Abstract
The mode of inheritance of wheat leaf rust resistance gene Lr45 was studied at seedling stage under greenhouse conditions against leaf rust race 77-5 in two F2 populations derived from the crosses between Thatcher (Tc)+Lr45 and two susceptible cultivars Agra Local and NI5439. The genetic analysis in F2:3 progeny validated the F2 results which unambiguously showed segregation for a single dominant gene. Genetic analysis in F2 and BC1F1 generations against five other leaf rust races confirmed the single dominant gene inheritance of Lr45. Mapping was carried out with 92 microsatellite markers specific to chromosome 2A on the F2 population of the cross Agra Local × Tc+Lr45. Out of seven markers linked to the gene, four (gwm372, gwm275, gpw3167 and gwm122) were co-dominant and the other three (cfd168, cfd6 and gwm249) showed dominance, amplifying the allele only in the susceptible parent. The genetic map of 13.1 cM was constructed based on the results in 140 homozygous resistant and homozygous susceptible plants. cfd168 was the closest marker linked to Lr45, followed by gwm372. These markers were validated on the NI5439 × Tc+Lr45 F2 population, 12 different backcross lines carrying Lr45 and near-isogenic lines, mostly in Tc background isogenic for 46 different Lr genes belonging to both native and alien species. The marker gwm122 was found to be monomorphic. The closest co-dominant marker gwm372 showed reduced polymorphism. Two sequence-based primer pairs, G372 94 and G372 185 , were designed and validated. Hence, the markers G372 94 and G372 185 closely linked to the gene can serve as robust co-dominant markers for utilization of Lr45 in wheat improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Globally, wheat (Triticum aestivum L.) rusts occupy a prominent place among the fungal diseases, causing severe damage to the crop and leading to huge losses in grain production. Leaf rust caused by Puccinia triticina Eriks. is present wherever wheat is grown in the world. Although rusts can be controlled by fungicide sprays, the use of genetic resistance in cultivars is a more cost-effective and environment friendly method.
Of the 71 documented leaf rust resistance (Lr) genes, approximately 38 have been transferred to bread wheat from related and distantly related species and genera, including Aegilops, Agropyron, Secale cereale, Triticum, Thinopyrum and Elymus. Several of them have been deployed to protect crops in different wheat-growing regions of the world (McIntosh et al. 2011; Herrera-Foessel et al. 2012; Singh et al. 2012; Tomar et al. 2014). Rye (S. cereale L.) is one of the important donors of disease and pest resistance to wheat. Among the many genes transferred from rye, only those on the short arm of chromosome 1R have been widely exploited. The 1BL.1RS wheat–rye translocation carrying the resistance genes for leaf, stem and stripe rusts and powdery mildew has contributed substantially to world wheat production. However, many of the alien genes, including those carried by IBL.1RS translocation, no longer confer resistance due to the appearance of new virulent races of P. triticina. This therefore necessitates the search for new resistance genes.
In this context, the wheat–rye translocation lines developed by Mukade et al. (1970) for rust resistance through spontaneous and X-ray irradiation assume great importance. One of the derivatives, a putative spontaneous translocation stock ST-1, showed a high degree of leaf rust resistance. Later it was subjected to cytogenetic analysis, C-banding and in situ hybridization studies by McIntosh et al. (1995) and the structure of the translocation chromosome was defined as T2AS-2RS.2RL. This translocation appeared to be carrying a new gene for leaf rust resistance and it was designated Lr45. However, the monosomic analysis of the F2 population for leaf rust resistance showed deviation from the monogenic inheritance and an excess of susceptible segregants was observed. The gene Lr45 confers a high degree of seedling and adult plant resistance to leaf rust but to date has not been utilized (Friebe et al. 1996; Tomar and Menon 2001; Bhardwaj et al. 2010) in wheat improvement. Hence, it is important to understand the mode of inheritance of Lr45 before its utilization in breeding programmes.
Earlier, the gene Lr45 was mapped using amplified fragment length polymorphism (Zhang et al. 2006) and sequence characterized amplified region (Fein et al. 2009) markers. However, the unavailability of complete information along with the dominant nature of inheritance, high costs and labour involved restrict the use of these markers in marker-assisted breeding. Therefore, simple sequence repeat (SSR) markers are used more frequently than other markers due to advantages associated with co-dominance, accuracy, high repeatability, high level of polymorphism, chromosome specificity and ease of manipulation. The present investigation was therefore undertaken to study the mode of inheritance and to map the leaf rust resistance gene Lr45 using SSR markers.
Here we report the mode of inheritance of Lr45 in different segregating populations against multiple races of leaf rust pathogen. This study also reports the identification of SSR markers linked to the gene Lr45, and the development of new markers from the locus gwm372 to provide ease of selection for marker-assisted breeding.
Materials and methods
Plant materials
A near-isogenic line (NIL) of Thatcher (Thatcher*7/ST-1 = RL6144) carrying leaf rust resistance gene Lr45 (Tc+Lr45) and two susceptible cultivars, Agra Local and NI5439, were used to study the mode of inheritance and mapping of Lr45. The line Tc+Lr45 was crossed with Agra Local and NI5439. The F1 plants were selfed and harvested separately to get the F2 seed. The F1 (Agra Local × Tc+Lr45) was backcrossed to Agra Local to get BC1F1 seeds. For screening the F3 families, single selfed spikes with at least 20 seeds were utilized from each resistant and susceptible F2 plant to confirm their genotype. The inheritance of Lr45 was studied in both F2 and BC1F1 generations, whereas the molecular mapping was done with the F2 population from the cross Agra Local × Tc+Lr45.
Validation of the markers linked to Lr45 was performed on a subset of the population comprising 92 F2 plants from the cross NI5439 × Tc+Lr45. The efficiency of the linked markers was checked on backcross lines of wheat carrying leaf rust resistance gene Lr45 in the genetic background of 12 different wheat cultivars. The specificity of the markers was also tested on a set of 46 NILs of leaf rust resistance genes, mostly in Thatcher background carrying 23 leaf rust resistance genes derived from related/alien species and 23 leaf rust resistance genes derived from bread wheat.
Leaf rust races
The pure inoculum of the six different races of P. triticina, i.e. 77-5 (syn. 121R63-1), 12-1 (syn. 5R37), 162 (syn. 93R7), 104-2 (syn. 21R31), 107(45R3) and 108 (syn. 13R27), was obtained from the Directorate of Wheat Research, Regional Station, Flowerdale, Shimla, India. The races were maintained and multiplied on susceptible cultivar Agra Local under greenhouse conditions at the Division of Genetics, IARI, New Delhi, India. Spores were sprayed as a suspension in water with a drop of Tween20 (0.75 μl/ml). The parents, F1, F2 and F3 populations from both the crosses were tested for infection type (IT) against the most virulent race 77-5 of P. triticina. The seedlings of F2 and BC1F1 populations, along with the parents, were also screened against five other races mentioned above in addition to race 77-5.
Screening the population for rust resistance
The seeds were sown in trays containing soil mixed with one fifth of farmyard manure. About 10-day-old seedlings were inoculated by spraying with hand sprayer and were incubated for 48 h in humid glass chambers. After incubation, the seedlings were shifted to greenhouse benches at temperatures ranging between 20 and 25 °C under ambient light and relative humidity conditions. Later, the individual seedlings were scored for the rust reaction (IT) at 12 days after inoculation, using the 0–4 scale as described by Stakman et al. (1962).
Genomic DNA isolation
The leaf samples were collected from 40 to 45-day-old plants of the above mentioned plant material. The fresh leaf samples were utilized for DNA isolation using the CTAB method (Murray and Thompson 1980) with minor modifications (10 % of CTAB was used in extraction buffer and CsCl density gradient was not measured). Purified DNA was checked for quality and quantity using agarose gel electrophoresis and diluted to a final concentration of 20 ng/μl for PCR analysis.
Molecular analysis
Since the gene Lr45 has been reported to be located on chromosome 2A, a parental polymorphism survey was conducted on parental lines Tc+Lr45 and Agra Local using a total of 92 SSR markers which were specific to chromosome 2A. Genomic SSR markers (comprising CFD, GWM, CFA, WMC, GDM and BARC series) were tested for polymorphism and used in bulked segregant analysis (BSA). The primer sequences of all the SSR markers used in this study were obtained from the GrainGenes website http://www.graingenes.pw.usda.gov. The SSR marker analyses were carried out in 10-µl reaction volumes containing 4 mM Tris- HCl (pH 8.0), 20 mM KCl, 0.8 mM MgCl2, 200 µM of each dNTP (MBI Fermentas, Germany), 1.0 unit Taq DNA polymerase (Bangalore Genei Pvt Ltd, India), 5 pM each of forward and reverse primers and 20 ng of genomic DNA. The polymerase chain reaction (PCR) was carried out in 96-well PCR plates with thermal seal in an Eppendorf thermal cycler (model Mastercycler pro S, Hamburg, Germany; www.eppendorf.com) with the following thermal profile: initial denaturation step of 94 °C for 4 min, followed by 45 cycles of 94 °C for 1 min (denaturation), 50–60 °C for 1 min (primer annealing) and 72 °C for 1 min (primer extension), with a final extension of 72 °C for 10 min. MetaPhor™ (Lonza) agarose gel (3.5 %) was used to resolve the amplified PCR products in 1X TBE buffer and they were visualized by ethidium bromide staining. Gel photographs were documented using Syngene G:Box Geldoc system (www.syngene.com).
Bulked segregant analysis
Bulked segregant analysis (Michelmore et al. 1991) was employed to identify SSR markers linked to the leaf rust resistance gene Lr45. The homozygous resistant and homozygous susceptible F2 plants identified based on F3 progeny testing were used for BSA. Equal amount of DNA from 10 homozygous resistant and 10 homozygous susceptible F2 plants was pooled to constitute the contrasting resistant and susceptible bulks, respectively. The resistant and susceptible bulks, along with parents, were tested with polymorphic SSR markers to identify putatively linked SSR markers.
Development of sequence-specific markers
To provide an easily detectable polymorphic marker, the most tightly linked co-dominant SSR marker gwm372 band was excised from agarose gel and purified using Qiagen Gel Extraction Kit. The purified PCR product was sequenced using the forward primer of gwm372 at Saf Lab Private Limited (www.saflabs.com) and Xcelris Private Limited (www.xcelrisgenomics.com) using the Sanger sequencing protocol. Later, these sequences were used to develop multiple sequence alignment (MSA) using BioEdit and ClustalW programs (http://www.ebi.ac.uk/tools/clustalw2/index.html). The MSA files were analyzed for the presence of deletions and a pair of 20-mer oligonucleotide primers was designed flanking the major deletion from within the sequence information of the marker fragment. Amplification of genomic DNA was done using all combinations of newly designed forward and reverse primers along with the primer pair of gwm372 at various annealing temperatures. The PCR reaction and amplified product separation was carried out as mentioned earlier.
Linkage analysis
The putative markers identified in BSA were used to genotype the entire F2 population to determine the number of recombinants produced by each marker, whereas, for linkage analysis involving both co-dominant and null allele dominant SSR markers, only homozygous resistant and homozygous susceptible F2 plants were used, as described by Gupta et al. (2006). Linkage analysis was done using the software MAPMAKER version 3.0 (Lander et al. 1987). The recombination frequencies were converted to map distances in centimorgans using the option of Kosambi’s function in the software.
Statistical analysis
The segregation ratios in F2 and BC1F1 plants scored as resistant and susceptible were subjected to Chi-squared (χ 2) analysis to test the goodness of fit to the theoretically expected Mendelian segregation ratios. In F3 analysis, the number of segregating and non-segregating families were also subjected to Chi-square test to confirm the F2 genetic ratio, using χ 2 = ∑(O − E)2/E, where O = observed number of individuals and E = expected number of individuals.
Results
Genetics of leaf rust resistance gene Lr45
The parental plants of Tc+Lr45 were distinguishable by resistance characterized by zero fleck (IT 0;) seedling reaction from susceptible response (IT 33+ to 4) of Agra Local and NI5439 against the leaf rust race 77-5. The rust reaction on F1 plants of the crosses Agra Local × Tc+Lr45 and NI5439 × Tc+Lr45 also exhibited IT 0; indicating the dominant nature of the gene Lr45. Out of the 323 F2 seedlings of the cross Agra Local × Tc+Lr45, 251 seedlings were resistant and 72 susceptible. Similarly, in the population NI5439 × Tc+Lr45, out of 335 F2 seedlings 260 showed resistant reaction and 75 seedlings were susceptible (Table 1). The segregation showed a goodness of fit to a 3:1 resistant:susceptible ratio with a P value of 0.2608 at χ 23:1 = 1.26 in the cross Agra Local × Tc+Lr45, whereas in the cross NI5439 × Tc+Lr45 a χ 23:1 value of 1.22 was obtained with P-value of 0.2505. This indicated that the resistance conferred by Lr45 is dominant and monogenic. The F3 segregation ratio of 1:2:1 (non-segregating resistant:segregating resistant:non-segregating susceptible) validated the results of F2 analysis (Table 1).
To confirm the results obtained in both the crosses against race 77-5, the F2 and BC1F1 populations of the cross Agra Local × Tc+Lr45 were also screened against five other races, 12-1, 162, 104-2, 107 and 108. For all the races tested on F2 and BC1F1 populations, monogenic ratios of 3:1 and 1:1, respectively, were obtained, confirming the segregation of a single dominant gene for resistance (Table 1).
Identification of SSR markers linked to Lr45
Out of 92 SSR markers specific to chromosome 2A, 34 were polymorphic between the parents Agra Local and Tc+Lr45. However, only seven markers located on the short arm were found putatively associated with Lr45 in the bulked segregant analysis (Table 2). Four markers, namely gwm372, gwm122, gwm275 and gpw3167, behaved as co-dominant, while three markers, namely cfd168, cfd6 and gwm249, were dominant, amplifying the marker fragment only in susceptible plants. The seven SSR markers differentiating the two contrasting bulks were screened for the respective marker fragment on the entire F2 population. In the case of co-dominant SSR markers, the lowest number of recombinants was observed for gwm372 (6/323) (Supplementary Table S1). Of the three null allele markers, cfd168 showed only one recombinant out of 323 F2 plants tested (Supplementary Table S2). Both the gene and the markers in combination deviated significantly from the expected independent assortment ratio, suggesting that the two loci involved were tightly linked to each other. The χ 2 test for linkage also confirmed that the leaf rust resistance gene Lr45 and the other markers were linked (P = 0.00 in each case) (Supplementary Table S1 and Table S2).
Development of genetic linkage map of Lr45 locus
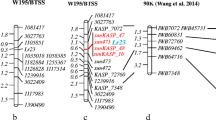
Three-point and multi-point analysis using MAPMAKER on 70 homozygous resistant and 70 homozygous susceptible plants fixed the seven SSR markers within a distance of 13.1 cM from the gene Lr45 (Fig. 1). No recombinant was observed for the marker cfd168 in a mapping population of 140 homozygous individuals and it was placed on the gene Lr45 with a logarithm of odds (LOD) score of 84.5. The closest co-dominant marker gwm372 was located at 0.7 cM distant from the gene Lr45 with a LOD score of 80.32. A linkage map of Lr45 was synthesized, where all the markers linked to the gene were aligned towards one side of the locus Lr45 (Fig. 1).
Validation of the SSR markers
SSR markers linked to the gene Lr45 in the F2 population Agra Local × Tc+Lr45 were validated on 92 plants of the second F2 population derived from the cross NI5439 × Tc+Lr45. One of the markers gwm122, however, was found monomorphic between NI5439 and Tc+Lr45. The polymorphic fragments produced by the rest of the markers were of the same size in NI5439 as found in the susceptible parent Agra Local, except for the marker gwm372. These five SSR markers were tested on 92 F2 plants of the cross NI5439 × Tc+Lr45. The minimum number of recombinants was observed for null allele marker cfd168 (1/92), followed by cfd6 (6/92) and gwm249 (7/92). However, co-dominant markers gwm275 (8/90) and gpw3167 (11/92) showed relatively higher frequencies of recombination between marker loci and Lr45. The marker gwm372 was found to be the closest co-dominant marker linked to Lr45, producing a fragment of 285 bp in Agra Local and 307 bp in Tc+Lr45. However, the same marker amplified a fragment of approximately 311 bp in NI5439, thus making it difficult to distinguish it from the Tc+Lr45 fragment on a MetaPhor agarose gel (Supplementary Fig. S3). Hence, it was difficult to differentiate the alleles produced by gwm372 in the parents Tc+Lr45 and NI5439.
Validation of SSR markers linked to Lr45 on backcross lines of wheat cultivars carrying Lr45 was also carried out. However, the marker gwm372 failed to distinguish four backcross lines of Lr45 from their respective recurrent parents, HD2329, Kalyansona, NI5439 and WH147 (Supplementary Fig. S4). The specificity of these SSR markers to the gene Lr45 was confirmed on a set of NILs in Thatcher background carrying 46 different leaf rust resistance genes derived from alien and native wheat germplasm. Five SSR markers, cfd168, cfd6, gwm249, gwm275 and gpw3167, amplified the expected marker fragments only in Tc+Lr45. However, the marker gwm372 failed to distinguish the NILs carrying different Lr genes from Lr45 (Supplementary Fig. S5).
Development of sequence-specific markers and validation
The marker fragment amplified by gwm372 in Tc+Lr45, Agra Local and NI5439 was sequenced. Later, the multiple sequence alignment showed the presence of a large unique region of 185 bp adjacent to the simple sequence repeats in the resistant parent Tc+Lr45 (Fig. 2). A common deletion of 32 bp was found in both the susceptible parents Agra Local and NI5439, while an additional 26 bp deletion was specific to Agra Local in the unique flanking region. These deletions resulted in flanking regions of 165 and 141 bp in NI5439 and Agra Local, respectively. Marker G372 94 was designed by a pair of 20-mer oligonucleotide primers (forward GACGTTAGCTCACGCAACCG, reverse CTCTTGAAACACAAAGCACA) flanking the 32-bp deletion region which amplified a fragment of 94 bp in the resistant parent Tc+Lr45 lacking the deletion region and a fragment of 62 bp in both the susceptible parents Agra Local and NI5439. Another marker G372 185 , developed by combining the reverse primer of G372 94 along with the forward primer of original marker gwm372, amplified the whole unique region of 185 bp adjacent to the repeat region in Tc+Lr45. It produced the expected amplicon of 153 bp in the parent NI5439 and a fragment of 127 bp in Agra Local which carries two deletions of 32 bp and 26 bp (Fig. 3).
Multiple sequence alignment using nucleotide sequence of marker gwm372 from Tc+Lr45, Agra Local and NI5439 indicating the forward and reverse primers of G372 94 . Lr45-a, AL-a and NI-a are sequencing results of Tc+Lr45, Agra Local and NI5439, respectively, from Saf Lab Pvt Ltd and Lr45-b, AL-b and NI–b are sequencing results of Tc+Lr45, Agra Local and NI5439, respectively, from Xcelris Pvt Ltd
When the two newly developed markers G372 94 and G372 185 were tested on 92 F2 individuals of the cross NI5439 × Tc+Lr45, genotyping results for both the primer pairs were found to be same as that of gwm372 (Supplementary Table S1). They were also validated on backcross lines of Lr45 in Indian wheat cultivars (Supplementary Fig. S6) and also on NILs carrying different leaf rust resistance genes, mostly in Thatcher background (Supplementary Fig. S7). Results showed that markers G372 94 and G372 185 produced 94-bp and 185-bp alleles, respectively, only in the backcross lines carrying Lr45 and not in the original recurrent parents or NILs of Thatcher carrying alien or native leaf rust resistance genes.
Discussion
A large number of alien rust resistance genes have been transferred into wheat (Friebe et al. 1996) but many of them have not been used in commercial cultivars. S. cereale-derived linked rust resistance genes Lr26/Sr31/Yr9 were deployed globally, though the wheat–rye translocation 1BL.1RS carrying these genes involved the entire short arm of the rye chromosome (Zeller 1973). Thus, in some cases a large size of alien segment may not necessarily have an adverse effect. The structure of Lr45 carrying translocation T2AS-2RS.2RL suggests that the translocation is a compensating type involving homoeologous chromosomes from group 2 only. However, the effect of this translocation on various agro-morphological traits needs to be studied. Genetic analysis of Lr45 in F2, F3 and BC1F1 generations of two crosses involving susceptible cultivars Agra Local and NI5439 against the most virulent race 77-5 showed segregation for a single dominant gene. The present findings are in contrast to the earlier report (McIntosh et al. 1995) that gametes carrying Lr45 had poor transmission resulting in segregation distortion as more susceptible plants were recorded in the F2 generation. However, in the present study, the monogenic inheritance of Lr45 was also confirmed against five other races of leaf rust, viz. 12-1, 162, 104-2, 107 and 108. Thus, like other alien genes, the S. cereale-derived resistance gene (Lr45) may also prove useful in wheat breeding.
Lr45-based resistance has not been utilized in marker-assisted selection because of the non-availability of markers. In the present study the leaf rust resistance gene Lr45 was mapped using seven SSR markers spanning a distance of 13.1 cM (Fig. 1). The marker order is similar to the high-density SSR consensus map developed by Somers (2004), with one exception, that cfd6 mapped proximal to gwm372 on the map by Somers (2004), whereas cfd6 was located distal to gwm372 in our map. Even though the SSR markers are expected to generally inherit as co-dominant loci, in the present study three SSR markers inherited as dominant loci amplifying only in susceptible plants and showing null allele in the resistant plants. The T2AS-2RS.2RL translocation, being alien to the T. aestivum genomic region, could have caused disturbances in the primer annealing sites of these SSR primers at the breakage point, resulting in the null alleles as reported in other alien gene-transferred wheat lines (Liu et al.2002; Brown-Guerdira et al. 2003; Malik et al. 2003; Adhikari et al. 2004; Vikal et al. 2004; Gupta et al. 2006; Singh et al. 2011).
The nearest SSR marker, cfd168, behaved as a dominant marker amplifying an allele specific to the susceptible parent. Although it produced one recombinant out of 323 F2 plants, no recombinant was identified in 140 homozygous individuals used for mapping analysis. Another marker, gwm372, behaved as co-dominant, but the allele produced in susceptible parent NI5439 was difficult to distinguish from the allele linked to Lr45. Hence, an effort was made to convert the less polymorphic gwm372 into a clearly distinguishable polymorphic marker by sequencing the PCR product, and designing the primers from the unique adjacent regions of the repeat sequences. One of the markers designed, G372 94 , produced a fragment difference of 32 bp between the resistant gentoype and all the genotypes devoid of the gene Lr45. Another primer marker G372 185 amplified the complete adjacent unique region of 185 bp in Tc+Lr45. The genetic map of Lr45 involving six molecular markers on the 40 homozygous resistant and susceptible F2 plants of the population NI5439 × Tc+Lr45 covered a distance of 10.1 cM (Fig. 1), and was almost similar to the genetic map of Agra Local × Tc+Lr45. The order of the markers remained the same and the difference found between the two genetic maps was very small, which shows that tight linkage existed between the marker and the gene Lr45. This small difference between the two maps may be because of the difference in the size of the F2 populations.
Marker validation, a process of examining the behavior of markers and the associated polymorphism, is necessary before applying a marker in breeding (Langridge and Chalmers 1998). Gupta et al. (1999) suggested that the validity of a molecular marker linked with any trait should be examined in crosses other than the ones in which the marker was developed. One of the SSR markers, gwm122, turned out to be monomorphic in the second F2 population used for validation. The validation study conducted by Sharp et al. (2001) showed that the STS marker developed by Naik et al. (1998) linked to Lr28 was ineffective in marker-assisted breeding due to the poor linkage of marker to gene. A SCAR marker developed for leaf rust resistance gene Lr24 by Schachermayr et al. (1995) and Dedryver et al. (1996) was found to be monomorphic in Indian sources, as they have a shortened alien segment relative to the original source, Agent, and have therefore been found to be ineffective in marker-assisted selection (Prabhu et al. 2004).
Both the sequence-specific markers developed were tested on two different sources that were used for validation of other linked markers. When validated on the 12 resistant backcross lines carrying Lr45 and their susceptible recurrent parents, the allele specific to Tc+Lr45, i.e. 94 bp in the case of G372 94 and 185 bp in the case of G372 185 , was amplified in all the 12 NILs carrying gene Lr45. In 12 recurrent parental genotypes, an allele of 64 bp was produced by the marker G372 94 . In the case of G372 185 , recurrent parents HD2329, Kalyansona, NI5439 and WH147 produced the allele of 153 bp and the rest produced the allele of 127 bp. Although the marker G372 185 produced two distinct alleles in susceptible recurrent parents, the allele size is sufficient to differentiate them easily from the allele of the resistant parent, indicating that both the sequence-specific markers are highly specific to gene Lr45 and can be effectively used for the selection of this gene in different genetic backgrounds. These markers were also validated for their uniqueness and specificity to Lr45, because they amplified the resistance allele only in the Thatcher NIL carrying gene Lr45 and not in the Thatcher NILs carrying other 46 alien or native Lr genes in wheat.
Molecular markers can predict the presence of a specific gene with very high probability without the need for disease evaluation, and thus aid the transfer of several resistance genes into adapted materials to pyramid genes in one plant. The markers reported in this study can be effectively utilized for the selection of this gene in different genetic backgrounds, as indicated by successful validation on divergent Lr genes and wheat cultivars. The newly designed co-dominant markers G372 94 and G372 185 were able to identify the heterozygotes, and would serve as an important tool to rapidly transfer this gene into other wheat cultivars.
References
Adhikari TB, Yang X, Cavaletto JR, Hu X, Buechley G, Ohm HW, Shaner G, Goodwin SB (2004) Molecular mapping of Stb1, a potential durable gene for resistance to Septoriatritici blotch in wheat. Theor Appl Genet 109:944–953
Bhardwaj SC, Prashar M, Jain SK, Kumar S, Sharma YP (2010) Physiologic specialization of Puccinia triticina on wheat (Triticum species) in India. Indian J Agric Sci 80:805–811
Brown-Guerdira GL, Singh S, Fritz AK (2003) Performance and mapping of leaf rust resistance to wheat from Triticum timopheevii sub spp. armeniacum. Phytopathology 93:784–789
Dedryver F, Jubier MF, Thouvenin J, Goyeau H (1996) Molecular markers linked to the leaf rust resistance gene Lr24 in different wheat cultivars. Genome 39:830–835
Fein YH, Yan L, Gang GS, Xiang YW, Qun LD (2009) A SCAR marker for leaf rust resistance gene Lr45. Sci Agric Sin 42:124–129
Friebe B, Jiang J, Raupp WJ, McIntosh RA, Gill BS (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87
Gupta PK, Varshney RK, Sharma PC, Ramesh B (1999) Molecular markers and their applications in wheat breeding. Plant Breed 118:369–390
Gupta SK, Charpe A, Prabhu KV, Haque QMR (2006) Indentification and validation of molecularmarkers linked to the leaf rust resistance gene Lr19 in wheat. Theor Appl Genet 113:1027–1036
Herrera-Foessel SA, Singh RP, Huerta-Espino J, Rosewarne GM, Periyannan SK, Viccars L, Calvo-Salazar V, Lan C, Lagudah ES (2012) Lr68: a new gene conferring slow rusting resistance to leaf rust in wheat. Theor Appl Genet 124:1475–1486
Lander ES, Green P, Abrahamson J, Barlow A, Daly MJ, Lincoln SE, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Langridge P, Chalmers K (1998) Techniques for marker development. In: Slinkard AE (ed) Proceedings of the 9th international wheat genetics symposium, vol 1. University Extension Press, University of Saskatchewan, Saskatoon, Canada, pp 107–117
Liu XM, Smith CM, Gill BS (2002) Identification of microsatellite markers linked to Russian wheat aphid resistance genes Dn4 and Dn6. Theor Appl Genet 104:1042–1048
Malik R, Brown-Guedira GL, Smith CM, Harvey TL, Gill BS (2003) Genetic mapping of wheat curl mite resistance genes Cmc3 and Cmc4 in common wheat. Crop Sci 43:644–650
McIntosh RA, Friebe B, Jiang J, The D, Gill BS (1995) Cytogenetical studies in wheat XVI. Chromosome location of a new gene for resistance to leaf rust in a Japanese wheat-rye translocation line. Euphytica 82:141–147
McIntosh RA, Dubcovsky J, Rogers J, Morris C, AppelsR, Xia X (2011) Catalogue of gene symbols for wheat: 2011 supplement. http://www.shigen.nig.ac.jp/wheat/komugi/genes/macgene/supplement2011.pdf
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Mukade K, Kamio M, Hosoda K (1970) The transfer of leaf rust resistance from rye to wheat by intergeneric addition and translocation. Gamma field symposia no. 9. Mutagenesis in relation to ploidy level, pp 69–87
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight DNA. Nucleic Acids Res 8:4321–4325
Naik S, Gill KS, Rao VSP, Gupta VS, Tamhankar SA, Pujar S, Gill BS, Ranjekar PK (1998) Identification of a STS marker linked to an Aegilops speltoides-derived leaf rust resistance gene Lr28 in wheat. Theor Appl Genet 97:535–540
Prabhu KV, Gupta SK, Charpe A, Koul S (2004) SCAR marker tagged to the alien leaf rust resistance gene Lr19 uniquely marking the Agropyron elongatum-derived gene Lr24 in wheat: a revision. Plant Breed 123:417–420
Schachermayr G, Messmer MM, Feuillet C, Winzeler H, Winzeler M, Keller B (1995) Identification of molecular markers linked to the Agropyron elongatum-derived leaf rust resistance gene Lr24 in wheat. Theor Appl Genet 90:982–990
Sharp PJ, Johnston S, Brown G, McIntosh RA, Pallotta M, Carter M, Bariana HS, Khatkar S, Lagudah ES, Singh RP, Khairallah M, Potter R, Jones MGK (2001) Validation of molecular markers for wheat breeding. Aust J Agric Res 52:1357–1366
Singh A, Pallavi JK, Gupta P, Prabhu KV (2011) Identification of microsatellite markers linked to leaf rust resistance gene Lr25 in wheat. J Appl Genet 53(1):19–25. doi:10.1007/sl13353-011-0070-0
Singh D, Mohler V, Park RF (2012) Discovery, characterisation and mapping of wheat leaf rust resistance gene Lr71. Euphytica 190(1):131–136. doi:10.1007/s10681-012-0786-x
Somers DJ (2004) A high-density wheat microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Stakman EC, Stewart DM, Loegering WQ (1962) Identification of physiological races of Puccinia graminis var. tritici. USDA. ARS Bull. E617 (Revised), pp 53
Tomar SMS, Menon MK (2001) Genes for disease resistance in wheat. Indian Agricultural Research Institute, New Delhi, p 152
Tomar SMS, Singh SK, Sivasamy M, Vinod (2014) Wheat rusts in India: resistance breeding and gene deployment—a review. Indian J Genet 74:129–156
Vikal Y, Chhuneja P, Singh R, Dhaliwal HS (2004) Tagging of an Aegilops speltoides derived leaf rust resistance gene Lr28 with a microsatellite marker in wheat. J Plant Biochem Biotechnol 13:47–49
Zeller FJ (1973) 1B/1R wheat-rye chromosome substitutions and translocations. In: Sears ER, Sears LMS (eds) Proceedings of the fourth international wheat genetics symposium. Agricultural Experiment Station, University of Missouri, Columbia, Missouri, USA, pp 209–221
Zhang N, Yang WX, Yan H, Liu D, Chu D, Meng Q, Zhang T (2006) Molecular markers for leaf rust resistance gene Lr45 in wheat based on AFLP analysis. Agric Sci China 5:938–943
Acknowledgments
The senior author is grateful to Post Graduate School, Indian Agricultural Research Institute and the Indian Council of Agricultural Research, New Delhi, India for financial assistance given as SRF during the course of study. The authors are grateful to the Directorate of Wheat Research, Regional Station, Flowerdale, Shimla, India, for providing pure inoculum of leaf rust pathotypes and the Indian Agricultural Research Institute, New Delhi for facilitating the experiments.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Naik, B.K., Vinod, Sharma, J.B. et al. Molecular mapping and validation of the microsatellite markers linked to the Secale cereale-derived leaf rust resistance gene Lr45 in wheat. Mol Breeding 35, 61 (2015). https://doi.org/10.1007/s11032-015-0234-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-015-0234-4