellular Vesicles. J Extracell Vesicles. 2014;3:26913.
Lobb RJ, Becker M, Wen SW, Wong CS, Wiegmans AP, Leimgruber A, Moller A. Optimized exosome isolation protocol for cell culture supernatant and human plasma. J Extracell Vesicles. 2015;4:27031.
Soares Martins T, Catita J, Martins Rosa I, ABdCES O, Henriques AG. Exosome isolation from distinct biofluids using precipitation and column-based approaches. PLoS One. 2018;13(6):e0198820.
Yamashita T, Takahashi Y, Nishikawa M, Takakura Y. Effect of exosome isolation methods on physicochemical properties of exosomes and clearance of exosomes from the blood circulation. Eur J Pharm Biopharm. 2016;98:1–8.
Bari E, Perteghella S, Catenacci L, Sorlini M, Croce S, Mantelli M, Avanzini MA, Sorrenti M, Torre ML. Freeze-dried and GMP-compliant pharmaceuticals containing exosomes for acellular mesenchymal stromal cell immunomodulant therapy. Nanomedicine (Lond). 2019;14(6):753–65.
Cheruvanky A, Zhou H, Pisitkun T, Kopp JB, Knepper MA, Yuen PS, Star RA. Rapid isolation of urinary exosomal biomarkers using a nanomembrane ultrafiltration concentrator. Am J Physiol Renal Physiol. 2007;292(5):F1657-1661.
Baranyai T, Herczeg K, Onodi Z, Voszka I, Modos K, Marton N, Nagy G, Mager I, Wood MJ, El Andaloussi S, Palinkas Z, Kumar V, Nagy P, Kittel A, Buzas EI, Ferdinandy P, Giricz Z. Isolation of Exosomes from Blood Plasma: Qualitative and Quantitative Comparison of Ultracentrifugation and Size Exclusion Chromatography Methods. PLoS One. 2015;10(12):e0145686.
Stranska R, Gysbrechts L, Wouters J, Vermeersch P, Bloch K, Dierickx D, Andrei G, Snoeck R. Comparison of membrane affinity-based method with size-exclusion chromatography for isolation of exosome-like vesicles from human plasma. J Transl Med. 2018;16(1):1.
Greening DW, Xu R, Ji H, Tauro BJ, Simpson RJ. A protocol for exosome isolation and characterization: evaluation of ultracentrifugation, density-gradient separation, and immunoaffinity capture methods. Methods Mol Biol. 2015;1295:179–209.
Li M, Lou D, Chen J, Shi K, Wang Y, Zhu Q, Liu F, Zhang Y. Deep dive on the proteome of salivary extracellular vesicles: comparison between ultracentrifugation and polymer-based precipitation isolation. Anal Bioanal Chem. 2021;413(2):365–75.
Milane L, Singh A, Mattheolabakis G, Suresh M, Amiji MM. Exosome mediated communication within the tumor microenvironment. J Control Release. 2015;219:278–94.
Skotland T, Sagini K, Sandvig K, Llorente A. An emerging focus on lipids in extracellular vesicles. Adv Drug Deliv Rev. 2020;159:308–21.
Yuan T, Huang X, Woodcock M, Du M, Dittmar R, Wang Y, Tsai S, Kohli M, Boardman L, Patel T, Wang L. Plasma extracellular RNA profiles in healthy and cancer patients. Sci Rep. 2016;6:19413.
Mashouri L, Yousefi H, Aref AR, Ahadi AM, Molaei F, Alahari SK. Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer. 2019;18(1):75.
Mathivanan S, Fahner CJ, Reid GE, Simpson RJ. ExoCarta 2012: database of exosomal proteins. RNA and lipids Nucleic Acids Res. 2012;40(Database issue):D1241-1244.
Pathan M, Fonseka P, Chitti SV, Kang T, Sanwlani R, Van Deun J, Hendrix A, Mathivanan S. Vesiclepedia 2019: a compendium of RNA, proteins, lipids and metabolites in extracellular vesicles. Nucleic Acids Res. 2019;47(D1):D516-d519.
Li S, Li Y, Chen B, Zhao J, Yu S, Tang Y, Zheng Q, Li Y, Wang P, He X, Huang S. exoRBase: a database of circRNA, lncRNA and mRNA in human blood exosomes. Nucleic Acids Res. 2018;46(D1):D106-d112.
Cheng L, Zhang K, Qing Y, Li D, Cui M, Jin P, Xu T. Proteomic and lipidomic analysis of exosomes derived from ovarian cancer cells and ovarian surface epithelial cells. J Ovarian Res. 2020;13(1):9.
Zheng R, Du M, Wang X, Xu W, Liang J, Wang W, Lv Q, Qin C, Chu H, Wang M, Yuan L, Qian J, Zhang Z. Exosome-transmitted long non-coding RNA PTENP1 suppresses bladder cancer progression. Mol Cancer. 2018;17(1):143.
Kitai Y, Kawasaki T, Sueyoshi T, Kobiyama K, Ishii KJ, Zou J, Akira S, Matsuda T, Kawai T. DNA-Containing Exosomes Derived from Cancer Cells Treated with Topotecan Activate a STING-Dependent Pathway and Reinforce Antitumor Immunity. J Immunol. 2017;198(4):1649–59.
Torralba D, Baixauli F, Villarroya-Beltri C, Fernandez-Delgado I, Latorre-Pellicer A, Acin-Perez R, Martin-Cofreces NB, Jaso-Tamame AL, Iborra S, Jorge I, Gonzalez-Aseguinolaza G, Garaude J, Vicente-Manzanares M, Enriquez JA, Mittelbrunn M, Sanchez-Madrid F. Priming of dendritic cells by DNA-containing extracellular vesicles from activated T cells through antigen-driven contacts. Nat Commun. 2018;9(1):2658.
Takahashi A, Okada R, Nagao K, Kawamata Y, Hanyu A, Yoshimoto S, Takasugi M, Watanabe S, Kanemaki MT, Obuse C, Hara E. Exosomes maintain cellular homeostasis by excreting harmful DNA from cells. Nat Commun. 2017;8:15287.
Jeppesen DK, Fenix AM, Franklin JL, Higginbotham JN, Zhang Q, Zimmerman LJ, Liebler DC, Ping J, Liu Q, Evans R, Fissell WH, Patton JG, Rome LH, Burnette DT, Coffey RJ. Reassessment of exosome composition. Cell. 2019;177(2):428–445.e18.
Raiborg C, Stenmark H. The ESCRT machinery in endosomal sorting of ubiquitylated membrane proteins. Nature. 2009;458(7237):445–52.
Li SP, Lin ZX, Jiang XY, Yu XY. Exosomal cargo-loading and synthetic exosome-mimics as potential therapeutic tools. Acta Pharmacol Sin. 2018;39(4):542–51.
Xiao J, Pan Y, Li XH, Yang XY, Feng YL, Tan HH, Jiang L, Feng J, Yu XY. Cardiac progenitor cell-derived exosomes prevent cardiomyocytes apoptosis through exosomal miR-21 by targeting PDCD4. Cell Death Dis. 2016;7(6):e2277.
Zhang X, Shi H, Yuan X, Jiang P, Qian H, Xu W. Tumor-derived exosomes induce N2 polarization of neutrophils to promote gastric cancer cell migration. Mol Cancer. 2018;17(1):146.
Choi H, Kim Y, Mirzaaghasi A, Heo J, Kim YN, Shin JH, Kim S, Kim NH, Cho ES, In Yook J, Yoo TH, Song E, Kim P, Shin EC, Chung K, Choi K, Choi C. Exosome-based delivery of super-repressor IκBα relieves sepsis-associated organ damage and mortality. Sci Adv. 2020;6(15):eaaz6980.
Escrevente C, Keller S, Altevogt P, Costa J. Interaction and uptake of exosomes by ovarian cancer cells. BMC Cancer. 2011;11:108.
Kanchanapally R, Deshmukh SK, Chavva SR, Tyagi N, Srivastava SK, Patel GK, Singh AP, Singh S. Drug-loaded exosomal preparations from different cell types exhibit distinctive loading capability, yield, and antitumor efficacies: a comparative analysis. Int J Nanomedicine. 2019;14:531–41.
Melzer C, Rehn V, Yang Y, Bahre H, von der Ohe J, Hass R. Taxol-loaded MSC-derived exosomes provide a therapeutic vehicle to target metastatic breast cancer and other carcinoma cells. Cancers (Basel). 2019;11(6):798.
Koritzinsky EH, Street JM, Star RA, Yuen PS. Quantification of Exosomes. J Cell Physiol. 2017;232(7):1587–90.
Wiklander OP, Nordin JZ, O’Loughlin A, Gustafsson Y, Corso G, Mager I, Vader P, Lee Y, Sork H, Seow Y, Heldring N, Alvarez-Erviti L, Smith CI, Le Blanc K, Macchiarini P, Jungebluth P, Wood MJ, Andaloussi SE. Extracellular vesicle in vivo biodistribution is determined by cell source, route of administration and targeting. J Extracell Vesicles. 2015;4:26316.
Song H, Li X, Zhao Z, Qian J, Wang Y, Cui J, Weng W, Cao L, Chen X, Hu Y, Su J. Reversal of Osteoporotic Activity by Endothelial Cell-Secreted Bone Targeting and Biocompatible Exosomes. Nano Lett. 2019;19(5):3040–8.
Tian Y, Li S, Song J, Ji T, Zhu M, Anderson GJ, Wei J, Nie G. A doxorubicin delivery platform using engineered natural membrane vesicle exosomes for targeted tumor therapy. Biomaterials. 2014;35(7):2383–90.
Zhang H, Wang Y, Bai M, Wang J, Zhu K, Liu R, Ge S, Li J, Ning T, Deng T, Fan Q, Li H, Sun W, Ying G, Ba Y. Exosomes serve as nanoparticles to suppress tumor growth and angiogenesis in gastric cancer by delivering hepatocyte growth factor siRNA. Cancer Sci. 2018;109(3):629–41.
Kang JY, Park H, Kim H, Mun D, Park H, Yun N, Joung B. Human peripheral bloodderived exosomes for microRNA delivery. Int J Mol Med. 2019;43(6):2319–28.
Jeong K, Yu YJ, You JY, Rhee WJ, Kim JA. Exosome-mediated microRNA-497 delivery for anti-cancer therapy in a microfluidic 3D lung cancer model. Lab Chip. 2020;20(3):548–57.
Didiot MC, Hall LM, Coles AH, Haraszti RA, Godinho BM, Chase K, Sapp E, Ly S, Alterman JF, Hassler MR, Echeverria D, Raj L, Morrissey DV, DiFiglia M, Aronin N, Khvorova A. Exosome-mediated Delivery of Hydrophobically Modified siRNA for Huntingtin mRNA Silencing. Mol Ther. 2016;24(10):1836–47.
Gomari H, Forouzandeh Moghadam M, Soleimani M. Targeted cancer therapy using engineered exosome as a natural drug delivery vehicle. Onco Targets Ther. 2018;11:5753–62.
Kalimuthu S, Gangadaran P, Rajendran RL, Zhu L, Oh JM, Lee HW, Gopal A, Baek SH, Jeong SY, Lee SW, Lee J, Ahn BC. A New Approach for Loading Anticancer Drugs Into Mesenchymal Stem Cell-Derived Exosome Mimetics for Cancer Therapy. Front Pharmacol. 2018;9:1116.
Yim N, Choi C. Extracellular vesicles as novel carriers for therapeutic molecules. BMB Rep. 2016;49(11):585–6.
Yim N, Ryu SW, Choi K, Lee KR, Lee S, Choi H, Kim J, Shaker MR, Sun W, Park JH, Kim D, Heo WD, Choi C. Exosome engineering for efficient intracellular delivery of soluble proteins using optically reversible protein-protein interaction module. Nat Commun. 2016;7:12277.
Sterzenbach U, Putz U, Low LH, Silke J, Tan SS, Howitt J. Engineered Exosomes as Vehicles for Biologically Active Proteins. Mol Ther. 2017;25(6):1269–78.
Aqil F, Munagala R, Jeyabalan J, Agrawal AK, Gupta R. Exosomes for the Enhanced Tissue Bioavailability and Efficacy of Curcumin. AAPS J. 2017;19(6):1691–702.
Fuhrmann G, Serio A, Mazo M, Nair R, Stevens MM. Active loading into extracellular vesicles significantly improves the cellular uptake and photodynamic effect of porphyrins. J Control Release. 2015;205:35–44.
Pomatto MAC, Bussolati B, D’Antico S, Ghiotto S, Tetta C, Brizzi MF, Camussi G. Improved Loading of Plasma-Derived Extracellular Vesicles to Encapsulate Antitumor miRNAs. Mol Ther Methods Clin Dev. 2019;13:133–44.
Wang P, Wang H, Huang Q, Peng C, Yao L, Chen H, Qiu Z, Wu Y, Wang L, Chen W. Exosomes from M1-Polarized Macrophages Enhance Paclitaxel Antitumor Activity by Activating Macrophages-Mediated Inflammation. Theranostics. 2019;9(6):1714–27.
Wei H, Chen J, Wang S, Fu F, Zhu X, Wu C, Liu Z, Zhong G, Lin J. A Nanodrug Consisting Of Doxorubicin And Exosome Derived From Mesenchymal Stem Cells For Osteosarcoma Treatment In Vitro. Int J Nanomedicine. 2019;14:8603–10.
Kim MS, Haney MJ, Zhao Y, Mahajan V, Deygen I, Klyachko NL, Inskoe E, Piroyan A, Sokolsky M, Okolie O, Hingtgen SD, Kabanov AV, Batrakova EV. Development of exosome-encapsulated paclitaxel to overcome MDR in cancer cells. Nanomedicine. 2016;12(3):655–64.
Niu F, Hu D, Gu F, Du Y, Zhang B, Ma S, Pan W. Preparation of ultra-long stable ovalbumin/sodium carboxymethylcellulose nanoparticle and loading properties of curcumin. Carbohydr Polym. 2021;271:118451.
Liu Y, Yang G, Jin S, Xu L, Zhao CX. Development of High-Drug-Loading Nanoparticles. ChemPlusChem. 2020;85(9):2143–57.
Yuan D, Zhao Y, Banks WA, Bullock KM, Haney M, Batrakova E, Kabanov AV. Macrophage exosomes as natural nanocarriers for protein delivery to inflamed brain. Biomaterials. 2017;142:1–12.
Richter M, Vader P, Fuhrmann G. Approaches to surface engineering of extracellular vesicles. Adv Drug Deliv Rev. 2021;173:416–26.
Alvarez-Erviti L, Seow Y, Yin H, Betts C, Lakhal S, Wood MJ. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat Biotechnol. 2011;29(4):341–5.
Bellavia D, Raimondo S, Calabrese G, Forte S, Cristaldi M, Patinella A, Memeo L, Manno M, Raccosta S, Diana P, Cirrincione G, Giavaresi G, Monteleone F, Fontana S, De Leo G, Alessandro R. Interleukin 3- receptor targeted exosomes inhibit in vitro and in vivo Chronic Myelogenous Leukemia cell growth. Theranostics. 2017;7(5):1333–45.
Tian T, Zhang HX, He CP, Fan S, Zhu YL, Qi C, Huang NP, Xiao ZD, Lu ZH, Tannous BA, Gao J. Surface functionalized exosomes as targeted drug delivery vehicles for cerebral ischemia therapy. Biomaterials. 2018;150:137–49.
Qi H, Liu C, Long L, Ren Y, Zhang S, Chang X, Qian X, Jia H, Zhao J, Sun J, Hou X, Yuan X, Kang C. Blood Exosomes Endowed with Magnetic and Targeting Properties for Cancer Therapy. ACS Nano. 2016;10(3):3323–33.
Sun D, Zhuang X, Xiang X, Liu Y, Zhang S, Liu C, Barnes S, Grizzle W, Miller D, Zhang HG. A novel nanoparticle drug delivery system: the anti-inflammatory activity of curcumin is enhanced when encapsulated in exosomes. Mol Ther. 2010;18(9):1606–14.
Han D, Wang K, Zhang T, Gao GC, Xu H. Natural killer cell-derived exosome-entrapped paclitaxel can enhance its anti-tumor effect. Eur Rev Med Pharmacol Sci. 2020;24(10):5703–13.
Zhang Q, Zhang H, Ning T, Liu D, Deng T, Liu R, Bai M, Zhu K, Li J, Fan Q, Ying G, Ba Y. Exosome-Delivered c-Met siRNA Could Reverse Chemoresistance to Cisplatin in Gastric Cancer. Int J Nanomedicine. 2020;15:2323–35.
Lin X, Wang L, Zhao L, Zhu Z, Chen T, Chen S, Tao Y, Zeng T, Zhong Y, Sun H, Wang Z, Zheng W, Zhang Y, Wu W, Nan K, Chen T. Curcumin micelles suppress gastric tumor cell growth by upregulating ROS generation, disrupting redox equilibrium and affecting mitochondrial bioenergetics. Food Funct. 2020;11(5):4146–59.
Wu X, Chen X, Zhou Q, Li P, Yu B, Li J, Qu Y, Yan J, Yu Y, Yan M, Zhu Z, Liu B, Su L. Hepatocyte growth factor activates tumor stromal fibroblasts to promote tumorigenesis in gastric cancer. Cancer Lett. 2013;335(1):128–35.
Pan S, Pei L, Zhang A, Zhang Y, Zhang C, Huang M, Huang Z, Liu B, Wang L, Ma L, Zhang Q, Cui D. Passion fruit-like exosome-PMA/Au-BSA@Ce6 nanovehicles for real-time fluorescence imaging and enhanced targeted photodynamic therapy with deep penetration and superior retention behavior in tumor. Biomaterials. 2020;230:119606.
Xin L, Yuan YW, Liu C, Zhou LQ, Liu L, Zhou Q, Li SH. Preparation of Internalizing RGD-Modified Recombinant Methioninase Exosome Active Targeting Vector and Antitumor Effect Evaluation. Dig Dis Sci. 2021;66(4):1045–53.
Iyikesici MS. Feasibility study of metabolically supported chemotherapy with weekly carboplatin/paclitaxel combined with ketogenic diet, hyperthermia and hyperbaric oxygen therapy in metastatic non-small cell lung cancer. Int J Hyperthermia. 2019;36(1):446–55.
Kenmotsu H, Yoh K, Mori K, Ono A, Baba T, Fujiwara Y, Yamaguchi O, Ko R, Okamoto H, Yamamoto N, Ninomiya T, Ogura T, Kato T. Phase II study of nab-paclitaxel + carboplatin for patients with non-small-cell lung cancer and interstitial lung disease. Cancer Sci. 2019;110(12):3738–45.
Nie H, Xie X, Zhang D, Zhou Y, Li B, Li F, Li F, Cheng Y, Mei H, Meng H, Jia L. Use of lung-specific exosomes for miRNA-126 delivery in non-small cell lung cancer. Nanoscale. 2020;12(2):877–87.
Kim MS, Haney MJ, Zhao Y, Yuan D, Deygen I, Klyachko NL, Kabanov AV, Batrakova EV. Engineering macrophage-derived exosomes for targeted paclitaxel delivery to pulmonary metastases: in vitro and in vivo evaluations. Nanomedicine. 2018;14(1):195–204.
Aqil F, Munagala R, Jeyabalan J, Agrawal AK, Kyakulaga AH, Wilcher SA, Gupta RC. Milk exosomes - Natural nanoparticles for siRNA delivery. Cancer Lett. 2019;449:186–95.
Spuch C, Navarro C. Liposomes for Targeted Delivery of Active Agents against Neurodegenerative Diseases (Alzheimer’s Disease and Parkinson’s Disease). J Drug Deliv. 2011;2011:469679.
Knudsen KB, Northeved H, Kumar PE, Permin A, Gjetting T, Andresen TL, Larsen S, Wegener KM, Lykkesfeldt J, Jantzen K, Loft S, Møller P, Roursgaard M. In vivo toxicity of cationic micelles and liposomes. Nanomedicine. 2015;11(2):467–77.
Zhuang X, Xiang X, Grizzle W, Sun D, Zhang S, Axtell RC, Ju S, Mu J, Zhang L, Steinman L, Miller D, Zhang HG. Treatment of brain inflammatory diseases by delivering exosome encapsulated anti-inflammatory drugs from the nasal region to the brain. Mol Ther. 2011;19(10):1769–79.
Wang H, Sui H, Zheng Y, Jiang Y, Shi Y, Liang J, Zhao L. Curcumin-primed exosomes potently ameliorate cognitive function in AD mice by inhibiting hyperphosphorylation of the Tau protein through the AKT/GSK-3beta pathway. Nanoscale. 2019;11(15):7481–96.
Yang T, Martin P, Fogarty B, Brown A, Schurman K, Phipps R, Yin VP, Lockman P, Bai S. Exosome delivered anticancer drugs across the blood-brain barrier for brain cancer therapy in Danio rerio. Pharm Res. 2015;32(6):2003–14.
Qu M, Lin Q, Huang L, Fu Y, Wang L, He S, Fu Y, Yang S, Zhang Z, Zhang L, Sun X. Dopamine-loaded blood exosomes targeted to brain for better treatment of Parkinson’s disease. J Control Release. 2018;287:156–66.
Kuroda H, Tachikawa M, Yagi Y, Umetsu M, Nurdin A, Miyauchi E, Watanabe M, Uchida Y, Terasaki T. Cluster of Differentiation 46 Is the Major Receptor in Human Blood-Brain Barrier Endothelial Cells for Uptake of Exosomes Derived from Brain-Metastatic Melanoma Cells (SK-Mel-28). Mol Pharm. 2019;16(1):292–304.
Morad G, Carman CV, Hagedorn EJ, Perlin JR, Zon LI, Mustafaoglu N, Park TE, Ingber DE, Daisy CC, Moses MA. Tumor-Derived Extracellular Vesicles Breach the Intact Blood-Brain Barrier via Transcytosis. ACS Nano. 2019;13(12):13853–65.
Jia G, Han Y, An Y, Ding Y, He C, Wang X, Tang Q. NRP-1 targeted and cargo-loaded exosomes facilitate simultaneous imaging and therapy of glioma in vitro and in vivo. Biomaterials. 2018;178:302–16.
Zhu Q, Ling X, Yang Y, Zhang J, Li Q, Niu X, Hu G, Chen B, Li H, Wang Y, Deng Z. Embryonic Stem Cells-Derived Exosomes Endowed with Targeting Properties as Chemotherapeutics Delivery Vehicles for Glioblastoma Therapy. Adv Sci (Weinh). 2019;6(6):1801899.
Aqil F, Jeyabalan J, Agrawal AK, Kyakulaga AH, Munagala R, Parker L, Gupta RC. Exosomal delivery of berry anthocyanidins for the management of ovarian cancer. Food Funct. 2017;8(11):4100–7.
Gong C, Tian J, Wang Z, Gao Y, Wu X, Ding X, Qiang L, Li G, Han Z, Yuan Y, Gao S. Functional exosome-mediated co-delivery of doxorubicin and hydrophobically modified microRNA 159 for triple-negative breast cancer therapy. J Nanobiotechnology. 2019;17(1):93.
Yu M, Gai C, Li Z, Ding D, Zheng J, Zhang W, Lv S, Li W. Targeted exosome-encapsulated erastin induced ferroptosis in triple negative breast cancer cells. Cancer Sci. 2019;110(10):3173–82.
Kamerkar S, LeBleu VS, Sugimoto H, Yang S, Ruivo CF, Melo SA, Lee JJ, Kalluri R. Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature. 2017;546(7659):498–503.
Saari H, Lazaro-Ibanez E, Viitala T, Vuorimaa-Laukkanen E, Siljander P, Yliperttula M. Microvesicle- and exosome-mediated drug delivery enhances the cytotoxicity of Paclitaxel in autologous prostate cancer cells. J Control Release. 2015;220(Pt B):727–37.
Li Y, Gao Y, Gong C, Wang Z, Xia Q, Gu F, Hu C, Zhang L, Guo H, Gao S. A33 antibody-functionalized exosomes for targeted delivery of doxorubicin against colorectal cancer. Nanomedicine. 2018;14(7):1973–85.
Robbins PD, Morelli AE. Regulation of immune responses by extracellular vesicles. Nat Rev Immunol. 2014;14(3):195–208.
Maeda H, Nakamura H, Fang J. The EPR effect for macromolecular drug delivery to solid tumors: Improvement of tumor uptake, lowering of systemic toxicity, and distinct tumor imaging in vivo. Adv Drug Deliv Rev. 2013;65(1):71–9.
Xin Y, Huang M, Guo WW, Huang Q, Zhang LZ, Jiang G. Nano-based delivery of RNAi in cancer therapy. Mol Cancer. 2017;16(1):134.
Meng W, He C, Hao Y, Wang L, Li L, Zhu G. Prospects and challenges of extracellular vesicle-based drug delivery system: considering cell source. Drug Deliv. 2020;27(1):585–98.
Lara P, Palma-Florez S, Salas-Huenuleo E, Polakovicova I, Guerrero S, Lobos-Gonzalez L, Campos A, Munoz L, Jorquera-Cordero C, Varas-Godoy M, Cancino J, Arias E, Villegas J, Cruz LJ, Albericio F, Araya E, Corvalan AH, Quest AFG, Kogan MJ. Gold nanoparticle based double-labeling of melanoma extracellular vesicles to determine the specificity of uptake by cells and preferential accumulation in small metastatic lung tumors. J Nanobiotechnology. 2020;18(1):20.
Wan Z, Zhao L, Lu F, Gao X, Dong Y, Zhao Y, Wei M, Yang G, Xing C, Liu L. Mononuclear phagocyte system blockade improves therapeutic exosome delivery to the myocardium. Theranostics. 2020;10(1):218–30.
Yang Z, Shi J, Xie J, Wang Y, Sun J, Liu T, Zhao Y, Zhao X, Wang X, Ma Y, Malkoc V, Chiang C, Deng W, Chen Y, Fu Y, Kwak KJ, Fan Y, Kang C, Yin C, Rhee J, Bertani P, Otero J, Lu W, Yun K, Lee AS, Jiang W, Teng L, Kim BYS, Lee LJ. Large-scale generation of functional mRNA-encapsulating exosomes via cellular nanoporation. Nat Biomed Eng. 2020;4(1):69–83.
Jhan YY, Prasca-Chamorro D, Palou Zuniga G, Moore DM, Arun Kumar S, Gaharwar AK, Bishop CJ. Engineered extracellular vesicles with synthetic lipids via membrane fusion to establish efficient gene delivery. Int J Pharm. 2020;573:118802.
Jang SC, Kim OY, Yoon CM, Choi DS, Roh TY, Park J, Nilsson J, Lötvall J, Kim YK, Gho YS. Bioinspired exosome-mimetic nanovesicles for targeted delivery of chemotherapeutics to malignant tumors. ACS Nano. 2013;7(9):7698–710.
Lunavat TR, Jang SC, Nilsson L, Park HT, Repiska G, Lasser C, Nilsson JA, Gho YS, Lotvall J. RNAi delivery by exosome-mimetic nanovesicles - Implications for targeting c-Myc in cancer. Biomaterials. 2016;102:231–8.
Wang S, Li F, Ye T, Wang J, Lyu C, Qing S, Ding Z, Gao X, Jia R, Yu D, Ren J, Wei W, Ma G. Macrophage-tumor chimeric exosomes accumulate in lymph node and tumor to activate the immune response and the tumor microenvironment. Sci Transl Med. 2021;13(615):eabb6981.
Zhang KL, Wang YJ, Sun J, Zhou J, Xing C, Huang G, Li J, Yang H. Artificial chimeric exosomes for anti-phagocytosis and targeted cancer therapy. Chem Sci. 2019;10(5):1555–61.
Funding
This work was supported in part by the National Natural Science Foundation of China (No. 31900976 and No. 32071360 to H.C.), by Research Starting Funding of South China University of Technology (D6181910, D6201880, K5180910, and K5204120 to Y.D.), by Research Agreement between South China University of Technology and Guangzhou First People’s Hospital (D9194290 to Y.D., ky13010037 to H.C.).
Author information
Authors and Affiliations
Contributions
YD and HC designed and supervised the whole work. WZ wrote the paper, and YD, HC and ZW revised it. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All co-authors have no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zeng, W., Wen, Z., Chen, H. et al. Exosomes as Carriers for Drug Delivery in Cancer Therapy. Pharm Res 40, 873–887 (2023). https://doi.org/10.1007/s11095-022-03224-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-022-03224-y