Abstract
Purpose
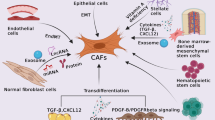
Cancer-associated fibroblasts (CAFs) are major components of tumor microenvironment that stimulate ESCC and GC progression. The LncRNA-CAF, FLJ22447, is located in the vicinity of HIF1A, while their association remains unclear. This study aims to assess the FLJ22447 expression in the ESCC and GC patients and evaluate its association with the HIF1A gene.
Methods
Fresh ESCC and GC tumor samples and their adjacent non-tumor tissues were collected from patients who underwent surgery in Imam Khomeini Hospital, Tehran, Iran. The expression of FLJ22447, HIF1A, and VEGF was evaluated using qRT-PCR test. The association of their expression with tumor clinicopathological features in ESCC patients was assessed. System biology tools were then applied for the possible biological subsequences of the FLJ22447.
Results
A significant reduction in FLJ22447 expression was observed in ESCC and GC tissues than adjacent non-tumor tissues, while, the expression of HIF1A and VEGF were increased. Low expression of FLJ22447 was significantly correlated with HIF1A (P = 2.4e–73, R = 0.63) and VEGF (P = 0.00019, R = 0.15) expression. A significant relationship was detected between the high expression of HIF1A and tumor stages (I–II) and it was related to the reduced survival of ESCC patients. Conversely, increased VEGF expression was linked to the advanced stages (III–IV) and metastasis in ESCC. The analysis of FLJ22447-interacted proteins showed that MYC, JUN, SMRCA4, PPARG, AR, FOS, and CEBPA are the hub genes. These proteins were implicated in the cancer related pathways. Among them, SPI1, E2F1, TCF7L2, and STAT1 were significantly expressed in esophageal and gastric cancers that were functionally involved in the proliferation, apoptosis, and angiogenesis pathways in cancer.
Conclusion
The results suggested that FLJ22447 may have a regulatory function on the HIF1A expression. We identified the FLJ22447-interacted proteins and their molecular function in cancer pathogenesis. Further research emphasis is to realize the association of FLJ22447 with its protein partners in progression of cancer. These may provide an insight into the FLJ22447 activity that could introduce it as a potential value in tumor gene therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer disease is characterized by the genomic abnormalities which threaten the people’s health worldwide. It has become the second leading cause of death after heart disease. Hypoxia and angiogenesis are known as common features of solid tumors [1, 2]. They are critical events for tumor cell growth and progression of cancer [3, 4]. Hypoxia is a key microenvironment condition that is modulated by transcription factors such as hypoxia-inducible factor 1 alpha (HIF1A). The HIF1A expression activates transcription of several genes that regulate proliferation of cell, metabolism of tumor cells, and angiogenesis processes [1, 5]. It is inhibited by the prolyl hydroxylase enzymes and the factor inhibiting HIF-1 oxygen sensors [6]. The HIF family members play an important role in the metabolism of cells in low oxygen condition. Previous research has documented the influence of HIF1A and HIF2A on the angiogenesis, cell metabolism, proliferation, and extracellular matrix remodeling [7].
Also, the HIF target genes influence all aspects of cancer biology including cell survival by TGF-β, bFGF, and p53 [8], metabolism via the GLUT-1 and glycolytic enzymes [9], and angiogenesis by the vascular endothelial growth factor (VEGF) [10]. In addition, the strong prognostic impact of the HIF1A and its association with the VEGF expression has been elucidated in pancreatic [11, 12] and breast [13] cancers.
Likewise, existing research recognizes the significant role played by the VEGF in angiogenesis in which promotes tumor growth and metastasis by blood vessels formation [14]. It is well known as an angiogenic factor that causes proliferation, migration, and permeability of endothelial cells. The high expression of VEGF causes to develop solid tumors by increasing the permeability of blood vessels and promoting angiogenesis [15]. The high expression of this gene also contributes to the other vascular diseases including neurodegeneration, retinal, and cancer [16].
Moreover, the impact of tumor microenvironment in cancer progression and metastasis has been demonstrated [17]. The cancer-associated fibroblasts (CAFs) are major components of tumor microenvironment that play a complex role in progression of cancer and are modulated by LncRNAs [18, 19]. Also, they can be formed upon stimulation of cytokines, growth factors, hypoxia, and non-coding RNAs [18]. The LncRNA-CAF, FLJ22447 (NR_039985), is recently identified in reprograming process of fibroblasts to mediate the growth of oral squamous cell carcinoma (OSCC) [19]. It is positioned on human chromosome 14q23 in the vicinity of the HIF1A gene. To date, the regulatory function of non-coding RNAs (ncRNA) in the expression of neighboring genes have been widely reported [17, 20, 21]. However, the association of LncRNA-CAF in the expression of HIF1A remains unclear. Additionally, research on the FLJ22447 has been delimited to the RNA-seq data, in which this ncRNA was highly expressed in testicular, lung, thyroid, skin, and prostate tissues, respectively [22]. This study, therefore, was designed to assess the FLJ22447 expression in the esophageal squamous cell carcinoma (ESCC) and gastric cancer (GC) patients and evaluate its association with the HIF1A and the VEGF genes.
Materials and methods
Clinical samples
This research was officially permitted by the Research Ethics committee (IR.GOUMS.REC.1397.006) to perform on the patients undergoing surgical resection without adjuvant therapy from 2016 to 2017. The informed consent forms were received from participants by the Ethical Committee of Tumor Bank of Cancer Institute, Imam Khomeini Hospital, Tehran, Iran. The tissue samples of esophageal cancer (n = 40) and gastric cancer (n = 20) were instantly frozen in liquid nitrogen after surgery and stored at − 80 °C for further experiments. The margin of tumor tissue was histopathologically examined and considered as a non-tumor tissue.
RNA isolation and cDNA synthesis
In order to isolate the RNA of frozen tissues, TRIZOL reagent (Invitrogen, Life technology, USA) was used based on the manufacturer’s directions. The quality and quantity of the isolated RNAs were measured by PicoDrop. The complementary DNAs were synthesized from the isolated RNA using Thermo Fisher Scientific cDNA synthesis kit and random hexamer primer according to the procedure.
Quantitative real-time PCR (qRT-PCR)
In order to examine the mRNA expression level of the FLJ22447, the HIF1A, and the VEGF, the qRT-PCR was performed using SYBER Green PCR master mix kit (TAKARA, Japan). The cDNA products were applied as templates for denaturing and annealing/extension steps at 95 °C and 60 °C, respectively. The melt curve analysis was performed for evaluating the specificity of PCR products. Table 1 presents the characteristics of primers applied in this assay, in which the GAPDH was considered as houdsekeeping gene. Ultimately, the fold change expression level of genes was calculated by the 2–∆∆CT method.
Interaction analysis of FLJ22447
To identify the interaction of biological macromolecules with the FLJ22447, RNA association interaction database, the RAID v.2 database (www.rna-society.org/raid), was applied. The LncRNADisease (http://cmbi.bjmu.edu.cn/lncrnadisease) and the Lnc2Cancer (http://www.bio-bigdata.net/lnc2cancer) databases were screened out to investigate the biological function of irregular expression of the FLJ22447 in different types of cancer.
Systems biology analysis
The possible biological consequences of the FLJ22447 were determined by exploring the molecular mechanisms of its protein partners through the bioinformatics tools.
The protein–protein interaction network (PPI) of the predicted protein partners of the FLJ22447 was retrieved by the STRING (v 11.0) database with at least medium confidence (score 0.4). Then, the Cytoscape software was applied to identify the hub proteins by visualizing the protein interaction relationship network in network analyzer tool. The hub genes were subsequently recognized according to the degree parameter. Moreover, the functional annotation and significant pathways as a signature for predicting the clinical outcome were recognized by the PANTHER (v 14.0) and the KEGG resources, respectively.
Survival analysis
To assess whether the expression of HIF1A and VEGF is associated with the survival rate of patients with ESCA and STAD, the OncoLnc database (http://www.oncolnc.org/) was applied. The Kaplan–Meier analysis were performed for 72 ESCA patients and 189 STAD patients by the log rank comparison.
Statistical analysis
The statistical significance was determined using one-way analysis of variance (ANOVA) and Student’s t-test by the SPSS software (V. 16). All data were presented as mean ± SD and the significant value of P was considered less than 0.05.
Results
HIF1A expression in clinical ESCC and GC samples
The expression level of HIF1A was significantly increased in clinical ESCC specimens compared to the adjacent non-tumor tissues (P < 0.05, Fig. 1). No difference was observed in the HIF1A expression across GC samples and their adjacent non-tumor tissues (P > 0.05).
Table 2 represents the association of HIF1A expression with the clinicopathological factors in ESCC samples. A significant correlation was observed between the expression of HIF1A and tumor stage. The high expression of HIF1A was detected in stages I–II and its low expression in the stages III–IV.
FLJ22447 expression in clinical ESCC and GC samples
The expression of FLJ22447 was evaluated by qRT–PCR assay in the ESCC and the GC specimens. As can be seen in Fig. 1, the expression of FLJ22447 was significantly down-regulated in both ESCC and GC tissues than adjacent non-tumor tissues (P < 0.05).
VEGF expression in clinical ESCC and GC samples
The VEGF expression was significantly increased in both ESCC and GC samples than their adjacent non-tumor tissues (P < 0.05, Fig. 1). Table 3 shows the association of the VEGF expression with the clinicopathological features of the ESCC patients. A remarkable link was recognized between the high expression of VEGF and advanced stages (III–IV) and metastasis (P < 0.05).
In silico expression profile of FLJ22447, HIF1A, and VEGF
The expression pattern of target genes in esophagus (ESCA) and stomach (STAD) carcinomas samples was explored by the GEPIA database (Fig. 2A). The FLJ22447 was highly expressed in ESCC than STAD. However, in comparison with the adjacent non-tumor tissues, its expression reduced in ESCC, whereas, increased in STAD. Interestingly, the transcription of HIF1A was significantly heightened in both ESCA and STAD than their adjacent non-tumor tissues (P < 0.05). The mean expression of the VEGF and the FLJ22447 in ESCA were lower than the adjacent non-tumor ones.

adopted from GEPIA resources. Log2 (TPM + 1) was used for log-scale. ESCA esophageal carcinoma; TPM Transcripts per million
Comparison of the expression profile of FLJ22447, HIF1A, and VEGF in ESCA and STAD carcinoma. A The RNA seq data in GEPIA indicated that the expression of HIF1A was increased in ESCA tissues than normal tissues. In contrast, the mean expressions of VEGF and FLJ22447 were lower than in ESCA tissues. B Correlation of FLJ22447, HIF1A, and VEGF gene expression in both ESCA and STAD were considered. The analysis of spearman correlation revealed that low expression of FLJ22447 was significantly correlated with the HIF1A and the VEGF expression in esophagus and stomach carcinomas, respectively. Additionally, a positive correlation was observed between the HIF1A and the VEGF genes in both carcinomas. C The Kaplan-Meier survival curves for the patients harboring ESCC and STAD with regard to high and low expression of HIF1A and VEGF are demonstrated. Patients with high HIF1A expression have less survival. Data
Moreover, down-regulation of the FLJ22447 in esophagus and stomach carcinomas was significantly correlated with the HIF1A (P = 2.4e–73, R = 0.63; P = 4.7e–36, R = 0.55) and the VEGF (P = 0.00019, R = 0.15; P = 0.044, R = 0.096) expression, respectively (Fig. 2B, C). Also, a positive correlation was observed between the HIF1A and the VEGF genes in esophagus and stomach carcinomas (p = 7.6e–28, R = 0.41; p = 2.7e–15, R = 0.36).
Patients’ survival analysis
Figure 3 illustrates the survival rate of patients with high vs low expression levels of VEGF and HIF1A. The Kaplan–Meier survival curve indicated that 10-years survival rate of ESCA and STAD patients was approximately 30%. No significant difference was observed in terms of high and low expression of HIF1A and VEGF with the outcome of ESCA and STAD patients. The analysis of patients’ survival showed that the ESCA patients with low expression of HIF1A achieved better survival than those with high expression.
Interaction analysis of FLJ22447 in human tumors
The term of sequence ontology for the FLJ22447 is sense_intronic_ncRNA in the LNCipedia database.
It contains three transcripts with the length of 1368, 1312, and 577 bp. According to the LncRNA Disease database, the impact of FLJ22447 in cervical, lymphoma, glioma, stomach, thyroid, and bladder cancers has been predicted through the TAM method. Also, the interaction of FLJ22447 with 67 protein coding-genes and RNAs was revealed in the RAID v.2 resource (Table 4). These predicted results provide important insights into FLJ22447 function in the processes of biologic. However, further research is demanded.
Enrichment analysis of interacted proteins with FLJ22447
Molecular function and biological process of the FLJ22447-interacted proteins were explored by the PANTHER database (Fig. 4). The enrichment analysis suggested that 23 and 20 genes participate in two major molecular function categories including binding (GO: 0005488; 47.9%) and transcription regulator activity (GO: 0140110; 41.7%), respectively. From the pie chart above, we can see that these proteins were mostly involved in metabolic process (GO: 0008152; 30.1%), biological regulation (GO: 0065007; 26.5%), and cellular process (GO: 0009987; 16.9%).
Further, the conceivable biological function of FLJ22447 was demonstrated by finding the most important nodes across its interaction protein partners (Fig. 5A). Using network analyzer tool of Cytoscape, MYC (24), JUN (21), SMRCA4 (20), PPARG (18), AR (17), FOS (17) and CEBPA (16) protein-coding genes were detected as the hub genes according to the degree method (Fig. 5B). According to the network analysis of the hub genes, MYC and AR mostly act as an activator of other hub genes. While, PPARG and CEBPB are activated by others (Fig. 5C). Moreover, the result of Enrichr analysis revealed that these proteins were implicated in thyroid cancer (PPARG, MYC), acute myeloid leukemia (MYC, CEBPA), colorectal and breast cancers (MYC, JUN, FOS) (Fig. 5D). As shown in the above figure, all the hub genes were involved in the cancer related pathways (map05200).
Functional classification of FLJ22447-interacted proteins. A The molecular action mode of FLJ22447 protein partners with medium score (0.4) was nominated by STRING (V 11.0). The line colors indicate binding (blue), activation (green), inhibition (red), reaction (black), post-translational modification (pink), and transcriptional regulation (green mint). The un-specified, positive and negative effects were illustrated by dot, arrowhead, and bar, respectively. B Network of FLJ22447-interacted proteins was constructed by application of CytoHubba in Cytoscape software. C The network of hub genes was depicted according to the degree method. MYC, JUN, SMRCA4, PPARG, AR, FOS and CEBPA protein-coding genes were identified as the hub genes. D The clustergram of KEGG 2019 human pathway section in Enrichr demonstrated that these proteins were involved in thyroid cancer, acute myeloid leukemia, colorectal, and breast cancers. It is interesting to note that all the hub genes were involved in pathways in cancer (map05200). (Color figure online)
Discussion
Prior researches have noted that tumor microenvironment particularly its major component CAFs promote ESCC and GC initiation and progression [23, 24]. Among the components of the CAFs, the functional impact of non-coding RNAs has been recently considered in cancer biology. The lncRNAs participate in tumor-stromal crosstalk and provide a suitable tumor microenvironment [25]. The FLJ22447, referred to LncRNA-CAF, was introduced as a reprogramming factor of CAF in OSCC [19]. This lncRNA is cytologically located in the vicinity of HIF1A. Nevertheless, very little was found on the association of FLJJ22447 with the HIF1A and VEGF. Consequently, the present study was set out to identify whether the FLJJ22447 expression level can influence the vicinity gene expression of HIF1A in the ESCC and GC patients.
Our data showed that the FLJJ22447 was lower expressed in the human ESCC and GC. The interesting point of expression assay was that the most notable reduction in the FLJJ22447 expression was observed in ESCC samples; at the same condition that the expression of HIF1A increased. The value of FLJJ22447 suggests that it could be a major factor, if not the only one, regulating its neighbor gene, HIF1A.
These findings further support the idea of FLJJ22447 involvement into the cancer-associated pathways through the enrichment analysis of FLJ22447-interacted protein-coding genes (Fig. 5A). We also identified that MYC, JUN, SMRCA4, PPARG, AR, FOS and CEBPA genes as critical nodes that take part in pathways of thyroid, colorectal, and breast cancers (Fig. 5D). Additionally, functional interactions network of these proteins suggested that MYC and AR genes act as an activator in this biological network (Fig. 5C).
The underlying biological mechanisms of FLJJ22447 have not been fully elucidated. It has been demonstrated that the FLJJ22447 increased the IL-33 stability by preventing autophagic degradation and promotes tumor proliferation [18]. They also mentioned that the FLJ22447 promotes CAF activation and tumor progression in OSCC. Moreover, high expression of FLJ22447 was associated with a high TNM stage and poor patient survival, suggesting it as a novel therapeutic target in OSCC [19].
Assessment of RNA-seq data in the GEPIA revealed that the expression of SPI1, E2F1, TCF7L2, and STAT1 genes among the FLJ22447-interacted proteins were significantly higher than non-tumor tissue in ESCA and STAD (Fig. 6). They contribute in proliferation, evading apoptosis, and sustained angiogenesis pathways in cancer (hsa05200). From the oncology point of view, the over-expression of these genes is contributed to the tumor progression. The SPI1 and E2F1 are known as oncogenic transcription factors [26]. It is possible that the FLJ22447 expression in ESCC and GC is associated with these genes, at least in part. Further research on the current topic are therefore recommended. These results indicated that the FLJ22447 may have several biological functions and participate in cancer-associated pathways, suggesting its potential value in tumor gene therapy.
Expression profile analysis of the FLJ22447-interacted proteins. A Among the FLJ22447-interacted proteins, SPI1, E2F1, TCF7L2, and STAT1 genes were significantly expressed than non-tumor tissue in ESCA and STAD. B They contribute in proliferation, evading apoptosis, and sustained angiogenesis pathways in cancer (hsa05200). Data retrieved from the GEPIA, KEGG, and Enrichr databases
We have also found a positive link between the HIF1A and VEGF expression in ESCC tissues than adjacent non-tumor tissues. This result is consistent with other studies who suggested that these two angiogenic factors are essential in progression of squamous cell carcinoma of the esophagus [27,28,29].
In addition, HIF1A acts as a transcription factor for the VEGF [28]. In fact, there is a HIF1A binding site in the VEGF promoter that results in the regulating of the VEGF expression during hypoxia [27, 30].
In terms of clinical view, the correlation of these angiogenic factors with the histopathological parameters in breast cancer, pancreatic adenocarcinoma, and esophagus carcinoma has been demonstrated [2, 12, 28].
We found that low expression of HIF1A was related to the tumor stage, however, this association was not previously reported in ESCC tumor specimens [27]. Whereas, they demonstrated a correlation among high expression of HIF1A and venous invasion.
From this point of vision, increased expression of the VEGF in the present study was associated with high tumor grade (III–IV) and metastasis. In accordance with this result, previous researches have demonstrated that the VEGF expression was correlated with tumor stage in ESCC and gastric adenocarcinoma [29, 31]. The association of these molecules with tumor grade, metastasis, and vascular invasion suggested their contribution to the survival of tumor cells. HIF1A and VEGF are important determinants of hypoxia and angiogenesis phenomena in cancer [18, 32]. Recently, their implication in CAFs through lncRNAs, tumor suppressor genes, and oncogenes have been demonstrated [33, 34].
With respect to the survival rate of ESCA and STAD patients, there were no significant differences in terms of high and low expression of HIF1A and VEGF. However, high expression of VEGF in patients with gastric adenocarcinoma indicated a poor prognosis for overall survival [31]. The same result has been reported in the patients harboring breast cancer and ESCC with high expression of HIF1A [27, 35].
Conclusion
This investigation purposed to determine the association of FLJ22447 with the HIF1A as its neighbor gene in the ESCC and GC patients. Herein, we demonstrated low expression of FLJ22447 in the patients with ESCC and GC that was linked to the HIF1A expression. It is possible, therefore, that the vicinity of HIF1A with FLJ22447 may affect the regulation of its expression. Also, the relationship between HIF1A and VEGF was demonstrated in the ESCC and GC patients. The enrichment analysis of FLJ22447-interacted proteins revealed that this ncRNA may have various roles in the biological processes and may be involved in the pathways attributed to the cancer. Association of the oncogenic transcription factors with the FLJ22447 and their contribution in proliferation, evading apoptosis, and sustained angiogenesis pathways in cancer, suggesting its potential value in tumor gene therapy. Notwithstanding our research limitations, this study provides insights into the FLJ22447 interactions and its protein partners.
Data availability
Data will be available while editor requested.
References
Plavetić D, Plavetić ND, Barić MB, Bradić LB, Kulić A, Pleština S (2014) Hypoxia in solid tumors: biological responses to hypoxia and implications on therapy and prognosis. Period Biol 116(4):361–364
Banys-Paluchowski M, Witzel I, Riethdorf S, Pantel K, Rack B, Janni W, Fasching PA, Aktas B, Kasimir-Bauer S, Hartkopf A (2018) The clinical relevance of serum vascular endothelial growth factor (VEGF) in correlation to circulating tumor cells and other serum biomarkers in patients with metastatic breast cancer. Breast Cancer Res Treat 172(1):93–104
Spirina L, Usynin Y, Yurmazov Z, Slonimskaya E, Kolegova E, Kondakova I (2017) Transcription factors NF-kB, HIF-1, HIF-2, growth factor VEGF, VEGFR2 and carboanhydrase IX mRNA and protein level in the development of kidney cancer metastasis. Mol Biol 51(2):328–332
Bahramian S, Shamsabadi F, Fazel A, Delshad E, Amini A, Memari F, Shafiee M (2020) Evaluation of Arylsulfatase D (ARSD) and long noncoding RNA ARSD-AS1 gene expression in breast cancer patients and their association with oncogenic transcription factors. J BUON 25:1805–1813
Muz B, de la Puente P, Azab F, Azab AK (2015) The role of hypoxia in cancer progression, angiogenesis, metastasis, and resistance to therapy. Hypoxia 3:83
Ivan M, Kaelin WG Jr (2017) The EGLN-HIF O(2)-sensing system: multiple inputs and feedbacks. Mol Cell 66(6):772–779
Hashimoto T, Shibasaki F (2015) Hypoxia-inducible factor as an angiogenic master switch. Front Pediatr 3:33–33
AM Roberts, IR Watson, AJ Evans, DA Foster, MS Irwin, M Ohh (2009) Suppression of hypoxia-inducible factor 2α restores p53 activity via Hdm2 and reverses chemoresistance of renal carcinoma cells. Cancer Res 0008-5472. CAN-09-1770.
Nagao A, Kobayashi M, Koyasu S, Chow CCT, Harada H (2019) HIF-1-dependent reprogramming of glucose metabolic pathway of cancer cells and its therapeutic significance. Int J Mol Sci 20(2):238
Ladeira K, Macedo F, Longatto-Filho A, Martins SF (2018) Angiogenic factors: role in esophageal cancer, a brief review. Esophagus 15(2):53–58
Jin X, Dai L, Ma Y, Wang J, Liu Z (2020) Implications of HIF-1α in the tumorigenesis and progression of pancreatic cancer. Cancer Cell Int 20(1):1–11
Zhao X, Gao S, Ren H, Sun W, Zhang H, Sun J, Yang S, Hao J (2014) Hypoxia-inducible factor-1 promotes pancreatic ductal adenocarcinoma invasion and metastasis by activating transcription of the actin-bundling protein fascin. Cancer Res 74(9):2455–2464
Jin F, Brockmeier U, Otterbach F, Metzen E (2012) New insight into the SDF-1/CXCR4 axis in a breast carcinoma model: hypoxia-induced endothelial SDF-1 and tumor cell CXCR4 are required for tumor cell intravasation. Mol Cancer Res 10(8):1021
Viallard C, Larrivée B (2017) Tumor angiogenesis and vascular normalization: alternative therapeutic targets. Angiogenesis 20(4):409–426
Bramhachari PV, Prathyusha A, Reddy DRS (2017) Overview of transcription factors in esophagus cancer, role of transcription factors in gastrointestinal malignancies. Springer, Singapore, pp 31–42
Becker S, Wang H, Simmons AB, Suwanmanee T, Stoddard GJ, Kafri T, Hartnett ME (2018) Targeted knockdown of overexpressed VEGFA or VEGF164 in Müller cells maintains retinal function by triggering different signaling mechanisms. Sci Rep 8(1):2003
Gutschner T, Diederichs S (2012) The hallmarks of cancer: a long non-coding RNA point of view. RNA Biol 9(6):703–719
Ahn Y-H, Kim JS (2020) Long non-coding RNAs as regulators of interactions between cancer-associated fibroblasts and cancer cells in the tumor microenvironment. Int J Mol Sci 21(20):7484
Ding L, Ren J, Zhang D, Li Y, Huang X, Hu Q, Wang H, Song Y, Ni Y, Hou Y (2018) A novel stromal lncRNA signature reprograms fibroblasts to promote the growth of oral squamous cell carcinoma via LncRNA-CAF/interleukin-33. Carcinogenesis 39(3):397–406
Schmitt AM, Chang HY (2016) Long noncoding RNAs in cancer pathways. Cancer Cell 29(4):452–463
Rupaimoole R, Slack FJ (2017) MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov 16(3):203
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45(W1):W98–W102
Fang Z, Xu J, Zhang B, Wang W, Liu J, Liang C, Hua J, Meng Q, Yu X, Shi S (2020) The promising role of noncoding RNAs in cancer-associated fibroblasts: an overview of current status and future perspectives. J Hematol Oncol 13(1):1–21
Du X, Xu Q, Pan D, Xu D, Niu B, Hong W, Zhang R, Li X, Chen S (2019) HIC-5 in cancer-associated fibroblasts contributes to esophageal squamous cell carcinoma progression. Cell Death Dis 10(12):1–16
Zhou L, Zhu Y, Sun D, Zhang Q (2020) Emerging roles of long non-coding RNAs in the tumor microenvironment. Int J Biol Sci 16(12):2094
Lambert M, Jambon S, Depauw S, David-Cordonnier M-H (2018) Targeting transcription factors for cancer treatment. Molecules 23(6):1479
Kimura S, Kitadai Y, Tanaka S, Kuwai T, Hihara J, Yoshida K, Toge T, Chayama K (2004) Expression of hypoxia-inducible factor (HIF)-1α is associated with vascular endothelial growth factor expression and tumour angiogenesis in human oesophageal squamous cell carcinoma. Eur J Cancer 40(12):1904–1912
Takala H, Saarnio J, Wiik H, Ohtonen P, Soini Y (2011) HIF-1α and VEGF are associated with disease progression in esophageal carcinoma. J Surg Res 167(1):41–48
Tzao C, Lee S-C, Tung H-J, Hsu H-S, Hsu W-H, Sun G-H, Yu C-P, Jin J-S, Cheng Y-L (2008) Expression of hypoxia-inducible factor (HIF)-1α and vascular endothelial growth factor (VEGF)-D as outcome predictors in resected esophageal squamous cell carcinoma. Dis Markers 25(3):141–148
Kaelin WG Jr, Ratcliffe PJ, Semenza GL (2019) Out of breath: molecular description of cellular responses to hypoxia–2019 Nobel prize for physiology or medicine. Curr Sci 117(9):1418
Wei B, Tai Y, Tong H, Wen S-L, Tang S-H, Huan H, Huang Z-Y, Liu R, Tang Y-M, Yang J-H (2017) Correlations between VEGF-A expression and prognosis in patients with gastric adenocarcinoma. Int J Clin Exp Pathol 10(8):8461
Ham I-H, Lee D, Hur H (2019) Role of cancer-associated fibroblast in gastric cancer progression and resistance to treatments. J Oncol 2019:1–11
Ma Z, Chen M, Yang X, Xu B, Song Z, Zhou B, Yang T (2018) The role of cancer-associated fibroblasts in tumorigenesis of gastric cancer. Curr Pharm Des 24(28):3297–3302
Sewell-Loftin MK, Bayer SVH, Crist E, Hughes T, Joison SM, Longmore GD, George SC (2017) Cancer-associated fibroblasts support vascular growth through mechanical force. Sci Rep 7(1):1–12
Dales JP, Garcia S, Meunier-Carpentier S, Andrac-Meyer L, Haddad O, Lavaut MN, Allasia C, Bonnier P, Charpin C (2005) Overexpression of hypoxia-inducible factor HIF-1α predicts early relapse in breast cancer: Retrospective study in a series of 745 patients. Int J Cancer 116(5):734–739
Acknowledgements
We gratefully acknowledge all persons who helped us in this study in Molecular Genetic lab which was supported by Golestan University of Medical Sciences, Gorgan, Iran.
Funding
This research was supported in part by Golestan University of Medical sciences (Project Grant No. 960704177). The study sponsor had no role in the study design, collection, analysis and interpretation of the data or the writing of the report.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
No potential conflicts of interest were disclosed.
Ethical approval
This research was approved in Ethics Committee of by IR.GOUMS.REC.1397.006.
Consent for publication
Not Applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bahramian, S., Sahebi, R., Roohinejad, Z. et al. Low expression of LncRNA-CAF attributed to the high expression of HIF1A in esophageal squamous cell carcinoma and gastric cancer patients. Mol Biol Rep 49, 895–905 (2022). https://doi.org/10.1007/s11033-021-06882-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06882-0