Abstract
The marine clam Lutraria rhynchaena is gaining popularity as an aquaculture species in Asia. Lutraria populations are present in the wild throughout Vietnam and several stocks have been established and translocated for breeding and aquaculture grow-out purposes. In this study, we demonstrate the feasibility of utilising Illumina next-generation sequencing technology to streamline the identification and genotyping of microsatellite loci from this clam species. Based on an initial partial genome scan, 48 microsatellite markers with similar melting temperatures were identified and characterised. The 12 most suitable polymorphic loci were then genotyped using 51 individuals from a population in Quang Ninh Province, North Vietnam. Genetic variation was low (mean number of alleles per locus = 2.6; mean expected heterozygosity = 0.41). Two loci showed significant deviation from Hardy–Weinberg equilibrium (HWE) and the presence of null alleles, but there was no evidence of linkage disequilibrium among loci. Three additional populations were screened (n = 7–36) to test the geographic utility of the 12 loci, which revealed 100 % successful genotyping in two populations from central Vietnam (Nha Trang). However, a second population from north Vietnam (Co To) could not be successfully genotyped and morphological evidence and mitochondrial variation suggests that this population represents a cryptic species of Lutraria. Comparisons of the Qang Ninh and Nha Trang populations, excluding the 2 loci out of HWE, revealed statistically significant allelic variation at 4 loci. We reported the first microsatellite loci set for the marine clam Lutraria rhynchaena and demonstrated its potential in differentiating clam populations. Additionally, a cryptic species population of Lutraria rhynchaena was identified during initial loci development, underscoring the overlooked diversity of marine clam species in Vietnam and the need to genetically characterise population representatives prior to microsatellite development. The rapid identification and validation of microsatellite loci using next-generation sequencing technology warrant its integration into future microsatellite loci development for key aquaculture species in Vietnam and more generally, aquaculture countries in the South East Asia region.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Molluscs are the most important group of animals after fish contributing to world aquaculture products. However, they have seen far less attention in terms of genetic improvement programs, the characterisation and description of genetic variability and genetic-based management programs [1]. Microsatellites have been the molecular markers of choice for estimating genetic variability in natural and domesticated populations of animal and plants [2, 3]. Despite their popularity and utility, a major limitation to the use of microsatellites for non-model species is that, in general, loci have to first be identified and developed specifically for that species, which is a time and resource consuming process [2, 4]. However, next-generation sequencing (NGS) platforms are being increasingly used to support microsatellite marker identification and development [5–7], greatly decreasing the time and cost of this tedious process [8]. In addition, NGS approaches facilitate the identification of loci suitable for multiplexing, further increasing the efficiency of genotyping [2], which is also now being undertaken using NGS platforms [9, 10].
In addition to their growing importance for aquaculture, clams are important components of near-shore soft sediment environments and many are of significant commercial value as fisheries. The family Mactridae contains a diverse group of approximately 350 species of ecologically and commercially important clams with a global distribution [11, 12]. Species live by burrowing into sandy or gravelly substrates usually below the low water mark and there is increasing evidence of cryptic speciation in the group [12, 13]. In many countries, especially in the tropics, knowledge of genetic structure of species is lacking due to limited genetic resources [1].
Species of snout otter clams placed within the genus Lutraria are sought after by commercial fishers and aquaculture methods are increasingly being developed in Asian countries for several species [11, 14]. The species Lutraria rhynchaena is becoming an important aquaculture species in countries such as Vietnam [11]. However, there is a lack of genetic resources available for this species and the genus more generally [12, 15, 16], which constrains the effective development of stocks for aquaculture and the development of management plans for translocation of broodstock and seedstock.
Panels of microsatellite loci have been developed for a number of commercially or ecologically important bivalve species using both traditional approaches [17, 18] and more recently, NGS-based approaches [16, 19].
The objectives of our study were to: (1) sequence and assemble the partial genome of the commercially important snout clam Lutraria rhynchaena, (2) identify and validate microsatellite loci from the assembled partial genome using amplicon-based next generation sequencing, and (3) assess the potential of the identified microsatellite loci for population genetic studies.
Materials and methods
Sampling
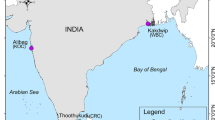
A total of 104 clam samples representing wild and cultured populations from north and central Vietnam were used in this study (Table 1).
Partial genome sequencing
Approximately 1 µg of genomic DNA was extracted from a muscle sample of a clam from Van Don Province (VD) using DNAeasy Blood and Tissue Kit (Qiagen, Hilden, Germany). The purified genomic DNA was quantified with Qubit HS (Invitrogen, USA) and normalized to 2 ng/μL and subsequently processed using Nextera-based library preparation (Illumina, San Diego, CA) following the manufacturer’s instructions. Quantification and size estimation of the library was performed on a Bioanalyzer 2100 High Sensitivity DNA chip (Agilent, Santa Clara, CA). Next, the library was normalized to 2 nM and sequenced on the MiSeq Benchtop Sequencer (2 × 250 bp paired-end reads) (Illumina, USA). The reads were assembled de novo into contigs using IDBA-UD (–mink 31 –maxk 251 setting) [20].
Microsatellite isolation and characterization
The open-source QDD version 3 [21] was used to identify contigs possessing microsatellite motifs as well as to design primer pairs suitable for the amplification of these loci. Primers were subsequently filtered based on suggestions by the authors of the software [21]. A selection of contigs including di-, tri-, and tetra-nucleotide repeats were used for subsequent analysis. A total of 48 loci were initially screened for amplification success and for presence of polymorphism using template DNA from eight clams, representing four sampling location from north and central Vietnam. Primers were pooled for the co-amplification of suitable loci by multiplex PCR using a QIAGEN multiplex kit and an Eppendorf MastercyclerS gradient PCR machine following the protocol described by Blacket et al. [22]. Illumina adapters were attached to the purified amplicons using NEXTflex DNA preparation kit (BiooScientific, Austin, TX).
Preliminary sequencing results identified a subset of 12 suitable polymorphic loci. The multiplex primers were re-designed to contain partial Illumina adapter, allowing for rapid and economical library construction using a 2-step PCR method. Briefly, a multiplex PCR was performed on each clam sample and the amplicons were purified and size-selected using Ampure Bead XP (0.9 × volume ratio). A second PCR was carried out using Illumina Nextera-based barcode primers to generate amplicons containing the complete Illumina adapter and unique barcode. After the second PCR, the amplicons were purified using Ampure Bead XP (0.8 × volume ratio), quantified using KAPA Library Quantification kit (KAPA Biosystems, Cape Town, South Africa), normalized, pooled and sequenced on the MiSeq (2 × 250 bp paired-end run).
Raw sequence quality control and bioinformatics processing
The raw paired-end reads were adapter-trimmed and overlapped using Trimmomatic [23] and PEAR (setting: −q 15, −m 150) [24], respectively. Then, the processed reads were filtered for reads containing both the complete forward and reverse primer sequences using a grep command. Next, the reads were mapped to the assembled contigs using Bowtie2 (with—very-sensitive option) [25]. PCR amplicon lengths were genotyped for variation in repeat motif from alignments using Geneious v.7.0.4 [26] by summarising the read length distributions for each locus in frequency histograms and using established criteria for genotyping microsatellite loci (see pages 599 and 603 in [2]).
The software GeneAIEx (http://biology-assets.anu.edu.au/GenAlEx/Welcome.html) was then used to estimate expected (H E) and observed (H O) heterozygosities and number of alleles (NA), while conformity to Hardy–Weinberg equilibrium (HWE) expectations, inbreeding coefficient (F IS) and linkage disequilibrium estimates between all pairs of loci were examined using the open-source GENEPOP on the web v4 [27]. Bonferroni corrections [28] were used to adjust significance values for multiple comparisons. Lastly, loci were assessed for null alleles and scoring errors using MICRO-CHECKER [29]. Pairwise comparisons of allelic frequencies between populations VDT (north Vietnam) and NTT (central Vietnam) were carried out using the G-test of independence also implemented by GENEPOP [27].
Results and discussion
Next-generation sequencing and de novo genome assembly
A total of 4,214,000 paired-end genomic reads consisting of 622.5 Mb of data were obtained from the library using the MiSeq platform. These reads were submitted to the Sequence Read Archive [SRA: ERR955910]. An assembly of these reads produced 22,193 contigs with a N50 of 661 bp.
Microsatellite isolation and characterisation
Using the assembled contigs, a total of 916 contigs possessing microsatellite motifs were identified by QDD analysis, of which 48 contigs contained priming sites for loci with an estimated melting temperature of approximately 58°. A subset of 12 of these loci were then selected for further genotyping based on the degree of polymorphism and the strength and consistency of amplification using an initial set of eight samples. The details of these loci and accession numbers are provided in Table 2.
The set of loci was found to have low to moderate genetic variation, with an average of 2.6 alleles per locus (range = 2–4 alleles) and heterozygosity estimates ranging between 0.14 and 0.77 (mean = 0.34; Table 2). There was no strong evidence of linkage between pairs of loci with only one pairwise comparisons being significant, and this became non-significant after Bonferroni correction. With the exception of loci mumLr11 and mumLr12, all loci conformed to Hardy–Weinberg expectations. These 2 loci remained significant after Bonferroni adjustment and showed an excess of homozygotes, as indicated by high F IS values (Table 2) and the primers for locus mumLr12 may have been amplifying a second locus in some individuals as three and sometimes four different length variants at high frequencies were apparent. Analysis using MICRO-CHECKER detected the presence of null alleles at these same two loci and also at locus mumLr8, which was marginally non-significant for HW equilibrium without Bonferroni corrections (p = 0.0518). All loci amplified strongly in two of the three additional populations examined (n = 7–36). Only a few individuals and loci were scorable in clams from site CTT, which also possessed divergent mitochondrial COI haplotypes (unpublished data), suggesting the presence of a cryptic species at this location. Pairwise comparisons of allelic frequencies between population VDT (north Vietnam) and NTT (central Vietnam) for the 10 loci giving non-significant HWE results produced four significant results (Table 2), with two comparisons giving P-values <0.001. Thus, these loci have the potential to be used to address questions relating to population differentiation and gene flow in this species.
The allelic diversity and heterozygosity in L. rhynchaena is lower than in a number of other bivalve mollusc species [18, 30]. This may reflect either the small sampling of loci, an inherent characteristic of the species or the population genetic history of the sample that could have suffered a population bottleneck or be the result of human translocation. It is noteworthy that a number of recent studies have noted reduced allelic variation in clam species [31, 32] including one on the surf clam species, Mactra chinensis, from the same family [16]. Only 2 of the 12 loci showed deviation from HWE, which is a lower proportion than in many similar studies of bivalve molluscs [18, 33]. In general, utilizing NGS on any given species enables the identification of hundreds or sometimes thousands (depending on sequencing depth) of candidate microsatellite loci. A high number of candidate loci ensure sufficient loci are retained after strict loci filtration for quality such as pure microsatellite motifs, non-hairpin forming sequences, compatible primer annealing temperature for multiplexing and low or zero hit to transposable elements. Further, the use of NGS platforms for subsequent genotyping not only removes the limitations associated with fragment analysis but also allows for more accurate genotyping, given that amplicon length estimation is based on absolute quantification (base-by-base) instead of relative quantification based on a size standard. Additionally, the ability to differentiate loci bioinformatically allows for the use of loci with overlapping sizes to be amplified in a single PCR reaction, thereby reducing laboratory cost and time [18, 33].
Conclusion
In this study we successfully identified over 900 potential microsatellite loci for L. rhynchaena from contigs assembled from 622.5 Mb of NGS data using the Illumina MiSeq platform. From these contigs, we were able to rapidly identify and characterise 12 polymorphic microsatellite markers suitable for multiplexing with all but two conforming to HWE expectations. Based on pairwise statistical tests, we showed that the microsatellite loci could effectively differentiate clam populations in Vietnam. Thus, we also demonstrated the feasibility of using NGS platforms as a faster, more accurate and potentially cheaper approach for genotyping.
References
Astorga MP (2014) Genetic considerations for mollusk production in aquaculture: current state of knowledge. Front Genet 5:435. doi:10.3389/fgene.2014.00435
Guichoux E, Lagache L, Wagner S, Chaumeil P, Léger P, Lepais O, Lepoittevin C, Malausa T, Revardel E, Salin F (2011) Current trends in microsatellite genotyping. Mol Ecol Resour 11(4):591–611
Sunnucks P (2000) Efficient genetic markers for population biology. Trends Ecol Evol 15(5):199–203. doi:10.1016/S0169-5347(00)01825-5
Castoe TA, Poole AW, de Koning APJ, Jones KL, Tomback DF, Oyler-McCance SJ, Fike JA, Lance SL, Streicher JW, Smith EN, Pollock DD (2012) Rapid microsatellite identification from illumina paired-end genomic sequencing in two birds and a snake. PLoS One 7(2):e30953. doi:10.1371/journal.pone.0030953
Berman M, Austin CM, Miller AD (2014) Characterisation of the complete mitochondrial genome and 13 microsatellite loci through next-generation sequencing for the New Caledonian spider-ant Leptomyrmex pallens. Mol Biol Rep 41(3):1179–1187. doi:10.1007/s11033-013-2657-5
Luo W, Nie Z, Zhan F, Wei J, Wang W, Gao Z (2012) Rapid development of microsatellite markers for the endangered fish Schizothorax biddulphi (Günther) using next generation sequencing and cross-species amplification. Int J Mol Sci 13(11):14946–14955
Peñarrubia L, Sanz N, Pla C, Vidal O, Viñas J (2015) Using massive parallel sequencing for the development, validation, and application of population genetics markers in the invasive bivalve zebra mussel (Dreissena polymorpha). PLoS One 10(3):e0120732. doi:10.1371/journal.pone.0120732
Gardner MG, Fitch AJ, Bertozzi T, Lowe AJ (2011) Rise of the machines—recommendations for ecologists when using next generation sequencing for microsatellite development. Mol Ecol Resour 11(6):1093–1101. doi:10.1111/j.1755-0998.2011.03037.x
Van Neste C, Van Nieuwerburgh F, Van Hoofstat D, Deforce D (2012) Forensic STR analysis using massive parallel sequencing. Forensic Sci Int: Genet 6(6):810–818. doi:10.1016/j.fsigen.2012.03.004
Zavodna M, Bagshaw A, Brauning R, Gemmell NJ (2014) The accuracy, feasibility and challenges of sequencing short tandem repeats using next-generation sequencing platforms. PLoS One 9(12):e113862. doi:10.1371/journal.pone.0113862
Luca M, Nam DX (2011) Hatchery techniques applied for the artificial production of snout otter clam (Lutraria rhynchaena) in small scale farms in Nha Trang City, Vietnam. Advancing Aquaculture Around the World, p 25
Ni L, Li Q, Kong L, Huang S, Li L (2012) DNA barcoding and phylogeny in the family Mactridae (Bivalvia: Heterodonta): evidence for cryptic species. Biochem Syst Ecol 44:164–172. doi:10.1016/j.bse.2012.05.008
Kong L, Li Q (2009) Genetic evidence for the existence of cryptic species in an endangered clam Coelomactra antiquata. Mar Biol 156(7):1507–1515. doi:10.1007/s00227-009-1190-5
Liu H, Zhu JX, Sun HL, Fang JG, Gao RC, Dong SL (2006) The clam, Xishi tongue Coelomactra antiquata (Spengler), a promising new candidate for aquaculture in China. Aquaculture 255(1–4):402–409. doi:10.1016/j.aquaculture.2005.12.027
Gan HM, Tan MH, Thai BT, Austin CM (2016) The complete mitogenome of the marine bivalve Lutraria rhynchaena Jonas 1844 (Heterodonta: Bivalvia: Mactridae). Mitochondrial DNA 27(1):335–336. doi:10.3109/19401736.2014.892104
Li H, Zhang J, Li H, Gao X, He C (2014) Development and characterization of 13 polymorphic microsatellite markers for the Chinese surf clam (Mactra chinensis) through Illumina paired-end sequencing. Conserv Genet Resour 6(4):877–879
Kim E, An H, Kang J, An C, Dong C, Hong Y, Park J (2014) New polymorphic microsatellite markers for the Korean manila clam (Ruditapes philippinarum) and their application to wild populations. Genet Mol Res: GMR 13(4):8163
Nantón A, Arias-Pérez A, Méndez J, Freire R (2014) Characterization of nineteen microsatellite markers and development of multiplex PCRs for the wedge clam Donax trunculus (Mollusca: Bivalvia). Mol Biol Rep 41(8):5351–5357. doi:10.1007/s11033-014-3406-0
Duan C-X, Li D-D, Sun S-L, Wang X-M, Zhu Z-D (2014) Rapid development of microsatellite markers for Calloso bruchus chinensis using Illumina paired-end sequencing. PLoS One 9(5):e95458. doi:10.1371/journal.pone.0095458
Peng Y, Leung HCM, Yiu SM, Chin FYL (2012) IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28(11):1420–1428. doi:10.1093/bioinformatics/bts174
Meglécz E, Costedoat C, Dubut V, Gilles A, Malausa T, Pech N, Martin J-F (2010) QDD: a user-friendly program to select microsatellite markers and design primers from large sequencing projects. Bioinformatics 26(3):403–404. doi:10.1093/bioinformatics/btp670
Blacket MJ, Robin C, Good RT, Lee SF, Miller AD (2012) Universal primers for fluorescent labelling of PCR fragments—an efficient and cost-effective approach to genotyping by fluorescence. Mol Ecol Resour 12(3):456–463. doi:10.1111/j.1755-0998.2011.03104.x
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. doi:10.1093/bioinformatics/btu170
Zhang J, Kobert K, Flouri T, Stamatakis A (2014) PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30(5):614–620. doi:10.1093/bioinformatics/btt593
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359. doi:10.1038/nmeth.1923
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28(12):1647–1649. doi:10.1093/bioinformatics/bts199
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86(3):248–249
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43(1):223–225. doi:10.2307/2409177
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) micro-checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4(3):535–538. doi:10.1111/j.1471-8286.2004.00684.x
Cavaleiro NP, Solé-Cava AM, Lazoski C, Cunha HA (2013) Polymorphic microsatellite loci for two Atlantic oyster species: Crassostrea rhizophorae and C. gasar. Mol Biol Rep 40(12):7039–7043. doi:10.1007/s11033-013-2823-9
Chen X, Li Z, Chen L, Cao Y, Li Q (2012) Isolation and characterization of new microsatellite markers in the pen shell Atrina pectinata (Pinnidae). Genet Mol Res 11(3):2884–2887
Kang J-H, Kim B-H, Park J-Y, Lee J-M, Jeong J-E, Lee J-S, Ko H-S, Lee Y-S (2012) Novel microsatellite markers of Meretrix petechialis and cross-species amplification in related taxa (Bivalvia: Veneroida). Int J Mol Sci 13(12):15942–15954
Chacón G, Arias-Pérez A, Méndez J, Insua A, Freire R (2012) Development and multiplex PCR amplification of microsatellite markers in the commercial clam Venerupis rhomboides (Mollusca: Bivalvia). Mol Biol Rep 40(2):1625–1630. doi:10.1007/s11033-012-2211-x
Acknowledgments
Funding for this project was provided by the Government of Vietnam through its Aquaculture Biotechnology programs (606a/HD-KHCN-CNSH).The authors also wish to acknowledge the support of Monash University Malaysia through its Tropical Medicine and Biology Multidisciplinary Platform. We would like to thank Fisheries College for support in this project and Mr. Nguyen Thanh Nhon and Mr. Nguyen Van Ha who helped to collect the samples.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Thai, B.T., Tan, M.H., Lee, Y.P. et al. Characterisation of 12 microsatellite loci in the Vietnamese commercial clam Lutraria rhynchaena Jonas 1844 (Heterodonta: Bivalvia: Mactridae) through next-generation sequencing. Mol Biol Rep 43, 391–396 (2016). https://doi.org/10.1007/s11033-016-3966-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-016-3966-2