Abstract
Historical records suggest that Chiriguano tribe is the result of a genetic admixture event. The process involved the arrival of Guaraní tribesmen descending from Amazonian region of Brazil along with groups of Arawak origin that inhabited the foothill plains of Bolivia. Later they arrived in Argentina at the beginning of the twentieth century. Aiming to test the historical records, we analysed a set of 46 samples collected at San Ramon de la Nueva Orán, Province of Salta, Argentina. A wide set of uni- and biparentally transmitted genetic markers were analysed, including 23 autosomal STRs; 46 AIM-DIPs and 24 AIM-SNPs all located at diverse autosomal chromosome locations; 23 Y-STRs and the entire mtDNA D-Loop sequence. Ancestry informative markers allowed for the detection of a strong Native American component in the genomes (> 94%), while all mtDNA haplotypes showed Native American characteristic motives, and 93% of Y-haplotypes belonged to the Q1a3a Y-haplogroup. The analysis of mitochondrial haplotypes and Y chromosome, although they did not match other populations, revealed a relationship between the Chiriguano and other groups of Guaraní and Arawak origin inhabiting Brazil and Bolivia, confirming, at least in part, the historical records describing the origins of Chiriguano tribal settlements in northwestern Argentina.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chiriguano is the name given to a Guaraní-speaking ethnic group inhabiting southern Bolivia, western Paraguay and northwestern Argentina. The first historical references of the tribe date back to early sixteenth century [1,2,3].
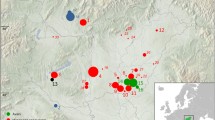
Amazonian peoples belonging to the Tupí-Guaraní linguistic branch advanced southward from the Amazon basin to the south and east of the South American subcontinent. This migratory movement was interpreted as a search for the “Land without Evil” (a philosophical concept of Guaraní mythology expressed as ivy imaraä in the Guaraní language). The Guaraní migrations towards the Andean foothills began in the pre-Hispanic period in the late fifteenth century because of pressure due to over-population, religious reasons or search for metals [3, 4]. They arrived at the Bolivian Andean foothills from the Paraguayan-Brazilian plains in the sixteenth century and inhabited hills and valleys between Grande and Pilcomayo rivers (Fig. 1).
a Map indicating the area occupied by the Chiriguano tribesmen (in red) until nineteen century, the main rivers (Río Grande, Pilcomayo and Bermejo) and location of San Ramon de la Nueva Oran city, Salta province (23°08′10″S 64°19′20″W); b relative position of the region within South America. (Color figure online)
According to the chronicles written by Díaz de Guzmán in 1612 [5], the Guaraní reached the region between the Grande (Guapay) and Pilcomayo rivers, currently part of Bolivia, having been previously recruited as a working force by the Portuguese Alejo García in Paraguay. García was a castaway from a caravel of Juan de Solís, who ran aground on the Brazilian coast in 1516, and arrived in Paraguay and there recruited 2000 Guaraní tribesmen with whom he crossed the plain towards the Andes. His objective was to push back the Incas, who controlled the region in order to acquire agricultural and mining resources after having conquered the Chané-Arawak inhabitants. After being defeated, Garcia and his Guaraní army returned to Paraguay, where he was killed by his own troops. The Guaraní then returned to the Andean foothills, where they settled and received the pejorative name of Chiriguano. The Chiriguano seized the lands conquered by the Incas, ruling over the Arawak.
Ava-Guaraní (Ava means “man” in the Guaraní language) is the name currently adopted for Guaraní-Arawak admixture descendants, who were previously called Chiriguano by other Native American tribes. This name (formerly Chiriguanaes: “those who have a wife chané”) is a pejorative nickname given by Quechua-speaking tribesmen to the members of Avá-Guaraní. The name was originally derived from two Quechua words: chiri (‘cold’) and wañuq (“those who die”). It was then adopted and modified by the Spaniard conquerors, resulting in the name Chiriguano.
During this period, the admixture and ethnogenesis of Chiriguanos occurred. This process involved a reduced number of Guaraní speaking aboriginals subduing another group of Indians belonging to the Chané tribe (Arawak speakers). The Chané people had settled earlier in the Chaco Boreal and North Central Chaco ecological region. According to the chronicles, the Guaraní were such fierce warriors that approximately 50 of them could beat 1000–3000 Chané [3, 6]. A particular feature exhibited by these warriors was the practice of cannibalism against their rivals [2]. Guaraní warriors enslaved Chané populations, forcibly taking their women and relegating the Chané to slavery.
As mestizos, resulting from the admixture among the Guaraní minority and Chané women of Arawak origin (named “Guana” in the northern Chaco and “Chané” in the Andean foothills), the Chiriguano built up their own identity distinct from whites (called “karai”) or indigenous ethnic groups (called “tapuy”) [7].
The admixed population acquired distinct cultural traits. The Chané had developed a sedentary way of life in large communal houses, working in basketry and pottery, onto which the Guaraní imposed their language, belief system, social organization and practices of hunting, fishing and war. Thus, their traditional way of livelihood was based on a combination of horticulture (corn, beans, and cassava) and hunting, gathering, and fishing imposed by Guaraní influence.
Upon arrival of the Spanish in the sixteenth century, the Chiriguano occupied the headwaters of Pilcomayo and Río Grande rivers and were mainly at war against the Aymara and Quechua ethnic groups. They were, militarily, almost unconquerable by the Spaniards, although the Catholic missions were quite successful with them, mainly in the valley of the Parapetí River.
The start of the Spanish-Chiriguano war took place in 1564. In 1574, the viceroy of Peru, Francisco de Toledo, personally led a punitive campaign against the Chiriguano, which was unsuccessful. King Felipe II of Spain ordered by royal decree on May 20, 1584, an unusual declaration of war, ordering the enslavement of any Chiriguanos that were captured. The Chiriguanos offered fierce resistance to the conquerors until the nineteenth century, in a war that lasted for 328 years. The last battle fought by the Chiriguano against the Bolivian army in 1892, which the Chiriguano lost, was called “the last race to death” by Thierry Saignes in his Historia del pueblo Chiriguano [7], when their leader Apiaguaiki Tumpa was assassinated.
During the twentieth century, the desperate economic situation and the Chaco War (1932–1935) forced the tribe members to move and find refuge in the Argentinean Chaco region that includes the Salta province, while some communities moved to western Paraguay.
Currently, the “Chiriguano” still maintain a position of prominence in small communities, with the Chané as the “minority shareholders” in Chiriguano-Chané populations.
The last census of indigenous people in Argentina (ECPI 2004–2005) [8] registered 2099 individuals who declared their ethnicity as Chiriguano, inhabiting the Salta province in northwestern Argentina. In San Ramón de la Nueva Orán, Salta province, there are 15 communities belonging to Guaraní, Tupí-Guaraní and Avá Guaraní ethnicities.
Taking into account the geographical region occupied by the Chiriguano until their displacement to northern Argentina and the west of Paraguay, we selected previously published results of several populations from Bolivia, Peru and Brazil who belonged to Tupí-Guaraní or Arawak linguistic groups for comparison with the Chiriguano results described herein.
Given the genesis and history of the Chiriguano, the aim of this study was to characterize the population by means of uni- and bi-parentally transmitted genetic markers in order to establish potential links between the historical and genetic information. In addition, autosomal ancestry informative markers (AIMs) including single nucleotide polymorphisms (SNPs), deletion-insertion polymorphisms (DIPs) and a set of short tandem repeats (STRs), gold standards for human identification applications, showed the possible admixture that could have occurred in the genesis of this tribal group, at present settled in northwestern Argentina.
Materials and methods
Individuals
Blood samples were collected during the 1980s as part of an immunological survey and have been stored since then at the Immunology Service at the Buenos Aires University Hospital “José de San Martín” in the city of Buenos Aires. Sampling occurred at San Ramón de la Nueva Orán, Orán Department (23°08′10″S–64°19′20″W), Salta province, Argentina. The sample set included 46 liquid blood samples (32 females and 14 males) anti-coagulated with 0.5% (v/v) ethilendiaminetetracetic acid disodium salt (EDTA-Na2). Donors declared their birthplace as either the Salta province in Argentina (89%) or Bolivia (11%). The Ethical Committee of the School of Pharmacy and Biochemistry (Res. 744085/2008) approved the project. These samples have a historical value, and although they were collected in the early 1980s before of the Helsinki Declaration [9], the individuals participated voluntarily, by oral consent.
DNA extraction and quantification
DNA extraction was carry out by conventional approaches [10] from blood samples. Quantification was performed by real time PCR [11], using a Rotor Gene 6000 (Corbet, Brisbane, Australia).
Genetic analysis
Autosomal STRs
Samples were amplified with a set of STRs routinely used for forensic identification purposes, including: D3S1358, D1S1656, D2S441, D10S1248, D13S317, PENTA E, D16S539, D18S51, D2S1338, CSF1PO, PENTA D, TH01, VWA, D21S11, D7S820, D5S818, TPOX, D8S1179, D12S391, D19S433, SE33, D22S1045 and FGA. This marker set constitutes the commercial kit PowerFusion 6C System (Promega Corp. Madison, USA) and was analysed according to the manufacturer’s protocol. Some samples were amplified with PowerPlex 16 kit (Promega Corp. Madison, USA).
Ancestry informative markers (AIM)
A panel of 24 autosomal SNPs (AIM-SNPs) and 46 AIM-DIPs were amplified as previously described [12, 13].
Y-chromosome STRs
A set of 23 Y-STRs included in the commercial kit Power Y23 (Promega Corp. Madison, USA) were typed according to the procedure manual. All haplotypes reported in this study have been deposited in the Y-Chromosome Haplotype Reference Database (YHRD, http//http://www.yhrd.org) under accession number YA004299, R56.
mtDNA control region sequencing
The complete D-Loop region was amplified using primers L15971 and H00639 [14]. Amplicons were purified with ExoSap (USB Corp, USA) and sequencing reactions were performed with the Big Dye Terminator System v3.1 (Life Technologies, Applied Biosystems, Foster City, USA) according to the supplier’s protocol. Sequencing products were purified from residual dye terminators by means of alcohol precipitation. All sequences were performed with both forward and reverse primers (with at least six primers for each sample); electropherograms were visualized and edited with Sequencher v5.3 software (Gene Codes Corporation, USA).
Capillary electrophoresis
Electrophoresis analysis was carried out in an ABI3500 (Applied Biosystems, Foster City, USA) sequencer either for fragment or sequence analysis.
Haplogroup specific SNPs detection
The Y-SNP M3-Q3 (Q1a3a1 Y haplogroup-“Hg”), as well as mtDNA-SNPs 8027 G/A and 12,007 G/A (HgA2); 3547 A/G (HgB2); 14,308 T/C (HgC) and 2092 C/T (HgD2) were analysed by real time PCR followed by high resolution melting analysis (HRM) as previously described [15].
Statistical analysis
Allele frequency, Hardy–Weinberg equilibrium (HW), gene diversity and heterozygosity were calculated with Arlequin v3.5 [16] along with haplotype frequencies, molecular diversity indices and genetic distances. A neighbour joining (NJ) genetic distance tree (autosomal STRs) and evolutionary analysis was constructed with Mega v7.0.21 [17]. Median-joining networks were obtained with Network v 5.0.0.3 program (http://www.fluxus-engineering.com).
For Y-chromosome analysis, the length of DYS389II, expressed in number of repeat units, was subtract from that of DYS389I [18]. A normalized Slatkin genetic distance matrix (Arlequin v3.5) was graphed as a multi-dimensional scaling plot (MDS) by using XLSTAT (Addinsoft Corp) software. The YHRD database was use to search worldwide haplotype frequency distribution.
Admixture analysis (autosomal data) was performed using Structure v 2.3.4 software [19]. Genotypes of parental population including Europe, Africa, and Native American were obtained from published data of CEPH-panel samples [20, 21]. AIM-SNPs, AIM-DIP and autosomal STRs were analysed for genetic structure by means of structure [19]. In Structure, jobs were set for five iterations rounds. Parental populations (k) were set from two to five. Monte Carlo-Markov chain simulation, including a burning step of 10,000 followed by 20,000 iterations for data gathering were performed for each round. Start at pop info was set for parental populations. An admixture model and independent allele frequencies were used. The most likely value for the number of parental populations (k = 3) was determined using the program STRUCTURE HARVESTER, which enables the Evanno method to be implemented [22]. Data analysis was further refined using CLUMPP [23], and a bar plot was created using DISRUPT [24].
Admixture analysis involving Y-haplotype data was carried out by means of ADMIX 2.0 [25, 26]. Two parental populations were selected: Guaraní from Brazil [27] and Arawak from Bolivia [28]. Only samples belonging to the Q1a3a1 haplogroup were considered for the analysis. The mutation rate was taken from Zhivotovsky et al. [29]. Time of admixture event was 500 years ago.
Results
Analysis of kinship relationship among sample donors
This investigation used 46 samples (32 females and 14 males) belonging to unrelated individuals according to the information as recorded at the time of sampling. All samples were typed by means of 15–23 autosomal STRs; mtDNA D-Loop sequencing; 23 Y-STRs; 46 autosomal AIM-DIPs; 24 autosomal AIM-SNPs and 4–5 specific haplogroup’s SNPs.
Although declared as unrelated, the combined analysis of the different genetic markers (mtDNA, Y-STRs and autosomal STRs) showed some degree of biological relationships among donors. This finding led to a re-definition of the sample set to be considered for statistical analysis. For example, samples #37, #39 and #27 (female and two males, respectively), shared the same mtDNA haplotype (Hg A2); autosomal STRs profiles showed compatibility of the maternity of #37 with that of #39 and #27, but #39 and #27 had a different Y-haplotype. In consequence, in this case, only one sample was taken into account for mitochondrial analysis, two samples for Y-haplotype analysis, and only sample #37 for autosomal STRs analysis. Another pair of samples (#59 and #49, female and male, respectively) showed similar results and in this case only #49 was included in the analysis. These findings reduced the overall sample size for each type of marker.
Autosomal STRs
All 23 STRs loci investigated met HW expectations. The average heterozygosity was 0.742 (s.d. 0.133) and average gene diversity over loci was 0.744573 ± 0.373340. The Garza Williamson test indicated that there is no evidence of a recent bottleneck (values > 0.75). The lowest heterozygosity was observed in locus TH01 (0.577), and the highest value (0.884) was in loci D13S317, PentaE, D8S1179 and SE33. Although observed heterozygosity was similar, PentaE and SE33 showed the largest size range (16 different alleles vs 6) justified by their high intra-locus diversity (Online Resource 1).
A microvariant FGA/24.2 was observed in one profile. This microvariant is rare in the Argentinean population database (24.2 = 0.0003, http://www.ffyb.uba.ar/SHDG database) and was not observed in other aboriginal samples from Argentina previously investigated by our research group (i.e., Mapuche, Tehuelche, Mataco-Guaycurú speakers or Mbyá-Guaraní) [30, 31].
To evaluate genetic distance relationship between Chiriguano and other Native American groups of Argentina, an NJ tree was constructed based on a set of 15 autosomal STRs (included in the Power16 commercial kit, Promega Corp.) (Fig. 2). The analysis included Mataco-Guaycurú speakers (Wichi, Pilagá, Toba, and Mocoví settled in central and northern Argentina); Guaraní speakers (Mbyá-Guaraní inhabiting Misiones province, North Eastern Argentina) and Mapudungun speakers (Mapuche and Tehuelche, inhabiting Patagonia, Southern Argentina). Genetic distances between Chiriguano were statistically significant regarding all the groups; the lowest was observed with Toba (Rst: 0.00781, p = 0.00 ± 0.00) and highest with Wichi (Rst: 0.03847, p = 0.00 ± 0.00) (both Mataco-Guaycurú speaker groups). Although the Mbyá population from Misiones belongs to the Tupí-Guaraní linguistic branch, no relationship was observed with the Chiriguano (Rst: 0.03482, p = 0.00 ± 0.00). This fact could be explained by different evolutionary scenarios: Mbyá-Guaraní began its expansion in South America from Amazonian lands following a south-eastern direction and arrived to the present-day province of Misiones (north-eastern Argentina) approximately 1000 years ago [32]. The Guaraní ancestors of the Chiriguano began their displacement later, in the thirteenth century; this displacement followed a south-western direction, with originally Bolivia settled, and finally ended in the territory occupied at present in Salta province (Northwestern Argentina) in the early XX century.
Autosomal ancestry informative markers
To investigate the ancestry of Chiriguano tribe members, a set of 24 AIM-SNPs and 46 AIM-DIPs were characterized. Regarding AIM-DIP, all systems met HW expectations, although one locus (MID 1607, rs2307803) met HW equilibrium only after Bonferroni correction [33]. Two loci were monomorphic: MID 3072, rs34611875 (genotype 1–1, frequency = 1) and MID 593, rs1160852 (genotype: 2–2, frequency = 1). Both DIPs were reported with high frequency at the same alleles in Native American CEPH panel [13]. Observed heterozygosity ranged from 0.023 (MID 2929, rs33974167) to 0.558 (MID 406, rs25630) (mean 0.27998, s.d. 0.16609).
All AIM-SNPs met HW equilibrium. Six loci (rs136929, rs139168, rs144848, rs180808, rs72135 and rs384377) [20] were monomorphic in concordance with the Native American CEPH panel (at the same loci, same allele). Observed heterozygosity ranged from 0.069 (rs205276) to 0.536 (rs147878).
Ancestry estimation of the Chiriguano was carried out considering three parental populations: Native South Americans (NA-CEPH), Europeans (E-CEPH) and West Sub-Saharan Africans (A-CEPH). The estimation employed autosomal STRs, autosomal AIM-SNPs and autosomal AIM-DIPs.
Genetic structure was evaluated by means of Structure, whose results showed a high Native American (NA) component, NA = 94.83% (AIM-SNPs) and NA = 94.67% (AIM-DIPs); the European (E) component was 4.77 and 4.73%, respectively; meanwhile the African (A) component was 0.6 and 0.4%. The Structure analysis with 15 autosomal STRs, used for identification purposes, was in good agreement with the abovementioned results using AIMs, although lower sensitivity (NA = 90.31%; E = 8.45% and A = 1.23%) was observed. Figure 3 shows bar plots obtained with the three sets of markers.
Structure analysis for assessing ancestry estimation of Chiriguano (Chi) assuming three parental populations (k = 3). We used as reference samples those included in the panel of the Centre d’Etudes de Polymorphisme Humaine (CEPH). Parental samples included; African (A-CEPH in green), European (E-CEPH in purple) and Native American (and NA-CEPH in orange). a Based on 15 autosomal STRs; b based on AIM-SNP and c based on AIM-DIP. (Color figure online)
Mitochondrial DNA
Complete D-Loop sequence was obtained from 45 samples, with one sample failing to amplify, and 41 were used for statistical analyses (after removing maternally related samples). Of the 41 haplotypes, all of them belonged to Native American haplogroups; 31 different haplotypes were observed; 24 were unique and the rest were observed two or three times. Haplogroup distribution was as follows: 21.95% HgA2; 41.46% HgB; 26.83% HgC and 9.75% HgD. The mean number of pairwise differences was 15.322880 ± 6.984319, and nucleotide diversity (average over loci) was 0.014677 ± 0.007430 (insertions at 309 and 315 were not taken into account). Mutations were as follows: transition = 68, transvertions = 5 and indels = 6.
Sub-haplogroup prediction was estimated with the EMPOP Query tool (http://www.empop.org), and diagnosis of point mutations was checked in Phylotree mtDNA tree Build 17 (http://www.phylotree.org). Specific SNPs in coding region were analysed by PCR real time followed by HRM [15]. The results were in accordance with the predicted haplogroup (data not shown). Online Resource 2 summarizes haplotype results.
Table 1 summarizes the total number of polymorphic sites per haplogroup. In general, HVRI was more polymorphic than HVRII-III. HgC samples displayed the highest polymorphism at HVRI, with 18 mutated positions over 27 observed within the haplogroup (considering the entire control region). Heteroplasmy was observed in two haplotypes (position T16217Y–HVRI and T195Y–HVRII; B4b hg in both cases) and was confirmed by sequencing in both directions. Position 195 was described as a fast evolving site [34], and the heteroplasmy T16217Y was also observed in some other population studies [35]. Haplotypes were submitted to EMPOP (accession number EMP00693). These haplotypes were not observed in 26127 of the EMPOP entire database (http://www.empop.org, Release 11).
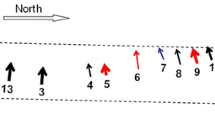
Figure 4 shows the median joining network for the four Native American haplogroups. No reticulations or complex structures were observed. Networks of haplogroups B and C showed highest diversity compared to those observed in A and D hgs. As can be observed, Chiriguano haplotypes show a high proportion of private mutations (positions highlighted in red).
Median joining network of Native American mitochondrial DNA haplogroups. Grey node: nodal haplogroup A2, B2, C1 and D1 respectively. Green nodes: median vectors. Black nodes: sub-haplogroups, sizes are proportional to frequency. Private mutations are indicated in red, insertion − 16282.1C (HgD) in blue. Insertions at 309 and 315, as well as heteroplasmic positions (HgB), were not included. Transversions were weighted three times as high as transitions. (Color figure online)
Y-STR and Y-SNPs
Online Resource 3 shows haplotypes from male samples. The haplotypes were validated by YHRD validation system and submitted to be incorporated in the database (accession number YA004299). All haplotypes were unique (gene diversity: 1.0000 ± 0.0270) and 13/14 showed M3/Q3-T variant, characteristic of founder Native American haplogroup Q1a3a. Only one haplotype (CHY-1, see Online Resource 3) showed the ancestral condition of M3/Q3-C and belongs to haplogroup R1b. Two haplotypes (CHY-7 and CHY-14) differed in one STR, DYS570. Although this STR was described as a rapid mutation Y-STR (mutation rate 1.33e−02 [36]), autosomal profiling allows for the rejection of a direct biological relation between donors (i.e., father /son) and mitochondrial DNA rejected a maternal relationship (i.e. half-brothers). Using Familias software v3.2 [37] the likelihood ratio (LR) obtained for indirect relations (such as half-brother, grandfather–grandson or uncle–nephew; kinship coefficient k2 = 0; 2k1 = 0.5; k0 = 0.5) was LR = 5.15. Employing a simulation tool for this type of degree relation, the hypothesis should be discarded with a probability of 97% (Familias v3.2, simulation with 1000 replicates, seed 12,345).
Other pair of haplotypes (CHY-3–CHY-12; and CHY-9–CHY-11) differed in the DYS458 marker by one repeat (mutation rate 6.36e−03, http://www.yhrd.org). Autosomal profiling allowed for the rejection of a first-degree relationship and mitochondrial DNA haplotypes a common maternal lineage.
None of these haplotypes was found in the YHRD database (release 55, including 39,414 haplotypes) in whole or meta-population databases.
We compiled a large list of published Y-haplotypes (N = 760, 16 populations) in order to compare our data. Most of the populations selected belong to the same linguistic group, Equatorial-Tucanoan [38], including Arawak from Bolivia, Brazil, Peru and Venezuela, and Tupí-Guaraní and Tupí from Brazil. Tupí-Guaraní is a branch of the Tupí language, spoken by a high proportion of indigenous tribes inhabiting the Amazonian basin. In addition, geographically related groups from Peru and Bolivia were included in the analysis (Table 2) [27, 28, 39,40,41,42], Slatkin’s normalized genetic distance matrix was computed by means of Arlequin V 3.5.2. For this analysis, haplotypes including 16 Y-STRs (DYS19, DYS389I, DYS389II; DYS390; DYS391, DYS392, DYS393, DYS385, DYS437, DYS438, DYS439; DYS448; DYS456; DYS458; YGATAH4 and DYS635) were considered in order to include several populations previously investigated. All haplotypes belonged to the Q1a3a Native American haplogroup.
Figure 5 depicts a multidimensional-scaling plot of genetic distances between groups. Except for one group (Pará Brazil, PaBr), Brazilian populations (AmBr, TuBr, MaBr and MgBr, belonging to Tupí, Tupí Guaraní and Arawak linguistic groups, see Table 2) occupy the most distant places among themselves and among the rest of the groups investigated. Since these samples correspond to very isolate Native American groups inhabiting Brazil, the result obtained is not surprising. In this figure, Chiriguano shared a central position together with the rest of populations included in the analysis.
MDS plot analysis of Slatkin linearized genetic distance matrix, based on Y-STR haplotypes of 17 populations (see Table 2 for population codes). Analysis included 16 Y-STRs haplotypes from Native American patrilineages (Hg Q1a3a). MDS was obtained with XLSTAT software. Kruskal stress: 0.099
Detailed relationships between populations can be observed if the genetic distant Brazilian groups are excluded from the analysis. Figure 6 shows an MDS representation of Y-STRs haplotypes for the closest related populations (geographical or linguistic), including Arawak from Venezuela, Peru, Bolivia and Brazil; Tupí-Guaraní from Pará, Brazil; Andeans populations from Perú; and Aymara and Quechua groups from Perú and Bolivia.
MDS plot of genetic distances between 13 populations (excluding MaBR; AmBR, TuBr and MgBr, see Table 2 for population codes). Analysis included 16 Y-STRs haplotypes from Native American patrilineages (Q1a3a hg). Kruskal stress 0.132
Chiriguano’s smallest genetic distance was observed with PaBr (Tupí-Guaraní, Brazil) and a native group from Beni, Bolivia (MoBo, Arawak) and could be explained taking into account the origin and evolution of Chiriguano (see introduction). On the other hand, it is closely related to other groups from Bolivia (ScBo) and Perú (CaPe; HuPe, UrPe and YaPe) (genetic distance matrix and significant matrix are shown in Online Resource 4). The group coded as MoBo corresponds to samples analysed by Tirado et al. [28]. These authors analysed 87 samples from indigenous settlements, most of which belong to the Mojeño ethnic group, which are an Arawak speaking people. Some of these populations were referred to as “allies” of the Chiriguano [43]. Among the haplotypes included in the comparison, only one Chiriguano haplotype (conformed by 16 Y-STRs) matched with an individual classified as Trinitario, Mojeño dialect, included by Tirado et al. [28] in his database. In addition, the genetic distance between Chiriguano and Cajamarca from Peru [40] was also not significant. According to the authors, there is evidence of a strong migration from Cajamarca to Amazonas (but not vice versa) during the second half of the last century. With the aim of demonstrating the relationship between Chiriguano and Tupí-Guaraní and Arawak people, an admixture analysis was carried out employing the groups of PaBr and MoBo as parental populations. The results showed that the contribution of parental population 1 (Tupí-Guaraní from Brazil) was 72.4% while the contribution of parental population 2 (Arawak from Bolivia) was 27.5%.
Figure 7 is a network representation of Y-haplotypes considering the closest related populations. This figure shows a complex network of haplotypes relationship where Chiriguano are closely interrelated with all the groups, occupying a central position in the reticulation.
Discussion
In this study, we describe the genetic attributes of a group of biologically unrelated individuals self-declared as Chiriguano of the Avá Guaraní linguistic group. Ava Guaraní represent 2.9% of the overall aboriginal population of Argentina [8], 2.1% of the aboriginal population in Bolivia [44], and 15.7% of the aboriginal population in Paraguay [45].
This small but representative sample allowed us to detect high genetic variation, in both uni- and bi-parentally transmitted genetic markers. Matrilineage and patrilineage analyses demonstrated that these people carried specific Native American inheritance. AIM-SNPs and AIM-DIP analyses showed a high native genetic component in their genomes. Structure analysis carried out with a panel of autosomal STRs employed in human identification revealed that these data was well correlated with the AIM markers.
Mitochondrial DNA haplotypes were not found in EMPOP worldwide database, with only infrequent haplotypes detected. Only some of these haplotypes could be associated with other South American aboriginal people. Some particular haplotypes observed in Chiriguano were:
A2ah: the haplotype T16097C, T16098G, C16111T, C16167T, C16223T, C16290T, G16319A, T16324C, T16362C, C64T, A73G, T146C, A153G, A234G, A235G; A263G, 309.1C, 315.1C, 523DEL, 524DEL (diagnosis mutations in bold) was found in two samples. Posterior analysis of autosomal STRs analysis allowed us to suggest some degree of relationship between both donors, and consequently, only one sample was used in the statistical analyses. This haplotype belongs to A2ah sub-Hg, displaying the diagnosis mutations T16097C and A16098G and three private mutations (C16167T, T16324C and A234G).
The A2ah sub-Hg was first described by Taboada-Echalar et al. [46] in two samples from Bolivia (Beni and Santa Cruz, respectively). The authors postulate that this motif could be restricted to Bolivia, more specifically to the “llanos” geographical area. Recently, it was described in an individual declared as “Chanka” on the pacific coast of Perú. Chanka were closely related to their neighbours who spoke Amazonian languages, with inter-population migrations between the Andes and Amazonia previously postulated [47]. Alves-Silva et al. [48] reported two samples (HVRI–HVRII sequences) in Minas Gerais in southeastern Brazil, showing the diagnosis of mutations. Finally, it was also found in a sample belonging to the Toba tribe in Chaco, Argentina [49].
It is important to mention that all these samples exhibited private mutations and in consequence showed different haplotypes.
A2q (T16209C) was found in two Chiriguano samples. Both haplotypes are not identical but present the same private mutations 16301T and 238G and a lack of mutation T16362C (16362T!). This haplogroup was described by Behar et al. [50] (GenBank: JQ705973.1 and JQ705557.1); Achilli [51] (GenBank: EU431081.1); and Kumar in 2011 [52] (GenBank: HQ012132.1) (Phylotree mtDNA Tree build 17, http://www.phylotree.org) and was not found in other Argentinean ethnic groups. Alves-Silva et al. [48] described this haplogroup (based on HVRI–HVRII sequence) in two samples from southern and southeastern Brazil and Taboada Echalar [46] in eight samples from the llanos area of Bolivia (based only on HVRI sequence). The same private mutation in HVRI (16301T) was found in two out of eight samples of Santa Cruz de la Sierra, Bolivia. However, since only HVRI was analysed in these samples, the haplotype identity cannot be established.
B2b2 (G16145A): This sub-Hg was observed in one sample and was previously described by Taboada-Echalar (GenBank: KC503926.1) in Beni, Bolivia [46].
B2y: This sub-Hg was observed in one sample with a C16261T diagnosis mutation and three private mutations (A197G, G207A and T292C) that distinguishes it from those already reported (GenBank: JQ705259.1 and EU597534.1) [50, 53]. B2y sub-Hg was found in two samples from Bolivia [46], one sample in Santa Cruz de la Sierra (HVRI sequence) and another in Chuquisaca (HVRI–HVRII, sequences), although HVRII sequence shows dissimilarity with Chiriguano sequence.
D4h3a: This sub-Hg was observed in one Chiriguano haplotype, exhibiting a diagnosis mutation at A16241G and three private mutations: an insertion at − 16282.1T and the substitution T16311C and T58C. This sub-Hg is in Phylotree with accession number FJ168754.1 [54] and EU095531.1 [55].
D4j4a: was observed in one sample (mutation motif T16263C) and described previously by Sukernik et al. [56], in Even, Siberia (HM044856.1 and FJ858887.1).
Other particular haplotypes were inferred as belonging to A2w; D1e; C1c or D1a sub-Hgs (based on D-Loop sequences). However, further studies of diagnostic mutations located in the coding region should be done to confirm the haplogroup assignment and detect novel findings that will allow a re-classification of the present ones. Described mitochondrial sub-Hgs were not found either in other Native American groups investigated by our team [30, 31, 49] nor in an Argentinean population database [57] with the exception of D4h3a previously found in Mapuche and Tehuelche ethnic groups, Patagonia, Argentina [31] and A2ah in Toba, north Argentina [49].
Comparative analysis of Y-chromosome haplotypes from several related populations allowed us to find a close relationship between Chiriguano and other Guaraní or Arawak groups (see “Results”). Admixture analysis demonstrate a strong Y-chromosome parental contribution from Tupí Guaraní (Brazil) and in less proportion of Arawak from Bolivia. These finding reinforce the historical records about the ethnogenesis of this group, where groups of migrant Guaraní from the Amazon conquered the Arawak that inhabited Bolivia, enslaving them and appropriating their women.
Conclusions
Mitochondrial haplotypes, as well as Y-STRs haplotypes, demonstrated a genetic relationship between the Chiriguano and groups of inhabitants from Brazil, Bolivia and, in lesser degree, Peru. These findings correlate well with historical information about their origin and interrelations with other people during their displacement from the Amazon. Historical records reported that the Chiriguano were fierce warriors who subjugated other peoples in their migration from the Amazon to northern Argentina. The present study relates this group to other Native American groups from South America of Arawak and Tupí–Guaraní speakers. The anthropological knowledge about them could be correlated with the genomic information of the extant population inhabiting Argentina. Their evolutionary history has been short in relation to other groups. However, their genetic imprint is evident and correlates them with other groups of speakers of the same linguistic branches.
References
Pifarré F (1989) Los Guaraní-Chiriguanos 2. Historia de un pueblo. Ed. CIPCA, La Paz
Martinez Sarasola C (2005) Nuestros paisanos los indios. Emece Edts, Buenos Aires
Combès I (2005) Etno-historias del Isoso: chané y chiriguanos en el Chaco boliviano. Siglos XVI al XX. Fundación PIEB/IFEA Eds, La Paz
Renard-Casevitz F-M, Saignes T (1988) Al Este de los Andes: relaciones entre las sociedades amazónicas y andinas entre los siglos XV y XVII (I). IFEA, La Paz. http://www.ifea.org.pe
Diaz de Guzman R (2001) Historia Argentina del descubrimiento, población y conquista de las provincias del Río de la Plata/escrita por Ruy Díaz de Guzmán, en el año de 1612. Biblioteca Virtual Miguel de Cervantes, Alicante. http://www.cervantesvirtual.com
Oliveto G (2010) Chiriguanos: la construcción de un estereotipo en la política colonizadora del sur andino. Mem Am 18(1):43–69
Cornbès I (2007) Thierry Saignes, Historia del pueblo Chiriguano. Compilación, introducción y notas. Instituto Francés de Estudios Andinos, Plural editores, La Paz
ECPI (2004–2005) ECPI, Encuesta Complentaria de Pueblos Indigenas. Instituto de Estadísticas y Censos, Buenos Aires. https://www.indec.gob.ar/
World Medical Association (2013) World Medical Association Declaration of Helsinki: ethical principles for medical research involving human subjects. JAMA 310(20):2191–2194
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold spring harbor laboratory, New York
Ginart S, Caputo M, Alechine E, Corach D, Sala A (2016) Development of a quantitation approach for total human and male DNA based on real time PCR followed by high resolution melting analysis. Electrophoresis 37(21):2734–2741
Corach D, Lao O, Bobillo MC, van Der Gaag K, Zuniga S, Vermeulen M, van Duijn K, Goedbloed M, Vallone PM, Parson W, deKnijff P, Kayser M (2010) Inferring continental ancestry of Argentineans from autosomal, Y-chromosomal and mitochondrial DNA. Ann Hum Genet 74:65–76
Pereira R, Phillips C, Pinto N, Santos C, dos Santos SEB, Amorim A, Carracedo A, Gusmao L (2012) Straightforward inference of ancestry and admixture proportions through ancestry-informative insertion deletion multiplexing. PLoS ONE 7(1):e29684
Brandstätter A, Niederstatter H, Parson W (2004) Monitoring the inheritance of heteroplasmy by computer-assisted detection of mixed basecalls in the entire human mitochondrial DNA control region. Int J Legal Med 118:47–54
Zuccarelli G, Alechine E, Caputo M, Bobillo MC, Corach D, Sala A (2011) Rapid screening for Native American mitochondrial and Y-chromosome haplogroups detection in routine DNA analysis. Forensic Sci Int Genet 5:105–108
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Kayser M, Caglia A, Corach D, Fretwell N, Gehrig C, Graziosi G, Heidorn F, Herrmann S, Herzog B, Hidding M, Honda K, Jobling M, Krawczak M, Leim K, Meuser S, Meyer E, Oesterreich W, Pandya A, Parson W, Penacino G, Perez-Lezaun A, Piccinini A, Prinz M, Schmitt C, Roewer L et al (1997) Evaluation of Y-chromosomal STRs: a multicenter study. Int J Legal Med 110:125–133
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Lao O, van Duijn K, Kersbergen P, de Knijff P, Kayser M (2006) Proportioning whole-genome single-nucleotidepolymorphism diversity for the identification of geographic population structure and genetic ancestry. Am J Hum Genet 78:680–690
Santos C, Phillips C, Oldoni F, Amigo J, Fondevila M, Pereira R, Carracedo Á, Lareu MV (2015) Completion of a worldwide reference panel of samples for an ancestry informative Indel assay panel. Forensic Sci Int Genet 17:75–80
Earl DA, von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Rosenberg N (2004) Distruct: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Bertorelle G, Excoffier L (1998) Inferring admixture proportions from molecular data. Mol Biol Evol 15:1298–1311
Dupanloup I, Bertorelle G (2001) Inferring admixture proportions from molecular data: extension to any number of parental populations. Mol Biol Evol 18:672–675
Roewer L, Nothnagel M, Gusmão L, Gomes V, González M, Corach D, Sala A, Alechine E, Palha T, Santos N, Ribeiro-Dos-Santos A, Geppert M, Willuweit S, Nagy M, Zweynert S, Baeta M, Núñez C, Martínez-Jarreta B, González-Andrade F, Fagundes de Carvalho E, da Silva DA, Builes JJ, Turbón D, Lopez Parra AM, Arroyo-Pardo E, Toscanini U, Borjas L, Barletta C, Ewart E, Santos S, Krawczak M (2013) Continent-wide decoupling of Y-chromosomal genetic variation from language and geography in native South Americans. PLoS Genet 9(4):e1003460
Tirado M, López-Parra AM, Baeza C, Bert F, Corella A, Pérez-Pérez A, Turbón D, Arroyo-Pardo E (2009) Y-chromosome haplotypes defined by 17 STRs included in AmpFlSTR Yfiler PCR Amplification Kit in a multi ethnical population from El Beni Department (North Bolivia). Leg Med 11:101–103
Zhivotovsky LA, Underhill PA, Cinnioğlu C, Kayser M, Morar B, Kivisild T, Scozzari R, Cruciani F, Destro-Bisol G, Spedini G, Chambers GK, Herrera RJ, Yong KK, Gresham D, Tournev I, Feldman MW, Kalaydjieva L (2004) The effective mutation rate at Y chromosome short tandem repeats, with application to human population-divergence time. Am J Hum Genet 74:50–61
Sala A, Argüelles CF, Marino ME, Bobillo C, Fenocchio A, Corach D (2010) Genetic analysis of six communities of Mbyá-Guaraní inhabiting North Eastern Argentina by means of nuclear and mitochondrial polymorphic markers. Hum Biol 82:433–456
Sala A, Corach D (2014) Analysis of admixture and genetic structure of two Native American groups of Southern Argentinean Patagonia. Mol Biol Rep 41:1533–1543
Tarragó MN (2000) Nueva Historia Argentina. Los pueblos originarios y la conquista. Edts Sudamericana, Buenos Aires
Bonferroni CE (1936) Teoria statistica delle classi e calcolo delle probabilita. Pubblicazioni del R Istituto Superiore di Scienze Economiche e. Commerciali di Firenze 8:3–62
Irwin JA, Saunier JL, Niederstätter H, Strouss KM, Sturk KA, Diegoli TM, Brandstätter A, Parson W, Parsons TJ (2009) Investigation of heteroplasmy in the human mitochondrial DNA control region: a synthesis of observations from more than 5000 global population samples. J Mol Evol 68(5):516–527
Li M, Schönberg A, Schaefer M, Schroeder R, Nasidze I, Stoneking M (2010) Detecting heteroplasmy from high-throughput sequencing of complete human mitochondrial DNA genomes. Am J Hum Genet 87(2):237–249
Ballantyne KN, Goedbloed M, Fang R, Schaap O, Lao O, Wollstein A, Choi Y, van Duijn K, Vermeulen M, Brauer S, Decorte R, Poetsch M, von Wurmb-Schwark N, de Knijff P, Labuda D, Vézina H, Knoblauch H, Lessig R, Roewer L, Ploski R, Dobosz T, Henke L, Henke J, Furtado MR, Kayser M (2010) Mutability of Y-chromosomal microsatellites: rates, characteristics, molecular bases, and forensic implications. Am J Hum Genet 10(3):341–353 87(
Kling D, Tillmar AO, Egeland T (2014) Familias 3—extensions and new functionality. Forensic Sci Int Genet 13:121–127
Greenberg JH (1987) Language in the Americas. Stanford University Press, Stanford
Cardenas JM, Heinz T, Pardo-Seco J, Alvarez-Iglesias V, Taboada-Echalar P, Sanchez-Diz P, Carracedo A, Salas A (2015) The multiethnic ancestry of Bolivians as revealed by the analysis of Y-chromosome markers. Forensic Sci Int Genet 14:210–218
Guevara EK, Palo JU, Guillén S, Sajantila A (2016) MtDNA and Y-chromosomal diversity in the Chachapoya, a population from the northeast Peruvian Andes-Amazon divide. Am J Hum Biol 28(6):857–867
Sandoval JR, Lacerda DR, Jota MS, Salazar-Granara A, Vieira PP, Acosta O, Cuellar C, Revollo S, Fujita R, Santos FR, Genographic Project Consortium (2013) The genetic history of indigenous populations of the Peruvian and Bolivian Altiplano: the legacy of the Uro. PLoS ONE 8(9):e73006. https://doi.org/10.1371/journal.pone.0073006
Barbieri C, Heggarty P, Yang Yao D, Ferri G, De Fanti S, Sarno S, Ciani G, Boattini A, Luiselli D, Pettener D (2014) Between Andes and Amazon: the genetic profile of the Arawak-speaking Yanesha. Am J Phys Anthropol 155(4):600–609. https://doi.org/10.1002/ajpa.22616
Fischermann B (2010) Pueblos Indígenas y Nacionales Originarios en Bolivia Tierras Bajas: Pueblo Mojeño. En: Atlas Territorios Indígenas y Originarios en Bolivia, MDRyT-Viceministerio de Tierra, La Paz, pp 55–56
INE (2012) Instituto Nacional de Estadística, Gobierno de Bolivia. Censo Nacional de Población y Vivienda 2012. https://www.ine.gob.bo
DGEEC (2012) Dirección General de Estadística, Encuestas y Censo, Gobierno de Paraguay. III Censo Nacional de Población y Viviendas para Pueblos Indígenas 2012. http://www.dgeec.gov.py
Taboada-Echalar P, Alvarez-Iglesias V, Heinz T, Vidal-Bralo L, Gomez-Carballa A, Catelli L, Pardo-Seco J, Pastoriza A, Carracedo A, Torres-Balanza A, Rocabado O, Vullo C, Salas A (2013) The genetic legacy of the pre-colonial period in contemporary bolivians. PLoS ONE 8(3):E58980
Sandoval JR, Lacerda DR, Acosta O, Jota MS, Robles-Ruiz P, Salazar-Granara A, Vieira PP, Paz-Y-Miño C, Fujita R, Santos FR, Genographic Project Consortium (2016) The genetic history of Peruvian Quechua-Lamistas and Chankas: uniparental DNA patterns among Autochthonous Amazonian and Andean populations. Ann Hum Genet 80(2):88–101
Alves-Silva J, da Silva Santos M, Guimaraes P, Ferreira A, Bandelt H-J, Pena S, Ferreira Prado V (2000) The ancestry of Brazilian mtDNA lineages. Am J Hum Genet 67:444–461
Sala A, Alechine E, Bobillo C, Merini L, Ayala C, Acosta Ferreira JC, Corach D (2009) Mitochondrial DNA control region sequence analysis of Mataco–Guaicuru speaking tribes from Argentina. Forensic Sci Int 2:331–333
Behar DM, van Oven M, Rosset S, Metspalu M, Loogvali EL, Silva NM, Kivisild T, Torroni A, Villems RA (2012) ‘Copernican’ reassessment of the human mitochondrial DNA tree from its root. Am J Hum Genet 90(4):675–684
Achilli A, Perego UA, Bravi CM, Coble MD, Kong QP, Woodward SR, Salas A, Torroni A, Bandelt HJ (2008) The phylogeny of the four pan-American MtDNA haplogroups: implications for evolutionary and disease studies. PLoS ONE 3(3):e1764
Kumar S, Bellis C, Zlojutro M, Melton PE, Blangero J, Curran JE (2011) Large scale mitochondrial sequencing in Mexican Americans suggests a reappraisal of Native American origins. BMC Evol Biol 7:11:293
Hartmann A, Thieme M, Nanduri LK, Stempfl T, Moehle C, Kivisild T, Oefner PJ (2009) Validation of microarray-based resequencing of 93 worldwide mitochondrial genomes. Hum Mutat 30(1):115–122
Perego UA, Achilli A, Angerhofer N, Accetturo M, Pala M, Olivieri A, Hooshiar Kashani B, Ritchie KH, Scozzari R, Kong QP, Myres NM, Salas A, Semino O, Bandelt HJ, Woodward SR, Torroni A (2009) Distinctive Paleo-Indian migration routes from Beringia marked by two rare mtDNA haplogroups. Curr Biol 19(1):1–8
Tamm E, Kivisild T, Reidla M et al (2007) Beringian standstill and spread of native american founders. PLoS ONE 2(9):e829. https://doi.org/10.1371/journal.pone.0000829
Sukernik RI, Volodko NV, Mazunin IO, Eltsov NP, Dryomov SV, Starikovskaya EB (2012) Mitochondrial genome diversity in the Tubalar, Even, and Ulchi: contribution to prehistory of native Siberians and their affinities to Native Americans. Am J Phys Anthropol 148(1):123–138
Bobillo MC, Zimmermann B, Sala A, Huber G, Röck A, Bandelt H-J, Corach D, Parson W (2010) Amerindian mitochondrial DNA haplogroups predominate in the population of Argentina: towards a first nationwide mitochondrial DNA sequence database. Int J Legal Med 124:263–268
Acknowledgements
This work was partly supported, by Grants UBACyT 20020130100783BA and CONICET PIP11220150100349 CO.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that the are no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sala, A., Caputo, M., Ginart, S. et al. Historical records under the genetic evidence: “Chiriguano” tribe genesis as a test case. Mol Biol Rep 45, 987–1000 (2018). https://doi.org/10.1007/s11033-018-4246-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-018-4246-0