Abstract
Myostatin (MSTN) is a strong inhibitor of skeletal muscle growth in human and other vertebrates. Its transcription is controlled by a proximal promoter/enhancer (Mstn P/E) containing a TATA box besides CREB, NF-Y, MEIS1 and FXR transcription factor binding sites (TFBSs), which are conserved throughout evolution. The aim of this work was to investigate the role of these TFBSs on Mstn P/E activity and evaluate the potential of their putative ligands as Mstn trans regulators. Mstn P/E mutant constructs were used to establish the role of conserved TFBSs using dual-luciferase assays. Expression analyses were performed by RT-PCR and in situ hybridization in C2C12 myoblasts and E10.5 mouse embryos, respectively. Our results revealed that CREB, NF-Y and MEIS1 sites are required to balance Mstn P/E activity, keeping Mstn transcription within basal levels during myoblast proliferation. Furthermore, our data showed that NF-Y site is essential, although not sufficient, to mediate Mstn P/E transcriptional activity. In turn, CREB and MEIS1 binding sites seem to depend on the presence of NF-Y site to induce Mstn P/E. FXR appears not to confer any effect on Mstn P/E activity, except in the absence of all other conserved TFBS. Accordingly, expression studies pointed to CREB, NF-Y and MEIS1 but not to FXR factors as possible regulators of Mstn transcription in the myogenic context. Altogether, our findings indicated that CREB, NF-Y and MEIS1 conserved sites are essential to control basal Mstn transcription during early myogenesis, possibly by interacting with these or other related factors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The embryo has the challenging task of unfolding different cellular phenotypes from a common repertoire of genes shared by all cells of its developing body. Success in this enterprise requires gene regulation, which enables different genes to be turned on/off as development proceeds. Transcriptional control of gene expression involves regulatory DNA sequences such as promoters, enhancers, silencers and insulators that cooperate to establish the spatial/temporal expression pattern of specific genes. RNA polymerase II, pre-initiation complex and tissue specific transcription factors bind at specific sites on these regulatory elements to drive gene expression [1]. A finely orchestrated interaction among DNA elements and regulatory proteins is therefore essential for the generation of different tissues during development.
Myostatin (MSTN) is a signaling molecule that plays an essential role in regulating development, homeostasis and repair of skeletal muscles [2]. This protein belongs to the transforming growth factor-β (TGF-β) superfamily and presents highly conserved structure and function. Mstn knockout in mice was shown to cause a dramatic increase in skeletal muscle mass due to both hyperplasia and hypertrophy [2]. Accordingly, natural mutations in this molecule have been associated to a hyper-muscular phenotype in many species [2,3,4,5]. In contrast, increased levels of Mstn expression or circulating protein were described to reduce muscle size and lead to muscle atrophy [6, 7] while inhibition of MSTN improves skeletal muscle regeneration in mouse models for muscle dystrophies [8]. In accordance to its role, Mstn is expressed in the myotome of developing somites, in fetal and adult muscles as well as in satellite cells [2, 9, 10]. However, variations in expression patterns are known to occur among different species of vertebrates (reviewed by Rodgers and Garikipati [11]). For instance, in mouse embryos Mstn expression seems to be limited to differentiating myoblasts of somites and limbs, while in chicken several other embryonic tissues express Mstn, such as testis and ovaries, suggesting yet unknown developmental functions [2, 12]. In mammals, several adult tissues express Mstn, such as heart, mammary gland, adipocytes, olfactory neurons and brain [11, 13,14,15,16,17]. Furthermore, Mstn expression changes under different nutritional states [18], aging [19], practice of physical exercises [20] and diseases associated to muscle wasting, such as the acquired immunodeficiency syndrome—AIDS [21], cirrhosis [22, 23], chronic obstructive pulmonary disease [24] and cachexia, a degenerative syndrome associated to several kinds of cancer [25, 26].
Its dynamic expression suggests that Mstn transcription is finely regulated, making the study of its promoter and other regulatory elements of particular interest. In a previous work, we have characterized a proximal promoter/enhancer (P/E) for Mstn using a phylogenetic approach and functional assays [27]. This proximal element is 260-bp-long and is conserved from human to fish. As about 10% of gene promoters, Mstn P/E has a canonical TATA box, which possibly recruits the TATA-binding protein (TBP) subunit of the TFIID complex, as a first step to form the RNA polymerase II preinitiation complex [28, 29].
Besides TATA-box, four other conserved transcription factor binding sites (TFBS) were found within Mstn P/E. These sites possibly interact with the factors CREB, NF-Y, MEIS1 and FXR, or related proteins [27]. The first TFBS corresponds to a cAMP response element binding (CREB) site, a cAMP responsive element (CRE) potentially bound by members of the CREB family of Basic Leucine Zipper transcription factors, which also includes activating transcription factor (ATF) and cAMP response element modulator (CREM) [30]. These factors bind to DNA as homo or heterodimers to activate transcription upon phosphorylation by protein kinase A (PKA) [31]. The second TFBS is for nuclear transcription factor Y (NF-Y), which is a widely expressed heteromeric protein composed by the subunits NF-YA, B and C. This complex binds to CCAAT boxes, which are ubiquitous elements found in several gene promoters, especially in those which also possess a CRE site [32, 33], as is the case of Mstn P/E. The third binding site is for the homeobox protein myeloid ecotropic viral integration site 1 (MEIS1), which belongs to the three aminoacid loop extension (TALE) subclass of transcription factors [34]. MEIS1 is known to heterodimerize with pre-B-cell leukemia homeobox 1 (PBX1), another TALE protein, to induce or repress transcription in several cellular contexts [35]. Finally, the fourth TFBS is likely bound by farnesoid X receptor (FXR), a member of the nuclear receptor superfamily of transcription factors. FXR is activated by ligands such as farnesol metabolites or bile acids, and is capable of mediating transcriptional activation or repression through a large variety of FXR response elements (FXREs), either as a monomer or as a FXR/RXR heterodimer [36].
Considering that the conserved TFBSs compose an essential part of vertebrate Mstn P/E architecture, in this work we aimed at establishing the transcriptional role of these sequences and evaluating the potential of CREB, NF-Y, MEIS1 and FXR factors as regulators of Mstn expression during myogenesis.
Experimental procedures
Cell culture
C2C12 mouse myoblast cell line was cultivated in growth medium composed by Dulbecco’s modified eagle’s medium (DMEM) supplemented with 10% fetal bovine serum containing the antibiotics penicillin (100 U/mL) and streptomycin (100 µg/mL). Cells were maintained in 6-wells plates at 37 °C in humidified atmosphere containing 5% CO2. Once achieving ~60% confluence, myoblasts were trypsinized and used to perform the RT-PCR assays.
RT-PCRs
Total RNA from proliferating C2C12 myoblasts was obtained using Trizol reagent (Invitrogen) and cDNA was synthesized with RevertAid H Minus First Strand cDNA Synthesis kit (Fermentas). Gene-specific primers, annealing temperatures and predicted PCR product sizes are detailed in Online Resource Table 1. PCRs were performed in 20 µL reactions, using 1 U of GOTaq® Polymerase (Promega), 300–450 ng of template cDNA, 0.3 mM dNTP mix, 0.5 mM of each primer, 1× Green GOTaq® reaction buffer with the addition of MgCl2 to 1.9 mM final concentration. Cycling conditions started with an initial denaturation at 94 °C for 1 min, followed by 35–40 cycles of denaturation at 95 °C for 1 min, annealing at 55–57 °C for 30 s, extension at 68 °C for 1 min. A final extension was performed at 72 °C for 10 min. Transcripts identity was confirmed by sequencing.
Site-directed mutagenesis
The tE4 construct, which contains the mouse Mstn P/E driving the expression of luciferase reporter gene (as previously described elsewhere [27]) was used as template to mutate specific TFBSs. Site-directed mutagenesis assays were performed using QuikChange II Site-Directed Mutagenesis kit (Agilent). Primers used to introduce single or multiple mutations in TFBS (Online Resource Table 2) were designed with QuikChange Primer design program. Details about the site-directed mutagenesis strategy are available in Fig. 1 of the Online Resource. All mutant constructs were sequenced to confirm the mutagenesis success.
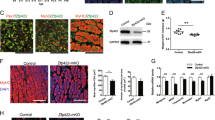
Conserved TFBSs affect Mstn P/E basal levels of expression during C2C12 myoblast proliferation. a Mstn and its putative regulators are co-expressed in proliferating C2C12 myoblasts; myogenic (MyoD and Myogenin) and endogenous (16S rRNA) controls were also detected. b In vitro activity of Mstn P/E-Luc constructs containing different mutations. Mutant constructs were transfected into C2C12 cells for measurement of luciferase reporter gene activity, which was normalized to the wild-type control. Wild-type Mstn P/E-Luc activity is indicated by the dashed line. Bars indicate means ± SEM of the independent experiments. (a) p < 0.05 for FXR+ versus ΔMstnP/E-Luc; (b) p < 0.05 for CREB+/NF-Y+ versus ΔMstnP/E-Luc; (c) p < 0.05 for NF-Y+/MEIS1+/FXR+ versus MEIS1+/FXR+; (#) CREB+/NF-Y+/MEIS1+ is the only construct with equivalent activity to the wild-type control
Transfections and dual luciferase assay
Mutant constructs were transfected into C2C12 mouse myoblasts in order to measure differences in luciferase activity. On the day before transfection, C2C12 cells cultured in proliferating conditions were trypsinized and plated in semi-confluency (~60%). Transfections were performed using 10 µg PEI (polyethyleneimine) for 10 µg of test DNA (wild-type and each of the eleven mutant constructs of Mstn P/E). In all assays, 5 µg of the control vector pRL-TK (Promega) DNA was co-transfected to normalize the transfection efficiency. Myoblasts were harvested 18 h after transfection for reporter gene luciferase assay. Transfections were always performed in duplicate and each experiment was repeated four times. The Dual-Luciferase Reporter Assay System kit (Promega) was used to measure Firefly and Renilla luciferase enzymatic activities.
Statistical analysis
After normalization with Renilla luciferase, the activity of Firefly luciferase of each tested construct was calculated as fold change in comparison to the wild-type Mstn P/E. Data were presented as mean ± standard error of the mean (SEM). One-way ANOVA followed by Tukey’s multiple comparisons test was performed using GraphPad Prism version 6.01 for Windows, GraphPad Software.
Synthesis of antisense RNA probes
Templates to generate antisense RNA probes were prepared by RT-PCR. Primers were initially used to amplify gene specific products using E10.5 mouse embryos cDNA. A second PCR was performed to add a T7 RNA polymerase promoter tail in synthesized amplicons. All primers used to produce probe templates are listed in Online Resource Table 1. T7 RNA polymerase (Invitrogen) and Digoxigenin RNA labeling mix (Roche) were used to synthesize probes, which were purified with SigmaSpin™ Post-Reaction Purification Clean-up kit (Sigma).
In situ hybridizations
Animal care was carried out in accordance with the Ethical Principles for Animal Research established by the Brazilian College for Animal Experimentation (COBEA). The protocol was approved by the Committee for Ethics in Animal Research of the State University of Campinas (CEUA/UNICAMP), under the permit number 3366-1.
E10.5 mouse embryos were fixed in 4% paraformaldehyde/PBS and dehydrated in methanol. Specimens were permeabilized by treatment with 10 µg/mL proteinase K/PBS at 37 °C during 20 min, followed by proteinase K inactivation in 2 µg/mL glycine/PBS for 10 min and post-fixation. Hybridizations with gene specific digoxigenin labelled probes were conducted at 70 °C overnight. Post-hybridization washes, antibody incubation and signal revelation were conducted as described by Wilkinson [37]. To detail gene expression patterns, some embryos were embedded in 20% gelatin and sectioned in vibratome (50 μm).
Results
Creb, Nf-Y, Meis1 and Fxr are co-expressed with Mstn during C2C12 myoblast proliferation
We started this work by investigating whether Creb, Nf-Y, Meis1 and Fxr gene expression profiles are related to that of Mstn during in vitro myogenesis. For that, RT-PCR was carried out in proliferating C2C12 myoblasts, a well-established model to study myogenesis [38]. Besides Mstn and its putative regulators, expression of MyoD and Myogenin (myogenic controls) as well as 16S rRNA (endogenous control) was analyzed (Fig. 1a). Our data showed that all tested molecules were expressed in C2C12 myoblasts. Therefore, Mstn transcription occurs in proliferating C2C12 myoblasts concomitantly with Creb, Nf-Y, Meis1 and Fxr mRNAs, indicating that proteins encoded by these genes are available to interact with their respective binding sites in the Mstn P/E.
Conserved TFBSs for CREB, NF-Y and MEIS1 cooperate to drive Mstn P/E basal activity
Having established the expression profiles of Mstn potential regulators during C2C12 myoblast proliferation, we next worked to elucidate the role of CREB, NF-Y, MEIS1 and FXR sites. For that, we investigated the effects of single or multiple mutations in the conserved TFBSs of Mstn P/E by measuring luciferase activity after transfection of proliferating C2C12 myoblasts with different constructs (Fig. 1b). For each construct, the relative values of activity were calculated as fold change in comparison to the wild-type Mstn P/E. It is worth noting that TATA-box was preserved in all constructs in order to allow transcription.
Taken as a whole, the site-directed mutagenesis assays showed that the individual or combined disruption of any of the conserved TFBS can cause changes in the basal activity of Mstn P/E in proliferating C2C12 myoblasts. In the more drastic scenario, disruption of all four TFBS (ΔMstnP/E-Luc) caused a 50% reduction in reporter expression relative to wild-type P/E. This result was used as reference for comparisons with the effects in Mstn P/E caused by four combinations of triple mutations, in order to mimic the restoration of each one of the TFBS to ΔMstnP/E-Luc (Fig. 1b; Online Resource Fig. 2).
Expression patterns indicate that CREB, NF-Y and MEIS1 are good candidates to regulate Mstn expression during in vivo myogenesis. Transverse sections through somites of E10.5 mouse embryos at interlimb levels show prominent expression of a Mstn and f MyoD in the myotome (arrows). b Creb and c Nf-Y present similar diffuse expression patterns that comprise the developing somites. d Meis1 transcripts are strongly upregulated in the dermomyotome (arrowhead). e Fxr is not expressed in myogenic regions. Scale bar = 200 µm
The single recovery of CREB, NF-Y or MEIS1 sites did not change P/E activity in relation to ΔMstnP/E-Luc (Fig. 1b). In contrast, restoration of FXR site caused a significant increase in reporter activity (a, P < 0.05), which was 1.94-fold higher than that observed for the wild-type P/E (Fig. 1b). Therefore, in a background lacking other conserved sites, most TFBSs were unable to modulate transcription by themselves, except for FXR, which mediated a positive effect.
Subsequently, analysis of double mutations showed that when only MEIS1 and FXR sites were intact, P/E activity was maintained equivalent to that observed for ΔMstnP/E-Luc and for the construct containing only MEIS1 site intact (Fig. 1b; Online Resource Fig. 2). This result also revealed an antagonistic interaction between MEIS1 and FXR sites, since in the presence of MEIS1, FXR site failed to induce P/E activity. In contrast, the presence of CREB and NF-Y binding sites significantly raised reporter expression in comparison to ΔMstnP/E-Luc (b, P < 0.05), reaching levels 1.45-fold higher than wild-type MstnP/E-Luc (Fig. 1b). Since the effect of CREB and NF-Y sites in Mstn P/E transcription changed from repression, when alone, to upregulation, when acting together (Online Resource Fig. 2), these sites probably interact synergistically.
Finally, MstnP/E-Luc constructs containing three intact conserved TFBS (or, in other words, constructs with single TFBS disrupted) were evaluated. In this set of assays, we initially analyzed the restoration of CREB or NF-Y in the MEIS1+/FXR+ construct, and then, conversely, the restitution of MEIS1 or FXR in the CREB+/NF-Y+ construct (Fig. 1b). After rescuing NF-Y in the MEIS1+/FXR+ background, there was an induction instead of repression (c, P < 0.05), leading to transcription levels 1.53-fold higher than those of wild-type MstnP/E construct (Fig. 1b). In contrast, CREB restoration had no effect in comparison to the transcription levels of MEIS1+/FXR+ construct (Fig. 1b). Taken together, these results show that MEIS1 and CREB sites cannot change the downregulation status of Mstn P/E without the NF-Y site (Online Resource Fig. 2). Meanwhile, only the restitution of MEIS1 in the CREB+/NF-Y+ background was able to reestablish wild-type expression levels, indicating that the CREB, NF-Y and MEIS1 sites are sufficient to control Mstn P/E activity during myoblast proliferation.
In short, our data revealed that in the context of myoblast proliferation, NF-Y site is essential, although not sufficient, to mediate Mstn P/E activity. In turn, CREB and MEIS1 binding sites seem to depend on the presence of NF-Y site to induce Mstn P/E activity. FXR appears not to confer any effect on the transcriptional activity of Mstn P/E, except in the absence of all other conserved TFBS. Taken together, our results showed that cooperation among CREB, NF-Y and MEIS1 TFBSs is crucial to balance Mstn P/E basal activity during myoblast proliferation.
Expression patterns indicate that CREB, NF-Y and MEIS1 are good candidates to regulate Mstn expression during in vivo myogenesis
After identifying the cooperation among the conserved TFBS in the regulation of Mstn P/E in vitro, we decided to investigate whether Creb, Nf-Y, Meis1 and Fxr gene expression patterns could also be temporally and spatially related to Mstn during in vivo myogenesis. For that, we evaluated the expression of Mstn and that of its putative regulators in E10.5 mouse embryos (Fig. 2), a stage in which Mstn is known to be strongly expressed [2]. MyoD expression was also used as a marker for skeletal myogenesis. Our results showed that Mstn transcripts (Fig. 2a) are located in the myotome of developing somites, overlapping with sites of MyoD expression (Fig. 2f). Creb (Fig. 2b) and Nf-Y (Fig. 2c) mRNAs are diffuse and distributed in a dorsal to ventral gradient with expression levels declining from dermomyotome towards myotome. Meis1 expression (Fig. 2d) is highly upregulated in the central dermomyotome, avoiding the myotome, Mstn main domain of expression. In turn, Fxr transcripts (Fig. 2e) were not associated to myogenic sites. Since Creb, Nf-Y and Meis1 are expressed in myogenic compartments of developing somites, the proteins encoded by these genes are possibly being synthesized in vivo in proper position and time to regulate Mstn expression.
Discussion
In the current study we carried out expression analysis and functional assays to evaluate the potential of CREB, NF-Y, MEIS1 and FXR as regulators of Mstn transcription, since TFBSs for these proteins have been conserved in Mstn P/E throughout vertebrate evolution [27]. Considering that Mstn is expressed mainly in skeletal muscles and its progenitors during development, and given its crucial function in determining muscle fiber number and size [2, 9], we focused our studies in the myogenic context, both in vitro and in vivo.
Using C2C12 as an in vitro model, we identified basal levels of Mstn expression in proliferative myoblasts. Expression data in C2C12 myoblasts are conflicting, as some works report no expression [38, 39] while others describe basal levels of Mstn expression [40, 41]. This heterogeneity probably reflects variations in the sensitivity of the techniques used to evaluate Mstn expression. Even so, it is consensus that Mstn expression in vitro is reduced during proliferation in comparison to differentiation [38, 40]. Thus, the Mstn expression detected in our work correlates to this scenario where Mstn is strictly downregulated to allow C2C12 myoblasts to undergo several rounds of cell division before differentiation begins. Along with Mstn, we found Creb, Nf-Y, Meis1 and Fxr expression, indicating that Mstn P/E may be regulated by the proteins encoded by these genes.
In this context, we further evaluated the role of Mstn potential regulators performing a systematic study of Mstn P/E-Luc constructs containing different sets of mutations on conserved TFBSs. Our data revealed that disruption of these binding sites interferes with Mstn P/E activity. It is relevant to mention that the conserved TATA box was left intact in all Mstn P/E-Luc constructs. TATA boxes have been commonly associated with highly regulated and strong tissue-specific promoters [28, 42]. This characteristic, however, is not due to TATA box itself, but most likely conferred by adjacent TFBSs. In fact, without any of the conserved TFBSs neighboring the TATA box in Mstn P/E, expression levels decrease substantially.
As an essential part of Mstn P/E, the conserved TFBSs seem to act in concert to balance Mstn expression. NF-Y site was shown to be necessary, although not sufficient, to promote Mstn P/E activity. In fact, recent works revealed that NF-Y causes epigenetic modifications that change chromatin conformation to facilitate the binding of other transcription factors to DNA [43, 44]. Moreover, CCAAT boxes are typically flanked by at least one functionally important promoter element, and the interaction between them is essential for the function of the regulatory unit [32]. In the Mstn P/E, this essential DNA element seems to be the adjacent CREB binding site (CRE site), which increases Mstn P/E activity whenever in the presence of NF-Y site. Interestingly, a similar promoter organization was also described for Hexokinase II (HKII), a gene that encodes an important enzyme for glucose metabolism in skeletal muscle [45, 46]. Therefore, NF-Y and CREB sites compose an important regulatory module of Mstn P/E.
It is important to highlight that CREB functional status changes after phosphorylation by PKA, as in the transition from proliferation to differentiation of C2C12 myoblasts [47]. In this context, phosphorylated CREB (P-CREB) was shown to interact with p300 to recruit MYOD on Retinoblastoma promoter, an important gene involved in cell cycle arrest [47]. P-CREB was also shown to modulate Mstn activity in differentiating C2C12 cells treated with IGF-1 factor [48]. In contrast, in the absence of phosphorylation by PKA, CREB is capable of inhibiting promoters of several genes [49,50,51,52]. Taken together, these data indicate that cAMP/PKA signaling pathway may be relevant in the regulation of Mstn transcriptional activity.
Another factor related to cAMP signaling is MEIS1, which also presents a transcriptional activation domain responsive to PKA [53]. Our data revealed that MEIS1 site contributes to Mstn P/E function in proliferating C2C12 myoblasts and, as CREB, requires NF-Y to mediate its transcriptional effects. In fact, MEIS1 interacts with PBX1, another HOX–TALE transcription factor, to recruit the heterodimer MyoD/E12 to a noncanonical E-box located in Myogenin promoter, activating its transcription [35]. Since MyoD participates in Mstn regulation [54], it may be speculated that MEIS1 could mediate non canonical binding of this factor to Mstn P/E.
Differently from the other TFBSs, FXR presented a neutral role on Mstn P/E activity except when it is the only intact TFBS. Although Fxr transcripts were detected in proliferating C2C12 cells, they were not found in mouse embryos somites where myogenic precursor cells are located. Furthermore, FXR is not expressed in adult skeletal muscle, but it is detected in liver, intestine and adipose tissue [55, 56]. As there has been growing evidence of roles for Mstn in the above mentioned sites of Fxr expression [57], it is possible that FXR has a more active post-natal role on the regulation of Mstn in a tissue-specific manner, acting in other locations such as liver and adipose tissue, where Mstn is also expressed [58].
Except for FXR, Mstn regulators studied in vitro are also expressed in myogenic territories of developing somites in E10.5 mouse embryos. At this stage, most somites have two compartments committed to skeletal muscle formation: the medially/laterally located dermomyotome, which contains proliferating myoblasts and, beneath it, the myotome, bearing differentiating cells [59]. Our in situ hybridization assays unraveled a complementary nature of Mstn expression, restricted to the myotome, in comparison to Meis1 expression, strongly upregulated in the dermomyotome. This fact suggests a negative effect of MEIS1 on Mstn transcription in vivo. In turn, Creb and Nf-Y present identical patterns with diffuse expression encompassing both the myotome and dermomyotome. In this scenario, it can be speculated that the proteins encoded by these genes may interact in vivo to regulate Mstn activity in both compartments. In the future, it would be interesting to investigate the role of CREB, NF-Y and MEIS1 on Mstn regulation in vivo to elucidate how these factors participate in the Mstn transcription when integrated to its complete regulatory network.
In conclusion, in the current paper we showed that CREB, NF-Y and MEIS1 conserved TFBSs mediate Mstn P/E transcriptional activity in the myogenic context. Future studies are needed to confirm the physical interaction between the conserved sites of Mstn P/E with the transcription factors CREB, NF-Y and MEIS1. It would be interesting to test whether overexpression or knockdown of these transcription factors in vivo or in vitro modulate Mstn expression. Besides, as both CREB and MEIS1 are affected by cAMP signaling it would be important to evaluate the effects of this pathway on Mstn transcription. It is also important to consider that there may be other non-conserved TFBSs influencing Mstn P/E activity and also working to keep an appropriate balance of Mstn expression. A good candidate to be investigated in the future is MYOD, a myogenic factor that has already been related to Mstn regulation. Finally, it is important to remember that probably besides the promoter there is a complex network of cis-regulatory elements working to determine when, where and at what levels Mstn transcription occurs during development and at different postnatal physiological contexts.
References
Mora A, Sandve GK, Gabrielsen OS, Eskeland R (2015) In the loop: promoter-enhancer interactions and bioinformatics. Brief Bioinform. doi:10.1093/bib/bbv097
McPherron AC, Lawler AM, Lee SJ (1997) Regulation of skeletal muscle mass in mice by a new TGF-beta superfamily member. Nature 387:83–90. doi:10.1038/387083a0
Grobet L, Martin LJ, Poncelet D, Pirottin D, Brouwers B, Riquet J, Schoeberlein A, Dunner S, Menissier F, Massabanda J, Fries R, Hanset R, Georges M (1997) A deletion in the bovine myostatin gene causes the double-muscled phenotype in cattle. Nat Genet 17:71–74. doi:10.1038/ng0997-71
Hu W, Chen S, Zhang R, Lin Y (2013) Single nucleotide polymorphisms in the upstream regulatory region alter the expression of myostatin. In Vitro Cell Dev Biol Anim 49:417–423. doi:10.1007/s11626-013-9621-5
Kambadur R, Sharma M, Smith TP, Bass JJ (1997) Mutations in myostatin (GDF8) in double-muscled Belgian Blue and Piedmontese cattle. Genome Res 7:910–916. doi:10.1101/gr.7.9.910
Reisz-Porszasz S, Bhasin S, Artaza JN, Shen R, Sinha-Hikim I, Hogue A, Fielder TJ, Gonzalez-Cadavid NF (2003) Lower skeletal muscle mass in male transgenic mice with muscle-specific overexpression of myostatin. Am J Physiol. doi: 10.1590/1414-431X20143858
Zimmers TA, Davies MV, Koniaris LG, Haynes P, Esquela AF, Tomkinson KN, McPherron AC, Wolfman NM, Lee SJ (2002) Induction of cachexia in mice by systemically administered myostatin, vol 296. Science, New York, pp 1486–1488. doi:10.1126/science.1069525
Joulia-Ekaza D, Cabello G (2007) The myostatin gene: physiology and pharmacological relevance. Curr Opin Pharmacol 7:310–315. doi: 10.1016/j.coph.2006.11.011
Amthor H, Huang R, McKinnell I, Christ B, Kambadur R, Sharma M, PateL K (2002) The regulation and action of myostatin as a negative regulator of muscle development during avian embryogenesis. Dev Biol 251:241–257. doi:10.1006/dbio.2002.0812
Zhang Y, Li W, Zhu M, Li Y, Xu Z, Zuo B (2016) FHL3 differentially regulates the expression of MyHC isoforms through interactions with MyoD and pCREB. Cell Signal 28:60–73. doi:10.1016/j.cellsig.2015.10.008
Rodgers BD, Garikipati DK (2008) Clinical, agricultural, and evolutionary biology of myostatin: a comparative review. Endocr Rev 29:513–534. doi:10.1210/er.2008-0003
Kubota K, Sato F, Aramaki S, Soh T, Yamauchi N, Hattori M (2007) Ubiquitous expression of myostatin in chicken embryonic tissues: its high expression in testis and ovary. Comp Biochem Physiol A 148:550–555. doi:10.1016/j.cbpa.2007.07.004
George I, Bish LT, Kamalakkannan G, Petrilli CM, Oz MC, Naka Y, Sweeney HL, Maybaum S (2010) Myostatin activation in patients with advanced heart failure and after mechanical unloading. Eur J Heart Fail 12:444–453. doi:10.1093/eurjhf/hfq039
Iwasaki S, Miyake M, Watanabe H, Kitagawa E, Watanabe K, Ohwada S, Kitazawa H, Rose MT, Aso H (2013) Expression of myostatin in neural cells of the olfactory system. Mol Neurobiol 47:1–8. doi:10.1007/s12035-012-8342-1
Li F, Yang H, Duan Y, Yin Y (2011) Myostatin regulates preadipocyte differentiation and lipid metabolism of adipocyte via ERK1/2. Cell Biol Int 35:1141–1146. doi:10.1042/CBI20110112
Manickam R, Pena RN, Whitelaw CB (2008) Mammary gland differentiation inversely correlates with GDF-8 expression. Mol Reprod Dev 75:1783–1788. doi:10.1002/mrd.20918
Sharma M, Kambadur R, Matthews KG, Somers WG, Devlin GP, Conaglen JV, Fowke PJ, Bass JJ (1999) Myostatin, a transforming growth factor-beta superfamily member, is expressed in heart muscle and is upregulated in cardiomyocytes after infarct. J Cell Physiol 180:1–9
Ruan J, Zhang Y, Yuan J, Xin L, Xia J, Liu N, Mu Y, Chen Y, Yang S, Li K (2016) A long-term high-fat, high-sucrose diet in Bamaminipigs promotes lipid deposition and amyotrophy by up-regulating the myostatin pathway. Mol Cell Endocrinol 425:123–132. doi:10.1016/j.mce.2016.02.001
Siriett V, Platt L, Salerno MS, Ling N, Kambadur R, Sharma M (2006) Prolonged absence of myostatin reduces sarcopenia. J Cell Physiol 209:866–873. doi:10.1002/jcp.20778
Hjorth M, Pourteymour S, Görgens SW, Langleite TM, Lee S, Holen T, Gulseth HL, Birkeland KI, Jensen J, Drevon CA, Norheim F (2016) Myostatin in relation to physical activity and dysglycemia and its effect on energy metabolism in human skeletal muscle cells. Acta Physiol 217:45–60. doi:10.1111/apha.12631
Gonzalez-Cadavid NF, Taylor WE, Yarasheski K, Sinha-Hikim I, Ma K, Ezzat S, Shen R, Lalani R, Asa S, Mamita M, Nair G, Arver S, Bhasin S (1998) Organization of the human myostatin gene and expression in healthy men and HIV-infected men with muscle wasting. Proc Natl Acad Sci USA 95:14938–14943. doi:10.1073/pnas.95.25.14938
García PS, Cabbabe A, Kambadur R, Nicholas G, Csete M (2010) Brief-reports: elevated myostatin levels in patients with liver disease: a potential contributor to skeletal muscle wasting. Anesth Analg 111:707–709. doi:10.1213/ANE.0b013e3181eac1c9
Qiu J, Thapaliya S, Runkana A, Yang Y, Tsien C, Mohan ML, Narayanan A, Eghtesad B, Mozdziak PE, McDonald C, Stark GR, Welle S, Naga Prasad SV, Dasarathy S (2013) Hyperammonemia in cirrhosis induces transcriptional regulation of myostatin by an NF-κB-mediated mechanism. Proc Natl Acad Sci USA 110:18162–18167. doi:10.1073/pnas.1317049110
Ju C-R, Chen R-C (2012) Serum myostatin levels and skeletal muscle wasting in chronic obstructive pulmonary disease. Respir Med 106:102–108. doi:10.1016/j.rmed.2011.07.016
Argiles JM, Almendro V, Busquets S, Lopez-Soriano FJ (2004) The pharmacological treatment of cachexia. Curr Drug Targets 5:265–277. doi: 10.2174/1389450043490505
Jespersen J, Kjaer M, Schjerling P (2006) The possible role of myostatin in skeletal muscle atrophy and cachexia. Scand J Med Sci Sports 16:74–82. doi: 10.1111/j.1600-0838.2005.00498.x
Grade CV, Salerno MS, Schubert FR, Dietrich S, Alvares LE (2009) An evolutionarily conserved Myostatin proximal promoter/enhancer confers basal levels of transcription and spatial specificity in vivo. Dev Genes Evol 219:497–508. doi:10.1007/s00427-009-0312-x
Baumann M, Pontiller J, Ernst W (2010) Structure and basal transcription complex of RNA Polymerase II core promoters in the mammalian genome: an overview. Mol Biotechnol 45:241–247. doi:10.1007/s12033-010-9265-6
Yang W, Zhang Y, Li Y, Wu Z, Zhu D (2007) Myostatin induces cyclin D1 degradation to cause cell cycle arrest through a phosphatidylinositol 3-kinase/AKT/GSK-3b pathway and is antagonized by insulin-like growth factor 1. J Biol Chem 282:3799–3808. doi:10.1074/jbc.M610185200
Brindle PK, Montminy MR (1992) The CREB family of transcription activators. Curr Opin Genet Dev 2:199–204. doi: 10.1016/S0959-437X(05)80274-6
Chen AE, Ginty DD, Fan CM (2005) Protein kinase A signalling via CREB controls myogenesis induced by Wnt proteins. Nature 433:317–322. doi: 10.1038/nature03126
Mantovani R (1999) The molecular biology of the CCAAT-binding factor NF-Y. Gene 239:15–27
Zhang X, Odom DT, Koo SH, Conkright MD, Canettieri G, Best J, Chen H, Jenner R, Herbolsheimer E, Jacobsen E, Kadam S, Ecker JR, Emerson B, Hogenesch JB, Unterman T, Young RA, Montminy M (2005) Genome-wide analysis of cAMP-response element binding protein occupancy, phosphorylation, and target gene activation in human tissues. Proc Natl Acad Sci USA 102:4459–4464. doi: 10.1073/pnas.0501076102
Steelman S, Moskow JJ, Muzynski K, North C, Druck T, Montgomery JC, Huebner K, Daar IO, Buchberg AM (1997) Identification of a conserved family of Meisl-related homeobox genes. Genome Res 7:142–156. doi:10.1101/gr.7.2.142
Heidt AB, Rojas A, Harris IS, Black BL (2007) Determinants of myogenic specificity within MyoD are required for noncanonical E box binding. Mol Cell Biol 27:5910–5920. doi:10.1128/MCB.01700-06
Claudel T, Staels B, Kuipers F (2005) The farnesoid X receptor: a molecular link between bile acid and lipid and glucose metabolism. Arterioscler Thromb Vasc Biol 25:2020–2031. doi:10.1161/01.ATV.0000178994.21828.a7
Wilkinson DG (1992) Whole mount in situ hybridization: a practical approach. IRL Press, Oxford, pp 75–83
Rios R, Carneiro I, Arce VM, Devesa J (2001) Myostatin regulates cell survival during C2C12 myogenesis. Biochem Biophys Res Commun 280:561–566. doi:10.1006/bbrc.2000.4159S0006291X00941597
Artaza JN, Bhasin S, Mallidis CON, Taylor W, Ma KUN, Gonzalez-Cadavid NF (2002) Endogenous expression and localization of Myostatin and its relation to Myosin heavy chain distribution in C2C12 skeletal muscle cells. 179:170–179. doi:10.1002/JCP.10044
Budasz-Świderska M, Jank M, Motyl T (2005) Transforming growth factor-β1 upregulates myostatin expression in mouse C2C12 myoblasts. J Physiol Pharmacol 56:195–214
McFarlane C, Vajjala A, Arigela H, Lokireddy S, Ge X, Bonala S, Manickam R, Kambadur R, Sharma M (2014) Negative auto-regulation of myostatin expression is mediated by Smad3 and MicroRNA-27. PLoS ONE 9:e87687. doi:10.1371/journal.pone.0087687
Louis E, Raue U, Yang Y, Jemiolo B, Trappe S (2007) Time course of proteolytic, cytokine, and myostatin gene expression after acute exercise in human skeletal muscle. J Appl Physiol 103:1744–1751. doi:10.1152/japplphysiol.00679.2007
Gurtner A, Fuschi P, Magi F, Colussi C, Gaetano C, Dobbelstein M, Sacchi A, Piaggio G (2008) NF-Y dependent epigenetic modifications discriminate between proliferating and postmitotic tissue. PLoS ONE 3:e2047. doi:10.1371/journal.pone.0002047
Oldfield AJ, Yang P, Conway AE, Cinghu S, Freudenberg JM, Yellaboina S, Jothi R (2014) Histone-fold domain protein NF-Y promotes chromatin accessibility for cell type-specific master transcription factors. Mol Cell 55:708–722. doi:10.1016/j.molcel.2014.07.005
Lee MG, Pedersen PL (2003) Glucose metabolism in cancer: Importance of transcription factor-DNA interactions within a short segment of the proximal region of the type II hexokinase promoter. J Biol Chem 278:41047–41058. doi:10.1074/jbc.M307031200
Osawa H, Robey RB, Printz RL, Granner DK (1996) Identification and characterization of basal and cyclic AMP response elements in the promoter of the rat hexokinase II gene. J Biol Chem 271:17296–17303. doi: 10.1074/jbc.271.29.17296
Magenta A, Cenciarelli C, De Santa F, Fuschi P, Martelli F, Caruso M, Felsani A (2003) MyoD stimulates RB promoter activity via the CREB/p300 nuclear transduction pathway. Mol Cell Biol 23:2893–2906. doi:10.1128/MCB.23.8.2893
Zuloaga R, Fuentes EN, Molina A, Valdés JA (2013) Biochemical and biophysical research communications the cAMP response element binding protein (CREB) is activated by insulin-like growth factor-1 (IGF-1) and regulates myostatin gene expression in skeletal myoblast. Biochem Biophys Res Commun 440:258–264. doi:10.1016/j.bbrc.2013.09.067
Giono LE, Varone CL, Canepa ET (2001) 5-Aminolaevulinate synthase gene promoter contains two cAMP-response element (CRE)-like sites that confer positive and negative responsiveness to CRE-binding protein (CREB). Biochem J 353:307–316. doi: 10.1042/bj3530307
Lamph WW, Dwarki VJ, Ofir R, Montminy M, Verma IM (1990) Negative and positive regulation by transcription factor cAMP response element-binding protein is modulated by phosphorylation. Proc Natl Acad Sci USA 87:4320–4324. doi: 10.1073/pnas.87.11.4320
Ofir R, Dwarki VJ, Rashid D, Verma IM (1991) CREB represses transcription of fos promoter: role of phosphorylation. Gene Expr 1:55–60
Vallejo M, Gosse ME, Beckman W, Habener JF (1995) Impaired cyclic AMP-dependent phosphorylation renders CREB a repressor of C/EBP-induced transcription of the somatostatin gene in an insulinoma cell line. Mol Cell Biol 15:415–424. doi: 10.1128/MCB.15.1.415
Goh S, Looi Y, Shen H, Fang J, Bodner C, Houle M, Ng AC, Screaton RA, Featherstone M (2009) Transcriptional activation by MEIS1A in response to protein kinase A signaling requires the transducers of regulated CREB family of CREB co-activators. J Biol Chem 284:18904–18912. doi: 10.1074/jbc.M109.005090
Salerno MS, Thomas M, Forbes D, Watson T, Kambadur R, Sharma M (2004) Molecular analysis of fiber type-specific expression of murine myostatin promoter. Am J Physiol Cell Physiol 287:C1031–C1040. doi:10.1152/ajpcell.00492.200300492.2003
Cariou B, van Harmelen K, Duran-Sandoval D, van Dijk TH, Grefhorst A, Abdelkarim M, Caron S, Torpier G, Fruchart JC, Gonzalez FJ, Kuipers F, Staels B (2006) The farnesoid X receptor modulates adiposity and peripheral insulin sensitivity in mice. J Biol Chem 281:11039–11049. doi: 10.1074/jbc.M510258200
Rizzo G, Disante M, Mencarelli A, Renga B, Gioiello A, Pellicciari R, Fiorucci S (2006) The farnesoid X receptor promotes adipocyte differentiation and regulates adipose cell function in vivo. Mol Pharmacol 70:1164–1173. doi:10.1124/mol.106.023820.induction
Allen DL, Hittel DS, McPherron AC (2011) Expression and function of myostatin in obesity, diabetes, and exercise adaptation. Med Sci Sports Exerc 43:1828–1835. doi:10.1249/MSS.0b013e3182178bb4
Sundaresan NR, Saxena VK, Singh R, Jain P, Singh KP, Anish D, Singh N, Saxena M, Ahmed KA (2008) Expression profile of myostatin mRNA during the embryonic organogenesis of domestic chicken (Gallus gallus domesticus). Res Vet Sci 85:86–91. doi: 10.1016/j.rvsc.2007.09.014
Buckingham M, Rigby PWJ (2014) Gene regulatory networks and transcriptional mechanisms that control myogenesis. Dev Cell 28:225–238. doi:10.1016/j.devcel.2013.12.020
Kang JH, Noh ES, Park JY, An CM, Choi JH, Kim JK (2015) Rapid origin determination of the Northern Mauxia Shrimp (Acetes chinensis) based on allele specific polymerase chain reaction of partial mitochondrial 16S rRNA gene. Asian-Australas J Anim Sci 28(4):568. doi:10.5713/ajas.14.0613
Acknowledgements
We thank Dr. Mônica Senna Salerno, from AgResearch, New Zealand, for providing the tE4 construct. Special gratitude is due to Dr. Ajeesh Balakrishnan-Renuka, for his help with cell culture, Rana Houmany and Swantje Wulf, for technical assistance at Ruhr-Universität Bochum, Germany.
Funding
This work was supported by National Counsel of Technological and Scientific Development - CNPq [480960/07-0; 142034/2009-6], Fundação de Apoio ao Ensino, à Pesquisa e Extensão - FAEPEX [1039/07], DeutscherAkademischerAustauschdienst - DAAD/CNPq [290068/2010-0], Forschungsförderung Ruhr-Universität Bochum MedizinischenFakultät - FoRUM [6308100907] and Programa Institucional de Bolsas de Iniciação Científica - PIBIC/CNPq [123066/2013-1]. All grant sponsors had no involvement in the conduct of the research and/or preparation of the article.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2017_4126_MOESM1_ESM.tif
Online Resource Fig. 1 Strategy of site-directed mutagenesis used to generate a series of single or multiple mutations in the MstnP/E-Luc construct. Round 1: the four constructs containing single mutated TFBSs were obtained using the wild-type MstnP/E-Luc vector as template; Rounds 2, 3 and 4: multiple binding site mutations were sequentially generated using previously mutated constructs as templates, as schematized above. In our strategy we chose to insert restriction sites (EcoRI, ClaI, PstI, SmaI or NotI, underlined) to disrupt the TFBS (changed nucleotides shown in red) and to facilitate the screening of the clones. All the mutant MstnP/E-Luc constructs were sequenced to confirm the mutagenesis success. The construct obtained in Round 4 with all TFBS disrupted was called ΔMstnP/E-Luc. (TIF 741 KB)
11033_2017_4126_MOESM2_ESM.tif
Online Resource Fig. 2 Analysis of the effects caused by the disruption of TFBS on Mstn P/E activity. After transfection of a series of mutant MstnP/E-Luc constructs into C2C12 mouse myoblasts, effects of single or multiple disruption of TFBS on Mstn P/E were evaluated by changes in luciferase activity in comparison to the wild-type MstnP/E control (black square). Results of upregulation (green up arrows) or downregulation (red down arrows) in P/E activity are shown for each construct. Results were analyzed starting with the construct containing all four TFBS disrupted (ΔMstnP/E-Luc) until getting to the constructs with single TFBS mutated, in order to mimic the effects of the progressive restoration of one, two or three sites to ΔMstnP/E. (TIF 216 KB)
11033_2017_4126_MOESM3_ESM.docx
Online Resource Table 1 Gene specific forward (F) and reverse (R) primers, annealing temperatures and predicted sizes of products generated by RT-PCRs assays for expression analysis in C2C12 cells. The same set of primers was used to generate templates for RNA probes synthesis with the addition of a T7 tail sequence (5´-TAATACGACTCACTATAGGGAGA-3´) to the 5’-end of R primers. Mstn F1 and R1, Myogenin and 16S primers were employed only for RT-PCRs. The second pair of Mstn primers (F2 and R2) was used exclusively to produce templates for RNA probe synthesis. *16Sbr and 16Sar are universal primers that generate a 570 bp PCR product of the 16S rRNA mitochondrial gene used here as endogenous control [60]. (DOCX 23 KB)
11033_2017_4126_MOESM4_ESM.docx
Online Resource Table 2 Primers used for site-directed mutagenesis of the Mstn P/E expression construct. Nucleotides replaced during site-directed mutagenesis are in red; nucleotides substituted in a previous mutagenesis round are shown in blue. For details see Fig.1 of the Online Resource. F: Forward primer; R: Reverse primer. (DOCX 20 KB)
Rights and permissions
About this article
Cite this article
Grade, C.V.C., Mantovani, C.S., Fontoura, M.A. et al. CREB, NF-Y and MEIS1 conserved binding sites are essential to balance Myostatin promoter/enhancer activity during early myogenesis. Mol Biol Rep 44, 419–427 (2017). https://doi.org/10.1007/s11033-017-4126-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-017-4126-z