Abstract
Glioma is among aggressive and common brain tumors, with a low survival rate, in its highest grade. Invasive methods, i.e., biopsy and spinal tap are clinically used to determine the grades of glioma. Depending upon the findings of these methods, treatment is planned to improve the life expectancy of the controls. Magnetic resonance imaging (MRI), the most widely used medical imaging modality to diagnose a brain tumor, is producing a huge volume of MRI data. A reliable, automatic, and noninvasive method of glioma grading are always required as an alternative to these invasive methods. In this research, a model has been proposed using Convolutional Neural Networks to classify low and high-grade glioma. A locally organized dataset, developed in the Department of Radiology (Diagnostics), Bahawal Victoria Hospital, Bahawalpur, Pakistan has been used for research and experiments. Additionally, results have also been validated on a publicly available benchmarked dataset, i.e., BraTS-2017. The proposed method demonstrated significant achievement in terms of classification rates, i.e., the accuracy of 98.93% (for low-grade glioma) and 98.12% (for high-grade glioma). Experimental results proved that the proposed model is accurate (98.52%) and is efficient in glioma grade identification.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
With an invention of a new technology barrier, i.e., deep learning, demand for embedded processing capabilities has also increased. Diagnosis through medical traits using automated manners has become a demanding area, in which early-stage detection of abnormalities can avoid life threats [6]. It necessitates the existence of a more accurate, automated, and non-invasive diagnosis protocol. To date, clinically recognized standard protocols executed for brain tumor grading are invasive.
The human body is built upon tissues, and organs, composed of tiny masses called cells [13]. When the normal working of cells is disturbed, their rapid division into more and more cells in an uncontrolled manner is known as tumor [8]. Cells analysis under a microscope determines their several states, including presence, benign or malignant, primary or secondary, low or high grade, extent, etc., overall, this examination is called a biopsy [14]. Since it requires resection of cells, therefore, is riskier and requires time to recover. Resultantly, it can cause bleeding in the brain, swelling, infections, stroke, coma, or seizures [22]. If affected tissues do not possess cancerous cells are benign, otherwise, malignant [27]. If cancerous cells remain in the organ, in which they originate are primary, otherwise secondary. Similarly, if infected cells have a low growth rate, fall under the lower grade category, otherwise in a higher grade.
According to World Health Organization (WHO), brain tumor has almost 120 types [9]. Depending upon their growth rates, brain tumors are assigned their grades, ranging from Grade I to Grade IV [12]. The human brain comprises almost 86 billion nerve cells, called gray matter or neurons, while almost billions of nerve fibers, called dendrites and axons, or white matter [1]. The glial cells are present surrounding the neurons to provide support [2]. Brain tumors located in glial cells, called glioma, are classified as low-grade glioma (LGG), and high-grade glioma (HGG). Clinically, gliomas of Grade-I and Grade-II are classified as LGG, while gliomas of Grade-III and Grade-IV as HGG.
Besides several other medical imaging modalities, MRI with T1 weighted (T1W), T2 weighted (T2W), proton density weighted (PDW), and fluid attenuated inversion recovery (FLAIR) have an important role in diagnosis, surveillance, characterization, and therapeutic monitoring of a glioma. MRI enables the clinicians to scan patients in sagittal, coronal, and axial orientation These MRI sequences and views help a medical practitioner to visualize tissues in a better way, so that tissues be, interpreted and analyzed meaningfully. Each sequence has its importance in the clinical environment in connection with a detailed study of anatomy, monitoring of pathology-related changes, evaluation, etc. Although mostly, T1w helps in the study of normal anatomy, however, with the use of intravenous contrast, active tumor areas, and necrotic are easily recognized using this sequence. Edema, a brain tumor surrounding area becomes easier to distinguish using the T2W sequence. Most of the time, FLAIR is used to suppress intraventricular cerebrospinal fluid (CSF) signals and it plays a vital role in separating and studying CSF and edema. Similarly, multiple sclerosis (MS) lesions are evaluated mostly with PDW sequence. Further to this, gliomas have a distinctive appearance on MRI scans, which enable clinicians to determine a possible diagnosis. Figures 1 and 2 represent images of T1w Axial, T2w Axial, T1 Sagittal, T2 Sagittal, and FLAIR Axial for each low-grade and high-grade glioma respectively.
The literature reviewed established that most of the work performed for glioma grading has classification accuracies to up to 96%, which is not up to the mark, and are trained on a limited volume of the dataset. Additionally, the models, they reported are not evaluated on cross datasets. To address these issues, a light weighted CNN model with less number of parameters has been proposed to classify glioma grades through MRI images in a non-invasive way. This study is conducted in continuation of the previously conducted work [9], in which astrocytoma grades were classified using CNN model, designed specifically to that study. The difference between these studies is the proposal of CNN models, designed specifically for their dedicated tasks, i.e., glioma and astrocytoma grade identification. Overall, these are modules of our ongoing big project ‘brain radiology’, designed and developed for the Department of Radiology (Diagnostics), Bahawal Victoria Hospital, Bahawalpur (BVHB), Pakistan.
The rest of the draft is organized as follows; the second section contains reviewed literature; a proposed architecture for glioma grading is positioned in the third section. The results with their discussion are part of section four, while the conclusion is ordered in section five.
2 Literature reviewed
The authors [25] have proposed a method for glioma grading from multi-parametric MRI images. They used radiomic features along with classifiers, i.e., Support Vector Machines (SVM) and Sequential minimal optimization (SMO) for grading of glioma. They classified high-grade and low-grade glioma using their reported method in a noninvasive way. The radiomic features, they extracted include statistical, texture, relative Cerebral Blood Volume (rCBV), and Magnetic Resonance Spectroscopy (MRS) features. A dataset of a total of 40 patients (20 low- and high-grade glioma each) has been used for research and experiments. Results have been obtained with SMO and SVM classifiers and found that the linear kernel SMO algorithm demonstrated better classification performance in comparison with ‘LibLinear’ and ‘LibSVM’. Finally, the reported method achieved the performance as per evaluation measures accuracy (95.5%), sensitivity (95%), specificity (96%), and Area under the Receiver Operating Characteristic (ROC) Curve (95.5%). The researchers [11] have reported a method for glioma grade identification from MR images. They used two classifiers, i.e., Weighted Neighbor Distance Compound Hierarchy Algorithm (WNDCHRM) and a CNN-based pre-trained model called ‘VGG-19’. Experiments have been performed on a locally developed dataset containing glioma of grades. The features selected for the classification of glioma grades using WNDCHRM are Chebyshev statistics & Fourier features Radon transform features, texture features (Gabor, Haralick, and Tamura), first 4 statistical moments, multiscale histogram, object & edge statistics features, and Zernike features. With the WNDCHRM classifier (on 557 images), the reported method achieved its performance, i.e., 92.86%, 92.89%, 91.75%, 97.48%, and 92.32 against accuracy, precision, sensitivity, specificity, and F1-Score respectively. With the VGG-19 classifier, the reported method achieved performance with accuracy as 94.64%, precision of 94.71%, sensitivity of 92.72%, specificity of 98.13%, and F1-Score of 93.71. Upon increasing the no. of test images, while keeping other parameters the same, the overall achieved accuracy was 98.25%.
The investigators [18] have also suggested a noninvasive method of glioma grading using the information scanned through MRI and molecular data of the patients. Different radiomic texture features including multifractal detrended fluctuation analysis, dynamic texture analysis, and multi-resolution fractal Brownian motion have been calculated from the information obtained through structural MRI. The results have been validated against two datasets, i.e., BRATS-2017 and the cancer imaging archive (TCIA). BRATS-2017 dataset contains low-grade glioma and high-grade glioma scans, while the TCIA repository contains scans of glioblastoma multiform (GBM) and low-grade glioma. Results have been validated as per (ROC, which is 0.88 for a BraTS dataset. For the TCIA repository, classification results for both MRI and digital pathology (DP) images were 0.90 and 0.93, respectively against ROC.
The authors [3] reported a glioma grade prediction based on radiomics and scattering wavelet transform features. MICCAI-2017 dataset has been used in research and experiments. Multimodal MRI scans of 285 patients (75 low-grade gliomas and 210 high-grade gliomas) having intratumoral and peritumoral have been labeled and used in the experiments. ROC has been used to evaluate the performance of their reported method of glioma grade prediction, which is up to 0.99 for both intratumoral and peritumoral features of MRI images. The researchers [26] have reported a method for glioma grading. Dataset collected from Shandong Provincial Hospital affiliated with Shandong University containing 146 cases of glioma (49 cases of grade II, 45 cases of grade III, and 52 cases of grade IV) has been used in the experiments. They calculated both visual (based on morphology), and sub-visual (first, and second order) parameters of whole slide images. SVMs have been used as a machine learning tool with overall achieved accuracy of 90%.
The researchers [16] have proposed neural networks and Convolutional Neural Networks (CNN) based brain tumor grading method. BRATS-2014 dataset consisting of MRI scans of 213 patients has been used in the experiments. Experiments proved that CNN has more grading performance over neural networks. The authors [5] have reported an automated grading of gliomas, they used digital pathology images and CNN. They used The Cancer Genome Atlas (TCGA) for research and experiments. Their reported approach achieved the highest accuracy (96%) in the case of classification of GBM and LGG.
In a recently published study [23], classification of gliomas grades has been performed. Residual networks are used for feature representations, while Dempster-shafer Theory is used for classification purpose. Data augmentation is also utilized to avoid over-fitting. The reported method achieves 95.87% accuracy. Similarly, another recently conducted research [19] performed glioma grades classification from magnetic resonance imaging images using Gaussian CNN. Authors performed experiments on to classify three grades of glioma, i.e., Grade-II-IV. Their reported approach achieved an accuracy of 97.14%.
3 Materials and methods
In the last decades, most researchers used traditional approaches for brain tumor detection, classification, and grading, which include feature selection, extraction, reduction, and classification through these features. The main issue with these methods is their time consumed in features engineering. Further to this, these conventional methods have performance measures as low. To cope with such issues, deep-learning-based architectures have been explored. The potential of deep features motivated us in the investigation of CNNs architectures.
3.1 Dataset
Experiments have been performed on two datasets, i.e., a benchmarked dataset BraTS-2017 and a locally developed dataset from the Department of Radiology (Diagnostics), BVHB, Pakistan. The benchmarked BraTS-2017 dataset consists of MRI images of 243 (108 LGG, and 135 HGG) patients. While locally developed dataset consists of MRI slices belonging to 34 subjects, 17 each diagnosed with low- and high-grade glioma. Low-grade glioma patients include glioma of both grade-I, and grade-II, while high-grade glioma includes glioma of both grade-III and grade-IV. Each patient has undergone a biopsy test from the department of pathology, Quaid-e-Azam Medical College, Bahawalpur, Pakistan, for his/her confirmation about glioma grades. For every subject of a dataset, three MRI sequences, i.e., T1W, T2W, and FLAIR have been used). Axial and sagittal orientations of both T1W, and T2W, while the only axial orientation of FLAIR sequence has been used in the experiments. Scanning was performed on Philips Medical Systems and was based on identical scanning parameters. Anatomical scans were acquired with 3D sequences (TR = 2s, TE = 30 ms, FOV = 20 cm, 512 × 512 matrix, 25 slices of 5 mm thickness without gap). Each subject has 120 slices (25 slices of T1W Axial, T2W Axial, FLAIR Axial, T1W Sagittal, and T2W Sagittal each). Hence, 2040 slices for each of the low-, and high-grade glioma have been used for experimental purposes.
3.2 Experimental setup
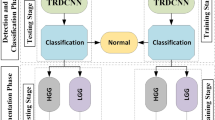
Three types of experiments, i.e., Exp. No. 01, Exp. No. 02, and Exp. No. 03 has been performed in this research, against each of the datasets. In Exp. No. 01, as it is, whole MRI slices (without skulled stripped or removing any related information) have been used for training, testing, and validation purpose. Figure 3(a) shows a sample brain MRI slice (belongs to a locally developed dataset), which has been used in this experiment. In Exp. No. 02, brain MRI slices after skulled stripped and any other related removed information) have been used for training, testing, and validation purpose. Figure 3(b) shows a sample brain MRI slice, which has been used in this experiment. In Exp. No. 03, brain MRI slices having only tumorous regions extracted through Mask R-CNN segmentation algorithm [10] have been used for training, testing, and validation purpose. Figure 3(c) shows a sample brain MRI slice, which has been used in this experiment.
3.3 Preprocessing
MRI images can be influenced by bias field distortion, which makes the intensity of identical tissues vary across an image. N4ITK method [24] has been used to correct bias field distortion present in MRI images. It is radiologically documented that the intensity distribution of a tissue type varies even if the image of the same subject is acquired using the same scanner in different time frames. Therefore, to have the intensity ranges and contrast similar across acquisitions and subjects, intensity normalization proposed by [15] has been applied to each sequence. Resultantly, the histogram of each sequence is similar across subjects. All training sample tumors have been examined to learn a clinical criterion for the glioma grading. Slices of each patient have been manually skulled stripped. From all training samples, blocks containing tumor regions have been extracted using k-mean clustering with no. of clusters equal to three.
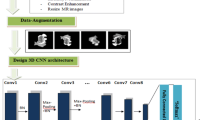
3.4 The proposed CNN architecture
In this research, to auto classify the grades of glioma, CNN has been used. Because CNN is a state-of-the-art area of machine learning inspired by the human brain. Neuroscientists discovered that the neocortex, a part of the cerebral cortex performing hearing and sight-related functions in mammals, processes the sensory signals after propagating these signals through a complex hierarchy. CNN works like the human visual system and are designed based upon the assumption that raw data consists of two-dimensional images, which enables certain properties to be encoded. So, CNNs have been used, which (with kernels) convolute on an image to have feature maps. Kernel weights connect feature map units to their previous layers. Because all units have used the same kernels, so, convolutional layers have fewer weights to train. Information about the neighborhood has been considered using kernels. The first layer has enhanced edges. The following are the important context deployed with use the of CNN to achieve the target of glioma grading.
-
1)
Initialization: This important function has been used to achieve convergence. Activations and gradients have been maintained using this function.
-
2)
Activation Function: This function has been used to transform the data in a non-linearly fashion. Rectifier linear units (ReLU) are defined in Eq. (1).
$$ f(x)=\max \left(0,\mathrm{x}\right) $$(1) -
3)
Pooling: This layer has been used to combine spatially nearby features in feature maps. To join features, it is more common to use max-pooling or average-pooling. However, in this proposal, max pooling has been used.
-
4)
Loss Function: It is a function to minimize during training. Categorical Cross-entropy represented in Eq. (2) has been used.
$$H=-{\sum}_{j{\in}voxel}{\sum}_{k{\in}classes}cj,k \ \text{log}({c}^{\prime}j,k)$$(2) -
5)
Architecture: It was the main objective that the proposed model be a reliable grading model. Since brain tumors have variability in intra-tumoral structures, hence, making grading, is a challenging task. This complexity has been reduced by designing the proposed CNN-based model tuned to the intensity normalization transformation of each tumor grade. To achieve invariance and eliminate irrelevant details, pooling has been used. However, pooling has also eliminated important details. Therefore, overlapped pooling with 2 × 2 receptive fields and 2 × 2 strides have been applied to keep information about a location. In convolutional layers explained in Eq. 3, feature maps have been padded before their convolution. Use of padding ensured feature maps of the same dimensions. The proposed CNN architecture has been depicted in Fig. 4.
$$S\left(i,j\right)=\left(I*K\right)\left(i,j\right)={S}_{m}{S}_{n}I\left(m,n\right)K(i-m,j-n)$$(3)where I is a 2D array containing segmented brain tumor and K is a kernel convolution function.
The proposed architectures for glioma grade identification are presented in Table 1.
Hyper-parameters of the proposed architectures have been shown in Table 2. These hyperparameters have been selected and tuned after several experiments (empirically).
4 Results and discussion
Classification of images containing glioma grades required several steps from preprocessing to recognition. Dataset has been randomly divided into 60%, 20%, and 20% for training, cross-validation, and testing sets, respectively. It has been ensured that slices belonging to different patients are used for each training, cross-validation, and testing purpose. The proposed CNN model has been implemented using MATLAB version 2020b, on a machine with specification, i.e., Core i9, 7th generation, 1 TB SDD (hard disk drive), and 16GB of RAM. The results obtained with the proposed method have also been compared with state-of-the-art methods. The performance plot representing accuracy and loss is shown in Fig. 5.
4.1 Results and discussions
Since, in this research work three experiments, i.e., Exp. No. 01, Exp. No. 02, and Exp. No. 03 has been performed to classify glioma grades. The results obtained on the locally developed dataset have been obtained and compiled accordingly as shown in Table 3. Table 3(a), 3(b), and 3(c) show results obtained against Exp. No. 01, Exp. No. 02, and Exp. No. 03 respectively. When Exp. N0. 01 has been performed, LGG and HGG achieved classification accuracies of 92.36% and 94.86% respectively, with an overall classification accuracy of 93.61%, as shown in Table 3(a). As these achieved measures were not so high, so, Exp. No. 02 has been performed after removing unnecessary information from MRI slices. As shown in Table 3(b), Exp. No. 02 achieved classification accuracies against LGG and HGG as 97.98% and 95.61% respectively, with an overall accuracy of 96.79%. It is evident from Table 3(a) and 3(b) that classification rates have been slightly improved, but still, achieved measures are not up to the mark. To achieve more classification rates, Exp. No. 03 has been performed after removing all other information except the tumor region. In this experiment, LGG and HGG have been classified with accuracies of 98.93%, and 98.12% respectively, while the overall accuracy achieved is 98.52%.
It is established from our previous study [7] that there is a little bit of change in the medical traits of locals, so to check the robustness of the proposed model, experiments have also been performed on a publicly available benchmarked dataset BraTS-2017. The results of these experiments are shown in Table 4.
Table 4(a), 4(b), and 4(c) show results obtained against Exp. No. 01, Exp. No. 02, and Exp. No. 03 respectively, when experiments were performed on the BraTS-2017 dataset. When Exp. N0. 01 has been performed, LGG and HGG achieved classification accuracies of 92.96% and 92.04% respectively, with an overall classification accuracy of 91.50%, as shown in Table 4(a). As these achieved measures were not so high again, so, Exp. No. 02 has been performed after removing unnecessary information from MRI slices. As shown in Table 4(b), Exp. No. 02 achieved classification accuracies against LGG and HGG as 94.08% and 93.85% respectively, with an overall accuracy of 94.97%. It is evident from Table 4(a) and 4(b) that classification rates have slightly improved, but still, achieved measures are not up to the mark. To achieve more classification rates, Exp. No. 03 has been performed after removing all other information except the tumor region. In this experiment, LGG and HGG have been classified with accuracies of 95.87%, and 96.92% respectively, while the overall accuracy achieved is 96.40%.
4.2 Comparison with state-of-the-art methods
A comparison of the results obtained through the proposed method with the results obtained from the state-of-the-art methods has been presented in Table 5. The research activity conducted by M. G. Ertosun and D. L. Rubin [5] has addressed the problem of low-grade glioma (glioma of Grade-I and Grade-II) and glioblastoma (Grade-IV) classification. Their proposed model attained an accuracy of 96.0% for low-grade glioma and 71.0% for glioblastoma, which is low in the case of glioblastoma classification rates. Another method reported by Priya, K Mohana et al. [17] classified astrocytoma grades with an overall classification accuracy of 78.26%, which is very low. Similarly, M. Monica Subashini et al. [21] have classified low-grade astrocytoma (Grade-I and Grade-II) and high-grade astrocytoma (Grade-III and Grade-IV) with overall achieved accuracy of 91.67%, which can be reasonable. Similarly in the study conducted by X. Wang et al. [26], a method of glioma grading has been reported with an overall accuracy of 90.0%, which is also a low classification rate. In the research work conducted by Q. Chen et al. [3], a method for low-grade glioma and high-grade glioma grading has been reported with an overall accuracy of 94.0%, which is reasonable.
In another recent study conducted by M. Sajjad et al. [20], grades of brain tumors have been classified, with accuracies of 95.54%, 92.66%, 87.77%, and 86.71 for brain tumors of Grade-I, Grade-II, Grade-III, and Grade-IV respectively. The overall accuracy achieved against the reported method is 90.67%. In the research work reported by A. Vamvakas et al. [25], conducted for low and high grade glioma, overall accuracy, i.e., 95.50% has been claimed, which is also appreciated. To classify low-grade glioma and glioblastoma, a recent approach reported by S. M. Reza et al. [18] has evaluated their model as per ROC, which is 0.90 (for MRI dataset) and 0.93 (for digital pathology images). To classify all grades of glioma, an overall accuracy, i.e., 98.25% has been claimed by A. M. KV and V. Rajendran, [11], which is the highest among all previously reported techniques.
The last row of Table 5 shows accuracies for each low and high grade of glioma and an overall accuracy obtained through the proposed system. The proposed system achieved 98.95% and 98.12% individual accuracies for both LGG and HGG respectively, which is high as compared to the previously reported methods. The suggested approach overwhelmed existing approaches and achieved the highest overall accuracy, i.e., 98.52%, which are the best results achieved so far as per our knowledge.
5 Conclusions
The results of the experiments for glioma as low/high grades are reasonable. In this study, a CNN-based glioma grade classification system has been proposed. A CNN-based model is fine-tuned. Results proved the effectiveness of the proposed system. The future work of this study is to enhance the proposed model to deal with all types of brain tumors for their classification and WHO grading. However, the current version of the reported model can be integrated (as an indigenous solution) into existing state-of-the-art MRI scanners almost available in every district-level hospital in Pakistan. The proposed system as a demo version is currently integrated with the image retrieval & archival system of the Radiology Department, Bahawal Victoria Hospital, Bahawalpur, Pakistan, and is helping in decision-making in a real environment.
References
Attique M et al (2012) Colorization and automated segmentation of human T2 MR brain images for characterization of soft tissues. PLoS ONE 7(3):e33616
Bartheld CS, Bahney J, Herculano-Houzel S (2016) The search for true numbers of neurons and glial cells in the human brain: a review of 150 years of cell counting. J Comp Neurol 524(18):3865–3895
Chen Q et al (2019) Glioma grade predictions using scattering wavelet transform-based radiomics. arXiv preprint arXiv:1905.09589
Chen Q et al (2020) Glioma grade prediction using wavelet scattering-based radiomics. IEEE Access 8:106564–106575
Ertosun MG, Rubin DL (2015) Automated grading of gliomas using deep learning in digital pathology images: A modular approach with ensemble of convolutional neural networks. In: AMIA Annual Symposium Proceedings. American Medical Informatics Association
Gilanie G et al (2013) Object extraction from T2 weighted brain MR image using histogram based gradient calculation. Pattern Recognit Lett 34(12):1356–1363
Gilanie G et al (2018) Classification of normal and abnormal brain MRI slices using Gabor texture and support vector machines. Signal Image Video Process 12(3):479–487
Gilanie G et al (2019) Computer aided diagnosis of brain abnormalities using texture analysis of MRI images. Int J Imaging Syst Technol 29(3):260–271
Gilanie G et al (2021) Risk-free WHO grading of astrocytoma using convolutional neural networks from MRI images. Multimed Tools Appl 80(3):4295–4306
He K et al (2017) Mask r-cnn. In: Proceedings of the IEEE international conference on computer vision
KV AM, Rajendran V (2019) Glioma tumor grade identification using artificial intelligent techniques. J Med Syst 43(5):113
Louis DN et al (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114(2):97–109
Majno G, Joris I (2004) Cells, tissues, and disease: principles of general pathology. Oxford University Press, Oxford
Mankin HJ, Lange TA, Spanier S (1982) The hazards of biopsy in patients with malignant primary bone and soft-tissue tumors. JBJS 64(8):1121–1127
Nyúl LG, Udupa JK, Zhang X (2000) New variants of a method of MRI scale standardization. IEEE Trans Med Imaging 19(2):143–150
Pan Y et al (2015) 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC). IEEE
Priya KM, Kavitha S, Bharathi B (2016) Brain tumor types and grades classification based on statistical feature set using support vector machine. In: Intelligent Systems and Control (ISCO), 10th International Conference on. 2016. IEEE
Reza SM et al (2019) Glioma grading using structural magnetic resonance imaging and molecular data. J Med Imaging 6(2):024501
Rizwan M et al (2022) Brain tumor and glioma grade classification using gaussian convolutional neural network. IEEE Access
Sajjad M et al (2019)Multi-grade brain tumor classification using deep CNN with extensive data augmentation. J Comput Sci 30:174–182
Subashini MM et al (2016) A non-invasive methodology for the grade identification of astrocytoma using image processing and artificial intelligence techniques. Expert Syst Appl 43:186–196
Tearney GJ et al (1997) In vivo endoscopic optical biopsy with optical coherence tomography. Science 276(5321):2037–2039
Tripathi PC, Bag S (2022) A computer-aided grading of glioma tumor using deep residual networks fusion. Comput Methods Programs Biomed 215:106597
Tustison NJ et al (2010) N4ITK: improved N3 bias correction. IEEE Trans Med Imaging 29(6):1310–1320
Vamvakas A et al (2019) Imaging biomarker analysis of advanced multiparametric MRI for glioma grading. Physica Med 60:188–198
Wang X et al (2019) Machine learning models for multiparametric glioma grading with quantitative result interpretations. Front Neurosci 12:1046
Weiss S, Langloss J, Enzinger F (1983) Value of S-100 protein in the diagnosis of soft tissue tumors with particular reference to benign and malignant Schwann cell tumors. Lab Invest 49(3):299–308
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gilanie, G., Bajwa, U.I., Waraich, M.M. et al. An automated and risk free WHO grading of glioma from MRI images using CNN. Multimed Tools Appl 82, 2857–2869 (2023). https://doi.org/10.1007/s11042-022-13415-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11042-022-13415-9