Abstract
Various viruses cause viral infection, and these viruses have different microscopic sizes, genetic material, and morphological forms. Due to a viral infection, the host body induces defense mechanisms that activate the innate and adaptive immune system. sncRNAs are involved in various biological processes and play an essential role in antiviral response in viruses including ZIKV, HCV, DENV, SARS-CoV, and West Nile virus, and regulate the complex interactions between the viruses and host cells. This review discusses the role of miRNAs, siRNAs, piRNAs, and tiRNAs in antiviral response. Cellular miRNAs bind with virus mRNA and perform their antiviral response in multiple viruses. However, the chemical modifications of miRNA necessary to avoid nuclease attack, which is then involved with intracellular processing, have proven challenging for therapeutic replacement of miRNAs. siRNAs have significant antiviral responses by targeting any gene of interest along the correct nucleotide of targeting mRNA. Due to this ability, siRNAs have valuable characteristics in antiviral response for therapeutic purposes. Additionally, the researchers noted the involvement of piRNAs and tiRNAs in the antiviral response, yet their findings were deemed insignificant.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The most common infections among diseases that require around 60% of medical care globally are caused by viruses [1]. Viral infections are caused by different types of viruses having various microscopic sizes (from 20 to 900 nm) and morphological forms, consisting of genetic material (can be positive or negative sense), single (ss) or double-stranded (ds), deoxyribonucleic acid (DNA) or ribonucleic acid (RNA) and are surrounded by a coating that is based on proteins, lipids or glycoproteins [2]. An extensive array of defense mechanisms is induced in the host body by viral infections activating the host body’s innate and adaptive immune systems [3]. For instance, some notable viruses and their infections include the human respiratory syncytial virus (HRSV), influenza viruses, human rhinovirus (HRV), human parainfluenza virus (HPIV), and human metapneumovirus (hMPV) are some of the most common respiratory viruses that are found in humans. HRSV causes lower respiratory tract infections (RTIs) in different age groups and is a highly contagious virus. Secondly, influenza A viruses, the causative agents of flu pandemics, have caused severe and massive pandemics with high mortality rates. Furthermore, HSV causes the upper RTIs and some lower RTIs in children, while HPIV is the pathogen that causes acute upper and lower RTIs with different levels of severity. The hMPV causes upper and lower RTIs in young children, the elderly, and immunocompromised individuals [4]. Moreover, viral infections induce non-coding RNAs, especially sncRNAs, as an immune response. The viral infections may be involved in either promoting the viral progression by manipulating the sncRNAs of the host cells or inhibiting the viral infections by different mechanisms of the host’s sncRNAs [3].
Furthermore, the host cells alter their sncRNA expression as a defense mechanism against viral infection. At the same time, viruses can evade the host defense mechanism and propagate by affecting host cellular sncRNA expression or expressing viral sncRNAs [5]. Nonetheless, the non-coding RNAs generally regulate the host’s immunities (innate and adaptive) through host-virus interactions [6]. Although host cells develop various mechanisms to inhibit viral replication, but viruses evolve corresponding immune evasion strategies to counteract the host’s immunity [7,8,9].
The host’s first line of defense against pathogens is its innate immune system, in which the innate immune response restricts viral entry, replication, and assembly. It helps to identify and remove the infected cells and accelerate the development of the adaptive immune system. The cell surface recognizes the pathogen-associated molecular patterns (PAMPs), endosomal, and cytosolic pattern recognition receptors (PRRs) of innate immune cells, which triggers the inflammatory response and programmed cell death that inhibits the viral infection [10]. The Toll-like receptors (TLRs), retinoic acid-inducible gene I (RIF-I-)-like receptors (RLRs), nucleotide-binding oligomerization domain (NOD)-like receptors (NLRs), C-type lectin receptors and absent in melanoma 2 (AIM2)-like receptors are the five primary PRR families [11].
Innate immunity, when it becomes ineffective in eliminating pathogens, aids in developing adaptive immunity. Adaptive immunity distinguishes the “self” antigens from “non-self” antigens, eliminates the specific pathogens or pathogen-infected cells by the generation of pathogen-specific immunologic effector pathways, and also develops the immunological memory that can quickly eliminate a specific pathogen [12]. The adaptive immune cells consist of the antigen-specific T cells that determine the specificity of immune response to antigens in the body [13]. Moreover, it also consists of B cells that can produce antibodies in our body [14]. These immunities (innate and adaptive) are considerably regulated by the non-coding RNAs (ncRNAs) [13].
Small non-coding RNAs and their biogenesis
The ncRNAs are generated from the non-protein-coding regions of the genome yet play a significant role in regulating gene expression and protein function Fig. 1 [13]. One of the types of non-coding RNAs is small non-coding RNAs (sncRNAs) that have a size smaller than 300 nucleotides (nt) and consist of microRNA (miRNA), Piwi RNA (piRNA), transfer RNA-derived stress-induced RNA (tiRNA) and small interfering RNA (siRNA) [15]. The sncRNAs (miRNA, piRNA, tiRNA, siRNA) play important biological roles in the regulation of cellular mechanisms such as translation inhibition, gene expression regulation, mediate gene silencing at the post-transcriptional level and RNA interference (RNAi) pathway, necessary for cell viability [16]. Furthermore, numerous human diseases have been differentially regulated by these sncRNAs, making them a new class of biomarkers for further investigation. The role of sncRNAs in disease development can help identify potential therapeutic targets and develop novel treatment methods [17,18,19,20]. Many viral infections, including hepatitis C virus (HCV), dengue virus (DENV), Zika virus (ZIKV), yellow fever virus, West Nile virus, Ebola virus, and severe acute respiratory syndrome coronavirus (SARS-CoV), are significant public health concern globally. sncRNAs are essential in regulating the complex interactions between the viruses and host cells [21].
Diversity of RNA Types and Their Functionalities in a Cell. This comprehensive diagram illustrates the various types of RNAs produced within a cell, showcasing their distinct functionalities and roles. It highlights messenger RNA (mRNA) as the template for protein synthesis, transfer RNA (tRNA) responsible for delivering amino acids to the ribosome during translation, ribosomal RNA (rRNA) as a structural component of ribosomes, small nuclear RNA (snRNA) involved in RNA splicing, small nucleolar RNA (snoRNA) participating in ribosomal RNA modification, microRNA (miRNA) regulating gene expression, long non-coding RNA (lncRNA) contributing to diverse cellular processes, small interfering RNA (siRNA) engaging in RNA interference, and other RNA species with specialized functions, providing a comprehensive overview of the functional diversity of cellular RNA molecules
Biogenesis of microRNA (miRNA)
miRNA biogenesis is a two-step process with nuclear and cytoplasmic cleavage. The primary miRNA (pri-miRNA) is produced by transcribing the miRNA gene in the nucleus. The Drosha RNase III endonuclease cleaves the pri-miRNA, called the precursor miRNA (pre-miRNA) Fig. 2. The RNase III endonuclease Dicer matures pre-miRNA in the cytoplasm by recognizing the 5′ phosphate and 3′ overhang and cutting the double strand. The cleavage separates the loop structure and forms a miRNA: miRNA* complex, which ultimately releases the mature miRNA [22].
Intronic and Intergenic Pathways of miRNA-mRNA Processing and Targeting in a Cell. The diagram depicts the intricate pathways involved in the processing and targeting of microRNAs (miRNAs) to messenger RNA (mRNA) molecules within a cell. It highlights both the intronic and intergenic mechanisms through which miRNAs are produced and subsequently bind to their target mRNA sequences, leading to post-transcriptional regulation of gene expression
Biogenesis of Piwi RNA (piRNA)
In the biogenesis of piRNA, many piRNA precursors are produced from the piRNA clusters (genetic regions), which are divided into uni-strand and dual-strand clusters Fig. 1. As the transcription is performed, piRNA precursors are transported out of the nucleus. The RNA helicase Armitage (Armi) enzyme in the cytoplasm resolves the secondary structure of piRNA precursors and then further processes and produces primary piRNAs. The Ago3 and Aub proteins then amplify the generation of primary piRNAs, called the secondary biogenesis of piRNAs and the ping-pong cycle. The PIWI proteins are then bound with piRNAs and transported back to the nucleus to silence the target genes [23].
Biogenesis of transfer RNA-derived stress-induced RNA (tiRNA)
The biogenesis of tiRNA involves the cleavage of mature tRNA in the middle of the anticodon loop by angiogenin (ANG). Most often, the stress stimulations such as heat shock, cold shock, hypoxia, and oxidative stress trigger the production of tiRNA [24, 25]. The tiRNA is divided into two subclasses (5’ tiRNA and 3’ tiRNA) based on including 5’ sequences and 3’ sequences [26]. The 5’ tiRNA consists of the portion from the anticodon loop cleavage site to the mature 5’ end of the tRNA, while the 3’ tiRNA refers to the anticodon loop cleavage position to the 3’ end. 5’ tiRNAs also have the potential as circulating biomarkers [27].
Biogenesis of Small interfering RNA (siRNA)
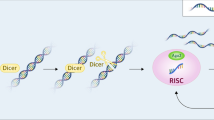
The siRNA biogenesis involves the canonical inducer of RNA interference (RNAi) that is long, linear, perfectly base-paired double-stranded RNA (dsRNA). When directly introduced into the cytoplasm or taken up from the environment, these (dsRNAs) are processed by Dicer (a member of the RNase III family) and produce siRNA Fig. 3. The siRNA duplex interacts with Argonaute 2 when incorporated into the RISC complex, resulting in the unwinding and degradation of the passenger strand, and the guide strand guides the RISC complex to the mRNA [28].
sncRNAs significance in immune response in viral immune response
sncRNAs are integral in a diverse range of biological processes. sncRNAs regulate the host-virus interaction, viral replication, and host responses to viral infection. The microRNAs (miRNAs), Piwi-Interacting RNAs (piRNAs), tRNA-derived RNA fragments (tRFs), and small interference RNAs (siRNAs) are the highly significant regulators of host-virus interaction [29]. Throughout evolution, multiple layers of antiviral mechanisms have been developed in the host cells against viruses. Cellular and humoral immunity, based on T, B, and other immune cells, play an essential role in mammals. Host immunity can be categorized as innate and adaptive immunity. Innate immunity establishes the first layer of immune responses and uses chemical, physical, molecular, and cellular mechanisms against viruses. Innate immunity uses the interferon I, II, and III pathways to defend host cells against viruses. In contrast, adaptive immunity develops antibodies and long-term memory primarily involving T and B immune cells [11].
miRNAs
miRNAs are essential in cellular differentiation, proliferation, development, and apoptosis. miRNAs have regulated viral infection and identified their targets and mode of action [30]. Furthermore, recent research has revealed that miRNA exhibits binding specificity to particular sequences within the 3’ untranslated region (UTR) of their target mRNAs, leading to the initiation of translational repression, mRNA deadenylating, and decapping. Moreover, the miRNA binding sites have been identified in different mRNA regions, such as the 5’ UTR and coding sequence; notably, miRNA interaction with the 5’ UTR abd coding sequence of mRNAs results in suppressive effects on gene expression [31]. In viral infection, cellular miRNAs bind with virus mRNA and change host mRNA’s stability, reducing the viral genome’s translation and the level of free miRNAs altered in cells [32]. This effect stabilized the viral transcripts and changed the genome expression of the host cell [33]. In the immune system, they act as gene expression regulators. They discharge the various inflammatory mediators and control antibody production. The abnormal expression and functioning of miRNA produced multiple diseases like inflammatory disorders, cancers, and allergic diseases in the immune system related to or initiated by viruses. They regulate the responses associated with both acquired and innate immune systems in humans [34].
piRNAs
piRNAs form a piRNA/piwi complex when bound with piwi proteins and influence transposon silencing, spermiogenesis, genome rearrangement, epigenetic regulation, protein regulation, and germ stem-cell maintenance. piRNAs mainly originate from loci that act as transposon traps and function as RNA-mediated adaptive immunity against transposon expansions and invasions. piRNAs are also involved in some somatic tissues, such as innate immunity regulation in the lungs [35]. Nevertheless, the piRNA/piwi complex employs base-pairing to recognize the target RNAs, subsequently recruiting co-factors to establish either transcriptional gene silencing (TGS) or inducing post-transcriptional gene silencing (PTGS) [36]. Most researchers have observed that piRNAs act as antiviral in different viral infections. However, 52 piRNAs have been observed in RSV exosomes, with 28 upregulated and 24 downregulated piRNAs through next-generation sequencing [37]. Surprisingly, in the context of viral infection, the function of piRNA has yet to be studied in detail.
siRNAs
siRNAs regulate gene expression through a phenomenon known as RNA interference (RNAi). siRNAs, a type of RNA, play a crucial role in cellular defense against viral infections by targeting and cleaving specific messenger RNAs (mRNAs), preventing protein translation, and reducing antiviral gene expression by degrading mRNAs encoding antiviral proteins, thereby suppressing the host’s immune response against the virus. siRNAs disrupt viral replication and transcription processes, inhibiting the synthesis and assembly of viral particles. They also modulate the host cell’s response to viral infection by targeting host factors, ultimately interfering with the virus-host interaction [38,39,40]. Moreover, the RNAi role in defense mechanisms still remains controversial [40].
tiRNAs
The tiRNAs control the transcription of protein-coding genes by targeting epigenetic silencing complexes. In the bloodstream, 5′-tiRNAs circulate and act as extracellular miRNAs in response to immune reactions and viral infections. tiRNAs also produce the stress-related granules of independent assembly, translating messenger ribonucleoproteins (mRNPs) that can temporarily silence mRNAs, helping cells survive. They involve many cellular and molecular mechanisms associated with stress responses and diseases such as cancer, and metabolic, and immune disorders [41]. In virus infection, ANG-induced tRNA halves have been used for cellular responses. Researchers identified that the introduction of tRNA halves was virus-specific, cleaved the subset of tRNAs, and cleavage occurred around anticodon of the tRNAs that showed 5′-tRNA half derived from tRNAGlu-CTC repressed the target mRNA in the cytoplasm and promotes replication of viruses in cells [42]. However, the tiRNA products can be produced by ANG when secreted by stressed cells due to paracrine signaling. The ANG usually elicits prosurvival signaling, and many resulting tiRNAs inhibit apoptosis and degrade mRNAs by facilitating the cellular response to stress by reprogramming translation [43].
Methodology
A comprehensive literature search was conducted to identify the relevant articles for this review. Different search terms were used for the selection of articles (viral immune response, significance of viral infections, human immune response, human viral infections, innate immune response, adaptive immune response, ncRNAs, relation of immunity with ncRNAs, small ncRNAs, role of sncRNAs in immunity in humans, biogenesis of sncRNAs in humans, biogenesis of miRNA, piRNA, tiRNA, and siRNA in humans, role of sncRNAs in viral immune response). Additional studies were also identified based on the referenced articles of the relevant articles. The articles that were included in this review had to meet specific criteria, such as the articles that were selected were published between 2017 and 2023, the role of sncRNAs (specifically miRNA, piRNA, tiRNA, and siRNA) in viral immune response, no conference papers were added in this review, and this review specifically focused on the viral immune response in humans. The Google Scholar and PubMed literature search engines were utilized.
This methodology was developed to establish an extensive search for relevant literature and select articles to include in this review systematically. The articles included in this review were of high quality based on the set criteria and provided a significant perception of the role of sncRNAs in human viral immune response. Author 1 actively contributed to writing the manuscript, explicitly addressing the functions and mechanisms of non-coding RNAs. They provided critical revisions to ensure content accuracy and clarity. Author 1 approved the article’s final version, taking responsibility for its integrity and accuracy. Author 2 made substantial contributions by providing expertise in experimental techniques and methodologies used in non-coding RNA research. They assisted in data analysis and interpretation, specifically focusing on experimental approaches. Author 2 critically reviewed the manuscript, offering valuable feedback to enhance scientific rigor and clarity. They approved the final version, ensuring accuracy and integrity. Author 3 contributed significantly by offering expertise in molecular biology and genomics, analyzing research trends and advancements in non-coding RNA studies. They actively participated in writing, specifically focusing on emerging applications and prospects. Author 3 provided critical review and intellectual input to ensure scientific soundness. They approved the final version, taking responsibility for overall accuracy and integrity.
Results
Role of miRNA in viral immune response
Host miRNAs play a role as proviral and antiviral Table 1. The interaction between viral and host miRNAs enables viral replication and infection, thus, proviral function. Proviral miRNAs elevate the viral infection and allow the virus to evade the host’s immune response by suppressing antiviral factors such as interferon (IFN) [44]. Moreover, host miRNAs also act as antiviral functions by regulating viral RNA production, stopping viral replication, and suppressing and entering a virus to a latent phase [45]. Furthermore, it is reported in several studies that miRNAs are involved in crucial mechanisms of cell biology, regulation of the immune system, and the onset and progression of numerous types of disorders, including viral infections [46].
The innate immune system employs miRNAs to target the NF-κB signaling pathway, a crucial regulator of innate immune responses Fig. 4. This pathway is activated by the MyD88 and TRIF-dependent pathways, triggering pro-inflammatory cytokines and interferon-stimulated genes essential for responding to pathogens and damaged cells [47]. In contrast, miRNAs also play a role in adaptive immune responses, influencing antigen-specific T-cell differentiation and B-cell development and maturation [48]. Notable miRNAs that recognize pathogens include miR-17-92, which enhances T-cell survival and suppresses pro-apoptotic proteins, critical in thymopoiesis. miR-17-92 also plays a vital role in developing T follicular helper cells, which are crucial for long-lasting antibody responses [47, 49].
Another example is miR-155, which intensifies both innate and adaptive immune responses. It inhibits replication in viruses like HBC, promotes viral latency, and is highly involved in HCV and HIV infections [50]. FOXP3, a transcription factor essential for T regulatory cells, regulates miR-155, which is crucial for T cell formation and function. In the mammalian immune system, miR-155 contributes to an optimal T-cell-independent antibody response [51]. MiR-150 has been investigated in reproductive and respiratory syndrome (PRRSV) infection, where its up-regulation suppresses PRRSV and activates IFN signaling [52]. MiR-223 plays a regulatory role in immune responses, modulating the activation of vital immune cells like neutrophils, macrophages, and dendritic cells. Its abnormal expression is associated with infectious diseases such as HIV-1, viral hepatitis, and tuberculosis, impacting macrophage function, neutrophil infiltration, dendritic cell maturation, and inflammasome activation [53].
Additionally, miR-146 regulates dendritic cell survival and inflammatory activation and is a negative regulator in immune pathways. It is implicated in viral infectious diseases and antiviral innate immune responses [54]. The let-7 family of miRNAs reduces the expression levels of specific target genes, bolstering the human immune system’s antiviral response. Let-7 may serve as a potential diagnostic marker due to its distinct expression in viral-infected patients compared to healthy individuals. Let-7f controls genes pivotal in host inflammatory responses, as observed in a study on A549 human alveolar epithelial cells infected with recombinant respiratory syncytial virus (RSV) [55]. Furthermore, viral-encoded miRNAs from pathogens like SV40 and KSHV contribute to immune evasion by regulating viral gene expression. The KSHV viral miRNA, for instance, regulates cytokine signaling by reducing the C/EBPβ p20 (LIP) expression, a negative regulator of IL6 and IL10 cytokines.
Additionally, miR-K12-9 directly targets IκB and regulates the NF-κB pathway [56]. These viral miRNAs modulate host cell signaling depending on the type of cell infected, crucial for controlling lytic and latent infections [57]. SARS-CoV-2’s miRNAs indirectly or directly coordinate tumor necrosis factor (TNF) and chemokine signaling immune pathways. SARS-CoV-2’s miRNAs modulate immune-inflammatory responses and host infection, highlighting specific interactions with host genes and proteins [58].
The viral miRNA shows the involvement in regulating different signaling pathways for proliferation, differentiation, and tumorigenesis. The major signaling hubs highlighted are Akt signaling in blue, MAPK signaling in dark blue, Wnt signaling in pink, and TGFβ signaling in brown), whereas the viral miRNAs are shown in red
Role of siRNAs in viral immune response
The short interfering RNAs (siRNAs) are crucial in regulating gene expression by RNA silencing. At the same time, its biogenesis and function involve the activity of a conserved group of enzymes [59]. The short interfering RNA (siRNA) stimulates the immune responses by pathogen recognition receptors Table 1. Toll-like receptor 3 (TLR3) is a subcellular component of target cells on the cell surface. In the endosomes and lysosomes of specialized immune cells, TLR7 and TLR8 are present and identified siRNA. Retinoic acid-inducible gene 1 protein (RIG-I) and a dsRNA-dependent protein kinase (PKR) detect and react with siRNAs. Each receptor released a unique type of signaling for immune response, increasing mRNA transcription. This mRNA produces Type I interferons and inflammatory cytokines [60]. In vitro and in vivo antiviral activity of siRNAs against viruses can be studied, and the first potential antiviral siRNAs studied in respiratory syncytial virus (RSV). ALN-RSV01 was the first siRNA that is 19 bp RNA duplex and two thymidine attached on both 3’ ends, which prevents its nuclease degradation. In RSV nucleocapsid protein, ALN-RSV01 targets a highly conserved region of mRNA.
Furthermore, it reduced the risk of bronchiolitis obliterans syndrome in double-blind, placebo-controlled trials in RSV-infected lung transplant patients [61]. siRNAs have been used against Zaire ebolavirus (ZEBOV). They protected the rhesus macaques from lethal infection by preventing the synthesis of Zaire ebolavirus polymerase, viral proteins 35 and 24. The modified siRNA, TKM-Ebola, was used to treat humans during the ZEBOV outbreak in West Africa in 2013–2016 [62]. ARC-520 and ARC-521 are two anti-HBV siRNAs that are used in the treatment of Hepatitis B Virus. In clinical trials, patients respond less to ARC-520 due to losing the target site when the HBV genome enters the host DNA. ARC-520 and ARC-521 have some drawbacks, so the following formulation, ARO-HBV, has been proposed, with significant antiviral activity [63]. LNP-siRNA can be studied in SARS-CoV-2 infections that are highly effective in mice. Multiple siRNAs have been screened against SARS-CoV-2, but only siHel1, siHel2, UC7, and siUTR3 siRNAs inhibit the virus either in combination with one another or alone. Scalable LNP-siRNA methods can treat human SARS-CoV-2 infections as soon as they are obvious [64].
Role of piRNA
piRNAs play a crucial role in maintaining the complexity and integrity of the genome by suppressing the mutations caused by transposable elements. However, it is also involved in regulating gene expression in somatic cells. However, very little is known about the possible functions in response to viral infections [37]. The role of piRNA in the antiviral defense is shown in Table 1. piRNAs bind with PIWI proteins and silence the target genes by producing an RNA-induced silencing complex (RISC). piRNA produces Immune responses when infection occurs with an exogenous virus or bacteria. In HIV-1, PIWIL2 activates CD4 + T cells to inhibit HIV-1 replication in anti-CD3/anti-CD28, increasing viral proteins by decreasing PIWIL2 [65]. piRNA-hsa-28,382 and piRNA-hsa-28,190 have been involved in the regulation of Herpes Simplex Virus type-1 (HSV-1) replication. Upregulated piRNA-hsa-28,382 has played an essential role in HSV-1 infection and performed antivirus proliferation by silencing the virus genes expression, but piRNA-hsa-28,190 regulates the HSV-1 infection [66]. Endogenous piRNA and synthetic piRNAs (spiRNAs) have been identified in the gRNA of SARS-CoV-2, which suppressed the multiplication of SARS-CoV-2. These spiRNAs and endogenous piRNA have fewer side effects on human 17,494 protein-coding genes, and all strains of SARS-CoV-2 can be controlled by them [67]. piR-57,640 targeted the STATS, which is required in RSV infection, and this virus-induced the type 1 interferon from airway epithelial infected cells. RSV repressed the expression of human STAT2 proteins and escaped the type 1 interferon response of the host [45]. piR-4095 targets the DDX4 and different DDX enzymes involved in the coronavirus replication [68].
Role of tiRNAs
The tiRNA regulates translational procedures under stress stimulation instead of decreasing mature tRNA levels of encoding mRNAs [69]. The tiRNAs play a role in NF-κB, IFN, and Wnt/β-Catenin pathways, which recommend they are also involved in the immune responses Table 1 [42]. Toll-like receptors (TLR) stimulate the membrane, activate the NF-kB pathway, and regulate the overexpression of ANG. Anticodon, loops of tRNAs, split by ANG and produce 5’ tiRNAs that emit the signaling molecules in the form of EVs (extracellular vesicles). In targeted cells, complexes of EV-5′-tiRNAs send into endosomes and activate the TLR-7, which assist the immune response [70]. Activation of the JAK/STAT pathway regulates the interferon expression. In viral infection, interferon expression reduces the expression of ANG gene, and tiRNAs change into antiviral tiRNAs production [71]. After activating ANG, RSV infection causes epithelial cells to produce tRNA fragments that resemble 5 abundantly’ -tiRNAs. At the post-transcriptional level, tRF5-GluCTC represses the expression of luciferase mRNA, downregulates RSV replication, and decreases the yield of RSV particles produced by infected cells [72].
Discussion
Viruses’ remarkable adaptability enables them to thrive and persist within their hosts’ intricate physiological and immune environments. Organisms can respond to environmental changes by undergoing genetic modifications during replication [73]. On the contrary, the conventional genome encompasses the entire proteins necessary for virus replication. The viral variants exhibit random mutations that may augment the virus’s capacity to adapt to novel circumstances [74]. One of the most notable accomplishments in modern medicine is the development of highly effective vaccines that specifically target viral pathogens.
Nevertheless, there is a lack of available vaccines for numerous hazardous viruses, including the Hepatitis C virus (HCV), Zika virus, and Human Immunodeficiency Virus (HIV). Small non-coding RNAs (sncRNAs), such as microRNAs (miRNAs), piwi-interacting RNAs (piRNAs), transfer RNA-derived fragments (tiRNAs), and small interfering RNAs (siRNAs), have been identified as critical regulators in host-virus interactions [51].
Researchers have found that miRNAs, piRNAs, siRNAs, and tiRNAs are highly involved in antiviral response. Mammalian vsiRNAs challenge conventional antiviral defenses. These 22-nucleotide RNAs, cleaved by Dicer, operate independently of the IFN signaling pathway. They efficiently target and degrade invading viruses, emphasizing the potency of RNA interference (RNAi) in innate immunity [75]. WNV, dengue, Zika, SARS-CoV, and SARS-CoV-2 all contain sncRNAs, including virus-encoded miRNAs (v-miRNAs). They regulate viral replication and immune signaling. KUN-miR-1 in WNV and DENV-vsRNA-5 in DENV influence viral replication at different infection stages of SARS-CoV-2, and v-miR-147-3 affects host factors. miRNA encoding potential has been suggested for ssRNA (−) viruses such as Filoviruses (e.g., Ebola virus) and Orthomyxoviruses (e.g., Influenza A viruses). H5N1 (influenza A virus) produces miR-HA-3p, which inhibits immune response-associated host factors. BLV-miR-B4-3p, a functional v-miRNA, influences cell proliferation and tumorigenesis. While HIV-1-encoded miRNAs are controversial, foamy viruses like Bovine and Simian Foamy Virus encode v-miRNAs essential for viral replication. These findings suggest that sncRNAs may help combat viruses, especially without vaccines [76].
microRNAs (miRNAs) are pivotal in the host’s antiviral response. Notably, miR-221/-222 demonstrates increased expression in response to TNFα, effectively targeting CD4 in HIV-1 and impeding viral entry [77]. In the DENV, miR-3614 levels rise upon viral infection, subsequently targeting host ADAR1 and reducing virus infectivity [78]. Additionally, miR-323 suppresses viral replication by targeting viral PB1 RNA, with its expression heightened post-IAV infection [79]. The downregulation of miR-340 following viral infection leads to increased antiviral immunity in IAV infection, as it targets host RIG-1 and OAS2 [80]. Moreover, miR-1/-30/-128/-196-296/-351/-431/-448 are potent regulators of HCV replication, with their expression induced by IFN signaling [81]. miR-122, downregulated by IFN signaling, targets the 5’ non-coding region of the viral genome, effectively suppressing viral genome replication [82].
The precise role of piRNAs in viral infection still needs to be explored. Target prediction analysis indicates that piR-38,587 targets the AP2 complex in ocular infections following human adenovirus species D type 37 (HAdV-D37) infection [83]. However, a comprehensive understanding of piRNAs in viral infections necessitates further investigation.
siRNAs have shown distinctive therapeutic applications among small non-coding RNAs. The effectiveness of these molecules stems from their ability to achieve complete Watson-Crick base pairing with target mRNA, offering a broad therapeutic scope, particularly for genes that are challenging to target using other techniques [84]. siRNAs can specifically target any potential gene, thus making them a valuable tool in antiviral therapy. Hereditary genetic diseases such as hATTR amyloidosis, Huntington’s disease, and muscular dystrophy are considered prominent applications. Phase 3 clinical trials are underway for siRNA-based drugs that treat cardiovascular diseases, explicitly targeting genes such as ApoB and PCSK9. Moreover, siRNA demonstrates regulation in glucose homeostasis and prevention of renal fibrosis, extending to cancer treatment, with ongoing research targeting various cancer types. siRNA-based and gene therapy are developing rapidly as an antiviral strategy against viruses such as SARS-CoV and HBV and are under exploration for diseases such as respiratory distress, neurological disorders, ocular diseases, and inflammatory conditions [85].
Despite our knowledge of miRNAs and siRNAs, tiRNAs’ role in viral infection still needs to be discovered. Pathological conditions involving tRNA fragments include cancer and neurodegenerative disorders. Crucial mutations in the ANG gene for tiRNA production are associated with neurodegenerative diseases. Diverse tRNA fragments regulate cancer cell proliferation and DNA processes. Glu-derived tiRNAs stimulate RSV replication in viral infections, indicating their role in host-virus interactions. A recent study revealed that tRF5-GluCTC targets APOER2 protein, revealing a novel antiviral defense mechanism [86]. Understanding the role of tiRNAs in immune responses to viral infections may lead to new antiviral therapies [84].
Although siRNAs show potential for therapeutic applications, miRNAs present specific difficulties because they are susceptible to degradation by nucleases. Chemical modification of miRNAs is required to improve their stability, which introduces additional intricacy to their intracellular processing. Advancements in nucleic acid medicine delivery have improved treatment outcomes. Synthetic surfactants like argininocalix[4]arene 1 and tetraargininocalix[4]arene [1] efficiently transfer miRNAs and anti-miRNA molecules to target cells. These carriers show high transfection efficiency and low cytotoxicity. Neuromag, a polymeric magnetic particle, is promising for delivering nucleic acids to pyramidal cells in the rat visual cortex [87]. These recent advancements in lipid nanoparticles and nucleic acid medicine delivery have addressed these challenges, offering therapeutics based on small non-coding RNAs (sncRNAs) [81].
Conclusion
A comprehensive literature search and systematic approach have unveiled the significant roles of miRNAs, piRNAs, siRNAs, and tiRNAs in the host’s defense mechanisms. MicroRNAs (miRNAs) significantly regulate viral infections, displaying both pro-viral and antiviral activities. They modulate NF-κB signaling in the innate immune system and have a significant role in antigen-specific T-cell differentiation and B-cell development in the adaptive immune response. The let-7 miRNAs have shown potential as diagnostic markers due to their distinct expression patterns in individuals infected with viruses. miRNAs also provide targeted regulatory functions against HIV-1, DENV, IAV, and HCV, suggesting therapeutic potential. siRNAs can potentially target viral pathogens and enhance our antiviral defense mechanism. Their precise base pairing with target mRNA offers a potent method for targeting a wide range of genes, including those that are challenging to target using alternative methods. Initial research suggests that PIWI-interacting RNAs (piRNAs) may play a role in antiviral defenses, particularly ocular infections. Though less explored, piRNAs have been found to play a crucial role in inhibiting viral replication and immune signaling during infections. tiRNAs, due to their regulatory function in translational processes, present an intriguing subject for future research. Nucleic acid medicine delivery to specific tissues is still tricky, but recent advances offer hope. As researchers continue investigating the complex relationship between sncRNAs and viral infections, new antiviral therapeutic approaches may be developed, including those without vaccines, providing hope for more effective viral pathogen treatments.
Data availability
All data in this manuscript are available.
References
Dronina J, Samukaite-Bubniene U, Ramanavicius A (2021) Advances and insights in the diagnosis of viral Infections. J Nanobiotechnol 19(1):348
Rojek A, Horby P, Dunning J (2017) Insights from clinical research completed during the West Africa Ebola virus Disease epidemic. Lancet Infect Dis 17(9):e280–e292
Klimenko OV (2022) Perspectives on the use of small noncoding RNAs as a therapy for severe Virus-Induced Disease manifestations and late Complications. Bionanoscience 12(3):994–1001
Weston S, Frieman MB (2019) Respiratory viruses. Encycl Microbiol. 85–101
Nunes A, Ribeiro DR, Marques M, Santos MAS, Ribeiro D, Soares AR (2020) Emerging roles of tRNAs in RNA virus Infections. Trends Biochem Sci 45(9):794–805
Sajjad N, Wang S, Liu P, Chen JL, Chi X, Liu S et al (2021) Functional Roles of Non-coding RNAs in the Interaction Between Host and Influenza A Virus. Front Microbiol [Internet]. [cited 2023 Oct 17];12. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fmicb.2021.742984
Albanese M, Tagawa T, Hammerschmidt W (2022) Strategies of Epstein-Barr virus to evade innate antiviral immunity of its human host. Front Microbiol 13:955603
Nelemans T, Kikkert M (2019) Viral innate Immune Evasion and the pathogenesis of emerging RNA virus Infections. Viruses 11(10):961
Wang C, Wang T, Duan L, Chen H, Hu R, Wang X et al Evasion of Host Antiviral Innate Immunity by Paramyxovirus Accessory Proteins. Front Microbiol [Internet]. 2022 [cited 2023 Oct 17];12. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fmicb.2021.790191
Diamond MS, Kanneganti TD (2022) Innate immunity: the first line of defense against SARS-CoV-2. Nat Immunol 23(2):165–176
Kanneganti TD (2020) Intracellular innate immune receptors: life inside the cell. Immunol Rev 297(1):5–12
Marshall JS, Warrington R, Watson W, Kim HL (2018) An introduction to immunology and immunopathology. Allergy Asthma Clin Immunol 14(2):49
Zhang L, Xu X, Su X (2020) Noncoding RNAs in cancer immunity: functions, regulatory mechanisms, and clinical application. Mol Cancer 19(1):48
Tsai DY, Hung KH, Chang CW, Lin KI (2019) Regulatory mechanisms of B cell responses and the implication in B cell-related Diseases. J Biomed Sci 26(1):64
Zhang Z, Zhang J, Diao L, Han L (2021) Small non-coding RNAs in human cancer: function, clinical utility, and characterization. Oncogene 40(9):1570–1577
Zhang P, Wu W, Chen Q, Chen M Non-Coding RNAs and their Integrated Networks. J Integr Bioinforma [Internet]. 2019 Sep 1 [cited 2023 Oct 17];16(3). Available from: https://www.degruyter.com/document/doi/https://doi.org/10.1515/jib-2019-0027/html
Watson CN, Belli A, Di Pietro V (2019) Small Non-coding RNAs: New Class of Biomarkers and Potential Therapeutic Targets in Neurodegenerative Disease. Front Genet [Internet]. [cited 2023 Oct 17];10. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fgene.2019.00364
Zhu L, Liu X, Pu W, Peng Y (2018) tRNA-derived small non-coding RNAs in human Disease. Cancer Lett 419:1–7
Poller W, Dimmeler S, Heymans S, Zeller T, Haas J, Karakas M et al (2018) Non-coding RNAs in Cardiovascular Diseases: diagnostic and therapeutic perspectives. Eur Heart J 39(29):2704–2716
Romano G, Veneziano D, Acunzo M, Croce CM (2017) Small non-coding RNA and cancer. Carcinogenesis 38(5):485–491
Bhatti GK, Khullar N, Sidhu IS, Navik US, Reddy AP, Reddy PH et al (2021) Emerging role of non-coding RNA in health and Disease. Metab Brain Dis 36(6):1119–1134
MiRNA Biogenesis and Regulation of Diseases : An Overview | SpringerLink [Internet]. 2023 [cited 2023 Oct 17]. Available from: https://springerlink.bibliotecabuap.elogim.com/protocol/10.1007/978-1-4939-6524-3_1
Wu X, Pan Y, Fang Y, Zhang J, Xie M, Yang F et al (2020) The Biogenesis and functions of piRNAs in Human Diseases. Mol Ther Nucleic Acids 21:108–120
Elkordy A, Mishima E, Niizuma K, Akiyama Y, Fujimura M, Tominaga T et al (2018) Stress-induced tRNA cleavage and tiRNA generation in rat neuronal PC12 cells. J Neurochem 146(5):560–569
Tao EW, Wang HL, Cheng WY, Liu QQ, Chen YX, Gao QY (2021) A specific tRNA half, 5’tiRNA-His-GTG, responds to hypoxia via the HIF1α/ANG axis and promotes Colorectal cancer progression by regulating LATS2. J Exp Clin Cancer Res 40(1):67
Tao EW, Cheng WY, Li WL, Yu J, Gao QY, tiRNAs: (2020) A novel class of small noncoding RNAs that helps cells respond to stressors and plays roles in cancer progression. J Cell Physiol 235(2):683–690
Zhu L, Li T, Shen Y, Yu X, Xiao B, Guo J (2019) Using tRNA halves as novel biomarkers for the diagnosis of gastric cancer. Cancer Biomark Sect Dis Markers 25(2):169–176
Alshaer W, Zureigat H, Al Karaki A, Al-Kadash A, Gharaibeh L, Hatmal MM et al (2021) siRNA: mechanism of action, challenges, and therapeutic approaches. Eur J Pharmacol 905:174178
Changes of small non-coding RNAs by severe acute respiratory syndrome coronavirus 2 infection | bioRxiv [Internet]. 2023 [cited 2023 Oct 17]. Available from: https://www.biorxiv.org/content/https://doi.org/10.1101/2021.12.16.472982v1
Girardi E, López P, Pfeffer S (2018) On the importance of host MicroRNAs during viral Infection. Front Genet 9:439
Rani V, Sengar RS (2022) Biogenesis and mechanisms of microRNA-mediated gene regulation. Biotechnol Bioeng 119(3):685–692
Abu-Izneid T, AlHajri N, Ibrahim AM, Javed MN, Salem KM, Pottoo FH et al (2021) Micro-RNAs in the regulation of immune response against SARS CoV-2 and other viral Infections. J Adv Res 30:133–145
Viral long non-coding RNA regulates virus life-cycle and pathogenicity - PMC [Internet]. [cited 2023 Oct 17]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC8929458/
Chandan K, Gupta M, Sarwat M (2019) Role of host and Pathogen-derived MicroRNAs in Immune Regulation during Infectious and Inflammatory Diseases. Front Immunol 10:3081
Wasserman GA, Szymaniak AD, Hinds AC, Yamamoto K, Kamata H, Smith NM et al (2017) Expression of Piwi protein MIWI2 defines a distinct population of multiciliated cells. J Clin Invest 127(10):3866–3876
Loubalova Z, Konstantinidou P, Haase AD (2023) Themes and variations on piRNA-guided transposon control. Mob DNA 14(1):10
Corsello T, Kudlicki AS, Liu T, Casola A (2022) Respiratory syncytial virus infection changes the piwi-interacting RNA content of airway epithelial cells. Front Mol Biosci [Internet]. [cited 2023 Oct 17];9. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fmolb.2022.931354
Qureshi A, Tantray VG, Kirmani AR, Ahangar AG (2018) A review on current status of antiviral siRNA. Rev Med Virol 28(4):e1976
Ketzinel-Gilad M, Shaul Y, Galun E (2006) RNA interference for antiviral therapy. J Gene Med 8(8):933–950
Mehta A, Michler T, Merkel OM (2021) siRNA therapeutics against respiratory viral infections-what have we learned for potential COVID-19 therapies? Adv Healthc Mater 10(7):e2001650
Sarais F, Perdomo-Sabogal A, Wimmers K, Ponsuksili S, tiRNAs (2022) Insights into their Biogenesis, functions, and future applications in Livestock Research. Non-Coding RNA 8(3):37
Pandey KK, Madhry D, Ravi Kumar YS, Malvankar S, Sapra L, Srivastava RK et al (2021) Regulatory roles of tRNA-derived RNA fragments in human pathophysiology. Mol Ther Nucleic Acids 26:161–173
Menegazzi M, Gotte G (2022) Role of the Ribonuclease ONCONASE in miRNA Biogenesis and tRNA Processing: Focus on Cancer and viral Infections. Int J Mol Sci 23(12):6556
Bruscella P, Bottini S, Baudesson C, Pawlotsky JM, Feray C, Trabucchi M (2017) Viruses and miRNAs: More Friends than Foes. Front Microbiol [Internet]. [cited 2023 Oct 17];8. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fmicb.2017.00824
Barbu MG, Condrat CE, Thompson DC, Bugnar OL, Cretoiu D, Toader OD et al (2020) MicroRNA involvement in Signaling pathways during viral Infection. Front Cell Dev Biol 8:143
Mirzaei R, Mahdavi F, Badrzadeh F, Hosseini-Fard SR, Heidary M, Jeda AS et al (2021) The emerging role of microRNAs in the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Infection. Int Immunopharmacol 90:107204
Nejad C, Stunden HJ, Gantier MP (2018) A guide to miRNAs in inflammation and innate immune responses. FEBS J 285(20):3695–3716
Tahamtan A, Teymoori-Rad M, Nakstad B, Salimi V (2018) Anti-inflammatory MicroRNAs and their potential for inflammatory Diseases treatment. Front Immunol 9:1377
Formosa A, Turgeon P, Dos Santos CC (2022) Role of miRNA dysregulation in sepsis. Mol Med Camb Mass 28(1):99
Jafarzadeh A, Naseri A, Shojaie L, Nemati M, Jafarzadeh S, Bannazadeh Baghi H et al (2021) MicroRNA-155 and antiviral immune responses. Int Immunopharmacol 101:108188 Pt A)
Dickey LL, Hanley TM, Huffaker TB, Ramstead AG, O’Connell RM, Lane TE (2017) MicroRNA 155 and Viral-Induced Neuroinflammation. J Neuroimmunol 308:17–24
Li S, Zhang X, Yao Y, Zhu Y, Zheng X, Liu F et al (2022) Inducible miR-150 inhibits Porcine Reproductive and Respiratory Syndrome Virus replication by targeting viral genome and suppressor of Cytokine Signaling 1. Viruses 14(7):1485
Yuan S, Wu Q, Wang Z, Che Y, Zheng S, Chen Y et al (2021) miR-223: An Immune Regulator in Infectious Disorders. Front Immunol [Internet]. [cited 2023 Oct 17];12. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fimmu.2021.781815
Nahand JS, Karimzadeh MR, Nezamnia M, Fatemipour M, Khatami A, Jamshidi S et al (2020) The role of miR-146a in viral Infection. IUBMB Life 72(3):343–360
Letafati A, Najafi S, Mottahedi M, Karimzadeh M, Shahini A, Garousi S et al (2022) MicroRNA let-7 and viral Infections: focus on mechanisms of action. Cell Mol Biol Lett 27(1):14
Mishra R, Kumar A, Ingle H, Kumar H The Interplay Between Viral-Derived miRNAs and Host Immunity During Infection. Front Immunol [Internet]. 2020 [cited 2023 Oct 17];10. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fimmu.2019.03079
Diggins NL, Hancock MH (2023) Viral miRNA regulation of host gene expression. Semin Cell Dev Biol 146:2–19
Circulating microRNAs as emerging regulators of COVID-19 - PMC [Internet]. [cited 2023 Oct 17]. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9800721/
Leonetti P, Ghasemzadeh A, Consiglio A, Gursinsky T, Behrens S, Pantaleo V (2021) Endogenous activated small interfering RNAs in virus-infected Brassicaceae crops show a common host gene‐silencing pattern affecting photosynthesis and stress response. New Phytol 229(3):1650–1664
Meng Z, Lu MRNA, Interference-Induced Innate Immunity, Off-Target Effect, or Immune Adjuvant? Front Immunol [Internet]. 2017 [cited 2023 Oct 17];8. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fimmu.2017.00331
Levanova A, Poranen MM RNA Interference as a Prospective Tool for the Control of Human Viral Infections. Front Microbiol [Internet]. 2018 [cited 2023 Oct 17];9. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fmicb.2018.02151
Cross RW, Mire CE, Feldmann H, Geisbert TW (2018) Post-exposure treatments for Ebola and Marburg virus Infections. Nat Rev Drug Discov 17(6):413–434
Wooddell C, Zhu R, Hamilton H, Chu Q, Sternard H, Schumacher J et al (2018) Development of subcutaneously administered RNAi therapeutic ARO-HBV for chronic Hepatitis B virus Infection. J Hepatol 68:S18–S19
Idris A, Davis A, Supramaniam A, Acharya D, Kelly G, Tayyar Y et al (2021) A SARS-CoV-2 targeted siRNA-nanoparticle therapy for COVID-19. Mol Ther J Am Soc Gene Ther 29(7):2219–2226
Ophinni Y, Palatini U, Hayashi Y, Parrish NF (2019) piRNA-Guided CRISPR-like immunity in eukaryotes. Trends Immunol 40(11):998–1010
Wang X, Huang P, Lei M, Ma Y, Chen H, Sun J et al (2023) Global expression and functional analysis of human piRNAs during HSV-1 Infection. Virus Res 328:199087
Akimniyazova A, Yurikova O, Pyrkova A, Rakhmetullina A, Niyazova T, Ryskulova AG et al (2022) In Silico Study of piRNA interactions with the SARS-CoV-2 Genome. Int J Mol Sci 23(17):9919
Taschuk F, Cherry S, DEAD-Box Helicases (2020) Sensors, regulators, and Effectors for Antiviral Defense. Viruses 12(2):181
Zong T, Yang Y, Zhao H, Li L, Liu M, Fu X et al (2021) tsRNAs: Novel small molecules from cell function and regulatory mechanism to therapeutic targets. Cell Prolif 54(3):e12977
Pawar K, Shigematsu M, Sharbati S, Kirino Y (2020) Infection-induced 5′-half molecules of tRNAHisGUG activate toll-like receptor 7. PLoS Biol 18(12):e3000982
Raftery N, Stevenson NJ (2017) Advances in anti-viral immune defence: revealing the importance of the IFN JAK/STAT pathway. Cell Mol Life Sci CMLS 74(14):2525–2535
Jiang P, Yan F (2019) tiRNAs & tRFs Biogenesis and Regulation of Diseases: a review. Curr Med Chem 26(31):5849–5861
Vignuzzi M, López CB (2019) Defective viral genomes are key drivers of the virus–host interaction. Nat Microbiol 4(7):1075–1087
Poirier EZ, Vignuzzi M (2017) Virus population dynamics during Infection. Curr Opin Virol 23:82–87
Dai H, Gu W (2020) Small RNA plays important roles in virus–host interactions. Viruses 12(11):1271
Ruivinho C, Gama-Carvalho M (2023) Small non-coding RNAs encoded by RNA viruses: old controversies and new lessons from the COVID-19 pandemic. Front Genet [Internet]. [cited 2023 Oct 17];14. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fgene.2023.1216890
Song J, Ouyang Y, Che J, Li X, Zhao Y, Yang K et al (2017) Potential Value of miR-221/222 as Diagnostic, Prognostic, and Therapeutic Biomarkers for Diseases. Front Immunol [Internet]. [cited 2023 Oct 17];8. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fimmu.2017.00056
Vuillier F, Li Z, Black I, Cruciani M, Rubino E, Michel F et al (2022) IFN-I inducible miR-3614-5p targets ADAR1 isoforms and fine tunes innate immune activation. Front Immunol [Internet]. [cited 2023 Oct 17];13. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fimmu.2022.939907
Regulation of influenza virus infection by microRNAs [Internet]. [cited 2023 Oct 17]. Available from: https://www.chinaagrisci.com/Jwk_zgnykxen/EN/https://doi.org/10.1016/S2095-3119(18)62134-3
Zhao L, Zhang X, Wu Z, Huang K, Sun X, Chen H et al (2019) The downregulation of MicroRNA hsa-mir-340-5p in IAV-Infected A549 cells suppresses viral replication by targeting RIG-I and OAS2. Mol Ther Nucleic Acids 14:509–519
Takahashi T, Heaton SM, Parrish NF (2021) Mammalian antiviral systems directed by small RNA. PLOS Pathog 17(12):e1010091
Panigrahi M, Thibault PA, Wilson JA (2022) MicroRNA 122 affects both the initiation and the maintenance of Hepatitis C Virus Infections. J Virol 96(4):e0190321
Zhang K, Lee YS, Lee I, Bao X, Editorial Small non-coding RNAs in diseases. Front Mol Biosci [Internet]. 2023 [cited 2023 Oct 17];9. Available from: https://www.frontiersin.org/articles/https://doi.org/10.3389/fmolb.2022.1086768
Rayford KJ, Cooley A, Rumph JT, Arun A, Rachakonda G, Villalta F et al (2021) piRNAs as modulators of Disease Pathogenesis. Int J Mol Sci 22(5):2373
Kurakula H, Vaishnavi S, Sharif MY, Ellipilli S (2023) Emergence of small interfering RNA-Based gene Drugs for various Diseases. ACS Omega 8(23):20234–20250
Ivanov P (2015) Emerging roles of tRNA-derived fragments in viral Infections: the case of respiratory Syncytial Virus. Mol Ther 23(10):1557–1558
Dasgupta I, Chatterjee A (2021) Recent advances in miRNA Delivery systems. Methods Protoc 4(1):10
Acknowledgements
The researchers express their sincere thanks and appreciation to the Department of Biotechnology, College of Science, University of Anbar, and the Department of Biology, College of Science, Tikrit University, for providing some of the research references.
Funding
There is no fund received for this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval and consent to participate
not applicable.
Consent for publication
not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Suleiman, A.A., Al-Chalabi, R. & Shaban, S.A. Integrative role of small non-coding RNAs in viral immune response: a systematic review. Mol Biol Rep 51, 107 (2024). https://doi.org/10.1007/s11033-023-09141-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11033-023-09141-6