Abstract
In recent years, it is popular that the combination of chaos and DNA is employed for image ciphers. There have been a great number of image ciphers which are designed based on chaos and DNA, but the corresponding cryptanalytic works are insufficient and in-depth study should be made. In this paper, we decipher an image cipher combining a 3-cell chaotic map with DNA. It was claimed that this cipher has high security and can resist different well-known attacks. However, we demonstrate that the claim is not reasonable, since the cipher can be broken by chosen plaintext attack with the complexity \(O({\alpha \beta })\), where \(\alpha \) and \(\beta \) represent the number of the image rows and columns, respectively.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Nowadays the usages of image data have become more and more widespread over various networks, thus the corresponding security has been receiving increasing attention. Chaos is one of the most common tools to protect image data since it has the sensitivity to initial condition and control parameter, which is similar to the basic properties of cryptography, confusion and diffusion. Meanwhile, the cryptanalysis of chaos-based image cipher is a vital work for its development. Alvarez and Li presented a general framework of basic guidelines involved with three main issues including implementation, key management and security analysis that a cryptosystem should follow [1]. In [2], chosen plaintext attack and chosen ciphertext attack were separately employed to break an image encryption algorithm based on hyper-chaos without any knowledge of the key value and with only three couples of plaintext/ciphertext. Rhouma and Belghith cryptanalyzed a chaos-based cipher for images and videos with two different attacks due to the weakness in the generation of the keystream [3]. The knowledge and significance of the cryptanalysis of chaotic ciphers have been discussed in [4, 5] in detail. Çokal and Solak applied chosen plaintext and known plaintext attacks to reveal the secret parameters of a chaos-based image encryption algorithm [6]. Two image encryption schemes consisting of a permutation operation and an XOR-like transformation of the shuffled pixels were cracked in [7] by use of a chosen plaintext attack. A differential cryptanalysis was leveraged by Li et al. [8] to aim at the Yen–Chen–Wu multimedia cryptography system with only seven chosen plaintexts. The image encryption cryptosystems based on multi-chaotic systems and improved hyper-chaotic sequences were also analyzed in terms of security in [9, 10], respectively. In addition, Bigdeli algorithm was broken by using Çokal and Solak attack in [11]. In recent years, it is very interesting to combine the biological characteristics like DNA with chaos system for image ciphers. So far, a large number of image ciphers based on DNA and chaos have been designed [12–25]. Their basic idea is to encode the original image into DNA sequence according to some mapping rule. The obtained DNA sequence is then processed in the some way with the help of the secret DNA keystream which is typically controlled by chaos. The processed DNA sequence is finally decoded into the cipher image. In addition, some chaos-based cryptographic features can also be embedded to enhance the security. On the other hand, a small number of cryptanalytic works for a few of these image ciphers have been done [26–32], which stimulates the further development of the image ciphers based on DNA and chaos. In spite of this, such cryptanalytic works are insufficient and in-depth study should be made.
In [33], the combination of a 3-cell chaotic map and DNA is employed for designing image cipher. Specifically, the original image is firstly converted to a DNA sequence. Then, the obtained DNA sequence is scrambled under the control of cycling chaos. After that, a secret DNA sequence is generated by using cycling chaos to mask the scrambled DNA sequence. The results are decoded to form the final cipher image. It was claimed that this cipher has high security and can resist different well-known attacks. In this paper, we demonstrate that the claim is not reasonable, since the cipher can be broken by known attacks. The rest of the paper is organized as follows. The next section overviews the original image cipher based on 3-cell chaotic map and biological operations. Section 3 gives the corresponding cryptanalysis for the original image cipher followed by some discussions in Sect. 4. The last section concludes this paper.
2 Overview of the image cipher based on 3-cell chaotic map and biological operations
This section firstly re-introduces the preparatory work for the cipher design and then re-describes the original image cipher.
2.1 The preparatory work
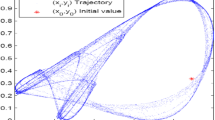
The 3-cell cyclic chaos map used in the image cipher is defined as
A 120-bit external key \(k=k_{120} ,k_{119} ,k_{118} ,\ldots ,k_1 \) is applied for providing the three initial values \(x_1 \), \(y_1 \), \(z_1 \) and the parameter p by the following formulas:
Each 8-bit pixel can be comprised of four 2-bits, i.e., 00, 01, 10 and 11, which can correspond to four nucleic acid bases T (thymine), A (adenine), C (cytosine), G (guanine), where A and T are complementary, while G and C are complementary, according to Watson–Crick complement rule, as shown in Table 1. The DNA algebraic operations including addition and subtraction are shown in Tables 2 and 3, respectively.
2.2 The encryption algorithm
The original image is firstly converted to a DNA sequence. Then, the obtained DNA sequence is scrambled under the control of cycling chaos. After that, a secret DNA sequence is generated by using cycling chaos to mask the scrambled DNA sequence. The results are decoded to form the final cipher image. The specific procedures are as follows:
-
Step 1: The DNA encoding phase
Encode each pixel of the original image into a DNA image according to the rule: “00”-A, “01”-C, “10”-G, and “11”-T.
-
Step 2: The scrambling phase
-
a.
Let the number of the image rows and columns be \(\alpha \) and \(\beta \), respectively, and divide the DNA image into N equal blocks.
-
b.
Generate the chaotic sequences \(X=x_0 ,x_1 ,x_2 ,\ldots ,x_\alpha \), \(Y=y_0 ,y_1 ,y_2 ,\ldots ,y_{4\beta } \), and \(Z=z_0 ,z_1 ,z_2 ,\ldots ,z_N \) with the initial values \(x_1 \), \(y_1 \), \(z_1 \) and the parameter p;
-
c.
Normalize the values in the sequence Z to the integers between 1 to N;
-
d.
Rearrange these integers into a new matrix \({Z}'\);
-
e.
Scramble the blocks using \({Z}'\);
-
f:
Normalize the values in X and Y to the integers between 1 to \(\alpha \) and between 1 to \(\beta \), respectively, rearrange these integers into row matrix and column matrix \({X}'\) and \({Y}'\);
-
g:
Scramble the nucleic acid base values using the ordered pair \(\left( {{x}',{y}'} \right) \in \left( {{X}',{Y}'} \right) \).
-
a.
-
Step 3: The mask phase
-
a.
Generate the chaotic sequence \(X=x_0 ,x_1 ,x_2 ,\ldots ,x_{4\alpha \,\beta } \) with the initial values \(x_1 \), \(y_1 \), \(z_1 \) and the parameter p;
-
b.
Construct the mask image M in which each pixel value can be generated using the following formula:
$$\begin{aligned} \left\{ {\begin{array}{l} A,-2<X\le 1.1 \\ C,-1.1<X\le -0.2 \\ G,0.2\le X<1.1 \\ T,1.1\le X<2 \\ \end{array}} \right. \end{aligned}$$(3) -
c.
Perform the DNA addition of the scrambled DNA image and the mask image.
-
a.
-
Step 4: The decoding phase
Decode the result into the final cipher image according to the same encoding rule: “00”-A, “01”-C, “10”-G, and “11”-T.
3 The deciphering process
After carefully observing the encryption algorithm, it seems that the security of the algorithm depends mainly on the 120-bit external key, which manipulates the generation of the three initial values \(x_1 \), \(y_1 \), \(z_1 \) and the parameter p for 3-cell cycling chaos map. However, with respect to different images, the same external key is utilized, thus keeping the same \(x_1 \), \(y_1 \), \(z_1 \) and p. Furthermore, the chaotic sequences remain unchanged, therefore \({X}'\), \({Y}'\), \({Z}'\) and M are fixed. The encryption algorithm possesses the architecture of scrambling mask. The encoding and decoding phases do not involve cryptographic behaviors. The scrambling phase contains two operations. One is to scramble the blocks using \({Z}'\) and the other is to process the whole bases using \({X}'\) and \({Y}'\). The mask phase only leverages the mask image M. The goal of deciphering the image encryption algorithm is to reveal \({X}'\), \({Y}'\), \({Z}'\) and M. In order to be understood for the proposed cryptanalysis, the encryption algorithm was simplified as:
-
Step 1: The DNA encoding phase
$$\begin{aligned}&P\left( {00,01,10,11} \right) \mathop {\xrightarrow {\qquad \qquad \mathrm{Encode}\qquad \qquad }}\limits _{\text {00-A,01-C,10-G,11-T}}\nonumber \\&\quad \times {P}'\left( {A,C,G,T} \right) \end{aligned}$$(4) -
Step 2: The scrambling phase
$$\begin{aligned}&{P}'\left( {A,C,G,T} \right) \mathop {\xrightarrow {\qquad \mathrm{1st \,Scramble}\qquad }}\limits _{{Z}'}\nonumber \\&\quad {P}'_{S^{1}} \left( {A,C,G,T} \right) \end{aligned}$$(5)$$\begin{aligned}&{P}'_{S^{1}} \left( {A,C,G,T} \right) \mathop {\xrightarrow {\qquad \mathrm{2st \,Scramble}\qquad }}\limits _{{X}',{Y}'}\nonumber \\&\quad \times {P}'_{S^{2}} \left( {A,C,G,T} \right) \end{aligned}$$(6) -
Step 3: The mask phase
$$\begin{aligned} {P}'_{S^{2}} \left( {A,C,G,T} \right) \mathop {\xrightarrow {\qquad \mathrm{Mask}\quad }}\limits _{M} {P}'_{M,S^{2}} \left( {A,C,G,T} \right) \end{aligned}$$(7) -
Step 4: The decoding phase
$$\begin{aligned}&{P}'_{M,S^{2}} \left( {A,C,G,T} \right) \mathop {\xrightarrow {\qquad \qquad \mathrm{Decode}\qquad \qquad }}\limits _{\text {A-00,C-01,G-10,}T\text {-11}}\nonumber \\&\quad \times C\left( {A,C,G,T} \right) \end{aligned}$$(8)
Our attack adopts chosen plaintext attack. The basic idea is to select one or more special images, which are not or regularly affected by the scrambling phase. So the attacker can infer the mask image M at first with the knowledge of one or more pairs of plain/cipher images. The specific procedures are the followings:
-
(1)
Select a special plain image, where each 8-bit pixel consists of the same four 2-bits, i.e.,
$$\begin{aligned}&P\left( {00,00,00,00} \right) , \quad P\left( {01,01,01,01} \right) ,\\&P\left( {10,10,10,10} \right) \quad \hbox {and}\quad P\left( {11,11,11,11} \right) , \end{aligned}$$corresponding to \({P}'\left( {A,A,A,A} \right) \), \({P}'\left( {C,C,C,C} \right) \), \({P}'\left( {G,G,G,G} \right) \) and \({P}'\left( {T,T,T,T} \right) \). Without loss of generality, \(P\left( {00,00,00,00} \right) \) is selected.
-
(2)
The two scrambling operations do not work for \({P}'\left( {A,A,A,A} \right) \), thus
$$\begin{aligned} {P}'\left( {A,A,A,A} \right)= & {} {P}'_{S^{1}} \left( {A,A,A,A} \right) \nonumber \\= & {} {P}'_{S^{2}} \left( {A,A,A,A} \right) \end{aligned}$$(9) -
(3)
The mask phase is changed as
$$\begin{aligned} {P}'\left( {A,A,A,A} \right) \mathop {\xrightarrow {\quad \mathrm{Mask}\quad }}\limits _{M} {P}'_M \left( {A,C,G,T} \right) \end{aligned}$$(10)The decoding phase is
$$\begin{aligned} {P}'_M \left( {A,C,G,T} \right) \mathop {\xrightarrow {\qquad \mathrm{Decode}\qquad }}\limits _{\text {A-00,C-01,G-10},T\text {-11}} C^{1}\left( {A,C,G,T} \right) \end{aligned}$$(11) -
(4)
\(P\left( {00,00,00,00} \right) \)and \(C^{1}\left( {A,C,G,T} \right) \) are known, so \({P}'\left( {A,A,A,A} \right) \quad {P}'_M \left( {A,C,G,T} \right) \) are also known. With the addition and subtraction operation rules in Tables 2 and 3 together with Eq. (10), the mask image M can be immediately calculated.
-
(5)
After revealing M, the resultant encryption algorithm turns into a scrambling-only framework. The effect of the second scrambling is obviously stronger than that of the first one. The first scrambling can be integrated into the second one. The two scrambling operations can be equivalent to only once nucleic acid base value scrambling. The equivalent matrices for \(\left( {{X}',{Y}'} \right) \) and \({Z}'\) which need to be revealed are marked as \(\left( {{X}'',{Y}''} \right) \).
-
(6)
Alter three 2-bits of four 2-bits as 01, 10, and 11 for the first pixel in P( 00, 00, 00, 00 ). After the scrambling, the positions of these three altered values can be found such that three ordered pairs in \(( {{X}'',{Y}''} )\) are obtained. Next, the same processing way for the following pixels is repeatedly performed till the whole ordered pairs in \(( {{X}'',{Y}''} )\) are revealed.
The above steps demonstrate that the encryption algorithm can be broken by using chosen plaintext attack. The mask phase requires an image, while the scrambling phase requires about \({4\alpha \beta }\big /3\) images. Thus, the complexities in the mask and scrambling phases are O( 1 ) and \(O( {{4\alpha \beta }\big /3} )\), respectively. The whole complexity is confined to \(O( {\alpha \beta })\), which is low. If there needs to be a blurry or rough image through the cryptanalysis, the number of the images when deciphering the scrambling phase can be further optimized by the methods [34, 35], in which the scrambling-only image ciphers have been thoroughly and systematically cryptanalyzed. Figure 1 shows a validation example, in which a–c represents the original test image, the corresponding encrypted image and the cryptanalyzed image, respectively. It is worth mentioning in the end that the algebraic attack against the image encryption algorithm consisting of chaos and DNA is an efficient tool of the cryptanalysis. For example, in [27], Belazi et al. came up with an algebraic analysis against a RGB image encryption algorithm based on DNA encoding and chaotic map by constructing some systems of linear equations. However, in [33], the introduction of permutation which has the linearity impedes the construction of linear equations. Thus, the algebraic attack does not take effect on the cipher in [33].
4 Discussion
In the following, some improvements and suggestions with respect to the encryption algorithm are given:
-
1.
There are eight kinds of coding schemes in total that satisfy the Watson–Crick complement rule. In the encryption algorithm, the coding rule is fixed and no cryptographic features are embedded in the encoding phase. Therefore, a new key can be added to control the encoding and decoding rules.
-
2.
For the DNA algebraic operations, another DNA XOR operation can also be introduced to change the base values in the mask phase. For example, one possible definite for DNA operation is shown in Table 4. The DNA XOR operation corresponds to the binary XOR operation, which has been widely used in the field of cryptography.
-
3.
The leading reason why the encryption algorithm is vulnerable to chosen plaintext attack is not related to plaintext information. The algorithm should depend on the plain image information during the encoding, scrambling and mask phases. As a result, different plain images can generate inconsistent keystreams.
-
4.
In the scrambling and mask phases, the same initial values and parameter should not be utilized. Once one part of keystream is revealed by the attacker, the other part may also be known at once. The keystream used in mask phase can be perturbed by the encryption result in the scrambling phase to avoid the same keys and enhance the security
-
5.
The A, C, G and T in the mask image are not uniform in terms of the distribution situation due to the fact that in general, the track of chaos system is not uniform. Thus, the encryption algorithm easily suffers from statistical attack. Some excellent DNA cipher techniques like [15] are worth referencing for better image cipher based on DNA and chaos.
5 Conclusion
An encryption algorithm based on a 3-cell chaotic map and DNA, including the encoding, scrambling, mask and decoding phases, has been analyzed in terms of security in this paper. The algorithm can be broken by chosen plaintext attack with a low complexity \(O( {\alpha \beta } )\). At the same time, some improvements and suggestions have also been proposed for the purpose of designing better image ciphers based on DNA and chaos and stimulating the further development.
References
Alvarez, G., Li, S.: Some basic cryptographic requirements for chaos-based cryptosystems. Int. J. Bifurc. Chaos 16(8), 2129–2153 (2006)
Rhouma, R., Belghith, S.: Cryptanalysis of a new image encryption algorithm based on hyper-chaos. Phys. Lett. A 372, 5973–5978 (2008)
Rhouma, R., Belghith, S.: Cryptanalysis of a spatio-temporal chaotic image/video cryptosystem. Phys. Lett. A 372, 5790–5794 (2008)
Arroyo, D., Li, S.: Lessons learnt from the cryptanalysis of chaos-based ciphers. In: Kocarev, L., Lian, S. (eds.) Chaos Based Cryptography Theory Algorithms and Applications, pp. 257–295. Springer, Berlin (2011)
Solak, E.: Cryptanalysis of chaotic ciphers. In: Kocarev, L., Lian, S. (eds.) Chaos Based Cryptography Theory Algorithms and Applications, pp. 227–256. Springer, Berlin (2011)
Çokal, C., Solak, E.: Cryptanalysis of a chaos-based image encryption algorithm. Phys. Lett. A 373, 1357–1360 (2009)
Arroyo, D., Li, C., Li, S., Alvarez, G., Halang, W.: Cryptanalysis of an image encryption scheme based on a new total shuffling algorithm. Chaos Solitons Fract. 41, 2613–2616 (2009)
Li, C., Li, S., Lo, K.-T., Kyamakya, K.: A differential cryptanalysis of Yen–Chen–Wu multimedia cryptography system. J. Syst. Softw. 83(8), 1443–1452 (2010)
Solak, E., Rhouma, R., Belghith, S.: Cryptanalysis of a multi-chaotic systems based image cryptosystem. Opt. Commun. 283, 232–236 (2010)
Özkaynak, F., Özer, A.B., Yavuz, S.: Cryptanalysis of a novel image encryption scheme based on improved hyper chaotic sequences. Opt. Commun. 285, 4946–4948 (2012)
Özkaynak, F., Özer, A.B., Yavuz, S.: Cryptanalysis of Bigdeli algorithm using Çokal and Solak attack. Int. J. Inf. Secur. Sci. 1(3), 79–81 (2012)
Babaei, M.: A novel text and image encryption method based on chaos theory and DNA computing. Nat. Comput. 12(1), 101–107 (2013)
Huang, X.L., Ye, G.D.: An image encryption algorithm based on hyper-chaos and DNA sequence. Multimed. Tools Appl. 72(1), 57–70 (2014)
Liu, L.L., Zhang, Q., Wei, X.P.: A RGB image encryption algorithm based on DNA encoding and chaotic map. Comput. Electr. Eng. 38(5), 1240–1248 (2012)
Liu, H.J., Wang, X.Y., Kadir, A.: Image encryption using DNA complementary rule and chaotic maps. Appl. Soft Comput. 12(5), 1457–1466 (2012)
Wei, X.P., Guo, L., Zhang, Q., Zhang, J.X., Lian, S.G.: A novel color image encryption based on DNA sequence operation and hyper-chaotic system. J. Syst. Softw. 85(2), 290–299 (2012)
Zhang, Q., Guo, L., Wei, X.P.: Image encryption using DNA addition combining with chaotic maps. Math. Comput. Model. 52(11–12), 2028–2035 (2010)
Zhang, Q., Guo, L., Wei, X.P.: A novel image fusion encryption algorithm based on DNA sequence operation and hyper-chaotic system. Optik 124(18), 3596–3600 (2013)
Zhang, Q., Liu, L.L., Wei, X.P.: Improved algorithm for image encryption based on DNA encoding and multi-chaotic maps. AEU-Int. J. Electron. Commun. 68(3), 186–192 (2014)
Zhang, Q., Wei, X.P.: A novel couple images encryption algorithm based on DNA subsequence operation and chaotic system. Optik 124(23), 6276–6281 (2013)
Jain, A., Rajpal, N.: A robust image encryption algorithm resistant to attacks using DNA and chaotic logistic maps. Multimed. Tools Appl. doi:10.1007/s11042-015-2515-7
Liu, Y., Wang, J., Fan, J., Gong, L.: Image encryption algorithm based on chaotic system and dynamic S-boxes composed of DNA sequences. Multimed. Tools Appl. doi:10.1007/s11042-015-2479-7
Enayatifar, R., Abdullah, A.H., Isnin, I.F.: Chaos-based image encryption using a hybrid genetic algorithm and a DNA sequence. Opt. Lasers Eng. 56, 83–93 (2014)
Kulsoom, A., Xiao, D., Rehman, A., Abbas, S.: An efficient and noise resistive selective image encryption scheme for gray images based on chaotic maps and DNA complementary rules, Multimed. Tools Appl. doi:10.1007/s11042-014-2221-x
Hermassi, H., Belazi, A., Rhouma, R., Belghith, S.: Security analysis of an image encryption algorithm based on a DNA addition combining with chaotic maps. Multimed. Tools Appl. 72(3), 2211–2224 (2014)
Özkaynak, F., Yavuz, S.: Analysis and improvement of a novel image fusion encryption algorithm based on DNA sequence operation and hyper-chaotic system. Nonlinear Dyn. 78(2), 1311–1320 (2014)
Belazi, A., Hermassi, H., Rhouma, R., Belghith, S.: Algebraic analysis of a RGB image encryption algorithm based on DNA encoding and chaotic map. Nonlinear Dyn. 76(4), 1989–2004 (2014)
Zhang, Y.: Cryptanalysis of a novel image fusion encryption algorithm based on DNA sequence operation and hyper-chaotic system. Optik 126(2), 223–229 (2015)
Xie, T., Liu, Y., Tang, J.: Breaking a novel image fusion encryption algorithm based on DNA sequence operation and hyper-chaotic system. Optik 125(24), 7166–7169 (2014)
Liu, Y., Tang, J., Xie, T.: Cryptanalyzing a RGB image encryption algorithm based on DNA encoding and chaos map. Opt. Laser Tech. 60, 111–115 (2014)
Zhang, Y.S., Wen, W., Su, M., Li, M.: Cryptanalyzing a novel image fusion encryption algorithm based on DNA sequence operation and hyper-chaotic system. Optik 125(4), 1562–1564 (2014)
Zhang, Y.S., Xiao, D., Wen, W., Wong, K.-W.: On the security of symmetric ciphers based on DNA coding. Inf. Sci. 289, 254–261 (2014)
SaberiKamarposhti, M., Mohammad, D., Rahim, M., Yaghobi, M.: Using 3-cell chaotic map for image encryption based on biological operations. Nonlinear Dyn. 75(3), 407–416 (2014)
Li, S.J., Li, C.Q., Chen, G.R., Bourbakis, N.G., Lo, K.T.: A general quantitative cryptanalysis of permutation-only multimedia ciphers against plaintext attacks. Signal Process. Image 23(3), 212–223 (2008)
Li, C.Q., Lo, K.T.: Optimal quantitative cryptanalysis of permutation-only multimedia ciphers against plaintext attacks. Signal Process. 91(4), 949–954 (2011)
Acknowledgments
The work was supported by the Fundamental Research Funds for the Central Universities (Grant Nos. SWU115016, XDJK2015C077, XDJK2015B030), the National Natural Science Foundation of China (Grant Nos. 61462032, 61402380, 61403313, 61374078), the Natural Science Foundation of Jiangxi Province (Grant No. 20142BAB217012), and the Natural Science Foundation Project of Chongqing CSTC (Grant No. cstc2014jcyjA40014).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, Y., Li, Y., Wen, W. et al. Deciphering an image cipher based on 3-cell chaotic map and biological operations. Nonlinear Dyn 82, 1831–1837 (2015). https://doi.org/10.1007/s11071-015-2280-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11071-015-2280-1