Abstract
Background
Metastasis is a major cause of death in Colorectal cancer (CRC) patients, and the Epithelial–mesenchymal transition (EMT) has been known to be a crucial event in cancer metastasis. Downregulated expression of AT-rich interaction domain-containing protein 1A (ARID1A), a bona fide tumor suppressor gene, plays an important role in promoting EMT and CRC metastasis, but the underlying molecular mechanisms remain poorly understood. Here, we evaluated the impact of ARID1A knockdown and overexpression on the expression of EMT‑related genes, E-cadherin and β-catenin, in human CRC cells.
Methods and results
The expression levels of ARID1A, E-cadherin and β-catenin in CRC cell lines were detected via real-time quantitative PCR (qPCR) and western blot. ARID1A overexpression and shRNA-mediated knockdown were performed to indicate the effect of ARID1A expression on E-cadherin and β-catenin expression in CRC cell lines. The effect of ARID1A knockdown on the migration ability of HCT116 cells was assessed using wound-healing assay. We found that the mRNA and protein expression of adhesive protein E-cadherin was remarkably downregulated in response to shRNA-mediated ARID1A knockdown in HCT116 and HT29 cells. Conversely, overexpression of ARID1A in SW48 cells significantly increased E-cadherin expression. In addition, ARID1A silencing promoted the migration of HCT116 cells. ARID1A knockdown and overexpression did not alter the level of β-catenin expression.
Conclusions
Our study demonstrates that E-cadherin levels were closely correlated with ARID1A expression. Thus, ARID1A downregulation may promote CRC metastasis through decreasing EMT‑related protein E-cadherin and promoting epithelial cell movement. ARID1A could represent a promising candidate therapeutic target for CRC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer (CRC) is the third most common cancer worldwide, and is a major cause of cancer mortality, accounting for approximately one-tenth of all tumor-related deaths in the world annually [1, 2]. Metastasis is the primary cause of treatment failure and death in CRC [3, 4]. Thus, a better understanding of the molecular mechanism underlying colorectal cancer metastasis may lead to new therapeutic strategies.
AT-rich interactive domain-containing protein 1A (ARID1A), a bona fide tumor suppressor gene, encodes a key subunit of the SWItch/Sucrose Non-Fermentable (SWI/SNF) chromatin-remodeling complex. The SWI/SNF complex modulates the DNA accessibility to co-regulatory and transcriptional machinery and acts as a transcriptional regulator of genes involved in many important cellular processes, including proliferation, development, differentiation and DNA damage repair [5]. As a subunit of SWI/SNF complex, ARID1A is assumed to play a crucial role in SWI/SNF complex targeting to enhancers and in maintaining their chromatin accessibility as it has DNA binding activity. Upon ARID1A inactivation, defects in targeting SWI/SNF and control of enhancer activity impair development and differentiation programs and cause extensive dysregulation of gene expression, thus driving tumor formation [5]. For instance, in ARID1A-deficient colon cancer, SWI/SNF is lost from thousands of enhancers with corresponding effects on gene expression [6].
ARID1A is one of the most frequently mutated tumor suppressors in various human cancers [7]. As data have emerged from the Cancer Genome Atlas project (TCGA), mutations in ARID1A have been found at high frequencies in CRC. With an estimated mutation rate of 10–40%, ARID1A is the third most significantly mutated gene in human CRC, following APC and P53 [6, 8]. Furthermore, mutations in ARID1A are generally inactivating (insertion/deletion) a characteristic of many tumor suppressors [7]. Apart from inactivating mutations, aberrant DNA methylation, has also been reported as an important mechanism for suppressing ARID1A expression in CRC [9]. Available data support an important role of ARID1A inactivation in promoting formation and metastasis of CRC. ARID1A knockout in mice per se, drives the formation of invasive colon tumors [6]. In addition, in a clinical study of CRC, Kishida et al. revealed that negative ARID1A expression was correlated with early onset and lymphatic invasion [10]. It has been found that ARID1A knockout can enhance the migratory activity of cancer cells by promoting Epithelial mesenchymal transition (EMT) [11,12,13,14]. EMT has been known to be a crucial event in cancer invasion and metastasis [15, 16]. During EMT, cancer cells lose their cell–cell adhesions and polarity and thereby acquire enhanced invasive and migratory properties [16, 17]. However, the role of ARID1A in expression of genes associated with EMT and cell–cell adhesion in CRC remains poorly understood. Therefore, in this study, we aimed to investigate the impact of ARID1A knockdown and overexpression on the expression of EMT‑related genes, E-cadherin and β-catenin, in human CRC cells. Since ARID1A expression loss is not rare in colorectal cancer tumors, correlation of ARID1A and E-cadherin expression in CRC tissue samples remains a goal for future studies.
Materials and methods
Cell culture
HEK293T cell line and six CRC cell lines (HCT116, HT 29, SW48, SW742, SW480 and LS180) were purchased from the National Cell Bank of Iran (Pasteur Institute, Tehran, Iran). HCT116, HT 29, SW48, SW742 and SW480 cells were cultured in RPMI 1640 medium (#11875093, Thermo Fisher Scientific, Inc., Bartlesville, OK, USA) and HEK293T and LS180 cells were cultured in DMEM medium (#12430062, Thermo Fisher Scientific, Inc., Bartlesville, OK, USA) containing 10% fetal bovine serum (Gibco-BRL), 1% penicillin(#16000044, Thermo Fisher Scientific, Inc., Bartlesville, OK, USA) and 1% streptomycin (HyClone; GE Healthcare Life Sciences, Logan, UT, USA), in a humidified CO2 (5%) incubator at 37 °C, and regularly checked and confirmed to be mycoplasma-free.
Plasmids and lentivirus production
Overexpression plasmid of pcDNA 6- ARID1A was obtained from addgene (#39311, Addgene plasmid) for the overexpression assay. Empty vector was used as a negative control. The short hairpin RNA (shRNA) lentiviral plasmid (pLKO.1) which contains a puromycin resistance gene was obtained from Addgene (#10878, pLKO.1 - TRC Cloning Vector, Addgene). The shRNA oligonucleotide against ARID1A gene (oligo ID: TRCN0000059090) with the sequence of 5'- CCTCTCTTATACACAGCAGAT -3' was cloned into the pLKO.1 vector. A pLKO.1-puro Empty Vector (#RHS4080, Open Biosystems) was also used to produce empty lentivirus as a control. The empty lentivirus and lentivirus encoding ARID1A shRNA were produced using HEK293T cells transfected with pLKO.1 plasmids and the second generation packaging system, pMD2.G (#12259, Addgene plasmid) and pSPAX2 (#12260, Addgene plasmid). Briefly, about 80% confluent HEK293T cells were co-transfected with 0.5 µg of the envelope protein–coding plasmid (pMD2.G), and 1.5 µg of the packaging construct (pSPAX2) together with 2 µg of pLKO.1 vector encoding ARID1A shRNA or negative control pLKO.1 Empty Vector using a standard polyethylenimine (PEI) mediated method [18]. After 48 h, the virus-containing media was collected, centrifuged (1250 rpm for 5 min), and stored in small aliquots at − 80 °C.
Generation of the HCT116 and HT29 cells stably expressing shRNA against ARID1A
HCT116 and HT29 cells were seeded in 6-well plates. After 24 h, the lentiviral supernatant containing either a lentivirus expressing ARID1A shRNA or an empty lentivirus as a control was added at MOI (multiplicity of infection) of 5–50 together with 8 μg/ml polybrene (#sc-134220, Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA). 24 h later, the transduction medium was replaced with fresh complete medium and puromycin (#sc-108071, Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA) selection (1 μg/ml) was started 72 h post transfection. Approximately, in every 2–3 days, the culture media was aspirated and replaced with freshly prepared selective media. After 3 weeks under puromycin selection the remaining resistant clones were assessed for knock down efficacy. ARID1A Knockdown efficacy was assessed by immunoblotting and qPCR.
ARID1A gene transfection into SW48 cells
SW48 cells were transfected with 2 μg pcDNA6- ARID1A or 2 μg pcDNA6 (as a negative control). Cell transfection was performed using Lipofectamine 2000 (#11668030, Invitrogen; Thermo Fisher Scientific, Inc. USA) according to the manufacturer’s instructions. The efficiency of the transfection was examined 48 h post-transfection using real-time quantitative PCR and Western blotting.
RNA extraction and real-time PCR assay
RNA of cultured cells was extracted using the Tripure RNA isolation kit (#11667165001, Roche Applied Science, Germany) based on the manufacturer’s guidelines. cDNA synthesis was conducted by RevertAid First Strand cDNA Synthesis Kit (#K1622, Thermo Fisher Scientific, USA). mRNA analysis of ARID1A was determined by real-time quantitative PCR (qPCR) using SYBR Green master mix (#4309155, Thermo Fisher Scientific, USA) on an ABI 7500 Sequence Detection System (Applied Biosystems, USA). GAPDH was used as an internal housekeeping gene. Relative gene expression levels were calculated using the 2−ΔΔCt method. The primers used in this study are listed in Table 1.
Western blot analysis
Cultured cells were lysed using ice‑cold lysis buffer in the presence of a protease inhibitor cocktail (Roche Applied Science) and the protein concentration was quantified using a Pierce BCA protein assay kit (#23225, Thermo Fisher Scientific, USA). Proteins with an equal amount were separated by 12% SDS‑PAGE and transferred onto nitrocellulose membranes. The membranes were then blocked using 5% skim milk/PBS for 1 h. After blocking, membranes were incubated with the following primary antibodies overnight at 4 °C: ARID1A (#sc-373784; Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA), β-catenin (#sc-7963; Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA), E‑cadherin (#sc-71009; Santa Cruz Biotechnology, Inc., Santa Cruz, CA, USA) and GAPDH (#ab181602; Abcam, UK) as an internal control. Each membrane was subsequently washed with TBST three times and incubated with a corresponding HRP-conjugated secondary antibody at room temperature for 1 h. Protein detection was performed using a Pierce™ enhanced chemiluminescence (ECL) western blotting substrate (#32209, Thermo Fisher Scientific, Inc., Bartlesville, OK, USA). Band intensity data was obtained using Image lab software (version 5.2, Bio-Rad).
Wound healing (scratch) migration assay
In order to determine the cell migration, the HCT116 cells were cultured in 6-well plates to confluence. Subsequently, the monolayer cells were scratched using 200 μl pipette tip from the top to the bottom of the culture plates. After the debris was removed with PBS, the cells were further maintained at 37 °C in a humidified incubator with 5% CO2. Wounded area was visualized after 0, 24 and 48 h using an Olympus CX31 light microscope (Olympus Corporation, Tokyo, Japan).and calculated by NIH ImageJ software (https://imagej.nih.gov/ij/). Cell motility was estimated through the quantification of the % of recovery using the equation: R (%) = [1 − (wound area at Tt/wound area at T0)] × 100, where T0 is the wounded area at 0 h and Tt is the wounded area after t.
Statistical analysis
All the experiments were repeated at in triplicate and all data were expressed as mean ± SD. The statistical package IBM SPSS Statistics (IBM Corp. Released 2017. IBM SPSS Statistics for Windows, Version 25.0. Armonk, NY: IBM Corp.) was used for all analyses. graphs were performed using Prism version 6.00 (GraphPad Software, Inc., San Diego, CA, USA). The one-way ANOVA, accompanied by Tukey’s multiple comparison tests were performed to analyze data. Differences with a P value ≤ 0.05 were statistically significant.
Results
The efficiency of overexpression and shRNA-mediated knockdown of ARID1A in CRC cell lines
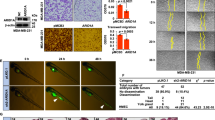
We examined the expression of ARID1A in a panel of CRC cell lines using qPCR (Fig. 1a). Among the six CRC cell lines, the highest expression level of ARID1A was in the HCT116 and HT29 cells, whereas SW48 cells lacked ARID1A expression (Fig. 1a). Thus, these cells were selected for subsequent knockdown and overexpression analyses, respectively. We stably silenced ARID1A in HCT116 and HT29 cells using ARID1A shRNA lentivirus and overexpressed ARID1A in SW48 cells by plasmid transfection. Successful knockdown and overexpression of ARID1A in these cells were confirmed using Western blot and qPCR analysis (Fig. 1b and c).
ARID1A relative mRNA expression in several colorectal carcinoma cell lines and efficiency of overexpression and shRNA-mediated knockdown of ARID1A. a Endogenous expression of ARID1A was examined in HCT116, HT29, LS180, SW742 by qPCR. SW48 cell line was utilized as a reference, with an expression level set to 1.0, and expressions in all other cell lines were presented as an n-fold difference, compared to the SW48. Analysis of western blot (b) and qPCR (c) showed reduction in ARID1A expression in HCT116 and HT29 cells transduced with ARID1A shRNA virus, compared to control virus transduced cells, while pCDNA6-ARID1A plasmid transfection markedly enhanced ARID1A levels in SW48 cells. The ARID1A expression was normalized relative to GAPDH. For panel A and C, Mean ± standard deviation (SD) of three experiments is reported. (**p < 0.01)
ARID1A silencing suppresses the expression of E-cadherin protein and mRNA, whereas the ARID1A overexpression exerts an opposite effect
Western blot analysis and RT‑qPCR indicated that the HCT116 and HT29 cells with stable knockdown of ARID1A by ARID1A shRNA virus exhibited a lower expression of E‑cadherin protein and mRNA as compared to cells transduced with control virus carrying pLKO.1. On the other hand, the SW48 cells transfected with pCDNA6-ARID1A exhibited a higher expression of E‑cadherin protein and mRNA as compared to the control group of cells lacking ARID1A expression (Fig. 2). The above results suggest that ARID1A may modulate E‑cadherin expression.
Effects of ARID1A on the protein and mRNA expression of E‑cadherin and β-catenin in HCT116, HT29 and SW48 cells. Western blot (a) and qPCR (b) demonstrates that ARID1A stable knockdown results in reduced expression of E‑cadherin in both HCT116 and HT29 cells, while induction of ARID1A expression leads to upregulation of E‑cadherin in SW48 cells. The β-catenin expression is not significantly affected following ARID1A knockdown or overexpression. Values are expressed as mean ± SD (n = 3). (**P < 0.01)
β-catenin remains stable regardless of ARID1A silencing or overexpression
To identify the role of ARID1A in the expression of β-catenin in CRC cells, protein and mRNA levels of β-catenin in CRC cells were analyzed after overexpression or stable knockdown of ARID1A. As shown in Fig. 2, the expression of β-catenin remained largely stable regardless of ARID1A silencing or overexpression in CRC cells.
Knockdown of ARID1A promotes HCT116 cells migration
The migration ability of HCT116 cells was examined by wound healing assay. Microscopic examination after wounding revealed that HCT116 cells transduced with ARID1A shRNA lentivirus have obviously increased ability of wound closure at all time points (up to 48 h post-scratch) compared to HCT116 cells transduced with control lentivirus (Fig. 3).
Knockdown of ARID1A promotes HCT116 cells migration. a The Effects of ARID1A knockdown on migratory activity of HCT116 cells was examined by wound healing assay. Representative images of wound healing assay at indicated time points (0, 24 and 48 h after the scratch) are shown. Dashed line indicates boundary of the scratched wound. b Cell motility was estimated through the quantification of the % of recovery using the equation detailed in section of Materials and Methods. Data were presented as means ± SD from three replicates. (*p < 0.05)
Discussion
EMT, an absolute requirement for tumor metastasis and invasion, plays a key role in cancer progression, including CRC [16, 17, 19]. Due to its emerging role as a pivotal driver of tumorigenesis, inhibiting EMT is an attractive therapeutic strategy in combating metastasis in cancer patients [20]. The hallmark of EMT is the decreased expression of the E-cadherin protein [21, 22]. When E-cadherin is downregulated, epithelial cells lose their polarity and cell–cell adhesion, and EMT then occurs. Therefore, the downregulation mechanism of E-cadherin is a key topic of EMT research [15, 21]. In recent years, an increasing number of studies have found that ARID1A downregulation is associated with EMT and tumor invasion via the repression of E-cadherin in several different types of cancer including some types of gastrointestinal cancers [11,12,13, 23]. Although, decreased expression of ARID1A protein has been associated with enhanced invasion and metastasis of colorectal carcinoma in mice and humans [6, 10]; however, there is limited evidence for the ARID1A role in regulating E-cadherin expression in CRC. To address this issue, we sought to delineate the impact of ARID1A knockdown and overexpression on the expression of E-cadherin protein and mRNAs in CRC cells. Here, we demonstrated that the expression of E-cadherin was strongly correlated with that of ARID1A. shRNA-mediated knockdown of ARID1A significantly downregulated E-cadherin expression in HCT116 and HT29/219 cells. Conversely, in the SW48 cells transfected with ARID1A overexpression vector, E-cadherin expression was significantly upregulated. These results are in good agreement with the findings from the only available experimental study in CRC, in which Mathur.et al. indicated the E-cadherin downregulation in HCT116 isogenic lines with monoallelic (ARID1A+/−) or biallelic (ARID1A−/−) deletion of ARID1A [6]. Importantly, our results were also consistent with data from the previous studies, which suggest that ARID1A knockdown decreased the expression of E-cadherin, whereas ARID1A overexpression increased the expression of E-cadherin in human gastric, breast and hepatocellular cancer cells [12, 13, 24]. Moreover, ARID1A knockdown strongly downregulated E-cadherin expression in cells derived from nasopharyngeal, neuroblastoma and pancreatic carcinomas [23, 25, 26]. Interestingly, Yan.et al. indicated that the expression of ARID1A was strongly correlated with that of E-cadherin in the analyzed gastric cancer tissues. Expression of ARID1A and E-cadherin were synergistically downregulated in 23.5% of analyzed gastric cancer tissues [13]. Notably, controversial report also exists. In a clinical study of esophageal squamous cell carcinoma, Ozawa et al. found that ARID1A expression status did not significantly correlate with E-cadherin expression levels of the patients [27]. The lack of rescue experiments to prove and justify the obtained results is one of the limitations of our study. Another limitation of this study was the lack of immunohistochemical analysis in clinical samples and animal experiments. Further investigations to provide more comprehensive mechanisms of the correlation between ARID1A and E-cadherin expression status in CRC tissues are under consideration for future study. However, additional studies would be helpful to explore the exact mechanism by which ARID1A regulates E-cadherin, such as ChIP-sequencing of histone markers of repression and activation coupled with ATAC-seq [28] will be useful to examine the interplay of ARID1A and chromatin accessibility of E-cadherin in CRC cells or detecting of ARID1A effects on transcriptional repressors of E-cadherin such as Slug, Zeb, Twist, Snail and some microRNAs especially the miR-200 family [15, 16].
Considering that E-cadherin downregulation marks the initiation of EMT and cancer cells invasion and metastasis, we postulated that E-cadherin downregulation by ARID1A silencing may partly account for enhanced migration activity of CRC cells. Therefore, using wound healing, we also noted that knockdown of ARID1A significantly enhanced the migration activity of the HCT116 cells, consistent with the data reported in pancreas, kidney, neuroblastoma, gastric, breast, and liver cancers, in which loss of ARID1A promoted cell migration and invasion [11, 13, 14, 24, 29, 30].
β-catenin, a major component of the canonical Wnt signaling pathway, has a crucial role in the negative regulation of E-cadherin expression and the EMT induction [19, 31,32,33]. Upon Wnt stimulation, β-catenin is stabilized, translocate to the nucleus where it triggers constitutive activation of Wnt target genes [32]. Aberrant activation of the Wnt pathway underlies many cancers including CRC [19]. In this study, ARID1A knockdown or overexpression did not alter the expression level of β-catenin. This finding is in agreement with a previous study in human gastric cells, in which the expression of β-catenin remained largely stable regardless of ARID1A silencing [13]. With respect to the ARID1A role in the regulation of β-catenin, the documented results to date are controversial. For instance, in the only available experimental study in CRC, Mathur.et al. found that Arid1afl/fl mice drive colorectal tumorigenesis by a mechanism that is independent of Wnt signaling and distinct from established genetic models of colon cancer [6]. In the mentioned research, β-catenin localized exclusively outside of the nucleus in mice colon tumors, indicating that ARID1A inactivation does not cooperate with aberrant Wnt signaling in driving tumorigenesis [6]. In contrast, ARID1A silencing in human gastric and neuroblastoma cells induced disassociation of the E-cadherin/ β-catenin adhesion complex at the membrane and a redistribution of β-catenin from the plasma membrane to the nucleus, where β-catenin triggers the EMT process [13, 25]. Additional studies are needed to elucidate the effect of the manipulation of ARID1A expression on nuclear translocation of β-catenin and the activation of Wnt signaling in CRC cells.
Conclusion
Our findings, together with those of previous studies, further demonstrate that ARID1A regulates E-cadherin expression in cancer cells. Considering that downregulation or loss of E-cadherin expression can activate EMT, therefore, our data suggest that ARID1A downregulation may contribute to enhancing the migratory and invasive abilities of CRC cells. In addition, we present evidence for the role of E-cadherin as one potential therapeutic target for ARID1A-deficient tumors as it was affected by ARID1A re-expression. Although, low ARID1A expression is not rare in colorectal cancer tumors [9, 34,35,36], the correlation between ARID1A and E-cadherin expression status in CRC tissue samples remains a goal for future studies.
Data availability
All data are included in this article and datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Bray F et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68(6):394–424
Siegel RL et al (2020) Colorectal cancer statistics, 2020. CA Cancer J Clin 70(3):145–164
Kranenburg O, Speeten KVD, Hingh ID (2021) Peritoneal metastases from colorectal cancer: defining and addressing the challenges. Front Oncol 11:639
Filip S et al (2020) Distant metastasis in colorectal cancer patients—do we have new predicting clinicopathological and molecular biomarkers? A comprehensive review. Int J Mol Sci 21(15):5255
Mathur R (2018) ARID1A loss in cancer: towards a mechanistic understanding. Pharmacol Ther 190:15–23
Mathur R et al (2017) ARID1A loss impairs enhancer-mediated gene regulation and drives colon cancer in mice. Nat Genet 49(2):296
Jones S et al (2012) Somatic mutations in the chromatin remodeling gene ARID1A occur in several tumor types. Hum Mutat 33(1):100–103
Network CGA (2012) Comprehensive molecular characterization of human colon and rectal cancer. Nature 487(7407):330
Erfani M et al (2020) Altered ARID1A expression in colorectal cancer. BMC Cancer 20:1–13
Kishida Y et al (2019) Associations between loss of ARID1A expression and clinicopathologic and genetic variables in T1 early colorectal cancer. Am J Clin Pathol 152(4):463–470
Somsuan K et al (2019) ARID1A knockdown triggers epithelial-mesenchymal transition and carcinogenesis features of renal cells: role in renal cell carcinoma. FASEB J 33(11):12226–12239
Wang T et al (2020) Role of ARID1A in epithelial-mesenchymal transition in breast cancer and its effect on cell sensitivity to 5-FU. Int J Mol Med 46(5):1683–1694
Yan H-B et al (2014) Reduced expression of the chromatin remodeling gene ARID1A enhances gastric cancer cell migration and invasion via downregulation of E-cadherin transcription. Carcinogenesis 35(4):867–876
Wang W et al (2019) ARID1A, a SWI/SNF subunit, is critical to acinar cell homeostasis and regeneration and is a barrier to transformation and epithelial-mesenchymal transition in the pancreas. Gut 68(7):1245–1258
Ramirez Moreno M, Stempor PA, Bulgakova NA (2021) Interactions and feedbacks in E-cadherin transcriptional regulation. Front Cell Dev Biol 9:1685
Brabletz T et al (2018) EMT in cancer. Nat Rev Cancer 18(2):128–134
Pastushenko I, Blanpain C (2019) EMT transition states during tumor progression and metastasis. Trends Cell Biol 29(3):212–226
Kuroda H et al (2009) Simplified lentivirus vector production in protein-free media using polyethylenimine-mediated transfection. J Virol Methods 157(2):113–121
Vu T, Datta P (2017) Regulation of EMT in colorectal cancer: a culprit in metastasis. Cancers 9:171
Ramesh V, Brabletz T, Ceppi P (2020) Targeting EMT in cancer with repurposed metabolic inhibitors. Trends Cancer 6(11):942
Loh C-Y et al (2019) The E-cadherin and N-cadherin switch in epithelial-to-mesenchymal transition: signaling, therapeutic implications, and challenges. Cells 8(10):1118
Druzhkova I et al (2019) E-cadherin in colorectal cancer: relation to chemosensitivity. Clin Colorectal Cancer 18(1):e74–e86
Yang Y et al (2019) Loss of ARID1A promotes proliferation, migration and invasion via the Akt signaling pathway in NPC. Cancer Manag Res 11:4931
He F et al (2015) Decreased expression of ARID1A associates with poor prognosis and promotes metastases of hepatocellular carcinoma. J Exp Clin Cancer Res 34(1):47
Li C et al (2017) ARID1A gene knockdown promotes neuroblastoma migration and invasion. Neoplasma 64(3):367–376
Tomihara H et al (2021) Loss of ARID1A promotes epithelial-mesenchymal transition and sensitizes pancreatic tumors to proteotoxic stress. Can Res 81(2):332–343
Ozawa Y et al (2015) Decreased expression of ARID1A contributes to infiltrative growth of esophageal squamous cell carcinoma. Tohoku J Exp Med 235(3):185–191
Buenrostro JD et al (2013) Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods 10(12):1213
Sun X et al (2017) Arid1a has context-dependent oncogenic and tumor suppressor functions in liver cancer. Cancer Cell 32(5):574–589.e6
Takao C et al (2017) Downregulation of ARID1A, a component of the SWI/SNF chromatin remodeling complex, in breast cancer. J Cancer 8(1):1
Kim WK et al (2019) β-catenin activation down-regulates cell-cell junction-related genes and induces epithelial-to-mesenchymal transition in colorectal cancers. Sci Rep 9(1):1–15
Kaszak I et al (2020) Role of cadherins in cancer—a review. Int J Mol Sci 21(20):7624
Basu S, Cheriyamundath S, Ben-Ze’ev A (2018) Cell–cell adhesion: linking Wnt/β-catenin signaling with partial EMT and stemness traits in tumorigenesis. F1000Research 7:1488
Wei X-L et al (2014) Clinicopathologic and prognostic relevance of ARID1A protein loss in colorectal cancer. World J Gastroenterol WJG 20(48):18404
Chou A et al (2014) Loss of ARID1A expression in colorectal carcinoma is strongly associated with mismatch repair deficiency. Hum Pathol 45(8):1697–1703
Lee SY et al (2015) Loss of AT-rich interactive domain 1A expression in gastrointestinal malignancies. Oncology 88(4):234–240
Acknowledgements
The authors would like to express their sincere gratitude to Shiraz University of Medical Sciences for financially supporting the study (No. 17288).
Funding
This work was supported by the Shiraz University of Medical Sciences [Grant number: 17288].
Author information
Authors and Affiliations
Contributions
PM and ME contributed to the study conception and design. Material preparation, data collection and analysis were performed by ME, MZ, SYH, ZM and SMS. The first draft of the manuscript was written by ME and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest.
Ethical approval
Ethics approval was obtained from the Ethics Committee of Shiraz University of Medical Sciences prior to this study (IR.SUMS.REC.1398.1390).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Erfani, M., Zamani, M., Hosseini, S.Y. et al. ARID1A regulates E-cadherin expression in colorectal cancer cells: a promising candidate therapeutic target. Mol Biol Rep 48, 6749–6756 (2021). https://doi.org/10.1007/s11033-021-06671-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-021-06671-9