Abstract
Long Non-Coding RNAs (lncRNAs), with diagnostic and therapeutic applications in malignancies, are newly described tumour-related molecules. Here, we reported the importance of circulating DSCAM-AS1 as the biomarker to detect Estrogen Receptor (ER)-positive breast cancer (BC) cases. Moreover, the expression of a BC-associated lncRNAs, namely DSCAM-AS1, was measured in tumoural and Paired Adjacent Non-Tumoral (PANT) tissue, as well as plasma, using Real-Time Polymerase Chain Reaction (RT-PCR). Besides, the correlations between gene expression and the clinicopathological features were analyzed. The diagnostic power of circulating DSCAM-AS1 in BC was estimated using the Area Under the Curve (AUC) value. Furthermore, we studied the DSCAM-AS1 associated with the network of competitive endogenous RNA (ceRNA) in BC using the literature review and in silico analysis. We found a significant increase in the expression levels of lncRNA in the tumour (P < 0.001) and in plasma (P < 0.001) of ER-positive BC patients. The sensitivity and specificity of DSCAM-AS1 in plasma for detection of BC from healthy controls were 100 and 97%, respectively (AUC = 0.98, P < 0.001). Accordingly, we suggest an elevated level of circulating DSCAM-AS1 as a candidate biomarker of ER-positive BC patients. Moreover, perturbation of DSCAM-AS1, as a ceRNA, acts in the tumor progression and drug resistance by affecting different cell signaling.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
According to the report of the Non-Communicable Diseases Research Center (NCDRC) of Iran, breast cancer (BC) is the most prevalent type of cancer among females in Kermanshah in western Iran [1].
The main drawbacks of BC therapy are the delayed diagnosis, tumour metastasis, and recurrence. Accordingly, early detection of BC could be a key factor in improving the prognosis and reducing the mortality rate. Blood-based testing is considered as an ideal strategy for discovering cancer biomarkers due to its easiness and non-invasive nature [2]. Though, measuring conventional serum biomarkers for BC diagnoses, such as measuring Carbohydrate Antigen (CA125) and Carcino-Embryonic Antigen (CEA), could not be employed for early diagnoses [3, 4]. That is why the identification of biomarkers is essential for early diagnosis and monitoring the BC progression.
Long Non-Coding RNAs (lncRNAs), with a length longer than 200 nucleotides, are known to be connected with various biological functions and known to have regulatory roles in the progression of multiple cancers, either as oncogenes or tumour suppressor genes [5, 6]. Evidence proposed lncRNAs could behave like a miRNA-sponge by competing with the endogenous circulatory RNAs (ceRNAs) activity [7]. Uncovering the crosstalk between this novel class of RNAs with other components of the gene regulatory system would bring insight into their role in development and disease [8].
DSCAM-AS1 is an estrogen receptor (ER)-dependent lncRNA that is located within the sequence of Down Syndrome Cell Adhesion Molecule (DSCAM) gene and expresses as an antisense intronic transcript. Knockdown of the DSCAM-AS1 gene shows similar effects as silencing of ER, which includes an increase in apoptosis, reduction of cell growth, and also induction of Epidermal-Mesenchymal-Transition (EMT) markers without effect on the ER expression [9].
In normal condition, estrogen receptor alpha (ERa) is expressed in about 10–15% of luminal epithelial breast cells. The activity of estrogen is mediated by ERa and ERb. Evaluation of ER levels has been used as a predictive and prognostic marker in the luminal BC subtype, which includes 75% of BC cases [10,11,12].
Currently, the most comprehensive endocrine therapy of BC is the prescription of Aromatase Inhibitors (AIs), which block the estrogen biosynthesis [13]. Despite its efficacy, therapeutic failure occurs in about 25% of breast cancer patients with positive ERa. The reason might be the emergence of de novo or acquired endocrine resistance [11]. Interestingly, a Super Enhancer (SE) has been located in the proximity of the DSCAM-AS1 which is occupied by apoERα. Mostly, the estrogen therapy does not affect the apoERα activity, though E2 (17β-estradiol) enhances the binding of ERα to the DSCAM-AS1 promoter [11].
The present study aimed to investigate the levels of circulating DSCAM-AS1 in plasma and tumour tissue of BC patients to find the related potential role as a novel biomarker. Furthermore, we constructed the DSCAM-AS1 associated ceRNA network and its possible role in breast cancer.
Materials and methods
Participants and sample collection
All the participants were female with Kurdish background from Kermanshah, the west of Iran. The mean age of BC patients was 49 years (range 30–68 years). Also, healthy volunteer women with the mean age of 48 years (26–64 years, P = 0.14) consisted of our control group. The BC cases were selected at the time of the diagnosis at the Bistoon Hospital, Kermanshah Province, Iran, from 2016 to 2017. This study was approved by the ethics committee of Kermanshah University of Medical Sciences, and informed consent was obtained from all subjects. Only histologically confirmed breast cancer cases by an expert pathologist, without any previously diagnosed malignancies, were entered in the study. The first group of samples consisted of 20 tumoural and Paired Adjacent Non-Tumoural (PANT) tissue of BC patients, which was taken at the time of BC diagnosis (from frozen samples). The second group was 40 patients who peripheral blood was taken from them during the period of diagnosis (before surgery and any therapy). Accordingly, the blood samples were provided from a control group that consisted of 40 healthy women (third group). Immunohistochemical tests were done to determine the Estrogen Receptor (ER) or Progesterone Receptor (PR) in the clinical laboratory of Bistoon hospital, then read and confirmed by an expert pathologist. Breast tumors were classified as ER-positive or PR-positive if staining was present in 1% or more of tumor nuclei.
Stock of plasma samples
The blood samples were kept in EDTA containing tubes. To separate cellular fraction from plasma, samples were centrifuged at 3500×g for 20 min, within about one hour after blood collection.
RNA extraction and complementary DNA (cDNA) synthesis
Fresh frozen tissues were powdered in liquid nitrogen by mortar and pestle and then total RNA was isolated using Qizol reagent (QIAGEN, US) according to the manufacturer’s procedure. Also, total RNA was extracted from 200 μl plasma using the miRNeasy Serum/Plasma Kit (QIAGEN, US) and was eluted into 14μl of elution solution. The concentration and purity of isolated RNA were measured by a Nanodrop spectrophotometer (Thermo Scientific, Waltham, MA). cDNA was synthesized from total extracted RNA using a QuantiTect Reverse Transcription Kit (QIAGEN, US) and the synthesized cDNA was stored at − 20 °C until further processing.
Quantitative Real-Time PCR (qRT-PCR)
The primers were chosen from the literature and then checked for their accuracy and specificity at the NCBI data bank (https://www.ncbi.nlm.nih.gov/). The sequences of selected primers used for qRT-PCR were as follow, AGAGCGAAACCCCATCTCAA (sense) and CTGAGAGATCCCCTGTAGCG (antisense) for DSCAM-AS1 [14], and CTCGCTTCGGCAGCACA (sense) and AACGCTTCACGAATTTGCGT (antisense) for U6-snRNA [15] as an internal control. The qRT-PCR assay was performed on a Rotor-gene machine (Corbett Research) with SYBR Premix Ex Taq II (TaKaRa, China). The PCR reaction conditions were: denaturation at 95 °C for the 30s as the initial step, followed by 45 PCR cycles at 95 °C for 10s, 60 °C for 15s and 72°C for 30s. All reactions were done in duplicate. The relative expression of the DSCAM-AS1 gene in each tumoural tissue was compared with its PANT. The cycle threshold (Ct) level is inversely proportional to the target gene expression. Relative expression of the genes was assessed by △Ct values (Ct reference gene − Ct target gene). Fold changes of the gene expression in the cancer samples relative to the healthy controls were determined by the “2−△△Ct method”.
Bioinformatics analysis and literature review
For finding miRNAs related to the DSCAM-AS1, we conducted an in silico search in the LncBase database (v.2) [16]. The prediction modules were set using the following search strategies: Filters: Threshold = 0.7, Tissue: mammary glands; Category: cancer/malignant and Experimental module were as follow: Filters: Tissue: mammary glands; miRNA Species: Homo sapiens; Validation type: direct; Validated as: positive; Source: LncBase v.2. Then, mRNAs targeted by the detected miRNAs were identified in the miRDB database [17, 18]. The prediction was limited to human species and gene targets with ‘target prediction score’ less than 90 were excluded. The predicted mRNAs then further enriched using the Enrichr database [19].
Furthermore, in order to add more data about DSCAM-AS1: miRNAs: mRNAs connections, we conducted a literature review in the PubMed and Scopus with these keywords: (TITLE-ABS-KEY (DSCAM-AS1) AND TITLE-ABS-KEY ("breast cancer") OR TITLE-ABS-KEY ("breast carcinoma")) for finding the relation between DSCAM-AS1 and breast cancer, and then separately we searched for the relation between breast cancer and each DSCAM-AS1 related miRNA with this pattern, for instance: (TITLE ("breast cancer") AND TITLE (mir-27a) OR TITLE (mirna-27a) OR TITLE (mir-27a-3p)). For more assurance, all searches were performed by two researchers, blindly.
KEGG pathway and gene ontology (GO) analysis
Functional enrichment can be useful to find out the functions of genes. The GO functional annotation and KEGG pathway analysis for miRNA target mRNAs was performed by the web-based tool, Enrichr (https://amp.pharm.mssm.edu/Enrichr). Gene ontology categorizes genes based on cellular component, biological process, and molecular function. Potential biological pathways were identified through the analysis of the enriched genes in the KEGG database (https://www.genome.jp/kegg/). Gene ontology terms and KEGG pathways with p < 0.05 set as the cut-off to pick out significantly enriched terms and pathways.
Statistical analysis
T-test was used to assess the significance of the difference in the expression of DSCAM-AS1 between tumoural vs. paired-adjacent tissues or between plasma of patients vs. healthy controls. The relation among tumour features and expression of DSCAM-AS1 was appraised using the Chi-square test. P-values <0.05 were assumed as significant. The diagnostic power of circulating DSCAM-AS1 was estimated using the Receiver Operating Characteristic (ROC) curve. The Area Under the Curve (AUC) value were used for the decision about if the proposed biomarker is suitable or not. SPSS software (version 16) was applied for statistical analysis.
Results
Some characteristics of the patients’ baseline profiles are indicated in Table 1.
The expression levels of DSCAM-AS1 was significantly higher in paired tumoural tissues compared with PANTs
Using qRT-PCR, the expression of DSCAM-AS1 was measured in 20 paired breast tumoural and adjacent non-tumoural tissues. Differences between each pair of the tumour and its normal margin tissues were significant by paired t-test (P < 0.001, Fig. 1).
The relative expression of the DSCAM-AS1 gene in tumoural tissue was compared with Paired Adjacent Non-tumoural (PANT) tissue. Relative expression was calculated by Delta Ct values (Ct reference gene − Ct target gene). Ct (Cycle threshold) level was inversely proportional to the amount of the target gene in the sample
The plasma levels of circulating DSCAM-AS1 was significantly higher in BC patients
The relative level of circulating DSCAM-AS1 was determined in plasma samples. We found that in the plasma samples of patients, the level of DSCAM-AS1 was significantly higher than in the cancer-free donors (P < 0.001). Moreover, it was higher in ER-positive BC patients compared to that of ER-negative patients (P < 0.001, Fig. 2).
Comparisons of circulating DSCAM-AS1 levels in the controls (Ctrl), all breast cancer patients (patients), Estrogen receptor-positive patients (ER+), Estrogen receptor-negative patients (ER-), progesterone receptor-positive patients (PR+), and human epidermal growth factor receptor 2-positive patients (HER2+). Relative Quantitation (RQ) was calculated by 2−△△Ct method
DSCAM-AS1 as a diagnostic tool to discriminate ER-positive from ER-negative BC patients
To determine the possibility of using plasma levels of DSCAM-AS1 as a diagnostic tool, ROC curves were used. As shown in Fig. 3, the value of the AUC was 0.98 for the cancer-free group compared to that of the BC group. In this model, optimal cutoff points of DSCAM-AS1 were 6.76 (with 100% sensitivity and 97% specificity). Comparing cancer-free group with ER-positive BC group, AUC was 0.99 and optimal cutoff points were found to be 6.76 (with 100% sensitivity and 97% specificity). In the comparison between ER-positive and ER-negative groups, the value of the AUC for DSCAM-AS1 was 1.00 and the cutoff point was 6.76, where both sensitivity and specificity were 100% (Table 2).
The ROC curve was plotted for assessment of the diagnostic power of circulating DSCAM-AS1 as a candidate biomarker in breast cancer patients (regardless of ER status) versus healthy controls/ ER-negative patients, and also ER-positive patients versus healthy controls/ ER-negative patients. ROC receiver operating characteristic, ER estrogen receptor
Bioinformatics analysis and literature review revealed six miRNAs related to DSCAM-AS1
The LncBase software (v.2) prediction module identified 145 miRNAs with binding sites on the DSCAM-AS1 transcript (Supplementary File 1). The Experimental module of LncBase software (v.2) showed that DSCAM-AS1 is a verified target for four miRNAs (hsa-miR-130a-3p, hsa-miR-27a-3p, hsa-miR-301a-3p, and hsa-miR-193a-3p). Pilli et al. (2014) experimentally validated these four miRNAs as potential targets of DSCAM-AS1 in the mammary gland tissue and MCF7 cell line, using an immunoprecipitation method, HITS-CLIP [20]. Although, in the literature review we found only a report for hsa-mir-137 [21] and hsa-mir-204-5p [22] as direct regulators of DSCAM-AS1 in breast cancer.
Supplementary file 2 contains the predicted genes targeted using these six miRNAs (miR-130a-3p, miR-27a-3p, miR-301a-3p, miR-193a-3p, miR-137, and miR-204-5p) according to the miRDB databank. In total, 910 unique genes are potential targets of these six miRNAs. Enrichment analysis for the predicted genes (with a prediction score > 60) conducted in the KEGG database (2016), showed that they are involved in different signaling pathways (Figs. 4, 5). In brief, mRNAs targeted by miR-130a-3p and miR-301a-3p are involved in the mTOR signaling pathway, and mRNAs targeted using miR-204-5p are involved in the estrogen signaling pathway. It was predicted that some proteins of the ErbB signaling pathway are targeted by miR-137, miR-27a-3p, and miR-193b-3p. Table 3 illustrates experimentally validated mRNAs targeted using the six miRNAs in the literature. In brief, ESR1 (estrogen receptor 1 which encodes Era) is the target of 3 miRNAs related to DSCAM-AS1 (miR-193b-3p, miR-301a-3p, and miR-27a-3p). In addition, miR-193b, miR-27a-3p, and miR-137 are involved in estrogen signaling through affecting AKR1C2, ZBTB10, and ERRA, respectively. Phosphatase and tensin homolog (PTEN) is another important gene in the development of breast cancer, which is targeted by miR-301a-3p and miR-130a-3p. Many of the proteins affected through miRNAs related to DSCAM-AS1 are involved in tumour growth and metastasis such as PTEN, RAB5B, FOSL1, SFRP1, SPRY2, and FSTL1.
ERα binds to super enhancers (SEs) upstream of the DSCAM-AS1 gene to regulate the expression of this gene. This study revealed six miRNAs which have binding sites on DSCAM-AS1. Each of these miRNAs has binding sites on some mRNAs which involved in a specific signaling pathway. Abbreviation: ERa, estrogen receptor alpha; SE, super enhancer; miR, micro RNA
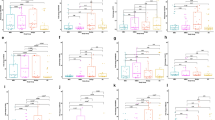
KEGG pathways and GO classification of miRNA target genes
The 910 target genes which had a prediction target score equal 90 or more were subjected to the KEGG pathway and GO analysis (Supplementary File 3). In the KEGG analysis, two significantly enriched pathways were including ‘ErbB signaling pathway’ (Combined Score (CS) = 81.5, P = 1.5 × 10-7) and ‘Proteoglycans in cancer’ (CS = 49.8, P = 3.7 × 10-7). The top 15 enriched KEGG pathways and GO items, based on the Enrichr combined score (CS), are displayed on Table 4. Through GO annotation and enrichment analysis, the roles of gene products from the cellular component, biological process, and molecular function were identified. The most significant term in GO cellular component was ‘RISC complex’ (GO:0016442, CS = 115.7 P = 1.4 × 10-4) and RNAi effector complex (GO:0031332, CS = 115.7 , P = 1.4 × 10-4), that in GO biological process was ‘positive regulation of transcription, DNA-templated’ (GO:0045893, CS = 68.7, P = 1.3 × 10-13), and that in GO molecular function was ‘transcription regulatory region DNA binding’ (GO:0044212, CS = 33.4 , P = 1.7 × 10-6). To better illustrate gene enrichment in the three mentioned GO categories, a clustergram was constructed consisting of the top 20 enriched GO terms, according to combined score (Fig. 6).
Discussion
The DSCAM-AS1 levels in tissue and plasma of BC patients were measured. The possible correlations between circulating DSCAM-AS1 and clinicopathological features were analyzed. Furthermore, the potential ceRNA network of DSCAM-AS1-miRNAs-mRNAs was investigated. According to our best knowledge, these results, for the first time, showed that the circulating level of DSCAM-AS1 might be useful as a potential novel biomarker for identifying ER-positive BC.
It is suggested that lncRNAs play an important role in cell differentiation and development [23, 24]. Significant correlations have been reported between malignancies and aberrant expression of lncRNAs [25, 26]. Since circulating lncRNAs are somehow stable in plasma/serum and their transcripts are the final functional production, assessing the lncRNA level could directly indicate the levels of the active product. Consequently, lncRNAs have a potential value as a bio-marker [23].
“Subtypes of breast tumour are defined using the immunohistochemical expression of Estrogen Receptor (ER), Progesterone Receptor (PR), and HER2” [27]. ERα-positive breast tumours have a good response to endocrine therapies. However, 25% of these cases show primary or acquired resistance which motivates researchers to develop novel biomarkers and therapeutic targets [28].
There are pieces of evidence indicating that the increased level of DSCAM-AS1 as a lncRNA is highly specific for luminal BC and, therefore, can be used as a biomarker for this subtype [11, 28]. Furthermore, it has been reported that increased expression of DSCAM-AS1 is related to worse overall survival rates among BC patients [29]. We found that in addition to an increase in the expression of DSCAM-AS1 in the ER-positive breast tumour tissues, the circulating DSCAM-AS1 level increased in the plasma of ER-positive BC patients compared to those healthy controls and ER-negative patients. Next, the ROC curve was used to find out if DSCAM-AS1 could take a role as a tumour biomarker. The ROC curve demonstrated the sensitivity and specificity of DSCAM-AS1 (100% and 96.77%) for ER-positive BC vs. ER-negative patients, respectively.
To further investigate the function of this transcript in the development of breast cancer, we employed bioinformatics approaches. Our findings suggested that lncRNA DSCAM-AS1 might be involved in mTOR, MAPK, estrogen, and ErbB signaling pathways through at least six miRNAs (hsa-miR-130a-3p, hsa-miR-27a-3p, hsa-miR-301a-3p, hsa-miR-193a-3p, has-miR137, and has-miR-204-5p). Four of them (hsa-miR-130a-3p, hsa-miR-27a-3p, hsa-miR-301a-3p, and hsa-miR-193a-3p) were validated as potential targets of DSCAM-AS1 by an immunoprecipitation method, namely HITS-CLIP, in mammary gland tissue, and MCF7 cell line [20]. Moreover, hsa-mir-137 and hsa-mir-204-5p as direct regulators of DSCAM-AS1 in breast cancer were introduced by Ma et al. [21] and Liang et al. [22], respectively. According to the theory of ceRNA [30, 31], lncRNA DSCAM-AS1 as a ceRNA can attenuate the activity of the miRNAs that interfere in the signaling pathways involved in breast cancer development. ErbB receptors control key pathways which regulate cellular processes. In tumours, activated ErbB2 stimulates several intracellular pathways, e.g. the mTOR, PI3K/Akt, and MAPK pathways [32]. There are interactions between the ER and PI3K/mTOR. Activation of the mTOR and ER pathways up-regulates cyclin D to trigger cell cycle progression [32]. Since the ectopic expression of DSCAM-AS1 in ER-positive BC patients could decrease the level of related miRNAs, probably upregulation of the predicted mRNAs leads to activation of the ErbB signaling pathway among these patients.
According to the literature, ESR1 is targeted using three DSCAM-AS1-related miRNAs (miR-193b-3p, miR-301a-3p, and miR-27a-3p) [33,34,35]. Pillai et al. identified miR-193a/b-3p as a novel regulator of the ER pathway, which targets multiple genes involved in ER signaling [20]. The miR193b-3p could suppress the local production of estrogens and other steroid hormones using the direct targeting of 5′-Un-Translated Region (5′-UTR) of AKR1C2 [33]. The miR-27a-3p targets ESR1 as well as indirectly control E2-responsiveness in MCF-7 cells using the repression of ZBTB10, which increases the expression of ERα [36]. These results can explain the peculiar correlation between ERα and DSCAM-AS1 expression. On the other hand, miR-301a-3p and miR-130a-3p through targeting PTEN, and miR-137 could trigger the Wnt/β-catenin pathway and thereby promote BC invasion and metastasis through targeting PTEN and Follistatin Like Protein 1 (FSLT1), respectively. Moreover, it could regulate stemness and chemo resistance [37, 38].
Our findings proposed a candidate circulating biomarker, namely DSCAM-AS1, for clinical assessment of ER-positive breast cancer patients. Presumably, perturbation of DSCAM-AS1, as a ceRNA, acts in the progression of breast cancer and drug resistance by affecting different cell signaling. However, to verify these results more studies with larger sample sizes and among other ethnic groups are necessary as a future suggestion.
Abbreviations
- BC:
-
Breast cancer
- ceRNA:
-
Competing endogenous RNA
- DSCAM-AS1:
-
Down syndrome cell adhesion molecule anti-sense 1
- EMT:
-
Epidermal-mesenchymal transition
- ER:
-
Estrogen receptor
- lncRNAs:
-
Long noncoding RNAs
References
Moradi M-T, Hatami R, Rahimi Z (2020) Circulating CYTOR as a potential biomarker in breast cancer. International Journal of Molecular and Cellular Medicine (IJMCM) 9 (1):0–0
Elmore JG, Armstrong K, Lehman CD, Fletcher SW (2005) Screening for breast cancer Jama 293(10):1245–1256
Molina R, Barak V, van Dalen A, Duffy MJ, Einarsson R, Gion M, Goike H, Lamerz R, Nap M, Sölétormos G (2005) Tumor markers in breast cancer–European Group on Tumor Markers recommendations. Tumor Biology 26(6):281–293
Marić P, Ozretić P, Levanat S, Orešković S, Antunac K, Beketić-Orešković L (2011) Tumor markers in breast cancer–evaluation of their clinical usefulness. Collegium antropologicum 35(1):241–247
Qiu M-T, Hu J-W, Yin R, Xu L (2013) Long noncoding RNA: an emerging paradigm of cancer research. Tumor Biology 34(2):613–620
Spizzo R, Almeida MIe, Colombatti A, Calin GA, (2012) Long non-coding RNAs and cancer: a new frontier of translational research? Oncogene 31(43):4577
Liu H, Zhang Z, Wu N, Guo H, Zhang H, Fan D, Nie Y, Liu Y (2018) Integrative analysis of dysregulated lncRNA-associated ceRNA network reveals functional lncRNAs in gastric cancer. Genes 9(6):303
Tay Y, Rinn J, Pandolfi PP (2014) The multilayered complexity of ceRNA crosstalk and competition. Nature 505(7483):344
Cerk S, Schwarzenbacher D, Adiprasito JB, Stotz M, Hutterer GC, Gerger A, Ling H, Calin GA, Pichler M (2016) Current status of long non-coding RNAs in human breast cancer. Int J Mol Sci 17(9):1485
Badve S, Turbin D, Thorat MA, Morimiya A, Nielsen TO, Perou CM, Dunn S, Huntsman DG, Nakshatri H (2007) FOXA1 expression in breast cancer—correlation with luminal subtype A and survival. Clin Cancer Res 13(15):4415–4421
Miano V, Ferrero G, Rosti V, Manitta E, Elhasnaoui J, Basile G, De Bortoli M (2018) Luminal lncRNAs regulation by ERα-controlled enhancers in a ligand-independent manner in breast cancer cells. Int J Mol Sci 19(2):593
Deroo BJ, Korach KS (2006) Estrogen receptors and human disease. J Clin Investig 116(3):561–570
Chumsri S, Howes T, Bao T, Sabnis G, Brodie A (2011) Aromatase, aromatase inhibitors, and breast cancer. The Journal of Steroid Biochemistry and Molecular BIOLOGY 125(1–2):13–22
Sun W, Li AQ, Zhou P, Jiang YZ, Jin X, Liu YR, Guo YJ, Yang WT, Shao ZM, Xu XE (2018) DSCAM-AS 1 regulates the G1/S cell cycle transition and is an independent prognostic factor of poor survival in luminal breast cancer patients treated with endocrine therapy. Cancer Med 7(12):6137–6146
Li D, Li J (2016) Association of miR-34a-3p/5p, miR-141-3p/5p, and miR-24 in decidual natural killer cells with unexplained recurrent spontaneous abortion. Med Sci Monit 22:922
Paraskevopoulou MD, Vlachos IS, Karagkouni D, Georgakilas G, Kanellos I, Vergoulis T, Zagganas K, Tsanakas P, Floros E, Dalamagas T (2015) DIANA-LncBase v2: indexing microRNA targets on non-coding transcripts. Nucleic Acids Res 44(D1):D231–D238
Wong N, Wang X (2014) miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Res 43(D1):D146–D152
Liu W, Wang X (2019) Prediction of functional microRNA targets by integrative modeling of microRNA binding and target expression data. Genome Biol 20(1):18
Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, Koplev S, Jenkins SL, Jagodnik KM, Lachmann A (2016) Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res 44(W1):W90–W97
Pillai MM, Gillen AE, Yamamoto TM, Kline E, Brown J, Flory K, Hesselberth JR, Kabos P (2014) HITS-CLIP reveals key regulators of nuclear receptor signaling in breast cancer. Breast Cancer Res Treat 146(1):85–97
Ma Y, Bu D, Long J, Chai W, Dong J (2019) LncRNA DSCAM-AS1 acts as a sponge of miR-137 to enhance Tamoxifen resistance in breast cancer. J Cell Physiol 234(3):2880–2894
Liang WH, Li N, Yuan ZQ, Qian XL, Wang ZH (2018) DSCAM‐AS1 promotes tumor growth of breast cancer by reducing miR‐204‐5p and up‐regulating RRM2. Molecular Carcinogenesis
Xu N, Chen F, Wang F, Lu X, Wang X, Lv M, Lu C (2015) Clinical significance of high expression of circulating serum lncRNA RP11–445H22. 4 in breast cancer patients: a Chinese population-based study. Tumor Biology 36 (10):7659–7665
Fatica A, Bozzoni I (2014) Long non-coding RNAs: new players in cell differentiation and development. Nat Rev Genet 15(1):7
Zhao Y, Guo Q, Chen J, Hu J, Wang S, Sun Y (2014) Role of long non-coding RNA HULC in cell proliferation, apoptosis and tumor metastasis of gastric cancer: a clinical and in vitro investigation. Oncol Rep 31(1):358–364
Zhang E-B, Han L, Yin D-D, Kong R, De W, Chen J (2014) c-Myc-induced, long, noncoding H19 affects cell proliferation and predicts a poor prognosis in patients with gastric cancer. Med Oncol 31(5):914
Yang XR, Chang-Claude J, Goode EL, Couch FJ, Nevanlinna H, Milne RL, Gaudet M, Schmidt MK, Broeks A, Cox A (2010) Associations of breast cancer risk factors with tumor subtypes: a pooled analysis from the Breast Cancer Association Consortium studies. J Natl Cancer Inst 103(3):250–263
Miano V, Ferrero G, Reineri S, Caizzi L, Annaratone L, Ricci L, Cutrupi S, Castellano I, Cordero F, De Bortoli M (2016) Luminal long non-coding RNAs regulated by estrogen receptor alpha in a ligand-independent manner show functional roles in breast cancer. Oncotarget 7(3):3201
Xu S, Kong D, Chen Q, Ping Y, Pang D (2017) Oncogenic long noncoding RNA landscape in breast cancer. Molecular Cancer 16(1):129
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (2011) A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146(3):353–358
Moradi MT, Fallahi H, Rahimi Z (2019) Interaction of long noncoding RNA MEG3 with miRNAs: A reciprocal regulation. J Cell Biochem 120(3):3339–3352
Cortés J, Im S-A, Holgado E, Perez-Garcia JM, Schmid P, Chavez-MacGregor M (2017) The next era of treatment for hormone receptor-positive, HER2-negative advanced breast cancer: triplet combination-based endocrine therapies. Cancer Treat Rev 61:53–60
Leivonen S-K, Rokka A, Östling P, Kohonen P, Corthals GL, Kallioniemi O, Perälä M (2011) Identification of miR-193b targets in breast cancer cells and systems biological analysis of their functional impact. Mol Cell Proteomics 110:005322
Lettlova S, Brynychova V, Blecha J, Vrana D, Vondrusova M, Soucek P, Truksa J (2018) MiR-301a-3p Suppresses estrogen signaling by directly inhibiting ESR1 in ERα positive breast Cancer. Cell Physiol Biochem 46(6):2601–2615
Ljepoja B, García-Roman J, Sommer A-K, Wagner E, Roidl A (2019) MiRNA-27a sensitizes breast cancer cells to treatment with Selective estrogen receptor modulators. The Breast 43:31–38
Li X, Mertens-Talcott SU, Zhang S, Kim K, Ball J, Safe S (2010) MicroRNA-27a indirectly regulates estrogen receptor α expression and hormone responsiveness in MCF-7 breast cancer cells. Endocrinology 151(6):2462–2473
Ma F, Zhang J, Zhong L, Wang L, Liu Y, Wang Y, Peng L, Guo B (2014) Upregulated microRNA-301a in breast cancer promotes tumor metastasis by targeting PTEN and activating Wnt/β-catenin signaling. Gene 535(2):191–197
Wei H, Cui R, Bahr J, Zanesi N, Luo Z, Meng W, Liang G, Croce CM (2017) miR-130a deregulates PTEN and stimulates tumor growth. Can Res 0530:2017
Acknowledgements
We wish to show our appreciation to Dr. Sh. Mostafaei and Mr. E. Zereshki for their biostatistician consulting.
Funding
This study has been funded by Kermanshah University of Medical Sciences, Iran [Grant No. 95680].
Author information
Authors and Affiliations
Contributions
M-TM: Performed all experiments, analyzed the data and wrote the initial draft of the manuscript. HF: Contributed to bioinformatic analysis and revised the manuscript. ZR: Contributed to concept and design, financial support, and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Moradi, MT., Fallahi, H. & Rahimi, Z. The clinical significance of circulating DSCAM-AS1 in patients with ER-positive breast cancer and construction of its competitive endogenous RNA network. Mol Biol Rep 47, 7685–7697 (2020). https://doi.org/10.1007/s11033-020-05841-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-020-05841-5