Abstract
Background
Uropathogenic Escherichia coli (UPEC) is a major pathogen of the urinary tract infection (UTI), and biofilm formation is crucial as it facilitates the colonization in the urinary tract. We aimed to investigate the antibiotic susceptibility pattern, biofilm formation capability, distribution of quinolone resistance genes, and phylogenetic groups among UPEC isolates from an Iranian inpatients’ community.
Methods and results
A collection of 126 UPEC obtained from hospitalized patients with symptomatic UTI at 3 teaching hospitals during 2016 were included. Antibiogram of all isolates against quinolone and fluoroquinolones was performed using the disk diffusion method. Phylogenetic groups and qnr A, B, and S genes were assessed by PCR. Susceptibility pattern showed that more than 50% and 81% of the isolates were resistant to fluoroquinolones and quinolones, correspondingly. The frequency of qnrS and qnrB genes was 22% and 13.5%, correspondingly. Our result indicated no significant association between the presence of fluoroquinolone genes and antibiotic resistance to them. The frequent common phylogroup was B2 (84.1%), followed by D (10.3%), A (3.2%) and B1 (2.4%) groups. Indeed, 80.2% of the isolates were biofilm producers, so that 42.1%, 16.7% and 21.4% of them were classified as weak, moderate and strong producers, respectively.
Conclusions
Our results showed considerable fluoroquinolone and quinolone resistance among UPEC along with a remarkable rate of biofilm-producing isolates from symptomatic hospitalized patients, making them a serious health concern in the region. This survey highlights the need for awareness on quinolone resistance and careful prescription of them by physicians.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Urinary tract infection (UTI) is one of the frequently recognized bacterial infections, striking 150 million humans every year. It results in high morbidity and mortality and also costly health and medical issues [1]. Among the numerous bacterial agents causing UTI, Uropathogenic Escherichia coli (UPEC) is one of the etiologic agents of UTIs, accounting for 80–90%, and 50% of the community and hospital-acquired UTIs, respectively [1,2,3,4,5]. Among the pathogenesis characterization of UPEC isolates, biofilm formation is crucial as it facilitates their colonization in the urethra [6,7,8]. Moreover, recurrence of UTI by UPEC isolates might be due to the capability of virulent strains in biofilm production [9]. Biofilm formation can affect both the activity of antimicrobial agents and the host immune response, leading to the persistence of UPEC in the urinary tract and emergence of severe symptoms and antimicrobial resistance [7,8,9,10]. Several antibiotics are used for treatment of UTI, including Nitrofurantoin, Trimethoprim/Sulfamethoxazole, Oral β-lactams, fosfomycin and Fluoroquinolones [11]. Nitrofurantoin is the best recommended treatment for cystitis. This drug inhibits bacterial enzymes involved in the synthesis of DNA, RNA, cell wall protein synthesis, and other metabolic enzymes [12]. Trimethoprim/Sulfamethoxazole has been considered as the standard for therapy of acute and recurrent UTIs. These two agents act synergistically in inhibiting folic acid synthesis [11]. Fosfomycin inhibits an enzymatic-catalyzed reaction in the first step of the synthesis of bacterial cell wall [13]. Quinolones, as one of the frequently used antibiotics, are critical drugs for therapy of UTIs caused by Gram-negative bacteria in clinical settings [14, 15]. The extensive and inappropriate use of quinolones and other antimicrobial agents for the management of bacterial infections has led to a remarkable increase in resistant isolates which are recognized as a great public health concern [16, 17]. Four main groups of qnr determinants, namely qnrA, qnrB, qnrC, and qnrS, have been diagnosed. These plasmid-mediated quinolone resistance (PMQR) determinants act through protection of DNA gyrase and topoisomerase IV from antibiotic action [10, 18]. The increased antibiotic resistance could be attributed to horizontal transfer of genes for formation of multi-drug resistant (MDR) strains; therefore, biofilms are considered as an ideal niche for this transfer especially plasmid exchange [19, 20]. To study the population genetics of UPEC isolates, researchers have extensively used phylogenetic groups for their simplicity [21]. Both virulent and commensal isolates of E. coli have been categorized into four major phylogenetic sets consisting of A, B1, B2, and D, which are common in humans. Accordingly, human virulent extraintestinal pathogenic Escherichia coli (ExPEC) are commonly classified as phylogroup B2 or D, while A or B1 groups mostly belong to commensal strains and less pathogenic strains [22, 23]. Determination of resistance pattern and phylogenetic groups and their association is important in characterization of the isolates [21]. Due to lack of studies on quinolone resistance genes, phylogenetic groups and biofilm formation among UPEC isolates in our region, this study aimed to investigate these characteristics among UPEC isolates from inpatients in Shiraz, southern Iran.
Materials and methods
Bacterial isolates and study population
In this study, a total of 126 non-duplicate UPEC which were recovered from a previous study in 2016 were used [16]. The participants were hospitalized patients with symptomatic UTI, including cystitis, pyelonephritis and urosepsis. Exclusion criteria were patients with asymptomatic UTI, and not taking antibiotics during 1 month ago. The Ethics Committee of Shiraz University of Medical Sciences approved this study (Approval No. EC IR.SUMS.REC.1397.688).
Quinolone susceptibility testing
The antibiotic susceptibility pattern was performed by standard disk diffusion method on Mueller–Hinton agar (Himedia, India) plate according to the Clinical and Laboratory Standards Institute (CLSI) recommendation for nalidixic acid, ciprofloxacin, levofloxacin, norfloxacin and ofloxacin (Mast Co., UK) [24]. In our study E. coli ATCC 25922 was used as the control strain.
Phylogenetic grouping and the presence of quinolone resistance genes
After recovery of the isolates from freeze − 70 °C by plating onto blood agar, DNA extraction of the studied isolates was conducted using the boiling method as described previously [25]. PCR for detection of qnrA, qnrB, and qnrS genes [26] was carried out on a T100™ thermal cycler (Bio-Rad, Hercules, CA, USA) in a total volume of 25 µL containing 3 µL DNA template, 2.5 µL PCR buffer (1×), 1 µL deoxyribonucleotide triphosphates solution (dNTPs, 200 µM), 1.5 µL MgCl2 (1.5 mM), 0.25 µL Taq DNA polymerase (1 Unit), and 1 µL each specific primer (1 µM). Indeed, PCR was used for determining the phylogenetic groups (A, B1, B2 and D) through amplification of chuA and yjaA genes and the DNA fragment TspE4.C2 [21]. PCR amplification consisted of initial-denaturation at 94 °C for 5 min, followed by 35 cycles of denaturation at 94 °C for 45 s, primer annealing at 54 °C and extension at 72 °C for 60 s, and a final extension at 72 °C for 7 min. The amplicons were analyzed on agarose gel 1.5%, stained with safe stain (CinnaGen Co. Iran) and finally visualized under ultraviolet light.
Biofilm formation assay
A microtiter plate method was performed to investigate the biofilm formation based on the protocol of O’Toole et al. [27]. Briefly, after adding 100 µL of the 1:100 dilutions of bacterial cultures into 96 well plates, they were incubated for 24 h at 37 °C. After washing of the wells with distilled water, the wells were stained with 0.1% solution of crystal violet (CV) and incubated for 15 min at room temperature. The microplates were rinsed and dried; then, 30% acetate was added as solvent of CV. After 15 of min incubation, the contents of wells were moved to a new microplate and their absorbance was read at 550 nm by ELISA reader. All tests were performed in triplicate.
DNA sequence analysis
To confirm the accuracy of amplified genes, the amplicons (four samples) were submitted for sequencing (Bioneer Co., Munpyeongseoro, Daedeok-gu, Daejeon, South Korea) and the sequences were compared using online BLAST software (http://www.ncbi.nlm.nih.gov/BLAST/).
Statistical analysis
The analysis was performed using SPSS™ software, version 21.0 (IBM Corp., USA). The Chi-square test was used and differences were considered significant when the P value was ≤ 0.05.
Results
Study population
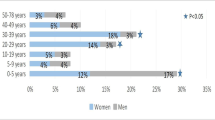
Our archive (126 UPEC) isolates were obtained from fifty females and seventy six males with an age range from 1 to 100 years old with a mean age of 48.9 ± 28.8 years. No significant association was shown in the age and gender of cases in relation to the three studied clinical groups (cystitis, pyelonephritis and urosepsis).
The wards of UPEC isolation were from Intensive Care Unit (ICU) with a frequency of 76 (60.4%), followed by Internal ward (36; 28.6%), Surgery ward (7; 5.6%), and Transplantation center (7; 5.6%). Furthermore, the frequency of the cases in different wards was as follows: cystitis (ICU = 23, Internal ward = 12, Surgery = 2, Transplantation = 5), pyelonephritis (ICU = 44, Internal ward = 22, Surgery = 5, Transplantation = 2) and urosepsis (ICU = 9, Internal ward = 2, Surgery = 0, Transplantation = 0).
Quinolone resistance among UPEC isolates
The results of antibiogram pattern revealed that the least susceptibility was against nalidixic acid (19%), followed by ofloxacin with 42.1% (Table 1).
Characterization of quinolone resistance genes
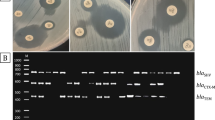
The analysis of PCR results showed that 22.2% (28/126) and 13.5% (17/126) of the isolates were positive for the qnrS and qnrB genes, respectively. Meanwhile, 2.4% of the isolates were positive for both qnrB and qnrS genes simultaneously, and qnrA gene was not detected in any of UPEC isolates. Descriptive results of the association between quinolone resistance and the presence of qnr genes showed that, among qnrS negative and positive isolates, the highest resistance rate was toward nalidixic acid, while the most effective antibiotic was ciprofloxacin. Statistical analysis indicated that there was no significant association between antibiotic resistance and the existence of qnr genes (Tables 2 and 3) (Fig. 1).
Gel electrophoresis of PCR products for qnrB, qnrS, chuA, TspE4C2 and yjaA genes in UPEC isolates. Lane 1 = chuA gene, Lane 2 = chuA Positive control, Lane 3 = TspE4C2 gene, Lane 4 = TspE4C2 Positive control, Lane 5 = yjaA gene, Lane 6 = yjaA Positive control, Lane M = DNA ladder (100 bp), Lane 7 = qnrS gene, Lane 8 = qnrS Positive control, Lane 9 = qnrB gene, Lane 10 = qnrB Positive control, Lane 11: Negative control
Phylogenetic grouping
PCR method for the investigation of phylogroups showed that UPEC isolates were separated into four phylogenetic groups (A, B1, B2, and D) and seven subgroups. Out of 126 UPEC isolates, the commonest phylogroup was group B2 (106 isolates; 84.1%), followed by group D (13, 10.3%), group A (4, 3.2%) and group B1 (3, 2.4%) (Fig. 1). Furthermore, the main phylogenetic groups were divided into seven subgroups, namely B23 (81%), D2 (7.8%), B22 (3.2%), B1 (2.4%), D1 (2.4%), A1 (2.4), and A0 (0.8%). Distribution of phylogroups according to the ward’s, clinical disease, and biofilm formation capability is shown in Table 4.
Biofilm production
Out of the 126 UPEC isolates, 80.2% (101/126) were positive to the ability of biofilm formation using the microtiter plate assay and were categorized into four groups based on their ability for biofilm production. According to the results, 42.1%, 16.7% and 21.4% of UPEC isolates were classified as weak, moderate and strong biofilm producers.
Discussion
Today, due to antibiotic resistance to first-line drugs, quinolones and fluoroquinolones are the preferential options for curing UTIs caused by UPEC isolates [28]. In this study, quinolone susceptibility pattern, biofilm formation and the distribution of quinolone resistance genes and phylogenetic groups among 126 UPEC isolated from a symptomatic population in southern Iran were assessed.
In the current survey, antibiotic susceptibility pattern showed that more than 50% and 81% of the isolates were resistant to fluoroquinolones and quinolones (nalidixic acid), respectively. In a recent survey from Shiraz, an increase in quinolone-resistant UPEC isolates (more than 40%) was reported; this in the same line with our study [29]. Shenagari and co-workers from north of Iran reported that 45.3% of UPEC isolates were resistant to norfloxacin, 48.9% to ofloxacin, 50.2% to ciprofloxacin, and 61.9% to nalidixic acid [30]. In the neighboring countries, reports indicate a high resistance to fluoroquinolone and quinolone. For instance, Muhammad et al. from Pakistan reported that the resistance rate to ciprofloxacin and nalidixic acid among UPEC isolates was 36.45% and 84.16%, respectively [31]. Because quinolones and fluoroquinolones are used for empirical therapy of complicated UTI, there is a major challenge for physicians to treat this infection [32].
Increased resistance to fluoroquinolones among Enterobacterales is associated with a high rate of PMQR genes which is related to diversity of PMQR genes or mutations in gyrA and parC genes, or both of them in PMQR-positive strains [33]. Accordingly, in the current study the frequency of qnrS and qnrB genes was found 22% and 13.5%, respectively, indicating no significant difference between the presence of fluoroquinolone genes and antibiotic resistance to them. In investigation of Malekzadegan et al. 33.1% and 12.4% of the isolates were qnrS and qnrB positive, respectively, and no significant difference was found between higher antibiotic resistance and qnr genes [29]. In contrast to our research, in studies conducted by Sedighi and colleagues [34] and Rezazadeh et al. [35]., low rates of qnrS and qnrB genes among quinolone-resistant UPEC isolates were identified, along with a significant association between qnr genes and quinolone resistance. In contrast, in a study from north of Iran, regarding to UPEC, a high prevalence of qnrB (71.3%) and qnrA (62.8%) genes together a considerable correlation with resistance to quinolones has been reported [36]. In the current work, no isolate harbored qnrA gene which is consistent with the result of the studies conducted by Sedighi and co-workers and Rezazadeh et al. [34, 35]. Although there is relative agreement on the high prevalence of qnrS and qnrB genes in UPEC isolates, the distribution of predominant genes varies in different areas.
According to Clermont classification, E. coli strains based on chuA, yjaA and TspE4.C2 genetic determinants have been divided into four phylogenetic groups, including A, B1, B2, and D [37].
In consistent with the literature, all of the major phylogroups were determined in the current work. In line with prior studies [8, 21], most of our isolates belonged to group B2, followed by D group. Conversely, in studies from Romania and India after B2 as the dominant group, B1 and A groups were the second phylogroup, respectively [38, 39]. Distribution of phylogenetic groups varies according to geographical regions, even within each country. In accordance with our results, Yazdanpour et al. and Mostafavi and colleagues revealed that among 248 their UPEC isolates, 67.3%, and 45.4% of the isolates were associated with the B2 phylogenetic group, followed by D (21.4%, 25.3%), A (6.5%, 10.5%) and B1 (4.8%, 16.8%) groups, respectively [40, 41]. Iranpour et al. and Derakhshan et al. also showed that the B2 group was the most prevalent group among UPEC isolates and phylogenetic group of D had a very low rate (2.9% and 3.3%) [42, 43]. Even though previous investigations demonstrate that commensal strains are associated with group A or group B1[37], Hashemizadeh and co-workers revealed that B2 and D were the frequent phylogroups in fecal isolates [44]. Therefore, distribution of various types of phylogroups among E. coli isolates may be caused by the heterogeneity of ExPEC isolates in various studies.
It has been suggested that B2 phylogenetic group isolates reveal higher quinolone-susceptibility than those of other groups [39]. However, in our survey the isolates belonging to B2 phylogroup demonstrated higher resistance against fluoroquinolones. Likewise, in a report from Iran, B2 and D phylogroups represented the most resistant UPEC isolates to antimicrobials [45].
In 80% of infections, biofilm formation in bacteria is documented to account for a serious problem in UTIs [46]. Biofilm production by UPEC strains in the urinary tract, as well as on the urinary catheters, is one of the major causes of the persistence of these isolates, so that in most cases it leads to recurrent infections [9]. Our results showed that 80.2% (101/126) of the isolates were positive for biofilm production, of which, 42.1%, 16.7% and 21.4% of UPEC isolates were classified as weak, moderate and strong positive in their ability to form biofilms, correspondingly. In the study of Tajbakhsh et al., out of 80 biofilm-producing E. coli strains, 15 (18.75%) were strong, 20 (25%) moderate, and 45 (56.25%) weak positive isolates [47]. Also, Ponnusami and colleagues observed that among 100 UPEC isolates, 17.23%, 26.3% and 50% were strong, intermediate and weak biofilm formers, respectively [48]. These findings are relatively similar to our results. Likewise, Poursina et al. studied the biofilm formation assay of 100 UPEC isolates, of which 80% had capability of this characteristic and among them 29% were strong producers [32]. Jomezadeh and co-workers recovered 98 UPEC isolates, 42.85% of which were phenotypically biofilm formers, among them the majority (38%) formed moderate biofilms [49]. Taken together, these results imply that due to the high rate of biofilm-producing UPEC isolates, it is necessary to pay attention to these isolates for fast treatment of UTIs. The current work had a limitation, so that we did not evaluate the correlation of UPEC virulence determinants within phylogenetic groups, because it was beyond the scope of this work.
Conclusion
In summary, UPECs isolated from symptomatic hospitalized patients were highly resistant to fluoroquinolone and quinolones and biofilm formation capability was considerable among them. Additionally, we determined a remarkable rate of the isolates belonging predominantly to B2 and D phylogenetic groups. These findings strengthen the importance of these types of investigations for epidemiological surveillance and the therapeutic or prophylactic purposes of nosocomial UTIs.
Data availability
All data associated with this manuscript is inclusive in this paper.
Abbreviations
- UTI:
-
Urinary tract infection
- UPEC:
-
Uropathogenic Escherichia coli
- PMQR:
-
Plasmid-mediated quinolone resistance
- MDR:
-
Multi-drug resistant
- ExPEC:
-
Extraintestinal pathogenic Escherichia coli
- CLSI:
-
Clinical and Laboratory Standards Institute
References
Flores-Mireles AL, Walker JN, Caparon M, Hultgren SJ (2015) Urinary tract infections: epidemiology, mechanisms of infection and treatment options. Nat Rev Microbiol 13(5):269–284
Terlizzi ME, Gribaudo G, Maffei ME (2017) Uropathogenic Escherichia coli (UPEC) infections: virulence factors, bladder responses, antibiotic, and non-antibiotic antimicrobial strategies. Front Microbiol 8:1566
Walker E, Lyman A, Gupta K, Mahoney MV, Snyder GM, Hirsch EB (2016) Clinical management of an increasing threat: outpatient urinary tract infections due to multidrug-resistant uropathogens. Clin Infect Dis 63(7):960–965
Wiles TJ, Kulesus RR, Mulvey MA (2008) Origins and virulence mechanisms of uropathogenic Escherichia coli. Exp Mol pathol 85(1):11–19
Karshenas AE, Salehi TZ, Adabi M, Asghari B, Yahyaraeyat R (2022) Prevalence of main quinolones and carbapenems resistance genes in clinical and veterinary Escherichia coli strains. Iran J Microbiol 14(6):841–849
Bien J, Sokolova O, Bozko P (2012) Role of uropathogenic Escherichia coli virulence factors in development of urinary tract infection and kidney damage. Int J Nephrol 2012:681473
Eberly AR, Floyd KA, Beebout CJ, Colling SJ, Fitzgerald MJ, Stratton CW et al (2017) Biofilm formation by uropathogenic Escherichia coli is favored under oxygen conditions that mimic the bladder environment. Int J Mol Sci 18(10):2077
Pootong A, Mungkornkeaw N, Norrapong B, Cowawintaweewat S (2018) Phylogenetic background, drug susceptibility and virulence factors of uropathogenic E. coli isolate in a tertiary university hospital in central Thailand. Trop Biomed 35(1):195–204
Soto S, Smithson A, Martinez J, Horcajada J, Mensa J, Vila J (2007) Biofilm formation in uropathogenic Escherichia coli strains: relationship with prostatitis, urovirulence factors and antimicrobial resistance. J Urol 177(1):365–368
Pompilio A, Crocetta V, Savini V, Petrelli D, Di Nicola M, Bucco S et al (2018) Phylogenetic relationships, biofilm formation, motility, antibiotic resistance and extended virulence genotypes among Escherichia coli strains from women with community-onset primitive acute pyelonephritis. PLoS ONE 13(5):e0196260
Kumar Shrestha B, Tumbahangphe M, Shakya J, Chauhan S (2022) Uropathogenic Escherichia coli in urinary tract infections: a review on epidemiology, pathogenesis, clinical manifestation, diagnosis, treatments and prevention. Nov Res Microbiol J 6(4):1614–1634
Squadrito FJ, del Portal D (2022) Nitrofurantoin. StatPearls Publishing, Florida
Dzib-Baak HE, Uc-Cachón AH, Dzul-Beh AdJ, Rosado-Manzano RF, Gracida-Osorno C, Molina-Salinas GM (2022) Efficacy of fosfomycin against planktonic and biofilm-associated MDR uropathogenic Escherichia coli clinical isolates. Trop Med Infect Dis 7(9):235
Minardi D, d’Anzeo G, Cantoro D, Conti A, Muzzonigro G (2011) Urinary tract infections in women: etiology and treatment options. Int J Gen Med 4:333–343
Kang C-I, Kim J, Park DW, Kim B-N, Ha U-S, Lee S-J et al (2018) Clinical practice guidelines for the antibiotic treatment of community-acquired urinary tract infections. Infect Chem 50(1):67–100
Malekzadegan Y, Khashei R, Sedigh Ebrahim-Saraie H, Jahanabadi Z (2018) Distribution of virulence genes and their association with antimicrobial resistance among uropathogenic Escherichia coli isolates from iranian patients. BMC Infect Dis 18(1):1–9
Varughese LR, Rajpoot M, Goyal S, Mehra R, Chhokar V, Beniwal V (2018) Analytical profiling of mutations in quinolone resistance determining region of gyrA gene among UPEC. PLoS ONE 13(1):e0190729
Hassan D, El-Moaafy MA, Saleh HH, Abdelrahman AA, Abdallah SE (2022) Characterization of quinolone resistance genes in uropathogenic Escherichia coli isolates, in Egypt. Rec Pharm Biomed Sci 6(2):163–172
Lerminiaux NA, Cameron AD (2019) Horizontal transfer of antibiotic resistance genes in clinical environments. Can J Microbiol 65(1):34–44
Stalder T, Top E (2016) Plasmid transfer in biofilms: a perspective on limitations and opportunities. NPJ Biofilms Microbiomes 2:16022
Clermont O, Christenson JK, Denamur E, Gordon DM (2013) The Clermont Escherichia coli phylo-typing method revisited: improvement of specificity and detection of new phylo‐groups. Environ Microbiol Rep 5(1):58–65
Carlos C, Pires MM, Stoppe NC, Hachich EM, Sato MI, Gomes TA et al (2010) Escherichia coli phylogenetic group determination and its application in the identification of the major animal source of fecal contamination. BMC Microbiol 10:161
Er DK, Dundar D, Uzuner H, Osmani A (2015) Relationship between phylogenetic groups, antibiotic resistance and patient characteristics in terms of adhesin genes in cystitis and pyelonephritis isolates of Escherichia coli. Microb pathog 89:188–194
CLSI (2020) Performance Standards for Antimicrobial Susceptibility Testing. CLSI supplement M100. PCaLSI, 30th edn. Clinical and Laboratory Standards Institute, Wayne
Gajamer VR, Bhattacharjee A, Paul D, Ingti B, Sarkar A, Kapil J et al (2020) High prevalence of carbapenemase, AmpC β-lactamase and aminoglycoside resistance genes in extended-spectrum β-lactamase-positive uropathogens from Northern India. J Glob Antimicrob Resist 20:197–203
Cattoir V, Poirel L, Rotimi V, Soussy C-J, Nordmann P (2007) Multiplex PCR for detection of plasmid-mediated quinolone resistance qnr genes in ESBL-producing enterobacterial isolates. J Antimicrob Chemothe 60(2):394–397
O’Toole GA (2011) Microtiter dish biofilm formation assay. JoVE 47:e2437
Hooton TM (2003) Fluoroquinolones and resistance in the treatment of uncomplicated urinary tract infection. Int J Antimicrob Agents 22:65–72
Malekzadegan Y, Rastegar E, Moradi M, Heidari H, Ebrahim-Saraie HS (2019) Prevalence of quinolone-resistant uropathogenic Escherichia coli in a tertiary care hospital in south Iran. Infect Drug Resist 12:1683–1689
Shenagari M, Bakhtiari M, Mojtahedi A, Roushan ZA (2018) High frequency of mutations in gyrA gene associated with quinolones resistance in uropathogenic Escherichia coli isolates from the north of Iran. Iran J Basic Med Sci 21(12):1226–1231
Muhammad I, Uzma M, Yasmin B, Mehmood Q, Habib B (2011) Prevalence of antimicrobial resistance and integrons in Escherichia coli from Punjab, Pakistan. Braz J Microbiol 42(2):462–466
Poursina F, Sepehrpour S, Mobasherizadeh S (2018) Biofilm formation in nonmultidrug-resistant Escherichia coli isolated from patients with urinary tract infection in Isfahan, Iran. Adv Biomed Res 7:40
Kim HB, Park CH, Kim CJ, Kim E-C, Jacoby GA, Hooper DC (2009) Prevalence of plasmid-mediated quinolone resistance determinants over a 9-year period. Antimicrob Agents Chemother 53(2):639–645
Sedighi I, Arabestani MR, Rahimbakhsh A, Karimitabar Z, Alikhani MY (2015) Dissemination of extended-spectrum β-lactamases and quinolone resistance genes among clinical isolates of uropathogenic Escherichia coli in children. Jundishapur J Microbiol 8(7):e19184
Rezazadeh M, Baghchesaraei H, Peymani A (2016) Plasmid-mediated quinolone-resistance (qnr) genes in clinical isolates of Escherichia coli collected from several hospitals of Qazvin and Zanjan provinces, Iran. Osong Public Health Res Perspect 7(5):307–312
Yousefi S, Mojtahedi A, Shenagari M (2018) A Survey of gyrA target-site mutation and qnr genes among clinical isolates of Escherichia coli in the North of Iran. Jundishapur J Microbiol 11(9):e67293
Clermont O, Bonacorsi S, Bingen E (2000) Rapid and simple determination of the Escherichia coli phylogenetic group. Appl Environ Microbiol 10(66):4555–4558
Cristea VC, Gheorghe I, Czobor Barbu I, Popa LI, Ispas B, Grigore GA et al (2019) Snapshot of phylogenetic groups, virulence, and resistance markers in Escherichia coli uropathogenic strains isolated from outpatients with urinary tract infections in Bucharest, Romania. BioMed Res Int. https://doi.org/10.1155/2019/5712371
Harwalkar A, Gupta S, Rao A, Srinivasa H (2014) Lower prevalence of hlyD, papC and cnf-1 genes in ciprofloxacin-resistant uropathogenic Escherichia coli than their susceptible counterparts isolated from southern India. J Infect Public Health 7(5):413–419
Yazdanpour Z, Tadjrobehkar O, Shahkhah M (2020) Significant association between genes encoding virulence factors with antibiotic resistance and phylogenetic groups in community acquired uropathogenic Escherichia coli isolates. BMC Microbiol 20(1):241
Shokouhi Mostafavi SK, Najar-Peerayeh S, Mohabbati Mobarez A, Kardoust Parizi M (2019) Serogroup distribution, diversity of exotoxin gene profiles, and phylogenetic grouping of CTX-M-1- producing uropathogenic Escherichia coli. Comp Immunol Microbiol Infect Dis 65:148–153
Iranpour D, Hassanpour M, Ansari H, Tajbakhsh S, Khamisipour G, Najafi A (2015) Phylogenetic groups of Escherichia coli strains from patients with urinary tract infection in Iran based on the new Clermont phylotyping method. Biomed Res Int 2015:846219
Derakhshan S, Ahmadi S, Ahmadi E, Nasseri S, Aghaei A (2022) Characterization of Escherichia coli isolated from urinary tract infection and association between virulence expression and antimicrobial susceptibility. BMC Microbiol 22(1):89
Hashemizadeh Z, Kalantar-Neyestanaki D, Mansouri S (2017) Association between virulence profile, biofilm formation and phylogenetic groups of Escherichia coli causing urinary tract infection and the commensal gut microbiota: a comparative analysis. Microb Pathog 110:540–545
Farajzadah Sheikh A, Goodarzi H, Yadyad MJ, Aslani S, Amin M, Jomehzadeh N et al (2019) Virulence-associated genes and drug susceptibility patterns of uropathogenic Escherichia coli isolated from patients with urinary tract infection. Infect Drug Resist 12:2039–2047
Delcaru C, Alexandru I, Podgoreanu P, Grosu M, Stavropoulos E, Chifiriuc MC et al (2016) Microbial biofilms in urinary tract infections and prostatitis: etiology, pathogenicity, and combating strategies. Pathogens 5(4):65
Tajbakhsh E, Ahmadi P, Abedpour-Dehkordi E, Arbab-Soleimani N, Khamesipour F (2016) Biofilm formation, antimicrobial susceptibility, serogroups and virulence genes of uropathogenic E. coli isolated from clinical samples in Iran. Antimicrob Resist Infect control 5:11
Ponnusamy P, Natarajan V, Sevanan M (2012) In vitro biofilm formation by uropathogenic Escherichia coli and their antimicrobial susceptibility pattern. Asian Pac J Trop Med 5(3):210–213
Jomehzadeh N, Ahmadi K, Nasiri Z (2022) Evaluation of biofilm formation and antibiotic resistance pattern in extended-spectrum β-lactamase-producing Escherichia coli strains. Biomed Biotechnol Res J (BBRJ) 6(2):175–179
Acknowledgements
This work is extracted from the M.Sc thesis of Elham Rastegar in partial fulfillment of the requirements for the Medical Microbiology Master’s degree. The authors wish to thank Dr. Shokrpour at the Research Consultation Center (RCC) of Shiraz University of Medical Sciences for her valuable assistance in editing of this manuscript.
Funding
This work was supported by Shiraz University of Medical Sciences, (Grant number 14941).
Author information
Authors and Affiliations
Contributions
RK designed the study, ER and YM conducted the experiments and analyzed the data, RK and ER wrote the original draft which was revised by RK, RK and NH who were the supervisors of the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This study was in accordance with the declaration of 1964 Helsinki and approved by the institutional Ethics Committee of Shiraz University of Medical Sciences (Approval No. IR. SUMS. REC. 1397.688). However, because we only used leftovers from clinical specimens, the institutional ethics committee waived the need for informed consent.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rastegar, E., Malekzadegan, Y., Khashei, R. et al. Quinolone resistance and biofilm formation capability of uropathogenic Escherichia coli isolates from an Iranian inpatients’ population. Mol Biol Rep 50, 8073–8079 (2023). https://doi.org/10.1007/s11033-023-08704-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08704-x