Abstract
Ventricular septal defect (VSD) is the most common type of congenital heart disease. HAND1 gene plays a crucial role in the development of the heart, but the role of the variants in the HAND1 gene promoter region in patients with VSD has not been explored yet. From 588 participants (300 with isolated and sporadic VSD and 288 healthy controls), DNA was extracted from blood samples. Variants at the HAND1 gene promoter region were analyzed through Sanger sequencing. Subsequently, cell functional validation was conducted through cell experiments, including dual-luciferase reporter gene analysis, electrophoretic mobility shift analysis, and bioinformatics analysis was also conducted. The promoter region of HAND1 gene had a total of 9 identified variant sites. Among them, 4 variants were exclusively found in VSD patients, and 1 variant (g.3631A>C) was newly discovered. Cell functional experiments indicated that all four variants decreased the transcriptional activity of HAND1 gene promoter with three of them reached statistical significance (p < 0.05). Subsequent analysis using JASPAR (a transcription factor binding profile database) suggests that these variants may alter the binding sites of transcription factors, potentially contributing to the formation of VSD. Our study for the first time identified variants in the promoter region of HAND1 gene in Chinese patients with isolated and sporadic VSD. These variants significantly decreased the expression of HAND1 gene, impacting transcription factor binding sites, and thereby demonstrating pathogenicity. This study offers new insights into the role of HAND1 gene promoter region, contributing to a better understanding of the genetic basis of VSD formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Congenital heart disease (CHD) is a type of anatomical, structural, and functional abnormality caused by obstacles or developmental anomalies in the formation of the heart and major blood vessels during embryonic development, or the failure of channels that should naturally close after birth [1]. In clinical terms, CHD can be classified into at least 21 types based on anatomical or hemodynamic criteria. This includes ventricular septal defect (VSD), atrial septal defect (ASD), patent ductus arteriosus (PDA), tetralogy of Fallot (TOF), among others. Among them, ASD, VSD, and TOF are the most common congenital heart disease [2]. CHD is the leading cause of death in infants with congenital defects, accounting for approximately 23.8% of deaths among newborns with congenital defects. With the widespread application of prenatal diagnosis and postnatal screening for CHD, the incidence of CHD is gradually increasing. It greatly impacts population quality and quality of life, bringing a heavy economic burden and psychological pressure to families and society. Therefore, research based on human genetics is crucial for improving the diagnosis and treatment of CHD patients. However, the underlying causes of CHD are not yet fully understood. Despite the long-standing belief that both genetic and environmental factors contribute to the pathogenesis of CHD, epidemiological findings suggest that genetic factors are the primary cause of CHD [3].
VSD is one of the most common CHD, accounting for 40% of all cardiac defects [4]. It has been reported that the incidence of VSD in newborns screened by high-sensitivity color Doppler echocardiography can reach up to 5% [5]. Genetic variants have been confirmed to be closely associated with VSD. However, many genes that influence VSD formation have yet to be discovered.
The gene known as heart and neural crest derivatives expressed (HAND), a member of the basic helix-loop-helix (bHLH) family, consists of HAND1 and HAND2. The transcription factor encoded by HAND1 (NG_052889.1) is crucial in trophoblast cell differentiation and cardiac development [6]. Located on chromosome 5q33, HAND1 encodes a transcription factor consisting of 215 amino acids. HAND1 can directly transactivate its downstream target genes and collaborate with transcription factors like GATA4 and Nkx25 to activate downstream target genes, such as the ANF promoter [7, 8]. Loss-of-function variants in HAND1 may impact normal cardiac development and structural remodeling, leading to CHD. HAND1 is expressed in the embryonic and adult heart in humans and vertebrates [9, 10], playing a crucial role in fetal myocardial cell proliferation, differentiation, cardiac morphogenesis, and exec. In mice, the biallelic knockout of HAND1 results in cardiac looping defects, impaired ventricular development, heart failure, abnormal cardiac septation, and embryonic death [11]. Specific deletion of HAND1 in the mouse heart may lead to atrioventricular valve malformations, cardiac outflow tract abnormalities, VSD, and abnormal ventricular growth and maturation [12]. Overexpression of HAND1 in the adult mouse heart can cause myocardial hypertrophy [13].
Our recent studies have revealed variants in the promoter region of the genes related to the heart development in VSD and other CHD. The promoter region was studied in VSD patients regarding ISL1 [14], MEF2C [15], MYH6 [16], CITED2 [17] and other CHD [18,19,20,21,22,23]. Given that the promoter region is the primary regulatory area of gene expression, we hypothesize that variants in HAND1 promoter may be closely related to the occurrence and development of VSD. This study aimed to discover the possible variants in the promoter region of HAND1in the Chinese patients with VSD. Further cellular functional studies were also performed to demonstrate the pathological role of the variants in the heart development.
Materials and method
Study participants
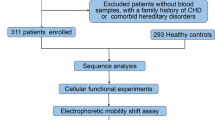
This study recruited a total of 588 participants, including 300 cases of sporadic and isolated VSD patients and a matched control group of 288 healthy individuals in terms of age and gender. All patients underwent VSD repair surgery at TEDA International Cardiovascular Hospital, Tianjin, China. Patients with a family history of VSD or CHD or other genetic diseases were excluded. The health controls were confirmed through cardiac Doppler ultrasound without CHD or other diseases. In accordance with the Helsinki Declaration, the study was approved by the Ethics Committee of TEDA International Cardiovascular Hospital (Clinical Research Ethics Review Number: [2021]-0715-4). Written informed consent was obtained from the parents or guardians of all participants after explaining the details. Figure 1 illustrates the flowchart of this study.
Genomic DNA extraction and sequence analysis
Using the blood genomic DNA extraction kit (centrifuge column type), DNA was extracted from 200 μl blood sample, resulting in a final 50 μl DNA product to be stored at − 20 °C. The promoter sequence of HAND1 gene (1681 bp, from − 1560 to + 121 bp to the transcription start site) was obtained from the GenBank database (NCBI, NG_052889.1). Two pairs of PCR primers were designed and the primer sequences are listed in Table 1. The PCR reaction cycle was set as follows: pre-denaturation at 95 °C for 5 min, denaturation at 95 °C for 30 s, annealing at 61 °C for 30 s, extension at 72 °C for 1 min/45 s for a total of 35 cycles, and final extension for 3 min. The PCR products obtained were subjected to Sanger sequencing, and the sequencing results of patients and healthy controls were compared by Chromas 2.6.5 and DNAMAN 6.0 software to screen for variants.
Plasmid construction, cell cultivation, and transfection
To clarify whether variants affect the promoter activity, the promoter region of HAND1 gene was inserted into the KpnI and HindIII cloning sites of the pGL3-Basic vector. Subsequently, the plasmid carrying the target gene was transformed into Escherichia coli DH5α chemically competent cells, followed by incubation in medium at 37 °C with agitation at 200 rpm for 16 h. Plasmids (pGL3-Basic, pGL3-WT, pGL3-V3592, pGL3-V3631, pGL3-V3658, pGL3-V3754) were extracted using the Star Prep rapid plasmid mini-prep kit (GenStar, China). On the day before transfection, cells were transferred to six-well plates and cultured in complete MEM/DMEM (10% fetal bovine serum + 1% penicillin–streptomycin). When cells reached 60–80% confluence, the pGL3-Basic-HAND1 promoter plasmid (2.5 ng) and the Renilla luciferase plasmid pRL-SV40 (0.25 ng) were co-transfected into HL-1 cells. pRL-SV40 served as an internal control, and an empty pGL3-Basic vector served as a negative control. Cells were further incubated in the six-well plate for 24–48 h, and their status was observed. Subsequently, dual-luciferase reporter gene assays were conducted.
Dual luciferase reporter gene assay
The transfected cells were collected and lysed, and an equal volume of reporter gene cell lysate was taken as a blank control. The RLU values of firefly fluoresceinase as well as sea kidney fluoresceinase were measured according to the instructions of the Dual Fluoresceinase Reporter Gene Detection Kit (Beyotime, China) using the specified Dual Fluoresceinase Reporter Gene Detection System (Beyotime Biotechnology, Beijing, China). The degree of activation of the target reporter gene was compared between different samples based on the ratios obtained. The experiment was independently repeated three times, with three replicates each time. Figure 2 shows the specific experimental process.
Dual luciferase reporter gene experiment workflow. In order to clarify whether variants have an impact on promoter activity, gene expression vectors containing wild-type and variant-type plasmids were constructed, and then transfected into mouse cardiac muscle cells (HL-1). The transcriptional activity of the wild-type and variant-type HAND1 gene promoters was assessed through a dual luciferase reporter gene experiment
Cell nucleus protein extraction and electrophoretic mobility shift experiment (EMSA)
Initially, Nuclear and cytoplasmic proteins were extracted from HL-1 cells using the Nuclear and Cytoplasmic Protein Extraction Kit (Beyotime, China). Subsequently, the protein concentration was determined using the enhanced BCA protein assay kit (Beyotime, China) and the protein was than stored at − 20 °C for further use. Biotinylated double-stranded oligonucleotides containing wild-type or variant sequences of HAND1 gene promoter (Table 1) were used as probes. The binding between HAND1 promoter and transcription factors was examined using the electrophoretic mobility shift assay (EMSA) kit (Beyotime, China) based on the chemiluminescence method.
Transcription factor binding site prediction
The JASPAR database was used to predict the transcription factor binding sites (TFBS) in HAND1 promoter region [24]. The relative threshold was set at 85%. The wild and variant-type nucleotide sequences are shown in Table 1.
Statistical analysis
All statistical analyses were conducted using SPSS 26.0. Standard Student's t-test was employed to compare quantitative experimental data. A value of p < 0.05 is considered statistically significant.
Results
Variants in HAND1 gene promoter region in VSD patients and healthy controls
Sanger sequencing was performed on the DNA of 588 participants (300 with VSD and 288 healthy controls), revealing a total of 9 variants. The mutation locations are illustrated in Fig. 3A. Among these, 4 variants [g.3592C>A(rs10062037), g.3631A>C, g.3658T>C(rs1287904093), g.3754T>C(rs1430611116)] were exclusive to VSD patients, as shown in sequencing chromatograms (Fig. 3B), while the remaining 5 variants were found in both VSD patients and healthy controls (Table 2). Notably, one of the four unique variants (g.3631A>C) has not been reported before (HAND1—SNP—NCBI https://www.ncbi.nlm.nih.gov/snp/?term=HAND1). Furthermore, the allele frequencies of these four variants in East Asian populations were < 0.001 in ALFA, 1000 Genomes, 1000 Genomes_30x, and gnomAD—Genomes databases (https://www.ncbi.nlm.nih.gov/snp/). Further validation through cellular functional experiments was conducted for these variants.
Location of variants in HAND1 gene promoter and sequencing chromatogram. A According to the genomic DNA sequence of HAND1 gene (NG_052889.1), variants are named based on the mutation points, with the transcription start site located at position 5,033 of the first exon. The diagram illustrates the positions of 9 promoter variants in HAND1 gene. B Sequencing chromatograms of 4 variants found exclusively in VSD patients [g.3592C>A(rs10062037), g.3631A>C, g.3658T>C(rs1287904093), g.3754T>C(rs1430611116)]. The upper trace represents the wild type, while the lower trace represents the mutated type, indicated by arrows
Dual luciferase reporter gene assay genetic test results
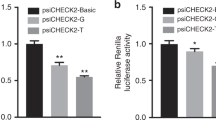
Wild and variant-type of HAND1 gene promoter were generated to construct a reporter gene expression vector (pGL3-Basic). This includes pGL3-Basic, pGL3-WT, pGL3-V3592, pGL3-V3631, pGL3-V3658, and pGL3-V3754. These vectors, along with pRL-SV40, were co-transfected into HL-1 cells, and after 48 h of cultivation, their dual luciferase activity was measured. The transcriptional activity of the wild-type HAND1 gene promoter was set as 100%. As shown in Fig. 4, the luciferase reporter gene detection activity of the four variants found exclusively in VSD patients was reduced and three of them reached statistical significance (p < 0.05).
Dual luciferase reporter gene assay results. The diagram illustrates the relative transcriptional activity data of the wild-type and variant-type HAND1 gene promoters in HL-1 cells, presented as the mean ± SD. The experiments were conducted three times, with three replicates each time (*p < 0.05, **p < 0.01, ***p < 0.001)
The results of EMSA
We used wild and variant-type biotin-labeled double-stranded probes for EMSA analysis of the four variants found exclusively in VSD patients. The biotinylated oligonucleotides for EMSA are listed in Table 1. As shown in Fig. 5, the arrows indicate bands with different brightness and darkness levels, suggesting the binding of different amounts of transcription factors at those positions. This verifies whether the mutated genes affect the binding of transcription factors. The results indicate that, compared to the wild type, probes with four variants [g.3592C>A(rs10062037), g.3631A>C, g.3658T>C(rs1287904093), g.3754T>C(rs1430611116)] exhibit a significantly reduced or increased binding capacity.
EMSA results. Biotin probes were separately subjected to EMSA analysis with four different variants of wild-type and variant-type. It was observed that four variants [g.3592C>A(rs10062037), g.3631A>C, g.3658T>C(rs1287904093), g.3754T>C(rs1430611116)] in cells could either induce or abolish the binding of transcription factors. EMSA electrophoretic mobility shift assay, WT wild type, VT variant type
The TFBS affected by regulatory variants
Variants identified through this study are predicted to potentially affect the binding sites of transcription factors in HAND1 gene promoter. The results indicate that the identified variants in this study may disrupt or create binding sites for transcription factors. As shown in Table 3, they generate 34 binding sites for factors such as HOXA5, PAX4, FOXA1, FOXD2, POU2F3, and disrupt 17 binding sites for factors like HNF4A, HNF4G, SP2, EHF, EGR3.
Figure 6 is a schematic diagram combining JASPAR database analysis, cell function experiments, and previous reports. The Figure depicts how variants in HAND1 gene promoter region could potentially influence VSD.
Discussion
This study for the first time in patients with sporadic and isolated VSD identified the variants in the promoter region of HAND1 gene and we have found that (1) among the 588 participants, a total of 9 variants were identified, with 4 variants [g.3592C>A(rs10062037), g.3631A>C, g.3658T>C(rs1287904093), g.3754T>C(rs1430611116)] found exclusively in patients with VSD. Importantly, the variant g.3631A>C was identified for the first time; (2) results from the dual-luciferase reporter gene assay indicated that all four variants decreased the transcriptional activity of HAND1 gene promoter with three of them reached statistical significance (p < 0.05); (3) through EMSA and bioinformatics analysis, it was discovered that these four variants are associated with changes in TFBS, and therefore the decreased transcriptional activity of HAND1 gene may be related to the occurrence of VSD.
The protein encoded by the human HAND1 gene contains an important structural domain (bHLH). HAND1 binds to downstream target gene promoters through this domain and also interacts with transcription factors. HAND1 gene is significantly expressed in human and vertebrate embryos as well as in the adult heart, playing a crucial role in fetal cardiac development and postnatal cardiac remodeling. It has been confirmed that variants in the HAND1 gene can cause VSD, left ventricular development defects, TOF, and dilated cardiomyopathy [25,26,27,28].
It is well known that the promoter is a critical regulatory region in gene transcription regulation. The mutation of the promoter can either decrease or increase the mRNA level, thereby altering the protein level [29]. In this study, the transcription activity of the wild-type and variant-type HAND1 gene promoters was examined using a dual luciferase reporter gene assay. The results indicate that all four variants in HAND1 gene promoter region significantly reduced the transcriptional activity of HAND1.
Transcription factors can regulate the expression of target genes by recognizing transcription binding sites within regulatory regions such as promoters and enhancers. They form specific interactions with DNA [30]. We also conducted bioinformatics analysis on four variants in the promoter region of HAND1 gene in VSD patients through EMSA and JASPAR database utilization. The analysis confirmed that these variants all lead to changes in cellular functions. Therefore, variants in the promoter region of HAND1 gene are highly likely to play a crucial role in reducing the expression of genes essential for cardiac morphogenesis during heart development. This, in turn, affects heart development and may ultimately lead to VSD.
Limitations
The interactions between the variants discovered in HAND1 gene promoter region and downstream genes need further validation. Further, the biological effects of these variants also need to be verified in animal models. These will be taken account into our future studies.
Conclusion
In conclusion, our study for the first time identified variants in the promoter region of HAND1 gene in Chinese patients with isolated and sporadic VSD. These variants significantly decreased the expression of HAND1 gene, impacting transcription factor binding sites, and thereby demonstrating pathogenicity. These pathological changes are likely related to the development of VSD. Therefore, this study provides new insights into the molecular mechanisms and potential causes of VSD related to genetic variants in the promoter region of the HAND1 gene.
Data availability
The individual SNP numbers are given in Table 2. The genetic variants described in this manuscript are available at https://www.ncbi.nlm.nih.gov/snp/. Data supporting the findings of this study are available upon reasonable request from the corresponding author.
References
Vincent SD, Mayeuf-Louchart A, Watanabe Y, Brzezinski JT, Miyagawa-Tomita S, Kelly RG, Buckingham M (2014) Prdm1 functions in the mesoderm of the second heart field, where it interacts genetically with Tbx1, during outflow tract morphogenesis in the mouse embryo. Hum Mol Genet 23:5087–5101. https://doi.org/10.1093/hmg/ddu232
Benjamin EJ, Blaha MJ, Chiuve SE, Cushman M, Das SR, Deo R, de Ferranti SD, Floyd J, Fornage M, Gillespie C et al (2017) Heart disease and stroke statistics-2017 update: a report from the American Heart Association. Circulation 135:e146–e603. https://doi.org/10.1161/CIR.0000000000000485
Oyen N, Poulsen G, Boyd HA, Wohlfahrt J, Jensen PK, Melbye M (2009) National time trends in congenital heart defects, Denmark, 1977–2005. Am Heart J 157:467–473. https://doi.org/10.1016/j.ahj.2008.10.017
Hammiri AE, Drighil A, Benhaourech S (2016) Spectrum of cardiac lesions associated with Isolated Cleft Mitral Valve and their impact on therapeutic choices. Arq Bras Cardiol 106:367–372. https://doi.org/10.5935/abc.20160053
Penny DJ, Vick GR (2011) Ventricular septal defect. Lancet 377:1103–1112. https://doi.org/10.1016/S0140-6736(10)61339-6
Vincentz JW, Barnes RM, Firulli AB (2011) Hand factors as regulators of cardiac morphogenesis and implications for congenital heart defects. Birth Defects Res A Clin Mol Teratol 91:485–494. https://doi.org/10.1002/bdra.20796
Zhou YM, Dai XY, Qiu XB, Yuan F, Li RG, Xu YJ, Qu XK, Huang RT, Xue S, Yang YQ (2016) HAND1 loss-of-function mutation associated with familial dilated cardiomyopathy. Clin Chem Lab Med 54:1161–1167. https://doi.org/10.1515/cclm-2015-0766
Morin S, Pozzulo G, Robitaille L, Cross J, Nemer M (2005) MEF2-dependent recruitment of the HAND1 transcription factor results in synergistic activation of target promoters. J Biol Chem 280:32272–32278. https://doi.org/10.1074/jbc.M507640200
Thattaliyath BD, Livi CB, Steinhelper ME, Toney GM, Firulli AB (2002) HAND1 and HAND2 are expressed in the adult-rodent heart and are modulated during cardiac hypertrophy. Biochem Biophys Res Commun 297:870–875. https://doi.org/10.1016/s0006-291x(02)02297-0
Knofler M, Meinhardt G, Vasicek R, Husslein P, Egarter C (1998) Molecular cloning of the human Hand1 gene/cDNA and its tissue-restricted expression in cytotrophoblastic cells and heart. Gene 224:77–86. https://doi.org/10.1016/s0378-1119(98)00511-3
Riley P, Anson-Cartwright L, Cross JC (1998) The Hand1 bHLH transcription factor is essential for placentation and cardiac morphogenesis. Nat Genet 18:271–275. https://doi.org/10.1038/ng0398-271
McFadden DG, Barbosa AC, Richardson JA, Schneider MD, Srivastava D, Olson EN (2005) The Hand1 and Hand2 transcription factors regulate expansion of the embryonic cardiac ventricles in a gene dosage-dependent manner. Development 132:189–201. https://doi.org/10.1242/dev.01562
Breckenridge RA, Zuberi Z, Gomes J, Orford R, Dupays L, Felkin LE, Clark JE, Magee AI, Ehler E, Birks EJ et al (2009) Overexpression of the transcription factor Hand1 causes predisposition towards arrhythmia in mice. J Mol Cell Cardiol 47:133–141. https://doi.org/10.1016/j.yjmcc.2009.04.007
Zheng SQ, Chen HX, Liu XC, Yang Q, He GW (2021) Identification of variants of ISL1 gene promoter and cellular functions in isolated ventricular septal defects. Am J Physiol Cell Physiol 321:C443–C452. https://doi.org/10.1152/ajpcell.00167.2021
Zeng ZH, Chen HX, Liu XC, Yang Q, He GW (2022) Functional significance of novel variants of the MEF2C gene promoter in congenital ventricular septal defects. Am J Med Genet a 188:2397–2405. https://doi.org/10.1002/ajmg.a.62871
Zuo JY, Chen HX, Liu ZG, Yang Q, He GW (2022) Identification and functional analysis of variants of MYH6 gene promoter in isolated ventricular septal defects. BMC Med Genom 15:213. https://doi.org/10.1186/s12920-022-01365-y
Zheng SQ, Chen HX, Liu XC, Yang Q, He GW (2021) Genetic analysis of the CITED2 gene promoter in isolated and sporadic congenital ventricular septal defects. J Cell Mol Med 25:2254–2261. https://doi.org/10.1111/jcmm.16218
Yin XY, Chen HX, Chen Z, Yang Q, Han J, He GW (2013) Genetic variants of ISL1 gene promoter identified from congenital tetralogy of Fallot patients alter cellular function forming disease basis. Biomolecules. https://doi.org/10.3390/biom13020358
Yin XY, Chen HX, Chen Z, Yang Q, Han J, He GW (2022) Identification and functional analysis of genetic variants of ISL1 gene promoter in human atrial septal defects. J Gene Med 24:e3450. https://doi.org/10.1002/jgm.3450
Zuo JY, Chen HX, Yang Q, Liu ZG, He GW (2023) Tetralogy of Fallot: variants of MYH6 gene promoter and cellular functional analyses. Pediatr Res. https://doi.org/10.1038/s41390-023-02955-x
Chen Z, Chen HX, Hou HT, Yin XY, Yang Q, He GW (2023) Identification and functional verification of CITED2 gene promoter region in patients with patent ductus arteriosus. Int J Mol Sci. https://doi.org/10.3390/ijms242216204
Chen Z, Chen HX, Hou HT, Yin XY, Yang Q, He GW (2022) Pathophysiological role of variants of the promoter region of CITED2 gene in sporadic tetralogy of Fallot patients with cellular function verification. Biomolecules. https://doi.org/10.3390/biom12111644
Chen Z, Chen HX, Hou HT, Yin XY, Yang Q, Han J, He GW (2022) Genetic variants of CITED2 gene promoter in human atrial septal defects: case-control study and cellular functional verification. J Cardiovasc Dev Dis. https://doi.org/10.3390/jcdd9100321
Castro-Mondragon JA, Riudavets-Puig R, Rauluseviciute I, Lemma RB, Turchi L, Blanc-Mathieu R, Lucas J, Boddie P, Khan A, Manosalva PN et al (2022) JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res 50:D165–D173. https://doi.org/10.1093/nar/gkab1113
Reamon-Buettner SM, Ciribilli Y, Inga A, Borlak J (2008) A loss-of-function mutation in the binding domain of HAND1 predicts hypoplasia of the human hearts. Hum Mol Genet 17:1397–1405. https://doi.org/10.1093/hmg/ddn027
Reamon-Buettner SM, Ciribilli Y, Traverso I, Kuhls B, Inga A, Borlak J (2009) A functional genetic study identifies HAND1 mutations in septation defects of the human heart. Hum Mol Genet 18:3567–3578. https://doi.org/10.1093/hmg/ddp305
Wang J, Lu Y, Chen H, Yin M, Yu T, Fu Q (2011) Investigation of somatic NKX2-5, GATA4 and HAND1 mutations in patients with tetralogy of Fallot. Pathology 43:322–326. https://doi.org/10.1097/PAT.0b013e32834635a9
Cheng Z, Lib L, Li Z, Liu M, Yan J, Wang B, Ma X (2012) Two novel HAND1 mutations in Chinese patients with ventricular septal defect. Clin Chim Acta 413:675–677. https://doi.org/10.1016/j.cca.2011.10.014
Oudelaar AM, Higgs DR (2021) The relationship between genome structure and function. Nat Rev Genet 22:154–168. https://doi.org/10.1038/s41576-020-00303-x
Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, Chen X, Taipale J, Hughes TR, Weirauch MT (2018) The human transcription factors. Cell 172:650–665. https://doi.org/10.1016/j.cell.2018.01.029
Vasicek R, Meinhardt G, Haidweger E, Rotheneder H, Husslein P, Knofler M (2003) Expression of the human Hand1 gene in trophoblastic cells is transcriptionally regulated by activating and repressing specificity protein (Sp)-elements. Gene 302:115–127. https://doi.org/10.1016/s0378-1119(02)01096-x
Acknowledgements
We thank the patients and their family members for their collaboration. The assistance of nursing staff at the Division of Pediatric Cardiac Surgery, Department of Cardiovascular Surgery is gratefully acknowledged.
Funding
This work was supported by the National Natural Science Foundation of China [82170353 & 82370350], Tianjin Municipal Science and Technology Commission (22ZYQYSY00020), the Non-profit Central Research Institute Fund of Chinese Academy of Medical Sciences (2020-PT310-007), and TEDA International Cardiovascular Hospital Internal Grant (2021-ZX-002), Tianjin Key Medical Discipline (Specialty) Construction Project [TJYXZDXK-019A].
Author information
Authors and Affiliations
Contributions
G.-W.H. and H.-X.C. concepted and designed of the studies; J.-L.Q., H.-X.C. and H.-T.H. conducted experiments; J.-L.Q., H.-X.C., H.-T.H., Q.Y. and G.-W.H. analyzed the data; Q.Y. and G.-W.H. explained the experimental results; J.-L.Q. prepared the figures; J.-L.Q. and G.-W.H. drafted the manuscript; the manuscript was edited and revised by G.-W.H.; G.-W.H. supervised the study; the final version of the manuscript was approved by J.-L.Q., H.-X.C., H.-T.H., Q.Y., Z.C. and G.-W.H. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of TEDA International Cardiovascular Hospital (Clinical Research Ethics Review Number: [2021]-0715-4).
Consent to participate
All subjects participating in the study have obtained written informed consent from their parents or guardians.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qi, JL., Chen, HX., Hou, HT. et al. Molecular and cellular role of variants of the promoter region of HAND1 gene in sporadic and isolated ventricular septal defect. Mol Cell Biochem (2024). https://doi.org/10.1007/s11010-024-05088-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11010-024-05088-9