Abstract
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) causing COVID-19 continues to mutate. Numerous studies have indicated that this viral mutation, particularly in the receptor-binding domain area, may increase the viral affinity for human angiotensin-converting enzyme 2 (hACE2), the receptor for viral entry into host cells, thereby increasing viral virulence and transmission. In this study, we investigated the binding affinity of SARS-CoV-2 variants (Delta plus, Iota, Kappa, Mu, Lambda, and C.1.2) on hACE2 using computational modeling with a protein–protein docking approach. The simulation results indicated that there were differences in the interactions between the RBD and hACE2, including hydrogen bonding, salt bridge interactions, non-bonded interactions, and binding free energy differences among these variants. Molecular dynamics simulations revealed that mutations in the RBD increase the stability of the hACE2-spike protein complex relative to the wild type, following the global stability trend and increasing the binding affinity. The value of binding-free energy calculated using molecular mechanics/Poisson–Boltzmann surface area (MM/PBSA) indicated that all mutations in the spike protein increased the contagiousness of SARS-CoV-2 variants. The findings of this study provide a foundation for developing effective interventions against these variants.
Graphical abstract

Computational modeling elucidates that the spike protein of SARS-CoV-2 variants binds considerably stronger than the wild-type to hACE2.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The COVID-19 (coronavirus disease 2019) pandemic has not yet been declared over. There are difficulties in reducing illness incidence because, despite widespread vaccination, the mutation of SARS-CoV-2, the cause of this disease, still occurs. Until now, the US government's SARS-CoV-2 Interagency Group (SIG) and the European Center for Disease Prevention and Control (ECDC) have continued to monitor the development of the emergence of new variants of this virus. They have classified SARS-CoV-2 variants into three categories as follows: variants of interest (VOI), variants of concern (VOC), and variants of high consequence (VOHC) [1]. The rapid spread of VOCs has become one of the remaining concerns due to their increased transmissibility, the severity of disease, significant reduction in antibody neutralization, and decreased treatment effectiveness [2,3,4,5,6].
The increased frequency of spike (S) protein mutations, particularly those affecting the receptor-binding domain (RBD), raises awareness. This is because if the mutation continues, it will cause a significant change in the structure of the RBD, resulting in a dramatic increase in the rate of reinfection and immunity evasion [7]. The S protein is composed of two subunits: Subunit 1 (S1), which contains the ACE2 receptor-binding domain (RBD), and Subunit 2 (S2), which contributes to the fusion process [8,9,10] (Fig. 1).
The VOC Delta (δ) Plus (AY.1 or lineage B.1.617.2.1), which evolved from Delta, possesses a distinct mutation profile. The Delta variant has L452R and T478K mutations in the receptor-binding domain (S1 subunit), while the δ Plus variant, besides having L452R and T478K, also has K417N mutations [11]. After the δ variant caused an increase in COVID-19 cases, a new VOI called Mu (μ) (also known as lineage B.1.621 or VUI-21JUL-1) has emerged, which has drawn the interest of the WHO. This variant was discovered in Colombia in January 2021 and contains several spike mutations, some of which are shared with other VOC (E484K, N501Y, and P681H), while others are new (R346K, Y144T, Y145S, and 146 N insertion) [12].
Another novel VOI within the lineage B.1.1.1 (also termed as C.37, or Lambda/λ variant) was detected in Peru. It has mutations L452Q and F490S in RBD [13]. The F490S mutation was suspected to be associated with decreased antibody neutralization susceptibility [14]. The Kappa (κ) variant (lineage B.1.617.1) was first discovered in India in 2020, and has been assigned as VOI. The mutation L452R in κ variant was associated with reduced antibody neutralization by disrupting the respective conformational epitopes [15]. The VOI Iota (ι) (lineage B.1.526), which became dominant in New York City in early 2021, contains the mutation E484K. It has been reported to be partially or wholly resistant to two currently used therapeutic monoclonal antibodies and is less susceptible to neutralization [16]. A novel variant, currently referred to as C.1.2, was first discovered in South Africa. It draws the attention of scientists because of the rapidity with which mutations have occurred. Although it shares some properties with VOC, it is currently included in variants under monitoring [17].
The development of safe and effective vaccines and drugs is urgently needed to end this pandemic. The S-protein has been identified as the most effective immunological target protein for COVID-19 vaccines [18, 19]. Several drug discovery efforts have been undertaken to prevent the virus from invading and infecting the host cells by targeting the S-protein from attaching to hACE2 (angiotensin-converting enzyme 2) on the surface of the host cells [20]. Some of them are repurposed antiviral medications that have been approved by the Food and Drug Administration (FDA) [21, 22]. Other initiatives include the search for drugs that are derived from medicinal plants [23,24,25].
Due to the dynamic nature of mutations in the RBD of SARS-CoV-2 that have been shown to change both viral affinity for the host receptor and infectivity, it is critical to perform rigorous investigations to determine the effect of mutations on the spike protein's binding to hACE2. In the present work, we assessed the structural alterations in RBD that affect its binding affinity for hACE2 using an in silico method that included protein–protein docking, molecular dynamics simulations, and binding-free energy calculations. The results of this evaluation will provide critical information about the dynamics of the RBD-hACE2 interaction, which will aid in the design of novel vaccines and the discovery of new therapeutics for the treatment of COVID-19.
Materials and methods
Multiple sequence alignment
The three-dimensional (3D) crystal structure of the SARS-CoV-2 receptor-binding domain (RBD) that binds to hACE2 was retrieved from the Protein Data Bank with PDB ID 6M0J (https://www.rcsb.org/structure/6M0J) [26]. This protein is hereinafter referred to as wild-type (WT). In an in silico mutagenesis of amino acid sequences of RBD, SARS-CoV-2 variants were modeled based on mutation sites. The UCSF Chimera package release 1.15 [27] was used to perform the multiple alignment analysis (MSA) of these amino acid sequences. The information on mutation sites in the RBD of SARS-CoV-2 variants was obtained from previously published articles and is shown in Table 1.
Homology modeling
The 3D structure of each variant was modeled using the SWISS-MODEL web server (https://swissmodel.expasy.org/; accessed on 12 Oct 2021) [30, 31]. To validate the protein structure, the overall quality of the modeled protein was verified using ProSA-web (https://prosa.services.came.sbg.ac.at/prosa.php; accessed on 12 Oct 2021) [32]. The sequence similarity was computed using a normalized BLOSUM62 [33] on the SWISS webserver. The accuracy of the protein model was analyzed on the PROCHECK server [34] (https://saves.mbi.ucla.edu/) and presented in a Ramachandran plot [35].
Molecular docking
The 3D structures of modeled RBD proteins were minimized in the SWISS-MODEL web server. Validation of the molecular docking was performed by redocking the WT RBD to hACE2 and quantified using the root-mean-square deviation (RMSD) value. The molecular docking simulation was performed in the HDOCK server (http://hdock.phys.hust.edu.cn/) [36, 37] to predict interactions between the RBD and hACE2. The predicted values for binding affinity (ΔG) kcal/mol) and dissociation constant (KD) (M) at 25.0 °C were obtained from the Prodigy website (https://wenmr.science.uu.nl/prodigy/; accessed on 17 Oct 2021) [38, 39]. PDBsum [40] was used to visualize the interaction network, including salt bridges, hydrogen interactions, and non-bonded contacts.
Molecular dynamics simulations
The following procedure was carried out following our previous study [1]. A molecular dynamics (MD) simulation was performed with the Gromacs 2019.2 version [41] to examine the dynamic behavior and binding interaction level of hACE2-RBD complexes. The topology of the hACE2-RBD proteins was created with the AMBER99SB-ILDN force fields [42] and the SCP water model. Triclinic box type was preferred for solvation at 10 Å from the protein–protein complex. The system was neutralized by adding 0.15 M NaCl. Energy minimization was performed in 5000 steps with the steepest descent integrator. It is equilibrated with 0.3 ns NVT and 0.3 ns NPT stages, a V-rescale thermostat, and a Parrinello-Rahman barostat [43], respectively. Molecular dynamics simulation of 1000 frames with 100 ns was performed at 2 fs with a leap-frog integrator. Trajectory analysis was performed with gmx rms, rmsf, gyrate and hbond scripts. GraphPad Prism 8 was used to plot all the analyses.
Binding free energy calculations
The molecular mechanics Poisson–Boltzmann surface area (MM-PBSA) [44] was utilized to perform the binding-free energy (BFE) calculation using the Gromacs g_mmpbsa script [45]. The calculation of BFE was performed using the following equation:
Results
Multiple sequence alignments
The MSA result for all RBD amino acid sequences is shown in Fig. 2, demonstrating that WT contains no mutations. The following are the names of the variants in the Greek alphabet and their mutation sites: δ417 (K417N, L452R, T478K), δ484 (L452R, T478K, E484Q), μ (R346K, E484K, N501Y), λ (L452R, F490S), κ (L452R, E484K), ι (E484K), and an unassigned Greek name C.1.2 (Y449H, E484K, N501Y).
Homology modeling
Table 2 shows the percentage sequence similarity of the RBD variants’ sequences with WT. The δ417, δ484, μ, and λ variants show 99.01% similarity, κ and ι have 99.51% similarity, while C.1.2 exhibits 98.52% similarity. Based on ProSA-web, the overall model quality of the 3D model of RBD variants is presented in Table 3, while the Z-score and plot of residue scores are displayed in Fig. 3. The Z-score ranges from −6.03 to −5.82. Because all Z-score values are negative, the modeling results appear to be valid. Additionally, validation was performed on the PROCHECK webserver and the results are presented in Ramachandran plots (Supplementary File 1). All models fall within the requirements because the residues in the most favored region are as follows: C.1.1 91.7%, δ484 91.7%, δ417 91.1%, ι 91.1%, κ 90.5%, λ 90.5%, μ 91.7%, and WT 89.3%. In addition, the percentage of non-glycine residues in the disallowed region of these models was less than 15%.
Protein–protein molecular docking
The WT RBD was re-docked to hACE2 for molecular docking validation. The RMSD value between the crystal structure (PDB: 6M0J) and the docked spike protein is 0.41 Å. which consequently revealed the validity of the docking protocol. The RMSD of the S protein of the variants is below 0.61 Å (Fig. 4).
The mechanism of interaction between RBD and hACE was elucidated using molecular docking on the HDOCK server. The results are presented in Table 4 and Fig. 5. The types of interactions are listed in Table 5. The predicted HDOCK score for the WT-hACE2 complex was −347.27 kcal/mol. This value is lower than the BFE values of the other variants except for C.1.2-hACE2, which is lower than WT-hACE2. Prodigy prediction for ΔG shows that the lowest value is μ-hACE2 (−14.2 kcal/mol), and the highest is κ-hACE2 (−12.8 kcal/mol), although the range of ΔG values is not too different between variants. The docking score is only a preliminary prediction of the affinity of the complex between RBD and h-ACE2. Following that, BFE values for MM/PBSA based on the results of molecular dynamics simulation studies are presented to provide a more accurate value. The highest KD value was observed in κ-hACE2 (3.9 × 10–10 M), and the lowest was in μ-hACE2 (3.6 × 10–11 M). The highest RMSD was observed in ι-hACE2 (0.61 Å) and the lowest was in δ484 (0.34 Å).
Except for C.1.2, all variants had an H-bond between Tyr449 and Asp38. With the exception of δ417, all variants have a Lys417-Asp30 salt bridge (indicated in bold). Except for δ484, H-bonds Gln493-Lys31 and Gln493-Glu35 (underlined) were found in all variants. The H-bond Gln498-Lys353 (indicated in italics) occurs in all but 484 and C.1.2, where these two variants only form the Gln498-Gln42 interaction (shown in italics); this interaction also occurs in other variants except WT, ι, κ, and δ417. Each RBD contains H-bonds with the residues Lys31 and Lys353, which act as hot spots in hACE2 (indicated by bold and underlined). Except for μ, all RBDs have three H-bonds with Lys31. The results of this molecular docking show there is a shift in interacting residues between WT RBD and hACE2 compared to other RBD variants, despite the fact that interactions continue to occur at critical residues on each protein.
Molecular dynamics simulations analysis
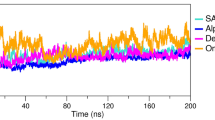
Analysis of the structural stability is a key parameter in the determination of the binding stability of the interacting partners. A common method for demonstrating the binding is to use simulation trajectory estimation for dynamic stability. Herein, employing root-mean-square deviation (RMSD) as a function of time, we computed the stability of each complex over the 100 ns simulation time (Fig. 6). The RMSD analysis of WT RBD revealed that the structure remained more stable than all the variants. It can be seen that the structure reached the equilibrium point at 5 ns and stabilized the structure at 0.18 nm. The RMSD of WT RBD remained uniform during the first 40 ns; however, the RMSD converged slightly between 41 and 50 ns and then decreased, repeating the pattern until 100 ns. On the other hand, the variants comparatively demonstrated unstable behavior during the simulation. In the case of δ417-ACE2 complex, the structure demonstrated significant unstable behavior. The RMSD deviated at different time intervals substantially, particularly between 5 and 75 ns. The mean RMSD for δ417-ACE2 complex was 0.260 nm. Moreover, the δ484-ACE2 complex also showed a destabilizing effect. This complex demonstrated stability drift of between 5 and 50 ns. The mean RMSD for δ484-ACE2 complex was 0.216 nm. Furthermore, the κ-ACE2 complex also had a destabilizing effect, with substantial convergence at different time intervals. As can be seen, the complex demonstrated significant convergence at the start of the simulation but then experienced a reduction in deviations in the final 50 ns. The ι-ACE2 and δ484-ACE2 complexes exhibited comparable behavior. The RMSD initially converged and then stabilized at the end of the simulation. The μ-ACE2 exhibited comparable behavior to WT, though it experienced structural perturbation at some points but remained stable. In addition, the λ-ACE2 and C.1.2-ACE2 variants also possessed unstable dynamics by showing deviations in the RMSDs.
Human ACE2 and wild-type spike and some mutant spike protein RBD S1b unit complexes molecular dynamics simulations. Root-mean-square deviation (RMSD) of ACE2 and RBD delta plus (δ417-ACE2 and δ484-ACE2), kappa (κ-ACE2), mu (μ-ACE2), iota (ι-ACE2), lambda (λ-ACE2), C.1.2 lineage (C.1.2-ACE2), and wild-type (WT-ACE2) complexes
Understanding the structural compactness of interacting molecules reveals critical information about their binding and unbinding events. To calculate the structural compactness, the radius of gyration (Rg) approach was used as a function of time, which revealed a stronger correlation with the RMSD results (Fig. 7). The wild type demonstrated significant structural compactness. Initially, during the 1–20 ns, the Rg values remained a little higher and then decreased to 3.1 nm. The Rg increased during 65–80 ns and then decreased. On the other hand, the δ417-ACE2 and δ484-ACE2 complexes demonstrated significant fluctuations in the Rg(s). These complexes exhibited a significant increase and a decrease in the Rg(s) during the start of the simulation, and then δ417-ACE2 particularly reported a uniform Rg after 60 ns. The δ484-ACE2 structure demonstrated a little higher Rg in the last part of the simulation, particularly after 40 ns. The κ-ACE2 complex demonstrated a gradual decrease in the Rg from the start of the simulation, and after reaching 70 ns, the Rg increased back for a short time period (until 90 ns) and then decreased back. The μ-ACE2, ι-ACE2, λ-ACE2, and C.1.2-ACE2 complexes showed comparable Rg(s) with no significant convergence, though they showed smaller deviations at different time intervals. The fluctuation in Rg values during the simulations of all the studied systems was associated with the binding and unbinding of one or another end of the spike RBD domain. An increase and decrease in the Rg pattern can also be pointed to 100 ns, although these variations occurred in WT and all the variants.
To understand the bonding pattern between the WT-ACE2 and variants, the total number of H-bonds in each complex was calculated, as follows: δ417-ACE2 (8.3), δ484-ACE2 (11.3), κ-ACE2 (13.9), μ-ACE2 (8.1), ι-ACE2 (10.4), λ-ACE2 (12.0), C.1.2-ACE2 (6.9), while 10.3 average H-bonds were reported in the WT complex (Fig. 8). Consequently, this shows the hydrogen reprogramming caused by the mutations in the RBD and induces a different approach to an increase in the transmissibility.
Overall, these findings indicate that the spike protein undergoes structural adjustments to bind efficiently to the ACE2 receptor and, in turn, increases entry into the host cells. The residual flexibility as root-mean-square fluctuation (RMSF) is shown in Fig. 9A. The RBDs apo demonstrated more similar residual fluctuations. In particular, the region between 350 and 400 displayed a higher fluctuation in the region 480–500 also reported a higher fluctuation, particularly the κ-ACE2 complex. These overall results suggest that possible evolutionary changes in the mutants have led to variations in their dynamic function. On the other hand, the ACE2 also demonstrated more similar fluctuations, except for minor variations in the residual flexibility (Fig. 9B). Comparatively, the variants exhibited higher fluctuations, particularly in the regions of 1–80, 210–280, 350–4.0, and 550–590. This shows the variation in the flexibility dynamics and, consequently, the conformational optimization, which in turn helps with better binding.
Protein–protein binding free energy calculations
The binding-free energy (BFE) value from docking results is frequently less precise. As a result, the MM/PBSA method was used to analyze BFE at the molecular level (Fig. 10). Table 6 displays the calculation result, which indicate that the BFE value of each variant’s complex is greater than the WT-hACE2 complex value (−1121.68 kcal/mol). The highest level of BFE (−2485.79 kcal/mol) was shown by the complex δ484-hACE2.
Discussion
Throughout the COVID-19 pandemic, the SARS-CoV-2 virus evolved, resulting in genetic heterogeneity in the population of circulating viral strains. The spike (S) protein on the surface of the viruses enables the entrance of the virus into cells by binding the receptor-binding domain to the hACE2 receptor on the host cell's surface [10]. A common mutation in spike appears to favor open conformations in the protein, which may allow the virus to enter cells more easily [46, 47]. Numerous studies have demonstrated that mutations alter the amino acids in RBD, affecting the viral infectivity [48], and also the ability of the host antibody to neutralize the virus [49]. Mutations in the spike, particularly in the receptor-binding domain, appear to have consequences for viral attachment to hACE2. Therefore, the current study examined the effect of RBD mutations on the binding affinity of several SARS-CoV-2 variants to hACE2.
The RBD amino acid sequences of the variants δ417, δ484, μ, λ, κ, ι, and C.1.2 were obtained based on information from various previous studies and aligned with WT. Based on these sequences, the RBD of each variant was modeled. The Z-score value reflects the overall quality of a protein model and is also a measure of the deviation of the total energy of the structure in relation to the energy distribution obtained from a random conformation. In general, a positive value of the Z-score indicates a problem or erroneous in a model [32]. The predicted Z-scores ranged from −6.03 to −5.82, which is consistent with native proteins and indicates very few incorrect structures. The score represents a highly reliable structure because it falls within the range of scores normally found in proteins of similar size [50]. Based on Ramachandran plots, the most favored region of the modeled proteins was in the range of 89.3–91.7%. Additionally, the percentage of non-glycine residues in the disallowed region was below 15%, indicating the good quality of the protein structures [51, 52].
The superimposed RMSD values recorded between variants and WT are important because they measure the kind of accuracy needed to interpret function. The difference in RMSD values was considerable, showing that the structures of the variants exhibited structural deviation, secondary structural element disturbance, and protein conformational changes. The structure of the viral protein plays an important role in its function, so any changes in the shape of the structure will affect its function, virulence, infectivity, and transmission [53]. Molecular docking of RBD and hACE2 was performed to better understand the impact of changes in the RBD protein structure of these variants. Protein–protein interactions are crucial in molecular biological events, and molecular docking facilitates the study of this interaction. Protein–protein docking is a method for predicting the interaction of two protein structures [54]. The changes in protein structure have an impact on the dynamics of protein–protein interactions between RBD and hACE2, which in turn affects the binding affinity of the two proteins. Mutations E484K and N501Y have binding affinity to hACE2 comparable to WT. According to previous findings, L452R increased ACE2 binding affinity to RBD [55]. The presence of mutations in the RBD amino acids K417, E484, L452, T478 and N501 also significantly increased the affinity of this protein for hACE2. This has a consequence where the α, β, γ, and δ variants will be more infective to host cells than WT [56]. The HDOCK docking score showed a higher C.1.2-ACE2 and μ-ACE2 complex value than WT-ACE2. This is also supported by the smallest KD of μ-ACE2 (3.6 × 10–11 M), where the lower the dissociation constant, the more tightly bound the ligand is, or the higher the ligand–protein affinity.
Critical residues in the SARS-CoV-2 RBD that recognize hACE include Gly446, Tyr449, Leu455, Phe486, Tyr491, Gln493, Gly496, Gln498, Thr500, Asn501, and Gly502 [26, 57,58,59]. Among these, Phe486, Gln493, and Asn501 allow hACE2 to easily recognize the RBD, making them the most important residues in RBD [60] [10]. Apart from being considered critical residues, Ala475 and Phe484 play a role in the antibody neutralization mechanism [61]. Meanwhile, when compared to SARS-CoV, the residues Leu455, Phe456, Phe486, Gln493, and Gln498 in SARS-CoV-2 RBD bind to hACE with a high degree of affinity. The ACE2-spike complex is stabilized by Lys31 and Lys353 of ACE2, which are located in the contact interface with the RBD and provide a significant amount of energy [62].
An interface analysis reveals that the highest number of H-bonds is found at C.1.2-hACE2 and the lowest number is found at μ-hACE2. Though H-bonding has shifted among the variants, the current findings demonstrate the consistency of important conserved interactions, including Lys417-Asp30, Tyr449-Asp38, Gly496-Asp38, and Gly496-Lys353. The salt bridge Lys417-Asp30 was observed in all variants, including WT-hACE, with the exception of δ417-hACE. The existence of variations in protein–protein interactions is something that needs to be taken into consideration when determining key residues that should be investigated further for therapeutic potential and, potentially, when developing a universal vaccine for COVID-19.
Despite the fact that the BFE value from the interaction between RBD and hACE2 via HDOCK and Prodigy was obtained, the BFE calculation using the MM-PBSA method to calculate in-depth atomic-level interaction energy analysis was still performed. This is due to the fact that the BFE value derived from docking results is less accurate [63]. The destabilizing effects of some variants have been previously reported to be associated with enhanced binding because of the fact that mutations with destabilizing effects produce radical functions [57, 64]. Additionally, the bonding rearrangement during the MD simulation holds and releases the two receptors, which causes a perturbation in the structural compactness [65].
Protein–protein association is mainly guided by a variety of factors, among which hydrogen bonds and hydrophobic interactions are key players. Interaction of protein interfaces is always occupied by water molecules which compete with the hydrogen bonding between the residues [66]. The processes behind protein–protein coupling as well as the implications of which hydrogen bonds play a role in this association are unknown [67]. Whether hydrogen bonds govern protein–protein docking, in particular, is a long-standing concern with poorly understood mechanisms [68, 69].
The BFE value of MM/PBSA based on the MDS study indicated that RBD variants have a higher affinity for hACE2 than RBD WT does. The increased affinity for hACE2 may be a contributing factor to the increased SARS-CoV-2's infectivity, which is associated with increased transmission [70]. This is consistent with our previous findings [1] and study of Suleman et al. [63] using a different variant, which demonstrated that the presence of mutations in RBD increased the spike binding affinity on hACE2. The complex δ484-hACE2 provides the highest level of BFE (−2485.79 kcal/mol).
Mutations at positions T478 and E484 affect residues at the immunodominant site of the RBD, resulting in virus-resistance antibody-mediated neutralization [71]. This has the potential to complicate vaccination programs as well as drug development, so these programs must pay close attention to ensure that vaccines and antiviral drugs can recognize these mutations. Similarly, mutations in other drug target sites in viruses must be carefully studied during the antiviral drug development process.
Conclusion
The infectivity of SARS-CoV-2 variants to humans is increasing day by day. This increase is caused by mutations in the spike protein of SARS-CoV-2 and stronger adhesion to hACE2. In this study, the binding of delta plus, kappa, mu, iota, lambda, C.1.2 lineage variants, and wild-type (WT-ACE2) spike proteins to hACE2 was analyzed by simulations. Based on the wild-type S1b (RBD) unit crystal structure, mutant RBD proteins were generated by homology modeling. These RBD and hACE2 were docked on a template-based basis. The behaviors of RBD-hACE2 complexes were investigated by molecular dynamics simulations with parameters such as RMSD, RMSF, Rg, and the number of hydrogen bonds between the two chains. Binding-free energy calculations revealed that, alarmingly, all variants showed a stronger affinity for hACE2 than the wild type.
References
Celik I, Yadav R, Duzgun Z et al (2021) Interactions of the receptor binding domain of SARS-CoV-2 variants with hACE2: Insights from molecular docking analysis and molecular dynamic simulation. Biology (Basel) 10:880. https://doi.org/10.3390/biology10090880
Liu Y, Liu J, Xia H et al (2021) Neutralizing activity of BNT162b2-elicited serum. N Engl J Med 384:1466–1468
Lopez Bernal J, Andrews N, Gower C et al (2021) Effectiveness of Covid-19 vaccines against the B.1.617.2 (Delta) variant. N Engl J Med 385:585–594. https://doi.org/10.1056/NEJMoa2108891
Planas D, Veyer D, Baidaliuk A et al (2021) Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature 596:276–280. https://doi.org/10.1038/s41586-021-03777-9
Plante JA, Mitchell BM, Plante KS et al (2021) The variant gambit: COVID-19’s next move. Cell Host Microbe 29:508–515. https://doi.org/10.1016/j.chom.2021.02.020
Sabino EC, Buss LF, Carvalho MPS et al (2021) Resurgence of COVID-19 in Manaus, Brazil, despite high seroprevalence. Lancet (London, England) 397:452–455. https://doi.org/10.1016/S0140-6736(21)00183-5
Shen L, Triche TJ, Bard JD, et al (2021) Spike Protein NTD mutation G142D in SARS-CoV-2 Delta VOC lineages is associated with frequent back mutations, increased viral loads, and immune evasion. medRxiv. https://doi.org/10.1101/2021.09.12.21263475
Tai W, He L, Zhang X et al (2020) Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol 17:613–620. https://doi.org/10.1038/s41423-020-0400-4
Tallei TE, Fatimawali YA et al (2021) An analysis based on molecular docking and molecular dynamics simulation study of bromelain as anti-SARS-CoV-2 variants. Front Pharmacol 12:2192. https://doi.org/10.3389/fphar.2021.717757
Shang J, Wan Y, Luo C et al (2020) Cell entry mechanisms of SARS-CoV-2. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.2003138117
Kannan SR, Spratt AN, Cohen AR et al (2021) Evolutionary analysis of the Delta and Delta Plus variants of the SARS-CoV-2 viruses. J Autoimmun 124:102715. https://doi.org/10.1016/j.jaut.2021.102715
Messali S, Bertelli A, Campisi G et al (2021) A cluster of the new SARS-CoV-2 B.1.621 lineage in Italy and sensitivity of the viral isolate to the BNT162b2 vaccine. J Med Virol. https://doi.org/10.1002/jmv.27247
Wink PL, Volpato FCZ, Monteiro FL et al (2021) First identification of SARS-CoV-2 Lambda (C.37) variant in Southern Brazil. Infect Control Hosp Epidemiol. https://doi.org/10.1017/ice.2021.390
Liu Z, VanBlargan LA, Bloyet LM et al (2021) Identification of SARS-CoV-2 spike mutations that attenuate monoclonal and serum antibody neutralization. Cell Host Microbe 29:477-488.e4. https://doi.org/10.1016/j.chom.2021.01.014
Yang T-J, Yu P-Y, Chang Y-C, et al (2021) Structure-activity relationships of B.1.617 and other SARS-CoV-2 spike variants. bioRxiv. https://doi.org/10.1101/2021.09.12.459978
Annavajhala MK, Mohri H, Wang P et al (2021) Emergence and expansion of SARS-CoV-2 B.1.526 after identification in New York. Nature 597:703–708. https://doi.org/10.1038/s41586-021-03908-2
Scheepers C, Everatt J, Amoako DG, et al (2021) Emergence and phenotypic characterization of C.1.2, a globally detected lineage that rapidly accumulated mutations of concern. medRxiv. https://doi.org/10.1101/2021.08.20.21262342
Salvatori G, Luberto L, Maffei M et al (2020) SARS-CoV-2 spike protein: an optimal immunological target for vaccines. J Transl Med 18:222. https://doi.org/10.1186/s12967-020-02392-y
Dai L, Gao GF (2021) Viral targets for vaccines against COVID-19. Nat Rev Immunol 21:73–82. https://doi.org/10.1038/s41577-020-00480-0
Zhang Q, Xiang R, Huo S et al (2021) Molecular mechanism of interaction between SARS-CoV-2 and host cells and interventional therapy. Signal Transduct Target Ther 6:233. https://doi.org/10.1038/s41392-021-00653-w
Gupta RK, Nwachuku EL, Zusman BE et al (2021) Drug repurposing for COVID-19 based on an integrative meta-analysis of SARS-CoV-2 induced gene signature in human airway epithelium. PLoS ONE 16:e0257784. https://doi.org/10.1371/journal.pone.0257784
Parvathaneni V, Gupta V (2020) Utilizing drug repurposing against COVID-19 - efficacy, limitations, and challenges. Life Sci 259:118275. https://doi.org/10.1016/j.lfs.2020.118275
Mahmud S, Paul GK, Afroze M et al (2021) Efficacy of phytochemicals derived from Avicennia officinalis for the management of COVID-19: a combined in silico and niochemical study. Molecules 26:2210. https://doi.org/10.3390/molecules26082210
Mousavi SS, Karami A, Haghighi TM et al (2021) In silico evaluation of Iranian medicinal plant phytoconstituents as inhibitors against main protease and the receptor-binding domain of SARS-CoV-2. Molecules 26:5724. https://doi.org/10.3390/molecules26185724
Tallei TE, Fatimawali NNJ et al (2021) A Comprehensive review of the potential use of green tea polyphenols in the management of COVID-19. Evid Based Compl Altern Med 2021:7170736. https://doi.org/10.1155/2021/7170736
Lan J, Ge J, Yu J et al (2020) Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581:215–220
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF Chimera - a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Wilhelm A, Toptan T, Pallas C, et al (2021) Antibody-Mediated Neutralization of Authentic SARS-CoV-2 B.1.617 Variants Harboring L452R and T478K/E484Q. Viruses 13:. https://doi.org/10.3390/v13091693
Laiton-Donato K, Franco-Muñoz C, Álvarez-D\’\iaz DA, et al (2021) Characterization of the emerging B.1.621 variant of interest of SARS-CoV-2. medRxiv. https://doi.org/10.1101/2021.05.08.21256619
Bienert S, Waterhouse A, de Beer TAP et al (2017) The SWISS-MODEL Repository-new features and functionality. Nucl Acids Res 45:D313–D319. https://doi.org/10.1093/nar/gkw1132
Waterhouse A, Bertoni M, Bienert S et al (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucl Acids Res 46:W296–W303. https://doi.org/10.1093/nar/gky427
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucl Acids Res 35:W407–W410. https://doi.org/10.1093/nar/gkm290
Henikoff S, Henikoff JG (1992) Amino acid substitution matrices from protein blocks. Proc Natl Acad Sci USA 89:10915–10919. https://doi.org/10.1073/pnas.89.22.10915
Laskowski RA, MacArthur MW, Moss DS, Thornton JM (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr 26:283–291. https://doi.org/10.1107/s0021889892009944
Kleywegt GJ, Jones TA (1996) Phi/Psi-chology: Ramachandran revisited. Structure 4:1395–1400. https://doi.org/10.1016/S0969-2126(96)00147-5
Yan Y, Zhang D, Zhou P et al (2017) HDOCK: a web server for protein-protein and protein-DNA/RNA docking based on a hybrid strategy. Nucl Acids Res 45:W365–W373. https://doi.org/10.1093/nar/gkx407
Yan Y, Tao H, He J, Huang SY (2020) The HDOCK server for integrated protein–protein docking. Nat Protoc 15:1829–1852. https://doi.org/10.1038/s41596-020-0312-x
Vangone A, Bonvin AMJJ (2015) Contacts-based prediction of binding affinity in protein–protein complexes. Elife. https://doi.org/10.7554/eLife.07454
Xue LC, Rodrigues JP, Kastritis PL et al (2016) PRODIGY: a web server for predicting the binding affinity of protein-protein complexes. Bioinformatics 32:3676–3678. https://doi.org/10.1093/bioinformatics/btw514
Laskowski RA, Jabłońska J, Pravda L et al (2018) PDBsum: structural summaries of PDB entries. Protein Sci 27:129–134. https://doi.org/10.1002/pro.3289
Abraham MJ, Murtola T, Schulz R, et al (2015) GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2:19–25. https://doi.org/10.1016/j.softx.2015.06.001
Lindorff-Larsen K, Piana S, Palmo K et al (2010) Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins Struct Funct Bioinforma 78:1950–1958. https://doi.org/10.1002/prot.22711
Parrinello M, Rahman A (1981) Polymorphic transitions in single crystals: a new molecular dynamics method. J Appl Phys 52:7182–7190. https://doi.org/10.1063/1.328693
Miller BR, McGee TD, Swails JM et al (2012) MMPBSA.py: an efficient program for end-state free energy calculations. J Chem Theory Comput 8:3314–3321. https://doi.org/10.1021/ct300418h
Kumari R, Kumar R, Lynn A (2014) G-mmpbsa -a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54:1951–1962. https://doi.org/10.1021/ci500020m
Wrapp D, Wang N, Corbett KS et al (2020) Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 367:1260–1263. https://doi.org/10.1126/science.abb2507
Mansbach RA, Chakraborty S, Nguyen K et al (2021) The SARS-CoV-2 Spike variant D614G favors an open conformational state. Sci Adv. https://doi.org/10.1126/sciadv.abf3671
Teng S, Sobitan A, Rhoades R et al (2021) Systemic effects of missense mutations on SARS-CoV-2 spike glycoprotein stability and receptor-binding affinity. Brief Bioinform 22:1239–1253. https://doi.org/10.1093/bib/bbaa233
Weisblum Y, Schmidt F, Zhang F et al (2020) Escape from neutralizing antibodies by SARS-CoV-2 spike protein variants. Elife. https://doi.org/10.7554/eLife.61312
Prajapat R, Bhattacharya I, Kumar A (2016) Homology modeling and structural validation of type 2 diabetes associated transcription factor 7-like 2 (TCF7l2). Trends Bioinforma 9:23–29. https://doi.org/10.3923/tb.2016.23.29
Lovell SC, Davis IW, Arendall WB et al (2003) Structure validation by Cα geometry: φ, ψ and Cβ deviation. Proteins Struct Funct Genet 50:437–450. https://doi.org/10.1002/prot.10286
Malleshappa Gowder S, Chatterjee J, Chaudhuri T, Paul K (2014) Prediction and analysis of surface hydrophobic residues in tertiary structure of proteins. Sci World J 2014:971258. https://doi.org/10.1155/2014/971258
Nguyen TT, Pathirana PN, Nguyen T et al (2021) Genomic mutations and changes in protein secondary structure and solvent accessibility of SARS-CoV-2 (COVID-19 virus). Sci Rep 11:3487. https://doi.org/10.1038/s41598-021-83105-3
Vakser IA (2014) Protein-protein docking: from interaction to interactome. Biophys J 107:1785–1793. https://doi.org/10.1016/j.bpj.2014.08.033
Deshpande A, Harris BD, Martinez-Sobrido L et al (2021) Epitope classification and RBD binding properties of neutralizing antibodies against SARS-CoV-2 variants of concern. Front Immunol 12:2185. https://doi.org/10.3389/fimmu.2021.691715
Sanches PRS, Charlie-Silva I, Braz HLB et al (2021) Recent advances in SARS-CoV-2 Spike protein and RBD mutations comparison between new variants Alpha (B.1.1.7, United Kingdom), Beta (B.1.351, South Africa), Gamma (P.1, Brazil) and Delta (B.1.617.2, India). J Virus Erad 7:100054. https://doi.org/10.1016/j.jve.2021.100054
Khan A, Zia T, Suleman M et al (2021) Higher infectivity of the SARS-CoV-2 new variants is associated with K417N/T, E484K, and N501Y mutants: an insight from structural data. J Cell Physiol. https://doi.org/10.1002/jcp.30367
Koley T, Madaan S, Chowdhury SR et al (2021) Structural analysis of COVID-19 spike protein in recognizing the ACE2 receptor of different mammalian species and its susceptibility to viral infection. 3 Biotech. https://doi.org/10.1007/s13205-020-02599-2
Yan R, Zhang Y, Li Y et al (2020) Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science. https://doi.org/10.1126/science.abb2762
Barh D, Tiwari S, Silva Andrade B et al (2020) Potential chimeric peptides to block the SARS-CoV-2 spike receptor-binding domain. F1000Research. https://doi.org/10.12688/f1000research.24074.1
Yi C, Sun X, Ye J et al (2020) Key residues of the receptor binding motif in the spike protein of SARS-CoV-2 that interact with ACE2 and neutralizing antibodies. Cell Mol Immunol 17:621–630. https://doi.org/10.1038/s41423-020-0458-z
Wan Y, Shang J, Graham R et al (2020) Receptor recognition by the novel coronavirus from Wuhan: an analysis based on decade-long structural studies of SARS coronavirus. J Virol. https://doi.org/10.1128/jvi.00127-20
Suleman M, Yousafi Q, Ali J et al (2021) Bioinformatics analysis of the differences in the binding profile of the wild-type and mutants of the SARS-CoV-2 spike protein variants with the ACE2 receptor. Comput Biol Med 138:104936. https://doi.org/10.1016/j.compbiomed.2021.104936
Hussain I, Pervaiz N, Khan A et al (2020) Evolutionary and structural analysis of SARS-CoV-2 specific evasion of host immunity. Genes Immun 21:409–419. https://doi.org/10.1038/s41435-020-00120-6
Khan A, Wei D-Q, Kousar K et al (2021) Preliminary structural data revealed that the SARS-CoV-2 B.1.617 variant’s RBD binds to ACE2 receptor stronger than the wild type to enhance the infectivity. Chem Bio Chem 22:2641–2649. https://doi.org/10.1002/cbic.202100191
Chen D, Oezguen N, Urvil P et al (2016) Regulation of protein-ligand binding affinity by hydrogen bond pairing. Sci Adv 2:e1501240. https://doi.org/10.1126/sciadv.1501240
Chodera JD, Mobley DL (2013) Entropy-enthalpy compensation: role and ramifications in biomolecular ligand recognition and design. Annu Rev Biophys 42:121–142. https://doi.org/10.1146/annurev-biophys-083012-130318
Patil R, Das S, Stanley A et al (2010) Optimized hydrophobic interactions and hydrogen bonding at the target-ligand interface leads the pathways of drug-designing. PLoS ONE 5:e12029. https://doi.org/10.1371/journal.pone.0012029
Olsson TSG, Ladbury JE, Pitt WR, Williams MA (2011) Extent of enthalpy-entropy compensation in protein-ligand interactions. Protein Sci 20:1607–1618. https://doi.org/10.1002/pro.692
Gan HH, Twaddle A, Marchand B, Gunsalus KC (2021) Structural modeling of the SARS-CoV-2 spike/human ACE2 complex interface can identify high-affinity variants associated with increased transmissibility. J Mol Biol 433:167051. https://doi.org/10.1016/j.jmb.2021.167051
Liu C, Ginn HM, Dejnirattisai W et al (2021) Reduced neutralization of SARS-CoV-2 B.1.617 by vaccine and convalescent serum. Cell 184:4220-4236.e13. https://doi.org/10.1016/j.cell.2021.06.020
Funding
The authors received no funding for the work they submitted. Aside from that, there are no known financial or personal relationships between the authors that might have influenced the findings reported in this article.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Celik, I., Khan, A., Dwivany, F.M. et al. Computational prediction of the effect of mutations in the receptor-binding domain on the interaction between SARS-CoV-2 and human ACE2. Mol Divers 26, 3309–3324 (2022). https://doi.org/10.1007/s11030-022-10392-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-022-10392-x