Abstract
Recent DNA sequencing projects and the establishment of high-throughput assays have provided an abundance of sequence information and data on nucleotide polymorphisms in rice. Based on previously identified single-nucleotide polymorphisms (SNPs) and insertions/deletions, we employed denaturing high-performance liquid chromatography (DHPLC) to genotype rice varieties using a chromatographic pattern-based strategy. In this study, 12 amplicons harboring multiple and informative SNPs were screened. PCR products of the 12 amplicons from 47 rice varieties were analyzed by DHPLC and DNA sequencing. Each homozygous sample with a single peak pattern in the initial DHPLC analysis was mixed with zhenshan97 for a second DHPLC analysis. The 12 amplicons were found to be polymorphic across the hybrids, and mixed homozygous samples with 43 distinct DHPLC elution profiles detected. Sequence analysis confirmed that the distinct DHPLC patterns corresponded to different DNA sequences. A set of distinct characteristic profiles in six amplicons differentiated between all of the hybrids, inbred lines, and restorer lines and produced unique a fingerprint for these lines. In addition, we found that the DHPLC pattern of the hybrid was in accordance with the results obtained by DHPLC analysis of a mixed sample of the two parents. These results demonstrate that DHPLC can be efficiently applied for the rapid and automated identification of diverse rice varieties and could possibly be utilized for seed genetic purity testing on a high-throughput scale.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa), which has two cultivated subspecies, indica and japonica, is one of the most important food crops throughout the world and is a staple for more than half the world’s population (Ohtsubo and Nakamura 2007). It is estimated that there are more than 100,000 rice varieties worldwide. Distinct rice genotypes used for cultivation and the quality of the seeds both have a strong effect on productivity. Elite cultivars are discriminated by commercial brands, and varietal adulteration with the intent to obtain premium-brand quality remains a serious problem at present. The owner of a cultivar requires the legal protection against adulteration as well as exclusive sales of a protected cultivar, and the developer of a variety needs to protect it from exploitation by others. Given the necessity of protecting cultivars and preventing unauthorized commercial use, the development of a stable and reliable method for cultivar identification is of great significance. However, because new cultivars normally arise from hybridizations between members of an elite group of genetically similar parents, the amount of genetic variability among newly developed cultivars is likely to become even smaller (Rahman et al. 2009) making it more difficult to distinguish the cultivars from one another (Zhu et al. 2012).
The success of hybrid rice production depends on the supply of genetically homogenous high-quality seeds to farmers. Because there is always a chance of contamination in a hybrid seed production plot due to sterile line selfing affected by low temperature, cross-pollination, and/or mechanical mixtures during subsequent handling of the harvested material, the assessment of seed purity is very important for quality control in hybrid seed production. It has been estimated that 1 % impurity in the hybrid seed decreases the potential yield of a hybrid by approximately 100 kg ha−1 (Mao et al. 1996).
Molecular markers based on the indirect detection of sequence-level polymorphism have proven to be powerful tools for the genetic discrimination among species and seed purity assessment. Several molecular markers including restriction fragment length polymorphisms (RFLPs) (Wang and Tanksley 1989; Zhang et al. 1992), random amplification of polymorphic DNA (RAPDs) (Saker et al. 2005; Yong et al. 1999), simple sequence repeats (SSRs) (Garris et al. 2005; Saker et al. 2005; Ni et al. 2002), inter-simple sequence repeats (ISSRs) (Blair et al. 1999), and amplified fragment length polymorphisms (AFLPs) (Saker et al. 2005) are presently available to identify variation at molecular level. However, little variation can be detected at the DNA level using techniques such as RAPD, AFLP, and RFLP.
Among the various molecular markers that are used, single-nucleotide polymorphisms (SNP) and insertions/deletions (INDELs) are the most abundant types of polymorphism, which can theoretically be found within every genomic sequence, and the polymorphism is strongly preserved in alternate generations (Giordano et al. 1999; Rafalski 2002a, b). SNPs and INDELs have been used as genetic markers for many genetic applications such as cultivar identification, the construction of genetic maps, the assessment of genetic diversity, the detection of genotype/phenotype associations, and marker-assisted breeding (Diego et al. 2007; Flint-Garcia et al. 2005; José et al. 2011; Simko et al. 2004; Szalma et al. 2005). In recent years, numerous sequencing projects in rice have generated an abundance of sequence information including SNPs and INDELs (Feltus et al. 2004; Huang et al. 2010; McCouch et al. 2010; McNally et al. 2009; Shen et al. 2004). Among these two basic types, SNP markers are very frequent in genomes, and recent studies have identified approximately nine SNP/kb across 517 resequenced rice varieties (Huang et al. 2010). Furthermore, the establishment and development of high-throughput genotyping methods have provided an abundance of valuable data on SNP frequency within and between germplasm groups, which make single-nucleotide polymorphisms (SNPs) highly attractive as genetic markers (Chen et al. 2011; Thomson et al. 2012; Yamamoto et al. 2010; Zhao et al. 2010, 2011).
Denaturing high-performance liquid chromatography (DHPLC) is an automated, highly sensitive and reliable genotyping platform for detecting SNPs and INDELs, based on the altered melting behavior of heteroduplexes versus homoduplexes in ion pair reverse-phase HPLC, under conditions of partial heat denaturation within a linear acetonitrile gradient (Hassan et al. 2013; Lehtokari et al. 2006; Li et al. 2008). In this technique, PCR product from a sample with heterozygous mutation or mixture of two PCR products (a reference and a test product) is denatured and annealed to form DNA heteroduplex molecules. On a chromatogram in the DHPLC system, DNA homoduplexes generally produce one peak, whereas DNA heteroduplexes elute earlier in one or more additional peaks. The sequence difference is thus translated into an altered elution profile (Nairz et al. 2002). Previous studies have demonstrated that the elution profile from DHPLC varies depending on the type of nucleotide change within the same fragment (Gross et al. 1999; Li et al. 2008; Wagner et al. 1999). Due to its ability to predict the sequence polymorphisms present in samples, DHPLC has been suggested as a highly effective method for genetic analyses (Gross et al. 1999).
DHPLC analysis provides high-throughput screening of samples in a 96-well format. In general, a single DHPLC analysis requires approximately 7–8 min, including column regeneration and re-equilibration to starting conditions. However, by using an accelerator, the running time per sample can be further minimized by 2–3 min, further improving the efficiency of the process (Raja et al. 2001).
In the last few years, DHPLC has become a widely employed method for the identification of SNPs in diagnostic testing for disease (Li et al. 2008) and high-resolution SNP mapping (Nairz et al. 2002; Raja et al. 2001). The application of DHPLC in the identification of crop species has not been reported to date.
In this study, we screened several amplicons containing multiple and informative SNPs based on the SNP data generated primarily from the OryzaSNP project (McNally et al. 2009) and the SNP research on 300 rice inbred lines (Chen et al. 2011). Further, we used DHPLC to genotype 47 rice varieties, test the variation within parental lines, and determine the hybrid heterozygosity by analyzing the screened amplicons. Our results demonstrate the power and usefulness of this technique for identifying rice varieties and assessing seed purity based upon elution profiles.
Materials and methods
Plant materials and DNA extraction
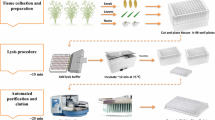
A total of 47 rice accessions representing the commercial rice germplasms used in China were chosen for DHPLC analysis. These rice accessions included 12 cytoplasmic male sterile (CMS) lines and 14 restorer (R) lines used as major parents in rice breeding programs in China, and 16 elite rice hybrids released for commercial cultivation in different regions of China. In addition, five inbred rice varieties were also analyzed (Supplementary Table S1). Among all rice varieties, 13 rice hybrids and their parental lines were included. Rice seeds of these varieties were obtained from Hunan Hybrid Rice Research Center (HHRRC), Changsha. DNA analysis of the varieties was conducted by randomly selecting several seeds per line and planting them in the paddy field in HHRRC. Rice leaves were collected for DNA extraction from approximately 10 plants of each genotype. Genomic DNA was extracted using the CTAB method of Murray and Thompson (1980).
Amplicons screening and PCR conditions
Because the elution profile from DHPLC varies depending on the type of nucleotide change, when detecting polymorphism by DHPLC, the analyzed amplicons should contain multiple and informative SNPs among the rice varieties. In this study, we screened amplicons within several chromosomes based on a set of 372 high-quality SNPs developed by Haodong Chen et al. (2011) and another SNP discovery set from the OryzaSNP project that provided comprehensive SNP data across 20 diverse rice varieties (McNally et al. 2009). The following procedure was used for amplicon screening: (1) select SNP loci on chromosomes 1–4 from the 372 SNPs set identified primarily by genotyping 300 rice accessions (Chen et al. 2011), and extract 1201 bp of genomic DNA sequence with 600 bp upstream and downstream of each selected SNP; (2) enter the 1201-bp sequence into Transgenomic WAVEMAKER™ Software for amplicon design, choose fragments with fairly uniform melt domains that range from 170 to 700 bp, and include the SNP loci selected in the previous step; (3) screen the fragments harboring two or more SNPs across four rice varieties (LTH, MH63, ZS97, SHZ2) that had been previously identified by the OryzaSNP project (McNally et al. 2009). From all screened fragments, 12 fragments distributed on four chromosomes were randomly selected for polymerase chain reaction (PCR) primer design.
PCR primers were designed for the 12 fragments using primer prime 5 software. PCRs consisted of 1× PCR buffer (Promega, USA), 5 pmol of each primer, 1.0 U of pfu DNA polymerase (Promega, USA), 250 μM dNTP, and 50 ng of genomic DNA, in a volume of 25 μL. The PCR cycles were carried out using a Veriti® Thermal Cycler with an initial denaturing step at 94 °C for 2 min, followed by 30 cycles of 30 s at 94 °C, 45 s at the respective annealing temperature (Supplementary Table S2), 1 min at 72 °C, and 10 min at 72 °C for final extension. Details regarding the 12 selected amplicons and their respective primers are given in Supplementary Table S2.
Heteroduplex formation
Prior to DHPLC analysis, unpurified PCR products of each rice variety underwent the denaturation/annealing step, which included denaturing at 95 °C for 5 min, followed by slow cooling to room temperature at a rate of 1 °C/min. The PCR products of the hybrids containing heterozygous sequence in the analyzed amplicon could form heteroduplex. In contrast, the PCR products of all R, CMS, and inbred lines, which are homozygous for any particular amplicon, and the PCR products of the hybrids, which are homozygous for the analyzed amplicon, could form homoduplex only. Therefore, it was necessary to perform an additional heteroduplex formation assay by mixing reference DNA and homoduplex to create a heteroduplex, to reveal sequence variants in the homozygous state by producing different DHPLC elution profiles. In the present study, PCR products from ZS97 were selected as reference DNA and added in equal proportions to each homozygous sample, and then, the denaturation/annealing step was performed. The DHPLC elution profile of the mixed homozygous samples would vary depending on the type of the sequence variation from ZS97.
In addition, equal amounts of the PCR products from the two parents of each hybrid were mixed together and hybridized with the denaturation/annealing step to create a heteroduplex. Due to the heterozygosity of the hybrid derived from the sequence variation between the parental lines, it is possible to determine hybrid seed purity by comparing the DHPLC elution profiles of the F1 hybrid with that of the mixed sample of the two parents.
DHPLC analysis
Analysis was performed using a WAVE DHPLC instrument (Transgenomic™). An aliquot (4–8 μL) of the products from the previous denaturation/annealing step was automatically injected into a DNASep column and analyzed at a flow rate of 0.90 ml/min by a linear acetonitrile gradient, achieved by mixing a buffer A (0.1 M TEAA, pH7.0) and a buffer B (0.1 M TEAA (Transgenomic, USA) with 25 % acetonitrile, pH7.0). The initial oven temperature and gradient conditions to be tested were determined using an algorithm provided by the WAVEMAKER (Transgenomic) software. When necessary, the temperatures and gradient condition were changed to obtain the optimal separation of heteroduplex molecules.
Direct sequencing
To identify the sequence polymorphisms among the rice varieties detected by DHPLC, the PCR products from each rice variety were further sequenced. To facilitate direct sequencing, universal M13 forward and M13 reverse promoter homologous sequences were added to each primer pair indicated in Supplementary Table S2 for re-amplifying each rice variety sequence. Amplified PCR products were purified and sequenced in both directions using universal M13 forward and M13 reverse primers by Sangon Biotech (Shanghai, China). DNA sequence data from both directions were checked for sequencing errors and aligned using Sequencher software (Gene Codes Corporation, USA). Sequences polymorphisms were identified manually.
Statistical analysis
To evaluate the repeatability and reproducibility of DHPLC analysis for rice genotyping, intra-day and inter-day assays were performed by analyzing each sequence variant of rice in 10 experiments per week on three consecutive weeks, under optimal DHPLC conditions. Buffer A and buffer B were replaced every week. The precision of DHPLC analysis was assessed by retention time differences and coefficients of variation. Statistical analyses were conducted with the SPSS 13.0 software program (SPSS, USA).
Results
Determination of temperature and separation gradient conditions
The temperature at which the samples are run is a critical parameter for the DHPLC procedure. In the present work, we first analyzed the PCR product from ZS97 at the temperate and separation gradient conditions predicted by WAVEMAKER software to obtain a homoduplex elution profile for each amplicon. As the homoduplex DNA, the PCR product from each amplicon from ZS97 should yield an elution profile with a single peak. The experimental results showed that the software-predicted temperature was suitable for the analysis of homoduplex DNA for only five amplicons. For example, at the software-deducted temperature (58.5 °C) for amplicon IV, the PCR product from ZS97 produced an elution profile with multiple peaks (Supplementary Fig S1a). This result may have occurred because the temperature was too high to partially melt the homoduplex DNA. Therefore, we analyzed the PCR product from ZS97 in 0.5 °C decrements over the range of predicted temperature until a single peak was detected. For DHPLC analysis of amplicon IV, the lower temperature (56.5 °C) was found to be appropriate for the analysis of homoduplex DNA (Supplementary Fig S1b).
The production of an elution profile with a single peak by homoduplex DNA is not the only criterion for temperature selection. For some amplicons, it was necessary to further adapt the temperature to resolve as many rice genotypes as possible. As shown in Fig. 1a, the temperature (59.4 °C) fit for the homoduplex DNA analysis was not optimal for the detection of sequence variants, due to the presence of a broadened signal as a result of an excessively high temperature, yielding inconsistent and irreproducible results. Under lower temperature (58.4 °C), the PCR product containing one type of sequence variant showed a single peak and was almost indistinguishable from homoduplex DNA (Fig. 1b). At increasing temperature, each genotype produced a unique elution profile that differed from the others with respect to the number and retention time of its DHPLC peaks (Fig. 1c). However, the peak for heteroduplex DNA was not apparent. To obtain a clear profile, we changed the linear gradient from a slope of 2 % increase in buffer B per minute to a 3 % increase, resulting in a narrow and enhanced signal, and finally obtained elution profiles with more apparent peaks (Fig. 1d). The optimal DHPLC gradient and temperature conditions for all the 12 amplicons are listed in Supplementary Table S3.
Impact of temperate and gradient conditions on DHPLC analysis. Profiles were produced by analyzing amplicon VII PCR products synthesized from three different rice genotypes under four separation conditions. a PCR products eluted with a slope of 2 % increase in buffer B per min and analyzed at 59.4 °C. b PCR products eluted with a slope of 2 % increase in buffer B per min and analyzed at 58.4 °C. c PCR products eluted with a slope of 2 % increase in buffer B per min and analyzed at 58.7 °C. d PCR products eluted with a slope of 3 % increase in buffer B per min and analyzed at 58.7 °C
DHPLC patterns
The PCR products of the 12 amplicons synthesized from 47 rice varieties were subjected to DHPLC analysis under the separation conditions listed in Supplementary Table S3. Their corresponding elution profiles were compared with the patterns from ZS97, which is a homozygous line selected as reference control. For each amplicon, the elution profiles of the restorer, CMS, and inbred lines were all highly similar to that of ZS97 and showed a single peak of homoduplex DNA. In contrast, different DHPLC elution profiles were detected upon analyzing the hybrids. In addition to profiles with a single peak, some hybrid PCR products produced elution profiles with multiple peaks due to the reduced column retention time of heteroduplex DNA, implying that these rice hybrids are heterozygous for the analyzed amplicon. In order to discriminate the homozygous samples based on their DHPLC elution patterns, each PCR product of the CMS, restore, inbred lines, and hybrids that produced single peak profiles in the initial DHPLC analysis was mixed with the PCR product of ZS97, respectively, and submitted for the additional DHPLC analysis. As expected, some of them exhibited altered DHPLC elution profiles and exhibited distinct patterns with multiple peaks.
By the continuous DHPLC analysis for all the PCR products and mixed PCR samples of a single amplicon, different DHPLC profiles could be detected and distinguished easily due to their significant differences in peak number and retention time. For each amplicon, some samples showed nearly identical profiles, which had the same peak number and similar retention times. As shown in Fig. 2, the CMS line (P64S), restore line (FH838), inbred lines (YJZ), and hybrid (YY9113) produced identical profiles with a single peak, and some hybrids (SY63, YLY-1, CLY343) produced two distinct profiles differing in peak number and retention times. By mixing with ZS97, FH838 and P64S produced multiple peaks which are highly similar to the patterns of hybrids SY63, YLY-1, and CLY343, respectively, while YJZ and YY9113 still produced a single peak.
DHPLC elution profiles of seven rice lines obtained by DHPLC analysis of amplicon III. Direct DHPLC analysis on PCR products of seven rice lines (YJZ, YY9113, FH838, P64S, SY63, YLY-1, CLY343) and additional DHPLC analysis on mixed PCR samples of four lines (YJZ, YY9113, FH838, P64S), which produced single peak profile in the initial DHPLC analysis. Variety names are labeled to the right of the chromatogram
Overall, the 12 amplicons selected in the present study exhibited a total of 43 different DHPLC elution profiles detected across the hybrids and mixed samples. Of the 12 amplicons, amplicon VII exhibited the highest polymorphism, producing six types of elution profiles (Fig. 3g), whereas the fewest polymorphic patterns were presented by amplicons I, II, and V, with only two types produced for all samples (Fig. 3a, b, e). The others included five types of elution profiles for amplicons XI and IX (Fig. 3k, i); four types for amplicons IV, VI, and XII (Fig. 3d, f, l); and three types for amplicons III, VIII, and X (Fig. 3c, h, j). For all the DHPLC profile patterns, 32 different types were detected in both the hybrids and mixed samples, seven types were detected only in the mixed samples, and four types were detected only in the hybrids.
Polymorphism of DHPLC profiles within a amplicon I, b amplicon II, c amplicon III, d amplicon IV, e amplicon V, f amplicon VI, g amplicon VII, h amplicon VIII, i amplicon IX, j amplicon X, k amplicon XI, and l amplicon XII. Elution profiles of the 12 amplicons were obtained by analyzing crude PCR products and mixed PCR products of rice varieties under respective DHPLC conditions. DHPLC profiles with distinct characteristics are superimposed to illustrate the differences. The retention times for homoduplex and heteroduplex fractions are indicated above the elution peaks
To assess whether it was possible to assign a specific elution profile to a sequence variant within the amplicon, the PCR products amplified from each rice variety were sequenced. The sequencing results validated that the CMS, R, inbred lines, and hybrids that produced single peak profiles in the initial DHPLC analysis were all homozygous for the analyzed amplicon. When these samples were mixed with ZS97, no nucleotide variation from ZS97 was detected in any of the samples for which the elution profile was the same with a single peak. Other DHPLC profiles with multiple peaks associated with heteroduplex DNA from hybrids or the mixed samples varied in accordance with the type of nucleotide change. As shown in Fig. 4, for nucleotide mutations at positions 50 and 104 of the sequence of amplicon IV (50G → A and 104T → C), a multi-peak elution profile (type A) was produced that was distinct from profile type B associated with the samples containing G → A alterations at base 50 and C → A alterations at base 110. In the case of the coexistence of two substitutions at positions 104 and 110 (104T → C and 110C → A) in the same amplicon, elution profile type C, which was noticeably different from types B and C, was observed.
DHPLC elution profiles for three sequence variant types within amplicon IV. a Elution profile for a hybrid carrying heterozygous mutations at positions 50 and 104. This profile pattern was defined as type A. The direct sequencing result illustrates the corresponding heterozygous G/A (50) and T/C (104) changes. b Elution profile for a hybrid carrying heterozygous mutations at positions 50 and 110. This profile pattern was defined as type B. The direct sequencing result illustrates the corresponding heterozygous G/A (50) and C/A (110) changes. c Homoduplex profile for ZS97 and heteroduplex profile for an inbred sample mixed with ZS97. The profile pattern of the mixed inbred sample was defined as type C. Comparison of the sequencing chromatograms from the inbred line and ZS97 illustrates the sequence differentiation between these lines with two base pair mismatches at positions 104 (T → C) and 110 (C → A), which lead to the formation of heteroduplex DNA by the denaturation/annealing step
Our sequencing results showed that the samples exhibiting nearly identical elution profiles harbor common sequence variants, and each sequence variant presented a unique elution profile for each of the 12 amplicons. All of the sequence variants involved single-nucleotide substitutions except the sequence variants associated with profile types C, D, and F for amplicon VII and a sequence variant associated with profile type C for amplicon X, which included deletions and insertions, in addition to several nucleotide substitutions (data not shown). Our results demonstrated that the polymorphism of the elution profiles was derived from the polymorphism in the positions and numbers of SNPs and INDELs within the amplicons among the different rice varieties.
As shown in Supplementary Fig S2, the profiles associated with a specific sequence variant exhibited the similar patterns and same peak quality by both intra-day and inter-day analyses. But significant inter-day assay variation in retention times was observed, which may be due to subtle changes in buffer composition, making profile characterization by retention time of peaks unreliable. However, it was noted that for each sequence variant, the difference in the retention time between each peak (ahead of the last peak) and the last peak (Delta-Peak, ∆P) just varied slightly by the intra-day and inter-day assays and stayed within a certain range (Supplementary Table S4). Therefore, we proposed that the peak number and the difference in the retention time between each peak and the last peak (∆P) could be used as the critical parameters to characterize DHPLC profiles that correlated with a specific sequence variant.
Identification of rice genotypes and assessment of hybrid purity by DHPLC analysis
In the 16 rice hybrids from which crude PCR products of the 12 amplicons were subjected to DHPLC analysis, the number of hybrids for which multiple peaks were detected averaged 9 per amplicon, with a minimum of 5 (amplicon IX) and a maximum of 14 (amplicon VI and VIII). It was easier to distinguish the hybrids producing profiles with multiple peaks from their homozygous parental lines as well as other hybrids that produced profiles with a single peak. In addition, the differences in peak number and ∆P values could help to distinguish these hybrids from each other. Interestingly, profile type B for amplicon VII, profile type B for amplicon VI, and type E for amplicon XI were all unique to their particular hybrid. These unique profiles could be the characteristic markers for the identification of these hybrids.
However, all R, CMS, and inbred lines and some hybrids that produced a single peak profiles in the first DHPLC analysis could not be distinguished from each other, because they could not be determined directly by the retention times. The additional heteroduplex-based assay of mixing using the PCR product from ZS97 was effective at changing the DHPLC profile pattern from monomorphism to polymorphism among these samples, allowing these samples to be discriminated based on specific profile patterns, resulting from their different sequence deviations from ZS97. In all elution profile patterns of mixed samples, type C for amplicon XI was specific to a restore line; type C for amplicon IV, type B for amplicon VIII, type B for amplicon IX, and type D for amplicon XII were detected in only LTH. In addition, of all the rice varieties analyzed in this study, only LTH showed an elution profile with multiple peaks for each of the 12 amplicons. We propose that this result was obtained because LTH was the only variety belonging to the japonica group in our study, whereas ZS97 and the other varieties all belong to the indica group.
After analyzing crude and mixed PCR products by DHPLC, we combined the elution profile types of the 12 amplicons for each rice variety and obtained a unique fingerprint for all of the hybrids. Therefore, DHPLC analysis was effective at discriminating between the 12 amplicons. The fingerprints of the inbred, restorer, and CMS lines were also distinct, with the exception of three CMS lines FY-A, TF-A, and J23A, which had identical profiles with respect to all the 12 amplicons analyzed in this study. Nevertheless, distinct characteristic profiles in six amplicons (IV, VI, VII, VIII, X, XI) together differentiated all the hybrids, inbred, and restorer lines with at least a single profile pattern difference (Supplementary Fig S3).
Hybrids display a characteristic pattern of DHPLC elution profile resulting from parental variation in allelic nucleotide sequence. Therefore, it is practical to test genetic purity of F1 hybrid seeds by comparing their DHPLC elution profiles with that of a mixed sample of their two parents. Our results showed three different cases: (1) As shown in Fig. 5a, if parental lines have identical profiles, their hybrid would show a profile with a single peak that is the same as that of a mixed parent sample. When mixed with ZS97, the hybrid showed a DHPLC profile pattern identical to those of the two parents. (2) If one of the parents produced a single peak, while the other one exhibited a different pattern with multiple peaks, their hybrid would produce a DHPLC profile pattern identical to the parent producing multiple peaks (Fig. 5b). (3) If the two parental lines each exhibited DHPLC profiles with multiple peaks that are distinct from each other, the mixed parental sample would produce a novel type of elution profile, as was also detected in their hybrid sample (Fig. 5c). Our results demonstrated that regardless of the type of pattern produced by the parents, the hybrid always showed a DHPLC profile pattern consistent with the mixed sample of the two parents. In case (1), the hybrid produced a DHPLC profile pattern highly similar to that of its two parents, by both the first DHPLC analysis on the PCR products and the second DHPLC analysis on the mixture of test sample and ZS97. This result makes it impossible to detect contamination of these hybrid samples with their parents. It also implies that only the amplicons that exhibited polymorphic DHPLC patterns between the hybrids’ respective parents could be selected as DHPLC markers for the hybrid seed purity assessment.
DHPLC analysis of hybrids and their parents. The male and female parents of the hybrid were mixed with PCR product from ZS97 to produce a characteristic DHPLC profile. Moreover, the two parents were mixed with each other in an equimolar ratio for further DHPLC analysis. Each PCR product of the hybrid was subjected to DHPLC for initial analysis. Then, the hybrids producing a single peak in first analysis were mixed with PCR product from ZS97 for a second DHPLC analysis. a Parental lines have identical characteristic profiles. The hybrid and mixed parent sample produced a single peak, whereas the hybrid mixed with ZS97 formed a heteroduplex, resulting in extra peaks identical to those of the parents. b Two parents produced different patterns of DHPLC profiles, one of which showed a single peak. Both the hybrid and the mixed parent sample produced DHPLC profiles with multiple peaks identical with the parent producing multiple peaks profile. c Two parents produced different profiles that showed multiple peaks and differed with respect to ∆P values. The hybrid and mixed parent sample produced a novel and identical elution profile pattern that was completely distinct from both parents
Discussion
Numerous types of DNA markers that are based on the indirect detection of sequence-level polymorphism have been developed (Henry 2001; Phillips and Vasil 2001). Sequence polymorphisms such as single-nucleotide polymorphisms (SNPs), nucleotide insertions, and deletions exploited as genetic markers can be detected by DHPLC analysis. In this study, we first applied DHPLC analysis to identify rice varieties and test the genetic purity of rice hybrids using DHPLC profile polymorphisms corresponding to various sequence variants. The sensitivity of the method is dependent on the temperature at which the analysis is performed. Because the temperature has a significant effect on the separation of hetero- and homoduplexes, the choice of operating temperature for DHPLC is critical for detecting different genotypes. Under the conditions established in this present study, DHPLC resolved 100 % of the sequence variants detected by sequencing. Although no nucleotide variation was detected by sequencing in any of the samples producing only a single peak, we suppose that it is possible that a few mutations could remain undetected by this method, because DHPLC analysis at a single temperature can fail to identify all alterations due to differences in the melting point of sequences flanking the SNP site. Therefore, to overcome this possibility, the amplicon should be designed to have a single and fairly uniform melt domain. Furthermore, we confirmed that amplicons vary significantly in the level of polymorphism of the DHPLC elution profile that is produced due to differences in the level of DNA sequence polymorphism. Thus, amplicon selection is another critical factor for this DHPLC assay. It is essential to select amplicons producing highly informative polymorphic elution profiles as the target fragments for DHPLC analysis.
This methodology yielded highly accurate and reproducible results in producing highly polymorphic chromatographic patterns from homoduplexes and heteroduplexes in the samples. We were able to genotype rice varieties solely based upon the combination of characteristic DHPLC profiles produced by two DHPLC analyses. In the first DHPLC analysis, the PCR products from all studied rice varieties were analyzed, and heterozygous genotypes of the hybrids could be determined due to their different characteristic DHPLC pattern corresponding to the type of nucleotide change. An additional mixing of the tested sample and ZS97 was conducted for the second DHPLC analysis when genotyping homozygotes that showed a single homoduplex peak. Only by the second DHPLC analysis, the different homozygous genotypes of R, CMS, inbred lines, and some hybrids, which showed single peak patterns in the first DHPLC analysis, can be discerned due to their characteristic DHPLC profile corresponding to their sequence variation from ZS97. In this study, we established a novel assay for identifying 44 rice varieties based on 12 amplicon-based DHPLC patterns. As the number of available amplicons grows, the required number of fragments to be analyzed is expected to decrease even for the identification of a greater number of rice varieties. It is feasible to automatically assign the identity of a rice variety using DHPLC profile patterns, if we can develop a software program capable of cataloging the different DHPLC elution profile patterns for each amplicon compiled into a DHPLC profile-based fingerprint database of rice.
Determining the genetic purity of hybrid seed is an essential requirement for its commercial use. In this study, we propose that as long as the amplicons that exhibited polymorphic DHPLC patterns between the hybrids’ respective parent were selected as markers, the DHPLC assay would detect multiple peak profile patterns highly similar to that of the mixed sample of the two parental lines in pure hybrids, whereas impurity could be identified by the presence of a single peak or other multiple peak patterns. In addition, genetic heterogeneity within the parental lines could lead to a lack of uniformity in the commercial F1 seed resulting in poor acceptance of the hybrid produced (Nandakumar et al. 2004). Due to its ability to readily distinguish between homozygosity and heterozygosity, DHPLC could also be used for testing genetic heterogeneity within the parental lines.
Frequently, highly informative SSR markers are preferred in cultivar identification and seed purity testing because of their codominant nature and high levels of polymorphism. However, the detection of SSR genotypes is often conducted using agarose or polyacrylamide gel electrophoresis or sequencers, which is time-consuming or costly. In addition, these markers have constraints in multiplexing, automation and limited abundance when saturating genomic regions. Although they have obvious advantages compared with SSRs in these respects, SNPs are less informative than SSR markers (Hamblin et al. 2007), because SNPs are bi-allelic markers that generally present only two alleles per SNP marker, whereas SSRs are multi-allelic markers capable of presenting a very large number of alleles per locus (Liu et al. 2003; Lu and Bernardo 2001). Consequently, it has been suggested that combining information from several SNPs within the same gene or locus may provide a partial solution to the disadvantage of SNP markers (Yan et al. 2010), and 3–4 SNP loci would be needed to provide the discriminatory power of one microsatellite locus (José et al. 2011). In this study, our results demonstrated the ability of DHPLC to examine and discriminate multiple SNPs within one amplicon. SNPs can be as informative as multi-allelic molecular markers when detected by DHPLC.
The strengths of DHPLC methods include ease of instrument operation, low reagent costs, and high sample throughput. Commercially available DHPLC instrumentation is as easy to use as are standard HPLC instruments. This system is semiautomatic and can process 2 × 96-well sample plate unattended for test, which is adequate for high-throughput analysis. And the cost to identify rice variety using this system would have been approximately $1.7 per test, which are $0.9 for high-fidelity PCR reagent, $0.4 for DHPLC buffers, and $0.4 for the DHPLC column. Due to these advantages, this genotyping platform might overtake SSR and be applicable for the rapid and automated genetic characterization of the diverse germplasm of rice. It might also be utilized to monitor seed genetic purity at both the parental and hybrid seed production stages.
References
Blair MW, Panaud O, McCouch SR (1999) Inter-simple sequence repeat (ISSR) amplification for analysis of microsatellite motif frequency and fingerprinting in rice (Oryza sativa L.). Theor Appl Genet 98(5):780–792
Chen HD, Hang H, Zou YJ, Chen W, Yu RB, Liu X, Yang Y, Gao YM, Xu JL, Fan LM, Li Y, Li ZK, Deng XW (2011) Development and application of a set of breeder-friendly SNP markers for genetic analyses and molecular breeding of rice (Oryza sativa L.). Theor Appl Genet 123(6):869–879
Diego L, José AC, Ana I, Virginia R, José MM-Z (2007) High throughput SNP discovery and genotyping in grapevine (Vitis vinifera L.) by combining a re-sequencing approach and SNPlex technology. BMC Genom 8:424
Feltus FA, Wan J, Schulze SR, Estill JC, Jiang N, Paterson AH (2004) An SNP resource for rice genetics and breeding based on subspecies indica and japonica genome alignments. Genome Res 14:1812–1819
Flint-Garcia SA, Thuillet AC, Yu J, Pressoir G, Romero SM, Mitchell SE, Doebley J, Kresovich S, Goodman MM, Buckler ES (2005) Maize association population: a high-resolution platform for quantitative trait locus dissection. Plant J 44:1054–1064
Garris AJ, Tai TH, Coburn J, Kresovich S, McCouch S (2005) Genetic structure and diversity in Oryza sativa L. Genetics 169:1631–1638
Giordano M, Oefner PJ, Underhill PA, Luca Cavalli Sforza L, Tosi R, Richiardi PM (1999) Identification by denaturing high-performance liquid chromatography of numerous polymorphisms in a candidate region for multiple sclerosis susceptibility. Genomics 56:247–253
Gross E, Arnold N, Goette J, Schwarz-Boeger U, Kiechle M (1999) A comparison of BRCA 1 mutation analysis by direct sequencing, SSCP and DHPLC. Hum Genet 105:72–78
Hamblin MT, Warburton ML, Buckler ES (2007) Empirical comparison of simple sequence repeats and single nucleotide polymorphisms in assessment of maize diversity and relatedness. PLoS ONE 2:e1367
Hassan D, Nazanin V, Majid F (2013) DNA sequence fragment containing C to A mutation as a convenient mutation standard for DHPLC. Iran J Basic Med Sci 16(8):946–949
Henry RJ (2001) Plant genotyping. The DNA fingerprinting of plants. CABI, Wallingford
Huang X, Wei X, Sang T, Zhao Q, Feng Q, Zhao Y, Li C, Zhu C, Lu T, Zhang Z, Li M, Fan D, Guo Y, Wang A, Wang L, Deng L, Li W, Lu Y, Weng Q, Liu K, Huang T, Zhou T, Jing Y, Li W, Lin Z, Buckler ES, Qian Q, Zhang QF, Li J, Han B (2010) Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42:961–967
José AC, Javier I, Diego L, Dolores V, Gema B, Virginia R, Iván C, Angelica MJ, Juan C, Leonor RG, Mark RT, José MM-Z (2011) A 48 SNP set for grapevine cultivar identification. BMC Plant Biol 11:153
Lehtokari VL, Pelin K, Sandbacka M, Ranta S, Donner K, Muntoni F et al (2006) Identification of 45 novel mutations in the nebulin gene associated with autosomal recessive nemaline myopathy. Hum Mutat 27:946–956
Li Q, Li LY, Huang SW, Li L, Chen XW, Zhou WJ, Xu XM (2008) Rapid genotyping of known mutations and polymorphisms in β-globin gene based on the DHPLC profile patterns of homoduplexes and heteroduplexes. Clin Biochem 41:681–687
Liu K, Goodman MM, Muse S et al (2003) Genetic structure and diversity among maize inbred lines as inferred from DNA microsatellites. Genetics 165:2117–2128
Lu H, Bernardo R (2001) Molecular diversity among current and historical maize inbreds. Theor Appl Genet 103:613–617
Mao CX, Virmani SS, Kumar I (1996) Technological innovations to lower the costs of hybrid rice seed production. In: Virmani SS, Siddiq EA, Muralidharan K (eds) Advances in Hybrid Rice Technology. Directorate of Rice Research, Hyderabad, pp 111–128
McCouch SR, Zhao K, Wright M, Tung CW, Ebana K, Thomson M, Reynolds A, Wang D, DeClerck G, Ali ML, McClung A, Eizenga G, Bustamante C (2010) Development of genome-wide SNP assays for rice. Breed Sci 60:524–535
McNally K, Childs K, Bohnert R, Davidson R, Zhao K, Ulat V, Zeller G, Clark R, Hoen D, Bureau T, Stokowski R, Ballinger D, Frazer K, Cox D, Padhukasahasram B, Bustamante C, Weigel D, Mackill D, Bruskiewich R, Rätsch G G, Buell C, Leung H, Leach J (2009) Genomewide SNP variation reveals relationships among landraces and modern varieties of rice. Proc Natl Acad Sci USA 106:12273–12278
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight DNA. Nucleic Acids Res 8:4321–4325
Nairz K, Stocker H, Schindelholz B, Hafen E (2002) High-resolution SNP mapping by denaturing HPLC. PNAS 99(16):10575–10580
Nandakumar N, Singh AK, Sharma RK, Mohapatra T, Prabhu KV, Zaman FU (2004) Molecular fingerprinting of hybrids and assessment of genetic purity of hybrid seeds in rice using microsatellite markers. Euphytica 136:257–264
Ni J, Colowit PM, Mackill DJ (2002) Evaluation of genetic diversity in rice subspecies using microsatellite markers. Crop Sci 42:601–607
Ohtsubo K, Nakamura S (2007) Cultivar identification of rice (Oryza sativa L.) by polymerase chain reaction method and its application to processed rice products. J Agric Food Chem 55:1501–1509
Phillips RL, Vasil IK (2001) DNA-based markers in plants, 2nd edn. Kluwer, Dordrecht
Rafalski A (2002a) Applications of single nucleotide polymorphisms in crop genetics. Curr Opin Plant Biol 5(2):94–100
Rafalski JA (2002b) Novel genetic mapping tools in plants: SNPs and LD-based approaches. Plant Sci 162(3):329–333
Rahman MS, Molla MR, Alam MS, Rahman L (2009) DNA fingerprinting of rice (Oryza sativa L.) cultivars using microsatellite markers. Aust J Crop Sci 3(3):122–128
Raja K, Markus W, Wolfgang M, Andreas G (2001) Application of denaturing high-performance liquid chromatography for mapping of single nucleotide polymorphisms in barley (Hordeum vulgare L.). Genome 44:523–528
Saker MM, Youssef SS, Abdallah NA, Bashandy HS, Sharkawy AM (2005) Genetic analysis of some Egyptian rice genotypes using RAPD, SSR and AFLP. Afr J Biotechnol 4(9):882–890
Shen YJ, Jiang H, Jin JP, Zhang ZB, Xi B, He YY, Wang G, Wang C, Qian L, Li X, Yu QB, Liu HJ, Chen DH, Gao JH, Huang H, Shi TL, Yang ZN (2004) Development of genome- wide DNA polymorphism database for map-based cloning of rice genes. Plant Physiol 135:1198–1205
Simko I, Haynes KG, Ewing EE, Costanzo S, Christ BJ, Jones RW (2004) Mapping genes for resistance to Verticillium alboatrum in tetraploid and diploid potato populations using haplotype association tests and genetic linkage analysis. Mol Genet Genomics 271(5):522–531
Szalma SJ, Buckler ES, Snook ME, McMullen MD (2005) Association analysis of candidate genes for maysin and chlorogenic acid accumulation in maize silks. Theor Appl Genet 110(7):1324–1333
Thomson MJ, Zhao KY, Wright M, McNally KL, Rey J, Tung CW, Reynolds A, Scheffler B, Eizenga G, McClung A, Kim H, Ismail AM, Ocampo M, Mojica C, Reveche MY, Dilla-Ermita CJ, Mauleon R, Leung H, Bustamante C, McCouch SR (2012) High-throughput single nucleotide polymorphism genotyping for breeding applications in rice using the BeadXpress platform. Mol Breed 29:875–886
Wagner T, Stoppa-Lyonnet D, Fleischmann E, Muhr D, Pages S, Sandberg T, Caux V, Moeslinger R, Langbauer G, Borg A, Oefner P (1999) Denaturing high-performance liquid chromatography detects reliably BRCA 1 and BRCA 2 mutations. Genomics 62:369–376
Wang ZY, Tanksley SD (1989) Restriction fragment length polymorphism in Oryza sativa L. Genome 32:1113–1118
Yamamoto T, Nagasaki H, Yonemaru J, Ebana K, Nakajima M, Shibaya T, Yano M (2010) Fine definition of the pedigree haplotypes of closely related rice cultivars by means of genome-wide discovery of single-nucleotide polymorphisms. BMC Genom 11:267
Yan JB, Yang XH, Shah T, Héctor SV, Li JS, Warburton M, Zhou Y, Crouch JH, Xu YB (2010) High-throughput SNP genotyping with the GoldenGate assay in maize. Mol Breed 25(3):441–451
Yong HJ, Sang NA, Hae CC, Tae RH, Huhn PM (1999) Identification of a RAPD marker linked to a brown planthopper resistance gene in rice. Euphytica 107(1):23–28
Zhang Q, Maroof MAS, Lu TY, Shen BZ (1992) Genetic diversity and differentiation of indica and japonica rice detected by RFLP analysis. Theor Appl Genet 83:495–499
Zhao K, Wright M, Kimball J, Eizenga G, McClung A, Kovach M, Tyagi W, Ali ML, Tung CW, Reynolds A, Bustamante CD, McCouch SR (2010) Genomic diversity and introgression in O. sativa reveal the impact of domestication and breeding on the rice genome. PLoS ONE 5:e10780
Zhao K, Tung CW, Eizenga GC, Wright MH, Ali ML, Price AH, Norton GJ, Islam MR, Reynolds A, Mezey J, McClung AM, Bustamante CD, McCouch SR (2011) Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2:467
Zhu YF, Qin GC, Hu J et al (2012) Fingerprinting and variety identification of rice (Oryza sativa L.) based on simple sequence repeat markers. Plant Omics J 5(4):421–426
Acknowledgments
This work is financially supported by the National Key Technology Research and Development Program of the Ministry of Science and Technology of China (2012BAC01B07).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Neither I nor any person with whom I have or had e personal or business relationship is engaged in any transaction or activity or has any relationship that may represent a potential competing or confliction interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Peng, Z., Yuan, X., Huang, Y. et al. Application of denaturing high-performance liquid chromatography for rice variety identification and seed purity assessment. Mol Breeding 36, 19 (2016). https://doi.org/10.1007/s11032-015-0429-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0429-8