Abstract
Background
Eukaryotic elongation factor 2 kinase (eukaryotic elongation factor 2 kinase, eEF2K) is a calcium calmodulin dependent protein kinase that keeps the highest energy consuming cellular process of protein synthesis under check through negative regulation. eEF2K pauses global protein synthesis rates at the translational elongation step by phosphorylating its only kown substrate elongation factor 2 (eEF2), a unique translocase activity in ekaryotic cells enabling the polypeptide chain elongation. Therefore, eEF2K is thought to preserve cellular energy pools particularly upon acute development of cellular stress conditions such as nutrient deprivation, hypoxia, or infections. Recently, high expression of this enzyme has been associated with poor prognosis in an array of solid tumor types. Therefore, in a growing number of studies tremendous effort is being directed to the development of treatment methods aiming to suppress eEF2K as a novel therapeutic approach in the fight against cancer.
Methods
In our study, we aimed to investigate the changes in the tumorigenicity of chordoma cells in presence of gene silencing for eEF2K. Taking a transient gene silencing approach using siRNA particles, eEF2K gene expression was suppressed in chordoma cells.
Results
Silencing eEF2K expression was associated with a slight increase in cellular proliferation and a decrease in death rates. Furthermore, no alteration in the sensitivity of chordoma cells to chemotherapy was detected in response to the decrease in eEF2K expression which intriguingly promoted suppression of cell migratory and invasion related properties.
Conclusion
Our findings indicate that the loss of eEF2K expression in chordoma cell lines results in the reduction of metastatic capacity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chordoma is a slowly developing, soft tissue type of cancer that typically originates from the notochordal remnants in the embryonic period and tends to localize at the base of the skull and around the sacrum in the axial skeletal system [1]. Chordoma was first reported by Virchow in 1857 as a clivus tumor [2]. According to SEER data, the rate of diagosis reported is 1 in 106 in the USA, and it mostly occurs between the ages of 40–70 and its incidence under the age of 20 is 1% [3]. Although basis for gender-based predisposition is not well understood, chordoma is seen approximately twice more in women than in men, [4,5,6,7]. One of the pathophysiological features of chordomas is the overexpression of the gene called T (BRACHYURY, mouse homolog), which is used as a biomarker for chordoma diagnosis [8]. Chordoma is highly resistant to traditional therapies like chemotherapy and relatively more responsive to radiotherapy which is -therefore- preferred as a follow up treatment option particularly ensuing radical surgeries [9,10,11]. Therefore, alternative treatment methods are needed because of the inadequacy of existing chemotherapy options. For that matter, understanding the molecular mechanisms driving chordoma carcinogenesis is extremely important for the discovery and development of novel targeted molecules of high efficacy. To study the molecular pathogenesis of the disease there have been several cell lines established either from primary or recurred tumors collected from patients. MUG-Chor1 and UM-Chor1 chordoma cell lines, both of which were the selected model systems for this study, are examples of such in vitro models. MUG-Chor1 originates from sacrococcygeal recurrent tumor tissue, while UM-Chor1 was established from clivial primary tumor tissue [12,13,14].
In an attempt to uncover a novel point of intervention for the treatment of chordoma tumors we undertook understanding the therapeutic potential of silencing of eEF2K which recently has emerged as a suitable molecular target in several solid tumors types such as breast, pancreas, lung, and brain cancer [15].
Protein synthesis is an essential cellular process for cell vialbility and is accomplished by the action of a humbling machinery comprised of several factors acting in a highly sophisticated and meticulously orchestrated fashion [16]. Furthermore, synthesis of nascent proteins dedicated to different functions to sustain cell viability claims 30–50% of cellular energy [15]. In that respect, eEF2K acts as a reversible break of this multi-member machinery to sustain metabolic energy until stress conditions are reverted back to normal [17]. eEF2K executes its break function to bring polypeptide chain elongation to a halt by phosphorylating its only known substrate, eEF2 (Eukaryotic Elongation Factor 2), which possesses a unique translocase activity in eukaryotic cells allowing the progression of ribosome on an mRNA being translated [18, 19]. In response to acute stress, the activity of eEF2K toward eEF2 increases so that the interaction of the phosphorylated eEF2 with the ribosome is abrogated, and thereby, protein synthesis becomes paused until the inhibitory phosphates are removed to allow its re-activation and re-access to the ribosome when normalcy in the growth conditions are re-attained [20].
Majority of the cancer cells in tumor bed have a higher division capacity as well as metabolic activity compared to normal cells which forms the basis for the severity of the living conditions in their microenviroment that is deficient in oxygen, nutrients and energy and is highly acidic. Initially, the association between high eEF2K expression in cancer cells and worse degree of malignancy appeared to be contradicting the prediction that a high expression of a negative regulator for protein synthesis does not reconcile with meeting the demand for the high protein synthesis rates. However, from the perspective of cytoprotective impact of eEF2K activity through energy preservation under cellular stress conditions, it becomes explicable why cancer cells of certain tissue types favors elevation of eEF2K levels in their deprived microenvironment where they have endure. In addition, in order provide high protein synthesis rates explotation of other regulators of protein translation machinery is a frequently encountered event in cancer cells where there is a strong propensity to increase the expression of those genes important for cancer development while decreasing the expression of others that prevent cancer formation [21]. Several lines of evidence obtained in different cancer models suggest that aberrant proliferation, angiogenesis, metastasis, pro-tumorigenic immune response and cancer energetics meets such pathological needs through aberrant protein translation [16].
Regulation of eEF2K function is highly complex under the control of several mitogenic signaling pathways, including PI3K (Phosphoinositide 3-kinase), mTOR (mammalian target of rapamycin-rapamycin in mammals) and MAPK (mitogen-activated protein kinase) the activity of which blocks eEF2K activity to allow protein synthesis in accordance with proliferation inducing stimuli [22]. In contrast, AMPK, the major energy sensor signaling activates eEF2K through direct phosphorylation. In several types of cancer, de-regulation of these pathways has been shown to be pivotal events driving the oncogenesis and determining the degree of malignancy whereby more aggressive phenotypes are more frequently associated with malfunction of these pathways executing key roles in cell division, differentiation, metabolism and motility. Strikingly, depending on the nutrient and energy status of the cell, signaling through these decision making pathways for the cell fate converge on eEF2K whose outcome activity directs protein synthesis accordingly [23].
Previous studies indicate that, silencing of eEF2K protein in glioma cells is shown to trigger Tumor Necrosis Factor (TNF)-related apoptosis-inducing ligand (TRAIL) dependent death mechanism [24] and reduces the proliferation of breast cancer cells by increasing sensitivity to chemotherapeutics such as doxorubicin [25]. Likewise, knockdown of higher expression of eEF2K compared to normal in esophageal squamous cell carcinoma (ESCC) slowed down migration and proliferation rate [34]. So far there has been no such study showing the association of high expression of eEF2K and bad prognosis in chordoma and whether eEF2K has any effect on the drug resistance [34].
Recent reports highlighted Afatinib, an Epidermal Growth Factor Receptor (EGFR) inhibitor, as a promising agent that inhibits the proliferation of chordoma cells via decreasing Akt/PKB pathway (negative regulator of eEF2K activity) and lowering the expression of T (BRACHYURY, mouse homolog) [26] suggesting that the antiproliferative effects of afatinib could be associated with re-activation of eEF2K as seen in the colon carcinoma in vitro and in vivo models [26,27,28,29]. This is contrasting to the data obtained from patient tumors and overall survival in TNBC, pancreatic, lung carcinoma and glioblastoma in the TGCA database as well as the experimental data from the in vitro and in vivo models of these cancer types where reduction in eEF2K expression results in decreased tumorigenicity [25].
To uncover the contributory role of eEF2K to the oncogenic processes in chordoma cells, we took a gene silencing approach where lowering eEF2K protein levels promoted an increase in cellular proliferation. Based on this observation and the commonly known fact that the failure of conventional chemotherapeutics in the treatment of chordoma that target DNA synthesis can be attributed to the slow proliferation kinetics of these cells, we hypothesized whether increased proliferation of chordoma cells in a eEF2K-silenced background could potentiate the standard chemotherapeutics.
In other words, it is worthwhile to address whether knock down of eEFK2 in combination with conventional therapeutics would result in improved efficacy of standard of care agents that elicit cell death through massive DNA damage.
Materials and methods
Cell culture
Chordoma cell lines, MUG-Chor1 and UM-Chor1 [30], were kindly gifted from the Chordoma Foundation and they were routinely checked for mycoplasma contamination and STR analysis. The cell lines were grown in glatin coated flasks with culture medium containing IMDM (Gibco cat no: 31980–022, ThermoScientific, USA), RPMI (11,875–093, ThermoSecientific, USA), 10% FBS and 1% Penicillin–Streptomycin-Anphoteracin (PSA).
Silencing of eEF2K expression via siRNAs)
Chordoma cells were transfected either with eEF2K-specific siRNA (4,392,420 Thermo Scientific) or negative control siRNA (AM4611 Thermo Scientific) using the liposomal-based carrier “Lipofectamine RNAi Max Transfection Reagent” (13,778,100, Thermo Scientific) according to the manufacturer’s instructions. Briefly, cells were grown in 6-well plates prior to the transfection and siRNAs –diluted in Opti-MEM (31,985,062, ThermoScientific, USA) with no serum and mixed with 3 µL Lipofectamine™ RNAiMAX- were introduced into the cells with a final concentration of 50 nM. Following a 36 h incubation with the siRNA coctails, cells were returned to fresh medium to be harvested at 72 h post-transfection.
Gene expression analysis and protein quantification
The silencing of eEF2K expression was investigated both at the mRNA and protein level on the 3rd day following siRNA transfection. GAPDH was used as an internal control. Total RNA was isolated using Trizol reagent (15,596,018, ThermoScietific, USA) according to the manfacturer’s protocol. Complementary DNA was synthesized from 1000 ng RNA with the “High-Capacity cDNA Reverse Transcription Kit”: (4,368,814, Thermo Fisher) in accordance with the manufacturer’s protocol. Taqman primer probes targeting eEF2K (Hs00179434_m1,) and GAPDH (Hs02786624_g1), and Universal Taqman Master Mix (4,440,038, Applied Biosystem) were purchased from Thermo Fisher to run the realtime PCR according to the manufacturer’s protocol. GAPDH was used as an internal control to normalize the results. As for the analysis 2−ΔΔCt method was used to find difference in fold change.
Proteins were isolated from chordoma cell lines, which were transfected with siRNAs, by using a RIPA buffer solution (9806, CST, USA). Briefly, upon transfection, cell pellets were lysed with RIPA buffer containing protease and phosphatase inhibitor cocktail (78,440, Thermo Fisher) on ice, and centrifuged for 30 min at 14,000 g. Protein concentration was determined by Pierce™ BCA Protein Assay Kit” (23,225, Thermo Fisher, USA). Proteins were loaded onto polyacrylamide gels, transferred onto the nitrocellulose membrane, blocked with skim milk, incubated with eEEF2K (ab45168, Abcam, USA) and GAPDH (5174, Cell Signaling Technology) antibodies (diluted in 1:1000), and finally visualized with a chemiluminescent solution containing hydrogen peroxide in the Bio Rad Chemidoc imaging system. Band lengths were normalized to GAPDH by the software and protein amounts were determined.
Proliferation assay
The effects of eEF2K silencing on the viability of chordoma cells were investigated on the 3rd day with a 3-(4,5-Dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium (MTS), viability assay (G3581, Promega, USA) which is based on the breakdown of tetrazolium salts into formazan crystals through the mitochondrial activity. Briefly, two days after the transfection, cells were seeded onto the four 96-well plates (CLS6509, Corning, USA) at a density of 5 × 103/well and treated with 10% MTS reagent and incubated in regular culture conditions for 1 h in dark. Absorbance values of each well were determined by a plate reader (ELx800, Biotek Instruments, USA) at wavelength 490 nm. Results were compared with negative controls.
Apoptosis assay
Early apoptotic cells were detected using the FITC Annexin V Apoptosis Detection Kit I (556,547, BD Biosciences, USA) according to manufacturer’s protocol. Briefly, cells were labeled with Propium Iodide (PI) and annexin V, which is an early apoptotic marker, and the resultant intensity of fluorescence was measured by using the BD Facs Calibur instrument [31].
Cell cycle analysis
Upon inhibition of eEF2K in cells with siRNA, the cell cycle profile in cells was determined using the BD Facs Calibur device using a previously established protocol [32]. This analysis was performed 3 days after siRNA transfection. The cells were harvested and fixed in cold in 70% ethanol at least 2 h prior to the analysis. The cells were then incubated with a solution containing 0.3 mg/ml RNAse A and 5 µg/ml PI and incubated at 37 °C for 30 min. The cell cycle analysis was performed by measuring the fluorescence intensity of DNA using the BD Facs Calibur device.
Invasion and migration assays
Chordoma cell lines transfected with siRNA were trypsinized and seeded at 2 × 105 cells by dissolving in 200 µl serum-free chordoma medium into the transwells, which was then placed in wells of a-24 well plate containing 1300 µL of regular chordoma medium and kept at 37 °C for 24 h. At the end of the incubation, cells were processed for the staining protocol applied in the invasion assay (described below). For the invasion assay, cells were seeded at 2 × 105 cells in matrigel-coated transwells (354,480, Corning, USA) according to the manufacturer’s protocol. After 24 h of incubation, the cells were fixed with 3.7% formaldehyde, permeabilized with 100% methanol, and stained with 0.1% Crystal Violet dye. The cells stained with the dye on the reverse side of the transwells were considered as the migratory and invasive cells, and compared to the unstained counterparts for the analysis.
Combinatorial treatment of eEF2K siRNA with chemotherapeutic agents
After the transfection with siRNAs, MUG-Chor1 and UM-Chor1 cells were treated for 72 h with conventional chemo agents such as Etoposide and Cisplatin (each at 10 µM as was used previously [33] to which they have resistance [34, 35]. The viability of the cells was determined by the “CellTiter 96® AQueous One Solution Cell Proliferation Assay (MTS)” according to the manufacturer's protocol described before. The combined treatment of cells silenced with eEF2K and the chemotherapeutics were compared to the negative counterparts. The percentages of the viability were determined the MTS assay.
Statistical analysis
Statistical analysis was performed with GraphPad Prism 5 (GraphPad Software, La Jolla, CA) and comparisons were made using unpaired t-test, post-test, and post hoc test where applicable two-way analysis of variance (ANOVA) and Bonferroni's test. A P value of < 0.05 was considered statistically significant.
Results
Transient silencing of the eEF2K in cells and the its confirmation
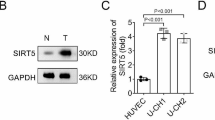
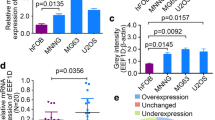
To verify that the endogenous levels of eEF2K in the UM-Chor1 and MUG-Chor1 chordoma cell lines are reduced with the gene silencing methodology used in the study, both total RNA and protein samples were prepared. Changes in gene expression both at the mRNA and protein levels were analyzed with qPCR and Western Blot Analysis. Interestingly, although MUG-Chor1 cells appeared to have higher remainder eEF2K mRNA compared to UM-Chor1 cell line under post-eEF2K-siRNA treatment conditions, the magnitude of reduction in eEF2K protein levels appear to be comparable in both MUG-Chor1 and UM-Chor1 cells. When MUG-Chor1 and UM-Chor1 cells were transiently transfected with siRNA against eEF2K, a significant decrease both in gene expression (Fig. 1A, B) and protein levels of eEF2K (Fig. 1C) was observed, which was accompanied with a reduction in its endogenous phosphorlyation target of eEF2 as indicated by the decrease in the signal from phospho-T56-eEF2 antibody (Supplemental figure). Differences in eEF2K amounts are calculated by their normalization to the GAPDH levels in each sample.
The effects of decrease in eEF2K levels on cell proliferation and cell cycle
To address the impact of reduced eEF2K expression on cell proliferation and distribution of cell cycle populations of chordoma cell lines, we used both an MTS-based assay and cell cycle analysis. The diminished levels of eEF2K did not significantly affect the cellular division rate in both cell lines (Fig. 2A, B). However, silencing of eEF2K, caused a mild increase in the number of cells in S and G2/M phases in MUG-Chor1 cells (Fig. 2C). On the other hand, there was not a substantial alteration in the phases of the cell cycle in UM-Chor1 cells upon eEF2K silencing (Fig. 2D).
The impacts of silencing eEF2K on cellular death
Next, we addressed whether silencing of eEF2K had any impact on cell death. To measure apoptosis Annexin V staining was performed on cells treated with either control or eEF2K siRNA. The data indicate that –albeit with minor differences- the silencing of eEF2K resulted in differential apoptotic or necrotic responses in MUG-Chor1 (Fig. 3A) and UM-Chor1 cells (Fig. 3B). For example, UMChor1 cells show a slightly higher percentage of living cells in both groups (control and sieEF2K) than the same conditions in MUG-Chor1 cells (Fig. 3A and B) and lower percentage of late and early apoptotic populations. The lower number of viable cells in the MUG-Chor1 cells is accompanied with slightly higher percentages of sub-apoptotic and necrotic populations. Although reasons for this difference is not investigated in-depth in this study, the mutational burden and variation in the genomic background between the two cell lines that are harvested from tumors found in different locations of the body could account for the discrepancies seen in death profiles.
The changes in the migration and invasion capacity of UM-Chor1 and MUG-Chor1 cells upon eEF2K silencing
To measure the potential changes in the migration and invasiveness of the two cell lines, control siRNA or eEF2K-siRNA treated cells were seeded in transwell plates and the number of cells that migrate across were determined through absobance readings in a Gimsa Stain-based protocol. Our findings indicate that the reduction of eEF2K expression was associated with a decrease in the number of cells that migrated across the transwell. For example, when normalized to the control cells (no eEF2K silencing) only 60% of both MUG-Chor1 (Fig. 4A) and UM-Chor1 (Fig. 4B) cells were able to migrate through the transwells. Similar results were also obtained in the invasion assay, where control or eEF2K-siRNA treated cells were allowed to migrate across a matrigel layer in same transwell setting. Accordingly, around 70% of MUG-Chor1 and 50% of UM-Chor1 (Fig. 4C and D) cells invaded the matrigel coated transmembrane in presence of eEF2K suppression.
Investigation of the changes in drug-resistance in response to eEF2K silencing
Transient administration of eEF2K siRNAs in both cell lines, chemosensitized these cells to both Etoposide and Cisplatin partially. However, the resultant reduction in cell viability obtained via the combination treatment did not exceed 50%. MUG-Chor1 cells that are silenced for eEF2K were treated with 10 µM Cisplatin and Etoposide. Nonetheless, lower percentage of cells were viable in groups co-treated with Cisplatin compared to those co-treated with Etoposide suggesting a more significant therapeutic outcome in the Cisplatin and eEF2K siRNA dual treatment (Fig. 5A). In contrast, response of UM-Chor1 to the identical treatment schemes indicated a higher percentage of viability when compared to the MUG-Chor1 cells (Fig. 5B). All together, the combination of eEF2K silencing and chemotherapeutics caused the viability to stay above 50%; therefore, this therapy regime is not considerably detrimental on both cells.
Discussion
The small interference-RNAs (si-RNAs) based gene silencing has become a promising tool in developing new methods in regard to elucidating the molecular mechanisms that underlie many diseases including cancer [36]. si-RNAs are essential tools in decipfering the function of particular genes as it promotes silencing by blocking translation of mRNAs into proteins [37]. In addition to being used as a tool for understanding the gene function, in recent years siRNAs have emerged as novel drug cancdidates due to their ability to silence expression of those genes involved in oncogenesis. In our study, physiological consequences of eEF2K silencing on malignant properties of chordoma cells were investigated via transient transfection. To our knowledge, this is the first study that addresses the link between chordoma malignancy and downregulation of eEF2K expression which has been reported to exert pro-tumorigenic effects both in vitro and in vivo models of breast, ovarian, lung, glioma, meduloblastoma, hepatocellular carcinoma, pancreatic cancer and prostate cancer [38]. Suppression of eEF2K expression in the slow-growing chordoma cell line model rather hindered migratory and invasiveness in terms of malignant characteristics, while having a mior effect on the proliferative properties.
eEF2K is an atypical Ca2 + /calmodulin-linked protein kinase and negatively regulates mRNA translation by phosphorylating the eEF2 protein that bocks its transocase activity [39]. Various stress stimuli such as nutrient depletion, hypoxia, energy crisis, and viral infections force the cells to adapt their new environment trigger eEF2K catalytic activity toward eEF2 to turn off translation which claims more 90% of the energy dedicated to protein synthesis [40]. It is known that the levels of eEF2K is increased in various rapidly dividing tumor cells and attenuation of eEF2K expression through gene silencing methodologies potentiatiates cancer cell death induced by chemotherapeutics via various cellular death mechanisms [25]. eE2FK is overexpressed in many metastatic cancer types and has been associated with poor prognosis [17, 41]. A recent study highlighted the importance of eEF2K as a metastatic as well as prognostic biomarker in people diagnosed with gastric cancer [42]. A similar study done by Xie et al. showed that si-RNA-mediated eEF2K silencing in human lung cancer cells suppressed the tumor growth and metastasis [43]. Silencing eEF2K reduced the number of proliferating esophageal squamous cell carcinoma cells [34]. In the same study Zhu and colleagues also showed that as a result of suppression of eEF2K in cells, there was a decrease in the number of cells in the S and M phases [44]. In contrast, our findings demonstrated that the proliferation rate of chordoma cells did not decrease upon the suppression of eEF2K, which -in fact- resulted in a mild increase in cell division. The cell cycle profiles for both lines were found to be similar except for a slight increase in S and G2/M phases in MUG-Chor1 cells.
As opposed to previous findings which suggest the inhibition of eEF2K expression in reducing the proliferation rate [45], other studies provide evidence that unleashing eEF2K activity via removal of the inhibitory upstream signaling exerts antitumorigenic effects including breast [46], intestinal, colorectal and lung cancer models [27, 47]. Particularly, translation-independent antitumorigenic effects of eEF2K attenuation detected in the non-small cell lung cancer model are interesting in the sense that an inhibitory phosphorylation of PKM2 directly by eEF2K ultimately results in suppression of glycolysis concomitant with STAT3 dependent transcription and decrease in c-Myc expression [48]. In contrast, high eEF2K expression is associated with poor patient survial and a more efficient siRNA-based silencing eEF2K in a panel of lung carcinoma cell lines resulted in a drastic suppression of colony formation, invasion and tumor volume in an earlier study [49]. Hence, it is worthwhile to understand whether differences in the gene silencing efficiency could account for the dual contrasting roles eEF2K in cancer.
The role of eEF2K in protecting cells with more proliferative capacity including stem cells was demonstrated in a study done by Liao et al. [50]. The authors claimed that in the absence of eEF2K expression in knock-out mice were protected against low dose ionizing radiation. In the same study, they also showed the reduction in death of stem cells of the bone marrow in the eEF2K knock out mice. The authors further explained that at lower doses of ionizing radiation (IR), the absence of eEF2K allows cells escape from (G2/M) checkpoint in the cell cycle accounting for the increased survival phenotype of the animals; however, at higher doses of IR eEF2K deficiency of the knock-out cells increases their susceptibility to undergo higher degree of mitotic catastrophe. This would explain why at least one type of cells are found arrested in S and G2/M phases of the cell cycle.
Therapeutics that target eEF2K are continued to be developed and become of great interest in the treatment of triple negative breast as therapeutic effect of eEF2K knock down was demonstrated both in cancer cell lines [51] as well as in in vivo models [52]. Silencing eEF2K increases the sensitivity against lapatinib in human nasopharyngeal carcinoma cells [53], and further sensitizes human glioma cells to agents such as tumor necrosis factor-related apoptosis-inducing ligand (TRAIL) and temozolomide [24]. Combined therapy of an eEF2K inhibitor –mitoxantrone- and mTOR targeting increased the efficiency of treatment method against breast cancer [54]. mTOR is highly expressed in chordoma cells and is a well known biomarker [55]. Targeting mTOR is not only promising for other cancer types, but also for chordoma [56]. Peng et al. showed that targeting mTOR sensitized the cells to Cisplatin [57]. Zhang et al. showed that silencing of eEF2K increased the sensitivity of hepatocellular carcinoma cells to Cisplatin [58]. In our study, MUG-Chor1 cells responded to Cisplatin better than they did to Etoposide. The cell lines used in this study originate from different parts of the spine and therefore show varying characteristics in their response as well as drug resistance to chemotherapy. The decrease in eEF2K levels might have increased the rate of proliferation and sensitizing the cells to chemotherapy considerably. This showed us that drug resistance in chordoma cells partially attenuated when silenced with eEF2K siRNAs. As suggested by our group that there is a population of cancer stem cells in chordoma [59] and hence they might be deprived of the protecting role of eEF2K due to the silencing [50]. Therefore, the partial death observed in the chordoma cells upon treatment with chemotherapeutics in presence of silencing eEF2K could be due to losing its protective role.
According to our findings, downregulation of eEF2K reduced the metastatic capacity of chordoma cells. A recent study suggests that targeting eEF2K could be a promising therapeutical approach in hindering melanoma progression [60].
Other studies show that there is an association between mTOR expression and migration [61]. Considering that both mTOR and eEF2K are highly expressed in chordoma and the findings of Guan and colleagues in the breast carcinoma model, it would be of interest to test whether inhibition of eEF2K could help potentiating the targeting mTOR pathway in the treatment of this cancer type in a future study. Nonetheless, based on the findings of the current work, we can conclude that chordoma cells may have lost their migratory capacity significantly due to the silencing of eEF2K.
Angiogenesis is the process in which newly formed tumor cells, become invasive and migrate into the other parts of the body. It is found that eEF2K induces an increase in the VEGF expression and, thereby, ultimately contributes to angiogenesis. A chain of events occur upon inhibition of eEF2K including a decrease in PI3K/Akt/STAT3 pathway, which is highly expressed in chordoma [11]. This might explain the reduced migratory capacity of chordoma cell when eEF2K is silenced. Ashour et al. showed that silencing of eEF2K in pancreatic cancer cells suppressed EMT and reduced metastasis and invasion [45]. Previously we have addressed genes that regulate the invasiveness of chordoma cells and reported that TWIST -a prominent marker of Epithelial Mesenchymal Transition (EMT)- also underlies invasiveness of chordoma [62]. In a similar study, Xie et al. delineated the molecular switches that control the metastatic capabilities of breast and lung cancer cells through suppression of eEF2K suggesting EMT-modulatory roles of eEF2K loss [43]. Therefore, we investigated the impact of eEF2K silencing on the metastasis and invasion capacities of the chordoma cells in the current study. Similar to the results in the lung [38] and breast cancer study [25], our data also point out that upon eEF2K suppression, a significant reduction in metastasis and invasion capabilities both MUG-Chor1 and UM-Chor1 cells is observed.
Chordoma is a rare type of bone tumor and yet classiffied as often presenting with malignant properties, including recurrence and ability for metastasis. Chordoma is also highly resistant to conventional drug-based treatment methods and therefore requires new strategies that can improves the clinical outcome. Proliferation of the cells are largely depend on the proper protein synthesis, but under suboptimal conditions, proliferation needs to be blocked until the environmental conditions go back to normal. eEF2K is an enzyme that negatively regulates the protein synthesis by phosphorylating eEF2 and causes an inhibition. When cells are exposed to energy depletion eEF2K becomes activated and trigger a reduction in overall translational rates. This mechanism is also adapted by the tumor cells as they are faced with various stress conditions including nutrient deprivation, high acidity and low oxygen levels in their poorly vascularized microenviroment, which account for these harsh living conditions as a consequence of exponential growth in the tumor mass. Chordoma cells express high eEF2K levels that might explain the slow proliferating nature of the cells. By inhibiting eEF2K we were able to force the cells to become more sensitive Cisplatin and Etoposide. Upon eEF2K depletion the chordoma cells significantly lost the metastatic and invasive capacities, which holds a promise for future preclinical studies and clinical trials.
Abbreviations
- eEF2:
-
Eukaryotic Elongation Factor 2
- eEF2K:
-
Eukaryotic Elongation Factor 2 Kinase
- siRNAs:
-
Silencing RNAs
- PI3K:
-
Phosphoinositide 3-kinase
- mTOR:
-
Mammalian target of rapamycin-rapamycin in mammals
- MAPK:
-
Mitogen-activated protein kinase
- AMPK:
-
Adenosine monophosphate-activated protein kinase
- STR:
-
Short tandem repeats
- TRAIL:
-
TNF-related apoptosis-inducing ligand
- TNF:
-
Tumor necrosis factor
- ESCC:
-
Esophageal squamous cell carcinoma
- EGFR:
-
Epidermal Growth Factor Receptor
- RPMI:
-
Roswell Park Memorial Institute
- PBK:
-
Protein Kinase B
- GAPDH:
-
Glyceraldehyde 3-phosphate dehydrogenase
- MTS:
-
3-(4,5-Dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium
- FITC:
-
Fluorescein isothiocyanate
- PI:
-
Propium Iodide
- NC:
-
Negative Control
- PKM2:
-
Pyruvate kinase isozymes M1/M2
- c-My:
-
C-myelocytomatosis
- Stat3:
-
Signal transducer and activator of transcription 3
- EMT:
-
Epithelial Mesenchymal Transition
- Twist1:
-
Twist Basic Helix-Loop-Helix Transcription Factor 1
References
Chugh R, Tawbi H, Lucas DR et al (2007) Chordoma: the nonsarcoma primary bone tumor. Oncologist 12:1344–1350. https://doi.org/10.1634/theoncologist.12-11-1344
Bell D, Raza SM, Bell AH et al (2016) Whole-transcriptome analysis of chordoma of the skull base. Virchows Arch 469:439–449. https://doi.org/10.1007/s00428-016-1985-y
Chambers KJ, Lin DT, Meier J et al (2014) Incidence and survival patterns of cranial chordoma in the United States. Laryngoscope 124:1097–1102. https://doi.org/10.1002/lary.24420
Campbell RG, Prevedello DM, Ditzel Filho L et al (2015) Contemporary management of clival chordomas. Curr Opin Otolaryngol Head Neck Surg 23:153–161. https://doi.org/10.1097/MOO.0000000000000140
Littman L, Reviews N (1856) Fletcher,14 Mabrey,28. 80–90
Healey JH, Lane JM (1989) Chordoma: a critical review of diagnosis and treatment. Orthop Clin North Am 20:417–426
McMaster ML, Goldstein AM, Bromley CM et al (2001) Chordoma: incidence and survival patterns in the United States, 1973–1995. Cancer Causes Contr 12:1–11. https://doi.org/10.1023/a:1008947301735
Vujovic S, Henderson S, Presneau N et al (2006) Brachyury, a crucial regulator of notochordal development, is a novel biomarker for chordomas. J Pathol 209:157–165. https://doi.org/10.1002/path.1969
Bailey CS, Fisher CG, Boyd MC, Dvorak MFS (2006) En bloc marginal excision of a multilevel cervical chordoma. Case report J Neurosurg Spine 4:409–414. https://doi.org/10.3171/spi.2006.4.5.409
Carrabba G, Dehdashti AR, Gentili F (2008) Surgery for clival lesions: open resection versus the expanded endoscopic endonasal approach. Neurosurg Focus 25:E7. https://doi.org/10.3171/FOC.2008.25.12.E7
Gulluoglu S, Turksoy O, Kuskucu A et al (2016) The molecular aspects of chordoma. Neurosurg Rev. https://doi.org/10.1007/s10143-015-0663-x
Brüderlein S, Sommer JB, Meltzer PS et al (2010) Molecular characterization of putative chordoma cell lines. Sarcoma. https://doi.org/10.1155/2010/630129
Hsu W, Mohyeldin A, Shah SR et al (2011) Generation of chordoma cell line JHC7 and the identification of Brachyury as a novel molecular target. J Neurosurg 115:760–769. https://doi.org/10.3171/2011.5.JNS11185
Rinner B, Froehlich EV, Buerger K et al (2012) Establishment and detailed functional and molecular genetic characterisation of a novel sacral chordoma cell line, MUG-Chor1. Int J Oncol 40:443–451. https://doi.org/10.3892/ijo.2011.1235
Ballard DJ, Peng H-Y, Das JK et al (2021) Insights ınto the pathologic roles and regulation of eukaryotic elongation factor-2 kinase. Front Mol Biosci. https://doi.org/10.3389/fmolb.2021.727863
Bhat M, Robichaud N, Hulea L et al (2015) Targeting the translation machinery in cancer. Nat Rev Drug Discov 14:261–278. https://doi.org/10.1038/nrd4505
Wang X, Xie J, Proud CG (2017) Eukaryotic Elongation factor 2 kinase (eEF2K) in cancer. Cancers (Basel). https://doi.org/10.3390/cancers9120162
Wang X, Regufe da Mota S, Liu R et al (2014) Eukaryotic elongation factor 2 kinase activity is controlled by multiple inputs from oncogenic signaling. Mol Cell Biol 34:4088–4103. https://doi.org/10.1128/MCB.01035-14
Temme L, Asquith CRM (2021) eEF2K: an atypical kinase target for cancer. Nat Rev Drug Discov 20:577. https://doi.org/10.1038/d41573-021-00124-5
Leprivier G, Rotblat B, Khan D et al (2015) Stress-mediated translational control in cancer cells. Biochim Biophys Acta 1849:845–860. https://doi.org/10.1016/j.bbagrm.2014.11.002
Silvera D, Formenti SC, Schneider RJ (2010) Translational control in cancer. Nat Rev Cancer 10:254–266. https://doi.org/10.1038/nrc2824
Roux PP, Topisirovic I (2012) Regulation of mRNA translation by signaling pathways. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a012252
Mendoza MC, Er EE, Blenis J (2011) The Ras-ERK and PI3K-mTOR pathways: cross-talk and compensation. Trends Biochem Sci 36:320–328. https://doi.org/10.1016/j.tibs.2011.03.006
Zhang Y, Cheng Y, Zhang L et al (2011) Inhibition of eEF-2 kinase sensitizes human glioma cells to TRAIL and down-regulates Bcl-xL expression. Biochem Biophys Res Commun 414:129–134. https://doi.org/10.1016/j.bbrc.2011.09.038
Tekedereli I, Alpay SN, Tavares CDJ et al (2012) Targeted silencing of elongation factor 2 kinase suppresses growth and sensitizes tumors to doxorubicin in an orthotopic model of breast cancer. PLoS ONE. https://doi.org/10.1371/journal.pone.0041171
Ng TH, Sham KWY, Xie CM et al (2019) Eukaryotic elongation factor-2 kinase expression is an independent prognostic factor in colorectal cancer. BMC Cancer 19:649. https://doi.org/10.1186/s12885-019-5873-0
Xie C-M, Liu X-Y, Sham KWY et al (2014) Silencing of EEF2K (eukaryotic elongation factor-2 kinase) reveals AMPK-ULK1-dependent autophagy in colon cancer cells. Autophagy 10:1495–1508. https://doi.org/10.4161/auto.29164
De Gassart A, Demaria O, Panes R et al (2016) Pharmacological eEF2K activation promotes cell death and inhibits cancer progression. EMBO Rep. https://doi.org/10.15252/embr.201642194
Liu R, Proud CG (2016) Eukaryotic elongation factor 2 kinase as a drug target in cancer, and in cardiovascular and neurodegenerative diseases. Acta Pharmacol Sin 37:285–294. https://doi.org/10.1038/aps.2015.123
Owen JH, Komarck CM, Wang AC et al (2018) UM-Chor1: establishment and characterization of the first validated clival chordoma cell line. J Neurosurg 128:701–709. https://doi.org/10.3171/2016.10.JNS16877
Jiang S-X, Qi B, Yao W-J et al (2017) Berberine displays antitumor activity in esophageal cancer cells in vitro. World J Gastroenterol 23:2511–2518. https://doi.org/10.3748/wjg.v23.i14.2511
Irazoqui AP, Gonzalez A, Buitrago C (2022) Effects of calcitriol on the cell cycle of rhabdomyosarcoma cells. J Steroid Biochem Mol Biol. https://doi.org/10.1016/j.jsbmb.2022.106146
Bayrak OF, Aydemir E, Gulluoglu S et al (2011) The effects of chemotherapeutic agents on differentiated chordoma cells. J Neurosurg Spine 15:620–624. https://doi.org/10.3171/2011.7.SPINE10798
Baldi GG, Lo Vullo S, Grignani G et al (2022) Weekly cisplatin with or without imatinib in advanced chordoma: a retrospective case-series analysis from the Italian rare cancers network. Cancer 128:1439–1448. https://doi.org/10.1002/cncr.34083
Yaniv D, Soudry E, Strenov Y et al (2020) Skull base chordomas review of current treatment paradigms. World J Otorhinolaryngol - head neck Surg 6:125–131. https://doi.org/10.1016/j.wjorl.2020.01.008
Ozcan G, Ozpolat B, Coleman RL et al (2015) Preclinical and clinical development of siRNA-based therapeutics. Adv Drug Deliv Rev 87:108–119. https://doi.org/10.1016/j.addr.2015.01.007
Hannon GJ, Rossi JJ (2004) Unlocking the potential of the human genome with RNA interference. Nature 431:371–378. https://doi.org/10.1038/nature02870
Zhang B, Zou J, Zhang Q et al (2021) Progress in the development of eukaryotic elongation factor 2 kinase (eEF2K) natural product and synthetic small molecule ınhibitors for cancer chemotherapy. Int J Mol Sci. https://doi.org/10.3390/ijms22052408
Carlberg U, Nilsson A, Nygård O (1990) Functional properties of phosphorylated elongation factor 2. Eur J Biochem 191:639–645. https://doi.org/10.1111/j.1432-1033.1990.tb19169.x
Wang X, Li W, Williams M et al (2001) Regulation of elongation factor 2 kinase by p90(RSK1) and p70 S6 kinase. EMBO J 20:4370–4379. https://doi.org/10.1093/emboj/20.16.4370
Pott LL, Hagemann S, Reis H et al (2017) Eukaryotic elongation factor 2 is a prognostic marker and its kinase a potential therapeutic target in HCC. Oncotarget. https://doi.org/10.18632/oncotarget.14447
Jiang M, Qi L, Jin K et al (2021) eEF2K as a novel metastatic and prognostic biomarker in gastric cancer patients. Pathol - Res Pract. https://doi.org/10.1016/j.prp.2021.153568
Xie J, Shen K, Lenchine RV et al (2018) Eukaryotic elongation factor 2 kinase upregulates the expression of proteins implicated in cell migration and cancer cell metastasis. Int J cancer 142:1865–1877. https://doi.org/10.1002/ijc.31210
Zhu H, Song H, Chen G et al (2017) eEF2K promotes progression and radioresistance of esophageal squamous cell carcinoma. Radiother Oncol J Eur Soc Ther Radiol Oncol 124:439–447. https://doi.org/10.1016/j.radonc.2017.04.001
Ashour AA, Abdel-Aziz A-AH, Mansour AM et al (2014) Targeting elongation factor-2 kinase (eEF-2K) induces apoptosis in human pancreatic cancer cells. Apoptosis 19:241–258. https://doi.org/10.1007/s10495-013-0927-2
Cheng Y, Ren X, Zhang Y et al (2013) Integrated regulation of autophagy and apoptosis by EEF2K controls cellular fate and modulates the efficacy of curcumin and velcade against tumor cells. Autophagy 9:208–219. https://doi.org/10.4161/auto.22801
Faller WJ, Jackson TJ, Knight JR et al (2015) mTORC1-mediated translational elongation limits intestinal tumour initiation and growth. Nature 517:497–500. https://doi.org/10.1038/nature13896
Xiao M, Xie J, Wu Y et al (2020) The eEF2 kinase-induced STAT3 inactivation inhibits lung cancer cell proliferation by phosphorylation of PKM2. Cell Commun Signal 18:25. https://doi.org/10.1186/s12964-020-0528-y
Bircan HA, Gurbuz N, Pataer A et al (2018) Elongation factor-2 kinase (eEF-2K) expression is associated with poor patient survival and promotes proliferation, invasion and tumor growth of lung cancer. Lung Cancer 124:31–39. https://doi.org/10.1016/j.lungcan.2018.07.027
Liao Y, Chu H-P, Hu Z et al (2016) Paradoxical roles of elongation factor-2 kinase in stem cell survival*. J Biol Chem. https://doi.org/10.1074/jbc.M116.724856
Comert Onder F, Siyah P, Durdagi S et al (2022) Novel etodolac derivatives as eukaryotic elongation factor 2 kinase (eEF2K) inhibitors for targeted cancer therapy. RSC Med Chem 13:840–849. https://doi.org/10.1039/d2md00105e
Chen X, Wang K, Jiang S et al (2022) eEF2K promotes PD-L1 stabilization through inactivating GSK3β in melanoma. J Immunother Cancer. https://doi.org/10.1136/jitc-2021-004026
Liu L, Huang P, Wang Z et al (2016) Inhibition of eEF-2 kinase sensitizes human nasopharyngeal carcinoma cells to lapatinib-induced apoptosis through the Src and Erk pathways. BMC Cancer 16:813. https://doi.org/10.1186/s12885-016-2853-5
Guan Y, Jiang S, Ye W et al (2020) Combined treatment of mitoxantrone sensitizes breast cancer cells to rapalogs through blocking eEF-2K-mediated activation of Akt and autophagy. Cell Death Dis 11:948. https://doi.org/10.1038/s41419-020-03153-x
Presneau N, Shalaby A, Idowu B et al (2009) Potential therapeutic targets for chordoma: PI3K/AKT/TSC1/TSC2/mTOR pathway. Br J Cancer 100:1406–1414. https://doi.org/10.1038/sj.bjc.6605019
Stacchiotti S, Marrari A, Tamborini E et al (2009) Response to imatinib plus sirolimus in advanced chordoma. Ann Oncol 20:1886–1894. https://doi.org/10.1093/annonc/mdp210
Peng D-J, Wang J, Zhou J-Y, Wu GS (2010) Role of the Akt/mTOR survival pathway in cisplatin resistance in ovarian cancer cells. Biochem Biophys Res Commun 394:600–605. https://doi.org/10.1016/j.bbrc.2010.03.029
Zhang C, Lei J-L, Zhang H et al (2017) Calyxin Y sensitizes cisplatin-sensitive and resistant hepatocellular carcinoma cells to cisplatin through apoptotic and autophagic cell death via SCF βTrCP-mediated eEF2K degradation. Oncotarget. https://doi.org/10.18632/oncotarget.19883
Aydemir E, Bayrak OF, Sahin F et al (2012) Characterization of cancer stem-like cells in chordoma. J Neurosurg 116:810–820. https://doi.org/10.3171/2011.12.JNS11430
Deng G, Zeng F, He Y et al (2022) EEF2K silencing inhibits tumour progression through repressing SPP1 and synergises with BET inhibitors in melanoma. Clin Transl Med. https://doi.org/10.1002/ctm2.722
Zhou H, Huang S (2011) Role of mTOR signaling in tumor cell motility, invasion and metastasis. Curr Protein Pept Sci 12:30–42. https://doi.org/10.2174/138920311795659407
Aydemir E, Kaşikci E, Coşkunçelebi B et al (2018) The effect of TWIST silencing in metastatic chordoma cells. Turkish J Biol. https://doi.org/10.3906/biy-1801-17
Acknowledgements
We would like to thank Fikrettin Sahin for kindly providing the chemicals and laboratory infrastructure to accomplish the study.
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have not disclosed any competing interests.
Ethical approval
This article does not contain any studies with human participants and/or animal participants performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Aydemir, E., Tüysüz, E.C., Bayrak, Ö.F. et al. Impact of silencing eEF2K expression on the malignant properties of chordoma. Mol Biol Rep 50, 3011–3022 (2023). https://doi.org/10.1007/s11033-023-08257-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-023-08257-z