Abstract
This study was conducted to further our understanding about the link between lipid peroxidation and protein carbonylation in rat brain slices incubated with the glutathione (GSH)-depletor diethyl maleate. Using this in vitro system of oxidative stress, we found that there is a significant lag between the appearance of carbonylated proteins and GSH depletion, which seems to be due to the removal of oxidized species early on in the incubation by the mitochondrial Lon protease. Upon acute GSH depletion, protein carbonyls accumulated mostly in mitochondria and to a lesser degree in other subcellular fractions that also contain high levels of polyunsaturated lipids. This result is consistent with our previous findings suggesting that lipid hydroperoxides mediate the oxidation of proteins in this system. However, these lipid hydroperoxides are not produced by oxidation of free arachidonic acid or other polyunsaturated free fatty acids by lipooxygenases or cyclooxygenases. Finally, γ-glutamyl semialdehyde and 2-amino-adipic semialdehyde were identified by HPLC as the carbonyl-containing amino acid residues, indicating that proteins are carbonylated by metal ion-catalyzed oxidation of lysine, arginine and proline residues. The present findings are important in the context of neurological disorders that exhibit increased lipid peroxidation and protein carbonylation, such as Parkinson’s disease, Alzheimer’s disease, and multiple sclerosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Carbonylation refers to the non-enzymatic addition of aldehydes or ketones to specific amino acid residues and constitutes the major and most common oxidative modification of proteins [1]. Protein carbonyls (PCOs) affect the function and/or metabolic stability of the modified proteins [2], and are likely to play an important role in the pathophysiology of disorders with considerable oxidative stress. Carbonylation of brain proteins has been implicated in the etiology and/or progression of several neurodegenerative disorders including Alzheimer’s disease [3], Parkinson’s disease [4], amyotrophic lateral sclerosis [5] and multiple sclerosis [6, 7].
To understand the process underlying the formation and accumulation of PCOs during oxidative stress, we utilized a brain slice system where oxidative stress and protein carbonylation are induced by acute depletion of cellular glutathione (GSH) with diethyl maleate (DEM) or with 1,2-bis(2-chloroethyl)-1-nitrosourea [8]. Under these conditions there is increased mitochondrial production of reactive oxygen species (ROS), which leads to extensive lipid peroxidation (LPO) and protein carbonylation. Later, we demonstrated that LPO is necessary for the carbonylation of most proteins since the hydrophobic antioxidants caffeic acid phenethyl ester and butylated hydroxytoluene were able to prevent DEM-induced protein oxidation [9]. Furthermore, evidence was obtained suggesting that the mechanism underlying LPO-mediated protein carbonylation is the direct oxidation of amino acid side-chains by lipid hydroperoxides (or their immediate decomposition products lipid peroxyl and lipid alkoxyl radicals) rather than the attachment of reactive carbonyl species (RCS) like 4-hydroxynonenal (4-HNE), malondialdehyde (MDA) and acrolein. This idea was based on the absence of RCS-protein adducts and the failure of several RCS-trapping agents (methoxylamine, pyridoxamine, carnosine, taurine and histidine hydrazide) to prevent protein carbonylation in rat brain slices incubated with DEM [10].
The present study was designed to fill in some gaps in our knowledge on the mechanism of carbonylation upon acute GSH depletion. Utilizing DEM-treated rat brain slices, we herein demonstrate that protein carbonylation occurs mostly in mitochondria and to a lesser degree in organelles that also contain high levels of polyunsaturated lipids by metal ion-catalyzed oxidation likely mediated by lipid hydroperoxides. However, these lipid hydroperoxides are not produced enzymatically by oxidation of free arachidonic acid or other polyunsaturated fatty acids by lipooxygenases (LOXs) or cyclooxygenases (COX). We also provide evidence suggesting that the mitochondrial Lon protease is responsible for removing most PCOs in this model of oxidative stress. Some of the results have been presented in abstract form [11].
Materials and Methods
Chemicals
Aristolochic acid (Aristo A), diethyl maleate (DEM), cycloheximide, 5,8,11,14-eicosatetraynoic acid (ETYA), fluoresceinamine isomer II (FlNH2), idebenone and rotenone were from Sigma (St. Louis, MO). Trifluoromethylketone arachidonic acid (AACOF3), docebenone (AA-861) and lactacystin (Lac) were from Enzo Life Sciences (Plymouth Meeting, PA). Bongkrekic acid (BkA) and cyclosporin A (CsA) were from Calbiochem (La Jolla, CA). Epoxomicin (Epo) was from Boston Biochemicals (Cambridge, MA). All other chemicals were of the highest purity available.
Incubation of Rat Brain Slices
Forty-day old Sprague-Dawley male rats were used throughout. Housing and handling of the animals as well as the euthanasia procedure were in strict accordance with the NIH Guide for the Care and Use of Laboratory Animals, and were approved by the Institutional Animal Care and Use Committee. Animals were killed by decapitation, and the brains were rapidly removed and sliced in two directions at right angle in sections 400 µm-thick using surgical grade, carbon steel, razor blades. Slices corresponding to ~ 80 mg of tissue were transferred to flasks containing 3 ml of Hank’s balanced salt solution supplemented with 10 mM d-glucose, and were incubated at 37 °C under 95% O2/5% CO2. DEM and other drugs were added at the beginning of the incubation period as indicated in the figure legends. After incubation, tissue sections were collected by low-speed centrifugation and rinsed twice with ice-cold saline solution. Slices were homogenized by probe sonication in HEN buffer (10 mM Hepes pH 7.0, 1 mM EDTA and 0.1 mM neocuproine) containing 1 mM 4,5-dihydroxy-1,3-benzene sulfonate and 0.5 mM dithiothreitol to prevent further protein oxidation. Homogenates were kept at − 80 °C until use. Protein concentration was assessed with the Bio-Rad protein assay (Bio-Rad Laboratories, Hercules, CA) using bovine serum albumin as a standard.
Measurement of Lipid Peroxidation Products
The concentration of MDA plus 4-HNE was measured with N-methyl-2-phenylindole [12]. Briefly, aliquots from brain homogenates were incubated in 3.2 mM N-methyl-2-phenylindole and 1.76 M methanesulfonic acid in acetonitrile:methanol (3:1, v/v) for 40 min at 45 °C. Aggregated material was removed by centrifugation at 10,000 g for 10 min and the absorbance of the supernatant was measured at 586 nm. The amount of MDA plus 4-HNE was calculated using a standard curve prepared with increasing amounts of MDA.
Assessment of Protein Carbonylation by Western Blotting
Protein carbonyl groups were measured with the OxyBlot™ protein oxidation detection kit (Intergen Co., Purchase, NY), following the protocol provided by the manufacturer. In brief, proteins were incubated with 2,4-dinitrophenyl-hydrazine to form the 2,4-dinitrophenyl (DNP) hydrazone derivatives. Proteins were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis on a 12% gel and blotted to polyvinylidene difluoride membranes. DNP-containing proteins were then immunostained using rabbit anti-DNP antiserum (1:500) and goat anti-rabbit IgG conjugated to horseradish peroxidase (1:2000). Blots were developed by enhanced chemioluminescence using the Western Lightning ECL™ kit from Perkin-Elmer (Boston, MA). The developed films were scanned in a Hewlett Packard Scanjet 4890 and the images quantified using the NIH image analysis program version 1.63. Lane intensities in the oxyblot were normalized by the corresponding commassie brilliant blue staining.
Subcellular Fractionation
Brain slices were homogenized in 0.30 M sucrose in HEN buffer containing 0.25 µM butylated hydroxytoluene using a glass–glass homogenizer. Subcellular fractions (myelin, synaptosomes, mitochondria, microsomes and cytosol) were prepared from an aliquot of the total homogenate by sucrose-gradient ultracentrifugation as described elsewhere [13]. Aliquots of these fractions were used for oxyblot and fatty acid analysis. Purity of the fractions was assessed by western blotting using the following antibodies: proteolipid protein (PLP, myelin), synaptosomal-associated protein 25 (SNAP-25, synaptosomes), Lon protease (LonP, mitochondria) and glyceraldehyde 3-phosphate dehydrogenase (GAPDH, cytosol).
Fatty Acid Analysis by GLC
Aliquots from various subcellular fractions were extracted with chloroform-methanol (2:1, v/v) containing 0.25% (v/v) 12 N HCl to ensure the complete extraction of acidic lipids. Extracts were rinsed with water, dried under a stream of nitrogen gas, and fatty acid methyl esters were prepared by alkaline methanolysis. Briefly, lipids were incubated overnight at room temperature in sealed tubes with chloroform: 0.2 N NaOH in methanol (2:1, v/v). The solution was then neutralized with acetic acid and washed twice with methanol:water (1:1, v/v). The resultant fatty acid methyl esters were dried under nitrogen, dissolved in hexane and analyzed by gas-liquid chromatography using a Hewlett Packard 5890 Series II Gas Chromatograph (Kennett Square, PA) equipped with a fused silica Megabore DB-225 column (15 m × 0.53 mm; J&W, Folsom, CA), a flame ionization detector and an integrator. The flow of carrier gas (helium) was set at 15 ml/min, and injection port and FID temperature were 250 °C. Oven temperature was raised from 150 to 220 °C in 14 min. Peaks were identified by the use of saturated, monounsaturated and polyunsaturated standard methyl esters. The area under each peak was considered proportional to the mass of each methyl ester within the sample. Membrane peroxidizability index (PI) was calculated according to the following formula: PI = (% monoenoic acids × 0.025) + (% dienoic acids × 1) + (% trienoic acids × 2) + (% tetraenoic acids × 4) + (pentaenoic acids × 6) + (hexaenoic acid × 8) [14].
Identification of Carbonylated Amino Acids
The identity of the carbonylated amino acid residues was determined by HPLC [15]. Briefly, proteins from tissue slices that had been incubated with DEM for 3 h were solubilized in 2-(N-morpholino) ethanosulfonate buffer pH 6.0 containing 1% (w/v) SDS and 0.1 mg/ml butylated hydroxytoluene. Carbonyl groups were then derivatized by incubation at 25 °C with 10 mM FlNH2 (in 0.5 M NaOH) and 20 mM sodium cyanoborohydride. After 2 h, proteins were precipitated with acetone and the pellets rinsed once with chloroform:methanol (1:1, v/v) to remove most of the reagents. FlNH2-derivatized proteins were hydrolyzed overnight in 6 N HCl at 110 °C. The dried hydrolysate was dissolved in 10 mM potassium phosphate pH 7.0 and filtered through a 0.2 µm nylon filter disk. A portion of the filtrate was injected onto a Discovery® reversed-phase C18 column (12.5 cm × 4 mm, 5 µm) (Supelco, Bellefonte PA) equilibrated with the same buffer and the labeled amino acids were eluted with a linear gradient from 0 to 70% acetonitrile in 35 min. Absorbance was followed at 490 nm. Bovine serum albumin was oxidized with ascorbic acid/FeCl3 before FlNH2 treatment and acid hydrolysis to generate the FlNH2 derivatives of γ-glutamyl semialdehyde (GGS) and 2-amino-adipic semialdehyde (AAS) [16].
Assay of Proteasomal Chymotrypsin-Like Activity
The chymotrypsin-like activity of the 20S proteasome was determined in the homogenates from control and DEM-treated slices using a fluorescence assay [16]. Briefly, 25 µg of protein were incubated for 1 h at 25 °C with 50 µM of the 7-aminomethyl-4-coumarin (AMC)-labeled peptide Suc-Leu-Leu-Val-Tyr-AMC in the absence or presence of 10 µM clasto-lactacystin-β-lactone. The activity of the 20S proteasome was calculated as the difference in fluorescence intensity at 460 nm between the samples without and with clasto-lactacystin-β-lactone using an excitation wavelength of 380 nm.
Statistical Analysis
All data is reported as mean ± SEM and statistical significance was determined utilizing the GraphPad Prism® (version 4) program (GraphPad Software Inc., San Diego, CA). Comparisons between two groups were done using the Student’s t test, while comparisons between multiple groups were carried out using analysis of variance (ANOVA).
Results
Lipid Peroxidation Precedes Protein Carbonylation in DEM-Treated Brain Slices
We have previously shown that treatment of brain slices with DEM leads to a rapid decline in the cellular levels of GSH causing lipid and protein oxidation [8]. Figure 1 depicts the appearance of PCOs and LPO products (MDA plus 4-HNE) as a function of time after addition of 10 mM DEM. Although GSH depletion reaches a maximum after 15 min of incubation (not shown), the buildup of LPO products does not begin until 60 min and accumulation of PCOs is not detected until the 90 min time point. These findings are not unique to this in vitro model of oxidative stress since almost identical results are obtained when oxidative stress is induced by addition of the mitochondrial toxin rotenone (Supplementary Figure 1). The time lag between the appearance of oxidized lipids and proteins could be due to the need for accumulating a considerable amount of lipid hydroperoxides before PCOs can be detected or most likely to the presence of proteolytic systems that remove carbonylated proteins. Over time, these systems may become overwhelmed by the constant production of oxidized protein substrates or just inactive.
Time-course of lipid and protein oxidation in DEM-treated slices. Rat brain slices were incubated with 10 mM DEM for various periods of time (0–180 min). After incubation, slices were homogenized in HEN buffer and aliquots of the homogenate were used to determine PCOs and LPO products (MDA + 4-HNE) as described under “Materials and Methods”. Values represent mean ± SEM of three experiments
Differential Effect of Epoxomicin and Lactacystin on Protein Carbonylation
Proteasomes and the Lon protease are known to remove oxidized proteins in the cytoplasm and mitochondria, respectively [17, 18]. To determine the role of these proteolytic systems in the removal of carbonylated proteins in GSH-depleted slices, we tested the effect of Epo (proteasome inhibitor) [19] and Lac (proteasome and Lon protease inhibitor) [20]. As depicted in Fig. 2, incubation with Epo has no effect on the accumulation of carbonylated proteins in DEM-treated slices while Lac more than doubles the buildup of oxidized proteins. Importantly, at the concentration used (5 µM), both drugs were found to reduce the chymotrypsin-like activity of the 20S proteasome by > 95% (data not shown). Thus, it is fair to conclude that the half-lives of most damaged proteins are relatively short and that their removal is carried out by the Lon protease. This finding agrees with the fact that most carbonylated proteins in this model are in mitochondria and not the cytoplasm (see below). In the absence of DEM, neither Epo nor Lac causes carbonyl accumulation, suggesting that basal oxidation is low.
Effect of proteasome inhibitors on DEM-induced protein carbonylation. Rat brain slices were incubated with 10 mM DEM for 2 h in the absence or presence of either 5 µM Epo or 10 µM Lac. After incubation, slices were homogenized in HEN buffer and aliquots of the homogenate were used to determine PCOs as described under “Materials and Methods”. Values represent the mean ± SEM of three experiments. Asterisk denotes the values that are significantly higher (p < 0.05) from those in DEM-treated slices
Inhibition of Protein Synthesis has no Effect on Protein Carbonylation
It has been suggested that carbonylation may serve as a tagging system to direct misfolded proteins, derived from transcriptional and translational errors, to the proteolytic apparatus [21]. To determine whether or not carbonylation occurs mostly on newly synthesized proteins, slices were incubated with 10 mM DEM in the absence or presence of 100 µM cycloheximide. At this concentration, cycloheximide reduces protein synthesis in brain sections by > 95% within 10 min [22]. As shown in Fig. 3, inhibition of protein synthesis does not have any appreciable effect on the oxidation of the bulk of proteins, indicating that carbonylation occurs mostly on pre-existing protein molecules.
Effect of cycloheximide on DEM-induced protein carbonylation. Rat brain slices were incubated with 10 mM DEM for 2 h in the absence or presence of 100 µM cycloheximide (CHX). After incubation, slices were homogenized in HEN buffer and aliquots of the homogenate were used to determine PCOs as described under “Materials and Methods”. Values are expressed as percent of controls and represent the mean ± SEM of three experiments. Asterisk denotes values that are significantly different (p < 0.05) from those in untreated (control) slices. NS not significant
GGS and AAS are the Major Carbonylated Amino Acids in Proteins
To identify the carbonyl-containing amino acid residues, proteins were treated with FlNH2 before acid hydrolysis and analysis by HPLC. As shown in Fig. 4, HPLC analysis of FlNH2-labeled amino acids derived from oxidized albumin gives two major peaks that have been previously identified as AAS and GGS [23]. These two amino acids, albeit in different proportions, are the only ones present in proteins from tissue slices incubated for 3 h with 10 mM DEM, indicating that metal catalyzed oxidation of lysine, arginine and proline is the major process by which proteins are carbonylated upon GSH depletion. This finding and our previous observation that protein oxidation is inhibited by LPO scavengers strongly suggest the occurrence of lipid hydroperoxide-mediated protein carbonylation.
Identification of the carbonylated amino acids by HPLC. Oxidized albumin and proteins from slices incubated with 10 mM DEM for 3 h were labeled with fluoresceinamine and, after acid hydrolysis, the amino acids were separated by HPLC as described under “Material and Methods”. The absorbance was measured at 490 nm and tracks the fluorescein-containing products. The peaks at 7.7 and 9.1 min are the decarboxylated fluoresceinamine derivatives of GGS and AAS, respectively. The peak at 10.9 min is decarboxylated fluoresceinamine. This peak was also observed when free FlNH2 was subjected to the HCl treatment used for peptide-bond hydrolysis
Inhibition of PLA2 and LOXs/COXs has no Effect on Protein Carbonylation
We next aimed to determine whether lipid hydroperoxides are formed by non-enzymatic oxidation of polyunsaturated fatty acids present in complex lipids or by enzymatic oxidation of free arachidonic acid via LOXs and COXs. As shown in Fig. 5, incubation of brain slices for 2 h with the PLA-2 inhibitors AACOF3 and Aristo A has no effect DEM-induced protein oxidation, indicating that the release of arachidonic acid from phospholipids is not required. We next tested the effect of LOX/COX inhibitor ETYA acid on protein carbonylation. This drug was used a concentration of 300 µM, which is much higher than those needed to produce 50% inhibition of 5-LOX (10 µM), 12-LOX (0.3 µM), 15-LOX (0.2 µM) and COX1/2 (8 µM) in cells [24, 25]. Under these conditions, the inhibitor has no effect on protein carbonylation, indicating that enzymatically-produced lipid hydroperoxides are not involved in the oxidation of most proteins. Surprisingly, the 5-LOX inhibitor AA-861 completely prevents protein carbonylation induced by GSH depletion. Because AA-861 structure is similar to that of ubiquinone, we hypothesized that this drug could also be reducing mitochondria-produced ROS and indirectly carbonyl formation. To address this issue, we studied the effect of increasing concentrations (1–25 µM) of idebenone, an AA-861 analogue that does not contain the ethynic bond and therefore lacks LOX/COX inhibitory activity (Supplementary Figure 2). As shown in Fig. 6, the concentration-dependent reduction in the amount of DEM-induced protein carbonyls caused by idebenone and AA-861 are identical, indicating that the latter acts mostly as an antioxidant.
Effect of PLA2 and LOX/COX inhibitors on DEM-induced protein carbonylation. Rat brain slices were incubated for 2 h with 10 mM DEM in the absence or presence of 200 µM Aristo A, 300 µM ETYA, 200 µM AACOF3 or 200 µM AA-861. After incubation, slices were homogenized in HEN buffer and aliquots of the homogenate were used to determine the PCOs by western blot as described under “Material and Methods”. Values are expressed as % of DEM-treated and represent the mean ± SEM of 3–4 experiments. Asterisk denotes values that are significantly different (p < 0.05) from those in DEM-treated slices
Effect of increasing concentrations of AA-861 and idebenone on DEM-induced protein carbonylation. Rat brain slices were incubated for 2 h with 10 mM DEM in the absence or presence of three different concentrations of idebenone or AA-861. After 2 h, slices were homogenized in HEN buffer and aliquots of the homogenate were used to determine the PCOs by western blot as described under “Material and Methods”. Values are expressed as % of DEM-treated and represent the mean ± SEM of three experiments. Asterisks denote values that are significantly different (p < 0.05) from those in DEM-treated slices
Most Carbonylated Proteins are Present in Mitochondria
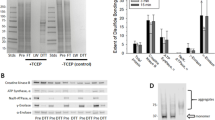
To determine whether the oxidized proteins are confined to specific organelles, slices were incubated for 3 h in the absence or presence of 10 mM DEM, followed by subcellular fractionation and analysis of protein carbonylation by western blotting. As shown in Fig. 7a, cellular mitochondria have the highest concentration of PCOs followed by the mitochondria- containing synaptosomal fraction (Fig. 7b) and microsomes, while cytosolic and myelin proteins are not carbonylated to any appreciable extent. A similar subcellular distribution of PCOs was obtained in rotenone-treated slices (Supplementary Figure 3a). The lack of significant protein oxidation in the cytosolic and myelin fractions is probably due to the absence of lipids in the former and the low levels of polyunsaturated acyl chains in the latter (Fig. 7c), which would lead to a diminished production of lipid hydroperoxides. While mitochondria, synaptosomes and microsomes all have elevated amounts of polyunsaturated fatty acids (Fig. 7c), mitochondrial proteins are the most oxidized. This is likely due to the fact that this organelle produces the most ROS (see below) and most importantly contains high amounts of transition metals. Indeed, when tissue slices are incubated with Fe2+, proteins from all three fractions are carbonylated to a similar degree (Supplementary Figure 3b).
Subcellular distribution of PCOs in DEM-treated slices. Rat brain slices were incubated for 3 h in the absence or presence of 10 mM DEM. After incubation, slices were collected and the various subcellular fractions were prepared by ultracentrifugation on a sucrose gradient. Aliquots from each subcellular fraction were used to determine the PCOs as described under “Materials and Methods”. TH total homogenate, My myelin, Syn synaptosomes, Mit mitochondria, Mic microsomes, Cyt cytosol. a Representative oxyblot. b distribution of organelle-specific markers in the various fractions. c membrane peroxidizability index calculated from the fatty acid values shown in Supplementary Table 1
Inhibition of the MPTP Prevents Protein Carbonylation
To determine the extent of the ROS that come from mitochondria, brain slices were incubated with DEM in the absence or presence of two different mitochondria permeability transition pore (MPTP) inhibitors: CsA and BkA. As shown in Fig. 8, addition of 5 µM CsA prevents protein carbonylation induced by GSH depletion almost completely, indicating that most free radicals in this experimental model of oxidative stress are coming from mitochondria. Surprisingly, addition of BkA has no effect on carbonylation. However, this toxin is known to have a biphasic effect on permeability transition, initially increasing the stability of mitochondrial membrane followed by pronounced opening of the pore [26]. Thus, it is fair to conclude that protein oxidation/aggregation occurs after MPTP opening.
Effect of MPTP inhibitors on DEM-induced protein carbonylation. Rat brain slices were incubated with 10 mM DEM for 2 h in the absence or presence of 50 µM BkA or 50 µM CsA. After incubation, slices were homogenized in HEN buffer and aliquots of the homogenate were used to determine PCOs as described under “Materials and Methods”. Values represent mean ± SEM of three experiments. Asterisks denote values that are significantly different (p < 0.05) from those in the absence of DEM
Discussion
The present work is part of a series of studies from our laboratory using DEM-treated brain sections to elucidate the mechanism of protein carbonylation induced by acute GSH depletion [8,9,10]. Herein, we show that (a) there is a significant lag between the appearance of carbonylated proteins and GSH depletion, which is likely due to removal of oxidized species early on in the incubation by the mitochondrial Lon protease, (b) the bulk of carbonylation occurs mostly on pre-existing proteins, (c) GGS and AAS are the only carbonyl-containing amino acid residues detected by HPLC, indicating that proteins are carbonylated by a direct (oxidative) mechanism of lysine, arginine and proline residues, (d) lipid hydroperoxides do not derive from enzymatic oxidation of arachidonic acid or other polyunsaturated fatty acids, (e) most of the carbonylated proteins are present in mitochondria, and (f) the majority of ROS in this paradigm arise from this organelle after permeability transition pore opening.
The mechanism linking GSH depletion and increased production of ROS by mitochondria has been studied extensively [27, 28]. A decline in the GSH/GSSG ratio is known to open the MPTP by oxidation of a critical dithiol in the voltage-sensing region of this protein complex [29]. When GSH falls below a certain level, permeability transition occurs followed by a collapse in mitochondrial membrane potential. This event causes enhanced production of superoxide either from the rise in redox cycling of ubiquinone within complex III [30] or from the reverse electron transport from succinate to NADH within complex I [31]. Loss of cytochrome c from mitochondria, after permeability transition and swelling, can also augment the production of oxygen free radicals by reducing the redox centers upstream of complex IV [32]. Our data are in agreement with this model since both MPTP inhibition by CsA and addition of ubiquinone analogs (idebenone and AA-861) prevent protein oxidation. These data also indicate that the majority of the ROS in GSH-depleted brain sections comes from mitochondria.
Proteolysis is considered the only process to eliminate PCOs since demonstrations of enzymatic systems capable of reducing protein-bound carbonyl groups are ambiguous at best [33]. Recent studies suggest that some proteins might be decarbonylated by a mechanism that involves the thioredoxin/thioredoxin reductase system [34] or glutaredoxins [35], and studies are underway in our laboratory to determine if such mechanisms also operate for the bulk of carbonylated protein in the tissue sections. However, the finding that Lac increases the accumulation of carbonylated proteins strongly supports the idea that proteolysis via the mitochondrial Lon protease is a major mechanism for disappearance of PCOs during severe GSH depletion. Lon protease is rapidly inactivated by peroxynitrite in vitro leading to electron transport failure [36], and dysfunctional Lon protease has been recently linked to accumulation of oxidized mitochondrial proteins in mice treated with the neurotoxin 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine [37]. Thus, it will be important to determine if this protease loses activity during the course of incubation with DEM and if that is what ultimately causes PCO accumulation in mitochondria. It is important to note that our findings do not minimize the role played by proteasomes and other proteolytic systems in the removal of oxidized proteins. Indeed, we have found that proteasomes are partially responsible for the removal of carbonylated proteins in lipopolysaccharide-treated astrocytes [38] and in PC12 cells incubated for 12 h with low concentrations (50 µM) of DEM [39].
Carbonyl groups are introduced into proteins by two distinct mechanisms: oxidative (direct) and non-oxidative (indirect). Oxidative mechanisms, which are metal ion-catalyzed, involve the direct reaction of hydrogen peroxide or lipid hydroperoxides with Lys, Arg, and Pro residues to produce AAS and GGS [33]. Non-oxidative carbonylation of proteins entails the reaction of the nucleophilic centers in Cys, His or Lys residues with RCS derived from the oxidation of lipids (e.g. 4-HNE, MDA, acrolein) and carbohydrates (e.g. glyoxal, methylglyoxal) [33]. Our present findings that AAS and GGS are the only modified amino acids in DEM-treated brain slices and the failure to detect RCS-protein adducts [10] imply that protein carbonylation in this model of oxidative stress occurs via a direct mechanism. Furthermore, the observations that (a) LPO scavengers prevent DEM-induced protein carbonylation [9], (b) incubation of brain proteins with the lipid hydroperoxide 15-hydroperoxy-eicosatetraenoic acid causes a marked increase in PCOs [9] and (c) protein oxidation is highest in organelles rich in polyunsaturated fatty acids (this study) strongly suggest that lipid hydroperoxides, and not hydrogen peroxide, are the oxidants. Lipid hydroperoxide-induced protein carbonylation was initially proposed by Refsgaard et al. [40], who discovered that metal ion-catalyzed oxidation of proteins is greatly enhanced by addition of polyunsaturated fatty acids. It was speculated that alkoxyl radicals, derived from metal-catalyzed heterolytic cleavage of lipid hydroperoxides [41], are responsible for the introduction of carbonyls into proteins. Lipid-derived alkoxyl radicals seem to be involved in the oxidation of retinal proteins [42] suggesting that this mechanism is more common than previously thought. Future studies will identify the lipid hydroperoxide species involved in the carbonylation of major brain proteins.
Abbreviations
- COX:
-
Cyclooxygenase
- DEM:
-
Diethyl maleate
- GSH:
-
Glutathione
- 4-HNE:
-
4-hydoxy-2-nonenal
- LOX:
-
Lipooxygenase
- LPO:
-
Lipid peroxidation
- MDA:
-
Malondialdehyde
- PCO:
-
Protein carbonyl
- RCS:
-
Reactive carbonyl species
- ROS:
-
Reactive oxygen species
References
Stadtman ER, Berlett BS (1997) Reactive oxygen-mediated protein oxidation in aging and disease. Chem Res Toxicol 10:485–494
Levine RL (2002) Carbonyl modified proteins in cellular regulation, aging, and disease. Free Radic Biol Med 32:790–796
Aksenov MY, Aksenova MV, Butterfield DA, Geddes JW, Markesbery WR (2001) Protein oxidation in the brain in Alzheimer’s disease. Neuroscience 103:373–383
Floor E, Wetzel MG (1998) Increased protein oxidation in human sustantia nigra pars compacta in comparison with basal ganglia and prefrontal cortex measured with an improved dinitrophenylhydrazine assay. J Neurochem 70:268–275
Ferrante RJ, Browne SE, Shinobu LA, Bowling AC, Baik MJ, MacGarvey U, Kowall NW, Brown RH, Beal MF (1997) Evidence of increased oxidative damage in both sporadic and familial amyotrophic lateral sclerosis. J Neurochem 69:2064–2074
Bizzozero OA, DeJesus G, Callahan K, Pastuszyn A (2005) Elevated protein carbonylation in the brain white matter and gray matter of patients with multiple sclerosis. J Neurosci Res 81:687–695
Hilgart AA, Bizzozero OA (2008) Carbonylation of major cytoskeletal proteins in multiple sclerosis. J Neurochem 104(Suppl.1):123
Bizzozero OA, Ziegler JL, DeJesus G, Bolognani F (2006) Acute depletion of reduced glutathione causes extensive carbonylation of rat brain proteins. J Neurosci Res 83:656–667
Bizzozero OA, Reyes S, Ziegler JL, Smerjac S (2007) Lipid peroxidation scavengers prevent the carbonylation of cytoskeletal brain proteins induced by glutathione depletion. Neurochem Res 32:2114–2122
Zheng J, Bizzozero OA (2010) Traditional reactive carbonyl scavengers do not prevent the carbonylation of brain proteins induced by acute glutathione depletion. Free Radic Res 44:258–266
Bizzozero OA, Zheng J (2010) Mechanism of protein carbonylation during glutathione depletion. Trans Am Soc Neurochem PSM01–P10
Gérard-Monnier D, Erdelmeier I, Régnard K, Chaudiére J (1989) Reactions of 1-methyl-2-phenylindole with malondialdehyde and 4-hydroxyalkenals. Chem Res Toxicol 11:1176–1183
Bizzozero OA, Leyba J, Nuñez DJ (1992) Characterization of proteolipid protein fatty acylesterase from rat brain myelin. J Biol Chem 267:7886–7894
Pamplona R, Portero-Otin M, Riba D, Ruiz C, Prat J, Bellmunt MJ, Barja G (1998) Mitochondrial membrane peroxidizability index is inversely related to maximum life span in mammals. J Lipid Res 39:1989–1994
Climent I, Tsai L, Levine RL (1989) Derivatization of γ-glutamyl semialdehyde residues in oxidized proteins by fluoresceinamine. Anal Biochem 182:226–232
Rodgers KJ, Dean RT (2003) Assessment of proteasome activity in cell lysates and tissue homogenates using peptide substrates. Int J Biochem Cell Biol 35:716–727
DeMartino GN, Slaughter CA (1999) The proteasome, a novel protease regulated by multiple mechanisms. J Biol Chem 274:22123–22136
Bender T, Leidhold C, Ruppert T, Franken S, Voos W (2010) The role of protein quality control in mitochondrial protein homeostasis under oxidative stress. Proteomics 10:1426–1443
Meng L, Mohan R, Kwok BHB, Eloffson M, Sin N, Crews CM (1999) Epoxomixin, a potent and selective proteasome inhibitor, exhibits in vivo anti-inflammatory activity. Proc Natl Acad Sci USA 96:10403–10408
Bayot A, Basse N, Lee I, Gareil M, Pirotte B, Bulteau AL, Friguet B, Reboud-Ravaux M (2008) Towards the control of intracellular protein turnover: mitochondrial Lon protease inhibitors versus proteasome inhibitors. Biochimie 90:260–269
Dukan S, Farewell A, Ballesteros M, Taddei F, Radman M, Nyström T (2000) Protein oxidation in response to increased transcriptional or translational errors. Proc Natl Acad Sci USA 97:5746–5749
Bizzozero OA, Soto EF, Pasquini JM (1983) Myelin proteolipid protein is not esterified at the site of synthesis. Neurochem Int 5:729–736
Daneshvar B, Frandsen H, Autrupand H, Dragsted LO (1997) γ–Glutamyl semialdehyde and 2-amino-adipic semialdehyde: biomarkers of oxidative damage to proteins. Biomarkers 2:117–123
Tobias LD, Hamilton JG (1979) The effect of 5,8,11,14-eicosatetraynoic acid on lipid metabolism. Lipids 14:181–193
Takami M, Preston SL, Behrman HR (2000) Eicosatetraynoic and eicosatriynoic acids, lipoxygenase inhibitors, block meiosis via antioxidant action. Am J Physiol 278:C646–C650
Gizatullina ZZ, Chen Y, Zierz S, Gellerich FN (2005) Effects of extramitochondrial ADP on permeability transition of mouse liver mitochondria. Biochim Biophys Acta 1706:98–104
Armstrong JS, Jones DP (2002) Glutathione depletion enforces the mitochondrial permeability transition and causes cell death in Bcl-2-overexpressing HL60 cells. FASEB J 16:1263–1265
Shen D, Dalton TP, Nebert DW, Shertzer HG (2005) Glutathione redox state regulates mitochondrial reactive oxygen production. J Biol Chem 280:25305–25312
Petronilli V, Costantini P, Scorrano L, Colonna R, Passamonti S, Bernardi P (1994) The voltage sensor of the mitochondrial permeability transition pore is tuned by the oxidation-reduction state of vicinal thiols. Increase of the gating potential by oxidants and its reversal by reducing agents. J Biol Chem 269:16638–16642
Chen Q, Vazquez EJ, Moghaddas S, Hoppel CL, Lesnefsky EJ (2003) Production of reactive oxygen species by mitochondria: central role of complex III. J Biol Chem 278:36027–36031
Lambert AJ, Brand MD (2004) Superoxide production by NADH:ubiquinone oxidoreductase (complex I) depends on the pH gradient across the mitochondrial inner membrane. Biochem J 382:511–517
Votyakova TV, Reynolds IJ (2005) Ca2+-induced permeabilization promotes free radical release from rat brain mitochondria with partially inhibited complex I. J Neurochem 93:526–537
Bizzozero OA (2009) Protein carbonylation in neurodegenerative and demyelinating CNS diseases. In: Lajtha A Banik N, Ray S (eds) Handbook of neurochemistry and molecular neurobiology, Springer, New York, pp 543–562
Wong CM, Cheema AK, Zhang L, Suzuki YJ (2008) Protein carbonylation as a novel mechanism in redox signaling. Circ Res 102:310–318
Wong CM, Marcocci L, Das D, Wang X, Luo H, Zungu-Edmondson M, Suzuki YJ (2013) Mechanism of protein decarbonylation. Free Radic Biol Med 65:1126–1133
Stanyer L, Jorgensen W, Hori O, Heales S (2008) Inactivation of brain mitochondrial Lon protease by peroxynitrite precedes electron transport chain dysfunction. Neurochem Int 53:95–102
Bulteau AL, Mena NP, Auchere F, Lee I, Prigent A, Lobsiger CS, Camadro JM, Hirsch EC (2017) Dysfunction of mitochondrial Lon protease and identification of oxidized proteins in mouse brain following exposure to MPTP: implication for Parkinson disease. Free Radic Biol Med 108:236–246
Zheng J, Bizzozero OA (2010) Accumulation of protein carbonyls within cerebellar astrocytes in murine experimental autoimmune encephalomyelitis. J Neurosci Res 88:3376–3385
Dasgupta A, Zheng J, Bizzozero OA (2012) Protein carbonylation and aggregation precede neuronal apoptosis induced by partial glutathione depletion. ASN Neuro 4(3):e00084. https://doi.org/10.1042/AN20110064
Refsgaard HF, Tsai L, Stadman ER (2000) Modification of proteins by polyunsaturated fatty acid peroxidation products. Proc Natl Acad Sci USA 97:611–691
Davies MJ, Slater TF (1987) Studies on the metal-ion and lipoxygenase-catalysed breakdown of hydroperoxides using electron-spin-resonance spectroscopy. Biochem J 245:167–173
Pennathur S, Ido Y, Heller JI, Byun J, Danda R, Pergola P, Wil-liamson JR, Heinecke JW (2005) Reactive carbonyls and poly-unsaturated fatty acids produce hydroxyl radical-like species. J Biol Chem 280:22706–22714
Acknowledgements
This work was supported by PHHS grant NS057755 from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zheng, J., Hu, CL., Shanley, K.L. et al. Mechanism of Protein Carbonylation in Glutathione-Depleted Rat Brain Slices. Neurochem Res 43, 609–618 (2018). https://doi.org/10.1007/s11064-017-2456-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11064-017-2456-9