Abstract
Purpose
Cyclocreatine, a creatine analog, is a candidate drug for treating patients with cerebral creatine deficiency syndromes (CCDSs) caused by creatine transporter (CRT, SLC6A8) deficiency, which reduces brain creatine level. The purpose of this study was to clarify the characteristics of cyclocreatine transport in HEK293 cells, which highly express endogenous CRT, in hCMEC/D3 cells, a human blood-brain barrier (BBB) model, and in CCDSs patient-derived fibroblasts with CRT mutations.
Methods
Cells were incubated at 37°C with [14C]cyclocreatine (9 μM) and [14C]creatine (9 μM) for specified periods of times in the presence or absence of inhibitors, while the siRNAs were transfected by lipofection. Protein expression and mRNA expression were quantified using targeted proteomics and quantitative PCR, respectively.
Results
[14C]Cyclocreatine was taken up by HEK293 cells in a time-dependent manner, while exhibiting saturable kinetics. The inhibition and siRNA knockdown studies demonstrated that the uptake of [14C]cyclocreatine by both HEK293 and hCMEC/D3 cells was mediated predominantly by CRT as well as [14C]creatine. In addition, uptake of [14C]cyclocreatine and [14C]creatine by the CCDSs patient-derived fibroblasts was found to be largely reduced.
Conclusion
The present study suggests that cyclocreatine is a CRT substrate, where CRT is the predominant contributor to influx of cyclocreatine into the brain at the BBB. Our findings provide vital insights for the purposes of treating CCDSs patients using cyclocreatine.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cerebral creatine deficiency syndromes (CCDSs) are the second most common cause of chromosome X-linked intellectual disability (1), where patients with CCDSs display signs of various neurodevelopmental disorders, including delays in speech, language, seizures, intellectual disability, autistic spectrum disorder, and epilepsy (2,3,4). Creatine plays a vital role in ATP storage by the creatine shuttle in vivo (5). The reduction in brain creatine levels in CCDSs patients is known to be caused by gene mutations in both L-arginine: glycine amidinotransferase (AGMT) and guanidinoacetate methyltransferase (GAMT), which are involved in creatine synthesis in the peripheral tissues and brain (6,7), and creatine transporter (CRT), which is involved in the supply of creatine to the brain from the bloodstream (8). In cases where CCDSs is caused by mutations of AGMT and GAMT, oral creatine supplementation can be effective, where this type of CCDSs is curable (9,10). However, in cases where CCDSs are caused by CRT deficiency, oral creatine supplementation is not effective, as the brain creatine levels have not been elevated (11,12,13). Currently, no effective treatment is available for patients with CCDSs caused by CRT deficiency.
CRT is a membrane protein encoded by the SLC6A8, which mediates the uptake of creatine, while being coupled with Na+ and Cl− (14). The reported number of missense gene mutations or deletions in CRT in CCDSs patients is at least 50 mutations (15). We and others have demonstrated, previously, that gene mutations of CRT in patients with CCDSs can result in the reduction of creatine transport in overexpressed cells (16,17,18). In the brain, CRT is mainly expressed in brain capillary endothelial cells, which form the blood-brain barrier (BBB) (8), as well as in neurons (19,20). We have reported that CRT can be a major supplier of creatine to the brain across the BBB (8). It was also reported that brain creatine levels in vivo were dramatically reduced in CRT knockout mice (21,22,23). Thus, the dysfunction of CRT in the BBB and neurons by missense gene mutations or gene deletion could cause the reduction of creatine in the brain.
A therapeutic strategy for patients with CCDSs caused by gene mutations or deletion in CRT is to supply creatine analogues that can cross the BBB, which allow the uptake into the neurons through the CRT independent transport system. Cyclocreatine (Fig. 1) is a promising creatine analogue for the treatment of patient with CCDSs, since cyclocreatine is able to function as the creatine shuttle in the brain (24), while cyclocreatine improves the cognitive behavior of the neuron-specific CRT knockout mice through oral administration (25). With that, cyclocreatine was detected in the mouse brain after oral administration in the wild type mouse as well as in the neuron-specific CRT knockout mouse (25). Cyclocreatine entry into the brain via BBB was still undetermined, where the uptake by neurons could be involved in terms of passive diffusion or via transporters other than CRT. Lipophilicity (Log n-octanol/Ringer distribution coefficient) of cyclocreatine and creatine were not significantly changed to explain the increase in the passive diffusion rate (−2.25 and − 2.16, respectively, referenced from the Chemistry Dashboard in United States Environmental Protection Agency). Thus, the transport mechanism involved in the cyclocreatine uptake was not fully identified yet.
Cyclocreatine and creatine contain a monocarboxylic acid group, which is normally transported via solute carrier (SLC) transporters (14,26). Hence, we proposed that cyclocreatine transport may be mediated via saturable transporters that include CRT (SLC6A8). In addition, it remains unclear how CRT mutations and deletion can affect the transport of cyclocreatine in patients with CCDSs.
Thus, the purpose of this study was to determine the characteristics of cyclocreatine uptake by HEK293 cells, which expresses highly endogenous CRT; by hCMEC/D3 cells, immortalized human brain capillary endothelial cells, as an in vitro human BBB model; and by CCDSs patient-derived fibroblasts with CRT mutations.
Materials and Methods
Materials
Cyclocreatine, creatine, β-guanidinopropionic acid (GPA), hydrocortisone, ascorbic acid, human basic fibroblast growth factor (bFGF), lithium chloride, Hank’s balanced salt solution (HBSS) buffer, and ammonium bicarbonate were purchased from Sigma-Aldrich (St. Louis, MO, USA). Chemically-defined lipid concentrate was purchased from Thermo Fisher Scientific (Waltham, MA, USA). Sodium deoxycholate, N-lauroylsarcosinate, dithiothreitol, iodoacetamide, trifluoroacetic acid, ethyl acetate and lysyl endopeptidase were purchased from Fujifilm Wako Pure Chemical Corporation (Osaka, Japan). Trypsin was purchased from Promega (Madison, WI, USA).
Fibroblast
Two fibroblasts from two CCDSs patients, as well as a fibroblast (CRT-WT) control sample from a person expressing wild type CRT, were obtained from the Division of Neurology, Kanagawa Children’s Medical Center, Yokohama, Japan. The first fibroblast (CRT-Deletion) was obtained from a CCDSs patient with a large 19 kb deletion that was encompassing exons 5 to 13 from SLC6A8 as well as exons 5 to 8 from the B cell receptor-associated protein (BAP31) gene (27). Another fibroblast (CRT-G561R mutation) was obtained from a CCDSs patient with a missense mutation in exon 12 from the SLC6A8 gene [c.1681G > C;p.Gly561Arg] (28). This study was approved by the Ethics Committees of the Graduate School of Pharmaceutical Sciences, Kumamoto University and the Kanagawa Children’s Medical Center.
Cell Culture
HEK293 cells were cultured in a Dulbecco’s modified Eagle medium (DMEM) high glucose medium (Thermo Fisher Scientific) containing 10% fetal bovine serum (FBS) (Thermo Fisher Scientific) and 1% penicillin-streptomycin (Fujifilm Wako Pure Chemical Corporation) in conditions of 95% air and 5% CO2 at 37°C until confluent. The hCMEC/D3 cells were cultured on collagen type I–coated dishes in endothelial cell basal medium-2 (EBM-2) (Lonza Walker sville, Inc., Waikersville, MD) that was supplemented with 5% FBS, 1.4 μM hydrocortisone, 5 μg/mL ascorbic acid, 1% chemically defined lipid concentrate, 1 ng/mL bFGF, 10 mM 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES) (Dojindo, Kumamoto, Japan), 10 mM lithium chloride, and 1% penicillin-streptomycin in conditions of 95% air and 5% CO2 at 37°C. Fibroblasts derived from the skin were cultured in a DMEM low glucose medium (Thermo Fisher Scientific) containing 10% FBS, 1% penicillin-streptomycin at 37°C in conditions of 95% air and 5% CO2.
[14C]Cyclocreatine and [14C]Creatine Uptake by Cell Lines
Cells were seeded on 24 well plates (Corning, NY, USA) at a density of 1.0 × 105 cells/well (HEK293) or 5.0 × 104 cells/well (fibroblast), which were cultured for 24 h at 37°C. The hCMEC/D3 cells were seeded on collagen type I-coated 24 well plates (CORNING) at a density of 3.5 × 104 cells/well, and then cultured for 24 h at 37°C. Cells were washed twice with HBSS buffer at 37°C, where the uptake was initiated by applying the HBSS buffer containing [14C]cyclocreatine (0.1 μCi, 9 μM) (Moravek, Ink., Mercury Lane Brea, CA) or [14C]creatine (0.1 μCi, 9 μM) (Moravek, Ink.) in the presence or absence of unlabeled cyclocreatine, creatine and GPA. Na+-free buffer was prepared by an equimolar replacement of NaCl, disodium phosphate, and NaHCO3 with choline chloride, dipotassium phosphate and potassium bicarbonate. Cl−-free buffer was prepared by an equimolar replacement of CaCl, KCl, and NaCl with calcium gluconate, potassium gluconate and sodium gluconate. Cells were washed three times with the HBSS buffer at 4°C. Then, the cells were solubilized with a 5 N NaOH solution and neutralized with 5 N HCl. Cell-associated protein was determined using the Micro BCA™ Protein Assay Kit (Thermo Fisher Scientific) with bovine serum albumin as a standard. The radioactivity in each sample was measured by liquid scintillation counting, where the degree of uptake by the cells was estimated by the radioactivity of the sample. This was expressed as a cell/medium ratio per milligram of cellular protein (μL/mg protein) or as a percentage of the control cells.
Knockdown with siRNA
Silencer Select Pre-designed siRNA SLC6A8 (ABI s12988) was used for the targeted silencing of the SLC6A8 gene, which was purchased from Thermo Fisher Scientific. For the negative control, the Silencer™ Negative Control No.1 siRNA (Thermo Fisher Scientific) was used. The cells were transfected with 10 nM siRNA using a 0.1% (v/v) Lipofectamine RNAiMAX (Thermo Fisher Scientific) according to the manufacture’s protocol. Transfection reactions were performed in serum-free OptiMEM (Thermo Fisher Scientific). The cell media was changed to serum containing media 6 h after transfection, when the cells were cultured for 24 h, and then used in the study.
Quantitative Real-Time PCR
Total RNA was isolated from cells at 24 h post-transfection by the RNeasy kit (Qiagen, Inc., Valencia, CA, USA). Single-stranded cDNA was prepared from 1 μg total RNA by ReverTraAce (Toyobo, Osaka, Japan) and an oligo dT primer. CRT and glyceraldehyde-3-phosphate dehydrogenase (GAPDH) mRNAs were quantified by real-time PCR (QuantStudio™ 3, Applied Biosystems, Foster City, CA, USA), where RT-qPCR was performed using a Prime Time qPCR assay (Integrated DNA Technologies, Inc., Coralvile, IA, USA). The forward primer 5’-GATCACCATGGAGGCGTAG-3′, the reverse primer 5’-TGATCGCCCTGGTTGGA-3′, and the probe 5′−/56-FAM/ATGTTCCAG/ZEN/ACATTGATGCTGCCG/3IAbkFQ/−3′ were designed for CRT. The forward primer 5’-ACATCGCTCAGACACCATG-3′, the reverse primer 5’-TGTAGTTGAGGTCAATGAAGGG-3′, and the probe 5′−/56-FAM/AAGGTCGGA/ZEN/GTCAACGGATTTGGTC/3IAbkFQ/−3′ were designed for GAPDH. Calibration curves were prepared using synthetic DNA from CRT and GAPDH ((Integrated DNA Technologies, Inc.). The cellular mRNA content was normalized to GAPDH.
Protein Quantification by Targeted Proteomics
Plasma membrane fractions were extracted from 1 × 107 of HEK293 cells, 1 × 106 of hCMEC/D3 cells or 1 × 107 of fibroblasts using the Plasma Membrane Protein Extraction Kit (BioVision, Milpitas, CA, USA). The fractionation was performed according to the instruction manual with modifications, as described previously (29). Total protein content was quantified using the Micro BCA Protein Assay Kit (Thermo Fisher Scientific). The fractionated protein samples were digested by the phase-transfer surfactant (PTS) method, as previously described (30). Briefly, the protein samples were denatured with 12 mM sodium deoxycholate, 12 mM N-lauroylsarcosinate, and 100 mM Tris-HCl (pH 9.0), which were heated at 95°C for 5 min. After sonication, the samples were reduced with 10 mM dithiothreitol, alkylated with 55 mM iodoacetamide, diluted 5-fold with 50 mM ammonium bicarbonate, and digested with lysyl endopeptidase at room temperature for 3 h, followed by a sequence-grade modified trypsin at 37°C for 16 h. For targeted quantitative proteomics, stable isotope-labeled internal standard peptides were spiked into the samples after trypsin digestion. Samples were acidified with trifluoroacetic acid, while ethyl acetate was added to remove the detergents. The lower phase was evaporated by vacuum centrifugation. Then, the residue was dissolved in 5% acetonitrile/0.1% trifluoroacetic acid, which was desalted by GL-Tip GC and SDB (GC: graphite carbon, SDB: styrene-divinylbenzene polymers) (GL Sciences, Tokyo, Japan). The samples were also desalted and dissolved in 0.1% trifluoroacetic acid, then analyzed by a micro-LC system with a ChromXP C18CL Column (Ekisigent, Redwood City, CA) that was coupled to an electrospray ionization triple-quadrupole mass spectrometer (QTRAP6500; SCIEX, Framingham, MA).
The proteomic analysis was performed according to previous work (31). Briefly, protein expression levels were quantified by the detection of specific peptides from the protein with multiple reaction monitoring (MRM) for targeted proteomics. In targeted proteomics, the product ion peaks from each targeted peptide, where the internal standard peptide corresponds to the targeted peptide is extracted from the MRM data. The peaks corresponding to the target peptides and the internal standard peptides are automatically identified by an in-house software written under contract by Mitsui Knowledge Industry (Tokyo, Japan). The protein expression level was determined as the average value of the peak ratios from 3 or 4 product ions. For CRT protein, the probe peptide and its 4 selected product ions / MRM transitions (internal standard peptide in parentheses) consisted of GDGPVGLGTPGGR (stable isotope-labeled on lysine (K) in the internal standard), with values of 570.3/714.4 (573.3/720.4), 570.3/657.4 (573.3/663.4), 570.3/544.3 (573.3/550.3), 570.3/386.2 (573.3/392.2), respectively. For the Na+/K+ ATPase protein, the probe peptide and its 4 selected product ions / MRM transitions (internal standard peptide in parentheses) consisted of AAVPDAVGK (stable isotope-labeled on arginine (R) in the internal standard), with values of 414.2/685.4 (417.2/691.4), 414.2/586.3 (417.2/592.3), 414.2/489.3 (417.2/495.3), 414.2/374.2 (417.2/380.3), respectively.
Statistical Analysis
Data are represented as the mean ± SEM. A two-tailed, unpaired Student’s t test and one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test were used to identify the significant differences between two or more groups, respectively. Graph Pad Prism 7 statistical software (GraphPad Software Inc. La Jolla, CA, USA) was used in all the analyses.
Results
Characteristics of [14C]Cyclocreatine Uptake by HEK293 Cells
As our previous study showed that [14C]creatine uptake by HEK 293 cells was predominantly mediated by CRT (16), the characteristics of cyclocreatine uptake by HEK293 cells was examined in this study. [14C]cyclocreatine was linearly taken up in a time-dependent manner up to 60 min by HEK293 cells (Fig. 2a), where this uptake exhibited saturable kinetics and a Km of 188 ± 25 μM, Vmax of 273 ± 9 pmol / (mg protein·min) as well as an influx clearance of 1.45 µL / (mg protein·min). The Eadie-Hofstee plot showed that the uptake of cyclocreatine by HEK293 cells was a monophasic kinetics (r2 = 0.909, Fig. 2b inset). In addition, the uptake of [14C]cyclocreatine was reduced in the absence of Na+ and Cl− by 97.4% and 96.5%, respectively (Fig. 2c). These results suggest that the uptake of cyclocreatine by HEK293 cells was predominantly mediated by a Na+ and Cl− dependent saturable transport systems, and not by passive diffusion. The involvement of CRT in the cyclocreatine transport by HEK293 cells was examined through the inhibition study. The uptake of [14C]cyclocreatine by HEK293 cells at 30 min was almost completely suppressed by 96.1% in the presence of 10 mM unlabeled creatine, while in the presence of 10 mM GPA, the CRT-selective inhibitor, was suppressed by 96.6%.
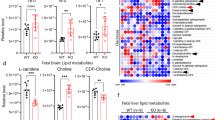
Transport property analysis of [14C]cyclocreatine transport in HEK293 cells. (a) Time-course of cyclocreatine uptake by HEK293 cells. [14C]Cyclocreatine uptake (0.1 μCi, 9 μM) was measured in HEK293 cells at 37°C, and expressed as cell/medium ratio. Each point represents the mean ± SEM (n = 4). (b) Concentration-dependence of cyclocreatine uptake by HEK293 cells exhibited by Michaelis-Menten equation. The inset graph shows the Eadie-Hofstee plot of the same data. The uptake was measured in the presence of the indicated concentrations of unlabeled cyclocreatine for 30 min. Each point represents the mean ± SEM (n = 4). The Km is 188 ± 25 μM and Vmax is 273 ± 9 pmol / (mg protein·min). (c) Effect of inhibitors on creatine transport by HEK293 cells. [14C]Cyclocreatine uptake (0.1 μCi, 9 μM) was measured in the absence of compounds (control), in the presence of either GPA (10 mM) or unlabeled creatine (10 mM), or in the absence of either Na+ or Cl− at 37°C for 30 min. Each column represents the mean ± SEM (n = 4). ****p < 0.0001, significantly different from control condition.
Effect of CRT Knockdown on [14C]Cyclocreatine and [14C]Creatine Uptake by HEK293 Cells
The involvement of CRT in the uptake of cyclocreatine by HEK293 cells was examined by the siRNA knockdown study (Fig. 3). CRT-targeting siRNA suppressed CRT mRNA expression by 70.0% (Fig. 3a) as well as protein expression by 41.6% (Fig. 3b) relative to the control siRNA. The uptake of [14C]cyclocreatine and [14C]creatine by the CRT-targeting siRNA treated HEK293 cells at 30 min was reduced by 33.4% and 37.3%, respectively, relative to the control sample (Fig. 3c-d). In addition, decreased CRT protein expression, cyclocreatine and creatine uptake were comparable in this knockdown condition, indicating that cyclocreatine could be a substrate of CRT.
Effect of CRT knockdown of CRT expression, [14C]cyclocreatine uptake and [14C]creatine uptake by HEK293 cells. (a) Quantitative analysis of mRNA expression using RT-qPCR in HEK293 cells transfected with CRT-targeted siRNA (CRT siRNA) or Silencer™ Negative Control No.1 siRNA (Control siRNA). Each column represents the mean ± SEM (n = 4). (b) The CRT protein expression measured by targeted proteomics in the plasma membrane fraction of HEK293 cells transfected with CRT-targeted siRNA (CRT siRNA) or Silencer™ Negative Control No.1 siRNA (Control siRNA). Each column represents the mean ± SEM (n = 3). The uptake of [14C]cyclocreatine (c) and [14C]creatine (d) by HEK293 cells transfected with CRT-targeted siRNA (CRT siRNA) or Silencer™ Negative Control No.1 siRNA (Control siRNA). The uptake was measured at 30 min. Each column represents the mean ± SEM (n = 4). **p < 0.01, ****p < 0.001 significantly different from control siRNA.
Characteristics of [14C]Cyclocreatine and [14C]Creatine Uptake by hCMEC/D3 Cells
How cyclocreatine is able to cross the human BBB still remains unclear. As shown in Fig. 4, uptake of [14C]cyclocreatine (7.04 μl/mg protein) and [14C]creatine (19.7 μl/mg protein) was observed at 30 min in hCMEC/D3 cells, the human BBB model cells. The uptake of [14C]cyclocreatine and [14C]creatine were reduced in the absence of Na+ by 90.5% and 86.1%, respectively, while in the absence of Cl− by 84.4% and 85.1%, respectively. The uptake of [14C]cyclocreatine and [14C]creatine was also inhibited in the presence of the 10 mM unlabeled cyclocreatine by 89.4% and 88.1%, respectively, while in the presence of 10 mM unlabeled creatine by 90.7% and 92.5%, respectively. These results suggest that cyclocreatine transport shares a Na+ and Cl− dependent saturable transporter of creatine in hCMEC/D3 cells.
Effect of inhibitors on [14C]cyclocreatine and [14C]creatine transport in hCMEC/D3 cells. Effect of inhibitors on [14C]cyclocreatine transport (a) and [14C]creatine transport (b) by hCMEC/D3 cells expressing endogenous CRT. [14C]cyclocreatine uptake (0.1 μCi, 9 μM) and [14C]creatine uptake (0.1 μCi, 9 μM) were measured in the absence of compounds (control), in the presence of either GPA (10 mM), unlabeled creatine (10 mM) or unlabeled cyclocreatine (10 mM), or in the absence of either Na+ or Cl− at 37°C for 30 min. Each column represents the mean ± SEM (n = 4). ****p < 0.0001, significantly different from control condition.
Targeted proteomics have demonstrated that CRT protein was also expressed in hCMEC/D3 cells, while the expression levels of CRT protein was 0.444 fmol/μg protein in the plasma membrane fraction of hCMEC/D3 cells (Table I). The uptake of [14C]cyclocreatine and [14C]creatine by hCMEC/D3 cells at 30 min was almost completely suppressed in the presence of 10 mM GPA by 90.9% and 92.0%, respectively (Fig. 4). These results suggest that CRT predominantly contributes to cyclocreatine and creatine transport in hCMEC/D3 cells.
Effect of CRT Knockdown on [14C]Cyclocreatine and [14C]Creatine Uptake by hCMEC/D3 Cells
To further examine the involvement of CRT, we examined cyclocreatine and creatine uptake by hCMEC/D3 cells treated with CRT-targeting siRNA (Fig. 5). CRT-targeting siRNA treatment suppressed the CRT mRNA expression by 96.9% compared to the control siRNA (Fig. 5a). The uptake of the [14C]cyclocreatine by the CRT-targeting siRNA-treated hCMEC/D3 cells at 30 min was reduced by 70.0% relative to the control cells (Fig. 5b). The uptake of [14C]creatine by the CRT-targeting siRNA-treated hCMEC/D3 cells was reduced by 63.2% compared to the control cells (Fig. 5c). The reduction in the uptakes of [14C]cyclocreatine and [14C]creatine was relatively the same. These results suggest that CRT is predominantly involved in cyclocreatine and creatine transport by hCMEC/D3 cells.
Effect of CRT knockdown of CRT mRNA expression, [14C]cyclocreatine and [14C]creatine uptake in hCMEC/D3 cells. (a) Quantitative analysis of mRNA expression using RT-qPCR in hCMEC/D3 cells transfected with CRT-targeted siRNA (CRT siRNA) or Silencer™ Negative Control No.1 siRNA (Control siRNA). Each column represents the mean ± SEM (n = 4). (b and c) [14C]Cyclocreatine uptake (b) and [14C]creatine uptake (c) by hCMEC/D3 cells transfected with CRT-targeted siRNA (CRT siRNA) or Silencer™ Negative Control No.1 siRNA (Control siRNA). The uptake was measured at 30 min. Each column represents the mean ± SEM (n = 4). ***p < 0.001, ****p < 0.0001 significantly different from control siRNA.
Characteristics of [14C]Cyclocreatine and [14C]Creatine Uptake by CCDSs Patient-Derived Fibroblasts
It is important to clarify the effect of the CRT deletion and mutation on the cyclocreatine transport. Thus, we examined the transport of cyclocreatine in two fibroblasts that were derived from patients with CCDSs. The first fibroblast harbored a large deletion in the SLC6A8 gene (CRT-Deletion), and the second harbored a single point mutation (CRT-G561R mutation) relative to the intact CRT expressed fibroblast (CRT-WT). The CRT expression level of CRT-WT in the plasma membrane was 0.287 fmol/μg protein (Table II). The inhibition study demonstrated that the uptake of [14C]cyclocreatine was predominantly mediated by CRT in the fibroblast as well as in HEK293 and hCMEC/D3 cells (Fig. 6). By contrast, in the CRT-Deletion, no detectable CRT protein in the plasma membrane was found, and no significant CRT-dependent [14C]cyclocreatine uptake was observed in the inhibition study. In the CRT-G561R mutation, the CRT expression level in the plasma membrane was found to be 0.197 fmol/μg protein (Table II), but no significant [14C]cyclocreatine uptake was observed. In the case of [14C]creatine, the uptake of [14C]creatine was observed only by the fibroblast expressed intact CRT. These results suggest that the fibroblasts from the CCDSs patients largely impair both creatine and cyclocreatine transport.
Effect of inhibitors on [14C]cyclocreatine and [14C]creatine transport in fibroblasts derived from healthy person and CCDSs patients. Effect of inhibitors on [14C]cyclocreatine transport in fibroblast (CRT-WT) of control from a person expressing wild type CRT (a), CCDSs patient with a large about 19 kb deletion (b), and CCDSs patient with a missense mutation (c). Effect of inhibitors on [14C]creatine transport in fibroblast (CRT-WT) of control from a person expressing wild type CRT (d), CCDSs patient with a large about 19 kb deletion (e), and CCDSs patient with a missense mutation (f). [14C]Cyclocreatine uptake (0.1 μCi, 9 μM) and [14C]Creatine uptake (0.1 μCi, 9 μM) were measured in the absence of compounds (control), in the presence of wither GPA(10 mM), unlabeled creatine (10 mM) or unlabeled cyclocreatine (10 mM), or in the absence of either Na+ or Cl− at 37°C for 30 min. Each column represents the mean ± SEM (n = 4). ****p < 0.0001, significantly different from control condition.
Discussion
Our present study demonstrates that cyclocreatine could be a substrate of CRT as well as creatine, which was determined by the in vitro reciprocal uptake studies and the gene knockdown studies in HEK293 cells. CRT seems to play an important role in supplying creatine into the brain across the BBB in mice (8). Hence, the in vitro uptake study by the hCMEC/D3 cells illustrated that CRT predominantly contributes to the uptake of cyclocreatine in the human BBB as well. In addition, the uptake study by the fibroblasts derived from patients with CCDSs also showed that CRT deletion and mutation dramatically reduced the uptake of both cyclocreatine and creatine. These findings suggest that CRT is a major player in the transport of cyclocreatine at the BBB as well as creatine, where the CRT gene mutation results in the reduction of cyclocreatine supply to the brain neurons across the BBB.
This study determined that the uptake of cyclocreatine is likely mediated by a saturable transport system. The lipophilicity (Log n-octanol/Ringer distribution coefficient) of cyclocreatine and creatine are −2.25 and − 2.16, respectively, suggesting that cyclocreatine and creatine both have low passive diffusion rates. In addition, cyclocreatine at pH 7.4 is zwitterionic in the blood (32). Based on the pH partition hypothesis, the zwitterionic form of cyclocreatine in the blood may not be able to penetrate the cell membrane by diffusion.
As CRT is able to transport cyclocreatine and creatine, the Km value of creatine transport by CRT was reported to be ~20 μM (33). Our results have shown that the Km value of cyclocreatine transport in HEK293 cells was determined as 188 μM (Fig. 2B), where the affinity of cyclocreatine for CRT was calculated to be 9.4 times lower than that of creatine. Blood creatine concentration is between 50 to 100 μM (34,35) in humans, and between 200 to 272 μM (34,36) in mice. Blood creatine levels were not altered in CCDSs patients (4,13). A pharmacokinetic study in mice reported that the Cmax of cyclocreatine in the plasma was 39.3 μM after a single oral administration (30 mg/kg PO) (37). Cyclocreatine was administered at 280 mg/kg/day PO for 9 weeks, which improved cognitive behavior in the neuron-specific CRT knockout mice (25). The plasma Cmax of cyclocreatine during the treatment is estimated to be greater than 350 μM with the linear assumption. Therefore, creatine and cyclocreatine levels in the blood were likely higher after the cyclocreatine treatment than their previous Km values against CRT (188 μM). In the studies conducted on WT mice, transport rates of both creatine and cyclocreatine were affected by their interaction in CRT. With that, brain creatine levels were decreased by ~23% after the administration of cyclocreatine (0.28 mg/g/day PO for 9 weeks) in WT mice (25).
Our hCMEC/D3 cells uptake study results suggest that CRT, in the capillary endothelial cells of human brain, could also be involved in cyclocreatine transport (Figs. 4 and 5). As is shown in Fig. 4, more than 88% of the uptake of both [14C]cyclocreatine and [14C]creatine, mediated by Na+ and Cl− dependent transport, was significantly inhibited by either creatine, cyclocreatine or GPA. These transport characteristics suggest that the cyclocreatine and creatine uptake by the hCMEC/D3 cells are primarily mediated by CRT, while the contribution of other transporter and/or passive diffusion is less than 12%. Furthermore, decreasing the expression of CRT by siRNA resulted in a similar attenuation of both cyclocreatine and creatine uptake (Fig. 5). These findings indicate that both creatine and cyclocreatine supply from the blood to the brain could be suppressed by the same extent when the transport activity of CRT at the BBB is limited.
As shown in Fig. 5, mRNA expression of CRT was suppressed by 97% by siRNA (Fig. 5a), while the uptake of [14C]cyclocreatine and [14C]creatine was not completely suppressed (Fig. 5b and c). The residual uptake is likely to be attributed to the remaining CRT proteins not suppressed by siRNA treatments, since the inhibition study suggests that those uptakes were primarily mediated by CRT (Fig. 4), and the siRNA suppression in HEK 293 cells affected the mRNA levels more than it affected protein levels (Fig. 3). The CRT protein suppression by siRNA treatments in hCMEC/D3 cells was not evaluated. Since the protein expression levels of CRT in hCMEC/D3 cells were low, and suppression levels cannot be accurately measured by targeted proteomics due to the sensitivity. Furthermore, antibody against endogenous human CRT is not commercially available for western blot quantification.
Our results indicate the possibility that cyclocreatine is hardly transported across the BBB and into the brain when CRT function is deficient. However, cyclocreatine has been reported to improve cognitive behavior by oral administration in the neuron-specific CRT knockout mice (25). The reason being that the CRT knockout mice were conditional, where the CRT gene was knocked out specifically in neurons by the Cre recombinase and CaMK2a promoter, resulting in CRT expression at the BBB that was not deficient. Indeed, 30% of CRT mRNA expression remained in the neuron-specific CRT knockout mice (25). Therefore, it is possible that cyclocreatine was transported from the blood to the brain by the CRT expressed at the BBB in the CRT knockout mice. Upon entering the brain, cyclocreatine might be distributed in the neural cells by the CRT-independent pathway. As the previous study used mouse hippocampal slices, it demonstrated that the uptake and phosphorylation of cyclocreatine in the slices was still detected to some extent despite the presence of GPA, where the uptake of cyclocreatine into the slices was largely inhibited (38).
In CCDSs, numerous single point mutation sites exist on the CRT gene that lead to single amino acid substitutions. In such cases, the mutated CRT protein is expressed in the cells of the patient. In this study, CRT protein was only detected in the fibroblasts from the patient with the single point mutation, and not with deletions (Table 2). The cyclocreatine uptake was decreased by more than 89% in both the CRT-deletion and the CRT-G561R mutation (Fig. 6b and c). The degree of decrease is relative to the creatine uptake (Fig. 6e and f). These findings indicate that cyclocreatine transport activity could be correlated with creatine transport activity rather than CRT expression in CCDSs patients. Furthermore, the uptake of [14C]cyclocreatine and [14C]creatine by CRT-deletion in the control samples were still comparable (Fig. 6b and e). In addition, the uptake of [14C]cyclocreatine by the CRT-G561R mutation in control conditions was found to be significantly lower than that of [14C]creatine (Fig. 6c and f). Hence, when CRT activity was almost absent, the uptake rate of cyclocreatine was not faster than the uptake of creatine in the fibroblasts. This suggests that passive diffusion of cyclocreatine is not faster than that of creatine. Taken together, it can be considered that cyclocreatine transport at the BBB in CCDSs patients is attenuated as well as creatine transport. Further studies using CRT knockout mice are required to evaluate the role of CRT in cyclocreatine transport in the BBB in vivo.
In conclusion, our results indicate that cyclocreatine is a CRT substrate, and CRT at the BBB likely plays a major role in the transport of cyclocreatine into the brain. Our study also suggests that cyclocreatine transport at the BBB is attenuated as well as creatine transport in CCDSs patients. The reason for discontinuing clinical trials of cyclocreatine is unknown, but the findings of this study may help understand the cause. Our findings have provided important insights for the treatment of CCDSs patients with creatine analogues, such as cyclocreatine. The therapeutic creatine analogues for CCDSs require an ATP energy storage function by creatine kinase and high permeability across the BBB for neuronal uptake regardless of CRT function. Therefore, during drug screening, confirming that the uptake of candidate compounds was mediated by CRT-independent transport mechanism or passive diffusion with the BBB model cells, neurons or CCDSs patient’s fibroblast is essential.
Abbreviations
- ADP:
-
Adenosine diphosphate
- AGAT:
-
L-Arginine: glycine amidinotransferase
- ATP:
-
Adenosine triphosphate
- BAP31:
-
B cell receptor-associated protein
- BBB:
-
Blood-brain barrier
- bFGF:
-
Basic fibroblast growth factor
- CaMK2a:
-
Ca2+/calmodulin-dependent protein kinase a
- CCDSs:
-
Cerebral creatine deficiency syndromes
- CRT:
-
Creatine transporter
- DMEM:
-
Dulbecco’s modified Eagle medium
- FBS:
-
Fetal bovine serum
- GAMT:
-
Guanidinoacetate methyltransferase
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- GC:
-
Graphite carbon
- GPA:
-
β-guanidinopropionic acid
- HBSS:
-
Hank’s balanced salt solution
- hCMEC:
-
Human cerebral microvascular endothelial cell
- HEPES:
-
4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
- MRM:
-
Multiple reaction monitoring
- PTS:
-
Phase-transfer surfactant
- SDB:
-
Styrene-divinylbenzene
- SLC6A8:
-
Solute carrier family 6 member 8
References
Rosenberg EH, Almeida LS, Kleefstra T, deGrauw RS, Yntema HG, Bahi N, et al. High prevalence of SLC6A8 deficiency in X-linked mental retardation. Am J Hum Genet. 2004;75:97–105.
Salomons GS, van Dooren SJ, Verhoeven NM, Marsden D, Schwartz C, Cecil KM, et al. X-linked creatine transporter defect: an overview. J Inherit Metab Dis. 2003;26:309–18.
deGrauw TJ, Cecil KM, Byars AW, Salomons GS, Ball WS, Jakobs C. The clinical syndrome of creatine transporter deficiency. Mol Cell Biochem. 2003;244:45–8.
Almeida LS, Verhoeven NM, Roos B, Valongo C, Cardoso ML, Vilarinho L, et al. Creatine and guanidinoacetate: diagnostic markers for inborn errors in creatine biosynthesis and transport. Mol Genet Metab. 2004;82:214–9.
Wyss M, Kaddurah-Daouk R. Creatine and creatinine metabolism. Physiol Rev. 2000;80:1107–213.
Mercimek-Mahmutoglu S, Stoeckler-Ipsiroglu S, Adami A, Appleton R, Araujo HC, Duran M, et al. GAMT deficiency: features, treatment, and outcome in an inborn error of creatine synthesis. Neurology. 2006;67:480–4.
Battini R, Leuzzi V, Carducci C, Tosetti M, Bianchi MC, Item CB, et al. Creatine depletion in a new case with AGAT deficiency: clinical and genetic study in a large pedigree. Mol Genet Metab. 2002;77:326–31.
Ohtsuki S, Tachikawa M, Takanaga H, Shimizu H, Watanabe M, Hosoya K, et al. The blood-brain barrier creatine transporter is a major pathway for supplying creatine to the brain. J Cereb Blood Flow Metab. 2002;22:1327–35.
Stockler-Ipsiroglu S, van Karnebeek C, Longo N, Korenke GC, Mercimek-Mahmutoglu S, Marquart I, et al. Guanidinoacetate methyltransferase (GAMT) deficiency: outcomes in 48 individuals and recommendations for diagnosis, treatment and monitoring. Mol Genet Metab. 2014;111:16–25.
Edvardson S, Korman SH, Livne A, Shaag A, Saada A, Nalbandian R, et al. L-arginine:glycine amidinotransferase (AGAT) deficiency: clinical presentation and response to treatment in two patients with a novel mutation. Mol Genet Metab. 2010;101:228–32.
Schulze A. Creatine deficiency syndromes. Mol Cell Biochem. 2003;244:143–50.
Poo-Arguelles P, Arias A, Vilaseca MA, Ribes A, Artuch R, Sans-Fito A, et al. X-linked creatine transporter deficiency in two patients with severe mental retardation and autism. J Inherit Metab Dis. 2006;29:220–3.
Cecil KM, Salomons GS, Ball WS Jr, Wong B, Chuck G, Verhoeven NM, et al. Irreversible brain creatine deficiency with elevated serum and urine creatine: a creatine transporter defect? Ann Neurol. 2001;49:401–4.
Rudnick G, Kramer R, Blakely RD, Murphy DL, Verrey F. The SLC6 transporters: perspectives on structure, functions, regulation, and models for transporter dysfunction. Pflugers Arch. 2014;466:25–42.
van de Kamp JM, Mancini GM, Salomons GS. X-linked creatine transporter deficiency: clinical aspects and pathophysiology. J Inherit Metab Dis. 2014;37:715–33.
Uemura T, Ito S, Ohta Y, Tachikawa M, Wada T, Terasaki T, et al. Abnormal N-glycosylation of a novel missense Creatine transporter mutant, G561R, Associated with Cerebral Creatine Deficiency Syndromes Alters Transporter Activity and Localization. Biol Pharm Bull. 2017;40:49–55.
Rosenberg EH, Martinez Munoz C, Betsalel OT, van Dooren SJ, Fernandez M, Jakobs C, et al. Functional characterization of missense variants in the creatine transporter gene (SLC6A8): improved diagnostic application. Hum Mutat. 2007;28:890–6.
Betsalel OT, Pop A, Rosenberg EH, Fernandez-Ojeda M, Jakobs C, Salomons GS. Detection of variants in SLC6A8 and functional analysis of unclassified missense variants. Mol Genet Metab. 2012;105:596–601.
Mak CS, Waldvogel HJ, Dodd JR, Gilbert RT, Lowe MT, Birch NP, et al. Immunohistochemical localisation of the creatine transporter in the rat brain. Neuroscience. 2009;163:571–85.
Dodd JR, Birch NP, Waldvogel HJ, Christie DL. Functional and immunocytochemical characterization of the creatine transporter in rat hippocampal neurons. J Neurochem. 2010;115:684–93.
Skelton MR, Schaefer TL, Graham DL, Degrauw TJ, Clark JF, Williams MT, et al. Creatine transporter (CrT; Slc6a8) knockout mice as a model of human CrT deficiency. PLoS One. 2011;6:e16187.
Baroncelli L, Molinaro A, Cacciante F, Alessandri MG, Napoli D, Putignano E, et al. A mouse model for creatine transporter deficiency reveals early onset cognitive impairment and neuropathology associated with brain aging. Hum Mol Genet. 2016;25:4186–200.
Baroncelli L, Alessandri MG, Tola J, Putignano E, Migliore M, Amendola E, et al. A novel mouse model of creatine transporter deficiency. F1000Res. 2014;3:228.
Woznicki DT, Walker JB. Formation of a supplemental long time-constant reservoir of high energy phosphate by brain in vivo and in vitro and its reversible depletion by potassium depolarization. J Neurochem. 1979;33:75–80.
Kurosawa Y, Degrauw TJ, Lindquist DM, Blanco VM, Pyne-Geithman GJ, Daikoku T, et al. Cyclocreatine treatment improves cognition in mice with creatine transporter deficiency. J Clin Invest. 2012;122:2837–46.
Halestrap AP. Monocarboxylic acid transport. Compr Physiol. 2013;3:1611–43.
Osaka H, Takagi A, Tsuyusaki Y, Wada T, Iai M, Yamashita S, et al. Contiguous deletion of SLC6A8 and BAP31 in a patient with severe dystonia and sensorineural deafness. Mol Genet Metab. 2012;106:43–7.
Kato H, Miyake F, Shimbo H, Ohya M, Sugawara H, Aida N, et al. Urine screening for patients with developmental disabilities detected a patient with creatine transporter deficiency due to a novel missense mutation in SLC6A8. Brain and Development. 2014;36:630–3.
Masuda T, Hoshiyama T, Uemura T, Hirayama-Kurogi M, Ogata S, Furukawa A, et al. Large-scale quantitative comparison of plasma Transmembrane proteins between two human blood-brain barrier model cell lines, hCMEC/D3 and HBMEC/cibeta. Mol Pharm. 2019;16:2162–71.
Masuda T, Tomita M, Ishihama Y. Phase transfer surfactant-aided trypsin digestion for membrane proteome analysis. J Proteome Res. 2008;7:731–40.
Nakamura K, Hirayama-Kurogi M, Ito S, Kuno T, Yoneyama T, Obuchi W, et al. Large-scale multiplex absolute protein quantification of drug-metabolizing enzymes and transporters in human intestine, liver, and kidney microsomes by SWATH-MS: comparison with MRM/SRM and HR-MRM/PRM. Proteomics. 2016;16:2106–17.
Pis-Diez R, Parajón-Costa BS, Franca CA, Piro OE, Castellano EE, González-Baró AC. Cyclocreatine, an anticancer and neuroprotective agent. Spectroscopic, structural and theoretical study. J Mol Struct. 2010;975:303–9.
Dai W, Vinnakota S, Qian X, Kunze DL, Sarkar HK. Molecular characterization of the human CRT-1 creatine transporter expressed in Xenopus oocytes. Arch Biochem Biophys. 1999;361:75–84.
Marescau B, De Deyn P, Wiechert P, Van Gorp L, Lowenthal A. Comparative study of guanidino compounds in serum and brain of mouse, rat, rabbit, and man. J Neurochem. 1986;46:717–20.
Harris RC, Soderlund K, Hultman E. Elevation of creatine in resting and exercised muscle of normal subjects by creatine supplementation. Clin Sci (Lond). 1992;83:367–74.
Marescau B, Deshmukh DR, Kockx M, Possemiers I, Qureshi IA, Wiechert P, et al. Guanidino compounds in serum, urine, liver, kidney, and brain of man and some ureotelic animals. Metabolism. 1992;41:526–32.
Wang AQ, Hughes E, Huang W, Kerns EH, Xu X. Quantification of cyclocreatine in mouse and rat plasma using hydrophilic-interaction ultra-performance liquid chromatography-tandem mass spectrometry. J Pharm Biomed Anal. 2017;145:629–33.
Enrico A, Patrizia G, Luisa P, Alessandro P, Gianluigi L, Carlo G, et al. Electrophysiology and biochemical analysis of cyclocreatine uptake and effect in hippocampal slices. J Integr Neurosci. 2013;12:285–97.
ACKNOWLEDGMENTS AND DISCLOSURES
This study was supported in part by JSPS KAKENHI Grant Number JP26293035 and JP18H02590, and by AMED under Grant Number JP19ek0109396. Ohtsuki S is a full professor at Kumamoto University and is also a director of Proteomedix Frontiers. The other authors declare no competing interests.
Author information
Authors and Affiliations
Contributions
All authors, Uemura T, Ito S, Masuda T, Shimbo H, Goto T, Osaka H, Wada T, Couraud PO, Ohtsuki S, contributed to study design and manuscript revision. Uemura T conducted the experiments and performed data analysis. Uemura T, Ito S. and Ohtsuki S wrote the manuscript. All the authors have provided their final approval for submission of this manuscript.
Corresponding author
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Uemura, T., Ito, S., Masuda, T. et al. Cyclocreatine Transport by SLC6A8, the Creatine Transporter, in HEK293 Cells, a Human Blood-Brain Barrier Model Cell, and CCDSs Patient-Derived Fibroblasts. Pharm Res 37, 61 (2020). https://doi.org/10.1007/s11095-020-2779-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11095-020-2779-0