Abstract
Psoriasis Severity and Area Index (PASI) is a gold standard scoring system for the assessment of Psoriasis skin disease. Generally, PASI scoring is done manually by expert dermatologists through visual and touch senses for psoriasis diagnosis and their treatment’s validation. This subjective approach raises several limitations and becomes unreliable. Many conventional and machine learning-based works are proposed for objective estimation of psoriasis area and severity from 2D RGB images. However, these works are validated on small datasets, require manual pre-processing, and rely heavily on hand-crafted features. In the proposed work, a fully automated system based on deep learning is designed for automated PASI scoring from raw 2D RGB images. This system contains a segmentation and three classification models for objective estimation of psoriasis area and severity scores for all three clinical symptoms of psoriasis, respectively. The psoriasis area is estimated by segmenting healthy and unhealthy regions simultaneously using a lightweight network as a backbone with UNet. After segmentation, the severity scores for each segmented lesion are automatically estimated by using a hybrid classification model. This model is developed by adopting a lightweight network for local feature extraction and integrating it with a vision transformer for learning global features. The psoriasis dataset used in the proposed work is self-prepared and contains 1,018 photographic images from different body regions of 212 psoriasis patients. The exhaustive performance analysis is done for the automatic estimation of each parameter of PASI. The proposed work achieves mean absolute error of 0.04, 0.23, 0.22, and 0.21 for objective estimation of Area, Redness, Scaliness, and Thickness scores, respectively. The mean absolute error obtained by the proposed system for automatic scoring of PASI is 1.02. The comparative studies with existing works further validate the efficacy of the proposed work. This work can further be improvised by using data from multi-centre and regions in a large population.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Psoriasis is a chronic, complex, autoimmune, and inflammatory skin disease affecting about 1–3% of the world’s population [1]. In psoriasis, skin cells grow more expeditious than in healthy human beings. For a healthy person, the development phase of skin cells is 28 to 30 days while it takes only 3 to 4 days in psoriasis patients [2]. It manifests as red, dry, itchy, raised, and scaly patches in the human skin. It generally occurs on the knees, scalp, elbows, and trunk, and further spreads to different parts of the body. It can develop in any human being regardless of age, gender, and community [3]. There is various form of psoriasis, but the most common form is plaque psoriasis [4]. The root cause of this skin disease is still unknown. However, the genetic fault is majorly evidenced as the cause of psoriasis [5]. Besides affecting the skin, it has an adverse impact on the quality of life because of its uncomfortable social presence and causes the risk of depression, stress, suicide, anxiety, etc. [6]. It is an incurable disease but can be controlled by regular and attentive treatments. Therefore, it’s considered a major concern to healthcare organizations and society [7, 8].
Psoriasis Area Severity Index (PASI) is used most often and accepted as a gold standard by dermatologists to measure the amount of existence and severity of plaque-type psoriasis in the patient [9]. It leads to an easy score for monitoring this disease and is standardized in daily practice [10]. It combines both psoriasis area and severity scores from all the body regions of a psoriasis patient. The percentage area of the body affected with psoriasis is scored from 0 to 6 as shown in Table 1. While, each clinical severity present in psoriasis (i.e. redness, scaliness, and thickness) is stratified on a scale of 0 to 4 score. The stratification details of these clinical severity parameters are given in Table 2 [1, 9].
All four major human body regions i.e., head and neck part (h), upper extremities part (ue), trunk part (t), and lower extremities part (le) from both front and rear sides are examined for overall PASI measurement in a psoriasis patient. Different weights (Wi) are assigned to consider body parts based on the area covered on the body surface. Hence, the weights Wh = 0.1, Wue = 0.2, Wt = 0.3, and Wle = 0.4 are included while measuring the overall PASI for a psoriasis patient (PASIpatient) that is represented by Eq. (1) [1, 9]. The maximum value of PASI for anybody region (i ∈ {h, ue, t, le}) is found to be equal to 72. Due to the consideration of the body part’s weights, the value of overall PASI is nearly continuous and ranges from 0 to 72.
In current practice, Dermatologists follow the visual inspection and the sense of touch to diagnose and further understand the existence and severity of this disease. This subjective approach required a highly skilled and experienced dermatologist. Also, manual assessment is usually a slow, hectic, and unreliable job for dermatologists. It also suffers from inter and intra-observer variability [11] and hence, fails to reproduce similar assessment reports. With so many subjective limitations, the psoriasis diagnosis, as well as monitoring, is a very tedious and challenging task [4, 12]. These subjective flaws can only be eliminated by adopting an objective approach [1]. Hence, the need for an objective assessment of psoriasis through a computer-aided diagnosis (CADx) system is clinically valuable and vital for dermatologists [13,14,15,16]. Dermatologists will benefit greatly from the CADx system for psoriasis since it will assist them with focused and reproducible diagnosis rapidly. This can also provide a good scope for researchers and medical experts to generalize behaviour or case studies, test hypotheses or theories, and discover new treatments as well [1, 17].
In this regard, many researchers have shown their interest and major contributions from objective segmentation of psoriasis lesions to its type and severity assessment from 2D digital images. To the best of our knowledge, a benchmark dataset of psoriasis is not available publicly to date. Therefore, every single research work on psoriasis uses a self-generated and private psoriasis image dataset [18, 19]. The objective approaches for the assessment of psoriasis lesions based on R, S, and T severity assessment are tabulated in Table 3. From Table 3, it can be observed that most of the existing research works are based on conventional image processing and/or machine learning (ML) algorithms [19]. In the ML paradigm, a manual feature engineering process is essentially required along with conventional image pre-processing steps. The manual extraction of relevant features is not an easy task and needs an expert from the domain of the dataset. This manual interaction makes ML approaches semi-automated and hence unreliable. The colour and texture features are used for redness and scaliness severity respectively in most of the ML approaches listed in Table 3 [20,21,22,23,24]. While not a single ML-based approach is developed for thickness severity scoring from 2D skin images. It can also be observed from Table 3 that these methods [20,21,22,23,24] are validated with less number of images.
Recent advancement of deep learning (DL) in computer vision and medical imaging domains has led to substantial improvement in performance and reliability over traditional ML techniques [19, 25, 26]. DL models extract different levels of reliable features automatically from digital images [27]. However, there is a limited amount of research work done based on a DL approach for psoriasis diagnosis [19]. In [28, 29], the DL model is adopted for objective assessment of psoriasis lesions based on R, S, and T severities (refer to Table 3). These approaches [28, 29] are validated using a required large number of cropped skin images of psoriasis lesions compared to other approaches [20,21,22,23,24] listed in Table 3.
PASI area scoring of lesions is done with reference to the healthy skin of the respective body region. Hence, PASI can’t be evaluated objectively only from cropped psoriasis lesions [1, 9, 16]. For this, the objective approach should be applied to images of different body regions for psoriasis patients. Only a few research works [30,31,32,33,34,35,36] have estimated A, R, S, T, and PASI from full-body 2D digital images of psoriasis patients using the ML or DL framework in the recent past. In [30], a two-stage ML algorithm is designed for measuring the psoriasis area where k-means clustering is implemented for segmentation on the images of two distinct colour spaces in two stages. A single-stage DL framework is designed in [31] for the automatic estimation of A using a modified UNet model. In [32], the conventional image processing algorithm is designed for objective scoring of each parameter of PASI and overall PASI too but validated on less number of test images. PSENet model based on DCNN with Siamese structure is proposed in [33]. This model directly estimates only PASI for a given input image without estimating any parameter of PASI. In [34], the PSENet model proposed in [33] is modified by introducing lesion attention modules (LAMs) for the measurement of PASI parameters which eventually gives the final PASI score for the image. In [35], a pre-trained ResNet18 model is adopted and fine-tuned to measure R, S, and T severities on an entire image. Moreover, the most important parameter of PASI i.e. A, is also estimated by the same classification model without segmenting psoriasis lesions from the image. Additionally, images with an area score of 6 and a severity score of 4 are not considered.
In this paper, a deep learning-based fully automated system for objective evaluation of PASI from 2D digital images by automatic severity assessment of segmented psoriasis lesions. A lightweight backbone network (i.e., MobileNetV2 [36]) is adopted in the UNet segmentation model with a transfer learning paradigm in this work for automatically segmenting the psoriasis lesion from the healthy skin for objective estimation of psoriasis area score (i.e., A). Further, the severity assessment of the segmented psoriasis lesion based on R, S, and T scores is performed automatically by the proposed classification model. The proposed classification model is developed by amalgamating a lightweight CNN model (i.e., MobileNetV2 [36]) and Vision Transformer (ViT) [37, 38]. The pre-trained MobileNetV2 model is adopted as a backbone CNN for local processing while ViT operates as a simple convolution for global processing. With this fusion, both local and global representations are learned effectively and efficiently by the proposed model. The class weighting, transfer learning, and data augmentation approaches are also utilized in the proposed model to deal with data imbalance and insufficiency issues [39,40,41,42,43,44,45]. These fusions of networks and approaches result in a better model for the intended task. The main contributions of the proposed work can be summed up as follows:
-
(i)
The proposed work is fully automated and designed for objective PASI scoring from 2D images along with the segmentation of healthy skin and psoriasis lesion regions.
-
(ii)
The experiments are conducted on the large self-prepared dataset with a considerable number of all PASI area and severity score images. It is validated with a five-fold cross-validation technique. The proposed classifier model is also found to be more potent and computationally efficient for the intended task when compared to other DL models.
-
(iii)
The result of each model is analyzed by computing various performance metrics with and without ± 1 tolerance. Intraclass Correlation Coefficient (ICC) is also computed which is an important statistical tool to measure the reliability of scoring.
-
(iv)
All the severity parameters of PASI are estimated by considering each plaque-psoriasis lesion in the digital body images of a psoriasis patient, which results in an adequate scoring for PASI when compared to other related works from the literature.
The remaining sections of the paper are organized as follows. Section 2 presents the methodology for objective scoring of PASI. Section 3 covers a detailed description of data and experimentations. The experimental results for each module of the proposed system with the evaluation of different performance metrics are demonstrated in Section 4. The ablation and comparative study analyses are done in Section 5. The research study is concluded in Section 6 and then, followed by recommendations for future research.
2 Methodology
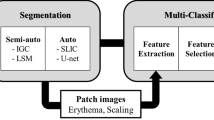
The proposed system is designed to estimate the PASI objectively from a 2D digital image of a plaque-psoriasis patient by predicting the area score (A) and three severity parameters (R, S, and T) scores. For this, the proposed system includes a segmentation model for the area and three classification models for each clinical severity. The overall flow diagram of the proposed methodology is illustrated in Fig. 1. The input used for the proposed work is in the form of raw 2D colour photographic images. The first stage of the proposed system consists of a segmentation module. In the segmentation module, psoriasis lesions from the skin region are automatically segmented. This automatically segmented image can be used for the objective estimation of A. Each segmented lesion from an input image of the first stage is fed as an input to the second stage. In the second stage, the severity scores based on R, S, and T factors are predicted automatically by the classification module. The classification module consists of three identical DL-based classification models for severity classification of R, S, and T. Lastly, the PASI for an input image is evaluated with the help of predicted scores of A, R, S, and T from the above methods.
2.1 Lesion segmentation module
In the first stage, a single-stage lesion segmentation module is used for the automatic calculation of the psoriasis lesion area percentage. The segmentation module is based on the architecture of a UNet model [46] having a lightweight MobileNetV2 network as a backbone. This model will perform multi-class segmentation of the image that will classify each pixel of the image into three different labels namely background, healthy skin, and psoriasis lesions simultaneously. The input of this segmentation model is a 2D RGB digital image of size 512 × 512 that results in an input shape of 512 × 512 × 3. Due to the symmetry nature of UNet, the output size is the same as the input size (i.e., 512 × 512). Therefore, the shape of the segmented output mask becomes 512 × 512 × 3 due to the multi-segmentation of three regions (i.e., background, healthy skin, and psoriasis lesion).
This module has two paths i.e. encoder and decoder that are connected U-shape. The encoder part of the model has a pre-trained MobileNetV2 network [36] for high-dimensional and relevant feature extraction which is further finely trained with the psoriasis dataset. MobileNetV2 network utilizes a linear bottleneck and inverted residual having depth-wise separable and pointwise convolution layers along with a residual connection. The use of depthwise separable convolution is not computationally expensive compared to standard convolution and hence makes the network lightweight. This versatile nature of the MobileNetV2 network allows easy training and makes it a popular backbone as a better and faster local feature extractor for several computer vision tasks [36]. The inner layer of this network exhibits the ability to switch the lower-level concepts like pixels to higher-level descriptors like image categories, while the bottlenecks encapsulate the network’s intermediary inputs and outputs. The short identity connection between the input and output of the block resembles a standard residual connection for improved performance. This backbone network comprises a sequence of convolution and bottleneck operations in five stages. In each stage, the receptive field of input is continuously increased whenever the input size is reduced by half. The details of layers in each stage with bottleneck and inverted residual design are stated in [36]. It performs local processing and extracts relevant and deep local feature representative maps of high spatial dimension. The decoder path expands the encoded features stage-wise to get the final segmented output of the same size as the input. The layer-wise details of the decoder path can be found in [31].
The percentage of skin area (i.e., Region of Interest, ROI) covered by psoriasis can be calculated with the help of Eq. (2). For this segmentation task, the total number of pixels belonging to psoriasis lesion region (NP) in the segmented mask output corresponds to the area of psoriasis lesion. The representation for skin region i.e., ROI area in the segmented mask output is the total number of pixels predicted as healthy skin (NH) or psoriasis lesion (NP). Therefore, Eq. (2) can be modified into Eq. (3) for the calculation of the percentage of psoriasis area (Ar) over the skin with the help of an automatically segmented mask. Based on the calculated percentage value of the psoriasis lesion area, the area score (A) parameter for PASI can be obtained using Table 1.
The automatically segmented psoriasis lesion image is obtained by overlaying the input RGB image with the segmented mask output. The area percentage of each psoriasis lesion (Arl) in the segmented mask output is also measured using Eq. (4) as shown below:
where; \({N}_{{P}_{l} }\) be the number of pixels belonging to each psoriasis lesion (l) and the sum of \({Ar}_{l}\) of all the individual segmented lesions in an image became equal to Ar. Further, the segmented lesions are cropped as per the closest bounding box to be fed as input to the second stage of the proposed work.
2.2 Lesion severity classification module
In the second stage, the objective assessment of the severity scores of lesions is done using a classification module. In this module, three identical DL-based classification model is designed and trained for each of the clinical severity parameters of PASI (R, S, and T). The model proposed for the severity classification task is developed and designed by combining lightweight CNN and ViT networks in series. In CNN, the properties of spatial inductive biases allow the network to learn local representations even with fewer parameters for different computer vision tasks [47]. One of the most popular lightweight networks that perform well for image classification tasks is MobileNetV2 [36]. Therefore, MobileNetV2 is selected as a lightweight backbone CNN to perform automatic local feature processing from the target dataset. ViT is based on multiple self-attention networks and shows a great performance for visual-related tasks due to input-adaptive weighting and global information learning [48]. ViT network in the proposed model operates as a standard convolution and thus, has convolution-like properties [38]. The proposed model incorporates properties of both CNN and ViT in the interest of building a high-performing and lightweight hybrid model for the intended task.
The architecture of the proposed model is represented in Table 4. The input RGB image of shape 128 × 128 × 3 is fed as input to the model. The input image is passed through a MobileNetV2 [36], which is pre-trained on the ImageNet dataset [49] (i.e., source) and fine-tuned with the psoriasis lesion dataset (i.e., target) for the intended task. The output shape of the local feature representative map generated automatically by the MobileNetV2 backbone network from the input image of 128 × 128 × 3 shape is 4 × 4 × 1280.
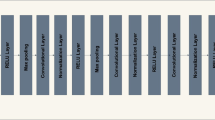
The high-dimensional local feature representative is projected to a point-wise convolution (having 64 filters) to capture and learn the linear combinations of input channels. Then, the convolved feature map is processed by ViT to capture long-range dependencies that represent global features [38, 50]. The structure of ViT to act as a simple convolution operation in the proposed model is illustrated in Fig. 2. ViT is used as a typical convolution involving unfolding, deeper global processing (replaces local processing in convolutions), and folding operations. The convolved feature map of resolution (H, W, k) is unfolded into small patches of considered sizes (i.e., 2 × 2) in a series. This results in N number of non-overlapping small patches of feature map having a resolution of (P, N, k) where P represents the area of small patch and N = (H×W)/P.
After being unfolded, this flattened vector of patches is then passed through a transformer block. This block is based on several self-attention layers that replicate long-range dependencies. It encodes the inter-patch relationships by interacting with each other to learn global representation. The global learning from the convolved feature map leads to better global representation [50]. The transformer block is generally made up of three significant processing components: Layer Normalization (LN), Multi-Headed Attention (MHA), and Multi-Layer Perceptron (MLP) [37]. LN controls the training process and enables the model to adjust to the differences between the training images. With MLP, attention maps are generated to keep the focus on important regions in the image. The input feature maps of the transformer block are added with concatenated attention maps before feeding forward to the MLP. MLP is used as an output of the transformer consisting of two fully connected layers to generate the encoded representations [37]. The final encoded global representation of the transformer block is obtained by adding the concatenated attention map output of MHA and the encoded representation output of MLP (refer to Fig. 2).
The local processing step of the convolution operation is replaced with deeper global processing in VIT. With this, the spatial information and patch order of pixels within each patch are not lost as happens in the case of standard ViT [37] and thus, the positional encoding operational layer of standard ViT is not required. The output of the transformer block is folded back again into a resolution of (H, W, k) to resemble a feature representation produced by a convolution operation. Thus, ViT operates more like a convolution despite a transformer and outputs a feature map having a respective field similar to the input feature map (i.e., H×W). This offers a different perspective to encode global representation that exhibits convolution-like properties such as spatial inductive biases, easy-to-learn/train/optimize, faster computation, etc. [38].
After the folding operation, the resolution of local and global feature representation maps became similar and both of these features are concatenated. The concatenated feature map is further fused by passing through a point-wise convolutional layer having filters equal to the channel size of the local or global feature i.e., 64 in this work. With this, the fused feature map efficiently encrypts both local and global representation with unchanged resolution. The fused feature map representation passes through a 2D global average pooling layer (GAP2D). GAP2D applies the average to H and W of the input feature map for all channels (k) [51, 52]. Lastly, a dense layer having the number of nodes equal to the total classes and softmax activation function is used in the output. The softmax helps in producing the final output of the model in terms of class probabilities within the range of [0,1] for each class.
2.3 Objective calculation of PASI
After the automated severity assessment of each segmented psoriasis lesion (l), the final severity scores (i.e., Rf, Sf, and Tf) of an input image are calculated based on the estimated area contribution and severity scores of each lesion present in the corresponding image. The mathematical equation for obtaining the final severity scores is represented by Eq. (5) which calculates the mean. The consideration of severity scores of each lesion based on its area leads to the effective calculation of the final severity score as well as PASI for an input image. In this way, all the parameters of PASI are estimated objectively and the objective calculation of PASI for an input image (PASIim) can be done using Eq. (6). With the proposed work, it is also possible to calculate the overall PASI for a psoriasis patient using Eq. (1). For overall PASI, PASIim should be calculated for both front and rear images of all four body regions of a psoriasis patient by the proposed system.
3 Data and experimentation details
3.1 Dataset details
For various skin diseases having lesions, dermatoscopy is used as a standardized image acquisition device [53, 54]. But for psoriasis, there is no such standardized image acquisition device or equipment exists. Hence, the psoriasis database is not available publicly anywhere. This left researchers with no option to self-prepare private psoriasis image databases using 2D digital cameras for psoriasis research [1, 16, 19].
In this work, the plaque-type psoriasis database was collected from the Psoriasis Clinic and Research Centre, Psoriatreat, Pune, Maharashtra, India. The digital images are obtained under the supervision of a team of dermatologists from Indian psoriasis patients irrespective of age, gender, and community. The database was generated with ethical approval and was anonymized. The digital images are acquired in an uncontrolled environment with an automatic setting using a Sony NEX-5 digital camera with a 22 mm lens and 350 dpi. The images are processed with an RGB colour depth of 24 bits per pixel in the Joint Photographic Expert Group (JPEG) format. All the major body regions of the patients i.e., upper extremities, trunk, and lower extremities from both front and rear sides are considered while capturing data. The sample images from the considered body region of psoriasis patients are shown in Fig. 3. However, head region images are not taken into account due to the patient’s privacy. For segmentation, the ground truth mask is labelled by the team of dermatologists for each image. Further, the severity of each psoriasis lesion is categorized into five different classes (i.e., none, mild, moderate, severe, and very severe). These annotations are also prepared with the help and support of a team of dermatologists based on all three severity parameters of PASI (i.e., R, S, and T). Figure 4 shows the sample skin images of psoriasis lesions under different severity scores of redness, scaliness, and thickness.
The self-prepared psoriasis image dataset contains 1,018 images of considered body regions from 212 psoriasis patients having a total of 5,075 psoriasis lesions. The distribution of lesions in each severity class of R, S, and T are tabulated in Table 5 which shows imbalanced data. Initially, 50 out of 212 patients were randomly selected as the test dataset for the proposed work that contains 350 images and 1,420 lesions. For the proposed severity classifier model, the remaining 162 patients that has 3,655 lesions from 668 images are considered for training and validation purpose using a stratified five-fold cross-validation technique [55]. With a five-fold stratified split, the ratio of training and validation data sets is 4:1. The simple pre-processing is done by resizing images of body regions into a common size of 512 × 512 for the segmentation task. The automatically segmented skin patch images of plaque psoriasis lesions are resized into a common size of 128 × 128 for the classification task.
3.2 System implementation details
All the models have been built using Python programming language [56] with the Keras API [57] and TensorFlow backend engine [58]. The experiment was conducted on a workstation having an Intel Xeon Silver CPU with 64GB RAM and NVIDIA Quadro P4000 GPU with 8GB memory. For efficient computation, pixel values of all the images are normalized into the range of [0,1]. The data augmentation and class weighting approaches are utilized to deal with data inadequacy and imbalance. The automatic data augmentation technique is applied randomly to generate augmented images during the training of the model using the Keras library [57]. The balanced class weight values [59] for our imbalanced psoriasis dataset are computed and assigned during model training. For further training/testing of the proposed severity classification module, the hyperparameter details are tabulated in Table 6. The hyperparameter details of the segmentation model are the same as described in [31].
4 Result
The performance is evaluated for each module of the proposed system and is presented in this section. For the segmentation module, the two potent segmentation performance metrics namely Dice-Coefficient Index (DI) and Jaccard Index (JI) are evaluated [62]. Due to the imbalanced number of lesions in different severity classes (refer to Table 5), the weighted averaging method is used for evaluating multi-class classification metrics (i.e., Precision, Recall, and F1-score) for the severity classification module [63, 64]. The multi-class Receiver Operating Characteristic (ROC) curve [65] analysis is also performed along with the computation of the area under the ROC curve (AUC) [66] for each severity class. In addition, the overall Accuracy (ACC) [63], Mean Absolute Error (MAE) [67, 68], and Intraclass Correlation Coefficient (ICC) [69] are also computed for evaluating the performance of the proposed system for automatic scoring of each parameter of PASI and PASI for a 2D RGB image of a psoriasis patient. ACC value close to 1 while an MAE value close to 0 indicates a better model’s performance [35]. ICC values lie in the range of [0,1]; where values lower than 0.5, between 0.5 and 0.75, between 0.75 and 0.9, and upper than 0.9 resembles the poor, moderate, good (acceptable), and high (adequate) level of agreement respectively [35, 69]. ACC and MAE metrics are also evaluated by considering the maximum variation of score 1 between automated and annotated scores (i.e., with ± 1 score tolerance) for each parameter of PASI [28, 29]. This is done in favour of concerning inter- and intra-observer variability issues from which psoriasis severity scoring suffers [1, 4, 11].
4.1 Result for segmentation module
The region-wise and overall segmentation result achieved by the segmentation model is tabulated in Table 7. The metrics value for the psoriasis lesion region is found to be less compared to the other two regions of healthy skin and background. This is due to the lower percentage distribution and high irregularity in the shape, size, number, dispersion, severity, noise, boundaries, etc. properties of the different lesions compared to the other two regions. However, the overall metrics result achieves values greater than 0.90 which represents good segmentation performance. Therefore, the automatic segmentation of psoriasis lesions by segmentation module is used for objective estimation of psoriasis area score (A). The final results achieved for the objective estimation of A using ACC, MAE, and ICC metrics are 0.9548, 0.0452, and 0.9490 respectively. With ± 1 score tolerance, ACC and MAE become 1.00 and 0.00 respectively as the value of absolute error observed is either 0 or 1 only. The visual segmentation results for sample two input images are presented in Fig. 5. These overall results confirm the better potential of the proposed segmentation model for the intended task.
4.2 Result for classification module
In this sub-section, the classification result is evaluated based on the severity prediction of each psoriasis lesion that is segmented automatically by the segmentation module of the proposed system. The performance of the proposed classifier model is acquired with 5-fold cross-validation for better generalizability. The average and fold-wise results of different performance metrics are tabulated in Tables 8, 9, and 10 for R, S, and T severities respectively. The average F1-score and overall ACC are greater than 0.75 and MAE is less than 0.30 for all three severity parameters in each fold. Also, the standard deviation among the folds is lower than 0.03. This shows the satisfactory and consistent performance of the proposed model in each fold for all severity parameters of PASI. The fold-average ACCs achieved for R, S, and T severities are 0.7903, 0.7764, and 0.8043 respectively. In addition, the values of the average F1-score and overall ACC are equal up to two decimals in each fold. Despite an imbalanced dataset, the proposed model is able to achieve stable performance for complex multi-classification without getting highly biased towards the majority class. Also, the ICC value is found to lie in the range of 0.75–0.90 in each case signifying overall good agreement between automated and manual severity scoring. The average value of ICC for R, S, and T severities achieved are 0.7997, 0.8084, and 0.8342 respectively. The severity results with ± 1 score tolerance for each case are tabulated in Table 11. From Table 11, it can be observed that the fold-average value of ACC became greater than 0.95 and MAE became less than 0.08 when ± 1 score tolerance is allowed. Hence, ACC and MAE values for R, S, and T severities are found to be significantly improved with ± 1 score tolerance.
The class-wise performance over all possible thresholds is summarized by plotting the ROC curve [65] and calculating AUC [66]. The ROC curves for the best fold of each severity are illustrated in Fig. 6. From Tables 8, 9, and 10, it can be observed that the folds f4, f1, and f0 for R, S, and T achieve the best classification performance respectively compared to other folds. Hence, Fig. 6a, b, and c represents the ROC curves of best fold f4 for R, f1 for S, and f0 for T respectively. From the ROC curves of Fig. 6, it can be observed that the average AUC score in each class of R, S, and T severity is greater than or equal to 0.9. This indicates the desirable performance of the proposed model in discriminating between the actual and other classes.
4.3 Result for objective estimation of PASI
The result presented in this sub-section is evaluated considering images of psoriasis patients from test datasets. Table 12 represents the performance for automatic estimation of each parameter of PASI and objective measurement of the final PASI for an image of a psoriasis patient. From Table 12, it can be observed that ACC achieved for automatic estimation of R, S, T, and A are 0.7971, 0.7943, 0.8000, and 0.9600 respectively. For PASI, the ACC can’t be considered an ideal performance metric because it is not possible to correctly predict the PASI whose numeric values are nearly continuous that ranges from 0 to 72. Therefore, the MAE metric is computed for PASI and achieved a value of 1.0200. The MAE values of 0.2314, 0.2200, 0.2086, and 0.0400 are achieved for R, S, T, and A respectively. With ± 1 score tolerance limit, a significant amount of improvement can be observed from Table 12 in the automatic scoring of each parameter of PASI. After considering ± 1 error, the value of ACC becomes [R:0.9714, S:0.9857, T:0.9914, A:1.0000] and MAE becomes [R:0.0571, S:0.0285, T:0.0171, A:0.0000]. For PASI, the value of MAE decreases from 1.0200 to 0.8029 with ± 1 score tolerance. The ICC values achieved for R, S, and T parameters are 0.7806, 0.8024, and 0.8205 which lie between the range of 0.75–0.9 indicating good agreement. The ICC value of 0.9630 (i.e., greater than 0.9) achieved for the most important parameter of PASI i.e., A indicates a high level of agreement. Therefore, the objective calculation of the PASI score from the proposed system also achieves an adequate agreement level with an ICC value of 0.9100. These results overall signify that the proposed system has significant potential for the objective evaluation of PASI.
5 Discussion
5.1 Ablation study on the proposed classification model
The effect of ViT in the proposed model is validated by removing it from the model. Without ViT, the model is left with only the MobileNetV2 network as the backbone CNN for local processing and decision-making layers having a GAP2D and a fully connected layer with a softmax activation function. For this, the implementation and experimentation of the model without and with ViT are performed on a similar computing platform and with the same train/test dataset (i.e., from fold f0). The comparison of performance achieved by both of these models is made in Table 13. From Table 13, the gain in the value of each performance metric with ViT can be observed for all three severity parameters of PASI. This validates that the integration of ViT in the model augments the classification performance for the intended task.
5.2 Impact of class weighting
The performance comparison of the proposed model with and without class weight for the lesion severity classification task with the same train/test dataset (i.e., from fold f0) is made in Table 14. From Table 14, it can be observed that there is a boost in the classification performance of the proposed model with class weighting. Due to class imbalance in the dataset, the normalized confusion matrix [70] is also plotted for better insights into the impact of class weighting in the multi-classification task. The normalized confusion matrix of the proposed model with and without class weight are illustrated in Fig. 7. The confusion matrix without class weight (refer to Fig. 7a, c, and e) shows the higher diagonal value (i.e. correct prediction rate) and lower off-diagonal value (i.e. misclassification/error rate) only for the classes having a higher number of observations. From Fig. 7b, d, and f, it can be observed that the diagonal values are very high compared to off-diagonal values in each class of the normalized confusion matrix of the model trained with class weight. This validates that the class-weighting addresses the class imbalance in the model and thus, leads to a good classification model despite imbalanced data.
5.3 Comparative study for psoriasis severity assessment
The proposed model for severity assessment of psoriasis lesions is compared with other existing DL models from the literature [16, 28, 29, 35, 71]. This comparison is made based on both multi-class classification performance and the number of trainable parameters in the model. Table 15 tabulates the comparison of the proposed model with different DL models implemented for the intended task. The implementation and experimentation of these models are performed on a similar computing platform and with the same train/test dataset (i.e., from fold f0). The class weight and data augmentation are adopted with all the models while training. The transfer learning concept is involved with standard CNN models (i.e., ResNet18 [35], ResNet50 [29], MobileNet [29]) by using pre-trained weights trained on ImageNet and fine-tuning them with our psoriasis lesion dataset. From Table 15, it can be observed that the performance of the proposed model is comparatively higher compared to that of other DL models. Also, the proposed model provides a good balance between performance and trainable parameters.
5.4 Comparative study for objective PASI scoring
The proposed work is compared with other existing recent works [30,31,32,33,34,35] for objective scoring of PASI or any one of the PASI parameters from 2D digital images of psoriasis patients in Table 16. From Table 16, it can be observed that only the proposed work utilizes both DL-based segmentation (i.e., MobileNetv2 + UNet) and classification (MobileNetv2 + ViT) models for automated scoring of lesion area (A) and severity (R, S, T), respectively. Also, the proposed work is the only single and fully automated research work in Table 16 that provides objective scoring of PASI by finding all the parameters of PASI with simultaneous automated segmentation of psoriasis and healthy regions. However, the existing works [30, 31] performs segmentation task for objective scoring of the A parameter only. While the remaining existing works [33,34,35] only performs classification task for objective scoring without segmenting the psoriasis lesion. Due to both segmentation and classification tasks, the performance achieved by the proposed work for objective scoring of PASI is better compared to other existing works. The objective severity assessment of an image by considering all the severity levels from each segmented psoriasis lesion present in that image is another important reason behind achieving better performance in the proposed work.
6 Conclusion
The proposed work develops an automated complete PASI scoring system for the assessment of psoriasis from raw 2D and RGB colour images of the body regions of patients. It is fully automated as it is based on a DL framework and doesn’t rely on hand-crafted features. This will be beneficial for dermatologists by providing an objective way for automatic estimation of different parameters of PASI i.e., psoriasis area (A) and clinical severity (R, S, and T) scores. In the proposed system, the segmentation task is performed using a UNet model with the MobileNetV2 network as a backbone. The severity classification task is carried out on each segmented lesion by a hybrid model having the fused strengths of both local and global processing from MobileNetV2 and ViT, respectively. The exhaustive experimental analysis with five-fold cross-validation is performed and different performance metrics are evaluated to show the adequacy of the proposed system. On the test dataset, the segmentation model in the proposed system achieves 0.0400MAE for psoriasis area scoring with an ICC of 0.9630. The proposed classification model developed for redness, scaliness, and thickness severity scoring achieves MAE of 0.2314, 0.2200, and 0.2086 with ICC of 0.7806, 0.8024, and 0.8205 respectively. However, the overall proposed system achieves MAE and ICC of 1.0200 and 0.9100, respectively for the objective calculation of the PASI score. However, these performances are getting further improved on the inclusion of ± 1 score tolerance limit such as MAE becomes 0.0571 for R, 0.0285 for S, 0.0171 for T, 0.0000 for A, and 0.8029 for PASI. These results of the proposed system are found outstanding when compared with other different fully automated approaches from the literature.
Thus, the proposed PASI scoring system has the potential to avoid the tedious and manual calculation of PASI in routine mode. It also offers quantitative and reproducible diagnosis and assessment for psoriasis treatment. Thus, it can easily be adaptable in large psoriasis research or clinical studies such as drug evaluation and therapy, etc. for psoriasis treatment. Also, the lightweight models used in the proposed system are mobile-friendly and they can be used by the concerned to monitor the disease severity remotely with ease.
The fusion of features from multi-scale and multi-view inputs with self-supervised DL models can be explored and incorporated in the future to improve the performance of automatic severity assessment of psoriasis lesions. Further, a better and more generalized PASI scoring system may be developed by utilizing a large psoriasis image dataset of multiple skin types annotated by multiple experts.
Data availability
The data that support the findings of this study are not openly available due to reasons of sensitivity and are available from the corresponding author upon reasonable request.
References
Shrivastava VK, Londhe ND, Sonawane RS, Suri JS (2015) First review on psoriasis severity risk stratification: an engineering perspective. Comput Biol Med 63:52–63
Nestle FO, Conrad C (2004) Mechanisms of psoriasis. Drug Discov Today: Dis Mech 1(3):315–319
Henseler T (1997) The genetics of psoriasis. J Am Acad Dermatol 37(2):S1–S11
Puzenat E, Bronsard V, Prey S, Gourraud PA, Aractingi S, Bagot M, ..., Aubin F (2010) What are the best outcome measures for assessing plaque psoriasis severity? A systematic review of the literature. J Eur Acad Dermatol Venereol 24:10–16
Chandran V, Raychaudhuri SP (2010) Geoepidemiology and environmental factors of psoriasis and psoriatic arthritis. J Autoimmun 34(3):J314–J321
Olivier C, Robert PD, Daihung DO, Urbà G, Catalin MP, Hywel W, ..., Gelfand JM (2010) The risk of depression, anxiety, and suicidality in patients with psoriasis: a population-based cohort study. Arch Dermatol 146(8):891–895
Huerta C, Rivero E, Rodríguez LAG (2007) Incidence and risk factors for psoriasis in the general population. Arch Dermatol 143(12):1559–1565
Menter A, Korman NJ, Elmets CA, Feldman SR, Gelfand JM, Gordon KB, Bhushan R (2011) Guidelines of care for the management of psoriasis and psoriatic arthritis: Sect. 6. Guidelines of care for the treatment of psoriasis and psoriatic arthritis: case-based presentations and evidence-based conclusions. J Am Acad Dermatol 65(1):137–174
Schmitt J, Wozel G (2005) The psoriasis area and severity index is the adequate criterion to define severity in chronic plaque-type psoriasis. Dermatology 210(3):194–199
Feldman SR, Krueger G (2005) Psoriasis assessment tools in clinical trials. Ann Rheum Dis 64(suppl 2):ii65–ii68
Fink C, Alt C, Uhlmann L, Klose C, Enk A, Haenssle HA (2018) Intra-and interobserver variability of image‐based PASI assessments in 120 patients suffering from plaque‐type psoriasis. J Eur Acad Dermatol Venereol 32(8):1314–1319
Chalmers RJ (2015) Assessing psoriasis severity and outcomes for clinical trials and routine clinical practice. Dermatol Clin 33(1):57–71
Maglogiannis I, Doukas CN (2009) Overview of advanced computer vision systems for skin lesions characterization. IEEE Trans Inf Technol Biomed 13(5):721–733
Chang W-Y, Huang A, Yang C-Y, Lee C-H, Chen Y-C, Wu T-Y (2013) Computer-aided diagnosis of skin lesions using conventional digital photography: a reliability and feasibility study. PLoS ONE 8(11):e76212
Razmjooy N, Somayeh Mousavi B, Soleymani F, Hosseini Khotbesara M (2013) A computer-aided diagnosis system for malignant melanomas. Neural Comput Appl 23:7–8
Dash M, Londhe ND, Ghosh S, Raj R, Sonawane RS (2020) A cascaded deep convolution neural network based CADx system for psoriasis lesion segmentation and severity assessment. Appl Soft Comput 91:106240
Morrow T (2004) Evaluating new therapies for psoriasis. Manag Care 13:34–40
Balestrieri E, Lamonaca F, Lembo S, Miele G, Cusano F, De Cristofaro GA (2019) Automatic psoriasis assessment methods: current scenario and perspectives from a metrologic point of view. In: 2019 IEEE International Symposium on Medical Measurements and Applications (MeMeA), IEEE, pp. 1–6
Yu K, Syed MN, Bernardis E, Gelfand JM (2020) Machine learning applications in the evaluation and management of psoriasis: a systematic review. J Psoriasis Psoriatic Arthritis 5(4):147–159
Lu J, Kazmiercazk E, Manton JH, Sinclair R (2012) Automatic scoring of erythema and scaling severity in psoriasis diagnosis. In: AI 2012: Advances in Artificial Intelligence: 25th Australasian Joint Conference, Sydney, Australia, December 4–7, 2012. Proceedings 25 (pp. 73–84). Springer Berlin Heidelberg
Banu S, Toacse G, Danciu G (2014) Objective erythema assessment of Psoriasis lesions for Psoriasis Area and Severity Index (PASI) evaluation. In: 2014 International Conference and Exposition on Electrical and Power Engineering (EPE), IEEE, pp. 052–056
Raina A, Hennessy R, Rains M, Allred J, Hirshburg JM, Diven DG, Markey MK (2016) Objective measurement of erythema in psoriasis using digital color photography with color calibration. Skin Res Technol 22(3):375–380
George Y, Aldeen M, Garnavi R (2018) Psoriasis image representation using patch-based dictionary learning for erythema severity scoring. Comput Med Imaging Graph 66:44–55
George Y, Aldeen M, Garnavi R (2019) Automatic scale severity assessment method in psoriasis skin images using local descriptors. IEEE J Biomedical Health Inf 24(2):577–585
Serte S, Serener A, Al-Turjman F (2022) Deep learning in medical imaging: a brief review. Trans Emerg Telecommun Technol 33(10):e4080
Li LF, Wang X, Hu WJ, Xiong NN, Du YX, Li BS (2020) Deep learning in skin disease image recognition: a review. IEEE Access 8:208264–208280
Mathew A, Amudha P, Sivakumari S (2021) Deep learning techniques: an overview. Adv Mach Learn Technol Appl: Proc AMLTA 2020:599–608
Pal A, Chaturvedi A, Garain U, Chandra A, Chatterjee R (2016) Severity grading of psoriatic plaques using deep CNN based multi-task learning. In: 2016 23rd International Conference on Pattern Recognition (ICPR), IEEE, pp. 1478–1483
Pal A, Chaturvedi A, Garain U, Chandra A, Chatterjee R, Senapati S (2018) Severity assessment of psoriatic plaques using deep cnn based ordinal classification. In: OR 2.0 Context-Aware Operating Theaters, Computer Assisted Robotic Endoscopy, Clinical Image-Based Procedures, and Skin Image Analysis: First International Workshop, OR 2.0 2018, 5th International Workshop, CARE 2018, 7th International Workshop, CLIP 2018, Third International Workshop, ISIC 2018, Held in Conjunction with MICCAI 2018, Granada, Spain, September 16 and 20, 2018, Proceedings 5 (pp. 252–259). Springer International Publishing
Tancharoen D, Tantawiwat P, Kovintavewat P (2019) Medical imaging using automatic region of interest segmentation for psoriasis diagnosis. In: 2019 34th International Technical Conference on Circuits/Systems, Computers and Communications (ITC-CSCC), IEEE, pp. 1–4
Raj R, Londhe ND, Sonawane RS (2021) Deep learning based multi-segmentation for automatic estimation of psoriasis area score. In: 2021 8th International Conference on Signal Processing and Integrated Networks (SPIN), IEEE pp. 1137–1142
Fink C, Fuchs T, Enk A, Haenssle HA (2018) Design of an algorithm for automated, computer-guided PASI measurements by digital image analysis. J Med Syst 42:1–8
Li Y, Wu Z, Zhao S, Wu X, Kuang Y, Yan Y, …, Wang Y (2020) PSENet:Psoriasis severity evaluation network. In: Proceedings of the AAAI Conference on Artificial Intelligence 34(01):800–807
Wu X, Yan Y, Zhao S, Kuang Y, Ge S, Wang K, Chen X (2021) Automatic severity rating for improved psoriasis treatment. In: Medical Image Computing and Computer Assisted Intervention–MICCAI 2021: 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part VII 24 (pp. 185–194). Springer International Publishing
Schaap MJ, Cardozo NJ, Patel A, De Jong EMGJ, Van Ginneken B, Seyger MMB (2022) Image-based automated psoriasis area severity index scoring by convolutional neural networks. J Eur Acad Dermatol Venereol 36(1):68–75
Sandler M, Howard A, Zhu M, Zhmoginov A, Chen LC (2018) Mobilenetv2: Inverted residuals and linear bottlenecks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 4510–4520
Vaswani A, Shazeer N, Parmar N, Uszkoreit, J, Jones L, Gomez AN, …, Polosukhin I (2017) Attention is all you need. Adv Neural Inf Process Syst 30
Mehta S, Rastegari M (2022) MobileViT: light-weight, general-purpose, and mobile-friendly vision transformer. Computer Vision and Pattern Recognition. https://doi.org/10.48550/arXiv.2110.02178
Fernández A, García S, Galar M, Prati RC, Krawczyk B, Herrera F, …, Herrera F (2018) Cost-sensitive learning. Learning from Imbalanced Data Sets 63–78
Johnson JM, Khoshgoftaar TM (2019) Survey on deep learning with class imbalance. J Big Data 6(1):1–54
Song B, Li S, Sunny S, Gurushanth K, Mendonca P, Mukhia N, …, Liang R (2021) Classification of imbalanced oral cancer image data from high-risk population. J Biomed Optics 26(10):105001–105001
Zhuang F, Qi Z, Duan K, Xi D, Zhu Y, Zhu H, …, He Q (2020) A comprehensive survey on transfer learning. Proc IEEE 109(1):43–76
Shorten C, Khoshgoftaar TM (2019) A survey on image data augmentation for deep learning. J big data 6(1):1–48
Anaya-Isaza A, Mera-Jiménez L (2022) Data augmentation and transfer learning for brain tumor detection in magnetic resonance imaging. IEEE Access 10:23217–23233
Rai R, Sisodia DS (2021) Real-time data augmentation based transfer learning model for breast cancer diagnosis using histopathological images. In: Advances in Biomedical Engineering and Technology: Select Proceedings of ICBEST 2018 (pp. 473–488). Springer Singapore
Ronneberger O, Fischer P, Brox T (2015) U-net: Convolutional networks for biomedical image segmentation. In: Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, October 5–9, 2015, Proceedings, Part III 18 (pp. 234–241). Springer International Publishing
Albawi S, Mohammed TA, Al-Zawi S (2017) Understanding of a convolutional neural network. In: 2017 International Conference on Engineering and Technology (ICET), IEEE, pp. 1–6
Han K, Wang Y, Chen H, Chen X, Guo J, Liu Z, …, Tao D (2022) A survey on vision transformer. IEEE Trans Pattern Anal Mach Intell 45(1):87–110
Russakovsky O, Deng J, Su H, Krause J, Satheesh S, Ma S, …, Fei-Fei L (2015) Imagenet large scale visual recognition challenge. Int J Comput Vis 115:211–252
Xiao T, Singh M, Mintun E, Darrell T, Dollár P, Girshick R (2021) Early convolutions help transformers see better. Adv Neural Inf Process Syst 34:30392–30400
Koffas S, Picek S, Conti M (2022) Dynamic backdoors with global average pooling. In: 2022 IEEE 4th International Conference on Artificial Intelligence Circuits and Systems (AICAS), IEEE, pp. 320–323
Kumar RL, Kakarla J, Isunuri BV, Singh M (2021) Multi-class brain tumor classification using residual network and global average pooling. Multimed Tools Appl 80:13429–13438
Errichetti E, Stinco G (2016) Dermoscopy in general dermatology: a practical overview. Dermatol Ther 6:471–507
Anand V, Gupta S, Nayak SR, Koundal D, Prakash D, Verma KD (2022) An automated deep learning models for classification of skin disease using dermoscopy images: a comprehensive study. Multimed Tools Appl 81(26):37379–37401
Lei J (2020) Cross-validation with confidence. J Am Stat Assoc 115(532):1978–1997
Python W (2021) Python. Python releases for windows, 24
Chollet F (2018) Keras: the python deep learning library. Astrophysics source code library, pp ascl–1806
Martín A, Ashish A, Paul B, Eugene B, Zhifeng C, Craig C, …, Matthieu D (2015) TensorFlow: Large-scale machine learning on heterogeneous systems. Software available from tensorflow.org
King G, Zeng L (2001) Logistic regression in rate events data, Harvard University. Center for Basic Research in the Social Sciences
Koidl K (2013) Loss functions in classification tasks. School of Computer Science and Statistic Trinity College, Dublin
Kingma DP, Ba J (2015) Adam: A method for stochastic optimization, 3rd International Conference for Learning Representations, San Diego. https://doi.org/10.48550/arXiv.1412.6980
Setiawan AW (2020) Image segmentation metrics in skin lesion: accuracy, sensitivity, specificity, dice coefficient, Jaccard index, and Matthews correlation coefficient. In: 2020 International Conference on Computer Engineering, Network, and Intelligent Multimedia (CENIM), IEEE, pp. 97–102
Grandini M, Bagli E, Visani G (2020) Metrics for multi-class classification: an overview. arXiv preprint arXiv:2008.05756
Mortaz E (2020) Imbalance accuracy metric for model selection in multi-class imbalance classification problems. Knowl Based Syst 210:106490
Hoo ZH, Candlish J, Teare D (2017) What is an ROC curve? Emerg Med J 34(6):357–359
Narkhede S (2018) Understanding auc-roc curve. Towards Data Science 26(1):220–227
Willmott CJ, Matsuura K (2005) Advantages of the mean absolute error (MAE) over the root mean square error (RMSE) in assessing average model performance. Climate Res 30(1):79–82
Chai T, Draxler RR (2014) Root mean square error (RMSE) or mean absolute error (MAE)?–Arguments against avoiding RMSE in the literature. Geosci Model Dev 7(3):1247–1250
Bartko JJ (1966) The intraclass correlation coefficient as a measure of reliability. Psychol Rep 19(1):3–11
Krstinić D, Braović M, Šerić L, Božić-Štulić D (2020) Multi-label classifier performance evaluation with confusion matrix. Computer Science & Information Technology 1. https://doi.org/10.5121/csit.2020.100801
Zivkovic M, Bacanin N, Antonijevic M, Nikolic B, Kvascev G, Marjanovic M, Savanovic N (2022) Hybrid CNN and XGBoost model tuned by modified arithmetic optimization algorithm for COVID-19 early diagnostics from X-ray images. Electronics 11(22):3798
Acknowledgements
We thank all the dermatologists and psoriasis patients of Psoriasis Clinic and Research Centre, Psoriatreat, Pune, Maharashtra, India who are involved in this research.
Funding
No funding was received for conducting this study.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Material preparation was performed by Ritesh Raj and Narendra Londhe. Data collection and analysis were performed by Ritesh Raj and Rajendra Sonawane. The first draft of the manuscript was written by Ritesh Raj and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Raj, R., Londhe, N.D. & Sonawane, R.S. Objective scoring of psoriasis area and severity index in 2D RGB images using deep learning. Multimed Tools Appl 83, 68253–68279 (2024). https://doi.org/10.1007/s11042-024-18138-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11042-024-18138-7