Abstract
N6-methyladenosine (m6A) serves as a major RNA methylation modification and impacts the initiation and progression of various human cancers through diverse mechanisms. It has been reported that m6A RNA methylation is involved in different physiological and pathological processes, including stem cell differentiation and motility, immune response, cellular stress, tissue renewal and viral infection. In this review, the m6A modification and its regulatory functions in a few major cancers is introduced. The detection approaches for the m6A sites identification are discussed. Additionally, the potential of the RNA m6A modification in clinical application is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
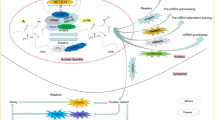
Chemical modifications of nucleobases, like methylation of adenine and cytosine, are critical for controls of gene expressions on different levels, which are important for cells to adapt to the environment or to develop to complex organisms from a single cell [1]. RNA methylation has been found in various RNAs including messenger RNA (mRNA) [2, 3], microRNA (miRNA) precursor [4] and long non-coding RNA (lncRNA) [5] transfer RNA [6,7,8,9], ribosomal RNA [10], small nuclear RNA [11]. The N6-methyladenosine (m6A) is one of the most common and abundant internal modifications on RNA molecules, which has been identified in eukaryotic mRNA [12,13,14] and virus nuclear RNA [15, 16] since 1970s. m6A modification is involved in almost all the stages of RNA life cycle, including RNA transcription, exporting through nuclear, translation and degradation [17,18,19,20], and has gained more and more attractions.
The adenosine methylation occurs preferentially within the degenerate consensus RRACH (R = G or A; H = A, C or U) in gene coding regions and 3′UTRs [21] or broader consensus motif DRACH (where D denotes A, G or U) [22]. The m6A modification has been considered as an epigenetic regulation similar to DNA and histone modifications, performing critical functions in important biological processes [23,24,25]. Accumulating evidences have supported the fact that the m6A RNA methylation impact on the initiation and progression of human cancers through diverse mechanisms [26,27,28]. It has been reported that the m6A RNA methylation is related to different processes, including stem cell differentiation and motility [29], immune response [30, 31], cellular stress [6], tissue renewal and pathology [32] and viral infection [33]. In the following sections, the m6A modification and its functional mechanism, and the m6A-associated modification landscapes in several major cancers are reviewed. The detection methods of m6A modification and the potential clinical application for RNA m6A modification are also discussed.
Survey methodology
In this review article, three steps of journal article searching were performed. First, the functional mechanism of RNA m6A methylation was identified using the search terms “RNA N6-methyladenosine” and terms “m6A writers, erasers, readers”. Moreover, terms “methyltransferase”, “demethylases” and “m6A-binding proteins” were also searched respectively. Then the functions of the RNA N6-methyladenosine modification in the cancer progression were further investigated by searching “RNA N6-methyladenosine” together with “cancer” and searching “RNA N6-methyladenosine” together with certain cancers. Finally, the development of the m6A modification detection was investigated by using the terms “m6A modification detection”. All relevant studies were included regardless of the year of publication. The research articles that were published before August 2018 based on the above search criteria were collected. In this publication survey, the PubMed databases were explored.
Functional mechanism of RNA m6A methylation
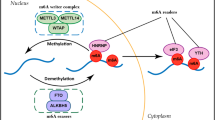
m6A modifications are dynamic and reversible, which is created by proteins called m6A “writers” and can be reversed by m6A “erasers” (Fig. 1). Some other proteins recognizing and binding m6A-containing mRNA serve as “readers” and regulate downstream molecular mechanisms accordingly.
Mechanism of m6A-based modification of RNA. m6A RNA methylation is regulated by its “writers,” “erasers,” and “readers.” Writers refer to the m6A methylase complex including METTL3, METTL14 and WTAP. Erasers are m6A demethylases involving FTO and ALKBH5. Readers are proteins that bind to m6A modifications, including YTH-domain containing proteins (YTHDF1, YTHDF2, YTHDF3, YTHDC1, YTHDC2), eIF3 and nuclear m6A reader protein, which can exert various functions in almost all stages of RNA life cycle. ALKBH5 a-ketoglutarate-dependent dioxygenase alkB homolog 5, eIF3 eukaryotic initiation factor 3, FTO fat mass and obesity-associated protein, m6A N6-methyladenosine, METTL3 methyltransferase like 3, METTL14 methyltransferase like 14, WTAP Wilms’ tumour 1-associating protein, YTH YT521B homology
m6A writers
The RNA methylation modification is catalyzed by the m6A writer complex [34, 35]. Methyltransferase like 3 (METTL3, also known as MT-A70) and methyltransferase like 14 (METTL14) both belong to the class I methyltransferase family30 with methyltransferase domains and efficiently catalyses methyl group transfer by forming a core catalytic complex [36, 37]. METTL3 appears to be a predominantly catalytic enzyme with a function reminiscent of N6-adenine methyltransferase systems [38], while METTL14 was reported to be a pseudomethyltransferase to stabilize METTL3 and recognize target RNA [39]. Wilms’ tumour 1-associating protein (WTAP) is a major regulatory component of the m6A methylation complex by interacting with METTL3 and METTL14 and helping their localization into nuclear speckles [40]. The METTL3 catalytic process has been considered as a common m6A pathway and modifies most m6A sites in mRNA [41]. Generally, the m6A writer complex can exert their catalytic function with or without other factors.
Additional subunits, like KIAA1429 (also known as vir-like m6A methyltransferase associated, or VIRMA) [42], RBM15 and its paralog RBM15B [43] are also associated with the methylation complex. Another methyltransferase METTL16 was recently found to be a m6A-forming enzyme in mRNA and mediate m6A formation in snRNA U6 by regulating methionine adenosyltransferase 2A intron retention in response to intracellular S-adenosylmethionine levels [44]. Meanwhile, the SpoU-TrmD RNA methyltransferase superfamily (including TrmH and TrmD) [45] and NSun6 [46] were reported to catalyze the transfer of the methyl group to tRNA.
m6A erasers
m6A can be demethylated by m6A demethylases like fat mass and obesity-associated protein (FTO) [47, 48] and a-ketoglutarate-dependent dioxygenase alkB homolog 5 (Alkbh5) [49, 50], which are described as m6A erasers. Both FTO and ALKBH5 belong to the AlkB family of nonheme Fe(II)/2-oxoglutarate dioxygenase. FTO is first shown as a human obesity susceptibility gene and was found to influences RNA processing through dynamic demethylation of m6A [51, 52]. ALKBH5 has catalytic domain that can demethylate both single stranded RNA (ssRNA) and single-stranded DNA (ssDNA), and specially catalyze the demethylation of m6A in ssRNA, supporting the reversible m6A modification on RNA [50, 53].
m6A readers
The m6A functions mainly with recruiting m6A-binding proteins. It can be recognized by proteins containing a YT521B homology (YTH) domain or by the translation initiation factor eukaryotic initiation factor 3 (eIF3) [41, 54]. The YTH domain characterized by 14 invariant residues within an α-helix/β-sheet structure recognizes m6A by a conserved aromatic cage sandwiched between a Trp and Tyr residue and with the methyl group pointed to another Trp residue [55, 56].
There are five proteins that contain the YTH domain in human cells, including YTH domain family (YTHDF1–3), YTH domain-containing 1 (YTHDC1) and YTH domain-containing 2 (YTHDC2). Among these proteins, YTHDF1–3 are cytoplasmic m6A-specific binders and highly similar to each other with a C-terminally located YTH domain [57]. Structural basis analysis revealed that YTHDF1 can recognize m6A without sequence preference and facilitates the translation of m6A-modifed transcripts [58]. Compared with YTHDF1, YTHDF2 was identified to selectively bind to mRNAs (also non-coding RNAs) with a conserved core motif of G(m6A)C, accelerating the decay of m6A-modifed RNAs [57, 59, 60]. YTHDF3, another member of the cytoplasmic YTH family, was reported to facilitate the translation of targeted mRNAs and affect methylated mRNA decay in synergy with YTHDF1, by binding to m6A-modifed mRNAs and interacting with 40S and 60S ribosome subunits [61, 62].
In contrast to the YTH domain family members, YTHDC1 is predominantly located in nuclear harboring a distinctly selective binding pocket of GG(m6A)C sequence. It prefers guanosine and disfavors adenosine at the position preceding the m6A nucleotide modification [58, 63]. YTHDC2 is a nucleocytoplasmic protein containing a DEAD-box RNA helicase domain. Recent studies have shown that YTHDC2 has a RNA-induced 3′ → 5′ RNA helicase activity and regulates m6A transcripts by recruiting the 5′/3′ exoribonuclease XRN1 via Ankyrin repeats [64, 65].
eIF3 has been considered as a m6A-binding protein as well. In the absence of the cap-binding factor eIF4E, mRNAs containing 5′ UTR m6A can be translated in a cap-independent manner by recruiting the 43S complex to initiate translation via directly binding to eIF3 [66]. However, unlike the YTH proteins which recognize m6A through a specific well-characterized YTH domain, the exact mechanism of eIF3-m6a recognition is not clearly understood yet. It probably relies not only on the sequence motif but also on adjacent RNA structures [67].
Another protein, heterogeneous nuclear ribonucleoprotein A2/B1 (HNRNPA2B1), has been identified as a nuclear reader of m6A, binding to RRAC containing sites on nuclear RNAs [68]. HNRNPA2B1 regulates the alternative splicing of exons, which is highly correlated with METTL3 [68]. Moreover, HNRNPA2B1 interacts with the pri-miRNA microprocessor complex component DGCR8 and modulates the nuclear processing of pri-miRNA [68].
m6A-mediated regulation of human cancer progression
Errors in the m6A pattern are associated with most types of cancers. Increasing studies show that m6A levels are related to cancer progression. As an example, m6A regulates the cancer-related RNA fate and functions including alternative splicing of pre-mRNA and the processing of miRNA and lncRNA [4, 69,70,71]. Relations between m6A and cancer progression in line with different cancer types are reviewed as follows.
Acute myeloid leukemia
Acute myeloid leukemia (AML), characterized by recurring chromosomal aberrations, gene mutations, and epigenetic modifications, is one of the most common cancers of the hematopoietic system [72, 73]. It has been proved that m6A modifications substantially contribute to the phenotype of leukemia cells [39, 74]. A strong association of mutations and/or copy number variations of m6A regulatory genes with the presence of TP53 mutations was concluded in AML patients by Cancer Genome Atlas Research Network datasets analysis [75]. Genetic alterations in m6A modifiers including METTL3, METTL14, YTHDF1, YTHDF2, FTO, and ALKBH5 together with TP53 and/or its regulator/downstream targets confer a worse survival with prognostic roles [75]. m6A demethylase FTO showed a significantly high expression in AMLs with t(11q23)/MLL-rearrangements, t(15;17)/PML-RARA, FLT3-ITD and/or NPM1 mutations and exerts its oncogenic role by regulating reducing the m6A levels of hematopoiesis regulators ASB2 and RARA transcripts [74]. Then METTL14, a m6A methyltransferase, was also found highly expressed in normal hematopoietic stem/progenitor cells (HSPCs) and t(11q23), t(15;17), or t(8;21) translocation subtype of AML and is decreased during myeloid differentiation. METTL14 suppresses HSPCs differentiation and promotes leukemogenesis by regulating the stability and translation of its mRNA targets via m6A modification [39]. It is very interesting that the elevated expressions of both writer and eraser may balance the perturbation of RNA m6A methylation contributing to AML progress. METTL14 appears to account for the stability of its mRNA target MYC through regulating m6A abundance mainly on the 3′-terminal exon [39], while FTO affects mRNA stability by inhibiting YTHDF2-mediated RNA decay via decreasing m6A abundance on the mRNA 5′-terminal and internal exons [76]. These data indicate that aberrant m6A modifiers on different transcript regions lead to distinct effects, probably due to the recognition of different readers. The underlying mechanisms need to be elucidated in further studies.
Cervical cancer
Cervical cancer is one of the most prevalent gynecological malignancies worldwide with poor prognosis [77]. In recent years, researchers have uncovered the important roles of RNA methylation in cervical cancer development. Dot blot assay results showed that the m6A levels were significantly decreased in cervical cancer samples compared with the adjacent normal tissues. And the reduction of m6A levels was significantly correlated with tumor progression (including tumor stage, tumor size, differentiation, lymph invasion) and cancer recurrence [78]. The m6A levels of certain genes mRNA also relate to the therapy resistance of cervical cancer. For example, m6A demethylase FTO, which shows higher expression in cervical squamous cell carcinoma tissues than respective adjacent normal tissues, enhances the chemo-radiotherapy resistance of cervical cancer by regulating the m6A levels of β-catenin mRNA transcripts [79]. The critical functions of m6A modifiers and their substrates in the regulation of chemoradiotherapy resistance may bear potential clinical implications for cervical cancer treatment.
Glioblastoma
Glioblastoma is characterized with stem-like cells at the apex of cellular hierarchies, which are self-renewing, resistant to conventional therapy, and sustaining long-term tumor growth leading to tumor recurrence [80]. It has been reported that m6A demethylase ALKBH5 is highly expressed in glioblastoma stem-like cells and can predict poor patient prognosis for glioblastoma patients [81]. Integrated transcriptome and m6A-seq analyses revealed that the nascent transcripts of transcription factor FOXM1 are substrates of ALKBH5 and can be demethylated by ALKBH5, causing an enhanced FOXM1 expression and reinstating the inhibition of tumor growth [81, 82]. Another study indicated that RNA methyltransferase METTL3 and METTL14 significantly promote the growth and self-renewal of glioblastoma stem-like cells by regulating the mRNA expression of glioblastoma related genes ADAM19 through mRNA m6A enrichment [83]. These data suggest a promising therapeutic tool for glioblastoma by targeting the m6A mRNA methylation.
Breast cancer
Breast cancer is a leading cause of cancer-related death and has a high incidence for women worldwide [84]. Recent studies have reported that the demethylation of NANOG mRNA in breast cancer cells can be induced by the exposure of hypoxia [85]. NANOG, which is a pluripotency factor promoting the breast cancer stem cell specification, is highly expressed due to the demethylation of m6A in its mRNA mediated by ALKBH5 [85]. Except for NANOG, another pluripotency factor KLF4 is also stimulated. In hypoxia condition, KLF4 is m6A demethylated in its mRNA and promotes the specification of breast cancer stem cell [86]. m6A RNA methylation can be also affected by anticancer chemicals. Sulforaphane, an isothiocyanate, is a dietary phytochemical and can promote genetic instability by diminishing m6A RNA methylation in breast cancer cells [87].
Liver cancer
Diverse and reversible m6A modification on RNAs emerges as a new layer of regulation in liver cancer. m6A modifiers YTHDF1 and METTL3 were reported to be significantly up-regulated in liver cancer and the overexpression of YTHDF1 and METTL3 are associated with poor prognosis of patients [88, 89]. By using transcriptome sequencing RNA m6A sequencing and m6A methylated RNA immuno-precipitation quantitative reverse-transcription polymerase chain reaction, METTL3 was identified to promote liver cancer progression through YTHDF2-dependent posttranscriptional silencing of cytokine signaling 2 by m6A modification [88].
Other kinds of cancers
Except for those mentioned above, m6A was also identified to regulate the progression of a variety of cancers, including nasopharyngeal carcinoma [90], prostate cancer [91], colorectal cancer [92], pancreatic cancer [93], lung cancer [94], gastric cancer [95], and renal cell carcinoma [96]. m6A in circulating tumor cells from lung cancer patients were discovered with a significant increase compared with whole blood cells, demonstrating that methylated RNA in body fluids may serve as biomarkers for lung cancer [94, 97]. In nasopharyngeal carcinoma, the alternative splicing of suppressor of variegation 3–9 homolog 1, a histone methyltransferase, is promoted by a natural compound baicalin hydrate via the enhancement of m6A RNA methylation [90]. m6A writer METTL3 acts as a tumor suppressor in renal cell carcinoma, promoting cell proliferation, migration and invasion and inducing G0/G1 arrest [96], whereas METTL3 only shows no morphologic and proliferative effects in pancreatic cancer but affects the sensitivity to anticancer reagents such as gemcitabine, 5-fluorouracil, cisplatin and irradiation [93]. METTL3 imparts radioresistance in glioma stem-like cells through SOX2-dependent enhanced DNA repair and enhanced sensitivity to γ-irradiation was observed when METTL3 was silenced [98]. m6A readers YTHDF1 and YTHDF2 have ontogenetic roles in promoting cell proliferation and migration by regulating m6A levels in colorectal cancer [92] and prostate cancer [91].
Detection of m6A modification
To better understand the important roles of m6A modification, more and more methods are developed for profiling precise m6A sites and locations. The crucial mechanism of m6A modification are further identified.
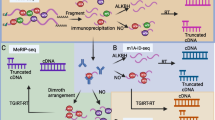
An affinity-based sequencing approach methylated RNA immunoprecipitation sequencing (MeRIP-Seq, also known as ‘m6A-seq’) has been developed and applied to survey the global mRNA m6A localizations [21, 99, 100]. In MeRIP-seq, mRNAs are first fragmented into approximately 100-nucleotide long fragments and then are immunoprecipitated with anti-m6A antibody followed by high throughput sequencing to form immunoprecipitation sequencing (IP) samples. In order to measure the background mRNA abundance for the IP experiments, input samples are additionally used, which are generated by sequencing the un-immunoprecipitated mRNA fragments. Finally, the reads enrichment in IP and input are assessed to predict m6A sites. Algorithms needed for MeRIP-seq-based m6A peak detection were subsequently developed. For example, the R/Bioconductor package exomePeak [101], the open source R package MeTCluster [102] and the graphical model-based MeTPeak [103]. By combining m6A-specific methylated RNA immunoprecipitation with high throughput deep sequencing, MeRIP-Seq has the potential to detect the transcriptome-wide distribution of m6A modifications. If the high-throughput assay is not available or necessary, a simple method based on high resolution melting analysis was introduced by Dontsova’s group, which allows a rapid and easy screening for the m6A presence at specific RNA position using total RNA sample and qPCR machine [104].
The mass spectrometry (MS) methodology, which has been used to identify posttranscriptional modifications of proteins, can also be developed to characterize the modified RNA. A promising alternative approach for RNA methylation is top-down MS, based on the RNA ionization in electrospray ionization, backbone cleavage in collisionally activated dissociation and electron detachment dissociation affected by specific nucleobase methylations [105]. However, samples for MS approaches need to be well prepared. By establishing a combined strategy for cell lysis, nucleic acids digestion, and nucleosides extraction in one-tube, Yuan’s lab successfully determined both DNA and RNA methylation in circulating tumor cells using liquid chromatography-electrospray ionization-tandem MS [97].
Different computational tools were also developed for the characteristics of m6A-associated variants, providing a resource for deeper analysis of m6A modifications and cancer disease at the epitranscriptomic layer of gene regulation. m6A-Driver, an algorithm developed by Huang’s group, are used for predicting m6A-driven genes and associated networks [106]. The m6A-driven network was built by integrating the Protein–Protein Interactions network and the predicted differential m6A methylation sites from MeRIP-Seq data using an algorithm called Random Walk with Restart [106]. These m6A-driven genes are likely to be actively modulated by m6A methylation under a specific condition, underlying the dynamic interactions between the methyltransferases and demethylase at the epitranscriptomic layer of gene regulation. Recently, Ren’s group reported a web server called ‘m6ASNP’ (http://m6asnp.renlab.org) [107] and a comprehensive database ‘m6Avar’ (http://m6avar.renlab. org) [108], which can be dedicated to identifying the m6A-associated genetic variants and their targeting m6A modification sites. These tools provided a useful resource for annotating genetic variants involved in m6A modification.
Conclusions
Given that m6A patterns in RNA transcripts play important roles in a variety of cancers, the m6A modification has great potential for clinical application by serving as diagnostic/prognostic targets or implicating an effective treatment strategy. There are also relevant studies focused on the inhibitors of RNA modifiers, giving clues for the rational design of potent and specific m6A inhibitors in medicine use. A natural product rhein was selected to competitively bind to the FTO active site by structure-based virtual screening and biochemical analyses and showed inhibitory effects on the activity on m(6)A demethylation [109]. Another FTO inhibitor, meclofenamic acid, which was approved by US Food and Drug Administration as a nonsteroidal anti-inflammatory drug, can compete with FTO for binding the m6A-containing nucleic acid [110]. Studies on RNA m6A patterns and their selective inhibitors will pave the way for research into epitranscriptomics in chemical biology and shed light on the discovery and development of m6A-specific probes and drugs.
Change history
14 July 2023
A Correction to this paper has been published: https://doi.org/10.1007/s11033-023-08633-9
References
Traube FR, Carell T (2017) The chemistries and consequences of DNA and RNA methylation and demethylation. RNA Biol 14:1099–1107. https://doi.org/10.1080/15476286.2017.1318241
Pan Y, Ma P, Liu Y, Li W, Shu Y (2018) Multiple functions of m(6)A RNA methylation in cancer. J Hematol Oncol 11:48. https://doi.org/10.1186/s13045-018-0590-8
Wang S, Sun C, Li J, Zhang E, Ma Z, Xu W, Li H, Qiu M, Xu Y, Xia W, Xu L, Yin R (2017) Roles of RNA methylation by means of N(6)-methyladenosine (m(6)A) in human cancers. Cancer Lett 408:112–120. https://doi.org/10.1016/j.canlet.2017.08.030
Alarcon CR, Lee H, Goodarzi H, Halberg N, Tavazoie SF (2015) N6-methyladenosine marks primary microRNAs for processing. Nature 519:482–485. https://doi.org/10.1038/nature14281
He Y, Hu H, Wang Y, Yuan H, Lu Z, Wu P, Liu D, Tian L, Yin J, Jiang K, Miao Y (2018) ALKBH5 inhibits pancreatic cancer motility by decreasing long non-coding RNA KCNK15-AS1 methylation. Cell Physiol Biochem 48:838–846. https://doi.org/10.1159/000491915
Blanco S, Dietmann S, Flores JV, Hussain S, Kutter C, Humphreys P, Lukk M, Lombard P, Treps L, Popis M, Kellner S, Holter SM, Garrett L, Wurst W, Becker L, Klopstock T, Fuchs H, Gailus-Durner V, Hrabe de Angelis M, Karadottir RT, Helm M, Ule J, Gleeson JG, Odom DT, Frye M (2014) Aberrant methylation of tRNAs links cellular stress to neuro-developmental disorders. EMBO J 33:2020–2039. https://doi.org/10.15252/embj.201489282
Gallo R (1971) Transfer RNA and transfer RNA methylation in growing and “resting” adult and embryonic tissues and in various oncogenic systems. Cancer Res 31:621–629
Gantt R (1971) In vitro transfer RNA methylation in paired neoplastic and nonneoplastic cell cultures. Cancer Res 31:609–612
Turkington R (1971) The regulation of transfer RNA methylation in normal and neoplastic mammary cells. Cancer Res 31:644–646
Erales J, Marchand V, Panthu B, Gillot S, Belin S, Ghayad SE, Garcia M, Laforets F, Marcel V, Baudin-Baillieu A, Bertin P, Coute Y, Adrait A, Meyer M, Therizols G, Yusupov M, Namy O, Ohlmann T, Motorin Y, Catez F, Diaz JJ (2017) Evidence for rRNA 2′-O-methylation plasticity: control of intrinsic translational capabilities of human ribosomes. Proc Natl Acad Sci USA 114:12934–12939. https://doi.org/10.1073/pnas.1707674114
Karijolich J, Yu Y-T (2014) Spliceosomal snRNA modifications and their function. RNA Biol 7:192–204. https://doi.org/10.4161/rna.7.2.11207
Adams J, Cory S (1975) Modified nucleosides and bizarre 5′-termini in mouse myeloma mRNA. Nature 255:28–33
Perry RP, Kelley DE, et al (1975) The methylated constituents of L cell messenger RNA: evidence for an unusual cluster at the 5′ terminus. Cell 4:387–394
Desrosiers R, Friderici K, Rottman F (1974) Identification of methylated nucleosides in messenger RNA from Novikoff hepatoma cells. Proc Natl Acad Sci USA 71:3971–3975
Aloni Y, Dhar R, Khoury G (1979) Methylation of nuclear simian virus 40 RNAs. J Virol 32:52–60
Beemon K, Keith J (1977) Localization of N6-methyladenosine in the Rous sarcoma virus genome. J Mol Biol 113:165–179
Adhikari S, Xiao W, Zhao YL, Yang YG (2016) m(6)A: signaling for mRNA splicing. RNA Biol 13:756–759. https://doi.org/10.1080/15476286.2016.1201628
Genenncher B, Durdevic Z, Hanna K, Zinkl D, Mobin MB, Senturk N, Da Silva B, Legrand C, Carre C, Lyko F, Schaefer M (2018) Mutations in cytosine-5 tRNA methyltransferases impact mobile element expression and genome stability at specific DNA repeats. Cell Rep 22:1861–1874. https://doi.org/10.1016/j.celrep.2018.01.061
Ke S, Alemu EA, Mertens C, Gantman EC, Fak JJ, Mele A, Haripal B, Zucker-Scharff I, Moore MJ, Park CY, Vagbo CB, Kussnierczyk A, Klungland A, Darnell JE Jr, Darnell RB (2015) A majority of m6A residues are in the last exons, allowing the potential for 3′ UTR regulation. Genes Dev 29:2037–2053. https://doi.org/10.1101/gad.269415.115
Ke S, Pandya-Jones A, Saito Y, Fak JJ, Vågbø CB, Geula S, Hanna JH, Black DL, Darnell JE, Darnell RB (2017) m6A mRNA modifications are deposited in nascent pre-mRNA and are not required for splicing but do specify cytoplasmic turnover. Genes Dev 31:990–1006. https://doi.org/10.1101/gad.301036.117
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, Cesarkas K, Jacob-Hirsch J, Amariglio N, Kupiec M, Sorek R, Rechavi G (2012) Topology of the human and mouse m6A RNA methylomes revealed by m6A-sEq. Nature 485:201–206. https://doi.org/10.1038/nature11112
Linder B, Grozhik AV, Olarerin-George AO, Meydan C, Mason CE, Jaffrey SR (2015) Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods 12:767–772. https://doi.org/10.1038/nmeth.3453
Fu Y, Dominissini D, Rechavi G, He C (2014) Gene expression regulation mediated through reversible m(6)A RNA methylation. Nat Rev Genet 15:293–306. https://doi.org/10.1038/nrg3724
Maity A, Das B (2016) N6-methyladenosine modification in mRNA: machinery, function and implications for health and diseases. FEBS J 283:1607–1630. https://doi.org/10.1111/febs.13614
Peer E, Rechavi G, Dominissini D (2017) Epitranscriptomics: regulation of mRNA metabolism through modifications. Curr Opin Chem Biol 41:93–98. https://doi.org/10.1016/j.cbpa.2017.10.008
Dai D, Wang H, Zhu L, Jin H, Wang X (2018) N6-methyladenosine links RNA metabolism to cancer progression. Cell Death Dis 9:124. https://doi.org/10.1038/s41419-017-0129-x
Jaffrey SR, Kharas MG (2017) Emerging links between m(6)A and misregulated mRNA methylation in cancer. Genome Med 9:2. https://doi.org/10.1186/s13073-016-0395-8
Wang S, Sun C, Li J, Zhang E, Ma Z, Xu W, Li H, Qiu M, Xu Y, Xia W, Xu L, Yin R (2017) Roles of RNA methylation by means of N6-methyladenosine (m6A) in human cancers. Cancer Lett 408:112–120. https://doi.org/10.1016/j.canlet.2017.08.030
Flores JV, Cordero-Espinoza L, Oeztuerk-Winder F, Andersson-Rolf A, Selmi T, Blanco S, Tailor J, Dietmann S, Frye M (2017) Cytosine-5 RNA methylation regulates neural stem cell differentiation and motility. Stem Cell Rep 8:112–124. https://doi.org/10.1016/j.stemcr.2016.11.014
Li H, Tong J, Zhu S, Batista P, Duffy E, Zhao J, Bailis W, Cao G, Kroehling L, Chen Y, Wang G, Broughton J, Chen Y, Kluger Y, Simon M, Chang H, Yin Z, Flavell R (2017) m6A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature 548:338–342
Li LJ, Fan YG, Leng RX, Pan HF, Ye DQ (2018) Potential link between m(6)A modification and systemic lupus erythematosus. Mol Immunol 93:55–63. https://doi.org/10.1016/j.molimm.2017.11.009
Blanco S, Frye M (2014) Role of RNA methyltransferases in tissue renewal and pathology. Curr Opin Cell Biol 31:1–7. https://doi.org/10.1016/j.ceb.2014.06.006
Tirumuru N, Zhao BS, Lu W, Lu Z, He C, Wu L (2016) N(6)-methyladenosine of HIV-1 RNA regulates viral infection and HIV-1 Gag protein expression. Elife. https://doi.org/10.7554/eLife.15528
Cao G, Li H, Yin Z, Flavell R (2016) Recent advances in dynamic m6A RNA modification. Open Biol 6:160003
Wang X, Huang J, Zou T, Yin P (2017) Human m(6)A writers: two subunits, 2 roles. RNA Biol 14:300–304. https://doi.org/10.1080/15476286.2017.1282025
Wang P, Doxtader KA, Nam Y (2016) Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Mol cell 63:306–317. https://doi.org/10.1016/j.molcel.2016.05.041
Wang X, Feng J, Xue Y, Guan Z, Zhang D, Liu Z, Gong Z, Wang Q, Huang J, Tang C, Zou T, Yin P (2016) Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature 534:575–578. https://doi.org/10.1038/nature18298
Barbieri I, Tzelepis K, Pandolfini L, Shi J, Millán-Zambrano G, Robson SC, Aspris D, Migliori V, Bannister AJ, Han N, De Braekeleer E, Ponstingl H, Hendrick A, Vakoc CR, Vassiliou GS, Kouzarides T (2017) Promoter-bound METTL3 maintains myeloid leukaemia by m6A-dependent translation control. Nature 552:126–131. https://doi.org/10.1038/nature24678
Weng H, Huang H, Wu H, Qin X, Zhao BS, Dong L, Shi H, Skibbe J, Shen C, Hu C, Sheng Y, Wang Y, Wunderlich M, Zhang B, Dore LC, Su R, Deng X, Ferchen K, Li C, Sun M, Lu Z, Jiang X, Marcucci G, Mulloy JC, Yang J, Qian Z, Wei M, He C, Chen J (2018) mettl14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m(6)A modification. Cell Stem Cell 22:191–205 e199. https://doi.org/10.1016/j.stem.2017.11.016
Ping XL, Sun BF, Wang L, Xiao W, Yang X, Wang WJ, Adhikari S, Shi Y, Lv Y, Chen YS, Zhao X, Li A, Yang Y, Dahal U, Lou XM, Liu X, Huang J, Yuan WP, Zhu XF, Cheng T, Zhao YL, Wang X, Rendtlew Danielsen JM, Liu F, Yang YG (2014) Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res 24:177–189. https://doi.org/10.1038/cr.2014.3
Meyer K, Jaffrey S (2017) Rethinking m6A readers, writers, and erasers. Annu Rev Cell Dev Biol 33:319–342
Schwartz S, Mumbach MR, Jovanovic M, Wang T, Maciag K, Bushkin GG, Mertins P, Ter-Ovanesyan D, Habib N, Cacchiarelli D, Sanjana NE, Freinkman E, Pacold ME, Satija R, Mikkelsen TS, Hacohen N, Zhang F, Carr SA, Lander ES, Regev A (2014) Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Rep 8:284–296. https://doi.org/10.1016/j.celrep.2014.05.048
Patil DP, Chen CK, Pickering BF, Chow A, Jackson C, Guttman M, Jaffrey SR (2016) m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537:369–373. https://doi.org/10.1038/nature19342
Pendleton KE, Chen B, Liu K, Hunter OV, Xie Y, Tu BP, Conrad NK (2017) The U6 snRNA m(6)A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell 169:824–835 e814. https://doi.org/10.1016/j.cell.2017.05.003
Hori H (2017) Transfer RNA methyltransferases with a SpoU-TrmD (SPOUT) fold and their modified nucleosides in tRNA. Biomolecules https://doi.org/10.3390/biom7010023
Long T, Li J, Li H, Zhou M, Zhou XL, Liu RJ, Wang ED (2016) Sequence-specific and shape-selective RNA recognition by the human RNA 5-methylcytosine methyltransferase NSun6. J Biol Chem 291:24293–24303. https://doi.org/10.1074/jbc.M116.742569
Jia G, Fu Y, Zhao X, Dai Q, Zheng G, Yang Y, Yi C, Lindahl T, Pan T, Yang YG, He C (2011) N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 7:885–887. https://doi.org/10.1038/nchembio.687
Mauer J, Luo X, Blanjoie A, Jiao X, Grozhik AV, Patil DP, Linder B, Pickering BF, Vasseur J-J, Chen Q, Gross SS, Elemento O, Debart F, Kiledjian M, Jaffrey SR (2016) Reversible methylation of m6Am in the 5′ cap controls mRNA stability. Nature 541:371–375. https://doi.org/10.1038/nature21022
Feng C, Liu Y, Wang G, Deng Z, Zhang Q, Wu W, Tong Y, Cheng C, Chen Z (2014) Crystal structures of the human RNA demethylase Alkbh5 reveal basis for substrate recognition. J Biol Chem 289:11571–11583. https://doi.org/10.1074/jbc.M113.546168
Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ, Vagbo CB, Shi Y, Wang WL, Song SH, Lu Z, Bosmans RP, Dai Q, Hao YJ, Yang X, Zhao WM, Tong WM, Wang XJ, Bogdan F, Furu K, Fu Y, Jia G, Zhao X, Liu J, Krokan HE, Klungland A, Yang YG, He C (2013) ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol Cell 49:18–29. https://doi.org/10.1016/j.molcel.2012.10.015
Loos RJ, Yeo GS (2014) The bigger picture of FTO: the first GWAS-identified obesity gene. Nat Rev Endocrinol 10:51–61. https://doi.org/10.1038/nrendo.2013.227
Zhao X, Yang Y, Sun BF, Shi Y, Yang X, Xiao W, Hao YJ, Ping XL, Chen YS, Wang WJ, Jin KX, Wang X, Huang CM, Fu Y, Ge XM, Song SH, Jeong HS, Yanagisawa H, Niu Y, Jia GF, Wu W, Tong WM, Okamoto A, He C, Rendtlew Danielsen JM, Wang XJ, Yang YG (2014) FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Res 24:1403–1419. https://doi.org/10.1038/cr.2014.151
Xu C, Liu K, Tempel W, Demetriades M, Aik W, Schofield CJ, Min J (2014) Structures of human ALKBH5 demethylase reveal a unique binding mode for specific single-stranded N6-methyladenosine RNA demethylation. J Biol Chem 289:17299–17311. https://doi.org/10.1074/jbc.M114.550350
Patil DP, Pickering BF, Jaffrey SR (2018) Reading m(6)A in the transcriptome: m(6)A-binding proteins. Trends Cell Biol 28:113–127. https://doi.org/10.1016/j.tcb.2017.10.001
Luo S, Tong L (2014) Molecular basis for the recognition of methylated adenines in RNA by the eukaryotic YTH domain. Proc Natl Acad Sci USA 111:13834–13839
Zhang Z, Theler D, Kaminska KH, Hiller M, de la Grange P, Pudimat R, Rafalska I, Heinrich B, Bujnicki JM, Allain FH, Stamm S (2010) The YTH domain is a novel RNA binding domain. J Biol Chem 285:14701–14710. https://doi.org/10.1074/jbc.M110.104711
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, Fu Y, Parisien M, Dai Q, Jia G, Ren B, Pan T, He C (2014) N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505:117–120. https://doi.org/10.1038/nature12730
Xu C, Liu K, Ahmed H, Loppnau P, Schapira M, Min J (2015) Structural basis for the discriminative recognition of N6-methyladenosine RNA by the human YT521-B homology domain family of proteins. J Biol Chem 290:24902–24913. https://doi.org/10.1074/jbc.M115.680389
Li F, Zhao D, Wu J, Shi Y (2014) Structure of the YTH domain of human YTHDF2 in complex with an m(6)A mononucleotide reveals an aromatic cage for m(6)A recognition. Cell Res 24:1490–1492. https://doi.org/10.1038/cr.2014.153
Zhu T, Roundtree IA, Wang P, Wang X, Wang L, Sun C, Tian Y, Li J, He C, Xu Y (2014) Crystal structure of the YTH domain of YTHDF2 reveals mechanism for recognition of N6-methyladenosine. Cell Res 24:1493–1496. https://doi.org/10.1038/cr.2014.152
Li A, Chen YS, Ping XL, Yang X, Xiao W, Yang Y, Sun HY, Zhu Q, Baidya P, Wang X, Bhattarai DP, Zhao YL, Sun BF, Yang YG (2017) Cytoplasmic m(6)A reader YTHDF3 promotes mRNA translation. Cell Res 27:444–447. https://doi.org/10.1038/cr.2017.10
Shi H, Wang X, Lu Z, Zhao BS, Ma H, Hsu PJ, Liu C, He C (2017) YTHDF3 facilitates translation and decay of N(6)-methyladenosine-modified RNA. Cell Res 27:315–328. https://doi.org/10.1038/cr.2017.15
Xu C, Wang X, Liu K, Roundtree IA, Tempel W, Li Y, Lu Z, He C, Min J (2014) Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain. Nat Chem Biol 10:927–929. https://doi.org/10.1038/nchembio.1654
Jain D, Puno M, Meydan C, Lailler N, Mason C, Lima C, Anderson K, Keeney S (2018) Ketu mutant mice uncover an essential meiotic function for the ancient RNA helicase YTHDC2. Elife 7:e30919
Wojtas MN, Pandey RR, Mendel M, Homolka D, Sachidanandam R, Pillai RS (2017) Regulation of m(6)A transcripts by the 3′ → 5′ RNA helicase YTHDC2 is essential for a successful meiotic program in the mammalian germline. Mol Cell 68:374–387 e312. https://doi.org/10.1016/j.molcel.2017.09.021
Meyer KD, Patil DP, Zhou J, Zinoviev A, Skabkin MA, Elemento O, Pestova TV, Qian SB, Jaffrey SR (2015) 5′ UTR m(6)A promotes cap-independent translation. Cell 163:999–1010. https://doi.org/10.1016/j.cell.2015.10.012
Genuth NR, Barna M (2018) Heterogeneity and specialized functions of translation machinery: from genes to organisms. Nat Rev Genet. https://doi.org/10.1038/s41576-018-0008-z
Alarcon CR, Goodarzi H, Lee H, Liu X, Tavazoie S, Tavazoie SF (2015) HNRNPA2B1 is a mediator of m(6)A-dependent nuclear RNA processing events. Cell 162:1299–1308. https://doi.org/10.1016/j.cell.2015.08.011
Li S, Mason CE (2014) The pivotal regulatory landscape of RNA modifications. Annu Rev Genom Hum Genet 15:127–150. https://doi.org/10.1146/annurev-genom-090413-025405
Liu N, Parisien M, Dai Q, Zheng G, He C, Pan T (2013) Probing N6-methyladenosine RNA modification status at single nucleotide resolution in mRNA and long noncoding RNA. RNA 19:1848–1856. https://doi.org/10.1261/rna.041178.113
Zhou KI, Parisien M, Dai Q, Liu N, Diatchenko L, Sachleben JR, Pan T (2016) N(6)-methyladenosine modification in a long noncoding RNA hairpin predisposes its conformation to protein binding. J Mol Biol 428:822–833. https://doi.org/10.1016/j.jmb.2015.08.021
Chen J, Odenike O, Rowley JD (2010) Leukaemogenesis: more than mutant genes. Nat Rev Cancer 10:23–36. https://doi.org/10.1038/nrc2765
Dohner H, Weisdorf DJ, Bloomfield CD (2015) Acute myeloid leukemia. N Engl J Med 373:1136–1152. https://doi.org/10.1056/NEJMra1406184
Li Z, Weng H, Su R, Weng X, Zuo Z, Li C, Huang H, Nachtergaele S, Dong L, Hu C, Qin X, Tang L, Wang Y, Hong G-M, Huang H, Wang X, Chen P, Gurbuxani S, Arnovitz S, Li Y, Li S, Strong J, Neilly MB, Larson RA, Jiang X, Zhang P, Jin J, He C, Chen J. (2017) FTO plays an oncogenic role in acute myeloid leukemia as a N6-methyladenosine RNA demethylase. Cancer Cell 31:127–141. https://doi.org/10.1016/j.ccell.2016.11.017
Kwok CT, Marshall AD, Rasko JE, Wong JJ (2017) Genetic alterations of m(6)A regulators predict poorer survival in acute myeloid leukemia. J Hematol Oncol 10:39. https://doi.org/10.1186/s13045-017-0410-6
Su R, Dong L, Li C, Nachtergaele S, Wunderlich M, Qing Y, Deng X, Wang Y, Weng X, Hu C, Yu M, Skibbe J, Dai Q, Zou D, Wu T, Yu K, Weng H, Huang H, Ferchen K, Qin X, Zhang B, Qi J, Sasaki AT, Plas DR, Bradner JE, Wei M, Marcucci G, Jiang X, Mulloy JC, Jin J, He C, Chen J (2018) R-2HG exhibits anti-tumor activity by targeting FTO/m 6 A/MYC/CEBPA signaling. Cell 172:90–105.e123. https://doi.org/10.1016/j.cell.2017.11.031
Siegel RL, Miller KD, Jemal A (2018) Cancer statistics, 2018. CA Cancer J Clin 68:7–30. https://doi.org/10.3322/caac.21442
Wang X, Li Z, Kong B, Song C, Cong J, Hou J, Wang S (2017) Reduced mA mRNA methylation is correlated with the progression of human cervical cancer. Oncotarget 8:98918–98930
Zhou S, Bai ZL, Xia D, Zhao ZJ, Zhao R, Wang YY, Zhe H (2018) FTO regulates the chemo-radiotherapy resistance of cervical squamous cell carcinoma (CSCC) by targeting beta-catenin through mRNA demethylation. Mol Carcinog 57:590–597. https://doi.org/10.1002/mc.22782
Lathia J, Mack S, Mulkearns-Hubert E, Valentim C, Rich J (2015) Cancer stem cells in glioblastoma. Genes Dev 29:1203–1217
Zhang S, Zhao BS, Zhou A, Lin K, Zheng S, Lu Z, Chen Y, Sulman EP, Xie K, Bögler O, Majumder S, He C, Huang S (2017) m6A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell 31:591–606.e596. https://doi.org/10.1016/j.ccell.2017.02.013
Dixit D, Xie Q, Rich JN, Zhao JC (2017) Messenger RNA methylation regulates glioblastoma tumorigenesis. Cancer Cell 31:474–475. https://doi.org/10.1016/j.ccell.2017.03.010
Cui Q, Shi H, Ye P, Li L, Qu Q, Sun G, Sun G, Lu Z, Huang Y, Yang C-G, Riggs AD, He C, Shi Y (2017) m6A RNA methylation regulates the self-renewal and tumorigenesis of glioblastoma stem cells. Cell Rep 18:2622–2634. https://doi.org/10.1016/j.celrep.2017.02.059
DeSantis CE, Ma J, Goding Sauer A, Newman LA, Jemal A (2017) Breast cancer statistics, 2017, racial disparity in mortality by state. CA Cancer J Clin 67:439–448. https://doi.org/10.3322/caac.21412
Zhang C, Samanta D, Lu H, Bullen JW, Zhang H, Chen I, He X, Semenza GL (2016) Hypoxia induces the breast cancer stem cell phenotype by HIF-dependent and ALKBH5-mediated m(6)A-demethylation of NANOG mRNA. Proc Natl Acad Sci USA 113:2047–2056. https://doi.org/10.1073/pnas.1602883113
Zhang C, Zhi WI, Lu H, Samanta D, Chen I, Gabrielson E, Semenza GL (2016) Hypoxia-inducible factors regulate pluripotency factor expression by ZNF217- and ALKBH5-mediated modulation of RNA methylation in breast cancer cells. Oncotarget 7:64527–64542. https://doi.org/10.18632/oncotarget.11743
Lewinska A, Adamczyk-Grochala J, Deregowska A, Wnuk M (2017) Sulforaphane-induced cell cycle arrest and senescence are accompanied by DNA hypomethylation and changes in microRNA profile in breast cancer cells. Theranostics 7:3461–3477. https://doi.org/10.7150/thno.20657
Chen M, Wei L, Law C, Tsang F, Shen J, Cheng C, Tsang L, Ho D, Chiu D, Lee J, Wong C, Ng I, Wong C (2017) RNA N6-methyladenosine methyltransferase-like 3 promotes liver cancer progression through YTHDF2-dependent posttranscriptional silencing of SOCS2. Hepatology 67:2254–2270
Zhao X, Chen Y, Mao Q, Jiang X, Jiang W, Chen J, Xu W, Zhong L, Sun X (2018) Overexpression of YTHDF1 is associated with poor prognosis in patients with hepatocellular carcinoma. Cancer Biomark 21:859–868. https://doi.org/10.3233/CBM-170791
Lai W, Jia J, Yan B, Jiang Y, Shi Y, Chen L, Mao C, Liu X, Tang H, Gao M, Cao Y, Liu S, Tao Y (2018) Baicalin hydrate inhibits cancer progression in nasopharyngeal carcinoma by affecting genome instability and splicing. Oncotarget 9:901–914
Li J, Meng S, Xu M, Wang S, He L, Xu X, Wang X, Xie L (2018) Downregulation of N-methyladenosine binding YTHDF2 protein mediated by miR-493-3p suppresses prostate cancer by elevating N-methyladenosine levels. Oncotarget 9:3752–3764
Nishizawa Y, Konno M, Asai A, Koseki J, Kawamoto K, Miyoshi N, Takahashi H, Nishida N, Haraguchi N, Sakai D, Kudo T, Hata T, Matsuda C, Mizushima T, Satoh T, Doki Y, Mori M, Ishii H (2018) Oncogene c-Myc promotes epitranscriptome mA reader YTHDF1 expression in colorectal cancer. Oncotarget 9:7476–7486
Taketo K, Konno M, Asai A, Koseki J, Toratani M, Satoh T, Doki Y, Mori M, Ishii H, Ogawa K (2018) The epitranscriptome m6A writer METTL3 promotes chemo- and radioresistance in pancreatic cancer cells. Int J Oncol 52:621–629. https://doi.org/10.3892/ijo.2017.4219
Lu Y, Li S, Zhu S, Gong Y, Shi J, Xu L (2017) Methylated DNA/RNA in body fluids as biomarkers for lung cancer. Biol Proc 19:2. https://doi.org/10.1186/s12575-017-0051-8
You Y, Liu L, Zhang M, Zhu Y, He L, Li D, Zhang J (2014) Genomic characterization of a Helicobacter pylori isolate from a patient with gastric cancer in China. Gut Pathog 6:5. https://doi.org/10.1186/1757-4749-6-5
Li X, Tang J, Huang W, Wang F, Li P, Qin C, Qin Z, Zou Q, Wei J, Hua L, Yang H, Wang Z (2017) The M6A methyltransferase METTL3: acting as a tumor suppressor in renal cell carcinoma. Oncotarget 8:96103–96116
Huang W, Qi C-B, Lv S-W, Xie M, Feng Y-Q, Huang W-H, Yuan B-F (2016) Determination of DNA and RNA methylation in circulating tumor cells by mass spectrometry. Anal Chem 88:1378–1384. https://doi.org/10.1021/acs.analchem.5b03962
Visvanathan A, Patil V, Arora A, Hegde AS, Arivazhagan A, Santosh V, Somasundaram K (2017) Essential role of METTL3-mediated m6A modification in glioma stem-like cells maintenance and radioresistance. Oncogene 37:522–533. https://doi.org/10.1038/onc.2017.351
Dominissini D, Moshitch-Moshkovitz S, Amariglio N, Rechavi G (2015) Transcriptome-wide mapping of N(6)-methyladenosine by m(6)A-SEq. Methods Enzymol 560:131–147. https://doi.org/10.1016/bs.mie.2015.03.001
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR (2012) Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149:1635–1646. https://doi.org/10.1016/j.cell.2012.05.003
Meng J, Lu Z, Liu H, Zhang L, Zhang S, Chen Y, Rao M, Huang Y (2014) A protocol for RNA methylation differential analysis with MeRIP-Seq data and exomePeak R/Bioconductor package. Methods 69:274–281
Cui X, Meng J, Zhang S, Rao MK, Chen Y, Huang Y (2016) A hierarchical model for clustering m(6)A methylation peaks in MeRIP-seq data. BMC Genom 17(Suppl 7):520. https://doi.org/10.1186/s12864-016-2913-x
Cui X, Meng J, Zhang S, Chen Y, Huang Y (2016) A novel algorithm for calling mRNA m6A peaks by modeling biological variances in MeRIP-seq data. Bioinformatics 32:i378–i385. https://doi.org/10.1093/bioinformatics/btw281
Golovina AY, Dzama MM, Petriukov KS, Zatsepin TS, Sergiev PV, Bogdanov AA, Dontsova OA (2014) Method for site-specific detection of m6A nucleoside presence in RNA based on high-resolution melting (HRM) analysis. Nucleic Acids Res 42:e27. https://doi.org/10.1093/nar/gkt1160
Glasner H, Riml C, Micura R, Breuker K (2017) Label-free, direct localization and relative quantitation of the RNA nucleobase methylations m6A, m5C, m3U, and m5U by top-down mass spectrometry. Nucleic Acids Res 45:8014–8025. https://doi.org/10.1093/nar/gkx470
Zhang S, Zhang S, Liu L, Meng J, Huang Y (2016) m6A-driver: identifying context-specific mRNA m6A methylation-driven gene interaction networks. PLoS Comput Biol 12:e1005287
Jiang S, Xie Y, He Z, Zhang Y, Zhao Y, Chen L, Zheng Y, Miao Y, Zuo Z, Ren J (2018) m6ASNP: a tool for annotating genetic variants by m6A function. GigaScience. https://doi.org/10.1093/gigascience/giy035
Zheng Y, Nie P, Peng D, He Z, Liu M, Xie Y, Miao Y, Zuo Z, Ren J (2018) m6AVar: a database of functional variants involved in m6A modification. Nucl Acids Res 46:D139–D145. https://doi.org/10.1093/nar/gkx895
Chen B, Ye F, Yu L, Jia G, Huang X, Zhang X, Peng S, Chen K, Wang M, Gong S, Zhang R, Yin J, Li H, Yang Y, Liu H, Zhang J, Zhang H, Zhang A, Jiang H, Luo C, Yang C-G (2012) Development of cell-active N6-methyladenosine RNA demethylase FTO inhibitor. J Am Chem Soc 134:17963–17971. https://doi.org/10.1021/ja3064149
Huang Y, Yan J, Li Q, Li J, Gong S, Zhou H, Gan J, Jiang H, Jia G-F, Luo C, Yang C-G (2015) Meclofenamic acid selectively inhibits FTO demethylation of m6A over ALKBH5. Nucleic Acids Res 43:373–384. https://doi.org/10.1093/nar/gku1276
Acknowledgements
We thank the PubMed database and its contributors for this valuable public data set. We also thank Dr Menglong Zhao from Dutch Institute for Fundamental Energy Research for helping to edit this manuscript.
Funding
This work was supported by the Key Project of Scientific Research Foundation for Colleges and Universities in Henan Province (Grant No. 16A320081) and National Natural Science Foundation of China (Grant No. 81802325).
Author information
Authors and Affiliations
Contributions
All authors participated in the preparation of the manuscript, read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, B., Li, Y., Song, R. et al. Functions of RNA N6-methyladenosine modification in cancer progression. Mol Biol Rep 46, 2567–2575 (2019). https://doi.org/10.1007/s11033-019-04655-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-04655-4