Abstract
Candida parapsilosis complex (CPC) is the third Candida species isolated in blood cultures of patients from our Hospital, following C. albicans and C. tropicalis. From 2006 to 2010, the median annual distribution of CPC was 8 cases/year. Records of 36 patients were reviewed. CPC were 31 (86.1 %) C. parapsilosis; 4 (11.1 %) C. orthopsilosis; and 1 (2.8 %) C. metapsilosis. Clinical characteristics were central venous catheter, 34 (94.4 %); parental nutrition, 25 (70 %); surgery, 27 (57.9 %); prior bacteremia, 20 (51.3 %); malignancy, 18 (50 %). General mortality was 47.2 %. Death was higher in immunosuppressed patients (17 vs. 11; p = 0.003). Three out four (75 %) patients with C. orthopsilosis and 14 out 31 (45.2 %) with C. parapsilosis died (p = 0.558). Thirty-nine individual isolates were tested for susceptibility to seven antifungal drugs, with MICs values showing susceptibility to all of them. Two isolates, one C. orthopsilosis and one C. parapsilosis, had fluconazole MIC = 4 μg/mL. Differentiation among CPC has implication in caring for patients with invasive candidiasis since there are differences in virulence, pathogenicity and drug susceptibility. A method targeting the topoisomerase II gene based on loop-mediated isothermal amplification (LAMP) was developed. LAMP emerges as a promising tool for the identification of fungal species due to the high sensitivity and specificity. LAMP can be performed at the point-of-care, being no necessary the use of expensive equipment. In our study, the method was successful comparing to the DNA sequencing and proved to be a reliable and fast assay to distinguish the three species of CPC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Invasive candidiasis is the most frequent, as well as life-threatening fungal infection in severely ill patients, either in medical, surgical or burn intensive care units (ICU) [1]. The prompt introduction of specific and highly effective antifungal therapy is of a major importance. However, significant epidemiological changes were witnessed during the past decades, either regarding the increasing prevalence of non-albicans Candida species, as well as the emergence of resistance to fluconazole [2, 3].
Despite the increase in the incidence of C. glabrata reported recently [2, 3], C. parapsilosis remains as the most frequent non-albicans Candida species in South American and Brazilian hospitals [3–6].
Several publications reported C. parapsilosis as a frequent causative agent of blood stream infection in the last decade [7–10] and as a major pathogen causing candidemia in neonates [11, 12], children [13] and onco-hematologic patients [14]. The ability of C. parapsilosis to adhere onto vascular catheters, prosthetics devices and the hands of health care workers [15, 16] represents a potential risk of the development of candidemia and a challenge for their control.
Since 2005, C. parapsilosis sensu lato has been recognized as a complex of three closely related species: C. parapsilosis sensu stricto, C. orthopsilosis and C. metapsilosis [17]. The three species of the Candida parapsilosis complex (CPC) are closely related even in their biochemical characteristics, which makes extremely difficult, if not impossible, differentiate the CPC members by phenotypic methods. Nevertheless, differentiation among CPC members has a potential implication in caring for patients with invasive candidiasis, since C. parapsilosis sensu stricto appears to be more resistant than C. orthopsilosis and C. metapsilosis for a variety of antifungal agents [7]. Based on in vitro experiments, there are also evidences that the three components of CPC have different pathogenicity, being C. parapsilosis sensu stricto more pathogenic than C. orthopsilosis which is, in turn, more pathogenic than C. metapsilosis [18]. Therefore, in order to determine and identify each species correctly, sequencing of ITS regions or other molecular techniques has been necessary. Molecular techniques have been applied for the identification of fungi, including DNA microarray [19, 20], real-time polymerase chain reaction (real-time PCR) [21] and DNA sequencing [22]. DNA microarray has been demonstrated as highly specific tool to simultaneously detect multiple pathogens on a 4-h runtime assay, which can be performed at the point-of care [19, 23]. Real-time PCR can quantify the DNA amount, showing a higher level of accuracy when compared to the standard PCR, though requires expensive equipment and skilled personnel to be performed [21, 24, 25]. Despite the high specificity and discrimination power that made DNA sequencing be chosen as the gold standard for fungi identification, this technique is a 12-h time-consuming assay, it is not widely available at the point-of-care and also requires skilled personnel and expensive supplies to be performed [22]. Since such works are tedious and time-consuming, more simple and rapid procedures are desired [26]. Recently, a DNA amplification technique, loop-mediated isothermal amplification (LAMP), was developed and has been applied to the detection of a variety of microorganisms. LAMP method consist of a PCR that uses a set of six primers, being two outer-inserted (forward and backward) in the 3′ position (F3; B3), two inner-inserted (forward and backward) in the ITS region (FIP; BIP), and two loop-inserted (loop-F; loop-B). Amplification starts with the pairing of the six primers with their counterpart, which allows a more specific identification of pathogen when comparing with conventional real-time PCR. Although LAMP can also quantify DNA amount in real time, the amplification can be visually checked by turbidity, making not necessary a dye labeling [27]. The simplicity and rapidness of LAMP have been confirmed [28–30]. In this paper, the usefulness of newly developed LAMP identification primers for the CPC was reported, and the performance of this method was compared with others [26, 31]. We also describe the clinical and epidemiological features of blood stream infections caused by CPC and the antifungal susceptibility testing.
Materials and Methods
This retrospective study was performed comprising 39 strains of CPC isolated from blood cultures of individual patients hospitalized in a Brazilian tertiary care university hospital. From January 2006 to December 2010, the epidemiological and clinical data from charts of 36 adult patients were reviewed. Blood cultures and yeast identification were performed by automated microbiological system BacT/ALERT 3D FA and PF bottles and Vitek 2YST card (bioMérieux Inc. Durham, NC, USA). The isolates were kept in sterile distilled water and, for the purpose of this study, the fungal isolates were cultured in CHROMagar (CHROMagar, Paris, France) and Sabouraud dextrose agar (SDA, Difco, Detroit, MI, USA) plate and kept on potato dextrose agar (PDA, Difco) slants before use.
Drug Susceptibility Tests
Drug susceptibility tests of 39 strains of CPC were performed by broth microdilution method according to Clinical Laboratory Standards Institute (CLSI) document M27-A2 [32] for flucytosine, itraconazole, fluconazole, voriconazole, micafungin and miconazole. Minimum inhibitory concentration (MIC) for amphotericin B was determined by E test [33]. C. parapsilosis ATCC 22019 was used as a reference strain. MICs at which 50 % (MIC50) and 90 % (MIC90) of strains were inhibited were reported. Interpretative criteria for fluconazole MICs were: ≤8 μg/mL: susceptible; 16–32 μg/mL: susceptible dose dependent and ≥64 μg/mL: resistant. MICs for micafungin were considered susceptible if ≤2 μg/mL and; however, an interpretative criterion for amphotericin B has not yet been established by CLSI, isolates with amphotericin B MIC ≤1 μg/mL were considered susceptible [34]. CLSI endpoints interpretation was compared to the recently suggested clinical endpoints for antifungal agents [35–38].
DNA Sequencing
DNA sequencing PCR were performed with universal fungus-specific primer pair ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) and ITS5 (5′-GGAAGTAAAAGTCGTAACAAGG-3′) (Sigma-Aldrich, Saint Louis, MO, USA) [39, 40]. The PCR products were purified with ExoSAP-IT (Affymetrix |USB, Cleveland, OH, USA) and then sequenced using Big Dye® terminator reagent kit (Applied Biosystem, Foster City, CA, USA) according to the manufacturer’s instructions in an ABI PRISM® 3100 Genetic analyzer (Applied Biosystems, Foster City, CA, USA). The sequence data were assembled by ATSQ software version 6.0.1 (Genetix Corporation, Tokyo, Japan) and then submitted to GenBank.
Molecular Identification by LAMP Method
For the molecular identification by LAMP method, following cultures were used as reference strains: Candida albicans ATCC 90028 (IFM 40213), Candida dubliniensis CBS 7987T (IFM 48313), Candida glabrata ATCC 2001T (IFM 46843), Candida guilliermondii ATCC 6260T (IFM 46823), Candida kefyr ATCC 4135T (IFM 5773), Candida krusei ATCC 62587T (IFM 46834), Candida parapsilosis ATCC22019T (IFM 46829) and Candida tropicalis ATCC 750T (IFM 5777). All fungal strains were grown on SDA slant at 27 or 37 °C for 4–7 days. Rapid preparation of DNA from fungi was performed by a modification of our previous report [41]. Small samples of fungal cells on SDA plate were suspended in a saline, and DNA was extracted with a DNA extraction kit (Dr. GenTLE, Takara Bio Inc., Shiga, Japan). Sets of LAMP primers were developed specifically for differentiating CPC, using sequence information of the topoisomerase II gene; known sequences of targeted genes were searched in the GenBank (http://www.ncbi.nlm.nih.gov/genbank/), Centraalbureau voor Schimmelcultures Fungal Biodiversity Centre (CBS; http://www.cbs.knaw.nl), and DNA Data Bank of Japan (DDBJ; http://www.ddbj.nig.ac.jp) databases. Then, six to ten sequences, each one having more than 600 base pairs (bp) and less than 800 bp, were aligned using the software Clustal-W (http://www.clustal.org). The final alignment sequence was used to design the specific primers for LAMP (PrimerExplorer V4, Eiken Chemical Co., Japan, http://primerexplorer.jp/e/). Primers were manufactured by Sigma-Aldrich Japan (http://www.sigmaaldrich.com/japan.html/). The LAMP reaction was performed with a Loopamp DNA amplification kit (Eiken Chemical Co., Tokyo, Japan) in reaction mixtures composed of 40 pmol each of primers FIP and BIP, 5 pmol each of primers F3 and B3, 20 pmol each primers LF and LB, 12.5 µL of 2× reaction mixture, 1 µL of Bst DNA polymerase, 2 µL DNA samples and distilled water up to a final volume of 25 µl. The mixtures were incubated at 63 °C for 60 min and then heated at 80 °C for 2 min to terminate the reaction. The turbidity was measured every 6 s using a real-time turbidity meter (LoopAmp EXIA; Eiken Chemical Co., Japan). Start of amplification of LAMP products at around 30 min suggested the positive reaction due to the presence of corresponding fungal DNA by specific LAMP primer sets [28, 41].
Results
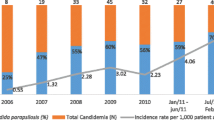
During the study period, 313 candidemia episodes were identified, and the CPC was the third Candida species isolated after C. albicans and C. tropicalis. Charts of 36 patients were reviewed. CPC were distributed as follows: C. parapsilosis sensu stricto: 31 (86.1 %), C. orthopsilosis: 4 (11.1 %) and C. metapsilosis: 1 (2.8 %). Annual distribution was similar during the period with a median of 8 (range 3–13) cases per year. Main characteristics of the patients are in Table 1. Fifty percent of the patients had cancer (onco-hematological: 7 or gastrointestinal: 10) as the main underlying disease. General mortality rate was 47.2 % (17 patients), and main risk factors were: ICU admission <48 h: 17 (47.2 %); surgery: 27 (57.9 %); parental nutrition: 25 (70 %); prior bacteremia: 20 (51.3 %); CVC: 34 (94.4 %); mean time from blood culture to antifungal treatment: 2.6 days. Antifungal treatment was administered to 28 (75 %) patients. The univariate analysis comparing patients’ outcomes (survival or death) showed a mean number of days from blood culture to outcome significantly higher in patients that survived (25 vs. 13 days; p = 0.04). Death was higher in patients with immunosuppressive conditions (17 vs. 11 patients; p = 0.003); three out of four patients with C. orthopsilosis died (75 %), while 14 of 31 patients with C. parapsilosis sensu stricto died (p = 0.558).
Drug susceptibilities of 39 isolates of CPC (C. parapsilosis: 33; C. orthopsilosis: 5; C. metapsilosis: 1) were compared with those of the reference C. parapsilosis ATCC 22019. MICs values are displayed in Table 2. Fluconazole MIC50/MIC90 were 0.5/1 μg/mL ranging from 0.5 to 4 μg/mL. One isolate of C. orthopsilosis and one isolate of C. parapsilosis showed MIC = 4 μg/mL for fluconazole. Most of 39 isolates showed lower MIC values against itraconazole (MIC50/MIC90: 0.06/0.125 μg/mL) and voriconazole (MIC50/MIC90: ≤ 0.015/0.06 μg/mL) when compared with those of reference C. parapsilosis ATCC 22019. MICs of amphotericin B (MIC50/MIC90: 0.5/1 μg/mL), flucytosine (MIC50/MIC90: 0.25/0.25 μg/mL) and itraconazole (MIC50/MIC90: 0.06/0.125 μg/mL) were very similar among the isolates. Micafungin MIC50/MIC90 was 0.5/0.5 μg/mL ranging from 0.125 to 1 μg/mL. These data suggest that our strains showed good susceptibility against all tested antifungal agents. No correlation was seen comparing mortality and the susceptibility to antifungal drugs. Fluconazole was administered to 23 patients and amphotericin B to 3 patients. In the group of patients that received fluconazole, 12 patients survived and 11 died.
The targets of the LAMP primers developed in this investigation were conserved region sequences inside the topoisomerase II. Our first screening works, which lead to specific LAMP primers, were repeated, and many possibly useful specific LAMP candidate primer sets for each species of the psilosis complex were prepared. Among their LAMP primer sets, the primers that showed a rising of the turbidity curves at around 30 min were selected. The primers’ sequence information, which showed a sharp DNA amplification in each species of the psilosis complex, was listed in Table 3. Usefulness of each of the three primer sets, thus prepared for the identification of C. parapsilosis sensu stricto, C. orthopsilosis and C. metapsilosis, was confirmed in the experiments as shown in Fig. 1a–c, respectively. Primer sets, which amplify the DNA of C. parapsilosis sensu stricto only, did not amplify the DNA of other eight Candida species including C. albicans, C. dubliniensis, C. glabrata, C. guilliermondii, C. kefyr, C. krusei and C. tropicalis. Also, the LAMP primer sets amplified DNA of three clinical isolates of C. parapsilosis sensu stricto. This primer set did not amplify the DNA of two isolates of C. orthopsilosis and one isolate of C. metapsilosis.
Accurate identification profiles for three respective Candida species (C. parapsilosis sensu stricto, C. orthopsilosis and C. metapsilosis) by LAMP assay using specific primers for each Candida species. Following Candida strains were used as reference Candida species: C. albicans IFM 40213, C. dubliniensis IFM 48313, C. tropicalis IFM 577, C. parapsilosis IFM 46829, C. glabrata IFM 46843, C. guilliermondii IFM 46823, C. krusei IFM 46834, and C. kefyr IFM 5773. Amplified products based on rising curves of turbidity indicate specific DNA amplification for identifying corresponding Candida species. a–c indicate identification of (a) C. parapsilosis sensu stricto, b C. orthopsilosis and c C. metapsilosis, respectively
LAMP primer sets designed for C. orthopsilosis and C. metapsilosis, respectively, could amplify DNA for respective species, but no cross-reactivity against C. parapsilosis was observed. Either LAMP primer sets for the two species did not show cross-reactivity with C. albicans, C. dubliniensis, C. tropicalis, C. glabrata, C. guilliermondii, C. krusei and C. kefyr. Furthermore, it was confirmed that newly developed LAMP primer sets did not amplify the any DNA samples from Cryptococcus neoformans, which is frequently isolated from clinical samples of yeast-like fungal infection patients.
Discussion
The current epidemiology of candidemia in our hospital stressed CPC as agents that led to high mortality rate. In our hospital, the prevalence of C. orthopsilosis (11 %) and C. metapsilosis (2 %) among the CPC was similar to studies previously reported of Brazilian and South America data [34] and conversely higher than many other countries [34, 42]. In a recent study [43], Wisplinghoff et al. reported that CPC were the second most frequent agent of candidemia, accounting for 210 (17.4 %) cases among 1,218 Candida isolates obtained between 1998 and 2006 from 52 geographically dispersed hospitals in the USA. In this study, C. albicans (50.7 %) and C. glabrata (16.7 %) were the first and third most common Candida species, respectively [43]. Mortality associated with CPC candidemia in our patients (47.2 %) was higher than the current literature [1, 8] but similar to other Brazilian publications [6, 44]. In addition, C. orthopsilosis in our hospital exhibited higher mortality rate (75 %) than C. parapsilosis sensu stricto. Mortality rate due to C. metapsilosis could not be calculated because only one strain was isolated from 2006 to 2010. Since the number of patients in the present study was not high, we need to continue monitoring the mortality rate for further comparative studies on the recently isolated CPC.
Candidemia due to CPC, in our hospital, was mainly in patients with malignancies (50 %), long term of hospitalization and relatively old age. Surgery was performed in 27 (75 %) patients, of whom 70 % were related to the gastrointestinal tract. Most of our patients had several risk factors and invasive procedures known as associated with candidemia. The worse outcome was associated with patients with immunosuppressive conditions (p = 0.003) and short-time survival days (p = 0.04). The high mortality rate of our patients might be secondarily to multiple variables.
Although there were some fluctuations in MIC values in each species, the CPC isolates were very susceptible, and their MIC values were within those of the reference strain or proposed MIC values (CLSI). No resistant strains were confirmed. CPC is known for its decreased susceptibility to echinocandins [45]; however, our strains were very susceptible to micafungin (MIC50/MIC90: 0.5/0.5 μg/mL; range 0.125–1 μg/mL) and that could be explained because during the present study, echinocandins were sporadic prescribed. Two isolates, one C. orthopsilosis and one C. parapsilosis, had MIC = 4 μg/mL, which is considered susceptible dose dependent to fluconazole, according to the proposed species-specific epidemiologic cut-off value for C. parapsilosis [32, 46].
Though closely related, the species of the CPC differ from each other not only in prevalence [34] but also in virulence [47] and pathogenicity [18]. Recently, Kasahara et al. [48] reported the use of LAMP to detect Candida spp. in dairy products. The authors started from 4 C. parapsilosis, 1 C. parapsilosis and 1 C. orthopsilosis belonging to one private Culture Collection, in order to design a set of common primers, which were able to identify the CPC. Conversely, in our study, we design three sets of primers that are able to distinctly identify each one of C. parapsilosis, C. orthopsilosis and C. metapsilosis. This differentiation among the components of the C. parapsilosis complex has significance in the medical field, since each species can present differences in virulence [47], pathogenicity [18] and drug susceptibility, as we demonstrated at least for fluconazole. Therefore, the search for a fast, cost-effective and reliable method that discriminated among the species would be an important aid for the choice of optimal antifungal treatment in patients with life-threatening diseases. Many scientists had investigated molecular techniques for rapid diagnosis of fungal infection. Still identification or classification by sequencing of pathogenic fungi so far has been considered to be a “gold standard” for Candida yeast identification [26, 49]. However, sequencing of fungi is time-consuming and labor-intensive and may have reproducibility limitations. Therefore, the need of the application of fast and appropriate identification to species level is still increasing. The high potential of LAMP for the development of improved DNA-based diagnostic kits was reported [50]. In general, LAMP was found to be either similar or superior to PCR and more specific [51]. Actually, LAMP based approaches have been applied to a wide range of samples, such as paraffin-embedded tissues, whole blood and swabs [50]. Since there is concern over the increasing rates of the CPC infections worldwide [7, 49, 52], our present experiments confirmed the usefulness of LAMP primers for correct identification of each the three species of the complex.
The use of PCR assay for detection or discrimination of the Candida psilosis complex has been reported [7, 53]. Recently, a new identification of C. parapsilosis (sensu lato) by use of mitochondrial DNA and real-time PCR was proposed [7]. Actually, the method can distinguish C. parapsilosis (sensu stricto), C. metapsilosis and C. orthopsilosis. However, LAMP has some advantages over PCR and real-time PCR, including isothermal conditions for amplification, and in addition LAMP does not need special thermo-cycler equipment [41].
In conclusion, we analyzed the clinical and epidemiological data of a 5 years surveillance of patients with candidemia due to CPC in our hospital and the antifungal susceptibility of the isolates. We proposed a novel molecular method, LAMP, for the identification of species of CPC. Since we identified two isolates with fluconazole MICs = 4 µg/mL and a high mortality rate among the patients there is a need for a fast, cost-effective and reliable molecular method that can distinguished the members of the CPC and to monitor the trends in antifungal susceptibility and clinical outcomes. In addition, LAMP method might be very useful method in discriminating the CPC in studies such as the clinical trials monitoring the changes in the species distribution and antifungal susceptibility.
References
Trofa D, Gácser A, Nosanchuk JD. Candida parapsilosis, an emerging fungal pathogen. Clin Microbiol Rev. 2008;21:606–25.
Asmundsdottir LR, Erlendsdottir H, Gottfredsson M. Nationwide study of candidemia, antifungal use, and antifungal drug resistance in Iceland, 2000 to 2011. J Clin Microbiol. 2013;51:841–8.
Moretti ML, Trabasso P, Lyra L, Fagnani R, Resende MR, de Oliveira Cardoso LG, Schreiber AZ. Is the incidence of candidemia caused by Candida glabrata increasing in Brazil? Five-year surveillance of Candida bloodstream infection in a university reference hospital in southeast Brazil. Med Mycol. 2013;51:225–30.
Azevedo AC, Bizerra FC, da Matta DA, de Almeida LP, Rosas R, Colombo AL. In vitro susceptibility of a large collection of Candida strains against fluconazole and voriconazole by using the CLSI disk diffusion assay. Mycopathologia. 2011;171(6):411–6.
Horn DL, Neofytos D, Anaissie EJ, Fishman JA, Steinbach WJ, Olyaei AJ, Marr KA, Pfaller MA, Chang CH, Webster KM. Epidemiology and outcomes of candidemia in 2019 patients: data from the prospective antifungal therapy alliance registry. Clin Infect Dis. 2009;48(12):1695–703.
Colombo AL, Nucci M, Park BJ, Nouér SA, Arthington-Skaggs B, da Matta DA, Warnock D, Morgan J. Brazilian Network Candidemia Study. Epidemiology of candidemia in Brazil: a nationwide sentinel surveillance of candidemia in eleven medical centers. J Clin Microbiol. 2006;44(8):2816–23.
Canton E, Pemán J, Quindós G, Eraso E, Miranda-Zapico I, Álvarez M, Merino P, Campos-Herrero I, Marco F, de la Pedrosa EG, Yagüe G, Guna R, Rubio C, Miranda C, Pazos C, Velasco D. FUNGEMYCA Study Group. Prospective multicenter study of the epidemiology, molecular identification, and antifungal susceptibility of Candida parapsilosis, Candida orthopsilosis, and Candida metapsilosis isolated from patients with candidemia. Antimicrob Agents Chemother. 2011;55:5590–6.
Pfaller M, Neofytos D, Diekema D, Azie N, Meies-Kriesche HU, Quan SP, Horn D. Epidemiology and outcomes of candidemia in 3,648 patients: data from the prospective antifungal therapy (PATH Alliance®) registry, 2004–2008. Diagn Microbiol Infect Dis. 2012;74:323–31.
Das I, Nightingale P, Patel M, Jumaas P. Epidemiology, clinical characteristics, and outcome of candidemia; experience in a tertiary referral center in the UK. Int J Infect Dis. 2011;15:759–63.
Falagas ME, Roussos N, Vardakas KZ. Relative frequency of albicans and the various non-albicans Candida spp among candidemia isolates from inpatients in various parts of the world: a systematic review. Int J Infect Dis. 2010;14(11):e954–66.
Oeser C, Vergnano S, Naidoo R, Anthony M, Chang J, Chow P, Clarke P, Embleton N, Kennea N, Pattnayak S, Reichert B, Scorrer T, Tiron I, Watts T, Sharland M, Heath PT. The Neonatal Infection Surveillance Network (neonIN). Neonatal invasive fungal infection in England 2004–2010. Clin Microbiol Infect. 2014;20(9):936–41. doi:10.1111/1469-0691.12578.
Juyal D, Sharma M, Pal S, Rathaur VK, Sharma N. Emergence of non-albicans Candida species in neonatal candidemia. N Am J Med Sci. 2013;5:541–5.
Dotes J, Prasad PA, Zaoutis T, Roilides E. Epidemiology, risk factors and outcome of Candida parapsilosis bloodstream infection in children. Pediatr Infect Dis J. 2012;31:557–60.
Martino P, Girmenia C, Micozzi A, Raccah R, Gentile G, Venditti M, Mandelli F. Fungemia in patients with leukemia. Am J Med Sci. 1993;306:225–32.
Kuhn DM, Mukherjee PK, Clark Ta, et al. Candida parapsilosis characterization in an outbreak setting. Emerg Infect Dis. 2004;10:1074–81.
Levin AS, Costa SF, Mussi NS, et al. Candida parapsilosis fungemia associated with implantable and semi-implantable central venous catheter and the hands of health care workers. Diagn Microbiol Infect Dis. 1998;30:243–9.
Tavanti A, Davidson AD, Gow NAR, Maiden MCJ, Odds FC. Candida orthopsilosis and Candida metapsilosis spp. nov. to replace Candida parapsilosis groups II and III. J Clin Microbiol. 2005;43:284–92.
Gago S, García-Rodas R, Cuesta I, Mellado E, Alastruey-Izquierdo A. Candida parapsilosis, Candida orthopsilosis, and Candida metapsilosis virulence in the non-conventional host Galleria mellonella. Virulence. 2014;5(2):278–85.
Sakai K, Trabasso P, Moretti ML, Mikami Y, Kamei K, Gonoi T. Identification of fungal pathogens by visible microarray system in combination with isothermal gene amplification. Mycopathologia. 2014;178(1–2):11–26.
Buelow DR, Gu Z, Walsh TJ, Hayden RT. Evaluation of multiplexed PCR and liquid-phase array for identification of respiratory fungal pathogens. Med Mycol. 2012;50(7):775–80.
Muraosa Y, Schreiber AZ, Trabasso P, Matsuzawa T, Taguchi H, Moretti ML, Mikami Y, Kamei K. Development of cycling probe-based real-time PCR system to detect Fusarium species and Fusarium solani species complex (FSSC). Int J Med Microbiol. 2014;304(3–4):505–11.
Hall L, Wohlfiel S, Roberts GD. Experience with the MicroSeq D2 large-subunit ribosomal DNA sequencing kit for identification of filamentous fungi encountered in the clinical laboratory. J Clin Microbiol. 2004;42(2):622–6.
Hung WT, Su SL, Shiu LY, Chang TC. Rapid identification of allergenic and pathogenic molds in environmental air by an oligonucleotide array. BMC Infect Dis. 2011;11:91. doi:10.1186/1471-2334-11-91.
Nguyen MH, Wissel MC, Shields RK, Salomoni MA, Hao B, Press EG, Shields RM, Cheng S, Mitsani D, Vadnerkar A, Silveira FP, Kleiboeker SB, Clancy CJ. Performance of Candida real-time polymerase chain reaction, β-d-glucan assay, and blood cultures in the diagnosis of invasive candidiasis. Clin Infect Dis. 2012;54(9):1240–8.
Fricke S, Fricke C, Schimmelpfennig C, Oelkrug C, Schönfelder U, Blatz R, Zilch C, Faber S, Hilger N, Ruhnke M, Rodloff AC. A real-time PCR assay for the differentiation of Candida species. J Appl Microbiol. 2010;109(4):1150–8.
Souza ARC, Ferreira RC, Goncalves SS, Quindos G, Eraso GQ, Birerra FC, Briones MRS, Colombo AL. Accurate identification of Candida parapsilosis (sensu Lato) by use of mitochondrial DNA and real-time PCR. J Clin Microbiol. 2012;50:2310–4.
Mori Y, Kitao M, Tomita N, Notomi T. Real-time turbidimetry of LAMP reaction for quantifying template DNA. J Biochem Biophys Methods. 2004;59(2):145–57.
Uemura N, Makimura K, Onozaki M, Otsuka Y, Shibuya Y, Yazaki H, Kikuchi Y, Abe S, Kudo S. Development of a loop-mediated isothermal amplification method for diagnosis Pneumocystis pneumonia. J Med Mycol. 2008;57:50–7.
Poon LL, Leung CS, Chan KH, Lee LH, Yuen KY, Guan Y, Peiris JS. Detection of human influenza A viruses by loop-mediated isothermal amplification. J Clin Microbiol. 2005;43:427–30.
Endo S, Komori T, Ricci G, Sano A, Yokoyama K, Ohori A, Kamei K, Franco M, Miyaji M, Nishimura K. Detection of gp43 of Paracoccidioides brasiliensis by the loop-mediated isothermal amplification (LAMP) method. FEMS Microbiol Lett. 2004;234:93–7.
Iida S, Imai T, Oguri T, Okuzumi K, Yamanaka A, Moretti ML, Nishimura K, Mikami Y. Genetic diversity of the internal transcribed spacers (ITS) and 5.8S rRNA genes among the clinical isolates of Candida parapsilosis in Brazil and Japan Jap. J Med Mycol. 2005;46:133–7.
Clinical Laboratory Standards Institute (CLSI). Reference method for broth dilution antifungal susceptibility testing of yeasts; Fourth Informational Supplement. CLSI Document M27-S4. Wayne, PA: Clinical and Laboratory Standards Institute; 2012.
Singla N, Gulati N, Kaistha N, Chander J. Candida colonization in urine samples of ICU patients: determination of etiology, antifungal susceptibility testing and evaluation of associated risk factors. Mycopathologia. 2012;174(2):149–55.
Lockhart SR, Messer SA, Pfaller MA, Diekema DJ. Geographic distribution and antifungal susceptibility of the newly described species Candida orthopsilosis and Candida metapsilosis in comparison to the closely related species Candida parapsilosis. J Clin Microbiol. 2008;46:2659–64.
Pfaller MA, Diekema DJ, Andes D, Arendrup MC, Brown SD, Lockhart SR, Motyl M, Perlin DS. The CLSI subcommittee for antifungal testing Clinical breakpoints for equinocandins and Candida revisited: integration of molecular, clinical, and microbiological data to arrive at species-specific interpretive criteria. Drug Res Updates. 2011;14:164–76.
Pfaller MA, Andes D, Arendrup M, Diekema DJ, Espinel-Ingroff A, Alexander BD, Brown SD, Chaturvedi V, Fowler CL, Ghannoum MA, Johnson EM, Knapp CC, Motyl MR, Ostrosky-Zeichner L, Walsh TJ. Clinical breakpoints for voriconazole and Candida spp. revisited: review of microbiologic, molecular, pharmacodynamics, and clinical data as they pertain to the development of species-specific interpretive criteria. Diagn Microbiol Infect Dis. 2011;70:330–43.
Pfaller MA, Boyken L, Hollis RJ, Kroeger J, Messer SA, Tendolkar S, Wild-type Diekema DJ, IC M. distributions and epidemiological cutoff values for posaconazole and voriconazole and Candida spp. as determined by 24-h CLSI broth microdilution. J Clin Microbiol. 2010;49:630–7.
Pfaller MA, Andes D, Diekema DJ, Espinel-Ingroff A, Sheehan D. The CLSI subcommittee for antifungal susceptibility testing Wild-type MIC distributions, epidemiological cutoff values and species-specific clinical breakpoints for fluconazole and Candida: time for harmonization of CLSI and EUCAST broth microdilution methods. Drug Res Updates. 2010;13:180–95.
Katsu M, Kidd S, Ando A, Moretti ML, Mikami Y, Nishimura K, Meyer W. The internal transcribed spacers and 5.8S rRNA gene show extensive diversity among isolates of the Cryptococcus neoformans species complex. FEMS Yeast Res. 2004;4:377–88.
Tamura M, Watanabe K, Mikami Y, Yazawa K, Nishimura K. Molecular characterization of new clinical isolates of Candida albicans and C. dubliniensis in Japan: analysis reveals a new genotype of C. albicans with group I intron. J Clin Microbiol. 2001;39(12):4309–15.
Matsuzawa T, Tanaka R, Horie Y, Gonoi T, Yaguchi T. Development of rapid and specific molecular discrimination methods for pathogenic Emericella species. Nihon Ishinkin Gakkai Zasshi. 2010;51(2):109–16.
Silva AP, Miranda IM, Lisboa C, Pina-Vaz C, Rodrigues AG. Prevalence, distribution, and antifungal susceptibility profiles of Candida parapsilosis, C. orthopsilosis, and C. metapsilosis in a tertiary care hospital. Clin Microbiol. 2009;47:2392–7.
Wisplinghoff H, Ebbers J, Geurtz L, Stefanik D, Major Y, Edmond MB, Wenzel RP, Seifert H. Nosocomial bloodstream infections due to Candida spp. in the USA: species distribution, clinical features and antifungal susceptibilities. Clin Microbiol. 2014;43(1):78–81.
Brito LR, Guimaraes T, Nucci M, Rosas RC, Paula Almeida L, Da Matta DA, Colombo AL. Clinical and microbiological aspects of candidemia due to Candida parapsilosis in Brazilian tertiary care hospitals. Med Mycol. 2006;44:261–6.
Garcia-Effron G, Canton E, Peman J, Dilger A, Romá E, Perlin DS. Epidemiology and echinocandin susceptibility of Candida parapsilosis sensu lato species isolated from bloodstream infections at a Spanish university hospital. J Antimicrob Chemother. 2012;67:2739–48.
Pfaller MA, Diekema DJ, Procop GW, Rinaldi MG. Comparison of the Vitek 2 yeast susceptibility system with CLSI microdilution for antifungal susceptibility testing of fluconazole and voriconazole against Candida spp., using new clinical breakpoints and epidemiological cutoff values. Diagn Microbiol Infect Dis. 2013;77:37–40.
Németh T, Tóth A, Szenzenstein J, Horváth P, Nosanchuk JD, Grózer Z, Tóth R, Papp C, Hamari Z, Vágvolgyi C, Gácser A. Characterization of virulence properties in the C. parapsilosis sensu lato species. PLoS ONE. 2013;8:e68704.
Kasahara K, Ishikawa H, Sato S, Shimakawa Y, Watanabe K. Development of multiplex loop-mediated isothermal amplification assays to detect medically important yeasts in dairy products. FEMS Microbiol Lett. 2014;357(2):208–16.
Ferrari M, Resende M, Sakai K, Muraosa Y, Lyra L, Gonoi T, Mikami Y, Tominaga K, Kamei K, Schreiber A, Trabasso P, Moretti ML. Accurate visual DAN-microarray for the molecular identification of non-albicans Candida species isolated from candidemia episodes. J Clin Microbiol. 2013;51(11):3826–9.
Inacio J, Flores O, Spencer-Matrins I. Efficient identification of clinical relevant Candida yeast species by use of an assay combining panfungal loop-mediated isothermal DNA amplification with hybridization to species-specific oligonucleotide probes. J Clin Microbiol. 2008;46:713–29.
Hara-Kudo Y, Yoshino M, Kojima T, Ikedo M. Loop-mediated isothermal amplification for the rapid detection of Slamonella. FEMS Micribiol Lett. 2005;255:155–61.
Pfaller MA, Andes DR, Diekema DJ, Horn DL, Reboli AC, et al. Epidemiology and outcomes of invasive candidiasis due to non-albicans species of Candida in 2,496 Patients: data from the Prospective Antifungal Therapy (PATH) Registry 2004–2008. PLoS ONE. 2014;9(7):e101510. doi:10.1371/journal.pone.0101510.
Prandini TH, Theodoro RC, Bruder-Nascimento AC, Scheel CM, Bagagli E. Analysis of inteins in the Candida parapsilosis complex for simple and accurate species identification. J Clin Microbiol. 2013;51:2830–6.
Acknowledgments
This project was approved on February 22, 2011 (No. 039/2011) by the Ethical Committee of the School of Medical Sciences of the University of Campinas. This study was partly supported by JICA (Japan International Cooperation Agency) and SATREPS (Science and Technology Research Partnership for Sustainable Development) Grant No. 02P-29548-09.
Conflict of interest
All authors declare no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
The authors are grateful to Mrs. Luzia Lyra and Eliene Pinheiro for their notable contribution.
Rights and permissions
About this article
Cite this article
Trabasso, P., Matsuzawa, T., Fagnani, R. et al. Isolation and Drug Susceptibility of Candida parapsilosis Sensu Lato and other Species of C. parapsilosis Complex from Patients with Blood Stream Infections and Proposal of a Novel LAMP Identification Method for the Species. Mycopathologia 179, 53–62 (2015). https://doi.org/10.1007/s11046-014-9830-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-014-9830-9