Abstract
The diagnosis of Talaromyces marneffei infection in HIV-negative patients remains challenging. There is an urgent need for rapid and convenient methods to diagnose this complicated disease. The aim of this study was to evaluate the diagnostic efficiency of metagenomic next-generation sequencing (mNGS) for talaromycosis in non-HIV-infected patients by comparing mNGS with traditional microbial culture. In total, 66 samples from 57 patients were analyzed via both mNGS and microbial culture. The ROC curve showed a sensitivity for mNGS of 97.22%, which was greater than that of microbial culture (61.11%). Samples from the respiratory tract, infectious skin lesions, and lymph nodes are recommended as routine samples for talaromycosis detection via mNGS. Furthermore, mNGS significantly reduced the diagnostic time compared to microbial culture. Overall, our study demonstrated that mNGS is a promising tool for rapid and accurate pathogenic detection in HIV-negative patients with talaromycosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Talaromyces marneffei is a dimorphic fungus that is predominantly endemic to Southeast Asia [1]. Infection with T. marneffei occurs mainly in immunocompromised hosts, especially HIV-infected patients, and is considered a major cause of HIV-related bloodstream infections and deaths in endemic regions [2]. In recent years, the incidence of talaromycosis in HIV-negative patients has gradually increased. Compared with that in HIV patients, talaromycosis in HIV-negative patients can manifest as variable and atypical systemic symptoms, which makes diagnosis relatively difficult [3]. In addition, these patients generally have a long duration of illness and a high mortality rate [4, 5].
The mainstay of diagnosis of T. marneffei infection is mycological culture of body tissues or fluids. However, diagnosis by culture is often delayed, as it can take up to 4 weeks to isolate and identify the pathogen from clinical specimens. To improve clinical outcomes, nonculture-based assays have been developed for the rapid detection of T. marneffei infection. For instance, the commercial (1,3)-β-d-glucan test (G test) and the galactomannan test (GM test) can be useful as screening tools and aids in diagnosis but cannot identify a specific genus [6, 7]. Although PCR-based assays and antigen detection assays targeting Mp1p may play a role in the rapid detection of T. marneffei [8,9,10,11], these methods require clinicians to have an initial suspicion of the pathogen, which limits its clinical application.
In recent years, metagenomic next-generation sequencing (mNGS) technologies have been increasingly applied in clinical laboratories for unbiased, culture-independent diagnosis. This technology does not require prior specific pathogen assumptions. The methods involve the extraction of nucleic acids from clinical samples and the application of a genomic approach to study the species and content of all microorganisms in the sample [12, 13]. In several published cases and clinical cohort studies, mNGS has been successfully applied to various samples, such as cerebrospinal fluid (CSF), respiratory specimens, blood and tissue samples. In addition, mNGS has provided great advantages in the diagnosis of complicated and severe infections [14]. The first case of talaromycosis diagnosed by NGS was reported in 2018 [15]. Subsequently, approximately 103 cases have been reported in the literature [16,17,18,19,20,21,22,23,24,25]. In 2022, a study reported that mNGS has good performance for diagnosing talaromycosis in HIV-infected patients [25]. However, the utility of mNGS for diagnosing talaromycosis in various samples from HIV-negative patients remains largely unexplored. The aim of this retrospective study was to investigate the diagnostic efficacy of the mNGS technique for T. marneffei by analyzing specimens from different affected sites of patients and evaluating its detection performance compared with that of conventional microbial culture.
Materials and Methods
Study Population
A retrospective study was conducted at the First Affiliated Hospital of Guangxi Medical University in Guangxi, China, from July 1, 2018, to November 30, 2021. Patients with suspected opportunistic infection who underwent both mNGS and microbiological culture were included. Patients who were HIV positive, had an incomplete medical history, had mNGS results not available for paired culture tests, or had noninfectious diseases were excluded (Fig. 1). Patients were classified into a T. marneffei infection (TM) group and a non-T. marneffei (non-TM) group according to the final diagnosis in the medical records. This study was approved by the ethics committee of the First Affiliated Hospital (number 2023-S812-01), and informed consent was obtained from the patients.
Categories
Given that T. marneffei is a described opportunistic pathogen, detection of T. marneffei in any clinical specimen should indicate infection. The results are in accordance with the 2020 EORTC/MSGERC revision and update of the consensus definitions of invasive fungal disease and expert consensus on clinical application [26] and refer to the diagnostic criteria for T. marneffei infection in this review [27]. Any of the following criteria can be defined as T. marneffei infection: T. marneffei identified by microbial culture of any clinical specimen from an affected site or a specific transverse septum in a dividing yeast cell found by histopathology or direct microscopy of clinical specimens from an affected site. Moreover, for multiple samples (≥ 2 types) from the same patient, T. marneffei was detected by mNGS. For other patients, T. marneffei was detected only once by mNGS at a low burden, whereas microbial culture was negative for any clinical specimen, considering that the concomitant clinical manifestations or imaging examinations were consistent with those of T. marneffei infections, we also defined these patients as having T. marneffei infections. All T. marneffei infections were included in the TM group. The non-TM group was defined as follows: the pathogen was confirmed to be the other microorganism by histopathology, direct microscopy, microbial culture or mNGS.
Sampling Protocols
For blood and bone marrow samples, 3–5 ml of the sample was collected using a tube containing the anticoagulant ethylenediaminetetraacetic acid (EDTA). At least 5 ml of bronchoalveolar lavage fluid (BALF), sputum, and CSF was collected in a sterile sealed tube for examination. Pus or secretions were obtained from abscesses, deep wounds or the base of ulcers in patients with skin involvement. For muscle, skin, lung tissue and lymph node tissue, biopsy was performed under sterile conditions, and the tissue was placed in a sterile container for examination. Bone tissue was obtained surgically by an orthopedic surgeon and placed in a sterile bottle. Blood samples were transported at room temperature, and nonblood samples were transported at low temperature with dry ice. The samples were delivered to the laboratory within 24 h of collection. The samples were then inactivated at 56 °C for 30 min for mNGS.
mNGS Procedure
All mNGS procedures followed the standardized guidelines of the Department of Laboratory Medicine of the Chinese Medical Association for NGS. The samples were placed in an automated NGS master workstation, which can carry out wet-laboratory procedures, including nucleic acid extraction, enzymatic fragmentation, end repair, A-tailing, ligation of sequencing adaptors and library purification. The finished libraries are then quantified and pooled for sequencing. The libraries were quantified by real-time PCR (KAPA), and shotgun sequencing was performed using the Illumina NextSeq high-throughput sequencing platform [28]. Approximately 20 million 75 bp single-end reads were generated from each library. The raw mNGS sequence data were analyzed using SURPI + as described in a previous report [29]; this procedure included filtering out human genome sequence data (GRCh38.p13) with Bowtie2 software and removing low-quality bases, linker sequences, and short sequences (< 35 bp). The remaining sequence data were aligned to microbial reference databases (the National Center for Biotechnology and Biotechnology Information [NCBI] GenBank and Pathogens Metagenomics Database [PMDB]) to determine microbial species and relative abundance.
Criteria for a Positive mNGS Result
Positive mNGS results were determined by referring to previously published studies, with slight modifications [30, 31]: (a) sequencing coverage of T. marneffei was among the top 10 pathogens detected; (b) the species-specific strictly mapped read number (SMRN) per million ratio (SMRN) of T. marneffei was ≥ 1; and (c) “other pathogens” was defined as in a previous study [25, 32]. mNGS was considered positive when one or both conditions were met.
Microbiological Culture
Collected samples (including peripheral blood, lymph nodes, bone tissue, BALF, lung tissue, sputum, skin tissue, pus, and secretions) were inoculated onto Sabouraud dextrose agar (SDA) and incubated at 25 °C and 37 °C for 30 days. At 25 °C, a light gray‒brown, membranous or yellowish fluffy mycelial phase was observed, and the back of the colony produced burgundy pigment (Fig. 2A). Microscopically, the mold formed a typical “broomstick” branch with hyaline septate hyphae and fruiting bodies composed of branched metulae and phialides, producing spherical conidia in chains (Fig. 2B). Light gray‒brown or cheese-colored moist yeast that formed at 37 °C and contained round, oval, and elongated yeast-like cells were identified as T. marneffei under the microscope (Fig. 2C).
Serum Galactomannan Test
A Platelia Aspergillus enzyme immunoassay (Bio-Rad, Marnesla-Coquette, France) was used for the GM test according to the manufacturer’s instructions. The content of serum galactomannan was detected by assessing the samples at 450 nm. The results were considered positive when the result was ≥ 0.5 OD.
Statistical Analysis
All statistical analyses and figures were generated using SPSS statistical package 26.0 software, GraphPad Prism 5 software and MedCalc 19.0 software. Numerical variables are expressed as medians and interquartile ranges and were compared by the Mann‒Whitney U test. Nominal variables are described by counts and percentages and were compared by the chi-square test. P values < 0.05 were considered to indicate statistical significance.
Results
Samples and Patient Characteristics
Fifty-seven HIV-negative patients, including 21 females and 36 males (the ratio of females to males was 1:1.71), were enrolled in the study. The mean age was 52.0 years (1.5–83 years). Twenty-eight patients were confirmed to have T. marneffei infection (the TM group). Twenty-nine patients were divided into the non-TM group and were diagnosed with tuberculosis (TB), nontuberculous mycobacterial (NTM) infections, or other invasive fungal infections (including histoplasmosis, cryptococcosis and invasive candidiasis). There were no significant differences in the demographic characteristics of the patients between the TM group and the non-TM group. The clinical and laboratory characteristics of the patients are shown in Table 1. The most common primary clinical manifestations in the TM group were respiratory symptoms (24/28, 85.7%), fever (22/28, 78.5%), lymphadenopathy (20/28, 71.4%), and weight loss (18/28, 64.3%). In the TM group, the median time to diagnosis of talaromycosis was 6.0 (1.8–10.0) months. A total of 66 specimens were collected from the 57 patients for both mNGS and microbiological culture. The predominant sample type was BALF (27/66, 40.9%), followed by skin tissue (13/66, 19.6%), lymph nodes (8/66, 12.1%), blood (5/66, 7.6%), pus (5/66, 7.6%), and swabs from skin lesions (2/66, 3.0%). CSF, sputum, bone, muscle, lung tissue, and BALF + lung tissue were collected from only 1 sample (Fig. 3). Talaromycosis was confirmed in 36 (54.6%) samples, whereas other pathogens were detected in 30 of the 66 samples (30/66, 45.4%).
Comparison of the Diagnostic Performance of mNGS and Traditional Methods
Comparison of Diagnostic Efficiency for Differentiating T. marneffei Patients from Non-TM Patients
A comparison of the results obtained by mNGS and traditional methods for 36 samples from patients with talaromycosis is shown in Fig. 4. The sensitivity and specificity of mNGS for diagnosing T. marneffei infection were 97.22% (95% CI: 85.5–99.9%) and 100.00% (95% CI: 88.4–100.0%), respectively, with positive predictive values (PPVs) and negative predictive values (NPVs) of 100.00% and 96.77%, respectively. The sensitivity and specificity of microbial culture for diagnosing talaromycosis were 61.11% (95% CI: 43.5–76.9%) and 100.00% (95% CI: 88.4–100.0%), respectively, with a PPV and NPV of 100.00% and 68.18%, respectively. In general, the false-negative rate (FNR) was 2.78% for the mNGS test and 38.89% for microbial culture. The results indicated that mNGS was more sensitive than conventional microbial culture for detecting T. marneffei (97.22% vs. 61.11%, P < 0.001). Both methods had high specificity for diagnosing T. marneffei infection (100.00% vs. 100.00%, P > 0.05).
Comparison of the diagnostic performance of mNGS and traditional methods for differentiating TM from non-TM lesions by the chi-square test. The results indicated that mNGS was more sensitive than conventional microbial culture for detecting T. marneffei (97.22% vs. 61.11%, P < 0.001). Both methods had high specificity for diagnosing T. marneffei infection (100.00% vs. 100.00%, P > 0.05). Contingency tables formatted in a 2 × 2 manner showing the diagnostic performance of mNGS and microbiological methods for differentiating TM from non-TM in 57 patients. Abbreviations: TM, Talaromyces marneffei; non-TM, non-Talaromyces marneffei; pos, positive; neg, negative; FPR, false positive rate; FNR, false negative rate; NPV, negative predictive value; PPV, positive predictive value
Concordance Between mNGS and Culture for Pathogen Detection
For the 36 (54.6%) samples in the TM group, the ROC curves in Fig. 5 show areas under the ROC curve (AUCs) for mNGS and microbial culture was 0.986 (95% CI: 92.1%–100.0%) and 0.806 (95% CI: 69.0%–89.3%), respectively, with P values < 0.01(MedCalc 19.0 software). Talaromyces marneffei was detected by both mNGS and culture in 21 samples (21/36, 58.3%). Among them, samples from two different sites were obtained from one patient; one site was positive by both mNGS and culture, while the other site was positive only by mNGS. Among the other 14 patients who were mNGS positive only, they were diagnosed with ongoing T. marneffei infection based on persistent signs and symptoms of infection and failure to respond to antibacterial treatment. However, mNGS failed to detect T. marneffei in 1 patient (2.8%), while microbial culture was positive for the sample from the same specimen site nearly 1 month later, which was considered a false negative for mNGS (Fig. 4). We also collected the serum GM test results of 51 patients, and the sensitivity of the GM test was 42.31% (Fig. 4). The AUC of the GM test was 0.508 (95% CI: 36.2–65.4%). The AUCs of mNGS were significantly greater than those of GM (0.986 vs. 0.508, P < 0.01) (Fig. 5).
Comparison Analysis at the Sample-Type Level
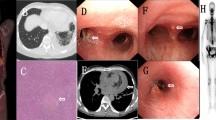
Clinical samples from different sources of suspected T. marneffei infection, including peripheral blood, lymph nodes, bone tissue, respiratory tract (lavage fluid, lung tissue, sputum), skin, swabs and muscle tissue, were tested. In the TM group, respiratory, cutaneous, lymphoid tissue and blood samples were most commonly used for detection (≥ 5 sample size). Among those sample types, for skin lesion tissue (n = 6), the sensitivities of mNGS and microbial culture were both 100.0% (6/6). There were papules with central necrosis, abscesses, and subcutaneous nodules. For the BALF samples (n = 13), the sensitivity of mNGS was 92.9% (13/14), and that of microbial culture was 64.3% (9/14). For the lymph node samples (n = 5), the sensitivity of mNGS was 100.0% (5/5), whereas that of microbial culture was 80.0% (4/5). For peripheral blood samples (n = 5), the sensitivity of mNGS was 100.0% (5/5), and that of microbial culture was 40.0% (2/5) (Table 2). We also detected low read lengths of T. marneffei in muscle, bone or lung tissue by mNGS, while microbial cultures were negative. However, due to the small sample size, the diagnostic value of mNGS could not be evaluated in those samples (Fig. 6).
Comparison of Turnaround Times
The time from the extraction of samples to the report was calculated for comparison. Overall, the turnaround times for cultures were significantly longer than those for mNGS, with cultures for lymph nodes, swabs, and bone and muscle samples requiring more time. Comparison revealed that the average turnover time was 2.8 days for mNGS and 9.8 days for microbial culture. There were significant differences between mNGS and culture in terms of average time consumption, and mNGS significantly reduced detection and diagnostic time (Fig. 7).
Discussion
Disseminated T. marneffei infection in HIV-negative patients usually presents with atypical symptoms and signs, including fever, weight loss, fatigue, hepatosplenomegaly, lymphadenopathy, skin lesions, and respiratory and gastrointestinal abnormalities [27]. These patients are often diagnosed inconclusively or misdiagnosed despite undergoing a series of tests and cultures in different hospitals. We have summarized 28 HIV-negative patients with talaromycosis in endemic areas over the past 3 years; 12 patients were misdiagnosed with nontuberculosis, and 5 were misdiagnosed with tuberculosis. The time between the onset of symptoms and the diagnosis of talaromycosis in these patients was up to 2 years.
Microbial culture is still the gold standard for the diagnosis of T. marneffei infection. However, the rate of positive culture results is low, especially in non-HIV-coinfected patients, as the fungal load of T. marneffei is often extremely low. A reliable diagnostic method for these patients is urgently needed to promote early diagnosis. PCR-based assays such as nested PCR and TaqMan real-time PCR have been developed to detect T. marneffei in whole blood, plasma or paraffin-embedded tissue samples. The studies revealed a high diagnostic specificity of the assay (100%), while the diagnostic sensitivity ranged from 67 to 91%. These assays provide useful tools for the rapid diagnosis of T. marneffei infection [8, 9, 33, 34]. However, these methods require physicians to suspect the pathogen before examination, which might limit their clinical application. In contrast to the above methods, mNGS provides an unbiased culture-independent technique for rapid etiological detection and has great advantages in the diagnosis of complicated and severe infections. Our study showed that mNGS has greater sensitivity (97.22%) than microbial culture (61.11%), which is consistent with the findings of other studies detecting T. marneffei in HIV-positive patients and other complex pathogen infections compared to microbial culture and pathology [25, 35,36,37].
Talaromycosis is a systemic infectious disease that can lead to multiple-organ damage. Choosing a suitable specimen for examination is crucial for improving the percentage of positive specimens and rapid diagnosis. Thus, we analyzed the percentage of positive specimens for diagnosis of talaromycosis. In our study, the most common samples were obtained from the respiratory tract (41.7%). This is likely because patients are initially infected through the airborne route by inhaling fungal spores, resulting in respiratory signs and symptoms [27, 38]. Our results showed that the percentage of positive BALF samples was significantly greater for mNGS than for culture (92.9% vs. 64.3%). Similar results were observed for the lymph node samples (mNGS 100% vs. culture 80%) and peripheral blood samples (mNGS 100.0% vs. culture 40.0%). The cutaneous system is also a common area affected by disseminated talaromycosis. Skin lesions can be the single symptom or the first symptom of disseminated talaromycosis [39, 40]. In our study, 48.3% of patients with a history of talaromycosis (14/28) had skin lesions, and 6 of those patients had papules with central necrosis, abscesses, or subcutaneous nodules. We conducted mNGS and microbial culture on these samples, and T. marneffei was detected in all the samples by mNGS and culture. These kinds of skin lesions are considered infective dermatoses and can be used for pathogen detection. However, reactive skin manifestations such as Sweet’s syndrome, erythema nodosum, and exanthematous pustulosis also occur in non-HIV-coinfected patients with talaromycosis, whereas T. marneffei cannot be detected in those reactive skin lesions [41]. Thus, it is not recommended to perform microbial detection on those lesions [42]. Therefore, we did not perform microbial detection on those reactive skin lesions in this study.
Disseminated talaromycosis in HIV-negative patients may lead to osteolytic damage manifesting as ostealgia, joint pain and local soft tissue swelling [43,44,45]. However, diagnosing talaromycosis using bone and joint samples in conventional microbial culture is challenging due to the rapid nature of the organism, biofilm formation on implant surfaces, prior antibiotic administration, or limited sample availability, resulting in a high rate of negative microbial culture results [46, 47]. In our study, approximately 30.56% of patients had ostalgia, and 66.67% of patients had bone destruction on imaging. With respect to a very low sequence read, T. marneffei could be captured in a bone tissue sample by mNGS, but culture at the corresponding site was negative. These results indicate that mNGS can be used as a supplemental method for diagnosing T. marneffei infections involving bone and joints [48].
The mNGS technique also has limitations. For instance, for a culture-positive T. marneffei BALF sample, the pathogen was not detected by mNGS. The reasons may be that T. marneffei is an intracellular fungus that releases little extracellular nucleic acid due to its intracellular growth characteristics and difficulty breaking through the cell wall during nucleic acid extraction. Therefore, the interpretation of mNGS results should receive sufficient attention. Combining mNGS with clinical manifestations, blood tests, and imaging examinations of patients is more scientific and appropriate for diagnosing T. marneffei infection if a low fungal load in the sample is reported.
Overall, our study has several limitations that should be considered. This was a retrospective analysis of data from a single institution with relatively few patients. Second, no further analysis was conducted on individuals with different immune statuses, especially in terms of infection characteristics and diagnostic efficiency of mNGS, between the HIV-infected group and the HIV-negative group. In addition, it is necessary to include more samples from special affected sites, such as CSF, bone, muscle and intestinal specimens, to accurately evaluate the diagnostic efficacy of mNGS on various types of samples. Nevertheless, this study may provide a valuable reference for the diagnosis of T. marneffei infection. In future research, prospective multicenter studies with larger sample sizes may provide additional information to evaluate the diagnostic efficacy of mNGS for patients with talaromycosis.
Conclusion
Our study demonstrates that mNGS is a potential tool for the rapid diagnosis of talaromycosis in HIV-negative patients. BMLF, lymph node and infectious skin lesion samples may be good options for mNGS and microbial culture to help in the early identification of T. marneffei in HIV-negative individuals.
References
Ashraf N, Kubat RC, Poplin V, Adenis AA, Denning DW, Wright L, McCotter O, Schwartz IS, Jackson BR, Chiller T, Bahr NC. Re-drawing the maps for endemic mycoses. Mycopathologia. 2020;185:843–65. https://doi.org/10.1007/s11046-020-00431-2.
Le T, Wolbers M, Chi NH, Quang VM, Chinh NT, Lan NP, Lam PS, Kozal MJ, Shikuma CM, Day JN, Farrar J. Epidemiology, seasonality, and predictors of outcome of AIDS-associated Penicillium marneffei infection in Ho Chi Minh City, Viet Nam. Clin Infect Dis. 2011;52:945–52. https://doi.org/10.1093/cid/cir028.
Kawila R, Chaiwarith R, Supparatpinyo K. Clinical and laboratory characteristics of penicilliosis marneffei among patients with and without HIV infection in Northern Thailand: a retrospective study. Bmc Infect Dis. 2013;13:464. https://doi.org/10.1186/1471-2334-13-464.
Zhang JQ, Yang ML, Zhong XN, He ZY, Liu GN, Deng JM, Li MH. A comparative analysis of the clinical and laboratory characteristics in disseminated penicilliosis marneffei in patients with and without human immunodeficiency virus infection. Zhonghua Jie He He Hu Xi Za Zhi. 2008;31:740–6.
Li HR, Cai SX, Chen YS, Yu ME, Xu NL, Xie BS, Lin M, Hu XL. Comparison of Talaromyces marneffei infection in human immunodeficiency virus-positive and human immunodeficiency virus-negative patients from Fujian, China. Chin Med J (Engl). 2016;129:1059–65. https://doi.org/10.4103/0366-6999.180520.
Poplin V, Smith C, Milsap D, Zabel L, Bahr NC. Diagnosis of pulmonary infections due to endemic fungi. Diagnostics (Basel). 2021;11(5):856. https://doi.org/10.3390/diagnostics11050856.
Huang YT, Hung CC, Liao CH, Sun HY, Chang SC, Chen YC. Detection of circulating galactomannan in serum samples for diagnosis of Penicillium marneffei infection and cryptococcosis among patients infected with human immunodeficiency virus. J Clin Microbiol. 2007;45:2858–62. https://doi.org/10.1128/JCM.00050-07.
Hien H, Thanh TT, Thu N, Nguyen A, Thanh NT, Lan N, Simmons C, Shikuma C, Chau N, Thwaites G, Le T. Development and evaluation of a real-time polymerase chain reaction assay for the rapid detection of Talaromyces marneffei MP1 gene in human plasma. Mycoses. 2016;59:773–80. https://doi.org/10.1111/myc.12530.
Li X, Zheng Y, Wu F, Mo D, Liang G, Yan R, Khader JA, Wu N, Cao C. Evaluation of quantitative real-time PCR and Platelia galactomannan assays for the diagnosis of disseminated Talaromyces marneffei infection. Med Mycol. 2020;58:181–6. https://doi.org/10.1093/mmy/myz052.
Thompson GR, Le T, Chindamporn A, Kauffman CA, Alastruey-Izquierdo A, Ampel NM, Andes DR, Armstrong-James D, Ayanlowo O, Baddley JW, Barker BM, Lopes BL, Buitrago MJ, Chamani-Tabriz L, Chan J, Chayakulkeeree M, Cornely OA, Cunwei C, Gangneux JP, Govender NP, Hagen F, Hedayati MT, Hohl TM, Jouvion G, Kenyon C, Kibbler CC, Klimko N, Kong D, Krause R, Lee LL, Meintjes G, Miceli MH, Rath PM, Spec A, Queiroz-Telles F, Variava E, Verweij PE, Schwartz IS, Pasqualotto AC. Global guideline for the diagnosis and management of the endemic mycoses: an initiative of the European Confederation of Medical Mycology in cooperation with the International Society for Human and Animal Mycology. Lancet Infect Dis. 2021;21:e364–74. https://doi.org/10.1016/S1473-3099(21)00191-2.
Pruksaphon K, Intaramat A, Ratanabanangkoon K, Nosanchuk JD, Vanittanakom N, Youngchim S. Diagnostic laboratory immunology for talaromycosis (penicilliosis): review from the bench-top techniques to the point-of-care testing. Diagn Microbiol Infect Dis. 2020;96: 114959. https://doi.org/10.1016/j.diagmicrobio.2019.114959.
Lecuit M, Eloit M. The diagnosis of infectious diseases by whole genome next generation sequencing: a new era is opening. Front Cell Infect Microbiol. 2014;4:25. https://doi.org/10.3389/fcimb.2014.00025.
Lefterova MI, Suarez CJ, Banaei N, Pinsky BA. Next-generation sequencing for infectious disease diagnosis and management: a report of the association for molecular pathology. J Mol Diagn. 2015;17:623–34. https://doi.org/10.1016/j.jmoldx.2015.07.004.
Han D, Li Z, Li R, Tan P, Zhang R, Li J. mNGS in clinical microbiology laboratories: on the road to maturity. Crit Rev Microbiol. 2019;45:668–85. https://doi.org/10.1080/1040841X.2019.1681933.
Zhu YM, Ai JW, Xu B, Cui P, Cheng Q, Wu H, Qian YY, Zhang HC, Zhou X, Xing L, Wu R, Li Y, Zhang WH. Rapid and precise diagnosis of disseminated T. marneffei infection assisted by high-throughput sequencing of multifarious specimens in a HIV-negative patient: a case report. Bmc Infect Dis. 2018;18:379. https://doi.org/10.1186/s12879-018-3276-5.
Tsang CC, Teng J, Lau S, Woo P. Rapid genomic diagnosis of fungal infections in the age of next-generation sequencing. J Fungi (Basel). 2021;7(8):636. https://doi.org/10.3390/jof7080636.
Chen Q, Qiu Y, Zeng W, Wei X, Zhang J. Metagenomic next-generation sequencing for the early diagnosis of talaromycosis in HIV-uninfected patients: five cases report. Bmc Infect Dis. 2021;21:865. https://doi.org/10.1186/s12879-021-06551-4.
Chen X, Jia L, Wu Y, Chang J, Zhang T, Ma Y, Zhang Y. A mass in the upper abdomen derived from Talaromyces marneffei infected lymphadenopathy: a case report. Bmc Infect Dis. 2021;21:750. https://doi.org/10.1186/s12879-021-06489-7.
Wang DM, Ma HL, Tan MQ, Wu YM, Wang SN. Next-generation sequencing confirmed the diagnosis of isolated central nervous system infection caused by Talaromyces marneffei in an immunocompetent patient. Chin Med J (Engl). 2020;133:374–6. https://doi.org/10.1097/CM9.0000000000000593.
Gao Y, Qu M, Song C, Yin L, Zhang M. Cerebral vasculitis caused by Talaromyces marneffei and Aspergillus niger in a HIV-positive patient: a case report and literature review. J Neurovirol. 2022;28:274–80. https://doi.org/10.1007/s13365-021-01032-5.
Ba H, Peng H, Cheng L, Lin Y, Li X, He X, Li S, Wang H, Qin Y. Case report: Talaromyces marneffei infection in a Chinese child with a complex heterozygous CARD9 mutation. Front Immunol. 2021;12: 685546. https://doi.org/10.3389/fimmu.2021.685546.
Yang A, Hu Y, Chen P, Zheng G, Hu X, Zhang J, Wang J, Wang C, Huang Z, Zhang Y, Guo Y. Diagnosis by metagenomic next-generation sequencing of a Talaromyces marneffei bloodstream infection in an HIV-negative child: a case report. Front Pediatr. 2022;10: 903617. https://doi.org/10.3389/fped.2022.903617.
Li D, Liang H, Zhu Y, Chang Q, Pan P, Zhang Y. Clinical characteristics, laboratory findings, and prognosis in patients with Talaromyces marneffei infection across various immune statuses. Front Med (Lausanne). 2022;9: 841674. https://doi.org/10.3389/fmed.2022.841674.
Liu L, Sun B, Ying W, Liu D, Wang Y, Sun J, Wang W, Yang M, Hui X, Zhou Q, Hou J, Wang X. Rapid diagnosis of Talaromyces marneffei infection by metagenomic next-generation sequencing technology in a Chinese cohort of inborn errors of immunity. Front Cell Infect Microbiol. 2022;12: 987692. https://doi.org/10.3389/fcimb.2022.987692.
Mao Y, Shen H, Yang C, Jia Q, Li J, Chen Y, Hu J, Huang W. Clinical performance of metagenomic next-generation sequencing for the rapid diagnosis of talaromycosis in HIV-infected patients. Front Cell Infect Microbiol. 2022;12: 962441. https://doi.org/10.3389/fcimb.2022.962441.
Donnelly JP, Chen SC, Kauffman CA, Steinbach WJ, Baddley JW, Verweij PE, Clancy CJ, Wingard JR, Lockhart SR, Groll AH, Sorrell TC, Bassetti M, Akan H, Alexander BD, Andes D, Azoulay E, Bialek R, Bradsher RW, Bretagne S, Calandra T, Caliendo AM, Castagnola E, Cruciani M, Cuenca-Estrella M, Decker CF, Desai SR, Fisher B, Harrison T, Heussel CP, Jensen HE, Kibbler CC, Kontoyiannis DP, Kullberg BJ, Lagrou K, Lamoth F, Lehrnbecher T, Loeffler J, Lortholary O, Maertens J, Marchetti O, Marr KA, Masur H, Meis JF, Morrisey CO, Nucci M, Ostrosky-Zeichner L, Pagano L, Patterson TF, Perfect JR, Racil Z, Roilides E, Ruhnke M, Prokop CS, Shoham S, Slavin MA, Stevens DA, Thompson GR, Vazquez JA, Viscoli C, Walsh TJ, Warris A, Wheat LJ, White PL, Zaoutis TE, Pappas PG. Revision and update of the consensus definitions of invasive fungal disease from the european organization for research and treatment of cancer and the mycoses study group education and research consortium. Clin Infect Dis. 2020;71:1367–76. https://doi.org/10.1093/cid/ciz1008.
Cao C, Xi L, Chaturvedi V. Talaromycosis (Penicilliosis) Due to Talaromyces (Penicillium) marneffei: insights into the clinical trends of a major fungal disease 60 years after the discovery of the pathogen. Mycopathologia. 2019;184:709–20. https://doi.org/10.1007/s11046-019-00410-2.
Luan Y, Hu H, Liu C, Chen B, Liu X, Xu Y, Luo X, Chen J, Ye B, Huang F, Wang J, Duan C. A proof-of-concept study of an automated solution for clinical metagenomic next-generation sequencing. J Appl Microbiol. 2021;131:1007–16. https://doi.org/10.1111/jam.15003.
Miller S, Naccache SN, Samayoa E, Messacar K, Arevalo S, Federman S, Stryke D, Pham E, Fung B, Bolosky WJ, Ingebrigtsen D, Lorizio W, Paff SM, Leake JA, Pesano R, DeBiasi R, Dominguez S, Chiu CY. Laboratory validation of a clinical metagenomic sequencing assay for pathogen detection in cerebrospinal fluid. Genome Res. 2019;29:831–42. https://doi.org/10.1101/gr.238170.118.
Geng S, Mei Q, Zhu C, Fang X, Yang T, Zhang L, Fan X, Pan A. Metagenomic next-generation sequencing technology for detection of pathogens in blood of critically ill patients. Int J Infect Dis. 2021;103:81–7. https://doi.org/10.1016/j.ijid.2020.11.166.
Long Y, Zhang Y, Gong Y, Sun R, Su L, Lin X, Shen A, Zhou J, Caiji Z, Wang X, Li D, Wu H, Tan H. Diagnosis of sepsis with cell-free DNA by next-generation sequencing technology in ICU patients. Arch Med Res. 2016;47:365–71. https://doi.org/10.1016/j.arcmed.2016.08.004.
Peng JM, Du B, Qin HY, Wang Q, Shi Y. Metagenomic next-generation sequencing for the diagnosis of suspected pneumonia in immunocompromised patients. J Infect. 2021;82:22–7. https://doi.org/10.1016/j.jinf.2021.01.029.
Lu S, Li X, Calderone R, Zhang J, Ma J, Cai W, Xi L. Whole blood nested PCR and real-time PCR amplification of Talaromyces marneffei specific DNA for diagnosis. Med Mycol. 2016;54:162–8. https://doi.org/10.1093/mmy/myv068.
Zeng H, Li X, Chen X, Zhang J, Sun J, Xie Z, Xi L. Identification of Penicillium marneffei in paraffin-embedded tissue using nested PCR. Mycopathologia. 2009;168:31–5. https://doi.org/10.1007/s11046-009-9195-7.
Miao Q, Ma Y, Wang Q, Pan J, Zhang Y, Jin W, Yao Y, Su Y, Huang Y, Wang M, Li B, Li H, Zhou C, Li C, Ye M, Xu X, Li Y, Hu B. Microbiological diagnostic performance of metagenomic next-generation sequencing when applied to clinical practice. Clin Infect Dis. 2018;67:S231–40. https://doi.org/10.1093/cid/ciy693.
Li H, Gao H, Meng H, Wang Q, Li S, Chen H, Li Y, Wang H. Detection of pulmonary infectious pathogens from lung biopsy tissues by metagenomic next-generation sequencing. Front Cell Infect Microbiol. 2018;8:205. https://doi.org/10.3389/fcimb.2018.00205.
Garnica M, Pierrotti LC, Oliveira PV, Mazzi M, Chebabo A. Metagenomic next-generation sequencing (mNGS) for diagnostically challenging infectious diseases in patients with acute leukemia. Braz J Infect Dis. 2021;25: 101548. https://doi.org/10.1016/j.bjid.2021.101548.
Narayanasamy S, Dougherty J, van Doorn HR, Le T. Pulmonary Talaromycosis: a window into the immunopathogenesis of an endemic mycosis. Mycopathologia. 2021;186:707–15. https://doi.org/10.1007/s11046-021-00570-0.
Hu Y, Zhang J, Li X, Yang Y, Zhang Y, Ma J, Xi L. Penicillium marneffei infection: an emerging disease in mainland China. Mycopathologia. 2013;175:57–67. https://doi.org/10.1007/s11046-012-9577-0.
Chan JF, Lau SK, Yuen KY, Woo PC. Talaromyces (Penicillium) marneffei infection in non-HIV-infected patients. Emerg Microbes Infect. 2016;5: e19. https://doi.org/10.1038/emi.2016.18.
Chan JF, Trendell-Smith NJ, Chan JC, Hung IF, Tang BS, Cheng VC, Yeung CK, Yuen KY. Reactive and infective dermatoses associated with adult-onset immunodeficiency due to anti-interferon-gamma autoantibody: Sweet’s syndrome and beyond. Dermatology. 2013;226:157–66. https://doi.org/10.1159/000347112.
Pattanaprichakul P, Leeyaphan C, Angkasekwinai N, Bunyaratavej S, Senawong S, Sereeaphinan C, Munprom K. Prevalence and clinical manifestations of cutaneous findings in patients with adult-onset immunodeficiency due to anti-interferon gamma autoantibodies: an eight-year retrospective study. Int J Dermatol. 2023;62:1506–10. https://doi.org/10.1111/ijd.16870.
Rammaert B, Gamaletsou MN, Zeller V, Elie C, Prinapori R, Taj-Aldeen SJ, Roilides E, Kontoyiannis DP, Brause B, Sipsas NV, Walsh TJ, Lortholary O. Dimorphic fungal osteoarticular infections. Eur J Clin Microbiol Infect Dis. 2014;33:2131–40. https://doi.org/10.1007/s10096-014-2149-0.
Qiu Y, Zhang J, Liu G, Zhong X, Deng J, He Z, Jing B. Retrospective analysis of 14 cases of disseminated Penicillium marneffei infection with osteolytic lesions. Bmc Infect Dis. 2015;15:47. https://doi.org/10.1186/s12879-015-0782-6.
Pun TS, Fang D. A case of Penicillium marneffei osteomyelitis involving the axial skeleton. Hong Kong Med J. 2000;6:231–3.
Thoendel M, Jeraldo P, Greenwood-Quaintance KE, Chia N, Abdel MP, Steckelberg JM, Osmon DR, Patel R. A novel prosthetic joint infection pathogen, Mycoplasma salivarium, identified by metagenomic shotgun sequencing. Clin Infect Dis. 2017;65:332–5. https://doi.org/10.1093/cid/cix296.
Fang X, Cai Y, Shi T, Huang Z, Zhang C, Li W, Zhang C, Yang B, Zhang W, Guan Z. Detecting the presence of bacteria in low-volume preoperative aspirated synovial fluid by metagenomic next-generation sequencing. Int J Infect Dis. 2020;99:108–16. https://doi.org/10.1016/j.ijid.2020.07.039.
Li N, Cai Q, Miao Q, Song Z, Fang Y, Hu B. High-throughput metagenomics for identification of pathogens in the clinical settings. Small Methods. 2021;5:2000792. https://doi.org/10.1002/smtd.202000792.
Funding
This study was supported by the Natural Science Foundation of Guangxi Province of China (2020GXNSFGA238001), the National Key Research and Development Program of China (2022YFC2504800), and the Foundation of Nanning Qingxiu District (2020016).
Author information
Authors and Affiliations
Contributions
J.L. and L.T.W. wrote the first draft of the manuscript. W.G. and C.C.W. designed the study. L.X.Y., P.K.S., Z.D.Y., W.L.L., L.C.Y. and J.Z.W. participated in the data collection. L.T.W. and H.Y. analyzed the data. All the authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Additional information
Handling Editor: Weida Liu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jiang, L., Liang, Tw., Al-Odaini, N. et al. Metagenomic Next-Generation Sequencing as an Effective Diagnostic Tool for Talaromycosis in HIV-Negative Patients. Mycopathologia 189, 63 (2024). https://doi.org/10.1007/s11046-024-00866-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11046-024-00866-x