Abstract
The current study aimed to characterize SNF5 expression and investigate the relationship between SNF5 and clinicopathological features in skull base chordoma. 48 patients diagnosed with skull base chordoma were enrolled in this study. Tissue microarray and immunohistochemistry were performed to evaluate the expression of SNF5 in skull base chordoma. Kaplan–Meier survival analysis was used to assess survival. Multivariable Cox regression analysis was used to identify risk factors affecting patient survival. The H-scores for cytoplasmic SNF5 ranged from 124.47 to 254.52. Low expression of SNF5 was correlated with shorter overall survival (OS) (p = 0.021). Patients with age > 55 years old had shorter progression free survival (PFS) and OS times than patients whose age ≤ 55 years old (p = 0.005 and 0.003, respectively). The gross total resection group showed longer PFS than the non-gross total resection group (p = 0.024). Females showed shorter PFS times than males (p = 0.033). Multivariable Cox regression analysis showed that age, extent of resection and sex were independent prognostic factors for PFS (p = 0.010, 0.013 and 0.042, respectively). Age was an independent prognostic factor for OS (p = 0.010). Our study indicate that low expression of SNF5 is associated with poor prognosis in skull base chordoma.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chordoma, a rare malignant neoplasm that comprises 1–4% of primary bone tumours, arises in the axial bones, and as much as 35% of chordoma locates in the clivus region [1]. In particular, chordoma is believed to originate from notochordal remnants [2]. The majority of chordoma occurs in adults between the ages of 50 and 60, with less than 5% occurring in infants and children [3]. Chordoma is a slow-growing tumor with low grade malignancy, although effective therapies for chordoma are lacking. Chordoma is often located close to critical neurovascular structures, particularly in skull base chordoma and it can invade surrounding tissues, making total resection difficult to achieve and contributing to the high local recurrence rate. Chordoma is highly resistant to conventional chemotherapy and radiotherapy, making therapy especially arduous. Recently, proton therapy, photon radiation and target therapy have been combined as a comprehensive treatment strategy to ameliorate chordoma progress [4]. However, we currently have a poor understanding of the tumorgenesis and molecular biological characteristics of chordoma.
The SWI/SNF complex is a 1.14 MDa multi-subunit ATP-dependent complex that regulates nucleosome positioning and chromatin remodelling [5, 6]. A growing body of evidence indicates that this complex plays a crucial role in many biological behaviours, including cell proliferation, differentiation and transcription [7]. The tumor suppressor gene SNF5 is the core subunit of the mammalian SWI/SNF chromatin remodeling complex. Paediatric rhabdoid tumours, which are a class of rare malignant tumors mostly occurring in brain with poor prognosis, are often characterized by the inactivation of SNF5. Additionally, germline mutations in SNF5 are found in familial rhabdoid tumours [8, 9]. Studies have also shown that SNF5 is deficient in cribriform neuroepithelial tumours, paediatric choroid plexus carcinomas and epithelioid sarcomas [10,11,12].
There is currently little known about SNF5 expression in skull base chordoma or the relationship between SNF5 expression and clinical features in skull base chordoma. Therefore, we chose to characterize expression of the tumour suppressor gene SNF5 in skull base chordoma.
Materials and methods
Patients
From January 2008 to September 2014, 48 patients who were admitted for surgery at Beijing Tiantan Hospital, Capital Medical University and were histopathologically confirmed as having chordoma were involved in this study. Every chordoma enrolled was located at the skull base. The average follow-up interval was 39.8 months (median 30months, range 9–98 months). This study was approved by the ethics committee of Beijing Tiantan Hospital, Capital Medical University.
Degree of resection
Preoperative and postoperative MR images and surgical records were provided to evaluate the extent of resection, which were defined as follows: gross total resection (visible tumour was removed and no remnant tumour in the postoperative MR images); and non-gross total resection, which contained subtotal resection (≥ 95% of tumour resection based on postoperative MR images) and partial resection (< 95% of tumour resection based on postoperative MR images) [13].
Tissue microarray
Formalin-fixed and paraffin-embedded tissue specimens from all 48 patients were assayed by tissue microarray (TMA) using the Tissue Array MiniCore (ALPHELYS, Plaisir, France). Two pathologists viewed the haematoxylin-eosin stained slides, and the two most representative 2-mm cores from every tissue slide were selected and removed to a new slide to perform the TMA. We used a Leica RM 2135 Rotary Microtome (Rankin, Wetzlar, Germany) to cut 4-mm sections from the TMA for immunohistochemical staining.
Immunohistochemistry and evaluation
The Leica BOND III automated system and Bond Polymer Refine Detection kit (Leica) were used for immunohistochemical staining. After baking at 65 °C for 0.5 h, the slides were deparaffinized with dimethylbenzene and then hydrated in a graded ethanol series followed by antigen retrieval. To block endogenous peroxide, the slides were treated with 0.3% H2O2. After incubation with the primary antibody (anti-SNF5 antibody, Abcam, 58209), the slides were treated with the Bond Polymer Refine Detection kit (Leica), according to the manufacturer’s protocol. Diaminobenzidine was used as the chromogen. The slides were then dehydrated, cleared and mounted. We replaced the primary antibody with phosphate-buffered saline for the negative control.
We used a cytoplasmic v9 automated algorithm to evaluate the immunostained slides after scanning with a Leica Aperio AT2 scanner and analysis with Leica Aperio ImageScope v12.3.0.5056. Immunostaining was scored as none (0), weak (1+), moderate (2+) or strong (3+), and the tumour regions were determined by a single pathologist. The H-score was calculated in the conventional manner using the formula: \({\text{H - scores}} = 1 \times \left( {{\text{percent of }}1 + {\text{cell}}} \right) + 2 \times \left( {{\text{percent of }}2 + {\text{cell}}} \right) + 3 \times ({\text{percent of }}3 + {\text{cell}})\), with values ranging from 0 to 300. We chose the median of the H-score as the cut-off value for separating patients into two groups: high or low SNF5 expression.
Statistical analysis
All statistical analyses were performed using SPSS 19.0 (IBM Corporation, USA). The χ2 test, 2 independent-samples t-test and Wilcoxon Mann–Whitney test were used to identify the relationships between SNF5 expression and clinicopathological characteristics. Kaplan–Meier curves and the log-rank test were applied for univariable survival analysis to find the differences between groups. Statistically significant variables were further analysed by multivariable Cox regression analysis. p < 0.05 was regarded as statistically significant. GraphPad Prism 5.0 (GraphPad, USA) was used to create the graphs.
Results
Clinical characteristics
In total, 35 males and 13 females (male: female = 2.69: 1) were included in this study. Only patients with skull base chordoma were enrolled in the present study. The age of the patients ranged from 13 to 69, with a mean age (± SD) of 44.25 (± 13.39). The most common complaints were diplopia (23/48), blurred vision (20/48) and headache (20/48). The mean duration was 12.5 months (median 4 months, range 0.5–120 months). 39 cases were primary tumours compared with 9 cases with prior operative history. 46 cases were conventional chordoma and the remaining 2 patients had chondroid chordoma; no dedifferentiated chordoma cases were investigated in this study. The endoscopic transphenoidal approach was used in 31 cases (64.6%), and the craniotomy approach was used in 17 cases (35.4%). Gross total resection was achieved in 10 (20.8%) patients and non-gross total resection was achieved in 38 (79.2%) patients. Surgery complications occurred in 12 cases. The most frequent complication was cranial nerve paralysis (6/48), followed by intracranial infection (5/48) (Table 1).
Relationship between SNF5 expression and clinicopathological features
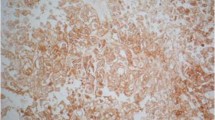
SNF5 expression was observed in the cytoplasm and the H-scores of SNF5 ranged from 124.47 to 254.52 (median 182.48) (Fig. 1). Furthermore, we analysed the relationship between SNF5 expression and clinical features. The mean age (± SD) in the high SNF5 expression group was 45.79(± 10.23) years, whereas in the low SNF5 expression group, the mean age (± SD) was 42.71(± 16.03) years (p = 0.431). There were 16 males (16/24) in the high SNF5 expression group and 19 males in the other group (p = 0.330). In total, 7/24 patients received gross total resection in the high SNF5 expression group whereas 3/24 received gross total resection in the low SNF5 expression group (p = 0.155). We found no significant relationships between SNF5 expression and blood supply, tumour texture, surgical approach, course of disease, or tumour volume. However, we note that patients with prior operative history tended to show lower SNF5 expression than patients without prior operative history (Table 1).
Factors affecting OS
For survival analysis, we defined the period between initial surgery and death or the date of the last follow up as the overall survival (OS) time. The mean OS in present study was 80.8 months. There were 8 patients who died during the follow-up period, 7 of which were in the low SNF5 expression group and 1 in the high expression group. Patients with high SNF5 expression showed longer OS [3-year survival rate: 95.7%; mean OS: 87.5 months, 95% confidence interval (CI) 80.9–94.2 months] compared with patients with low expression of SNF5 (3-year survival rate: 74.3%; mean OS: 67.5 months, 95% CI 49.9–85.2 months), with a p value of 0.021 (Fig. 2a).
Gross total resection was performed in ten patients, and none of those patients died during the follow-up. In the non-gross total resection group, 8/38 patients died (p = 0.108) (Fig. 2b).
To transform age into a categorical variable for analysis, 55 was chosen as the cut-off age, based on our previous study [14]. 5/9 patients died in the age > 55 group, whereas 3/39 patients died in the age ≤ 55 group. Consistent with previous studies, the mean OS in the age > 55 group was 49.5 months (95% CI 28.8–70.1 months), which was notably shorter than in the age ≤ 55 group (91.1 months, 95% CI 83.7–98.6 months); the 3-year survival rates were 64.8 and 94.7%, respectively. Additionally, the log-rank test had a p value of 0.003 between the two groups (Fig. 2c).
When considering OS, men showed 12 month longer mean OS times (83.4 months, 95% CI 71.8–95.0 months) than women (71.5 months, 95% CI 52.4–90.7 months), although this difference was not statistically significant (p = 0.354).
During the follow-up, 30 patients suffered from recurrence. Patients with recurrence of chordoma had significantly shorter OS times than in the no recurrence group (p = 0.037) (Table 2). Age, SNF5 expression and recurrence were subjected to Cox multivariable regression analysis, and we found that age was an independent prognostic factor for OS (p = 0.010) (Table 3).
Factors affecting PFS
Progression-free survival (PFS) was defined as the period from the first surgery to the date of progression or last follow-up. The median PFS in the present study was 24.0 months. For the age > 55 group, all patients (9/9) showed the recurrence of skull base chordoma, whereas in the age ≤ 55 group, the recurrence rate was 53.8% (21 cases). Kaplan–Meier survival analysis indicated that patients in the age > 55 group had shorter PFS periods than those in the age ≤ 55 group (median PFS 10 vs. 30 months, respectively, p = 0.005) (Fig. 2d).
We then analysed the relationship between SNF5 expression and PFS. The median PFS in the high SNF5 expression group was 58 months, which was 3.6 times as long as that in the low SNF5 expression group (16 months); however, no statistically significant difference was observed between the two groups (p = 0.290) (Table 2).
We next explored the relationship between sex and PFS. Interestingly, in the male group, 19/35 patients showed recurrence during the follow-up. By contrast, 11/13 female patients showed recurrence. The median PFS in male patients was 30 months, and the median PFS in female patients was 15 months (p = 0.033) (Fig. 2.e).
We also took surgery resection into consideration, and we found that 40.0% (4/10) of patients in the gross total resection group experienced progression. In the non-gross total group, the rate of progression was 68.4% (26/38). Additionally, the median PFS was 76 months for the gross total resection group, and 18 months for the non-gross total resection group (p = 0.024) (Fig. 2f).
Age, extent of resection and sex were included in the Cox multivariable regression analysis, and all three factors were found to be independent prognostic factors for PFS (p = 0.010, 0.013 and 0.042, respectively) (Table 3).
Discussion
Chordoma is rare, slow-growing, infiltrative malignancy arising from notochordal remnants, with an annual incidence of 1/1,000,000 [15]. Total surgery resection combined with adjuvant radiation therapy has been shown to prolong patient survival [16]. Studies on target therapies in chordoma (e.g., cetuximab and imatinib) have been more common in recent years, although these have involved small numbers of patients. Despite comprehensive therapy, patients with chordoma have an extremely poor prognosis for local recurrence and metastases [3]. Therefore, there is a pressing need to understand the mechanisms of chordoma tumourigenesis and to search for novel prognostic factors and therapies to improve the outcome of patients with chordoma.
It is still controversial whether sex is a risk factor for patient survival. Some studies have shown that male patients had a worse prognosis than females, whereas others showed the opposite, with women having shorter survival times [16]. We found that men had longer PFS than women, although sex was not statistically related to OS.
Radical resection of skull base chordoma is a suggested treatment, and previous studies have shown that radical surgical resection had a positive effect on patient prognosis [16, 17]. Consistent with those studies, we found a significant difference between the two groups, and patients with gross total resection had a longer PFS than those in the other group.
Increasing attention is being paid to the connection between age and patients prognosis in chordoma. Whether age can be used as a prognostic predictor remains debatable, likely due to the variety of cut-off values and differences in the age distribution between cohorts enrolled in previous studies. We found that patients with age > 55 had a shorter PFS and OS times than younger patients. Age was also an independent prognostic factor for both PFS and OS in skull base chordoma.
The SWI-SNF complex was first identified in yeast and is found in both prokaryotes and eukaryotes. SWI-SNF complex consists of a core complex with multiple subunit proteins, and it can remodel chromatin by hydrolysing ATP [7]. The SWI/SNF chromatin-remodelling complex is considered to play a critical role in the initiation and development of cancer because it is linked to a number of tumour suppressor genes and oncogenes [18]. Additionally, previous studies have shown that the SWI/SNF complex acts as a tumour suppressor in various malignancies, including ovarian carcinoma, rhabdoid tumours and pancreatic cancer [9, 18, 19]. SWI/SNF complex plays essential roles in cell differentiation and cell growth processes such as neuronal differentiation, myeloid differentiation and muscle development [20, 21]. The complex is also involved in DNA synthesis, DNA repair and mitosis [7].
SNF5, located on chromosome 22q11.2, is a component of all variants of the SWI/SNF complex and is required for the chromatin remodelling [22]. It has been demonstrated that SNF5 is inactivated in various tumours, such as rhabdoid tumours, hepatoblastoma and paediatric undifferentiated sarcoma [12, 23, 24]. One recent study showed that SNF5 negatively regulates the expression of transforming growth factor-β1 (TGF-β1), affecting the tumourigenesis of hepatocellular carcinoma [25]. Lin et al. reported that the expression of SNF5 is decreased in melanoma, a human skin malignancy characterized by high invasiveness and metastasis. Additionally, it was demonstrated that loss of SNF5 was related to poor survival in patients with melanoma [26].
However, the expression profile of SNF5, at the level of RNA or protein, in chordoma tissues is not clear, and the detailed mechanisms of SNF5 function in chordoma cell lines have not been well characterized. Hence, there is a need for studies assessing SNF5 expression and biological function in chordoma. In the present study, we found that low expression of SNF5 in chordoma was associated with shorter OS, which we explain as follows. First, several pathways crucial for tumour formation and development are regulated by SNF5, such as the well-known tumour suppressor gene p53, and loss of SNF5 function likely leads to the inhibition of p53 translation via decreased EIF4E expression [27]. Additionally, the Hedgehog pathway, which is aberrantly activated by the loss of SNF5, is thought to induce oncogenesis [28]. Second, aberrant cell proliferation, cell differentiation and cell apoptosis due to loss of SNF5 could be present in tumours with low SNF5 expression. Previous studies showed that SNF5 regulates the p16 and retinoblastoma genes, a member of the pocket protein family that acts as a tumour suppressor gene, eventually leading to cell cycle arrest via repression of cyclin D1 and E2F in various tumour cell lines [29]. In addition, DNA repair may be inhibited in chordoma with low SNF5 expression. However, these detailed mechanisms remain to be investigated.
Until now, there have been few studies of SNF5 expression in chordoma. Previous studies have demonstrated that SNF5 expression is decreased in undifferentiated chordoma. Additionally, Antonelli et al. found a loss of SNF5 in 3/7 classic chordoma [30,31,32,33]. However, there were relatively little conventional chordoma involved in these studies, with a greater emphasis placed on loss of SNF5 in undifferentiated chordoma and the distinction between poor differentiated chordoma and atypical teratoid/rhabdoid tumours. Additionally, these studies paid little attention to the correlations between SNF5 expression and the clinical features of chordoma. In the present study, we investigated SNF5 expression in 48 patients with skull base chordoma via immunohistochemistry and analysed the relationship between SNF5 expression and clinicalpathological characteristics. To the best of our knowledge, the present study is the largest single-centre study on SNF5 expression in chordoma. We demonstrated that low SNF5 expression was correlated with poor survival in patients with skull base chordoma even though SNF5 seemingly failed to be an independent prognostic factor for OS (p = 0.051), which may be explained by the limited numbers of patients in present study; further researches with larger number of patients are required.
Conclusion
Our study reveals that low SNF5 expression in skull base chordoma is correlated with poor patient prognosis, suggesting that SNF5 may be a novel biomarker for predicting chordoma prognosis.
Abbreviations
- OS:
-
Overall survival
- PFS:
-
Progression free survival
- TMA:
-
Tissue microarray
- ATP:
-
Adenosine triphosphate
- EIF4E:
-
Eukaryotic initiation factor 4E
- HR:
-
Hazard ratio
- CI:
-
Confidence interval
References
Heffelfinger MJ, Dahlin DC, MacCarty CS, Beabout JW (1973) Chordomas and cartilaginous tumors at the skull base. Cancer 32(2):410–420
Salisbury JR (1993) The pathology of the human notochord. J Pathol 171(4):253–255. https://doi.org/10.1002/path.1711710404
Walcott BP, Nahed BV, Mohyeldin A, Coumans JV, Kahle KT, Ferreira MJ (2012) Chordoma: current concepts, management, and future directions. Lancet Oncol 13(2):e69–e76. https://doi.org/10.1016/S1470-2045(11)70337-0
Stacchiotti S, Sommer J, Chordoma Global Consensus G (2015) Building a global consensus approach to chordoma: a position paper from the medical and patient community. Lancet Oncol 16(2):e71–e83. https://doi.org/10.1016/S1470-2045(14)71190-8
Kingston RE, Narlikar GJ (1999) ATP-dependent remodeling and acetylation as regulators of chromatin fluidity. Genes Dev 13(18):2339–2352
Smith CL, Horowitz-Scherer R, Flanagan JF, Woodcock CL, Peterson CL (2003) Structural analysis of the yeast SWI/SNF chromatin remodeling complex. Nat Struct Biol 10(2):141–145. https://doi.org/10.1038/nsb888
Roberts CW, Orkin SH (2004) The SWI/SNF complex–chromatin and cancer. Nat Rev Cancer 4(2):133–142. https://doi.org/10.1038/nrc1273
Eaton KW, Tooke LS, Wainwright LM, Judkins AR, Biegel JA (2011) Spectrum of SMARCB1/INI1 mutations in familial and sporadic rhabdoid tumors. Pediatr Blood Cancer 56(1):7–15. https://doi.org/10.1002/pbc.22831
Versteege I, Sevenet N, Lange J, Rousseau-Merck MF, Ambros P, Handgretinger R, Aurias A, Delattre O (1998) Truncating mutations of hSNF5/INI1 in aggressive paediatric cancer. Nature 394(6689):203–206. https://doi.org/10.1038/28212
Hasselblatt M, Oyen F, Gesk S, Kordes U, Wrede B, Bergmann M, Schmid H, Fruhwald MC, Schneppenheim R, Siebert R, Paulus W (2009) Cribriform neuroepithelial tumor (CRINET): a nonrhabdoid ventricular tumor with INI1 loss and relatively favorable prognosis. J Neuropathol Exp Neurol 68(12):1249–1255. https://doi.org/10.1097/NEN.0b013e3181c06a51
Weber M, Stockhammer F, Schmitz U, von Deimling A (2001) Mutational analysis of INI1 in sporadic human brain tumors. Acta Neuropathol 101(5):479–482
Modena P, Lualdi E, Facchinetti F, Galli L, Teixeira MR, Pilotti S, Sozzi G (2005) SMARCB1/INI1 tumor suppressor gene is frequently inactivated in epithelioid sarcomas. Cancer Res 65(10):4012–4019. https://doi.org/10.1158/0008-5472.CAN-04-3050
Gui S, Zong X, Wang X, Li C, Zhao P, Cao L, Zhang Y (2016) Classification and surgical approaches for transnasal endoscopic skull base chordoma resection: a 6-year experience with 161 cases. Neurosurg Rev 39(2):321–332. https://doi.org/10.1007/s10143-015-0696-1 (discussion 332–323).
Zhai Y, Bai J, Wang S, Du J, Wang J, Li C, Gui S, Zhang Y (2017) Differences in dural penetration of clival chordomas are associated with different prognosis and expression of platelet-derived growth factor receptor-beta. World Neurosurg 98:288–295. https://doi.org/10.1016/j.wneu.2016.07.096
Chambers KJ, Lin DT, Meier J, Remenschneider A, Herr M, Gray ST (2014) Incidence and survival patterns of cranial chordoma in the United States. Laryngoscope 124(5):1097–1102. https://doi.org/10.1002/lary.24420
Samii A, Gerganov VM, Herold C, Hayashi N, Naka T, Mirzayan MJ, Ostertag H, Samii M (2007) Chordomas of the skull base: surgical management and outcome. J Neurosurg 107(2):319–324. https://doi.org/10.3171/JNS-07/08/0319
Wang L, Wu Z, Tian K, Wang K, Li D, Ma J, Jia G, Zhang L, Zhang J (2017) Clinical features and surgical outcomes of patients with skull base chordoma: a retrospective analysis of 238 patients. J Neurosurg. https://doi.org/10.3171/2016.9.JNS16559
Wilson BG, Roberts CW (2011) SWI/SNF nucleosome remodellers and cancer. Nat Rev Cancer 11(7):481–492. https://doi.org/10.1038/nrc3068
Samartzis EP, Noske A, Dedes KJ, Fink D, Imesch P (2013) ARID1A mutations and PI3K/AKT pathway alterations in endometriosis and endometriosis-associated ovarian carcinomas. Int J Mol Sci 14(9):18824–18849. https://doi.org/10.3390/ijms140918824
de la Serna IL, Carlson KA, Imbalzano AN (2001) Mammalian SWI/SNF complexes promote MyoD-mediated muscle differentiation. Nat Genet 27(2):187–190. https://doi.org/10.1038/84826
Seo S, Richardson GA, Kroll KL (2005) The SWI/SNF chromatin remodeling protein Brg1 is required for vertebrate neurogenesis and mediates transactivation of Ngn and NeuroD. Development 132(1):105–115. https://doi.org/10.1242/dev.01548
Wang X, Lee RS, Alver BH, Haswell JR, Wang S, Mieczkowski J, Drier Y, Gillespie SM, Archer TC, Wu JN, Tzvetkov EP, Troisi EC, Pomeroy SL, Biegel JA, Tolstorukov MY, Bernstein BE, Park PJ, Roberts CW (2017) SMARCB1-mediated SWI/SNF complex function is essential for enhancer regulation. Nat Genet 49(2):289–295. https://doi.org/10.1038/ng.3746
Russo P, Biegel JA (2009) SMARCB1/INI1 alterations and hepatoblastoma: another extrarenal rhabdoid tumor revealed? Pediatr Blood Cancer 52(3):312–313. https://doi.org/10.1002/pbc.21893
Judkins AR (2007) Immunohistochemistry of INI1 expression: a new tool for old challenges in CNS and soft tissue pathology. Adv Anat Pathol 14(5):335–339. https://doi.org/10.1097/PAP.0b013e3180ca8b08
Sun H, Zhong X, Wang C, Wang S, Lin L, Zou R, Wu Y, Sun N, Sun G, Wen T, Chi ZH, Zhao Y (2016) SNF5 is Involved in Suppression of Hepatocellular Carcinoma Progression via TGF-Beta 1 Signaling. Anat Rec 299(7):869–877. https://doi.org/10.1002/ar.23357
Lin H, Wong RP, Martinka M, Li G (2009) Loss of SNF5 expression correlates with poor patient survival in melanoma. Clin Cancer Res 15(20):6404–6411. https://doi.org/10.1158/1078-0432.CCR-09-1135
Yan HX, Zhang YJ, Zhang Y, Ren X, Shen YF, Cheng MB, Zhang Y (2017) CRIF1 enhances p53 activity via the chromatin remodeler SNF5 in the HCT116 colon cancer cell lines. Biochim Biophys Acta 1860(4):516–522. https://doi.org/10.1016/j.bbagrm.2017.02.006
Jagani Z, Mora-Blanco EL, Sansam CG, McKenna ES, Wilson B, Chen D, Klekota J, Tamayo P, Nguyen PT, Tolstorukov M, Park PJ, Cho YJ, Hsiao K, Buonamici S, Pomeroy SL, Mesirov JP, Ruffner H, Bouwmeester T, Luchansky SJ, Murtie J, Kelleher JF, Warmuth M, Sellers WR, Roberts CW, Dorsch M (2010) Loss of the tumor suppressor Snf5 leads to aberrant activation of the Hedgehog-Gli pathway. Nat Med 16(12):1429–1433. https://doi.org/10.1038/nm.2251
Darr J, Klochendler A, Isaac S, Eden A (2014) Loss of IGFBP7 expression and persistent AKT activation contribute to SMARCB1/Snf5-mediated tumorigenesis. Oncogene 33(23):3024–3032. https://doi.org/10.1038/onc.2013.261
Antonelli M, Raso A, Mascelli S, Gessi M, Nozza P, Coli A, Gardiman MP, Arcella A, Massimino M, Buttarelli FR, Giangaspero F (2017) SMARCB1/INI1 involvement in pediatric chordoma: a mutational and immunohistochemical analysis. Am J Surg Pathol 41(1):56–61. https://doi.org/10.1097/PAS.0000000000000741
Hasselblatt M, Thomas C, Hovestadt V, Schrimpf D, Johann P, Bens S, Oyen F, Peetz-Dienhart S, Crede Y, Wefers A, Vogel H, Riemenschneider MJ, Antonelli M, Giangaspero F, Bernardo MC, Giannini C, Ud Din N, Perry A, Keyvani K, van Landeghem F, Sumerauer D, Hauser P, Capper D, Korshunov A, Jones DT, Pfister SM, Schneppenheim R, Siebert R, Fruhwald MC, Kool M (2016) Poorly differentiated chordoma with SMARCB1/INI1 loss: a distinct molecular entity with dismal prognosis. Acta Neuropathol 132(1):149–151. https://doi.org/10.1007/s00401-016-1574-9
Mobley BC, McKenney JK, Bangs CD, Callahan K, Yeom KW, Schneppenheim R, Hayden MG, Cherry AM, Gokden M, Edwards MS, Fisher PG, Vogel H (2010) Loss of SMARCB1/INI1 expression in poorly differentiated chordomas. Acta Neuropathol 120(6):745–753. https://doi.org/10.1007/s00401-010-0767-x
Yadav R, Sharma MC, Malgulwar PB, Pathak P, Sigamani E, Suri V, Sarkar C, Kumar A, Singh M, Sharma BS, Garg A, Bakhshi S, Faruq M (2014) Prognostic value of MIB-1, p53, epidermal growth factor receptor, and INI1 in childhood chordomas. Neuro Oncol 16(3):372–381. https://doi.org/10.1093/neuonc/not228
Acknowledgements
The study was supported by the Research Special Fund For Public Welfare Industry of Health (201402008); Supported by the National High Technology Research and Development Program of China (863 Program); Supported by the National Natural Science Foundation of China (30971005).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Drs. Mingxuan Li and Yixuan Zhai have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Li, M., Zhai, Y., Bai, J. et al. SNF5 as a prognostic factor in skull base chordoma. J Neurooncol 137, 139–146 (2018). https://doi.org/10.1007/s11060-017-2706-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-017-2706-3