Abstract
Natural genetic variation is randomly distributed and gene expression patterns vary widely in natural populations. These variations are an effect of multifactorial genetic perturbations resulting in different phenotypes. Genome-wide analysis can be used to comprehend the genetic basis governing this naturally occurring developmental variation. Secondary growth is a highly complex trait and systems genetics models are presently being applied to understand the molecular architecture of wood formation. In the present study, the natural variation in expression patterns of 18,987 transcripts expressed in the developing xylem tissues were documented across four phenotypes of Eucalyptus tereticornis with distinct holocellulose/klason lignin content. The differentially expressed genes across all the phenotypes were used to construct co-expression networks and sub-network 2 with 380 nodes and 17,711 edges was determined as the network of relevance, including 30 major cell wall biogenesis related transcripts with 2394 interactions and 10 families of transcription factors with 3360 interactions. EYE [EMBRYO YELLOW] was identified as major hub transcript with 173 degrees which interacted with known cell wall biogenesis genes. K-mean clustering was also performed for differentially expressed transcripts and two clusters discriminated the phenotypes based on their holocellulose/klason lignin content. The cluster based networks were enriched with GOs related to cell wall biogenesis and sugar metabolism. The networks developed in the present study enabled identification of critical regulators and novel transcripts whose expression variation could presumably govern the phenotypic variation in wood properties across E. tereticornis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genus Eucalyptus belonging to family Myrtaceae, consists of more than 900 species [1] which are native to Australia and its neighboring Islands. It is one of the widely planted hardwood species because of its superior growth, adaptability and wood properties and occupies 19.61 M hectares globally. India ranks second in area under eucalypts plantation (3.943 M ha) after Brazil (4.259 M ha) [2]. This genus is targeted world-wide for genetic improvement programs due to its high commercial value as raw material for paper and pulp industries.
Secondary growth leading to wood formation in plants is an exceptionally complex developmental event controlled by numerous gene families involved in the biosynthesis of polysaccharides, lignin and cell wall proteins. The molecular regulation of wood formation is reported in tree species like Populus [3–5], Pinus [6, 7], Picea [8] and Eucalyptus [9–16]. Most of these studies were limited to understanding the gene functions regulating the most relevant biological pathways governing secondary cell wall biogenesis. However, wood formation is an intricate process involving thousands of genes, most of which have unknown functions [17] and intricate cellular and molecular networks and cross talks govern the phenotypic variation [18]. Systems model for wood formation will thus enable an in depth understanding of important transcriptional and metabolic regulators in tress with industrially suitable phenotypes [19] and provide an insight into the role of individual genes, entire pathways, biological networks which are essential, redundant, auxiliary or unique to wood formation [20].
Gene networks specific to secondary cell wall formation is reported in Arabidopsis, poplar, pine, white spruce, rice and sugarcane [5, 21–31]. The present study was taken up to document the genome-wide natural variation of transcript expression in developing xylem tissues of four Eucalyptus tereticornis genotypes and develop gene co-expression networks to identify crucial regulators governing variation in wood properties in this tropical Eucalypt species.
Materials and methods
Plant material
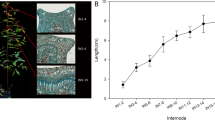
Four genotypes of E. tereticornis (SWMG-6, CW-8, KUP-14, NKR-49) were selected for expression profiling based on their holocellulose and klason lignin content (Table 1). The holocellulose and klason lignin content of the core samples was determined by NIR spectroscopy. Two wood core samples of approximately 2.0 cm length were collected at a height of ~1.3 m using increment borer (Haglof Inc., Sweden) from nineteen year-old standing trees available in the seed orchard established at Karunya Research Station, Coimbatore, India. The core samples were debarked, powdered, air dried and passed through 40–60 mesh screen (250–400 µm) to maintain uniform particle size. The duplicate powdered samples were pooled in equal quantity prior to sieving. The spectra were generated in the entire NIR range of 4000–12820 cm−1 using MPA FT-NIR Spectrometer (Bruker Optics, Ettlingen, Germany). Thirty-two scans per sample in diffuse reflectance mode were collected to obtain an average spectra. The spectral data was used to determine the holocellulose content using the model reported by Kothiyal et al. [32], while the klason lignin content was determined using the model developed by Kothiyal et al. (unpublished data).
Gene expression analysis was conducted in the same genotypes. The developing xylem from two wood core samples (after debarking 1.5 cm from periphery) was harvested and immediately frozen in liquid nitrogen and stored at −80 °C until RNA isolation.
RNA isolation
Total RNA was extracted from developing xylem tissues of duplicate wood core sample using Spectrum™ Plant Total RNA Kit (Sigma Aldrich, USA). The concentration and purity of the RNA was evaluated using NanoDrop ND-1000 UV–Vis Spectrophotometer (Thermo Scientific, USA) and RNA integrity was confirmed using 2100 Bioanalyzer (Agilent Technologies Inc., Santa Clara, CA). The total RNA from duplicate samples was pooled in equimolar concentration prior to labeling and hybridization.
Array design
A total of 18,987 transcripts were selected for the microarray analysis. Sequences were sourced from RNA-Seq data of differentiating xylem tissues [33] and transcript sequences identified from E. tereticornis [34] (Table 2). A 8x60 K microarray chip was custom-designed in Agilent platform (AMADID: 059849) consisting of 44,817 probes and the number of specific probes in the array was 81 % and the remaining 19 % probes were cross hybrid probes. The size of the probes was sixty base pairs and a minimum of two probes per transcript was designed.
Functional annotation of transcripts
Transcripts were functionally annotated and their position in chromosome, protein domains, biological pathways and gene ontology were defined based on the recent assembly of Eucalyptus grandis using Phytozome v10.1 (accessed on January 2015, E. grandis v1.1 Phytozome v10.1). The pathways were determined using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database [35, 36] and MAPMAN, a user-driven tool providing pathway and biological process information [37]. Further, Eucalyptus nucleotide sequences were used to search the complete protein sequences of Arabidopsis using BlastX with (e-value cut off of 1e−5) in the non-redundant database of NCBI and TAIR (v10) and the best hits (lowest e-value) was selected as Arabidopsis orthologs. Transcript IDs were given based on their Arabidopsis orthologs and transcripts which had no orthologs in Arabidopsis were given the E. grandis ID from Phytozome.
Sample labeling and hybridization
RNA sample preparation, labeling and hybridization was done using one-color microarray-based gene expression analysis with Tecan HS Pro protocol (Agilent Technologies, CA, USA) as per manufacturer’s protocol. Briefly, 500 ng each of RNA samples were converted to cDNA, primed by oligo dT with T7 polymerase by reverse transcription. Synthesized cDNA was then used as template for cRNA generation along with the incorporation of Cy3 CTP. Fluorescently labeled cRNA was purified using RNeasy mini kit (Qiagen, Valencia, USA), followed by assessment of quality and quantity using 2100 Bioanalyzer (Agilent Technologies, CA, USA). The Cy3 labeled cRNAs were fragmented and hybridized to the array at 65 °C for 16 h and the slides were scanned using Agilent Microarray Scanner G2505C and the features were extracted with the Feature Extraction Software (Agilent Technologies, v12).
Documentation of differential expression of transcripts across genotypes
Feature extracted data was analyzed using GeneSpring GX Version 12 software (Agilent Technologies, CA, USA). After background correction, the data was log transformed and normalized. Global normalization of the data was done in GeneSpring GX using the 75th percentile shift and normalization across samples was done using median values.
The log2 fold expression data was filtered for significantly regulated (up and down regulated) transcripts across all genotypes based on the criteria where in the cutoff for fold change was set at >0.8 and Flags “detected” in respective samples for up-regulated transcripts, while the cutoff for fold change <−0.8 and Flags “detected” was used for down-regulated transcripts. The differential expression of transcripts across all possible pair-wise combination was performed. Transcripts exhibiting ± 0.8 fold difference in expression with a statistical significance of p < 0.05 were considered as differentially regulated.
Validation of expression data by RT-qPCR
The expression pattern generated by the array experiment was validated by RT-qPCR for fifteen transcripts (Supplementary Table S1). The reactions were performed in ABI PRISM 7500 Step one plus Sequence Detection System (Applied Biosystems, USA) using the following program: one cycle of 95 °C for 10 min; 40 cycles of 95 °C for 15 s and 60 °C for 1 min. The PCR reaction for 25 μl included 200 ng of cDNA, 12.5 μl of SYBR Green PCR Master Mix (Applied Biosystems, USA), 200 nM of forward/reverse primer and Milli-q water. The primers used for the study were designed using Primer 3.0 software and their details are given in Supplementary Table S1. All reactions were conducted in triplicates. Melting curve analyses was done to ensure product specificity and to differentiate between the product and primer dimers. EtACT and EtSAND were used as reference genes for normalization of data [38]. The RQ value was calculated using the formula 2−ΔΔCT [39].
Construction of co-expression networks
The expression levels of differentially expressed transcripts (1130) was used to develop the co-expression network. Pearson correlation coefficient was used to measure the similarity of expression between pair-wise transcripts. Transcript pairs with r > 0.97 (positive co-expression) or r < −0.97 (negative co-expression) were considered as significantly co-expressed, based on the report by Freeman et al. [40]. Co-expression network was constructed with 1130 nodes and 49,786 edges using Cytoscape 3.2 (http://www.cytoscape.org) using default parameters [41]. Each node represented a transcript and the edge represented the strength of co-expression. Duplicated edges and self loops were manually removed from the network. The degree was calculated for each node based on the number of edges connected to a node and highly connected nodes (hub transcripts) were identified from the network.
Gene ontology enrichment and pathway identification
Assessment of over-representation of gene ontology (GO) was performed using ClueGO Cytoscape plugin (http://apps.cytoscape.org/apps/cluego). The statistical significance for all GOs for biological process (BP), molecular function (MF) and cellular component (CC) was evaluated with default parameters (kappa score 0.4). GO annotation terms were considered significant if the corrected p value (False discovery rate) was <0.05 and if there were at least 4 transcripts associated with the same annotation [42]. The major biological pathways of the transcripts represented in the network were determined as mentioned earlier.
Cluster analysis of differentially expressed transcripts
Differentially expressed transcripts were subjected K-mean clustering with J-Express Pro software [43] using Euclidean distance measure and Forgy initialization method.
Construction of cluster guided co-expression network
Co-expression network was constructed for each cluster using Cytoscape 3.2 and assessment of over-representation of GO within a cluster was performed using ClueGO Cytoscape plugin. The major biological pathways of the transcripts represented in the network were also determined.
Results
Functional annotation of transcripts
The Phytozome internal transcript ID for E. grandis, transcript ID, PFAM, Panther, KOG, KEGG, EC, KEGG Orthology, GO terms obtained for 18,987 transcripts represented in the array is presented in Supplementary Table 2. The transcripts spanned all the 11 chromosomes of the Eucalytpus genome. Chromosome 6 had a maximum of 2303 transcripts, while chromosome 4 had a minimum representation of 1112 transcripts.
GO mapping of the transcripts were performed to classify the functions and categorize them into cellular components, molecular function and biological process. The cellular component included 29,468 transcripts, while the number of transcripts grouped under biological process was 29,108. Molecular functions which describe the biochemical activity performed by the gene product grouped 29,629 transcripts in the present study. Gene Ontology categories overlap and therefore several transcripts are classified into more than one GO term. The categorization of transcripts in 45 GO categories is given in Supplementary Fig. 1.
One hundred and twenty-six pathways were determined using the KEGG and MAPMAN databases. The pathways and the number of transcripts under each pathway are listed in Supplementary Table 3. The major biologically relevant pathways represented in the array included carbon metabolism (147 transcripts); plant hormone signal transduction (104 transcripts); starch and sucrose metabolism (88 transcripts) and phenyl propanoid biosynthesis (44 transcripts).
RNA isolation and transcript profiling using customized array
Total RNA derived from developing xylem tissues of four genotypes of E. tereticornis was assessed for quality and quantity. The RNA concentration ranged from 189 to 305 ng/μl, with OD 260/280 = 1.71 to 1.95 and RNA Integrity Number ranged from 4.6 to 7.9. The expression patterns of 18,987 transcripts were determined using the custom-designed microarray chip. The expression patterns were determined within and between genotypes and their log2 fold expression ranged from −9.70 to 12.79. The expression data was submitted to NCBI’s Gene Expression Omnibus with accession number GSE73030.
Documentation of differentially expressed transcripts
Differential expression of transcripts across all possible pair-wise combination was performed. Across the four genotypes, significant up and down regulation was documented in 1822 and 2174 transcripts respectively. The fold expression of down-regulated transcripts ranged from −0.80 to −8.74, while the fold expression of up-regulated transcripts ranged from 0.80 to 8.45. All pair-wise comparison of differentially expressed transcripts is given in Fig. 1. The total number of transcripts which were differentially expressed across all genotypes (after removal of overlapping transcripts) was 1130.
Transcripts with significant differential expression were from different pathways including protein processing in endoplasmic reticulum (15), biosynthesis of amino acids (14), carbon metabolism (13), starch and sucrose metabolism (11), plant-pathogen interaction (11), plant hormone signal transduction (10), alanine, aspartate and glutamate metabolism (10), arginine and proline metabolism (8) and carbon fixation in photosynthetic organisms (8) (Supplementary table S4).
The major transcripts which differentially expressed across the four genotypes included HEX, INV and CesA from cellulose biosynthesis pathway; CAD, COMT, CCR, HCT from lignin biosynthesis pathway; cell wall related transcripts like myosin family protein; COBRA-like4, EXPA, XTH, FLA, TUA; enzymes including GH, XTH, UGT, pectin acetyl esterase, pectin methylesterase inhibitor, hydroxyproline-rich glycoprotein, pectate lyase, peroxidase, UDP glucose dehydrogenase, UDP-mannose dehydrogenase; heat shock proteins and transcription factor families from NAC, VND, MYB, HD-ZIP and Znf.
Validation of gene expression pattern using RT-qPCR
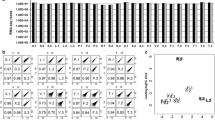
Fifteen major cell wall biosynthesis related transcripts were selected for validation of microarray generated expression pattern using RT-qPCR. The fold expression of the selected genes ranged from −3.54 to 7.88. Comparison of the expression patterns between microarray and RT-qPCR revealed that CAD, CCR, CCoAOMT, CesA4, CesA8, GAPC2, HCT, UGD3 had similar expression patterns, validating the microarray generated expression profiles (Fig. 2).
Development of co-expression network
The expression level of differentially expressed transcripts was used to construct the co-expression network. Initially, a correlation matrix was generated by calculating pair-wise Pearson correlation using normalized expression values across all samples. The number of significantly co-expressed transcripts (threshold >0.97) was 1130. The network was constructed with 1130 nodes (transcripts) and 49,786 edges using Cytoscape 3.2.0 and three sub-networks were generated. The GO enrichment was conducted for all sub-networks.
Sub-network 1 consisted of 381 nodes with 15,898 edges and the degree ranged from 6 to 141. This sub-network was significantly enriched with intrinsic component of plasma membrane (LYN2, XYP1, SWEET1, LPP2, VSR6, SA629); arginine and proline metabolism (ERD5, GDH1, PYD1); carbon fixation in photosynthetic organisms (GAPC2, GAPA2); cellular transition metal homeostasis (MT2A, ACA9); regulation of flavonoid biosynthetic process (PP2-A1, HCT, RPS2); negative regulation of cell death (LRR XI-23, PAD4); nucleotide transport (AAP3, NTF2B, LHT1) and microsporogenesis (AGL16, MOB1-like) (data not shown).
Sub-network 3 consisted of 369 nodes and 16,177 edges with degrees ranging from 4 to 152. The GOs in this network included phosphatidylglycerol biosynthetic process (CCD1, PHR2, CIP7, FBA2); cytoskeleton (TUA5, MAP70-5, CRK); cell wall pectin metabolic process (PERK1, QUA3); cellular biogenic amine biosynthetic process (SPDS1, TRP1); sodium ion trans membrane transport (HKT1, NHX1); regulation of signal transduction (CPL3, JAZ10, SDIR1, IBR5]; nitrate assimilation (NIA2, NRT1); auxin polar transport (ZIFL1, AVP1); regulation of cell morphogenesis (ASK2, RPN12, ROPGEF1) and membrane region (HMGR1, bZIP60, IRE1A) (data not shown).
The biologically relevant sub-network 2 consisted of 380 nodes, 17,711 edges and the degree ranged from 4 to 173 (Fig. 3). The top hub transcripts in this sub-network are given in Table 3. The sub-network included 30 major cell wall biogenesis related genes like CesA7 (MUR10), IRX15-L, IRX6, XTH9, ACO1, C3H, F5H, CCR-like25, COMT-like65, TUA5 with 2394 interactions. Similarly, 10 families of transcription factors including R2R3-MYB, NAC, WRKY, bHLH, GRAS, RAV, ABI3VP1, C3H, C2H2 and MADS -Type I were documented in this network with 3360 interactions. Hundred and sixty-four transcripts were found with degree greater than 100. Two hundred and seventy-five transcripts had degree greater than fifty, revealing extreme complexity of the network.
EYE (EMBRYO YELLOW), a conserved oligomeric golgi complex component-related transcript was identified as major hub transcript which interacted with 173 other transcripts. This transcript was found to interact with known cell wall biogenesis genes like CesA7, IRX15-L, IRX6, TUA5, TBL34, VND7, MYB20 and SCL14. Eucgr.C02641 (trans-membrane receptors; ATP binding) was identified as the second major hub transcript which interacted with 171 other transcripts. This transcript is yet to be functionally characterized in Arabidopsis. Annotation results indicated that it had a role in signal transduction, defense response (biological process), ATP binding (molecular function) and encodes a protein located in cytoplasm. It interacted with major transcripts like CesA7, IRX6, IRX15, MYB20 and TUA5.
The other major hub transcripts included a cold responsive transcript (SRC2), Eucgr.I00967.4 and dihydroneopterin aldolase (FOLB1) which documented 170, 167 and 166 interactions respectively. These transcripts shared common interactions with CesA7, IRX6, IRX15, XTH9, TUA5, TBL34, PERK10 and transcription factors like MYB20, VND7, FEM111 and SCL14. Eucgr.I00967.4 interacted with ATB2_a, CPK26, GRXC2, HA11, PNG1, RPN5B and STP14, while FOBL1 co-expressed with ACN1, DNAse I-like superfamily protein (Eucgr.E03549.1) and Xanthine/uracil permease family protein (Eucgr.K02873.1).
One of the major cell wall biosynthesis related transcript, cellulose synthase (CesA7) co-expressed with ninety transcripts including IRX15-L, XTH9, LGT6, ACO1, TBL34, ATB2, PERK1, AGL80, RAV1, HXK3 and MYB20. The monolignol biosynthesis pathway gene, cinnamoyl CoA reductase (CCR), which catalyze the conversion of the side-chain carboxyl to an alcohol group, co-expressed with 58 transcripts including ROP4, WRKY15 and UBQ5. Tubulin (TUA5) involved in deposition of cellulose microfibrils during cell wall formation in plants, co-expressed with 150 transcripts like IRX6, UGT76E2, UBQ5, BGAL12, sks5, ROP4 and transcription factors including WRKY15, VND7 and SCL14, while IRX6 (IRREGULAR XYLEM 6) or COBL4 (COBRA-like 4) had co-expression with 142 transcripts including VND7, UGT76E2, TUA5, sks5 and FAH1. Two major transcription factors VND7 and MYB20 were present in the network. VND7 interacted with 142 transcripts including structural genes like IRX6, F5H1, TUA5, sks5, TBL34 and transcription factors like AGL80, PAT1, HSL1, while MYB20 was found to interact with transcripts involved in cell wall biogenesis like CesA7, IRX15-L, SCL14, XTH9, BGAL12, FEM111 and sks5.
The GO enrichment confirmed that sub-network 2 was significantly enriched with cell wall biosynthesis related biological processes, molecular functions and cellular components. It included genes involved in lignin biosynthetic process (CYP98A3, FAH1, CAD1); glucuronoxylan metabolic process (TBL34, GAUT12, IRX15L); plant-type vacuole (ESL1, SRC2, ABCC3, GSTF12); sugar:hydrogen symporter activity (ESL1, INT2, G-TMT, STP14, G3Pp4); phosphatidylglycerol biosynthetic process (LSMT-L, MCA1, PMDH2); nucleotide transport (SRC2, NTF2B, AMT2, UTR6); hyper osmotic response (RABA1b, ACO1, BIG, PP2-A1, GS2, HVA22E, HAI1, LTP3) and serine family amino acid metabolic process (PSAO, D-CDES, PSAG, LHCA3, ALDH11A3, LHCB4.3, MCA1, CGL1, LHCB4.1, PMDH2) (Fig. 4).
Identification of novel transcripts in sub-network 2
The sub-network 2 included several unknown transcripts with significant expression variation across the four genotypes. EYE was identified as major hub gene and interacted with known cell wall biogenesis pathway transcripts. Till date there are no reports implicating this gene in secondary development in woody perennials. Similarly, transcripts like SRC2, protein kinase, SnRK3.16, exotosin, XIH (Myosin family protein) and O-fucosyltransferase which had significant expression patterns in developing xylem tissues were present in the network. Their function during wood formation is yet to be elucidated. Additionally, twelve Eucalyptus specific transcripts including Eucgr.B00912.1, Eucgr.C00456.2, Eucgr.C02271.1, Eucgr.C02390.1, Eucgr.E01070.1, Eucgr.E04290.1, Eucgr.F03259.2, Eucgr.G00681.1, Eucgr.H00168.2, Eucgr.H03075.1, Eucgr.H04013.3, Eucgr.I00224.6, Eucgr.J02973.2, Eucgr.K01518.2 with no orthologs from Arabidopsis was also documented in the network.
Cluster analysis of differentially expressed transcripts
K-mean clustering was performed with 1130 transcripts differentially expressed across genotypes, to identify specific groups that demonstrated the strongest expression correlation. Six clusters were identified from this analysis (Fig. 5). In cluster 1, 267 transcripts were up-regulated in CW-8 and NKR-49 and the corresponding transcripts were down-regulated in SWMG-6 and KUP-14. Cluster 2 comprising of 110 transcripts grouped the up-regulated transcripts in SWMG-6 and NKR-49 and down regulated transcripts in CW-8 and KUP-14. Cluster 3 with 259 transcripts consisted of up-regulated transcripts from CW-8 and KUP-14, while down-regulated in SWMG-6 and NKR-49. Cluster 6 (115 transcripts) had transcripts up-regulated in SWMG-6 and KUP-14 and down-regulated in CW-8 and NKR-49.
Two hundred and forty-eight transcripts with significantly up-regulated expression patterns in Cluster 4 was found in genotypes with high klason lignin content (KUP-14 and NKR-49) when compared to their corresponding expression in genotypes with low klason lignin content (SWMG-6 and CW-8). Cluster 5 with 132 transcripts consisted of transcripts up-regulated in genotypes with high holocellulose content (SWMG-6 and CW-8) and down-regulated in genotypes with low holocellulose content (KUP-14 and NKR-49). The two clusters could differentiate the genotypes based on their phenotypic attributes for klason lignin and holocellulose content.
Cluster based co-expression networks
Co-expression networks were constructed for all the six clusters to identify interactions between transcripts. The number of significantly co-expressed transcripts (threshold >0.97) varied for each cluster. The six cluster based networks were represented with 254, 110, 249, 235, 132 and 112 transcripts respectively.
Cluster 1 based network consisted of 254 nodes, 7907 edges and the degree ranged from 4 to 132. GO enrichment analysis revealed that the network consisted of transcripts with functions in intrinsic component of plasma membrane, nucleotide transport and substrate-specific channel activity. The cluster 2 based network consisted of 110 nodes, 1390 edges with degree ranging from 4 to 102. Transcripts involved in membrane related functions were represented in the network. The network derived from cluster 3 consisted of 249 nodes, 7872 edges and the degree ranged from 2 to 134. The major GO representations were from ABA signaling, sodium ion transport and carotenoid biosynthesis. Cluster 6 derived network consisted of 112 nodes, 1263 edges and the degree ranged from 2 to 50 and was enriched with transcripts involved in carboxypeptidase activity (Table 4). The top hub transcripts in these networks are presented in Table 5.
Cluster 4 and 5 based networks were enriched with GOs related to cell wall biogenesis and sugar metabolism. Cluster 4 based network consisted of 235 nodes, 7649 edges with degree ranging from 1 to 173 (Fig. 6). The GOs predominant in this network (specific in genotypes with high klason lignin and low holocellulose content) included transcripts involved in lignin biosynthetic process, regulation of flavonoid biosynthetic process, cell wall polysaccharide metabolic process, cell wall pectin metabolic process, regulation of anatomical structure morphogenesis and regulation of ion transport (Table 4; Fig. 6). The major transcripts in this network were CAD1, TUA5, SCL14, CesA7, IRX15-L, PERK1, LGT6, MYB20 and sks5. The top listed hub transcripts with degree of 173, 120 and 120 were NAD(P)-linked oxidoreductase superfamily (ATB2), EMBRYO YELLOW (EYE) and soybean gene regulated by cold (SRC2) respectively (Table 5).
Cluster 5 based network was constructed with 132 nodes and 1987 edges and the interactions ranged from 2 to 58 (Fig. 7). This network which grouped genotypes with high holocellulose content was enriched in GOs representing sugar:hydrogen symporter activity, fructose and mannose metabolism, glycolysis and glueconeogenesis, tyrosine alanine aspartate and glutamate metabolism and transcripts involved in carbon fixation (Table 4; Fig. 7). The major differentially expressed genes in this network included HXK3, PMDH2, WRKY, UGT76, UGT85, VND7, G3Pp4 and XIH. Heat shock protein (HSP101) was identified as top hub transcript with 58 degree (Table 5). The above results indicate that the two cluster based networks (4 and 5) were relevant to the phenotypic groups of E. tereticornis.
Discussion
Genome-scale expression analysis to understand secondary development
Wood formation is a complex dynamic process regulated by diverse metabolic pathways and significantly influenced by external stimuli like photoperiod, nutrient availability, moisture content and temperature [44, 45]. Genome scale studies on secondary cell wall development are reported from perennial species like Populus, Picea, Pinus and Eucalypts [8, 13, 16, 33, 46–49].
The present study was undertaken to identify the expression patterns of transcripts expressed in developing xylem of four genotypes of E. tereticornis with distinct wood property traits for holocellulose and klason lignin content and develop co-expression networks to understand the molecular processes determining the intra-specific variation in wood properties. A total of 18,987 transcripts expressed in the developing xylem tissues of different Eucalypt species were selected for designing the microarray. The chromosomal locations of all transcripts were derived using the recent assembly of E. grandis genome and they spanned all 11 chromosomes. This is in agreement with the recent report from E. grandis, wherein genome-wide occurrence of genes regulating cell wall lignification was documented [50]. The annotation of the E. grandis genome also revealed that gene families involved in cellulose and xylan biosynthesis pathways were randomly distributed in the genome [51]. Further, major transcription factor families regulating wood formation like R2R3 MYB and NAC were also localized on several chromosomes of E. grandis [52, 53].
Natural variation of transcript expression in wood forming tissues
Significant natural variation exists within tree species, which determine their adaptations to different environments [54]. Genome-wide analysis can be used to comprehend the genetic basis governing this naturally occurring developmental variation. The analyses of natural inter and intra-specific variation can aid in discovery of genes involved in trait expression and adaptation to different environments [55].
Wood properties are considered as a highly plastic trait, since several different types of wood can occur within a single tree. This variation can be attributed to the natural differences in expression patterns of genes/proteins regulating wood formation [44]. Intra-specific natural variations in expression of xylem-related genes has been reported from several woody perennials. In P. taeda, microarray analysis of genes expressed in developing xylem of two populations with difference in wood specific gravity revealed significant variation in expression of 131 ESTs, implicating the effect of gene expression on phenotypes [56]. In another study on P. taeda, gene expression variation in 400 clones representing the natural range of the species was reported. Hundred and six genes showed significant expression differences (fold expression ranging from 1.4 to 5.8) across the clones [22]. In E. nitens, Thavamanikumar et al. [57] reported significant gene expression variation ranging from −6.79 to 8.18 in genotypes with low and high kraft pulp yield. Recently, expression levels of genes involved in lignocellulose formation was reported in two Eucalyptus hybrid genotypes (E. urophylla x E. grandis) with different lignin content [49]. Considerable variation in gene expression was reported across the two genotypes specifically for the monolignol biosynthesis- related genes. In the present study, a significant expression variation in 1130 transcripts with log2 fold ranging from −9.70 to 12.79 was documented among the four E. tereticornis genotypes. The large amount of variation in expression observed in all the above studies support the idea that expression differences is an important factor governing variations in wood properties.
Intra and inter specific expression variation of transcripts regulating major cell wall biogenesis related pathways like cellulose, xylan and lignin are reported in several tree species. In E. nitens, genes like CesA3, HB1, Znf1 and NAM1 were differentially expressed across two genotypes selected for variable pulp yield [58], while CesA, COBL, FLA, expansin, pectin-lyase, plant invertase/pectin methylesterase inhibitor, glycosyl hydrolase, UDP glycosyl transferase and transcription factors including NAC and MYB were differentially expressed across two populations with low and high kraft pulp yield [57]. In E. grandis, microarray analysis was conducted for 1578 cDNA clones across seven different genotypes derived from two un-related open pollinated trials and the transcripts which registered significant expression variation included CCoAOMT, XET, UXS and COMT [12].
Similarly, in E. globulus, E. grandis and E. urophylla, inter-specific variations in gene expression was reported for CesA, EXPA, HCT, CCR, CAD and COMT [33]. In a recent study, the transcriptome of developing xylem tissues from two Eucalyptus hybrid genotypes differing in lignin content were compared and differentially expressed genes included INV, HEX from cellulose biosynthesis pathway, PAL, C4H, 4CL from lignin biosynthesis pathway and UXS, IRX10, GATL from xylan biosynthesis pathway. Four major transcription factors from NAC, WRKY, AP2/ERF and KNOX families were also reported to show variable expression across the genotypes [49].
In the present study, the expression of 1130 transcripts were significantly variable across the four genotypes and we documented homologous gene families reported from earlier studies to differentially express in E. tereticornis. The orthologs of major transcripts like HEX, INV CesA, CAD, COMT, CCR, HCT, COBRA-like4, EXPA, XTH, FLA, TUA, GH, XTH, UGT, pectin acetyl esterase, pectin, pectate lyase, peroxidise and transcription factor families from NAC, VND, MYB, HD-ZIP and Znf were differentially expressed across the genotypes. The above studies reveal that the functional homologs governing secondary development processes are analogous in both angiosperms and gymnosperms and genes regulating secondary cell wall biogenesis show high level of conservation in expression.
Cluster analysis: an explanatory technique for analysis of expression data
The complexity of biological networks has driven researchers to analyze large expression datasets to decipher gene interactions across different pathways. Cluster analysis is often employed to identify co-regulated groups of genes across multiple experimental samples. These gene sets are often over represented in specific clusters due to similar function [59]. There are several reports wherein this approach has been used to correlate transcript expression with traits of interest. In Rorippa spp., K-mean clustering was applied to identify gene clusters associated with extreme submergence tolerance [60], while in Arabidopsis, early and late ABA responsive gene clusters were identified in seeds [61]. In Populous tremula, anatomically homogenous cell layer specific gene clusters were reported from the meristem zone [3], while in E. nitens, hierarchical clustering was used to group genes expressed in high and low kraft pulp yielding trees [57]. In E. urophylla × E. grandis hybrids, two different groups of xylem related genes were expressed during nitrogen availability using cluster analysis [62]. In the present study, clustering of differentially expressed genes was conducted using K-mean algorithm and six clusters were generated. Two clusters ( 4 and 5) could differentiate the genotypes based on its phenotypic characters. The relevance of the clusters in terms of gene functions was validated using GO enrichment analysis. Enrichment of GOs in clusters with biologically relevant functions during xylogenesis are reported in Arabidopsis and Populus [21], E. nitens [58], E. grandis x E. urophylla [16] and P. trichocarpa [26].
Systems genetics approach to comprehend secondary development
Secondary development is an intricate process involving thousands of genes, most of which have unknown functions [17]. Systems model for wood formation are presently being used to obtain a comprehensive understanding of critical transcriptional and metabolic regulators of wood biosynthesis in mature trees with industrially suitable phenotypes [19]. These predictive models will provide an insight into the role of individual genes, entire pathways or biological sub-networks which are essential, redundant, auxiliary or unique to wood formation [20].
Gene co-expression networks and transcriptional networks relevant to wood formation reported till date have targeted secondary cell wall formation in species like Arabidopsis [21, 27, 30], poplars [5, 21, 26], P. taeda [22], sugarcane [31] and rice [29]. In Arabidopsis, major cell wall related gene families were represented in the network including cellulose synthases, glycoside hydrolases, glycosyl transferases, exostosin, kinase/LRR superfamily, plastocyanin-like family, xyloglucan endotransglucosylase/hydrolases, expansin and COBRA- like and transcription factor families like MYB, NAC, HB, and WRKY [21, 24, 27, 30, 63]. In Populus, genome wide co-expression network and transcriptional network related to cell wall biosynthesis comprised similar gene families like cellulose synthase family, glycoside hydrolase family, glycosyl transferase family, exostosin family, kinase/LRR superfamily, plastocyanin-like family and xyloglucan endotransglucosylase/hydrolase [XTH] family and TF families like MYB, NAC and HB [5, 21, 26]. Analogous gene families were represented in the cell wall related networks reported from gymnosperms like P. taeda [22]; Picea glauca [28] and from monocots like sugarcane [31] and rice [29].
In the present study, the sub-network 2 was enriched with 30 major cell wall biogenesis related genes, while cluster 4 and 5 based network were also enriched with GOs related to cell wall biogenesis and sugar metabolism. Several transcripts reported in cell wall biosynthesis of other plant species like Arabidopsis, poplar, pine and rice were also found in Eucalyptus co-expression networks including CesA7, IRX6 (COBL4), IRX15, XTH9, CCR, SND, MYB20 and VND7. All above mentioned studies reveal that genes regulating secondary cell wall biogenesis pathways are highly similar, indicating that biological pathways functionally relevant to secondary growth are conserved across gymnosperms and angiosperms [8, 9, 64–67].
However, several unknown transcripts like domain of unknown function (DUF231, DUF966, DUF3511), hypothetical proteins (Eucgr.H02217, Eucgr.I00888, Eucgr.H00218, Eucgr.H01945, Eucgr.I01788, Eucgr.C02248, Eucgr.J02099) and transcripts with no direct function in secondary development like SRC2, protein kinase, SnRK3.16, exotosin, XIH (Myosin family protein) and O-fucosyltransferase had significant expression levels in the developing xylem tissues and were present in the network. The under representation of important secondary cell wall transcripts in the network and presence of uncharacterized transcripts can be attributed to the transcript selection in the present study. Earlier studies targeted the use of genes specifically expressed during secondary cell wall development, while the present study considered transcripts differentially expressed in developing xylem tissues of E. tereticornis with different wood properties.
Population structure is a potential problem when testing for adaptive phenotypic differences and majority of inter and intra species variation in gene expression across phenotypes could be due to population history [68, 69]. The correlation between gene expression variation and genetic distance in populations is widely documented and the predominant factor determining gene expression is genetic drift [69–71]. Hence, gene expression studies on divergent populations with unknown genetic structures can generate false correlations. The potential limitation of the present study is the confounding effect of genetic structure of the selected genotypes on the transcript expression patterns. Further, significant unrelated genetic variation among the genotypes along with provenance and trait effect can have profound effect on transcript expression. The small sample size (two per phenotypic group) could not differentiate the effect of genotype, provenance and the trait of interest (wood composition) on the expression patterns, thus limiting the resolution of the co-expression networks generated in the present study. The resolution of the co-expression networks can be significantly improved by increasing the sample size to predict true interactions between transcripts governing wood formation and composition.
References
Brooker MIH, Kleing DA (2004) Field guide to Eucalyptus. Melbourne Blooming Books, Northern Australia
Iglesias Trabado G, Wilstermann D (2009) “Eucalyptus universalis. Global cultivated eucalypt forests map 2009”. GIT Forestry Consulting’s Eucalyptologics. http://www.git-forestry.com/. Accessed 3 May 2016
Schrader J, Nilsson J, Mellerowicz E, Berglund A, Nilsson P, Hertzberg M, Sandberg G (2004) A high-resolution transcript profile across the wood-forming meristem of poplar identifies potential regulators of cambial stem cell identity. Plant Cell 16:2278–2292
Andersson-Gunneras S, Mellerowicz EJ, Love J, Segerman B, Ohmiya Y, Coutinho PM, Nilsson P, Henrissat B, Moritz T, Sundberg B (2006) Biosynthesis of cellulose-enriched tension wood in Populus: global analysis of transcripts and metabolites identifies biochemical and developmental regulators in secondary wall biosynthesis. Plant J 45:144–165
Liu L, Ramsay T, Zinkgraf M, Sundell D, Street NR, Filkov V, Groover A (2015) A resource for characterizing genome-wide binding and putative target genes of transcription factors expressed during secondary growth and wood formation in Populus. Plant J 82(5):887–898
Yang SH, van Zyl L, No EG, Loopstra CA (2004) Microarray analysis of genes preferentially expressed in differentiating xylem of loblolly pine (Pinus taeda). Plant Sci 166:1185–1195
Paiva JA, Garces M, Alves A, Garnier-Gere P, Rodrigues JC, Lalanne C, Porcon S, Le Provost G, Perez Dda S, Brachm J, Frigerio JM, Claverol S, Barre A, Fevereiro P, Plomion C (2008) Molecular and phenotypic profiling from the base to the crown in maritime pine wood-forming tissue. New Phytol 178:283–301
Pavy N, Boyle B, Nelson C, Paule C, Giguère I, Caron S, Parsons LS, Dallaire N, Bedon F, Bérubé H, Cooke J (2008) Identification of conserved core xylem gene sets: conifer cDNA microarray development, transcript profiling and computational analyses. New Phytol 180:766–786
Kirst M, Basten CJ, Myburg AA, Zeng ZB, Sederoff RR (2005) Genetic architecture of transcript-level variation in differentiating xylem of a Eucalyptus hybrid. Genetics 169:2295–2303
Paux E, Tamasloukht M, Ladouce N, Sivadon P, Pettenati JG (2004) Identification of genes preferentially expressed during wood formation in Eucalyptus. Plant Mol Biol 55:263–280
Novaes E, Drost DR, Farmerie WG, Pappas GJ, Grattapaglia D, Sederoff RR, Kirst M (2008) High-throughput gene and SNP discovery in Eucalyptus grandis, an uncharacterized genome. BMC Genom 9:312
Barros E, Staden CA, Lezar S (2009) A microarray-based method for the parallel analysis of genotypes and expression profiles of wood-forming tissues in Eucalyptus grandis. BMC Biotechnol 9(51):1472–6750
Mizrachi E, Hefer CA, Ranik M, Joubert F, Myburg AA (2010) De novo assembled expressed gene catalog of a fast-growing Eucalyptus tree produced by Illumina mRNA-Seq. BMC Genom 11(1):681
Solomon OL, Berger DK, Myburg AA (2010) Diurnal and circadian patterns of gene expression in the developing xylem of Eucalyptus trees. S. Afr J Bot 76(3):425–439
Paiva JA, Prat E, Vautrin S, Santos MD, San-Clemente H, Brommonschenkel S, Fonseca PG, Grattapaglia D, Song X, Ammiraju JS, Kudrna D, Wing RA, Freitas AT, Berges H, Grima-Pettenati J (2011) Advancing Eucalyptus genomics: identification and sequencing of lignin biosynthesis genes from deep-coverage BAC libraries. BMC Genom 12:137
Hefer CA, Mizrachi E, Myburg AA, Douglas CJ, Mansfield SD (2015) Comparative interrogation of the developing xylem transcriptomes of two wood-forming species: populus trichocarpa and Eucalyptus grandis. New Phytol 206(4):1391–1405
Mewalal R, Mizrachi E, Mansfield SD, Myburg AA (2014) Cell wall-related proteins of unknown function: missing links in plant cell wall development. Plant Cell Physiol 55(6):1031–1043
Benfey PN, Mitchell-Olds T (2008) From genotype to phenotype: systems biology meets natural variation. Science 320(5875):495–497
Feltus FA (2014) Systems genetics: a paradigm to improve discovery of candidate genes and mechanisms underlying complex traits. Plant Sci 223:45–48
Mizrachi E, Myburg AA (2016) Systems genetics of wood formation. Curr Opin Plant Biol 30:94–100
Yang X, Ye CY, Bisaria A, Tuskan GA, Kalluri UC (2011) Identification of candidate genes in Arabidopsis and Populus cell wall biosynthesis using text-mining, co-expression network analysis and comparative genomics. Plant Sci 181(6):675–687
Palle SR, Seeve CM, Eckert AJ, Cumbie WP, Goldfarb B, Loopstra CA (2011) Natural variation in expression of genes involved in xylem development in loblolly pine (Pinus taeda L.). Tree Genet Genomes 7:193–206
Ruprecht C, Persson S (2012) Co-expression of cell-wall related genes: new tools and insights. Front Plant Sci 3:83
Wang S, Yin Y, Ma Q, Tang X, Hao D, Xu Y (2012) Genome-scale identification of cell-wall related genes in Arabidopsis based on co-expression network analysis. BMC Plant Biol 12(1):138
Hirano K, Aya K, Morinaka Y, Nagamatsu S, Sato Y, Antonio BA, Namiki N, Nagamura Y, Matsuoka M (2013) Survey of genes involved in rice secondary cell wall formation through a co-expression network. Plant Cell Physiol 54(11):1803–1821
Cai B, Li CH, Huang J (2014) Systematic identification of cell-wall related genes in Populus based on analysis of functional modules in co-expression network. PLoS One 9(4):e95176
Taylor-Teeples M, Lin L, de Lucas M, Turco G, Toal TW, Gaudinier A, Young NF, Trabucco GM, Veling MT, Lamothe R, Handakumbura PP (2015) An Arabidopsis gene regulatory network for secondary cell wall synthesis. Nature 517(7536):571–575
Lamara M, Raherison E, Lenz P, Beaulieu J, Bousquet J, MacKay J (2016) Genetic architecture of wood properties based on association analysis and co-expression networks in white spruce. New Phytol 210:240–255
Chandran AKN, Jeong HY, Jung KH, Lee C (2016) Development of functional modules based on co-expression patterns for cell-wall biosynthesis related genes in rice. J Plant Biol 59(1):1–15
Davin N, Edger PP, Hefer CA, Mizrachi E, Schuetz M, Smets EE, Myburg AA, Douglas CJ, Schranz ME, Lens F (2016) Functional network analysis of genes differentially expressed during xylogenesis in soc1ful woody Arabidopsis plants. Plant J. doi:10.1111/tpj.13157
Ferreira SS, Hotta CT, de Carli Poelking VG, Leite DCC, Buckeridge MS, Loureiro ME, Barbosa MHP, Carneiro MS, Souza GM (2016) Co-expression network analysis reveals transcription factors associated to cell wall biosynthesis in sugarcane. Plant Mol Biol 91:15–35
Kothiyal V, Jaideep Bhandari S, Ginwal HS, Gupta S (2015) Multi-species NIR calibration for estimating holocellulose in plantation timber. Wood Sci Technol 49:769–793
Salazar MM, Nascimento LC, Camargo ELO, Gonçalves DC, Neto JL, Marques WL, Teixeira PJPL, Mieczkowski P, Mondego JMC, Carazzolle MF, Deckmann AC (2013) Xylem transcription profiles indicate potential metabolic responses for economically relevant characteristics of Eucalyptus species. BMC Genom 14(1):201
Ghosh Dasgupta M, Dharanishanthi V, Agarwal I, Krutovsky KV (2015) Development of genetic markers in Eucalyptus species by target enrichment and exome sequencing. PLoS One 10(1):e0116528
Kanehisa M, Goto S (2000) KEGG: kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res 28(1):27–30
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M (2012) KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40(D1):D109–D114
Thimm O, Bläsing O, Gibon Y, Nagel A, Meyer S, Krüger P, Selbig J, Müller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37(6):914–939
Karpaga Raja Sundari B, Ghosh Dasgupta M (2012) Selection and validation of reference genes for real-time qRT-PCR normalization in different tissues of Eucalyptus tereticornis. Silvae Genetica 61:221–300
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2∆∆ CT Method. Methods 25:402–408
Freeman TC, Goldovsky L, Brosch M, Van Dongen S, Mazière P, Grocock RJ, Freilich S, Thornton J, Enright AJ (2007) Construction, visualisation, and clustering of transcription networks from microarray expression data. PLoS Comput Biol 3(10):e206
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13(11):2498–2504
Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman WH, Pagès F, Trajanoski Z, Galon J (2009) ClueGO: a Cytoscape plug-into decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25(8):1091–1093
Dysvik B, Jonassen I (2001) J-Express: exploring gene expression data using Java. Bioinformatics 17(4):369–370
Plomion C, Leprovost G, Stokes A (2001) Wood formation in trees. Plant Physiol 127:1513–1523
Groover AT, Nieminen K, Helariutta Y, Mansfield SD (2010) Wood formation in Populus. In: Jansson S, Bhalerao RP, Groover AT (eds) Genetics and Genomics of Populus. Springer, New York, pp 201–224
Prassinos C, Ko JH, Yang J, Han KH (2005) Transcriptome profiling of vertical stem segments provides insights into the genetic regulation of secondary growth in hybrid aspen trees. Plant Cell Physiol 46:1213–1225
Raherison E, Rigault P, Caron S, Poulin PL, Boyle B, Verta JP, MacKay J (2012) Transcriptome profiling in conifers and the Picea GenExpress database show patterns of diversification within gene families and interspecific conservation in vascular gene expression. BMC Genom 13(1):434
Carvalho A, Paiva J, Louzada J, Lima-Brito J (2013) The transcriptomics of secondary growth and wood formation in Conifers. Mol Biol Int. doi:10.1155/2013/974324
Shinya T, Iwata E, Nakahama K, Fukuda Y, Hayashi K, Nanto K, Rosa AC, Kawaoka A (2016) Transcriptional profiles of hybrid Eucalyptus genotypes with contrasting lignin content reveal that monolignol biosynthesis-related genes regulate wood composition. Front Plant Sci 7:443
Carocha V, Soler M, Hefer C, Cassan-Wang H, Fevereiro P, Myburg AA, Grima-Pettenati J (2015) Genome-wide analysis of the lignin toolbox of Eucalyptus grandis. New Phytol 206(4):1297–1313
Myburg AA, Grattapaglia D, Tuskan GA, Hellsten U, Hayes RD, Grimwood J, Jenkins J et al (2014) The genome of Eucalyptus grandis. Nature 510:356–362
Soler M, Camargo ELO, Carocha V, Cassan-Wang H, San Clemente H, Savelli B, Hefer CA, Paiva JAP, Myburg AA, Grima-Pettenati J (2015) The Eucalyptus grandis R2R3-MYB transcription factor family: evidence for woody growth-related evolution and function. New Phytol 206(4):1364–1377
Hussey SG, Saïdi MN, Hefer CA, Myburg AA, Grima-Pettenati J (2015) Structural, evolutionary and functional analysis of the NAC domain protein family in Eucalyptus. New Phytol 206(4):1337–1350
Linhart YB, Grant MC (1996) Evolutionary significance of local genetic differentiation in plants. Ann Rev Ecol Syst 27:237–277
Alonso-Blanco C, Mendez-Vigo B, Koornneef M (2005) From phenotypic to molecular polymorphisms involved in naturally occurring variation of plant development. Int J Dev Biol 49:717–732
Yang SH, Loopstra CA (2005) Seasonal variation in gene expression for loblolly pines (Pinus taeda) from different geographical regions. Tree Physiol 25:1063–1073
Thavamanikumar S, McManus LJ, Ades PK, Bossinger G, Stackpole DJ, Kerr R, Hadjigol S, Freeman JS, Vaillancourt RE, Zhu P, Tibbits JF (2014) Association mapping for wood quality and growth traits in Eucalyptus globulus ssp. globulus Labill identifies nine stable marker-trait associations for seven traits. Tree Genet Genomes 10(6):1661–1678
Bhuiyan N (2008) Identification of genes influencing wood fibre properties in Eucalyptus nitens. Thesis, University of Melbourne
Xiang Y, Zhang CQ, Huang K (2012) Predicting glioblastoma prognosis networks using weighted gene co-expression network analysis on TCGA data. BMC Bioinform 13(Suppl 2):S12
Sasidharan R, Mustroph A, Boonman A, Akman M, Ammerlaan AM, Breit T, Schranz ME, Voesenek LACJ, van Tienderen PH (2013) Root transcript profiling of two Rorippa species reveals gene clusters associated with extreme submergence tolerance. Plant Physiol 163(3):1277–1292
Costa MCD, Righetti K, Nijveen H, Yazdanpanah F, Ligterink W, Buitink J, Hilhorst HW (2015) A gene co-expression network predicts functional genes controlling the re-establishment of desiccation tolerance in germinated Arabidopsis thaliana seeds. Planta 242(2):435–449
Camargo ELO, Nascimento LC, Soler M, Salazar MM, Lepikson-Neto J, Marques WL, Alves A, Teixeira PJPL, Mieczkowski P, Carazzolle MF, Martinez Y, Deckmann AC, Rodrigues JC, Grima-Pettenati J, Pereira GAG (2014) Contrasting nitrogen fertilization treatments impact xylem gene expression and secondary cell wall lignification in Eucalyptus. BMC Plant Biol 14:256
Cassan-Wang H, Goue N, Saidi MN, Legay S, Sivadon P, Goffner D, Grima-Pettenati J (2013) Identification of novel transcription factors regulating secondary cell wall formation in Arabidopsis. Front Plant Sci 4:189
Groover AT (2005) What genes make a tree? Trends Plant Sci 10:210–214
Dejardin A, Laurans F, Arnaud D, Breton C, Pilate G, Leplé JC (2010) Wood formation in angiosperms. CR Biol 333(4):325–334
Spicer R, Groover A (2010) Evolution of development of vascular cambia and secondary growth. New Phytol 186(3):577–592
Hansen BO, Vaid N, Musialak-Lange M, Janowski M, Mutwil M (2014) Elucidating gene function and function evolution through comparison of co-expression networks of plants. Front Plant Sci 5(394):10–3389
Stone GN, Nee S, Felsenstein J (2011) Controlling for non-independence in comparative analysis of patterns across populations within species. Philosophical Transactions of the Royal Society of London. Series B Biol Sci 366:1410–1424
Kryvokhyzha D, Holm K, Chen J, Cornille A, Glémin S, Wright S, Lagercrantz U, Lascoux M (2016) The influence of population structure on gene expression and flowering time variation in the ubiquitous weed Capsella bursa-pastoris (Brassicaceae). Mol Ecol. doi:10.1111/mec.13537
Khaitovich P, Pääbo S, Weiss G (2005) Toward a neutral evolutionary model of gene expression. Genetics 170:929–939
Whitehead A, Crawford DL (2006) Variation within and among species in gene expression: raw material for evolution. Mol Ecol 15:1197–1211
Acknowledgments
The authors acknowledge Dr. Vimal Kothiyal, Scientist G, Indian Council of Forestry Research and Education, Dehradun, India for providing the phenotypic data of wood core samples of Eucalyptus tereticornis. The guidance given by Dr. Jennifer Dewoody, National Forest Genetics Laboratory, USDA Forest Service, Placerville, CA; Dr. Sucheta Tripathy, Structural Biology and Bio-informatics Division, Indian Institute of Chemical Biology, Kolkata, India; Dr. Saravanakumar Selvaraj, Centre for Pharmacology and Toxicology, Hannover Medical School, Hannover, Germany and Dr. Gayatri Ramakrishnan, Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India for developing gene regulatory networks is gratefully acknowledged. The funding for the research work was provided to MDG by Department of Biotechnology, Government of India under the research project with Grant Number BT/PR10055/PBD/16/772/2007. The funding support as research fellowship was provided to VD by Department of Biotechnology, Government of India.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dharanishanthi, V., Dasgupta, M.G. Construction of co-expression network based on natural expression variation of xylogenesis-related transcripts in Eucalyptus tereticornis . Mol Biol Rep 43, 1129–1146 (2016). https://doi.org/10.1007/s11033-016-4046-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-016-4046-3