Abstract
The spiro-oxindole derivatives were synthesized via a 1,3-dipolar cycloaddition approach and characterized by FT-IR, 1H, 13C NMR and mass spectral techniques. The single crystal XRD of 6d further validates the formation of compounds. DFT calculations indicated the reactive nature of compound 6d. Docking studies with 5LAW disclosed the minimum binding energy of − 10.83 kcal/mol for 6d. Furthermore, safe oral bioavailability was ensured by the physicochemical, pharmacokinetic, and toxicity predictions. The anticancer analysis of synthesized compounds showed substantial activity against A549 cells, notably with an IC50 value of 8.13 ± 0.66 µM for 6d compared to standard doxorubicin. 6d was also evaluated for cytotoxicity against L929 healthy cells and A549, showing selectivity towards A549 than healthy cells. AO/EB staining method showed early apoptotic cellular death in the A549 cell line in a dose-dependent manner.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
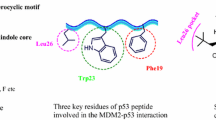
Lung cancer ranks as one among the most prevalent tumours contributing to mortality in humans. It is developed by the genetic changes in the normal healthy cells as a result of extrinsic stimuli such as smoking and air pollution [1, 2]. Two important molecular targets in lung carcinoma are EGFR (Epidermal Growth Factor Receptor) and ALK (Anaplastic Lymphoma Kinase). These targets play a significant role in progression and development of lung cancer. Targeting them with specific inhibitors has demonstrated substantial efficacy in treating patients suffering from Non-small cell lung cancer. However, due to resistance development, inadequate curative potential, high cost and side effects, new studies have explored targeting the interaction of MDM2-p53 as potential therapeutic strategy for lung cancer treatment. The cancerous growth suppressing protein p53, commonly referred as the “guardian of the genome,” is essential in preventing cancer. p53 protein functions as a factor that mediates transcription, managing the functioning of different genes involved in apoptosis DNA repair, senescence and controls cell cycle [3, 4]. When the DNA is damaged due to radiation, toxins or errors in DNA replication, p53 is activated and arrests the cell cycle. This interruption permits the cell to repair DNA damage before continuing the process of cell division. In case the DNA damage is significant and irreversible, p53 may trigger apoptosis in order to destroy the damaged cell. Cells with damaged DNA can replicate and multiply when p53 becomes inactive or flawed, raising the risk of tumor formation. p53 mutations are especially common in hostile and last stage cancer. MDM2 protein regulates the tumor suppressor protein p53. p53 and MDM2 interaction is critical for maintaining normal p53 levels and activity. MDM2 gene mutations or amplification could disturb this balance, resulting in elevated MDM2 levels and consequent p53 degradation. As a result, p53 tumor-suppressive actions are diminished and the probability of tumor growth increases [5,6,7,8,9,10]. The inhibition of the MDM2-p53 can significantly restore p53 functions and promotes tumor cell death and hence its interaction has been investigated as a potential therapeutic method for restoring p53 activity in cancer cells.
The spiro-oxindole ring system represents structural motifs of natural products with biological significance [11, 12]. Owing to their distinct structural characteristics and diverse chemical properties, they are attractive compounds for the development of novel pharmacological drugs. Spiro compounds, which are formed by fusing pyrrolidine and oxindole nuclei, have acquired prominence due to their intriguing biological activities such as anticancer [13,14,15], antifungal [16], antimalarial [17], antitubercular [18], antidiabetic [19], anti-inflammatory [20] and anti-HIV [21]. Some of the naturally occurring alkaloids bearing spiro ring system are shown in Fig. 1. In preclinical research, spiro-oxindole derivatives, such as MI-888 [22], MI-219 [23] and MI-43 [24] were employed as inhibitors of MDM2-p53 protein–protein interface and reactivation of p53 [25]. These compounds classically function by attaching to MDM2 hydrophobic pockets and preventing them from interacting with p53.
Computational chemistry adds substantial benefits to drug discovery by accelerating the process, guiding rational design, assisting with target identification and validation, forecasting ADMET properties and conserving both time and money. It has evolved into a necessary tool in the modern pharmaceutical business, aiding the identification and development of novel and effective therapeutic drugs [26,27,28,29,30,31,32,33,34].
The unique structural features of spiro-oxindoles allow them to bind effectively to MDM2-p53, which is crucial for effective inhibition. Most MDM2-p53 inhibitors often create resistance and mutations in the P53 pathway. The high selectivity of spiro-oxindoles with MDM2-p53 potentially reduces the development of resistance and addresses critical issues such as toxicity and poor pharmacokinetics. Hence, spiro-oxindole based MDM2-p53 inhibitors offer significant prospects in lung cancer treatment. The synthesis and purification of these spiro-oxindole derivatives were straightforward. More study is needed, though, to optimize their potency selectivity and pharmacokinetic features for clinical use. In considering the above aspects, novel spiro-oxindole compounds inhibiting MDM2-p53 protein–protein interaction have been developed. Spiro-oxindole derivatives were synthesized and then characterized via IR, Mass, 1H and 13C NMR spectral methods. These derivatives were examined for in vitro anticancer efficacy against lung cancer cells. Computational techniques such as Molecular Docking, DFT and ADMET parameters were utilized to investigate their optimized structures, effect on ADMET parameters and to validate experimental results.
Experimental
Materials
The chemicals used were procured from Sigma-Aldrich. Relitech melting point apparatus was used to determine Melting point. Elemental analyses (C, H, N) were performed in CHNS Elemental analyser-Perkin Elmer-2400 CHNS/O. FT-IR spectrum were recorded in SCHIMADZU Fourier Transformation Infra-Red spectrometer. NMR were analysed on Bruker Avance III HD Nanobay 400 MHz FT-NMR spectrometer by using DMSO-D6 and CDCl3 as solvent and TMS as standard. The LC-QTOF-HR Mass Spectrometer was employed to acquire mass spectra.
Synthesis
Synthesis of (2E)-1-([1,1′-biphenyl]-4-yl)-3-(4-aryl)prop-2-en-1-one (3a-3 h)
In an ethanolic medium, a proportionate amount of substituted aldehydes (1) and 4-acetyl biphenyl (2) were stirred in the presence of a catalytic dose of NaOH. At the end of the reaction, the mixture was transferred into crumpled ice and the coarse product was strained, parched, and recrystallized with ethanol.
Synthesis of 3′-([1,1′-biphenyl]-4yl)carbonyl-5′-(2-methylpropyl)4′-(aryl)spiro [indole-3,2′-pyrrolidin]-2(1H)-one (6a-6 h)
An equimolar mixture of chalcone (0.001 mol, 3a-3 h), leucine (0.001 mol, 4) and isatin (0.001 mol, 5) were dissolved in absolute ethanol and refluxed for about 5 h. TLC was utilised to observe the successful completion of the reaction, the reaction mixture was allowed to cool. Then the resultant crude product was retrieved by evaporation and recrystallized in ethanol to obtain the pure compound.
3′-([1,1′-biphenyl]-4yl)carbonyl-5′-(2-methylpropyl)4′-(phenyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6a)
White solid; Yield: 80.2%; M.P 210 °C; MF: C34H32N2O2; Elemental analysis: Cal (%) C 81.57; H 6.44; N 5.60; O 6.39%; found: C 81.55; H 6.42; N 5.57; O 6.40%; IR 1666.49 (CONH), 1710.50 (C=O), 2870.04 (Ar–H), 3124.62, 3271.25 (NH) (cm−1); 1H NMR (400 MHz) (CDCl3/TMS): §, 0.83 (d, 6H, (CH3)2, J = 7.6 Hz), 1.47 (t, 2H, CH2, J = 11.2 Hz), 1.58–1.67 (m, 1H, C–H), 2.03 (bs, 1H, NH), 3.05–3.11 (m, 1H, C-5′), 3.79 (t, 1H, C-4′, J = 10.4 Hz), 4.66 (d, 1H, C-3′, J = 10.8 Hz), 6.48–7.53 (m, 18H, Ar–H), 10.24 (s, 1H, NH) (ppm); 13C NMR (400 MHz) (CDCl3/TMS): §, 21.86, 23.66, 25.93 (Isopropyl C’s), 43.68 (C-4′), 55.89 (C-3′), 63.55 (Spiro C), 69.44 (C-5′), 123.04–145.21 (Ar–C), 181.87 (Amide C=O), 197.07 (C=O) (ppm); LC–MS (m/z) 501.26 [M + H]+
3′-([1,1′-biphenyl]-4yl)carbonyl-4′-(4-fluorophenyl)-5′-(2-methylpropyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6b)
White solid; Yield: 77.6%; M.P 212 °C; MF: C34H31FN2O2; Elemental analysis: Cal (%) C 78.74; H 6.02; N 5.40; O 6.17%; found: C 78.71; H 6.03; N 5.42; O 6.15%; IR 1667.39 (CONH), 1704.64 (C=O), 2980.02 (Ar–H), 3149.75, 3277.06 (NH) (cm−1); 1H NMR (400 MHz) (DMSO-d6/TMS): §, 0.84 (d, 6H, (CH3)2, J = 7.6 Hz), 1.46 (t, 2H, CH2, J = 11.2 Hz), 1.60–1.71 (m, 1H, C–H), 2.05 (s, 1H, NH), 3.09–3.17 (m, 1H, C-5′), 3.81 (t, 1H, C-4′, J = 10.4 Hz), 4.68 (d, 1H, C-3′, J = 10.8 Hz), 7.02–7.55 (m, 17H, Ar–H), 10.36 (s, 1H, NH) (ppm); 13C NMR (400 MHz) (DMSO-d6/TMS): §, 21.90, 23.85, 25.45 (Isopropyl C’s), 42.52 (C-4′), 54.76 (C-3′), 63.22 (Spiro C), 69.03 (C-5′), 120.06–144.38 (Ar–C), 181.54 (Amide C=O), 196.97 (C=O) (ppm); LC–MS (m/z) 519.20 [M + H]+
3′-([1,1′-biphenyl]-4yl)carbonyl-4′-(4-chlorophenyl)-5′-(2-methylpropyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6c)
Pale yellow solid; Yield: 79.1%; M.P 222 °C; MF: C34H31ClN2O2; Elemental analysis: Cal (%) C 76.32; H 5.84; N 5.24; O 5.98%; found: C 76.33; H 5.80; N 5.21; O 6.01%; IR 1662.64 (CONH), 1707.03 (C=O), 2924.09 (Ar–H), 3169.04, 3271.27 (NH) (cm−1); 1H NMR (400 MHz) (DMSO-d6/TMS): §, 0.79 (d, 6H, (CH3)2, J = 7.6 Hz), 1.18 (t, 2H, CH2, J = 10.8 Hz), 1.47–1.54 (m, 1H, C–H), 1.71 (s, 1H, NH), 3.25–3.32 (m, 1H, C-5′), 3.72 (t, 1H, C-4′, J = 10.4 Hz), 4.51 (d, 1H, C-3′, J = 10.4 Hz), 6.49–7.64 (m, 17H, Ar–H), 10.45 (s, 1H, NH) (ppm); 13C NMR (400 MHz) (DMSO-d6/TMS): §, 21.79, 23.74, 25.34 (Isopropyl C’s), 42.41(C-4′), 54.65 (C-3′), 63.11 (Spiro C), 68.92 (C-5′), 121.50–144.27 (Ar–C), 181.43 (Amide C=O), 196.86 (C=O) (ppm); LC–MS (m/z) 535.26 [M + H]+
3′-([1,1′-biphenyl]-4yl)carbonyl-4′-(4-bromophenyl)-5′-(2-methylpropyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6d)
White solid; Yield: 78.9%; M.P 228 °C; MF: C34H31BrN2O2; Elemental analysis: Cal (%) C 70.47; H 5.39; N 4.83; O 5.52%; found: C 70.44; H 5.38; N 4.81; O 5.55%; IR 1672.95 (C=N), 1708.93 (C=O), 2953.04 (Ar–H), 3095.71, 3284.76 (NH) (cm−1); 1H NMR (400 MHz) (DMSO-d6/TMS): §, 0.79 (d, 6H, (CH3)2, J = 7.6 Hz), 1.16 (t, 2H, CH2, J = 11.2 Hz), 1.45–1.52 (m, 1H, C–H), 1.68 (s, 1H, NH), 3.12–3.19 (m, 1H, C-5′), 3.67 (t, 1H, C-4′, J = 10.4 Hz), 4.49 (d, 1H, C-3′, J = 10.4 Hz), 6.47–7.62 (m, 17H, Ar–H), 10.43 (s, 1H, NH) (ppm); 13C NMR (400 MHz) (DMSO-d6/TMS): §, 22.08, 24.04, 25.64 (Isopropyl C’s), 42.71 (C-4′), 54.95 (C-3′), 63.41 (Spiro C), 69.22 (C-5′), 120.25–144.57 (Ar–C), 181.73 (Amide C=O), 197.16 (C=O) (ppm); LC–MS (m/z) 579.13 [M + H]+
3′-([1,1′-biphenyl]-4yl)carbonyl-4′-(4-methoxyphenyl)-5′-(2-methylpropyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6e)
White solid; Yield: 75.1%; M.P 219 °C; MF: C35H34N2O3; Elemental analysis: Cal (%) C 79.22; H 6.46; N 5.28; O 9.05%; found: 79.24; H 6.43; N 5.25; O 9.03%; IR 1668.41 (CONH), 1709.83 (C=O), 2881.69 (Ar–H), 3116.94, 3238.41 (NH) (cm−1); 1H NMR (400 MHz) (DMSO-d6/TMS): §, 0.78 (d, 6H, (CH3)2, J = 7.6 Hz), 1.27 (t, 2H, CH2, J = 11.2 Hz), 1.40–1.47 (m, 1H, C–H), 1.69 (s, 1H, NH), 2.98–3.04 (m, 1H, C-5′), 3.59 (t, 1H, C-4′, J = 10.4 Hz), 3.70 (s, 3H, OCH3), 4.44 (d, 1H, C-3′, J = 10.4 Hz), 6.88–7.62 (m, 17H, Ar–H), 10.38 (s, 1H, NH) (ppm); 13C NMR (400 MHz) (DMSO-d6/TMS): §, 22.16, 24.14, 25.68 (Isopropyl C’s), 42.91 (C-4′), 54.90 (C-3′), 55.39 (OCH3), 63.46 (Spiro C), 69.13 (C-5′), 121.79–158.47 (Ar–C), 181.88 (Amide C=O), 197.37 (C=O) (ppm); LC–MS (m/z) 531.22 [M + H]+
3′-([1,1′-biphenyl]-4yl)carbonyl-5′-(2-methylpropyl)4′-(4-thiomethylphenyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6f)
White solid; Yield: 72.5%; M.P 210 °C; MF: C35H34N2O2S; Elemental analysis: Cal (%) C 76.89; H 6.27; N 5.12; O 5.85%; found: C 76.91; H 6.24; N 5.09; O 5.88%; IR 1671.20 (CONH), 1710.85 (C=O), 2997.32 (Ar–H), 3112.35, 3234.61 (NH) (cm−1); 1H NMR (400 MHz) (DMSO-d6/TMS): §, 0.79 (d, 6H, (CH3)2, J = 7.6 Hz), 1.16 (t, 2H, CH2, J = 11.2 Hz), 1.42–1.48 (m, 1H, C–H), 1.61 (s, 1H, NH), 2.42 (s, 3H, SCH3), 3.01–3.09 (m, 1H, C-5′), 3.60 (t, 1H, C-4′, J = 10.4 Hz), 4.45 (d, 1H, C-3′, J = 10.4 Hz), 6.44–7.62 (m, 17H, Ar–H), 10.37 (s, 1H, NH) (ppm); 13C NMR (400 MHz) (DMSO-d6/TMS): §, 15.06 (SCH3), 22.11, 24.10, 25.65 (Isopropyl C’s), 42.81 (C-4′), 55.10 (C-3′), 63.39 (Spiro C), 69.17 (C-5′), 121.78–144.55 (Ar–C), 181.78 (Amide C=O), 197.25 (C=O) (ppm); LC–MS (m/z) 547.26 [M + H]+
3′-([1,1′-biphenyl]-4yl)carbonyl-4′-(4-N-dimethylphenyl)-5′-(2-methylpropyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6 g)
Yellow solid; Yield: 70.3%; M.P 224 °C; MF: C36H37N3O2; Elemental analysis: Cal (%) C 79.53; H 6.86; N 7.73; O 5.89%; found: C 79.54; H 6.84; N 7.69; O 5.92%; IR 1661.06 (CONH), 1709.64 (C=O), 2895.16 (Ar–H), 3159.02, 3278.91 (NH) (cm−1); 1H NMR (400 MHz) (DMSO-d6/TMS): §,0.79 (d, 6H, (CH3)2, J = 7.6 Hz), 1.23 (t, 2H, CH2, J = 11.2 Hz), 1.38–1.45 (m, 1H, C–H), 1.63 (s, 1H, NH), 2.82 (s, 6H, N(CH3)2), 3.04–3.13 (m, 1H, C-5′), 3.63 (t, 1H, C-4′, J = 10.4 Hz), 4.45 (d, 1H, C-3′, J = 10.8 Hz), 6.48–7.62 (m, 17H, Ar–H), 10.36 (s, 1H, NH) (ppm); 13C NMR (400 MHz) (DMSO-d6/TMS): §, 22.15, 24.17, 25.70 (Isopropyl C’s), 43.01 (C-4′), 54.99 (C-3′), 63.36 (Spiro C), 69.09 (C-5′), 121.76–149.69 (Ar–C), 181.93 (Amide C=O), 197.45 (C=O) (ppm); LC–MS (m/z) 544.31 [M + H]+
3′-([1,1′-biphenyl]-4yl)carbonyl-5′-(2-methylpropyl)4′-(3,4,5-trimethoxyphenyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6 h)
Reddish brown solid; Yield: 68.9%; M.P 216 °C; MF: C37H38N2O5; Elemental analysis: Cal (%) C 75.23; H 6.48; N 4.74; O 13.54%; found: C 75.20; H 6.45; N 4.71; O 13.52%; IR 1658.78 (CONH), 1709.78 (C=O), 2885.56 (Ar–H), 3159.41, 3251.98 (NH) (cm−1); 1H NMR (400 MHz) (DMSO-d6/TMS): §, 0.81 (d, 6H, (CH3)2, J = 7.6 Hz), 1.23 (t, 2H, CH2, J = 11.2 Hz), 1.43–1.49 (m, 1H, C–H), 1.69 (s, 1H, NH), 3.11–3.17 (m, 1H, C-5′), 3.61 (s, 3H, p–OCH3), 3.79 (s, 6H, m–OCH3), 3.92 (t, 1H, C-4′, J = 10.4 Hz), 4.48 (d, 1H, C-3′, J = 10.4 Hz), 6.45–7.62 (m, 17H, Ar–H), 10.35 (s, 1H, NH) (ppm); 13C NMR (400 MHz) (DMSO-d6/TMS): §, 22.14, 24.12, 25.72 (Isopropyl C’s), 42.93 (C-4′), 55.69, 56.33 (OCH3), 63.27 (Spiro C), 69.30 (C-5′), 121.74–153.31 (Ar–C), 181.70 (Amide C=O), 197.43 (C=O) (ppm); LC–MS (m/z) 591.31 [M + H]+
X-ray crystallographic studies
The colourless single crystal of compound 6d was acquired by sluggish evaporation from ethanol at ambient temperature. Data was collected on Bruker Kappa APEXII. Bruker SAINT (APEXII) was used for cell refinement and data reduction. The structure of 6d was solved through SHELXTL [35, 36]. The crystallographic information was uploaded at the Cambridge Crystallographic Data Centre under the depository number CCDC 2279234.
Computational and theoretical studies
Hirshfeld surface studies
Hirshfeld surfaces [37] and related 2D-fingerprint plots [38] have been used to investigate interactions in crystal structure via CrystalExplorer17.5 software [39, 40]. The crystallographic information file of 6d acquired from single crystal XRD was utilized as an input file to generate these interactions.
DFT studies
The crystal structure of compound 6d was selected for quantum chemical calculations. The CIF file from single crystal XRD was used as an input file for structure optimization in Gaussian 09 W software package [41]. GaussView 05 was used to view the findings of the computations. From the optimized structure of 6d molecular electrostatic potential, quantum chemical parameters, frontier molecular orbitals and Mulliken charges were calculated using Becke’s three-parameter hybrid function (B3) [42] and Lee–Yang–Parr (LYP) [43] in a basis set of 6–31 + G(d,p).
Molecular docking
The molecular docking was performed with target protein inhibitors of MDM2-p53 interaction saved from (www.rcsb.org) with protein ID: 5law. AutoDock tools 1.5.6 [44] was used to dock the compounds with target protein 5law. The structures of the ligands were generated using Chemsketch software [45] and saved as.mol files, which were then transformed to.pdb files using Open Babel software [46]. The protein molecule was optimized by removing HETATM. The energy minimization was carried out using Swiss-PDB Viewer [47]. The.pdbqt file of protein was generated by removing water molecules, introducing Kollman charges and hydrogen, while the.pdbqt file of ligand was generated by choosing a torsion tree from AutoDock. The Lamarckian Genetic Algorithm [48] was used to determine the optimum docking poses, which were then visualized using the Biovia Discovery Studio visualizer [49].
ADMET predictions
The in silico ADMET properties like physicochemical, pharmacokinetics and toxicity studies have been evaluated using online SwissADME [50] and ADMETlab 2.0 [51] software’s.
Biological studies
Cell culture and cell viability assay
A549 cells were chosen to evaluate in vitro cytotoxicity. The cells have been nurtured in DMEM media with 10% fetal bovine serum (FBS). A haemocytometer was used to gauge the trypsinized cell lines. 10,000 cells were seeded in each well and nurtured in 5% CO2 atmosphere at 37 °C for 24 h. Cell morphology was scrutinized at the termination of the cultivation period. The culture media in each well was replaced by sterilized DMEM medium. Compounds (6a-6 h) at varied concentrations were distributed onto the cells in triplicate and retained in a CO2 environment at 37 °C. Later, MTT (1 mg/mL) was injected into each well and the nurture period was extended for another 4–6 h. Subsequently, MTT were removed from each well, at the end of the incubation phase. The crystals of formazon were then dissolved in 100% DMSO and a photometer measurement at 570 nm was taken.
AO/EB nuclear staining
A medium containing MEM with 10% FBS was utilised to cultivate the A549 human lung cancer cells. To separate the cells, an appropriate amount of versene (0.1%) and Trypsin (0.25%)/EDTA (0.02%) were employed, while the cells were observed under the microscope to ensure complete detachment. Approximately 105 cells were seeded and cultured for 24 h at 37 °C. Specimens were supplied in triplicate at varied concentrations, while control cells were cultivated for 24 h without any samples. After incubation, the media was entirely removed from the plate surfaces and rinsed with PBS. An inverted phase-contrast microscope was employed to scrutinize the dye AO-EtBr (1 g/mL) in each well.
LDH assay
The A549 cell lines were cultivated in precisely the same manner as the MTT assay. Compound 6d was treated with 1 mL of buffer solution (lithium lactate in NaOH) and 0.1 mL of media, this combination was added to a set of test tubes and was gestated at 37 °C for 30 min. The incubation proceeded for an extra 30 min after injecting 0.2 mL of NAD solution. Upon arresting the reaction with 1.0 mL of DNPH mixture, the tubes were allowed to incubate for about 15 min at 37 °C. To each tube, 3.5 mL of 0.4 N NaOH was added. The colour produced was assessed using a spectrophotometer at 420 nm.
Statistical analysis
The results of anticancer activity were given as mean ± standard deviation (SD) with P < 0.05 value which is regarded as statistically significant.
Result and discussion
The current investigation includes the synthesis of various 3′-([1,1′-biphenyl]-4yl)carbonyl-5′-(2-methylpropyl)-4′-(aryl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one (6a-6 h) using substituted chalcone, leucine and isatin via two-step method as exposed in the Scheme 1. Solvent optimization for this three component reaction was investigated by the reaction of an equimolar mixture of chalcone (0.001 mol, 3a), leucine (0.001 mol, 4) and isatin (0.001 mol, 5) under reflux in different solvents such as toluene, acetonitrile, ethanol, methanol and methanol:water (1:1) which afforded highly functionalized 3′-([1,1′-biphenyl]-4yl)carbonyl-5′-(2-methylpropyl)4′-(phenyl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one 6a. Excellent yields were obtained when ethanol was used as a solvent and only moderate yield was observed with acetonitrile, methanol and methanol:water.
A plausible mechanism for the regioselectivity of the reaction is explained by the addition of the electron rich carbon of the dipole or azomethine ylide 4ʹ (decarboxylative condensation of the isatin 5 with leucine 4) to the electron-deficient β carbon of chalcone (dipolarophile) 3a (Track-1) which is more encouraging due to the presence of secondary orbital interaction (SOI) which is not in Track-2 (Scheme 2). Even though four new stereocenters have been generated, the stereoselectivity of the reaction is impressive with the formation of a single diastereomer. The only byproducts of this reaction were water and carbon dioxide, hence leading to a high atom economy.
The structure of all newly synthesized spiro-oxindole analogues 6a-6 h was fully characterized by elemental analysis, FTIR, Mass and NMR spectral studies (Supplemental data Figs. S1–S40). The FTIR spectra for the spiro-oxindole analogues showed stretching vibration for CONH at 1658.78–1672.95 cm−1 and C=O vibrations were observed at 1704.64–1710.85 cm−1. The vibrations around 2870.04–2997.32 cm−1 were ascribed to Ar–H. Broad peaks at 3095.71–3169.04 cm−1 and 3234.61–3284.76 cm−1 correspond to N–H stretching vibrations.
In 1H-NMR spectrum, six protons of (CH3)2 groups of the leucine resonated as a doublet at 0.79–0.84 ppm, a triplet and a multiplet peaks observed at 1.16–1.47 ppm and 1.38–1.71 ppm corresponds to CH2 and CH protons of the amino acid, respectively. A singlet at 1.61–2.05 ppm attributed to NH proton. C-5′ proton resonated as multiplet in the range of 2.98–3.32 ppm. Similarly, C-4′ and C-3′ protons appeared as triplet and doublet at 3.59–3.81 ppm and 4.44–4.68 ppm, respectively. The aromatic proton signals were observed at 6.44–7.64 ppm. The singlet peaks observed between 10.24–10.48 ppm correspond to NH proton. In 13C-NMR spectrum, isopropyl carbons were observed at 21.79–22.16 ppm, 23.66–24.17 ppm and 25.34–25.93 ppm. A peak at 54.65–55.89 ppm was assigned to C-3′. Spiro carbon was observed at 63.11–63.55 ppm, whereas a peak at 68.92–69.44 ppm was ascribed to C-5′. Aromatic carbons resonated at 120.06–158.47 ppm. The carbonyl carbons of CONH and C=O displayed signals in the region 181.43–181.93 ppm and 196.86–197.45 ppm.
X-ray crystallographic analyses
The X-ray crystalline structure of 6d was formed in the triclinic cubic system with P 1 space group Fig. 2a. From the structure refinement data Table 1, cell parameters of the unit cell were a = 6.7642(3) Å, b = 13.0580(5) Å, C=17.0927(7) Å and V = 1457.56(11) Å3. The spiro moieties pyrrolidine and oxindole were linked together via C17 asymmetric centre. The phenyl group was attached through C15 carbon, while biphenyl chalcone was connected via C14 carbon and L-leucine was interconnected via C16 chiral carbon. The spectral data and X-ray structure were in good agreement confirming the formation of spiro-oxindole compounds. Intermolecular hydrogen bonding of 6d was found between NH of pyrrolidine moiety and carbonyl oxygen N–H…..O and the interactions were shown in Fig. 2b. The chosen bond angles and bond lengths of 6d were tabulated in Table 2 and Table 3, respectively. The longest bond length was found to be C28-Br with a bond length of 1.875 (14) Å and both N1-H1 and N1-H2 showed the shortest bond length of 0.8600 Å. Similarly, C23-C24-C17 showed the highest bond angle of 132.8 (12)O and N1-C16-C15 showed the shortest bond angle of 103.2 (10)O.
Hirshfeld surface analysis
Hirshfeld surface analysis showed intermolecular interaction, the isosurface produced from the calculations indicated electron density partitioning between the molecules which defines the area of a molecule in the crystal by partitioning a molecule’s electron density into contributions from individual atoms [52]. Crystal Explorer is software used for analysing and visualising crystal structures and intermolecular interactions in materials [53, 54]. HSA of di, dnorm, de, curvedness, shape index and fragment patch were plotted in the region of − 0.3920 to 1.7745, − 0.7916 to 2.8598, − 0.9070 to 2.9150, − 1.0000 to 1.0000, − 4.0000 to 0.4000 and 0.0000 to 16.0000 shown in Fig. 3. The red spots in dnorm indicate the shortest contact and were localised in oxygen and nitrogen atoms, suggesting the presence of intermolecular hydrogen bonding while the blue and white regions correspond to longer bonds other than hydrogen bonding. The di signifies the distance between the surface and external atom and de indicates the distance of the surface to the internal atom. Fingerprint plots Fig. 4 acquired from HSA provide a quantitative summary of intermolecular interactions in a molecule. The most dominant interactions are H…H (54.7%) and C…H (10.1%). All the other interactions were less than 10%.
DFT calculations
Molecular geometry
All quantum chemical computations for compound 6d were performed using DFT methods by means of the B3LYP/6–31 + G(d,p) basis set. The compound 6d is a neutral molecule with 70 atoms and 300 electrons existing in a singlet system with a molecular formula of C34H31BrN2O2. The optimized and the 2D structure of compound 6d have been displayed in Fig. 5. The compound 6d was fully optimized and was visualized in GaussView 05 software. The molecule had two nitrogen, two oxygen atoms heteroatoms and one bromo atom. Carbon 1 was the spiro carbon, carbon atoms 1, 3, 4, 5 and N-2 were found in the pyrrolidine ring, whereas carbon atoms 30, 31, 33, 34, 35, 36 and 37, along with N-32 were located in the indole ring, carbon 6, 7, 8 and 9 was the aliphatic leucine carbons and the remaining carbon atoms were two phenyl ring carbons.
Molecular electrostatic potential
Molecular electrostatic potential (MEP) offers vital data regarding electrostatic interactions between molecules and the reactive sites based on the distribution of charges [55,56,57]. MEP is often depicted as an electrostatic potential map, which is a three-dimensional visualisation of the electrostatic potential at different spots in a space surrounding a molecule. MEP is represented in various colour codes. Positive potentials are portrayed in blue, whereas negative potentials are shown in red. Orange possesses a slight negative charge, yellow attributes to a slightly positive charge and green corresponds to neutral potential [58, 59]. The MEP map was envisioned in GaussView 05 software between − 7.592e-2 and 7.592e -2 and shown in Fig. 6. The red spot was localised in the oxygen atom of the indole ring O38 which is electron rich and susceptible to electrophilic attack. Similarly, the oxygen atom of the carbonyl group O12 is slightly electron rich. The blue area is broadly distributed over the H-atom throughout the surface of the molecules, subjected to nucleophilic attack.
Frontier molecular orbital
The FMO provides information about chemical reactivity, conductivity, kinetic stability and biological properties associated with the compound 6d. HOMO and LUMO are the highest and the lowest energy levels of a molecule. HOMO is about the ability of a molecule to act as an electron donor or nucleophile, while LUMO reveals its ability to act as an electron acceptor or electrophile. The energy gap is the energy required to promote an electron from HOMO to LUMO. If the energy gap is small then the molecule is more reactive because electrons can be more readily promoted from HOMO to the LUMO, facilitating electron transfer and bond formation in chemical reactions. If the energy gap is larger, then the molecule is stable and rigid [60,61,62]. In compound 6d, the electron cloud was distributed on the spiro ring while LUMO was localised in biphenyl rings. The calculated energy gap was found to be 0.0799 eV Fig. 7. The low energy gap value suggests the reactive nature of synthesized compound 6d.
Quantum chemical parameters
The physicochemical parameters were calculated from the HOMO and LUMO energy levels using the standard formulas [63] and tabulated in Table 4. The electron donating and accepting nature of compound 6d were illustrated based on ionization potential and electron affinity values. Chemical potential is exactly the opposite of electronegativity. The compound’s stability is connected to its chemical hardness, 6d showed a low value of 0.0399 eV suggesting the soft nature of the compound. The high global softness values also suggest 6d as a soft molecule i.e. high reactivity of the compound. The compound’s ability to accept and donate electrons was quantitatively evaluated using the electrophilicity and nucleophilicity indexes which were derived from chemical potential and chemical hardness values, respectively. Compound 6d displayed a strong electron donating nature with 5.2603 eV nucleophilicity index. The high chemical softness value 25.0626 eV also discloses the soft nature of 6d, indicating that these molecules can lose electrons readily and are polarizable. This confirmed the reactive nature of 6d.
Mulliken analysis
Mulliken analysis was performed to determine the distribution of electron density within the molecule and the results were summarised in Table 5 and the charge distribution was shown in Fig. 8. C33 exhibited a maximum positive charge of 0.482534 a.u and N32 possessed a minimal negative charge of − 0.777291 a.u. Both C33 and N32 were located inside the indole ring. The hetero atoms N2, N32, O12 and O38 are all negatively charged and act as electron donors. Among the hetero atoms N32 is more negatively charged and the least negatively charged is O12 with − 0.430457 a.u. Most of the carbon atoms are negatively charged. The bromine atom present in this compound Br39 possesses positive charge of 0.142207 a.u indicating it is an electron acceptor. All the hydrogen atoms showed a positive charge, H40 is more positively charged (0.378928 a.u) as it is directly attached to N2 nitrogen atom of the spiro ring followed by H66 (0.314379 a.u) which is attached to N32 of the indole ring and H68 (0.111935 a.u) is the less positive hydrogen atom, suggesting that they are electron acceptors. The Mulliken charge distribution demonstrated the positive and negative charge of compound 6d.
Molecular docking
The MDM2 protein structure was obtained from the RCSB protein data bank, along with its reference spiro[3H-indole-3,2-pyrrolidin]-2(1H)-one (6SJ) which was prepared according to standard etiquettes and docked with (6a-6 h) to identify the docking interactions [64, 65]. The docking results were validated by redocking the co-crystal 6SJ with the receptor and the results were concise in Table 6. From the binding affinity values, it was evident that, the synthesized compounds displayed better binding energy values of − 8.62 to − 10.83 kcal/mol in comparison to the standard reference 6SJ affinity value of − 8.06 kcal/mol. The halogens substituted derivatives 6b, 6c and 6d disclosed more potent interaction with the receptor than the other substituent. The compound 6d bearing bromo group in the para position of the phenyl ring displayed the best binding energy followed by 6c and 6b. Their 2D and 3D binding interactions were shown in Fig. 9 and the binding interactions of other compounds were given in supplementary Fig. S41. The compound 6d was stabilized by number of hydrophobic interactions with the residues; it displayed pi-alkyl and alkyl interactions with ILE61, VAL93, ILE99, LEU57, LEU54 and GLY58 with a bond distance of 5.88, 4.61, 5.24, 4.62, 5.00 and 6.31 Å. Similarly, pi-pi T shaped interactions such as HIS96, TYR100, TYR67 and HIS73 with a distance of 5.44, 6.96, 4.05 and 6.33 Å, a pi cation interaction with LYS94 with a distance of about 5.15 Å and halogen bond was found in MET62 at a distance of 3.81 Å. MET62 displayed another pi-sulfur interaction with at a distance of 7.69 Å. Likewise, 6c also disclosed pi-alkyl, alkyl, pi-pi stacked interactions with the amino acid residues. It also showed hydrogen bond interaction with LEU54 at a distance of 3.07 Å. On the other hand, along with pi-alkyl, alkyl and pi-sigma interactions, 6b also displayed halogen interaction between fluorine atom and GLY58 at a distance of 2.55 Å, pi-sulfur interaction with MET62 at 4.35 Å, pi-pi stacked interactions with TYR67 and HIS73 at a distance of 4.71 and 4.21 Å respectively. Also, an unfavourable donor-donor interaction with LYS94 at a bond distance of 2.47 Å.
Virtual ADMET screening
ADMET assessments are crucial for the effective drug discovery and development process, as they enable preliminary candidate selection, pharmacokinetic optimization, safety evaluation and regulatory compliance. They improve the possibility of identifying viable and safe drug candidates while reducing the cost associated with development [66, 67]. The oral bioavailability nature of the drug molecule is determined by its drug-likeness properties like total polar surface area, log P, molecular weight, hydrogen bond acceptor and donor [68]. All the compounds were assessed for ADMET properties, and the findings exposed that these compounds obey Lipinski’s rule with a couple of violations due to variations in the log P and molecular weight value as shown in Table 7. Pharmacokinetic studies are essential in predicting dose amount, dosing interval and route of delivery into the body. Understanding a drug’s pharmacokinetic behaviour enables us to optimize the outcomes of treatment [69]. For pharmacokinetic evaluation, parameters like gastrointestinal absorption (GI), P-glycoprotein (P-gp), Blood Brain Barrier permeation (BBB) and Cytochrome P450 were taken into consideration and their results were précised in Table 8. Most of the compounds disclosed high GI absorption except 6f, while these compounds could not penetrate through BBB. P-gp is an essential protein in drug absorption and elimination; however, these compounds act as non-substrate of P-gp. Cytochrome P450 enzymes are essential in the metabolism of pharmaceuticals, regulating pharmacological effectiveness and toxicity. Hence, hindering these enzymes has adverse impacts [70]. Most of the derivatives were non-inhibitors of CYPIA2 and CYP2C9, except 6 h and 6a. The methyl and methoxy substituted compounds 6e-6 h tends to inhibit CYP2C19 and CYP3A4, while the simple and halo group substituted derivatives showed positive outcomes. Whereas all the compounds were identified as inhibitors of CYP2D6. As excepted compounds 6b, 6c and 6d showed positive pharmacokinetic results. A molecule with the least skin permeant value is recommended for oral or transdermal administration, in accordance with compounds 6a-6 h exhibited promising results with negative log kp value.
Toxicity prediction is an effective tool in early-stage drug development, chemical safety evaluation and risk assessment. According to toxicology data, it was evident from Table 9, that these compounds revealed the tendency to constrain the hERG gene. Analyzing the hepatotoxicity descriptor of the derivatives, it was found that all the derivatives were soberly safe for the liver except 6 h, indicating that these compounds might be less prone to cause significant liver damage. In the case of AMES toxicity, except 6c and 6 h, the remaining derivatives were determined as mutagens. Rat oral acute toxicity results exposed these compounds as less toxic, suggesting that these compounds exhibit a lower degree of acute toxicity when delivered verbally to rats. These compounds were non-irritant to both the skin and ocular irritation suggesting the safe nature of the compounds. All the compounds were less-carcinogen, signifying that these compounds have lower potential to cause cancer, while 6 h is non-carcinogen. The potential of an agent to trigger liver discomfort or damage is referred to as respiratory toxicity. Nevertheless, these compounds have no negative effects on the respiratory organs.
Anticancer activity
Anticancer activity was assessed against the human lung cancer A549 cell line using the standard MTT assay protocol. The survival of the A549 cell line was reduced in a dose-responsive way by compounds 6a-6 h. The IC50 values were summarised in Table 10 and all the investigated derivatives demonstrated modest to substantial activity against A549 cells when compared to doxorubicin. Compounds 6d and 6c outperformed the other compounds in terms of antiproliferative efficacy. The compound 6d with IC50 value of 8.13 ± 0.66 was chosen for assessing cytotoxicity towards the L929 cell line. Figure 10 associates % cytotoxicity index of 6d in A549 and L929. Cytotoxicity of L929 and A549 cell lines proliferates with increasing concentration of 6d; nevertheless, it affects the viability of A549 cells in comparison with normal viable cells. Thus, signifying the efficacy of 6d towards lung cancer cell line.
Structural-activity relationship studies
The preceding section displayed the IC50 values of all six compounds (6a-6 h) as well as the positive control Doxorubicin. All the tested compounds demonstrated significant anticancer efficacy against the A549 cell line. However, the current investigation also explored the influence of substituents on its activity. Compound 6a, with no substituent on the phenyl ring, had the least activity among all the compounds. The inclusion of halogens (F, Cl and Br) enhanced its activity significantly; however, the bromo substituted molecule 6d demonstrated substantial anticancer activity, followed by chloro and fluoro substituted compounds. This may be due to the more polarisable nature of bromine which enhances the interaction between the receptor and the ligand. In contrast, the incorporation of electron donating groups (-OCH3, -N(CH3)2 and -(OCH3)3) resulted in a lower activity profile. Thus, compound 6d was found to be more potent compared to standard doxorubicin in targeting lung cancer cells and further studies should be conducted.
Cell morphology studies
The effect of compound 6d on cell survival might potentially be assessed by microscopic examination. An A549 lung cancer cell was treated with different concentrations of 6d namely 20–100 µL with 20 µL increment. The cell morphological changes were observed using an inverted phase contrast microscope. Figure 11 displayed that with the increase in the concentration of compound 6d cell viability of the A549 cell line decreased. Furthermore, cells that were medicated exhibited notable morphological alterations, such as cell shrinkage, in a dose-responsive way.
AO/EB fluorescent staining
Apoptosis is a sort of programmed destruction of cells that differs from necrosis [71]. Cells undergo a strictly regulated and controlled process of self-destruction during apoptosis. This procedure is crucial for eradicating damaged, unnecessary or hazardous cells from the body without swelling or damaging neighbouring tissues [72]. AO/EB staining is a typical fluorescence staining approach used to differentiate between living and dead cells based on membrane integrity [73]. In accordance with IC50 values against the A549 cell line, compound 6d was selected for the AO/EB nuclear staining approach. To evaluate the morphological modifications of the cell nucleus, cells were exposed to 6d, dyed with AO/EB colorant and viewed using an inverted phase contrast microscope. Figure 12 displays stained morphological changes of cells at different concentrations with the control cells. The bright green nuclei of treated cells displayed condensed chromatin, cell shrinkage and decreased cell size indicating that compound 6d stimulated early apoptotic cell destruction in A549 malignant cells.
LDH assay
Lactate dehydrogenase, a cytoplasmic enzyme facilitates the oxidation of lactate to pyruvate. The quantity of LDH seeping from the cytoplasm was measured to estimate the integrity and permeability of the plasma membrane [74]. When the A549 cells are damaged during the process of apoptosis, LDH is released. The control used in this study is sodium pyruvate. Compound 6d induced dose-dependent membrane damage similar to the control shown in Fig. 13, at 100 µg/mL concentration 6d exerted cytotoxicity via apoptosis.
Conclusion
In summary, a series of 3′-([1,1′-biphenyl]-4yl)carbonyl-5′-(2-methylpropyl)4′-(aryl)spiro[indole-3,2′-pyrrolidin]-2(1H)-one derivatives were synthesized and analyzed using FT-IR, mass, 1H and 13C NMR spectral techniques. Single crystal XRD of compound 6d validated the structural arrangement of the spiro compound. Intermolecular interaction was quantified from Hirshfeld surface analysis and major interaction was contributed from H…H interaction (54.7%). Furthermore, the structure of 6d was optimized using DFT calculations and their quantum chemical simulations correlated well with the outcomes of the experiments. The HOMO–LUMO energy gap was determined to be 0.0799 eV, indicating potential reactive nature of 6d. MEP map and Mulliken analysis disclosed reactive spots and charge distribution within the molecule. In contrast to the standard 6SJ affinity value was found to be − 8.06 kcal/mol, 6d excelled minimum binding affinity of − 10.83 kcal/mol from molecular docking analysis. ADMET studies suggest compounds tend to follow Lipinski’s rule, with some exceptions. Pharmacokinetic studies anticipate strong gastrointestinal absorption, non-inhibitors of CYPIA2 and CYP2C9 along with low skin permeant values. Toxicity prediction indicates most of these compounds constrain the hERG gene, are safe for the liver, less carcinogenic and have no effect on the respiratory organs. Compound 6d disclosed substantial anticancer activity against A549 cell line with IC50 value of 8.13 ± 0.66 µM compared to standard doxorubicin. When 6d was evaluated for cytotoxicity against A549 and L929, it specifically affected cancerous A549 cell line rather than healthy cells. Compound 6d showed early apoptotic cellular death in A549 cell line in AO/EB staining method. Lactose dehydrogenase assay evidenced the dose-dependent membrane damage by 6d. Based on the outcomes, compound 6d is endorsed as a potential anticancer agent targeting the human lung cancer A549 cell line.
Data availability
No datasets were generated or analysed during the current study.
References
Arun Y, Saranraj K, Balachandran C, Perumal PT (2014) Novel spirooxindole-pyrrolidine compounds: synthesis, anticancer and molecular docking studies. Eur J Med Chem 74:50–64. https://doi.org/10.1016/j.ejmech.2013.12.027
Wang Y, Kong H, Zeng X et al (2016) Activation of NLRP3 inflammasome enhances the proliferation and migration of A549 lung cancer cells. Oncol Rep 35(4):2053–2064. https://doi.org/10.3892/or.2016.4569
Gupta A, Behl T, Heer HR, Deshmukh R, Sharma PL (2019) Mdm2-P53 interaction inhibitor with cisplatin enhances apoptosis in colon and prostate cancer cells in vitro. Asian Pac J Cancer Prev 20:3341–3351. https://doi.org/10.31557/APJCP.2019.20.11.3341
Shangary S, Wang S (2008) Targeting the MDM2-p53 interaction for cancer therapy. Clin Cancer Res 14:5318–5324. https://doi.org/10.1158/1078-0432.CCR-07-5136
Ding K, Lu Y, Nikolovska-Coleska Z, Wang G et al (2006) Structure-based design of spiro-oxindoles as potent, specific small-molecule inhibitors of the MDM2—p53 interaction. J Med Chem 2:3432–3435. https://doi.org/10.1021/jm051122a
Rew Y, Sun D, De Turiso FG-L et al (2012) Structure-based design of novel inhibitors of the MDM2—p53 interaction. J Med Chem 55:4936–4954. https://doi.org/10.1021/jm300354j
Hai J, Sakashita S, ALlo G, Ludkovski O, Ng C, Shepherd FA, Tsao M (2015) Inhibiting MDM2-p53 interaction suppresses tumor growth in patient-derived non—small cell lung cancer. JTO Acquis 10(8):1172–1180. https://doi.org/10.1097/JTO.0000000000000584
Chessari G, Hardcastle IR, Ahn JS et al (2021) Structure-based design of potent and orally active isoindolinone inhibitors of MDM2-p53 protein—protein interaction. J Med Chem 64:4071–4088. https://doi.org/10.1021/acs.jmedchem.0c02188
Nietzold F, Rubner S, Berg T (2019) The hydrophobically-tagged MDM2–p53 interaction inhibitor Nutlin-3a-HT is more potent against tumor cells than Nutlin-3a. Chem Commun 1:14351–14354. https://doi.org/10.1039/c9cc07795b
Chène P (2003) Inhibiting the p53–MDM2 interaction: an important target for cancer therapy. Nature 3:102–109. https://doi.org/10.1038/nrc991
Babu ARS, Raghunathan R, Gayatri G, Sastry GN (2006) A highly regioselective synthesis of 1-N-methyl-spiro-[2,3′′′]-oxindole-spiro-[3,2″]indane-1″,3″-dione-4-arylpyrrolidines through 1,3-Dipolar cycloaddition protocol. ChemInform 5:1–6. https://doi.org/10.1002/chin.200713115
Jayashankaran J, Manian RDRS, Venkatesan R, Raghunathan R (2005) A regioselective synthesis of dispiro[oxindole-cyclohexanone]pyrrolidines and dispiro[oxindole-hexahydroindazole]pyrrolidines by sequential 1,3-dipolar cycloaddition and annulation through a microwave induced solvent-free approach. Tetrahedron 61(23):5595–5598. https://doi.org/10.1016/j.tet.2005.03.088
Girgis AS, Panda SS, Srour AM et al (2015) Rational design, synthesis and molecular modeling studies of novel anti-oncological alkaloids against melanoma. Org Biomol Chem 13(23):6619–6633. https://doi.org/10.1039/c5ob00410a
Islam MS, Ghawas HM, El-Senduny FF et al (2019) Synthesis of new thiazolo-pyrrolidine–(spirooxindole) tethered to 3-acylindole as anticancer agents. Bioorg Chem 82:423–430. https://doi.org/10.1016/j.bioorg.2018.10.036
Islam MS, Al-majid AM, El-senduny FF et al (2020) Synthesis, anticancer activity, and molecular modeling of new halogenated spiro[pyrrolidine-thiazolo-oxindoles] derivatives. Appl Sci 10:1–11. https://doi.org/10.3390/app10062170
Yeung BKS, Zou B, Rottmann M, Lakshminarayana SB et al (2015) Potent synergy between spirocyclic pyrrolidinoindolinones and fluconazole against Candida albicans. ChemMedChem 10(10):1672–1686. https://doi.org/10.1002/cmdc.201500271
Yeung BKS, Zou B, Rottmann M et al (2010) Spirotetrahydro β-carbolines (spiroindolones): a new class of potent and orally efficacious compounds for the treatment of malaria. J Med Chem 53(14):5155–5164. https://doi.org/10.1021/jm100410f
Kumar RS, Rajesh SM, Perumal S, Banerjee D, Yogeeswari P, Sriram D (2010) Novel three-component domino reactions of ketones, isatin and amino acids: synthesis and discovery of antimycobacterial activity of highly functionalised novel dispiropyrrolidines. Eur J Med Chem 45(1):411–422. https://doi.org/10.1016/j.ejmech.2009.09.044
Murugan R, Anbazhagan S, Sriman Narayanan S (2009) Synthesis and in vivo antidiabetic activity of novel dispiropyrrolidines through [3 + 2] cycloaddition reactions with thiazolidinedione and rhodanine derivatives. Eur J Med Chem 44(8):3272–3279. https://doi.org/10.1016/j.ejmech.2009.03.035
Crosignani S, Prêtre A, Jorand-Lebrun C et al (2011) Discovery of potent, selective, and orally bioavailable alkynylphenoxyacetic acid CRTH2 (DP2) receptor antagonists for the treatment of allergic inflammatory diseases. J Med Chem 54(20):7299–7317. https://doi.org/10.1021/jm200866y
Jiang T, Kuhen KL, Wolff K et al (2006) Design, synthesis and biological evaluations of novel oxindoles as HIV-1 non-nucleoside reverse transcriptase inhibitors. Part I Bioorganic Med Chem Lett 16(8):2105–2108. https://doi.org/10.1016/j.bmcl.2006.01.073
Zhao Y, Yu S, Sun W et al (2013) A potent small-molecule inhibitor of the MDM2-p53 interaction (MI-888) achieved complete and durable tumor regression in mice. J Med Chem 56(13):5553–5561. https://doi.org/10.1021/jm4005708
Zheng M, Yang J, Xu X, Sebolt JT, Wang S, Sun Y (2010) Efficacy of MDM2 inhibitor MI-219 against lung cancer cells alone or in combination with MDM2 knockdown, a XIAP inhibitor or etoposide. Anticancer Res 30(9):3321–3331
Shangary S, Ding K, Qiu S, Nikolovska-Coleska Z, Bauer JA et al (2008) Reactivation of p53 by a specific MDM2 antagonist ( MI-43) leads to p21-mediated cell cycle arrest and selective cell death in colon cancer. Mol Cancer Ther 7:1533–1543. https://doi.org/10.1158/1535-7163.MCT-08-0140
Barakat A, Islam MS, Ghawas HM et al (2019) Design and synthesis of new substituted spirooxindoles as potential inhibitors of the MDM2—p53 interaction. Bioorg Chem 86:1–29. https://doi.org/10.1016/j.bioorg.2019.01.053
Palermo G, Diego S (2015) Computational chemistry for drug discovery. Encycl Nanotechnol 66:15. https://doi.org/10.1007/978-94-007-6178-0
Moroy G, Martiny VY, Vayer P, Villoutreix BO, Miteva MA (2012) Toward in silico structure-based ADMET prediction in drug discovery. Drug Discov Today 17:44–55. https://doi.org/10.1016/j.drudis.2011.10.023
Olasupo SB, Uzairu A, Shallangwa G, Uba S (2020) QSAR modeling, molecular docking and ADMET/pharmacokinetic studies: a chemometrics approach to search for novel inhibitors of norepinephrine transporter as potent antipsychotic drugs. J Iran Chem Soc 17(8):1953–1966. https://doi.org/10.1007/s13738-020-01902-5
Sivanandhan M, Parasuraman A (2023) In silico molecular docking and ADMET predictions of pyrido[2,3-d]pyrimidine-2,4(1H,3H)-dione analogues as promising antimicrobial, antioxidant and anticancer agents. Polycycl Aromat Compd. https://doi.org/10.1080/10406638.2023.2191973
Elangovan N, Arumugam N, Almansour AI et al (2024) Synthesis, solvent role, absorption and emission studies of cytosine derivative. Heliyon 10:e28623. https://doi.org/10.1016/j.heliyon.2024.e28623
Arulmurugan S, Vennila JP, Kavitha HP et al (2024) Synthesis, solvent role in TD-DFT (IEFPCM model), fluorescence and reactivity properties, topology and molecular docking studies on sulfathiazole derivative. J Mol Liq 400:124570. https://doi.org/10.1016/j.molliq.2024.124570
Elangovan N, Sowrirajan S, Arumugam N, Almansour AI et al (2024) Synthesis, vibrational analysis, absorption and emission spectral studies, topology and molecular docking studies on sulfadiazine derivative. ChemistrySelect 9:e202303582. https://doi.org/10.1002/slct.202303582
Elangovan N, Arumugam N (2024) Solvent role in molecular structure level, structure and topology investigations, molecular docking and antimicrobial activity studies on sulpha drug derivatives. J Mol Struct 1303:137570. https://doi.org/10.1016/j.molstruc.2024.137570
Elangovan N, Thirumavalavan M, Ganesan TS, Sowrirajan S et al (2024) Comparison study (experimental and theoretical), hydrogen bond interaction through water, donor acceptor investigation and molecular docking study of 3,3-((1,2-phenylenebis(azaneylylidene))bis(methaneylylidene))diphenol. J Biomol Struct Dyn 1:1–16. https://doi.org/10.1080/07391102.2024.2333465
Sheldrick GM (2008) A short history of SHELX. Acta Crystallogr Sect A Found Crystallogr 64(1):112–122. https://doi.org/10.1107/S0108767307043930
Sheldrick GM (2015) SHELXT—Integrated space-group and crystal-structure determination. Acta Crystallogr Sect A Found Crystallogr 71(1):3–8. https://doi.org/10.1107/S2053273314026370
McKinnon JJ, Jayatilaka D, Spackman MA (2007) Towards quantitative analysis of intermolecular interactions with Hirshfeld surfaces. Chem Commun 37:3814–3816. https://doi.org/10.1039/b704980c
Spackman MA, McKinnon JJ (2002) Fingerprinting intermolecular interactions in molecular crystals. CrystEngComm 4(66):378–392. https://doi.org/10.1039/b203191b
Periyannan P, Beemarao M, Karthik K, Ponnuswamy S, Ravichandran K (2022) Crystal structure, DFT and Hirshfeld surface analysis of N-acetyl-t-3-methyl-r-2, c-6-diphenylpiperidine. Acta Crystallogr Sect E Crystallogr Commun 78:179–183. https://doi.org/10.1107/S2056989022000275
Periyannan P, Beemarao M, Karthik K, Ponnuswamy S, Ravichandran K (2020) Crystal structure, Hirshfeld surface analysis and DFT studies of 1-[r-2, c-6-diphenyl-t-3-(propan-2-yl)piperidin-1-yl]ethan-1-one. Acta Crystallogr Sect E Crystallogr Commun 76:377–381. https://doi.org/10.1107/S2056989020002042
Sert Y, El-Emam AA, Al-Deeb OA, Al-Turkistani AA, Ucun F, Çirak Ç (2014) The biomolecule, 2-[(2-methoxyl)sulfanyl]-4-(2-methylpropyl)-6-oxo-1,6- dihydropyrimidine-5-carbonitrile: FT-IR, Laser-Raman spectra and DFT. Spectrochim Acta—Part A Mol Biomol Spectrosc 126:86–97. https://doi.org/10.1016/j.saa.2014.01.140
Becke AD (1993) Density-functional thermochemistry: III—the role of exact exchange. J Chem Phys 98(7):5648–5652. https://doi.org/10.1063/1.464913
Lee C, Yang W, Parr RG (2011) Development of the Colic-Salvetti correlation-energy formula into a functional of the electron density. The Am Phy Soc 50(11):36–39. https://doi.org/10.1103/PhysRevB.37.785
Chugh A, Sehgal I, Khurana N et al (2023) Comparative docking studies of drugs and phytocompounds for emerging variants of SARS-CoV-2. Biotech 13(1):1–17. https://doi.org/10.1007/s13205-022-03450-6
Wdowiak K, Walkowiak J, Pietrzak R, Bazan-Woźniak A, Cielecka-Piontek J (2022) Bioavailability of hesperidin and its aglycone hesperetin—compounds found in citrus fruits as a parameter conditioning the pro-health potential (neuroprotective and antidiabetic activity). Nutrients 14:2647. https://doi.org/10.3390/nu14132647
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) Open Babel: an open chemical toolbox. J Cheminform 3(33):1–14
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18(15):2714–2723. https://doi.org/10.1002/elps.1150181505
Morris GM, Goodsell DS, Halliday RS et al (1998) Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem 19(14):1639–1662. https://doi.org/10.1002/(SICI)1096-987X(19981115)19:14%3c1639::AID-JCC10%3e3.0.CO;2-B
Shaweta S, Akhil S, Utsav G (2021) Molecular docking studies on the Anti-fungal activity of Allium sativum (Garlic) against Mucormycosis (black fungus) by BIOVIA discovery studio visualizer 21.1.0.0. Ann Antivirals Antiretrovir 5(1):028–032. https://doi.org/10.17352/aaa.000013
Daina A, Michielin O, Zoete V (2017) SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 7:1–13. https://doi.org/10.1038/srep42717
Dong J, Wang NN, Yao ZJ et al (2018) Admetlab: a platform for systematic ADMET evaluation based on a comprehensively collected ADMET database. J Cheminform 10(1):1–11. https://doi.org/10.1186/s13321-018-0283-x
Al WLH, Joubert J, Blacque O, Al SNH, El-Emam AA (2019) Crystal structure, Hirshfeld surface analysis and DFT studies of 5-(adamantan-1-yl)-3-[(4- chlorobenzyl)sulfanyl]-4-methyl-4H-1,2,4-triazole, a potential 11β-HSD1 inhibitor. Sci Rep 9:1–11. https://doi.org/10.1038/s41598-019-56331-z
Spackman MA, Spackman PR, Thomas SP (2021) Beyond Hirshfeld surface analysis: interaction energies, energy frameworks and lattice energies with CrystalExplorer. In: Complementary bonding analysis, pp 329–352. https://doi.org/10.1515/9783110660074-013
Saeed A, Ejazb SA, Aziz M, Channar PA et al (2023) Efficient synthesis of 1-(bromomethyl)-3,5-dimethoxybenzene: X-ray structure, Hirshfeld surface analysis, DFT AND Molecular modelling investigations as tyrosinase inhibitor. Quim Nova 47(5):1–10
Sasitha T, John WJ (2021) Design, docking, and DFT investigations of 2,6-bis(3,4-dihydroxyphenyl)-3-phenethylpiperidin-4-one. Heliyon 7(2):e06127. https://doi.org/10.1016/j.heliyon.2021.e06127
Kumar KS, Haridharan N, Ranjith S, Nataraj A (2023) Studies on the DFT calculations and molecular docking of versatile molecular sensor 1-(6-Aminopyridin-2-yl)-3-(4-nitrophenyl)urea. Chem Phys Impact 6:100139. https://doi.org/10.1016/j.chphi.2022.100139
Channar PA, Aziz M, Ejaz SA, Saeed A et al (2024) Synthesis, biochemical characterization and computational investigations of newly synthesized crystal structure of Dihydropyrimidin-2(1H)-thione. Results Chem 7:101537. https://doi.org/10.1016/j.rechem.2024.101537
Arulaabaranam K, Mani G, Muthu S (2020) Computational assessment on wave function (ELF, LOL) analysis, molecular confirmation and molecular docking explores on 2-(5-Amino-2- Methylanilino)-4-(3-pyridyl) pyrimidine. Chem Data Collect 29:100525. https://doi.org/10.1016/j.cdc.2020.100525
Sumithra M, Sundaraganesan N, Rajesh R et al (2023) Electron acceptor, excitation energies, oscillatory strength, spectroscopic and solvent effects on 5-amino-4,6-dichloro-2-( propylthio)pyrimidine—anticancer agent. Chem Phys Impact 6:100145. https://doi.org/10.1016/j.chphi.2022.100145
Khanna L, Singhal S, Jain SC, Khanna P (2020) Spiro-indole-coumarin hybrids: synthesis, ADME, DFT, NBO studies and in silico screening through molecular docking on DNA G-quadruplex. ChemistrySelect 5:3420–3433. https://doi.org/10.1002/slct.201904783
Attaullah HM, Ejaz SA, Channar PA, Saeed A et al (2024) Exploration of newly synthesized azothiohydantoins as the potential alkaline phosphatase inhibitors via advanced biochemical characterization and molecular modeling approaches. BMC Chem 18(47):1–15. https://doi.org/10.1186/s13065-024-01149-8
Ahmed A, Saeed A, Ejaz SA, Aziz M et al (2022) Novel adamantyl clubbed iminothiazolidinones as promising elastase inhibitors: design, synthesis, molecular docking, ADMET and DFT studies. RSC Adv 12:11974–11991. https://doi.org/10.1039/d1ra09318e
Kores JJ, Danish IA, Sasitha T, Stuart JG, Pushpam EJ, Jebaraj JW (2021) Spectral, NBO, NLO, NCI, aromaticity and charge transfer analyses of anthracene-9,10-dicarboxaldehyde by DFT. Heliyon 7:e08377. https://doi.org/10.1016/j.heliyon.2021.e08377
Lotfy G, El Ashry ESH, Said MM et al (2018) Regio- and stereoselective synthesis of new spirooxindoles via 1,3-dipolar cycloaddition reaction: anticancer and molecular docking studies. J Photochem Photobiol B Biol 180(2017):98–108. https://doi.org/10.1016/j.jphotobiol.2018.01.026
Islam MS, Al-Majid AM, Sholkamy EN et al (2023) Optimized spirooxindole-pyrazole hybrids targeting the p53-MDM2 interplay induce apoptosis and synergize with doxorubicin in A549 cells. Sci Rep 13(1):1–23. https://doi.org/10.1038/s41598-023-31209-3
Caceres EL, Tudor M, Cheng AC (2020) Deep learning approaches in predicting ADMET properties. Futur Med Chem 12:1995–1999. https://doi.org/10.4155/fmc-2020-0259
Kiriiri GK, Njogu PM, Mwangi AN (2020) Exploring different approaches to improve the success of drug discovery and development projects: a review. Futur J Pharm Sci 6(1):1–12. https://doi.org/10.1186/s43094-020-00047-9
Olasupo SB, Uzairu A, Shallangwa GA, Uba S (2020) Profiling the antidepressant properties of phenyl piperidine derivatives as inhibitors of serotonin transporter (SERT) via cheminformatics modeling, molecular docking and ADMET predictions. Sci African 9:e00517. https://doi.org/10.1016/j.sciaf.2020.e00517
Alqahtani MS, Kazi M, Alsenaidy MA, Ahmad MZ (2021) Advances in oral drug delivery. Front Pharmacol 12:1–21. https://doi.org/10.3389/fphar.2021.618411
Nandini Asha R, Durai Nayagam BR, Bhuvanesh N (2021) Synthesis, molecular docking, and in silico ADMET studies of 4-benzyl-1-(2,4,6-trimethyl-benzyl)-piperidine: potential Inhibitor of SARS-CoV2. Bioorg Chem 112:104967. https://doi.org/10.1016/j.bioorg.2021.104967
Verma R, Bairy I, Tiwari M, Bhat GV, Shenoy GG (2019) In silico studies, synthesis and anticancer activity of novel diphenyl ether-based pyridine derivatives. Mol Divers 23(3):541–554. https://doi.org/10.1007/s11030-018-9889-1
Li GY, Du KJ, Wang JQ et al (2013) Synthesis, crystal structure, DNA interaction and anticancer activity of tridentate copper(II) complexes. J Inorg Biochem 119:43–53. https://doi.org/10.1016/j.jinorgbio.2012.09.019
Duddukuri NK, Thatikonda S, Godugu C, Kumar RA, Doijad N (2018) Synthesis of novel thiophene-chalcone derivatives as anticancer- and apoptosis-inducing agents. ChemistrySelect 3(24):6859–6864. https://doi.org/10.1002/slct.201800613
Anbu S, Kandaswamy M, Kamalraj S, Muthumarry J, Varghese B (2011) Phosphatase-like activity, DNA binding, DNA hydrolysis, anticancer and lactate dehydrogenase inhibition activity promoting by a new bis-phenanthroline dicopper(II) complex. Dalt Trans 40(28):7310–7318. https://doi.org/10.1039/c1dt10277j
Acknowledgements
The authors thank PSGR Krishnammal College for Women and DST FIST, New Delhi, for infrastructural facilities, as well as the College of Excellence for their equipment and technical support.
Author information
Authors and Affiliations
Contributions
MS involved in the synthesis, characterization, application and specifically writing the initial draft. SR is responsible for docking and DFT studies. AT involved in analysis of crystal structure and ADMET studies. AP contributed to the supervision and leadership responsibility for the research activity planning and execution, including mentorship discussion, application and correcting the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sivanandhan, M., Ragupathy, S., Thangamani, A. et al. Synthesis, crystal structure, Hirshfeld surface, computational and biological studies of spiro-oxindole derivatives as MDM2-p53 inhibitors. Mol Divers (2024). https://doi.org/10.1007/s11030-024-10974-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11030-024-10974-x