Abstract
Natural root-knot nematode resistance genes are unique resources to control this major pest in pepper (Capsicum annuum). Although four genes (Me1, Me3, Me7 and N) conferring broad-spectrum resistance were mapped to a cluster in a 28-cm interval on chromosome P9, limited markers targeting this region were available. In the present study, the Me-gene cluster was structurally annotated for resistance genes to develop markers targeting the N gene. As a result, the Me-gene cluster (4.07 Mb in size) was found to contain three resistance gene hotspots. In addition, a SSR maker tightly linked to the N gene (0.8 cM away) was developed for marker-assisted selection in pepper.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Key message
This study revealed R gene hotspots in the Me-gene cluster on pepper chromosome P9 and identified a marker tightly linked to the N gene for root-knot nematode resistance.
Introduction
Root-knot nematodes (Meloidogyne spp.)(RKN) are an important biotic stress in pepper. RKN infect roots and use nutrients and water from the host plant. Thus, RKN infection causes decreased root and shoot growth which results in reduced pepper yield (Trudgill and Phillips 1997; Williamson and Kumar 2006). Although nematicides have been used to control the pest, these chemicals have negative effects on the environment and human health (Fazari et al. 2012; Van Gundy and McKenry 2012). Thus, development of pepper cultivars with natural resistance genes against RKN is the most effective and environmentally friendly way to control them.
There are ten genes for dominant resistance to RKN in pepper (Me1 to Me7, Mech1, Mech2 and N) (Djian-Caporalino et al. 2007; Fazari et al. 2012). These genes confer resistance based on gene-for-gene interaction mechanisms (Fazari et al. 2012). Four of the ten genes (Me1, Me3, Me7 and N) confer resistance to a wide range of RKN species (Djian-Caporalino et al. 1999). Thus, these genes are important resources for the development of resistant pepper cultivars. To date, none of these resistance genes have been isolated; however, they were mapped to a 28 cM cluster on pepper chromosome 9 (Djian-Caporalino et al. 2007; Fazari et al. 2012; Uncu et al. 2015). Comparative mapping studies showed that this cluster is syntenic to potato and tomato chromosome 12 which contains resistance genes such as Me(s), Bs2, Mi-3/Mi-5, Lv and Gpa2/Rx (Grube et al. 2000; Mazourek et al. 2009). Although the cluster was partially annotated in a study that identified the Bs2 gene conferring resistance to bacterial spot (Xanthomonas campestris pv. vesicatoria) (Tai et al. 1999), it has not been annotated for resistance genes. In addition, only a limited number of molecular markers including one RFLP (restriction fragment length polymorphism, CT135) and 12 SCAR (sequence characterized amplified region) markers (Djian-Caporalino et al. 2007; Fazari et al. 2012) targeting the cluster are available.
In the present study, the Me-gene cluster was structurally annotated for resistance genes using the genome and coding sequence (CDS) of pepper reported by Kim et al. (2014). In addition, simple sequence repeat (SSR) and cleaved amplified polymorphic sequence (CAPS) markers linked to the Me-gene cluster were identified. The second goal of the study was the development of a molecular marker tightly linked to the N gene for marker-assisted selection (MAS). Although the N gene was previously mapped in the Me-gene cluster, the distance between the gene and the closest linked marker was 2.3 cM (Wang et al. 2009; Fazari et al. 2012). Markers within 1 cM of the N gene are essential for efficient introgression of the gene to pepper cultivars via MAS. To identify such a marker, SSR and CAPS markers targeting the Me-gene cluster were tested in an F2 mapping population derived from a cross between RKN resistant Capsicum annuum cv. Carolina Wonder and RKN susceptible C. annuum cv. AZN-1. The segregation of the N gene in the F2 population was analyzed by pathogen tests performed with M. incognita race 2. As a result, a SSR marker tightly linked (within 1 cM) to the N gene was developed. Test of the SSR marker in an F4 population showed that the co-dominant marker can be used in MAS for RKN resistance breeding in pepper.
Materials and methods
Identification and physical mapping of NBS-coding resistance genes to Me-gene cluster
The location of the Me-gene cluster was identified by aligning flanking markers to the genome (Pepper.v.1.55). Flanking markers (At4g11120 and Bs2 gene) were selected from the linkage map of Djian-Caporalino et al. (2007). A total of 34,899 CDS (Pepper.v.1.55.CDS.fa) generated by Kim et al. (2014) were downloaded from the pepper genome platform (PGP) database (http://peppergenome.snu.ac.kr/download.php). All the CDS were functionally annotated using Blast2go software (Conesa et al. 2005). CDS with disease-resistance function were extracted for physical mapping to Me-gene cluster.
One of the aims of the study was to identify the genomic positions of the resistance genes in the Me-gene cluster. To achieve this aim, 436 CDS with disease-resistance function were aligned to the Me-gene cluster using spliced alignment programs such as Spidey (default parameters and plant gene model were used) (http://www.ncbi.nlm.nih.gov/spidey/) and NCBI Genome Workbench (default parameters were used) (http://www.ncbi.nlm.nih.gov/tools/gbench/). In addition to CDS, two cloned RKN resistance genes (CaRKNR and CaMi) were mapped using Spidey (Chen et al. 2007; Mao et al. 2015). Also marker C2At2g06530 which is linked to the Me1 gene was aligned to the pepper genome using the BLAST tool of the Pepper Genome Platform (PGP) database (http://peppergenome.snu.ac.kr/blast.php) (Uncu et al. 2015).
Plant material
For genetic linkage mapping, an F2 population (256 individuals) derived from the cross between RKN resistant C. annuum cv. Carolina Wonder and RKN susceptible C. annuum cv. AZN-1 was developed. The marker linked to the N gene was tested in an F4 population (40 individuals) derived from the same parents by selfing 40 F2 individuals.
RKN culture and resistance assay
For RKN culture, eggs of the M. incognita race 2 isolate identified by Devran and Sogut (2009, 2011) were propagated using tomato cv. Tueza (Multi Tohum, Antalya, Turkey). Egg masses were collected from infected roots at the true leaf stage using a small needle and hatched at room temperature (25 °C). Juveniles were used for inoculation of the tomato plants. The plants were grown in a growth chamber at 25 °C, and egg masses were harvested 8 weeks after the inoculation.
Pepper parents, F1 hybrids and F2 and F4 populations were tested for RKN resistance. For the assay, second stage juveniles (J2) harvested from tomato roots were counted under a microscope. The pepper plants were inoculated with a suspension of 1000 J2s at the fourth true leaf stage. During the assay, plants were grown in plastic pots containing 250 ml sandy soil under controlled conditions at day/night temperature of 25 °C, 65 % relative humidity and 14/10 h (light/dark) photoperiod. RKN resistant and susceptible plants were identified 8 weeks after inoculation. For RKN resistance evaluation, the plants were uprooted and rinsed under water. Egg masses were evaluated using a scale from 0 to 5 according to the publication of Hadisoeganda and Sasser (1982) as follows: 0 = no egg masses, 1 = 1–2 egg masses, 2 = 3–10 egg masses, 3 = 11–30 egg masses, 4 = 31–100 egg masses and 5 = more than 100 egg masses per root mass. Plants with scores of 0–2 were considered to be resistant, while the rest were considered to be susceptible.
Me-gene cluster targeted molecular marker development and linkage mapping of N gene
Physical mapping of CDS revealed hotspots for resistance in the Me-gene cluster. For N gene mapping, CAPS and SSR markers were developed from the hotspots. The study focused on one hotspot (called locus C) for development of SSR markers because this region contains an AFLP marker (N-SCAR-315) reported to be linked to the N gene (Wang et al. 2009). This hotspot was screened for SSRs using Batchprimer3 software (http://probes.pw.usda.gov/batchprimer3/) (You et al. 2008). The minimum number of repeats needed to identify perfect SSRs was: six for dinucleotide, four for trinucleotide and three for tetranucleotide, pentanucleotide and hexanucleotide repeats. Default settings were used for primer design with some adjustments: product size 100–300 bp, optimum 200 bp; primer size 18–23 nt, optimum 21 nt; primer Tm 50–60 °C, optimum 55 °C; primer GC content 40–50 %, optimum 45 %.
A total of 13 sequences (500 bp in size) from all hotspots were used for CAPS marker development with Batchprimer3 software. The default settings of the program were used with some adjustments: product size 400–600 bp, optimum 500 bp; primer size 18–23 nt, optimum 21 nt; primer Tm 50–60 °C, optimum 55 °C; primer GC content 40–50 %, optimum 45 %. Amplified PCR products of CAPS markers were digested with the restriction enzymes listed in Table S1.
Amplification of the SSR and CAPS primers was carried out in 25 μl reaction mixtures containing 1 X PCR buffer, 3 mM MgCl2, 0.125 mM deoxyribonucleotide triphosphates (dNTPs), 1 U Taq polymerase, 2 pmol forward and reverse primers and 80 ng template DNA. The PCR amplification protocol for primers consisted of one cycle of 5 min at 94 °C, followed by 35 cycles of 45 s at 94 °C, 1 min at 55 °C (annealing) and 45 s at 72 °C, with a final extension step of 5 min at 72 °C. PCR products of the primers were visualized by 3 % agarose gel electrophoresis.
In addition to markers developed in the present study, the Me-specific markers reported by Fazari et al. (2012) were tested in the parents according to the amplification protocols in the respective publication (Table S2). The genetic linkage map was constructed using JoinMap v 3.0 software (Ooijen and Voorrips 2001).
Results
Identification and physical mapping of NBS-coding resistance genes to Me-gene cluster
The flanking markers of the cluster (At4g11120 and the Bs2 gene) were blasted against the pepper genome. The results revealed that At4g11120 and Bs2 were positioned at 249.13 and 252.07 Mb, respectively, and that the cluster was located in an interval of 2.94 Mb (1.6 % of the whole chromosome). For further analysis, the cluster size was enlarged to 4.07 Mb (from position 248.00 to 252.07) to avoid possible mapping errors.
A total of 436 resistance CDS selected by functional annotation analysis were mapped to the Me-gene cluster. As a result, 50 CDS with more than three exons were mapped (Table S3) and analyzed in the splice alignment module of a genome browser (NCBI Genome Workbench) to determine orientation. Mapping showed that the cluster contained three loci (loci A, B and C) that were hotspots for R genes. Six CDS were found at locus B, two were found at locus C, and 42 were positioned at both loci A and C (Fig. 1). The A, B and C loci were 1.46 Mb, 265.87 and 390.35 kb in length, respectively.
Two cloned (CaRKNR and CaMi) and one mapped (Me1) RKN resistance genes were aligned to the genome to reveal whether these genes are located in the cluster. As a result, the Me1 gene was mapped to locus B at position 250.34 Mb. The cloned genes (CaRKNR and CaMi) were mapped to chromosome P6 at positions of 0.8 and 4.68 Mb with 5 and 32 exons, respectively.
RKN resistance assay
The parents, F1 hybrids and F2 population were tested for RKN (M. incognita race 2) resistance. The C. annuum cv. Carolina Wonder parent which carried the N gene was found to be resistant due to the absence of gall formation. The cv. AZN-1 parent was susceptible as expected with a mean of 258 ± 25.6 galls. In the F2 population, 186 plants (73.2 %) were resistant and the rest (26.8 %) were susceptible. These result indicated that N gene-based resistance was inherited dominantly with a segregation ratio of 3:1 (Chi-square goodness of fit test, p = 0.43).
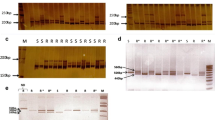
Me-gene cluster targeted molecular marker development and linkage mapping of N gene
The second aim of the study was development of markers tightly linked to the N gene. To achieve this, SSR and CAPS markers were developed from the three hotspot loci (A, B and C) identified in the Me-gene cluster. For the SSR markers, the study focused on only locus C because this region was found by Wang et al. (2009) to contain an AFLP marker (N-SCAR-315) linked to the N gene. As a result, 57 SSR primers were designed and used in linkage map construction (Table S4). While all except six markers showed clear PCR amplification, 12 primers generated more than one PCR fragment. Five markers were polymorphic (Table S4). Although four of them could not be mapped because they targeted multiple loci, CASSR37 located in locus C at a position of 251.09 Mb had clear PCR amplification and a Mendelian segregation pattern (Chi-square goodness of fit test, p = 0.9 (Fig. S1). Marker CASSR37 was found to be tightly linked to N at a distance of 0.8 cM (Fig. 2). An F4 population derived from the same parents as the mapping population was also tested for RKN resistance for verification of the usefulness of marker CASSR37 for MAS. As a result, 15 individuals were resistant and 25 individuals were susceptible in the F4 population. The marker was successful in distinguishing all resistant and susceptible plants. In addition to the SSR marker, a CAPS marker from locus B, CACAPS2 digested with DraI enzyme, was mapped 6.3 cM from N (Fig. 2). Seven SCAR (sequence characterized amplified regions) markers developed by Fazari et al. (2012) were also tested in the parents (Table S2). All the markers except SCARPM6a and SCARN amplified successfully. SCARPM6b and SSCPPM5 were found to be polymorphic and were tested in the F2 population. The two markers co-segregated at a distance of 3.2 cM from the N gene.
Discussion
Identification and physical mapping of NBS-coding resistance genes to Me-gene cluster
Functional and structural annotation of the CDS was previously reported by Kim et al. (2014). In the present study, CDS were re-annotated because the annotation results were not available in the PGP database. As expected, the numbers of resistance CDS identified by the present study and Kim et al. (2014) were similar.
This study identified the genomic position of the Me-gene cluster by physical mapping for the first time. Although the cluster was previously described as a 28 cM interval (13.5 % of the whole chromosome), this study revealed that the size was 4.07 Mb (1.6 % of the whole chromosome). Thus, this study decreased the cluster interval by 8.4-fold.
Mapping of 11.5 % of the resistance CDS to a small region (4.07 Mb) confirmed that the Me-gene cluster is a hotspot for resistance genes. Half of the CDS (54 % of the total CDS mapped to the cluster) located in the cluster were not assigned to any chromosome by Kim et al. (2014). Prediction of the pathogen targets of the resistance CDS was problematic because resistance genes for virus, fungus and nematode often belong to the same gene family (NBS-LLR gene family) (Jupe et al. 2012). The phylogenetic relationships of resistance CDS and the CaMi gene could not be identified because the high diversity of translated CDS prevented successful multiple sequence alignment. The genomic positions of the tomato RKN resistance gene Mi-1 homologs (CaMi and CaRKNR genes) in pepper showed that Mi-1-based homology evolved on P6, not on P9 as expected because chromosome P6 is syntenic to tomato chromosome T6 which contains the Mi-1 gene (Livingstone et al. 1999; Grube et al. 2000).
Our results showed that resistance CDS do not have uniform distribution. Thus, CDS mapped to certain loci (loci A, B and C) demonstrating that the Me-gene cluster has three intervals enriched for resistance genes. Physical mapping of CDS was problematic because most of the CDS (84 %) were positioned at both loci A and C. This may be due to evolutionary mechanisms such as recombination of the paralogous genes within the Me-gene cluster which caused in vivo DNA shuffling and duplications. Such rearrangements cause the dispersal of resistance genes (Song et al. 2002; Mazourek et al. 2009). Although it is known that R gene dispersal is associated with chromosome breakpoints, high-resolution comparative mapping studies between related genera need to be performed to estimate its effect on Me- gene cluster evolution (Ameline-Torregrosa et al. 2008; Mazourek et al. 2009). Precise location of the 2.12-Mb hotspots will be valuable information to develop markers for fine mapping resistance genes in the cluster.
This study identified the genomic positions of the two cloned (CaRKNR and CaMi) and two mapped (Me1, N) RKN resistance genes. To date, only two of the ten RKN resistance genes on chromosome P9, Me1 and N, have been mapped because linked markers have not been identified for the rest of the genes (present work, Uncu et al. 2015). Mapping of Me-1 and N to locus B and C, respectively, is consistent with resistance gene hotspots identified by CDS mapping. Other resistance genes (Me3 and Me7) might be located in one of the hotspot loci; however, linked markers needed to be developed to test this hypothesis. The genomic position of the CaRKNR gene identified in the present study was consistent with that reported by Mao et al. (2015). Mapping of the cloned genes CaMi and CaRKNR showed that chromosome P6 might also be rich for resistance genes similar to P9 (Mao et al. 2015).
RKN culture and resistance assay
Our work confirmed dominant inheritance of the N gene as reported by Fazari et al. (2012) and Wang et al. (2009). The mapping population of the present study contained more F2 individuals (256) than Fazari et al. (2012) (132 individuals) which increased the accuracy of linkage mapping.
Me-gene cluster targeted molecular marker development and linkage mapping of N gene
The subset of Me-gene cluster-specific SSR markers developed in the present study are valuable genetic tools for mapping of other resistance genes (Me7 and Me3). Parental polymorphism of the markers was low due to the self-pollinated, intraspecific nature of the population. The map positions of the SCAR markers (SCARPM6b and SSCPPM5) of the present study were consistent with previous results (Fazari et al. 2012). However, a new SSR marker, CASSR37, was found to be more tightly linked to the N gene. Test of CASSR37 in an F4 population demonstrated the high efficiency of this marker in MAS. Thus, this co-dominant and breeder-friendly marker can be used in breeding programs to develop pepper cultivars resistant to RKN.
In conclusion, this study revealed resistance gene hotspots in the Me-gene cluster on pepper P9 based on physical mapping. The work narrowed the interval of the cluster to 4.07 Mb in total. Within this cluster, a 2.12-Mb region was rich in potential resistance genes. Based on this research, a marker tightly linked to the N gene was developed for MAS in pepper breeding.
References
Ameline-Torregrosa C, Wang BB, O’Bleness MS, Deshpande S, Zhu H, Roe B, Young ND, Cannon SB (2008) Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula. Plant Physiol 146:5–21. doi:10.1104/pp.107.104588
Chen R, Li H, Zhang L, Zhang J, Xiao J, Ye Z (2007) CaMi, a root-knot nematode resistance gene from hot pepper (Capsium annuum L.) confers nematode resistance in tomato. Plant Cell Rep 26(7):895–905. doi:10.1007/s00299-007-0304-0
Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676. doi:10.1093/bioinformatics/bti610
Devran Z, Sogut MA (2009) Distribution and identification of root-knot nematodes from Turkey. J Nematol 41:128–133
Devran Z, Sogut MA (2011) Characterizing races of Meloidogyne incognita, M. javanica and M. arenaria in the West Mediterranean region of Turkey. Crop Prot 30:451–455. doi:10.1016/j.cropro.2010.12.008
Djian-Caporalino C, Pijarowski L, Januel A, Lefebvre V, Daubèze A, Palloix A, Dalmasso A, Abad P (1999) Spectrum of resistance to root-knot nematodes and inheritance of heat-stable resistance in pepper (Capsicum annuum L.). Theor Appl Genet 99:496–502. doi:10.1007/s001220051262
Djian-Caporalino C, Fazari A, Arguel MJ, Vernie T, VandeCasteele C, Faure I, Brunoud G, Pijarowski L, Palloix A, Lefebvre V, Abad P (2007) Root-knot nematode (Meloidogyne spp.) Me resistance genes in pepper (Capsicum annuum L.) are clustered on the P9 chromosome. Theor Appl Genet 114(3):473–486. doi:10.1007/s00122-006-0447-3
Fazari A, Palloix A, Wang L, Yan Hua M, Sage-Palloix AM, Zhang BX, Djian-Caporalino C (2012) The root-knot nematode resistance N-gene co-localizes in the Me-genes cluster on the pepper (Capsicum annuum L.) P9 chromosome. Plant Breed 131(5):665–673. doi:10.1111/j.1439-0523.2012.01994.x
Grube RC, Radwanski ER, Jahn M (2000) Comparative genetics of disease resistance within the Solanaceae. Genetics 155:873–887
Hadisoeganda WW, Sasser JN (1982) Resistance of tomato, bean, southern pea, and garden pea cultivars to root-knot nematodes based on host suitability. Plant Dis 66:145–150. doi:10.1094/PD-66-145
Jupe F, Pritchard L, Etherington GJ, Mackenzie K, Cock PJ, Wright F, Sharma SK, Bolser D, Bryan GJ, Jones JD, Hein I (2012) Identification and localisation of the NB-LRR gene family within the potato genome. BMC Genomics 13:75. doi:10.1186/1471-2164-13-75
Kim S, Park M, Yeom SI, Kim YM, Lee JM et al (2014) Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nat Genet 46(3):270–278. doi:10.1038/ng.2877
Livingstone KD, Lackney VK, Blauth JR, van Wijk R, Jahn MK (1999) Genome mapping in Capsicum and the evolution of genome structure in the Solanaceae. Genetics 152:1183–1202
Mao Z, Zhu P, Liu F, Huang Y, Ling J, Chen G, Yang Y, Feng D, Xie B (2015) Cloning and functional analyses of pepper CaRKNR involved in Meloidogyne incognita resistance. Euphytica 205(3):903–913. doi:10.1007/s10681-015-1438-8
Mazourek M, Cirulli ET, Collier SM, Landry LG, Kang B-C, Quirin EA, Bradeen JM, Moffett P, Jahn MM (2009) The fractionated orthology of Bs2 and Rx/Gpa2 supports shared synteny of disease resistance in the Solanaceae. Genetics 182:1351–1364. doi:10.1534/genetics.109.101022
Ooijen JW, Voorrips RE (2001) JoinMap Version 3.0: software for the calculation of genetic linkage maps. CPRO-DLO, Wageningen
Song R, Llaca V, Messing J (2002) Mosaic organization of orthologous sequences in grass genomes. Genome Res 12:1549–1555. doi:10.1101/gr.268302
Tai TH, Dahlbeck D, Clark ET, Gajiwala P, Pasion R, Whalen MC, Stall RE, Staskawicz BJ (1999) Expression of the Bs2 pepper gene confers resistance to bacterial spot disease in tomato. Proc Natl Acad Sci 96(24):14153–14158
Trudgill DL, Phillips MS (1997) Nematode population dynamics, threshold levels and estimation of crop losses. In: Maqbool MA (ed) Plant nematode problems and their control in the near east region (FAO Plant Production and Protection Paper, No. 144), Rome
Uncu AT, Celik I, Devran Z, Ozkaynak E, Frary A, Frary A, Doganlar S (2015) Development of a SNP-based CAPS assay for the Me1 gene conferring resistance to root knot nematode in pepper. Euphytica 206(2):393–399. doi:10.1007/s10681-015-1489-x
Van Gundy S, McKenry MV (2012) Action of nematicides. In: Horsfall JG (ed) Plant disease: an advanced treatise, 1st edn. Elsevier, pp 263–283
Wang LH, Gu XH, Hua MY, Mao SL, Zhang ZH, Peng DL, Yun XF, Zhang BX (2009) A SCAR marker linked to the N gene for resistance to root knot nematodes (Meloidogyne spp.) in pepper (Capsicum annuum L.). Sci Hortic 122(2):318–322. doi:10.1016/j.scienta.2009.04.011
Williamson VM, Kumar A (2006) Nematode resistance in plants: the battle underground. Trends Genet 22(7):396–403. doi:10.1016/j.tig.2006.05.003
You FMN, Huo N, Gu YQ, Luo MC, Ma Y, Hane D, Lazo GR, Dvorak J, Anderson OD (2008) BatchPrimer3: a high throughput web application for PCR and sequencing primer design. BMC Bioinform 9:253. doi:10.1186/1471-2105-9-253
Acknowledgments
This study was supported by grant 110O675 from the Scientific and Technological Research Council of Turkey (TUBITAK).
Authors contribution
IC analyzed all data, performed all bioinformatics analysis and laboratory experiments and drafted manuscript; MAS performed all nematode tests; EO developed mapping populations; SD designed the study; AF designed the study, received funding and revised manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Celik, I., Sogut, M.A., Ozkaynak, E. et al. Physical mapping of NBS-coding resistance genes to the Me-gene cluster on chromosome P9 reveals markers tightly linked to the N gene for root-knot nematode resistance in pepper. Mol Breeding 36, 137 (2016). https://doi.org/10.1007/s11032-016-0560-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-016-0560-1