Abstract
Triple-negative breast cancer (TNBC) is a highly aggressive form of breast cancer associated with poor prognosis, higher grade, and a high rate of metastatic occurrence. Limited therapeutic interventions and the compounding issue of drug resistance in triple-negative breast cancer warrants the discovery of novel therapeutic targets and diagnostic modules. To this view, in addition to proteins, lipids also regulate cellular functions via the formation of membranes that modulate membrane protein function, diffusion, and their localization; thus, orchestrating signaling hot spots enriched in specific lipids/proteins on cell membranes. Lipid deregulation in cancer leads to reprogramming of the membrane dynamics and functions impacting cell proliferation, metabolism, and metastasis, providing exciting starting points for developing lipid-based approaches for treating TNBC. In this review, we provide a detailed account of specific lipidic changes in breast cancer, link the altered lipidome with membrane structure and mechanical properties, and describe how these are linked to subsequent downstream functions implicit in cancer progression, metastasis, and chemoresistance. At the fundamental level, we discuss how the lipid-centric findings in TNBC are providing cues for developing lipid-inspired theranostic strategies while bridging existing gaps in our understanding of the functional involvement of lipid membranes in cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The plasma membrane is the most fundamental cellular component, which is now visualized, not just as a homogenous protein-enriched lipid bilayer providing structure but also as a platform for orchestrating the interactome of signaling proteins and lipids [1]. Various cellular processes such as membrane fusion, fission, endocytosis, protein trafficking, and exchanges with the external environment are initiated at the very heart of the plasma membrane within heterogeneous lipid domains [2]. Some of these domains are ordered and known as liquid-ordered (lo) lipid domains that reside within the sea of liquid-disordered lipid domains (ld). Lipid-ordered domains are also referred to as lipid rafts; however, due to conflicting views regarding the definition of lipid rafts, we prefer to use only the term ordered lipid domains throughout this work. Lipid domains house complex and dynamic protein-lipid structures, including distinct lipids and proteins [3]. Ordered lipid domains are relatively rigid compared to the surrounding region due to the presence of saturated lipids and cholesterol within their lipid pools. Casares et al. recently reviewed the role of membrane lipid composition within the plasma membrane (PM) and endomembrane compartments that are critical for sustaining organelle structure and function by regulating the abundance, size, and nature of lipid domains [4]. Hence, the lipid environment and proteins have an immense influence on most cellular functions, and the precise configuration and architecture of cell membrane and lipid domains delineate the fundamental pathophysiological aspects of cells. Pathological conditions, such as cancer or infection, bring about exceptional alterations in the lipid repertoire of cells [5,6,7] underpinning how membrane lipid composition changes, and their associated physicochemical properties, are significant enough to be regarded as biomarkers [8]. Aptly, these are considered potential targets for tangential therapeutic approaches [9, 10].
Breast cancer is a highly heterogeneous form of cancer known to have life-threatening consequences for women across the globe. Molecular classification of breast cancer based on therapy constitutes the following three major types (which are also further divided into subtypes): hormone-positive (estrogen (ER)- and progesterone (PR)-positive tumors), HER2-positive (positive for the human epidermal growth factor receptor 2 (HER2)) and triple-negative breast cancer (TNBC), which does not express ER, PR, or HER2 [11]. While there is some relief in the fact that the first two can be treated with hormonal therapies and chemotherapy, TNBC is exceptional in being an aggressive type of cancer, with a distinct phenotype expressing high invasiveness, high metastatic potential, predisposition to relapse, and poor prognosis [12]. TNBC can be subdivided into androgen receptor (AR)-positive TNBC and AR-negative TNBC, also known as quadruple-negative breast cancer (QNBC). QNBC is an even more aggressive form of cancer, and there is a dearth of prognostic biomarkers and therapeutic targets for it [12]. Compared with other subtypes of breast cancer, the survival time of TNBC patients is shorter, with a high probability and faster rate of metastasis to distant organs and a short relapse time [13]. Furthermore, TNBC tumors are not receptive toward endocrine therapy or molecular targeted therapy (as they lack ER, PR, and HER2 expression), and dedicated TNBC treatment protocols are still under-developed. Therefore, it is imperative to discover new biomarkers and therapeutic targets for breast cancer, particularly TNBC, to assist diagnosis and prognosis after surgery and to develop new and superior therapies.
Lipid alterations in cancer are known to vary for different stages of progression and cancer types [5]. Importantly, elaborating the ever-evolving role of lipids in reprogramming membrane dynamics for cancer proliferation, cell metabolism, and metastasis holds a huge promise for developing a lipid-based or lipid-targeted approach toward treating TNBC. This review describes how in recent years, cell membrane composition, organization, and nanomechanical properties have been correlated with the malignancy and invasiveness of breast cancer, and hence, are considered as orthogonal biomarkers for breast cancer diagnosis and prognosis. We also summarize some of the lipid-based/targeted therapeutic advancements in the last years. Overall, the lipidomic biomarkers can play a crucial role in prognostic characterization, diagnosis, and anticipation of the scope of available anticancer therapeutics.
Lipid composition, structural attributes, and biophysical properties of cellular membranes
Major lipid constituents of cell membranes
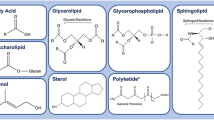
Major structural constituents of eukaryotic membranes are glycerophospholipids (GPL), sphingolipids (SL), and cholesterol (Chol) [4] (Fig. 1). Less abundant are the signaling-induced hydrolysis products of GPLs and SLs (such as lyso-P, and diacylglycerol, DAG); e.g., messenger lipids having distinct functions in signaling processes. GPLs are further classified on the basis of the phospholipid head group structure, which could be zwitterionic or charged. Chol is a crucial and abundant structure-forming unit of eukaryotic cell membranes. The rigid steroid backbone of Chol favors its interaction with SL [14], rendering Chol-SL platforms as important elements of some lipid domains [4] (Fig. 1). Chol and SL-enriched lipid domains harbor various receptor proteins, and these domains contribute to the specific membrane properties. As recently reviewed, such lipid domains are considered plausible therapeutic targets against different types of cancer [15, 16]. Variable levels of membrane order within lipid domains correlate to different levels of Chol-infusion that enable the regulation of membrane fluidity. The lipid domain surrounding the ld region has higher fluidity, less density, and increased dynamics due to the increased abundance of unsaturated phospholipids [17] (Fig. 1). Along with the structural role, lipids also influence membrane protein localization within membranes. Membrane proteins act as the receptors, channels, transporters [18] and are involved in signal transduction; some even act as enzymes [19, 20]. For the proper functioning of these proteins, the distinct interactions with membrane lipids (within lipid domains) are of utmost significance. Consequently, any destabilization of lipid domains is likely to impact membrane protein functions and hence lead to severe complexities and often a pathological condition [21,22,23,24].
Steady-state composition of cellular membranes, site of lipid synthesis, and lipid domain. The central panel shows the lipid composition of different cellular membranes, including plasma and internal membranes; data adapted from [37]. The site of synthesis of the major phospholipids (red: ER; brown: plasma membrane; gray: mitochondria; green: Golgi) along with lipid identity are shown. The most common glycerophospholipids assembled in the endoplasmic reticulum (ER) are phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylinositol (PI), phosphatidylserine (PS), and phosphatidic acid (PA). ER also synthesizes ceramide, galactosylceramide (GalCer), and cholesterol (chol). The Golgi is the site of synthesis of sphingomyelin (SM), complex glycosphingolipids (GSLs), and PC. Mitochondria synthesize almost 45% of phospholipids, including PE, PA, phosphatidylglycerol (PG), and cardiolipin (CL). The lipid composition of different membranes varies throughout the cell, and bar graphs show the composition of the total phospholipid for each membrane type in a mammalian cell. Plasma membranes consist of lipid domains enriched in saturated lipids, cholesterol, and proteins forming ordered lipid domains. Actin cytoskeleton underneath the plasma membrane is also involved in the stability of protein-lipid domains. These ordered domains are surrounded by fluid lipid domains. Another major characteristic feature of a normal and healthy plasma membrane is the lipid asymmetry between the two leaflets of the lipid bilayer, where charged lipids such as PS and PE are preferentially localized in the cytoplasmic leaflet. The lipid structure (relative size of the head and tail group area) determines its molecular geometry that dictates membrane curvature; membrane proteins also contribute substantially to the stabilization of membrane curvature. In the absence of proteins and in pure lipid bilayers, while PC forms membrane with no curvature, PE and lyso-lipids form negatively and positively curved membranes, respectively (bottom, inset). PI(3,5)P2 phosphatidylinositol-(3,5)-bisphosphate; PI(4,5)P2 phosphatidylinositol-(4,5)-bisphosphate; PI(3,4,5)P3 phosphatidylinositol-(3,4,5)-trisphosphate; PI4P phosphatidylinositol-4-phosphate; S1P sphingosine-1-phosphate; Sph sphingosine; R remaining lipids
The supramolecular assembly in lipid membranes involves the lamellar arrangement of lipids, which can get further organized into more structured varieties such as solid-ordered (so), gel-phase (Lβ), ripped-phase, or liquid-ordered phase (lo); the liquid crystalline/fluid phase (Lα) being the most physiologically relevant [25,26,27,28]. The lipid membrane can also acquire hexagonal or cubic lipid phases owing to the abundance of lipids such as phosphatidylethanolamine (PE), diacylglycerol (DAG), and acidic phosphatidylserine (PS) [3, 29, 30]. These lipids can induce curvature in the context of the cellular membrane (i.e., in vivo) as a result of their non-cylindrical shapes. Lipids such as lysophospholipids with big polar heads induce a positive curvature in the membranes, forming normal hexagonal monolayers (HI) or micelles [31, 32] (Fig. 1). DAG and PE induce negative curvature stress into the cell membrane and induce the formation of inverted hexagonal structure monolayers (HII) or inverted micelles that are known to play a crucial role in budding, fusion, and fission [2, 5, 30, 33, 34]. The presence of these specific regions can ultimately be responsible for the activation of key membrane proteins involved in various cell signaling pathways and their deregulation in pathological states.
Cellular lipid pool is organelle-specific
Membrane composition between organelles is highly divergent and mediates the organelle-specific function facilitated by the distinct conformation of the membranes. Thickness and stiffness are the two main physical parameters that invariably discriminate between the functions and identities of different secretory organelles like endoplasmic reticulum (ER), Golgi apparatus, and plasma membrane (PM) [4]. There exist a gradient of SL and Chol concentrations in ER and Golgi, with the highest abundance in PM that underlines its thickness and stiffness [5]. The ER provides a large percentage of membrane lipids for Golgi and PM [35] and produces significant levels of the structural phospholipids (PL) and Chol, together with non-structural lipids such as triacylglycerol cholesteryl esters and ceramide (Cer) [36]; Cer is the precursor for SL, Fig. 1. The rapid transference of Chol to other organelle decreases its concentration in ER, leading to enhanced ER membrane fluidity. Furthermore, the low Chol content-enabled loose lipid organization regulates ER function as it eases the insertion and the transport of lipids and proteins [37]. ER also houses minor lipids such as DAG, PA, and lysophospholipids [37] that induce membrane curvature, decrease the energy required for membrane fission, fusion, pore formation, and vesicle trafficking, and eventually assist in ER functions [34]. The trans-Golgi network, which produces complex sphingolipids utilizing Cer from ER, plays a significant role in the transition from the thin and loosely packed membrane found in the ER to the thick and rigid bilayer found at the PM [4]. It also transports proteins with variable transmembrane domains to the PM.
Mitochondria is a specialized organelle that derives its lipids mainly from mitochondria-associated membrane subfraction of the ER and is responsible for the synthesis of unique lipids like cardiolipin (CL), along with phosphatidylglycerol (PG), PE, PA, and DAG [38]. Mitochondrial lipid composition is marked by high levels of PC and PE (almost 80% of total PL), high CL, and low sterol and SL concentrations (Fig. 1). Mitochondrial membranes of the nervous, immune, and cardiovascular systems also contain PC and PE plasmalogens [39]. Plasmalogens are membrane structure lipids that contain a vinyl-ether and ester bond at the sn-1 and sn-2 positions, respectively, in the glycerol backbone [40]. This unique structural feature endows highly lipophilic characteristics to plasmalogens allowing them to generate inverse hexagonal phase structures that promote membrane fusion [39, 41]. Plasmalogens play an important role in the organization of stable lipid microdomains and Chol-rich membrane scaffolds and articulating cellular signaling while also serving as endogenous antioxidants and protecting other PL, lipid, and lipoprotein particles from oxidative stresses [40].
The plasma membranes of various cell types have different and unique lipid domains of variable order and fluidity [42,43,44,45,46,47]. While the ordered microdomains, sometimes also referred to as lipid rafts (lo), are rigid, tightly packed, and abundant in Chol, SL, and GSLs. Some of the disordered domains (ld) have PE-rich regions and are comprised of unsaturated FAs (Fig. 1); both of these lipid regions are enriched with a distinct set of membrane proteins. This culminates in enhanced membrane fluidity and reduced surface packing density in ld membrane regions reported both in vitro [48,49,50,51] and in cellular native membranes [47, 52, 53]. Various biological processes, including signal transduction, membrane trafficking, and immune responses are regulated by the specific properties of these lipid microdomains [54].
How do membrane properties govern cellular functions?
Lipid composition, spatiotemporal membrane arrangement, and membrane proteins dictate the biophysical properties of the cell membrane. One of the most significant properties is membrane fluidity, which is inversely related to membrane viscosity and has a huge impact on the dynamics of traversing molecules within the membrane [54, 55]. The optimal diffusion of proteins and lipids within the membrane plane enables efficient interactions between them and regulates lipid/protein clustering, which directly impacts the assembly of signaling hot spots. Membrane heterogeneities and lipid domains of distinct composition strongly control membrane fluidity, which in turn governs downstream cellular signaling, lipid-protein interactions, and cell communication [17, 37, 56]. Alterations in membrane fluidity have been reported in the pathogenesis of various diseases such as cancer, obesity, and neurodegenerative disorders by modulating numerous membrane-associated signaling pathways [57, 58]. Membrane fluidity also affects the interaction of chemotherapeutic drugs with membranes and has been linked to the ever-increasing problem of drug resistance in cancer cells [59].
PM has high heterogeneity and asymmetry in the lipid composition and spatial organization. The atomistic view of membrane heterogeneity or phase coexistence, phase transitions, and specific details of interactions within the lipid microdomains have been recently shown by Gu et al. [60]. They simulated binary (DPPC, DOPC) and ternary lipid mixtures (DOPC, DPPC, and Chol), showing Lα–gel/ripple, lo–ld, and gel–lo coexistence. This work also highlighted specific interaction geometries between Chol and PLs, which serve as new insights on possible driving forces for lipid phase separation. Experimentally, use of membrane sensitive probes [17, 61] and technologies with an advanced spatial and temporal resolution, such as super-resolution stimulated emission depletion (STED) microscopy [62], label-free Raman microscopy [63], high dimensional super-resolution imaging [64], and cryo-EM imaging [65] have opened new avenues for gauging the lateral heterogeneity in native membranes (both in vitro and in vivo) with unambiguous accuracy of their functional relevance.
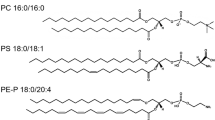
Asymmetry in lipid distribution between the two leaflets of the plasma membrane (PM) is well known and crucial for cellular processes [6, 42, 66]. While the intracellular leaflet of the PM is composed of higher concentrations of anionic phosphatidylserine (PS) and phosphatidylethanolamine (PE), the exoplasmic leaflet is rich in zwitterionic phosphatidylcholine (PC) and sphingomyelin (SM). Skotland and Sandvig reviewed the possible interactions between the very long-chain sphingolipids in the outer leaflet of the plasma membrane and the PS (18:0/18:1) in the inner leaflet and the role that cholesterol plays for such associations [67]. The inner leaflet is also enriched with signaling lipids like phosphatidylinositol (PI) or phosphatidic acid (PA). Membrane asymmetry causes a negative inner membrane surface charge that drives hydrolysis of PI, by means of phospholipase C, into inositol 1,4,5-triphosphate (IP3) and DAG second messengers [68]. Furthermore, lipid rafts, caveolae, receptors, and channel clusters of higher-order asymmetry facilitate cellular functioning by balancing electrostatic interactions and lateral diffusion within the cell membrane. The asymmetric distribution of PLs between both the leaflets of the membrane and, therefore, the analogous effect on membrane electrostatics are controlled by the Golgi apparatus. Finally, the asymmetric partitioning of lipids in the membrane is an active process and requires flippases, floppases, and scramblases [69]. These are proteins that differently translocate lipids from one leaflet to another. For example, flippases are ATP-consuming proteins that move the aminophospholipids PS and PE from the PM outer leaflet to the inner leaflet. Floppases are also ATP-dependent and transport substrates such as PC, SL, and Chol from the inner to outer PM leaflet [70]. Scramblases, on the other hand, are ATP-independent and Ca2 + -dependent transporters that facilitate the passive transit of lipids in either direction [69].
Peculiar case of Cancer Cell Membrane Structure
Membrane landscape in cancer
An altered lipid repertoire due to impaired lipid metabolism impacts the spatial–temporal arrangement of lipids within membranes. Foremost is the lipid asymmetry that undergoes major rearrangements, wherein its disorganization rewires cell signaling [25]. In several types of cancer, a loss of asymmetric lipid distribution has been observed, which results in the exposure of the negatively charged PS and PE on the outer surface of their membranes [68], Fig. 2A. A study by Vallabhapurapu et al. explored the mechanisms regulating the surface PS exposure in human cancer cells and found that differential flippase activity and intracellular calcium are the major regulators of surface PS exposure in viable human cancer cells [71]. Cancer cell lines with high surface PS exhibited low flippase activity and high intracellular calcium. The surface exposure of PS was observed to be of different extents in different cancer cell lines. Strikingly, breast cancer cells had a low exposure of PS, while the metastatic breast cancer cells had a higher surface PS, emphasizing that variable lipid compositions are observed during different stages of cancer progression. Due to the high levels of exposed PS in cancer cells, an immunosuppressive environment is created [72,73,74] that enables cells to evade the inflammatory immune response and promotes proliferation [75], a form of non-classical apoptosis mimicry [76].
Representation of altered lipid landscape in breast cancer. A Uptake of exogenous lipids and de novo lipid synthesis are major sources of cellular lipid pools in non-TNBC and TNBC cancers. Fatty acids (FAs) provide substrates for membrane synthesis and metabolic fuels through β-oxidation. In addition, specific to TNBC, expression of enzymes involved in acquiring exogenous FAS such as lipoprotein lipase (LPL), very low-density lipoprotein receptor (VLDLR) expression is elevated, contributing to deregulated lipidome. FA binding and activating proteins such as FABP, FABP7, and long-chain acyl CoA synthetase (ACSL)-4, respectively, are overexpressed, contributing to altered lipid landscape in TNBC. The excess lipids in tumors are stored in lipid droplets (LDs), and in the case of TNBC, the LDs are enriched with specific proteins such as Lipin-1, PLA2, and COX-2. Cancer cells display a loss of PM bilayer asymmetry, i.e., re-distributed amounts of PE and negatively charged PS lipids at the extracellular bilayer leaflet due to reduced flippase activity, eventually destroying asymmetric PE/PS distribution. Lack of PE at the cytoplasmic PM leaflet destabilizes membrane proteins, modulating signaling events in TNBC. PDB code of the depicted human PE binding proteins, 1BEH. PDB codes of transmembrane proteins shown are 2MFR (insulin receptor, red) and 4PYP (GLUT-1, blue). The enhanced PE abundance at the exoplasmic site provides sites for hypervesiculation in TNBC, leading to enhanced induction of extracellular membrane vesicles. ACC, acetyl-CoA carboxylase; ACLY, ATP citrate lyase; FABPs, fatty acid-binding proteins; FASN, fatty acid synthase; PUFA, polyunsaturated FA; SFA, saturated FA. B Representation of relative changes (not to scale) of the deregulated lipid classes in TNBC compiled from primary studies [107,108,109,110,111,112,113] to depict changes within various lipid classes on the same graph. C Schematic representation of the alteration in the plasma membrane organization and bilayer deformation in TNBC, concomitant with global lipid rewiring and implication in drug resistance. The loss of lipid asymmetry in cancer cell membranes due to the presence of negative charged phospholipids (PLs) impact membrane lipid domain organization. Reduced cholesterol (Chol) and SM levels modulate the membrane deformability and alter actin cytoskeleton. The reduced abundance of actin stress fibers that harbors actin-myosin interaction sites are reduced in cancer cells, subsequently modulating cell mechanical properties such as stiffness
The exposure of PE enhances membrane curvature that aids in hypervesiculation, leading to a release of extracellular membrane vesicles (MVs) implicated in cancer metastasis (Fig. 2A). PE also acts as a lipid chaperone by assisting in the folding of membrane proteins [77]. A lack of PE on the PM cytosolic side leads to the accumulation of unfolded/misfolded proteins and induces chronic endoplasmic reticulum (ER) stress, which further shifts the metabolism toward a cancerous phenotype [78,79,80]. PE is a fine regulator of autophagy, and its altered abundance affects the clearing process of malfunctioning organelles that eventually promote carcinogenesis [80]. Furthermore, as a part of the glycosylphosphatidylinositol (GPI) anchor, PE is responsible for binding proteins to the membrane, and impairment of this function leads to the destabilization of the membrane proteins [81] (Fig. 2A). As a result, downstream intracellular signaling events, metabolism, and communication are modulated, leading to cellular imbalance and carcinogenesis. In addition, lysophospholipids are also abundant in cancer cells which are attributed to the enhanced activity of phospholipase A2. The increased lysophospholipid content leads to the destabilization of the membrane and irregular membrane curvature [82] making it more susceptible to leakage and fusion.
Phosphatidylcholine (PC) is the most abundant structural component of the lipid bilayer and is mostly present in the outer PM leaflet [83]. It is also a precursor of signaling molecules such as DAG, arachidonic acid, and lyso-PC produced through a network of enzymatic reactions known as the phosphatidylcholine cycle. In the case of cancer, the PC content in the cell membrane increases by the induction of the Kennedy pathway, PE N-methyltransferase, and CDP-choline pathway [6, 84]. Moreover, its degradation to release signaling messengers for the proliferating cancer cells also increases by upregulation of enzymes such as choline kinase alpha (CHK@), phospholipase C (PLC), or phospholipase D (PLD) [85]. PC-specific phospholipase C (PC-PLC) acts as a chaperone for HER2 receptor, and its levels are 3- to sixfold elevated in breast cancer cells [86]. Confocal microscopy in vivo confirmed that PC-PLC extensively colocalizes with HER2 in the raft domains of SKBr3 breast cancer cell PM, and coimmunoprecipitation assays substantiated a physical association of PC-PLC with HER2 [87]. Therefore, PC-PLC inhibition comes across as a potential target against breast cancer as its inhibition leads to the substantial and long-lasting inhibition of HER2. DAG, a hydrolysis product of PC, has a profound effect on membrane structure and signaling. A local accumulation of DAG disorganizes phospholipid bilayer as the characteristic negative curvature of DAG, and its lack of charge induces unstable and asymmetric regions in the membrane bilayer. This facilitates fusion and fission interactions, induces the formation of microdomains, and alters protein-lipid interactions, de-routing the signaling cascades [88]. DAG and lyso-PA are also known to stimulate cell migration in cancer cells [4]. Finally, changes in the lipid composition are related to the malignancy of the tumor. For example, the pro-apoptotic Cer is reduced in tumors, and its levels are inversely associated with tumor progression. Other notable changes include the aberrant distribution of immunosuppressing gangliosides and the accumulation of esterified Chol levels, which are inadvertently associated with increased cell-cycle progression and tumor growth.
How does altered cell membrane impact drug resistance?
Lipid asymmetry is implicated in multidrug resistance in cancer cells [89] by virtue of altered lipid packing. Lipid packing is a balance between repulsive long-range forces between the lipid head groups and attractive-hydrophobic interaction between the lipid acyl chains. With negatively charged lipids, such as abundant PS in the inner leaflet, acidic pH decreases repulsion between their polar head groups and reduces the surface tension, leading to reduced area per lipid. This eventually forms a loosely packed membrane in non-transformed cancer cells. In the case of drug-resistant cells, a small increase in alkalinity at the cytoplasmic side reduces the neutralization of charges by H + ions and increases the repulsion of polar head groups, thereby increasing the surface tension and optimal area per lipid. This leads to tighter packing, eventually reducing permeability toward chemotherapeutics [89]. In a recent dynamic molecular simulation study of model membranes, the effect of lipid asymmetry and cholesterol content has been shown to reduce the cell permeability toward anticancer drugs such as cisplatin [90]. This study shows how these membrane lipid properties play a role in mechanically resisting anticancer drug uptake.
In an independent study, Peetla et al. investigated the role of lipids in drug transport in cancer chemotherapy to overcome drug resistance in breast cancer cells [91]. In this study, they isolated lipids from doxorubicin-sensitive (MCF7) and resistant (MCF7/ADR) breast cancer cells to characterize the biophysical properties of membrane lipids (particularly lipid packing and membrane fluidity) and to understand the role of the interaction of cell membrane lipids with drug/nanocarrier on drug uptake and efficacy. Resistant cell membrane lipids showed significantly different compositions and formed more condensed, less fluid monolayers than did lipids from sensitive cells. Similarly, the cancer cells which undergo metastasis reduce membrane Chol levels to enhance membrane fluidity and elasticity. This makes the cancer cells more resilient for permeation through blood vessels [5].
Rewiring of Lipid Metabolism in Cancer
Cancer cells, for their rapid proliferation, require membrane biogenesis which they achieve by reprogramming the inherent lipid metabolism. In response to the changes in the microenvironment, in terms of differential availability of oxygen and nutrients, most cancer cells exhibit aberrant stimulation of lipid metabolism. They respond to this metabolically challenging environment by striking a new balance between the endogenous synthesis and exogenous uptake of fatty acids in order to meet the growing need for membrane biogenesis, energy production, and protein modification [92, 93]. Alterations in lipid metabolism bring about changes in membrane composition and membrane dynamics which have several pathological implications. A detailed description of the fate of lipid metabolism through varied pathways has been demonstrated by Szalsa et al. in different types of carcinogenesis [6].
A predominant feature of lipid metabolic adaptation is the elevated de novo synthesis of fatty acids (FAs) in cancer cells which otherwise have a pathway of minor importance (with exceptions of liver and mammary cells) in healthy cells (Fig. 2A). This meets the increasing requirement of the proliferating cells and is a phenomenon observed in many different cancer types [94, 95]. In this context, it has been demonstrated that in breast tissue, the FA and PL profiles are altered not only in the tumor areas but also in the surrounding healthy areas [96]. This provides evidence that changes in PL composition can occur before morphological changes. The authors observed significantly higher amounts of the n-3 FA DHA in cancer tissue compared to the other tissues. Upregulation of FA synthase and the subsequent biogenesis of saturated fatty acids (SFA) initiating from palmitic acid (C16) play a key role in the synthesis of saturated membrane phospholipids that imparts rigidity to the membrane bilayer. The subsequent transformation to mono- and polyunsaturated fatty acids (MUFAs and PUFAs) by specific enzymatic machinery (desaturase and elongase) leads to the coordination of fluidity and permeability to the membrane structure. The balance between SFA, MUFA, and PUFA gets greatly altered in different cancer types, leading to a modified FA pool (Fig. 2A). In fact, the SFA-MUFA transformation is regarded as an important biomarker of cancer status [67].
Stearoyl CoA desaturase (SCD-1), an enzyme associated with SFA-MUFA transformation specifically acting on palmitic and stearic acid, is overexpressed in cancer cells and regulated by signaling cascades such as MAPK (mitogen-activated protein kinase) and AKT-favored in cancer [97, 98]. Elevated levels of SCD-1 are regarded as a biochemical signature of cancer cells and tumor tissues in lung, breast, colon, kidney, and prostate cancers [98, 99]. Additionally, lipogenesis is also upregulated due to increased mTORC2 activity, promoting tumorigenesis via increased production of glucosylceramide and CL as observed in hepatic cancer cells. Changes in the microenvironment, such as low oxygen availability or hypoxia, have several implications. Under a hypoxic environment, it has been reported that the ratio of MUFA (C18:1/C16:1) to SFA (C18:0/C16:0), also known as desaturation index, gets significantly reduced in many types of cancer cell lines (e.g., MDA-MB-468, HeLa, A549, MCF7,7 86–0, UMRC2, U87, and HCT116). The hypoxia-inducible factor (HIF1α) leads to an accumulation of lipid droplets which promote uptake of FAs through fatty acid-binding proteins rather than de novo FA synthesis [100]. It also induces the expression of acetyl-CoA synthetase (ACSS2), an enzyme that converts acetate to acetyl-CoA [101], for increasing the lipid availability in proliferating cancer cells. In addition to hypoxia, lipid depletion synergistically stimulates ACSS2 expression, depending upon the cancer cell-cycle stage [102].
Another strategy to increase the lipid mass in cancer cells is enhanced uptake of exogenous or dietary lipoproteins, driven by the upregulation of PI3K/AKT and subsequent storage of excess lipids within lipid droplets (LD; Fig. 2A). Lipid droplets are membrane-enclosed subcellular organelles acting as a reservoir for FAs and Chol in the form of triglycerides and cholesterol esters. The LD accumulation of cholesteryl esters in prostate cancer tissues and breast cancer cells has been associated with proliferation and aggressiveness [103, 104]. Apart from the usual route of SCD-1 dependent fatty acid desaturation, an SCD-1 independent and alternate desaturation pathway has recently been explored that generates the unusual FA, sapienate/sapienic acid (cis-6-C16:1) through activation of fatty acid desaturase-2 (FADS2) [105, 106]. Interestingly, in several cancer cell lines, including lung, breast, and prostate, desaturation through the FADS2 pathway has been observed in addition to SCD-1 dependent fatty acid desaturation. This pathway renders an alternative source of MUFAs and represents a novel route to expanding the pool of deregulated lipidome in cancer [95, 105]. The cumulated data underlines some of the most important alterations in FA compositions in various cancer types. Ferreri et al. have compiled a comprehensive FA landscape where they coordinated elevated levels of oleic acid, linolenic acid, arachidonic acid, pentadecanoic acid, heptadecanoic acid, 9C-C18:1 and 11c-C18:1 MUFAs, and n-6 C18:2 and C20:3 PUFAs with different carcinomas [95].
A substantial catabolic phenomenon that supports cancer cell metabolism is autophagy. Autophagy serves as lipolysis machinery for the degradation of lipids by recycling damaged organelles, cytosolic lipids, and thus supplies fatty acids, and acetyl-CoA for continued metabolism [107]. During autophagy, LDs encapsulated in double-membrane autophagosomes fuse with lysosomes, and undergoes degradation upon formation of autolysosomes. The degradation of LDs generates free-fatty acids that serve as a substrate for β-oxidation in mitochondria providing an efficient source of energy during cancer progression [108]. The autophagy-mediated selective degradation of lipids is called lipophagy and appears to be a well conserved mechanism of lipid degradation across several cell types. Although lipophagy plays a dual role (both pro and anticancer) in cancer metabolism upon regulation of lysosomal acid lipase expression, the relationship between lipophagy and cancer progression/ metastasis is, not fully understood and warrants more insights [109].
Lipid metabolic changes specific to breast cancer
Interactions between breast cancer and its microenvironment are highly complex given its heterogeneous nature, where the metabolic adaptions drive the metastatic process and chemoresistance. The expression level of various enzymes involved in different aspects of lipid metabolism is distinctly affected in different breast cancer types. For example, the de novo FA synthesis pathway, which is upregulated in most cancers, including HER2-positive breast cancer, appears not to be the chosen pathway in the case of TNBC [110]. Most enzymes of this pathway, such as ATP citrate lyase (ACLY), acetyl-CoA carboxylase alpha (ACACA), fatty acid synthase (FASN), and SCD-1, are overexpressed in HER2-enriched breast cancer cells, with extremely low levels in the case of TNBC (Fig. 2A). On the other hand, enzymes involved in acquiring exogenous FAS, such as lipoprotein lipase (LPL) and the very low-density lipoprotein receptor (VLDLR) expression, are elevated in TNBC (Fig. 2A). Similarly, FA binding proteins (FABP5 & FABP7) that bind long-chain FAs mediating their uptake into the cell and transportation to subcellular organelles, are overexpressed in TNBC and the basal-like subtype of breast cancer. Enzymes involved in FA activation, such as long-chain acyl CoA synthetase (ACSL-4), are also significantly overexpressed in TNBC and basal-like breast cancers, whereas ACSL-3 expression predominates in luminal and HER2-enriched subtypes. In fact, the tyrosine phosphatase, SHP2, which has been known to increase expression of ACSL-4 protein in MA-10 Leydig cells [111], also appears to be associated with breast cancer progression and increased motility in TNBCs [112, 113].
LD accumulation correlates well with increased breast cancer malignancy [114]. Raman imaging and spectroscopy have been used to detect the accumulation of LDs in MCF10A, MCF7, MDA-MB-231 cell lines [115]. Cytoplasmic LDs are considerably elevated in highly malignant MDA-MB-231 cells compared to moderately malignant MCF7 breast cells, with even lower levels in non-malignant MCF10A breast cells. The hypoxia-induced accumulation of lipids within LD is accompanied by an increase of LPIN1. LPIN1 regulates phospholipid synthesis by encoding lipin-1, a PA phosphatase (PAP). Lipin-1 is localized in LDs and drives the conversion of PA to DAG, the rate-limiting step in the PL and triglyceride synthesis. It was found to be highly upregulated in basal-like triple-negative breast cancer (TNBC; Fig. 2A). He et al. found that knockdown of LPIN (HCC1806 cells) significantly reduced PL synthesis as well as reduced CL, SM, and Cer levels and also considerably blocked tumor growth in an in vivo mouse xenograft tumor model [116]. Furthermore, knockdown of LIPIN-1 resulted in a reduced abundance of PLs having shorter acyl chains and an attenuated saturation index of membrane lipids. Taken together, this work suggests that PL synthesis can be a potential target for cancer therapy. Moreover, the other two enzymes known to be localized within LD and highly upregulated in TNBC are cytosolic PLA2α (PLA2G4A) and cyclooxygenase-2 (COX-2), both responsible for the synthesis of prostaglandins such as prostaglandin E2 (PGE2). The displacement of free-fatty acids from triglycerides in LDs is regulated by a class of proteins called perilipins (PLIN; Fig. 2A). It has been observed that PLIN2 is associated with increased LD formation, prolonged breast cancer cell survival in vitro [117], overexpression of basal-like TNBC, and downregulation of the ER, which increases expression.
Finally, elevated levels of PA have been associated with the activation of kinases and enhanced hypoxia-inducible factor 1-alpha (HIF1A) transcription, which promotes angiogenesis and cancer cell proliferation [118]. Hypoxic cells cause higher levels of glucose consumption, making the microenvironment acidic, which promotes invasiveness in cancer cells. Thus, different breast cancer cells and peculiarly TNBC cells rewire FA metabolism in most exclusive ways facilitating their survival, maintenance, and proliferation.
Transcriptional regulation of cancer lipid metabolism
Lipid homeostasis is transcriptionally regulated by various factors, which are modulated in various types of cancer. Carbohydrate-response element-binding protein (ChREBP), encoded by MLXIPL gene, acts as a transcriptional factor facilitating metabolism of excess glucose to FAs via formation of acetyl-CoA [119]. It also coordinates the carbohydrate induction of lipogenesis and is involved in pathogenesis of various cancers. For instance, ChREBP knockdown activates p53 to induce cell-cycle arrest and subsequently reduces colon cancer growth in vivo [120, 121]. In breast cancer, a positive correlation has been documented between ChREBP and lipogenic genes. Upstream of ChREBP is the liver X receptor (LXR) that directly regulates glycolysis and increases lipogenesis [122]. Removal of LXR reduces the abundance of key lipogenic genes and increases production of TGs [123]. LXRs are Chol sensors and LXR-driven genes that regulate Chol-efflux are less frequently expressed in breast cancer, leading to higher lipid contents and hence higher cancer cell viability [124]. In addition, thinner lipid rafts in the plasma membrane after LXR stimulation have been observed using AFM, consistent with its involvement in Chol modulation [125]. Activation of LXRs also decreases expression of lipogenic genes such as sterol regulatory element-binding proteins; SREBP1, SCD-1 and FASN in breast cancer [126]. Another transcriptional factor is farnesoid X receptor (FXR) that contributes toward maintenance of Chol homeostasis [127]. FXR has been shown to modulate the tumorigenic effect of lipids isolated from bone, impacting breast cancer metastasis [128]; however, more studies are required to explore the mechanisms behind FXR mediated lipid regulation in cancer. Another way of transcriptionally regulating Chol and other lipids is mediated by peroxisome proliferator-activated receptor (PPARs). PPAR downregulation decreases plasma TG levels, and increases lipase expression [129]. The delta isoform, PPARδ, directly impacts Chol levels by regulating the expression of genes that control Chol-efflux [130]. A rather ubiquitous transcriptional regulation of lipid metabolism is mediated by sterol regulatory element-binding proteins, SREBPs [131]; SREBPs activate expression of > 30 genes and contribute toward synthesis and uptake of Chol, FAs and TGs.
Lipidic Changes as Efficient Biomarkers in Breast Cancer
A significant variation is observed in the lipid composition and distribution among different cancer phenotypes compared to normal healthy cells; therefore, a lipidomic characterization is believed to serve as a reliable strategy to furnish effective biomarkers for detecting cancer. Lipidomics refers to the quantitative assessment of all the lipids in cells, tissue, body fluid, or organisms. Mass spectrometry (MS) coupled with soft ionization techniques such as MALDI and ESI, NMR, and Raman spectroscopy are key techniques employed for lipidomic characterization in cancer diagnostics. Breast cancer is the most commonly diagnosed cancer, and progress in characterizing the lipidome using MS has been reported earlier. Min et al., using nanoflow LC–ESI–MS-MS, analyzed four different categories of PLs (PS, PI, PG, and PA) from urine samples from breast cancer patients and healthy controls [132]. The total amounts of many PLs increased in cancer patients when compared to healthy controls, with a significant increase in PS species (18:1/18:1 and 18:2/18:0), while PI 18:0/20:4 was significantly reduced in breast cancer samples. Importantly, the PS levels were restored in the patients after surgery. The authors implied that the lipid composition found in the urine samples of breast cancer patients could serve as early diagnosis avenues.
Similarly, in order to identify lipid targets that play a role in breast cancer invasion, Wang et al. used MALDI-mass spectrometry imaging, compared lipidomic profiles of two poorly and two highly invasive breast cancer cells lines, and found upregulation of 31 lipids including PG and PA, and downregulation of 8 lipids, including SM [133]. The authors made a point that deciphering the correlation between cancer proliferation and lipid metabolism may assist in the exploration of diagnostic markers and therapeutic targets in breast cancer. In an earlier MALDI-MSI study of human breast cancer cells by Kawashima et al., accumulation of phosphatidylinositols (PIs) was observed in breast cancer tissues, and therefore PI was considered as a biomarker candidate [134]. They identified two different populations of cancer cells that predominantly expressed either PI18:0/18:1 or PI18:0/20:3, with the latter population believed to be associated with the invasiveness. Another MALDI-imaging mass spectroscopic study reveals that the PC (32:1) content of recurring TNBC is higher than the non-recurring TNBC tumors [135]; thus, it appears to be a predictive feature of recurrence and help in the improvement of the prognosis of TNBC patients. More recently, lipid profiling by MALDI-TOF MS by Silva et al. (2020) verified that the Lyso-PC/PC ratio, calculated from the MALDI-mass spectra, can be used as a marker for breast cancer progression [136]. The PC content is most likely affected by the changes in the phospholipase A2 activity, and its activity increases with the carcinoma stage. Contrary to earlier reports, Silva et al. confirmed that highly phosphorylated PIs are difficult to detect by MALDI MS, and even though PI levels have been established as reliable biomarkers in other studies, they could not be detected under given experimental settings for breast cancer cells.
Recently, Eiriksson et al. identified key differences between the lipidome of heterogeneous breast cancer subtypes with an aim to develop specific targeted therapeutic solutions [137]. Using LC–MS, an upregulation of triacylglycerols (TG) ≥ C48 with multiple unsaturation in fatty acyl chains and reduction in PE (C34 to C38) levels in cell lines exhibiting estrogen and progesterone receptor-positive tumor subtypes was observed. For the HER2-overexpressing tumor subtype, an elevated expression of TG (≤ C46), PC, and PE-containing short-chained (≤ C16) saturated or monounsaturated fatty acids were observed. Enhanced levels of PC ≥ C40 were also found in the cell lines of the TNBC subtype.
In another study, a comprehensive and comparative metabolic and lipidomic profiling of a human epithelial breast cell line (MCF10A), a slightly metastatic (MCF7), and a highly metastatic (MDA-MB-231) breast cancer cell line was done using gas chromatography-mass spectrometry (GC–MS) and direct infusion mass spectrometry (DI-MS) [138]. Enhanced levels of most phospholipids were observed in both metastatic groups, compared to normal cells. Coherent with earlier observations, the levels of phosphatidylserine (PS) 18:0/20:4, phosphatidylinositol (PI) 18:0/20:4, and phosphatidylcholine (PC) 18:0/20:4 were markedly higher, while those of phosphatidylethanolamine (PE) 18:1/18:1 and PI 18:0/18:1 were lower in MDA-MB-231 cells than in MCF7 cells.
A comprehensive lipidomic analysis through high-resolution LC–MS, specifically targeting the TNBC by Purwaha et al., were able to measure 684 named lipids across 15 lipid classes in 70 TNBC tumors [139]. Survival analysis across different races revealed that an enhanced level of sphingomyelins is closely associated with disease-free survival in TNBC patients. Undeniably, sphingomyelin levels serve as a potential prognostic marker and the enzymes implicated in sphingolipid metabolism as candidate therapeutic targets. An overview of varied changes in the global lipid classes in TNBC is represented in Fig. 2B for clarity.
Mechanical Signatures of Lipid Membranes in Invasive Breast Cancer: Future Biomarkers
Cell membranes are involved in the metabolic events of the cancer cells, as well as in metastasis. In this regard, cell stiffness is a crucial mechanical property closely related to cell motility, and this compliance is considered a biomarker of cancer metastasis. Biophysical measurements, such as micropipette aspiration [140] and predominantly Atomic Force Microscopy [141] for the determination of cell stiffness, yield a quantitative parameter designated as Young's modulus. In recent years, it has been consistently shown that cancerous cells are softer than normal cells [142]. However, whether metastatic cells are stiffer or softer than non-invasive cancer cells lacks consensus due to conflicting evidence showing that cells with higher motility are sometimes stiffer than their non-invasive counterparts [143].
The mechanical properties of cancer cells dynamically change during the metastatic process to successfully survive the harsh and variable environment of blood vessels, lymphatic vessels, and extracellular matrix [144]. Significantly, the whole metastatic cascade triggered by the accumulation of genetic modification, angiogenesis, and activation of complex signaling pathways leads to the modification of physicochemical properties of the cell; specifically, reduced intercellular adhesion and a morphological transition from cuboidal epithelial to mesenchymal. It is perceived that the physical transition to an invasive malignancy induces actin cortex rearrangement, which tends to make cancer cells softer and allows provisions for cell deformation for motility. Cell deformability is largely determined by myosin-driven cortical tension and actin fiber architecture at the cell cortex. Myosin II, an abundant force-producing protein, is critical for the mechanics of cell migration, while myosin IIB seems to have a preferential role in the mechanics of lamellar protrusion [145]. Using fluorescently labeled peroxisomes as microrheological probes, Smelser et al. (2015) compared the interior mechanical properties of normal breast cells with a triple-negative (metastatic) breast cell line, MDA-MB-231 [146]. They estimated the mechanical properties of cell cytoplasm from the motions of their peroxisomes while reducing the contribution of active cytoskeletal motions to peroxisome motion by means of myosin inhibiting and ATP depleting drugs. It was observed that the peroxisomes exhibited normal diffusion despite treatment with drugs and their mean squared displacements were significantly higher in metastatic cells than in normal cells, suggesting that these cells have differences in cytoplasmic mechanical properties. The measurement of the mean square displacement led to the inference that the MDA-MB-231 cells were significantly softer than normal cells.
AC Martin et al. used peak-force modulation atomic force microscopy for high-resolution topography and stiffness imaging of actin filaments of three epithelial breast cancer cells lines at the variable state of malignancy: healthy MCF10A, non-invasive MCF7, and invasive MDA-MB-231 living cells in order to derive a correlation between cytoskeleton organization and stiffness [147]. Actin filaments provide the scaffold for a cell body representing a significant part of the cytoskeletal structure and features such as stress fiber generation, cross-linking of fibers, and interaction with other proteins like myosin II that affect the mechanical properties of cells [140] (Fig. 2C). In healthy cells, local stiffness is maximum where filamentous actin is organized as well-aligned stress fibers (actomyosin fibers) resulting in apparent Young's modulus values up to one order of magnitude larger than those in regions where these structures are not observed. However, these organized actin fibers were barely observed in tumorigenic cells. The authors further investigated the cytoskeleton conformation in the above three cell lines by immunofluorescence confocal microscopy. The combination of both techniques determined that actin stress fiber is present at apical regions of healthy cells, while in tumorigenic cells, they appear only at basal regions where they cannot contribute to stiffness as probed by atomic force microscopy. These results substantiate that actin stress fibers provide a dominant contribution to stiffness in healthy cells, while the elasticity of tumorigenic cells appears to be not determined by these structures.
Apart from stiffness imaging, AFM single-cell force spectroscopy is a versatile method that allows the measurement of other nanomechanical properties of cells, such as forces of adhesion and tether extrusion, along with an assessment of elasticity. Tethers are nanotubes that get extended when the cell membrane is pulled away from the underlying cytoskeleton and can be immensely useful in differentiating any perturbation in membrane tension or membrane-cytoskeleton interactions arising as a result of changes in metabolic impairment, such as in cancer. Smolyakov et al. used AFM force measurements for assessment of invasive potential by comparing elasticity, adhesion, and tether extrusion of four breast cancer cell lines (SKBR3, MCF7, T47D, MDA-MB231) [148]. The authors observed that more aggressive is the invasive potential of cells, softer are the cells (lower young’s modulus) and exhibit higher adhesion forces. This is understandable because soft cells are more prone to deformation and present a higher surface area for adhesion, Fig. 2C. Tether analysis exhibited that more invasive cells tend to form more tethers, with a smaller tether rupture force per cell.
Yubero et al. recently published a study comparing the deformability of healthy (MCF10A), metastatic (MDA-MB-231), and non-invasive (MCF7) breast cancer cells [143]. They employed computational methods to obtain power-law parameters, namely, the elasticity modulus and fluidity from traditional AFM force vs. indentation curves and delineated the contributions of myosin II, actin cortex network, and energy metabolism to the elasticity of these cells. Strikingly, the value of Youngs’ Modulus revealed that invasive cancer cells are stiffer than non-invasive cancer cells, which imposes a challenge for whether the elasticity of cells can be used as a biomarker of metastatic potential. The plausible explanation for the higher stiffness of MDA-MB-231 cells as compared to MCF7 cells is that for migration and invasion into distant tissues, there is a need for a robust actomyosin network. For validation, the authors analyzed the effect of cytoskeletal drugs and energy starvation on the cell mechanical response. The authors reasoned that whereas the stiffness of healthy cells is due to ATP-driven actin polymerization, the metastatic cells use myosin II activity. The metabolic processes activate the actin cortex, and it is the attributes of these active processes that contribute toward cell stiffness. Thus, remodeling of the cytoskeleton and reprogramming of energy metabolism is a relevant cancer hallmark.
Additionally, a combination of shear assay and digital imaging correlation (DIC) techniques have also been used to measure the local viscoelastic properties of non-tumorigenic breast cells and TNBC cells at different stages of tumor progression subjected to constant shear stress [132, 133]. In recent work [149], the same group characterized statistical variations in the structure and the viscoelastic properties of the non-tumorigenic cell (MCF10A) and tumorigenic TNBC cells (MDA-MB-468 and MDA-MB-231). The results are consistent with the earlier findings where normal MCF10A breast cells exhibited a compact bundle of the actin cytoskeletal network, and the highly metastatic MDA-MB-231 TNBC cells had a more dispersed filament bundle of the actin cytoskeletal structure. Furthermore, the more metastatic MDA-MD-468 cells had a less dense actin filament bundle (Fig. 2C). The authors explained the variations in the viscoelastic properties of the nuclei and cytoplasm of non-tumorigenic and TNBC cells in terms of the statistical variations in pixel fluorescence intensity due to the local variations in the density of the actin filament bundles.
Hui et al. correlated malignancy and drug resistance, two significant characteristics of cancer cells originating from different cancer lines, with membrane resealing responses [150]. This work demonstrated the potential of using this quantity as a novel marker for future cancer diagnosis and drug resistance detection. In this regard, Bouvet et al. (2020) have shown that the disruption of membrane repair machinery involving Annexins (AnxA5 and AnxA6) in invasive MDA-MB-231 cells causes inhibition of metastasis and ultimately cell death [151]. Annexins are Ca2+ triggered phospho-lipid-binding proteins and emerge as important components of the plasma membrane repair system [152]. Due to upregulated annexin levels, cancer cells have an immense capacity for membrane repair, thus enhancing their survival against the disruptions of PM and pull-out of membrane fragments, processes that usually accompany migration of MDA-MB-231 cells on collagen I fibrils. In an earlier study, with metastatic MCF7 cells, it was observed that Ca2+ dependent annexin A2 and S100A11 proteins largely control the plasma membrane repair mechanism [153]. Annexins bind specific cytosolic S100 proteins through their NH2 terminal region along with Ca2+ to form a ternary complex that facilitates opposing of adjacent phospholipid membranes and accumulation of F-actin at the site of membrane injury. Through in vitro membrane repair assays, the authors have shown that silencing of the membrane repair machinery inhibits the migration of cancer cells and presents an encouraging avenue for inhibiting cancer metastasis.
The above studies highlight that the multifaceted approach toward understanding tumor mechanics is critical to reveal the underlying mechanism regulating the development and metastasis of tumors and is a promising methodology for diagnosis and targeted therapeutic interventions [154].
Oncological Scope of Membrane Targeted Therapies
Membrane-lipid therapy (MLT) is a relatively new strategy studied as a method of choice for treating not only cancer but also other anomalies like neurodegenerative diseases, diabetes, and stroke [155]. Conventionally, most cancer therapeutics are targeted against abnormally upregulated proteins and DNA; however, lipid therapy, which has its foundation on the characteristic differences in lipid composition among the healthy and cancer cell membranes, is an interesting and more sensitive alternative as a solution for multidrug resistance [5, 7]. However, this requires a thorough understanding of how therapeutic agents affect the membrane organization and composition by different mechanisms, which are discussed below.
Targeting PS and PE
Among the common targets of MLT, PS on the outer membrane of tumor cells is proposed to be an effective target for cancer therapy [156, 157]. The loss of PS asymmetry in cancer cells may be related to reduced activity of these ATP-dependent phospholipid translocases or to elevated activity of phospholipid scramblase due to high levels of intracellular calcium. The PS-targeting antibody bavituximab [158, 159] and the PS-binding peptide/peptoid hybrid PPS1D1 [160, 161] have shown significant cytotoxic effects in cancer cells (Fig. 3). Another approach involves the use of anticancer drugs encapsulated in specific lipid carriers. Anticancer drugs (e.g., camptothecin and doxorubicin) are entrapped in cationic phosphatidylcholine-stearylamine (PC-SA) liposomes which bind strongly to the membrane by direct interaction with the surface exposed PS and have been a successful anticancer strategy both in vitro and in mice models [162, 163] (Fig. 3). Similarly, the antibiotic cinnamycin, produced by Streptomyces sp., and the structurally related tetracyclic peptide duramycin, produced by some Gram-positive bacterium, including Streptoverticillium cinnamoneus, are among the PE-targeting drugs in cancer cells. Cinnamycin and duramycin belong to the family of lantibiotics. These toxins specifically bind to PE-containing membranes by changing membrane curvature and by inducing trans bilayer lipid movement, which consequently kills the cells [164]. A model-membrane-based study revealed that the binding of duramycin and cinnamycin to PE-containing liposomes is dependent on the membrane curvature, and the lantibiotics bind small vesicles more efficiently as compared to large liposomes. Electron microscopy and small-angle X-ray scattering studies showed that it involves tubulation of membranes in the course of binding the lantibiotics to multilamellar liposomes, suggesting that membrane deformation occurs upon binding [165]. Other PE-targeting molecules are cyclotides that disrupt PE-containing membranes by means of binding, followed by insertion and formation of a pore, eventually leading to cell death [166] (Fig. 3).
Summary of the drugs targeting lipids and membranes in breast cancer. Cinnamycin, duramycin, and cyclotides target exposed PE in the plasma membrane. Bavituximab and PPS1D1 target exposed PS. Statins, MTDBs, EPA, DHA, and Edelfosine modulate lipid raft structure and membrane cholesterol to downregulate raft-dependent processes. PC-SA-based liposomal drug delivery leads to efficient intracellular drug release by specifically binding to PS domains in the plasma membrane. Similarly, immunoliposomes specific to TNBC can deliver ncRNA and simvastatin to induce cell death. Resveratrol and azurin modify membrane properties that enable their use in combination therapy to facilitate uptake of anticancer agents such as doxorubicin and paclitaxel 1S, 3R-RSL3, and SAS inhibited GPX4 that activates ferroptosis cell death via membrane damage. HTO alters the composition and organization of the membrane by altering the abundance of SFA, PUFA, and Cer levels impacting membrane-associated signaling. PDB codes of depicted transmembrane proteins are 2MFR (insulin receptor, red) and 4PYP (GLUT-1, blue)
Targeting Lipid Domains
Another MLT target comprises lipid domains, including lipid rafts which disrupt either by altering Chol concentration (short-chain ceramides, statins, etc.) or by increasing the permeability of the membrane for other drugs–by means of changing membrane properties. The latter is achieved by drugs like resveratrol, which is commonly found in red wine [167], and azurin, a membrane-associated protein from Pseudomonas aeruginosa (Fig. 3). Azurin and its derived peptide p28 have been comprehensively studied as anticancerous and are known for down-regulating common signaling pathways controlled by membrane receptors, inducing alteration of lipid domains, decreases the stiffness of PM and ordered domains, and affects adhesion and invasiveness in vivo [68, 168, 169]. Bernades et al. observed that azurin uptake in cancer cells is mediated by lipid raft markers like caveolin one and ganglioside one and that its simultaneous administration along with anticancer therapeutic drugs like paclitaxel and doxorubicin enhance their anticancer potential [170]. Targeting the membrane cholesterol is another strategy proposed by Zhang et al. against the ErbB2-positive breast cancer subtype [171] in vivo (Fig. 3). Membrane cholesterol contents maintain cell surface distribution of ErbB2, a membrane-embedded receptor tyrosine kinase, by increasing the rigidity and decreasing the fluidity of cell membranes. Authors reported that lowering cholesterol levels by the use of drugs such as lovastatin assisted the internalization and degradation of ErbB2 along with the observation of synergistic effects of lovastatin with the ErbB2 inhibitor lapatinib, which were evaluated using an ErbB2-positive breast cancer xenograft mouse model. Lipophilic statins are also perceived as one of the promising adjuvants in cancer chemotherapy. Apart from their known function of lowering cholesterol, several membrane-based in vitro studies indicate that statins independently interact and induce modification of membrane dynamics in a depth-dependent manner [172, 173] along with alterations within membrane architecture by multi-domain formation [174] (Fig. 3). Translation of such an effect in cellular membranes in vivo remains to be fully investigated. Lipophilic statins, as compared to hydrophilic ones, are known to be associated with fewer side effects and lower cancer mortality [175,176,177]. Apart from therapeutics, the role of select membrane-targeted dietary bioactive (MTDBs) such as polyunsaturated fatty acids (PUFAs like eicosapentaenoic acid (EPA) and docosahexaenoic acid (DHA)) and linoleic acid has been implicated in reorganizing PM hierarchal domains and subsequently reduce cancer risk, especially in colon cancer [178] (Fig. 3).
State of the Art of Lipid-based Therapy in Breast Cancer: The Road Ahead
Breast cancer is the biggest cause of cancer-related morbidity among women across the globe. Unfortunately, complicated molecular mechanisms underlying breast cancer invasion and metastasis are not yet sufficiently understood. In order to improve the efficiency of the current antitumor therapies and overcome issues of drug resistance and cytotoxicity to normal cells, novel treatment approaches based on liposomal carrier agents are being considered as a promising solution. Statins, as mentioned above, are recognized as potential anticancer therapeutic agents. Alternatively, using statins formulated in different drug delivery systems has helped in improving their efficiency as antitumor agents. Augmented simvastatin cytotoxicity using optimized solutol-based lipid nanocapsules has been a potential breast cancer treatment [179]. Long circulating EGFR (epidermal growth factor receptor)-targeted immunoliposomes as a selective delivery system of simvastatin, a lipophilic statin, has been tested for its potential use in the treatment of TNBCs [180]. Similarly, treatment methods using lipid‑based nanoparticles (LBNPs) have markedly improved the delivery efficiency of therapeutic noncoding RNAs (ncRNAs), more specifically microRNAs (miRNAs) and small interfering RNAs (siRNAs) into tumor cells and tissues [181] (Fig. 3). ncRNAs are involved in several biological processes, including cell growth and proliferation, apoptosis, invasion, and metastasis. Recently, fascinating evidence has shown the impending contribution of ncRNAs toward breast carcinogenesis [182]. ncRNAs regulate the expression levels of oncogenes and tumor suppressor genes and hence, have emerged as attractive pharmacological targets for treating variegated malignancies like TNBC. A very recent example is the formulation of therapeutic RGD-PEG-ECO/siDANCR nanoparticles [183], which are targeted against oncogenic long noncoding RNA (lncRNA) DANCR one of the most overexpressed genes in TNBC and an emerging therapeutic target in human cancer [184]. The nanoparticle complexes were made via the self-assembly of multifunctional amino lipid ECO, cyclic RGD peptide-PEG, and siDANCR for systemic delivery. MDA-MB-231 and BT-549 cells treated with the therapeutic RGD-PEG-ECO/siDANCR nanoparticles, where they exhibited 80–90% reduction in the expression of DANCR for up to 7 days, indicated efficient intracellular siRNA delivery and sustained target silencing.
Very recently, Saraiva et al. demonstrated that edelfosine-based nanoemulsions (an anticancer drug and synthetic lipid member of the alkyl-lysophospholipid family) are effective against highly aggressive and invasive TNBC (MDA-MB-231) xenografts in zebrafish embryo animal models [185]. The nanoemulsions were composed of lipophilic edelfosine–an oil (Miglyol), and PC–a stable lipid with a neutral zeta potential and an average size of approximately 120 nm. Edelfosine, being an alkyl-lysophospholipid, has a long carbon chain, which gets incorporated into the oily core of the structure, while the phosphate and quaternary amine groups remain exposed on the surface, similar to PC (Fig. 3). The authors found that the nanoformulation was effectively internalized by cancer cells with a subsequent improved therapeutic effect (in vitro and in vivo) as compared to edelfosine alone. This is also the first time that the efficacy of edelfosine has been tested against TNBC, and the nano emulsion-based drug delivery has been claimed by authors to be a promising treatment against TNBC.
Another multifaceted approach in targeting TNBC has been demonstrated by Vita et al. [186]. To give context, the extracellular matrix (ECM) is an important component of the tumors microenvironment and regulates several tumor processes by means of certain enzymes such as lysyl oxidase 1 (LOX). LOX activity is responsible for the cross-linking of elastin and collagen fibers, resulting in the inflection of the architecture and stiffness of tumor ECM; this also coordinates cell migration and adhesion. Inhibition of LOX activity has been previously targeted as a promising approach against breast epithelial carcinogenesis, lung tumors, and prostate cancer. Vita et al. designed a poly-ethylene–glycol-based liposomal system functionalized with a LOX-antibody and encapsulating a breast anticancer drug, epirubicin (Lipo-LOX-EPI). Epirubicin is an anthracycline-based chemotherapeutic drug that suffers from drawbacks like limited tumor targeting and systemic toxicity. This work represents remarkable maneuvering of the inherent therapeutic activity of targeting LOX in the tumor ECM and discerningly enhancing the epirubicin concentration at the tumor site, with minimal systemic toxicity. The authors tested and compared Lipo-LOX-EPI, Lipo-EPI, and Lipo-LOX for the cell viability of MDA-MB-231 3D culture and found more than 80% cell death with Lipo-LOX-EPI. Concomitantly, a significant reduction of stiffness of the 3D-collagen-based scaffold, which mimics the tumor environment with features of ECM, was observed. Indeed, the development of such drug delivery vehicles opens up new avenues for the treatment of TNBC.
Doll et al. revealed that TBNC is more sensitive to ferroptosis than ER-positive breast cancer and hence, triggering ferroptosis cell death seems to be an effective treatment strategy for TNBC [187]. Ferroptosis is a recently discovered distinct type of regulated cell death caused by the accumulation of lipid-based reactive oxygen species (ROS), and targeting ferroptosis is proposed to be a promising anticancer strategy against breast cancer [188], Fig. 3. Ferroptosis is characterized by oxidation of polyunsaturated fatty acid-containing phospholipids, the presence of redox-active iron, and loss of lipid peroxide repairing ability by phospholipid hydroperoxidase glutathione peroxidase-4 (GPX4). The ferroptotic agent (1S, 3R)-RSL3 [189] and sulfasalazine (SAS) [190] inhibit the peroxidase activity of GPX4 in breast cancer cells. It has also been realized that drug-resistant breast cancer cells exhibit an augmented dependency on GPX4, which in turn relies on the presence of glutathione (GSH). This suggests that the inhibition of GPX4 and GSH is a potential measure to overcome drug resistance in breast cancer [191]. It has been shown that starvation of cystine, a substrate used to synthesize GSH to prevent ferroptosis, was able to induce cell necroptosis and ferroptosis in TNBC via the GCN2-eIF2α-ATF4-CHAC1 pathway [192].
Plant-derived pentacyclic triterpenic acids (TAs), namely betulinic acid (BA) and ursolic acid (UA), have been studied (1HNMR-based quantification and identification of cell-metabolome) and were found to modulate diverse pathways in carcinogenesis, pertaining to glucose and lipid metabolism in highly metastatic MDA-MB-231 cells [193]. TNBCs are marked by high rates of FA oxidation of DAG, triacylglycerides, and cholesteryl esters stored in the lipid droplets (LD), which are utilized for sustaining oxidative phosphorylation in the mitochondria through downregulation of acetyl-CoA and seem to play an extensive role in the cancer invasiveness [194]. NMR analysis of organic cell extracts displayed decreased levels of triglycerides and cholesteryl esters upon BA/UA treatment. Furthermore, MDA-MB-231 cells treated with TAs displayed significantly increased levels of free cholesterol and glycerophospholipids (GPL) (especially in the case of BA), which are known to be major constituents of the membrane. This induced reprogramming of lipid metabolism demonstrates a plausible therapeutic alternative in TNBC.
A novel anticancer drug, hydroxytriolein (HTO), has been found to regulate the composition and spatiotemporal organization of the membrane by direct interaction. This synergy in turn alters the downstream signaling events through transmembrane proteins embedded within the lipid bilayer that impairs the growth of TNBC cells, MDA-MB-231 and BT-549 [195] (Fig. 3). The authors suggested two possible mechanisms for the mechanism of action of HTO. First, HTO could possibly trigger the ERK (extracellular signal-regulated kinase) signaling pathway, and thus channelizing TNBC cell autophagy. Alternatively, or in addition, it could accelerate generation of dihydroceramide and ceramide that eventually inhibit Akt (protein kinase B) without EGFR activation. It was observed that the unsaturated/saturated fatty acid ratio increased in MDA-MB-231 cells following exposure to HTO, which inadvertently affects the biophysical properties of cell PM, such as its non-lamellar propensity, membrane surface packing density, or membrane domain organization and fluidity. Alterations of the structural properties of PM bring about significant changes in the membrane-binding interactions and activity of important membrane proteins that are critical for cell survival. Since HTO treatment does not exhibit any undesired side effects, this study puts forth a promising anticancer molecule that targets the lipid bilayer to tackle TNBC.
Among the approved liposomal formulations of doxorubicin are Lipo-Dox (DSPC, cholesterol, PEG 2000-DSPE (56:39:5), Doxil (HSPC:Cholesterol:PEG 2000-DSPE (56:39:5 molar ratio) and Myocet (EPC and cholesterol) designed to be more tolerable and more effective than free doxorubicin, targeted against breast cancer [196,197,198]. Myocet is a nonpegylated liposomal Dox that has been approved in combination with cyclophosphamide (CPM) for first-line treatment of patients with metastatic breast cancer [196]. MM-302 is another formulation of doxorubicin, a HER2-targeted antibody–liposomal doxorubicin conjugate that specifically targets HER2-overexpressing cells, increasing the delivery of doxorubicin to tumor cells and limiting exposure to healthy cells. MM-302 is yet awaiting approval. In 2016, Miller et al. (2016) used the MM-302 formulation plus trastuzumab in a phase II trial in patients with HER2-positive locally advanced/metastatic breast cancer [199]. Other Pegylated liposomal doxorubicin (PLD)-based formulations, either alone or in combination with other drugs targeted against MBC include Caelix, under phase II trials [200, 201] and PLD + CPM, under phase III clinical trials [202]. Specifically targeting against TNBC, the combination of PLD + bevacizumab + temsirolimus and PLD + bevacizumab + everolimus are under phase I clinical trials [203]. Endo-TAG (DOTAP, DOPC, PTX (50:47:3)) is a paclitaxel-based liposomal formulation against TNBC which is currently under clinical trials [204]. Paclitaxel-based approved formulation is that of Genexol-PM which is a PEGylated polymeric micelle formulation [205].
Conclusions
Our understanding of lipid reprogramming in cancer has steadily progressed over the years due to advances in lipidomics, analytical methods to sensitively detect and quantify cellular lipid classes, and a surge in cellular biophysical techniques for studying lipid membranes. The highly selective alteration in varied lipid classes in cancer (in general) and TNBC (in specific) culminate in the modification of cellular membrane biophysical properties predisposing changes in lipid membrane attributes as attractive diagnostic tools. These also serve as a validated platform to inspire the development of lipid-targeted therapy against TNBC and gauge the correlation of drug resistance with altered membrane structure, all eventually contributing to the common goal of exploring the potential of lipid-based theranostics in TNBC. Although promising, this field requires further investigations to validate the therapeutic and diagnostic potential of lipids and membranes in TNBC. This can be achieved by acutely and directly exploring the lipidome changes in membranes of TNBC patients using label-free and universal methods and scoring the same for disease progression and chemoresistance. Further, for lipidomics to be widely integrated into routine cancer diagnoses, an issue that needs urgent attention is the technicality of doing these investigations. These include sample collection, storage, access to mass spectroscopic instruments, and bio-informatic expertise to interpret changes in the patient lipidome with disease progression and in response to therapy with high confidence. A few ways to partially solve this problem are (a) the use of automatic high through-put mass spectrometers to save costs related to running experiments, transportation, and performing analysis within the limited sample shelf-life; (b) the development of highly sensitive bench-top mass spectrometer models which could be customized for specific lipid class detection for routine use in diagnoses; and (c) higher collaborations between clinicians and academic researchers to enable more exhaustive lipidome analysis in real-time. It can be foreseen that some of these goals will be achievable in the near future.
Availability of data and material
Not applicable.
Code availability
Not applicable.
References
Yang NJ, Hinner MJ (2015) Getting across the cell membrane: an overview for small molecules, peptides, and proteins. Methods Mol Biol 1266:29–53
Escribá PV, González-Ros JM, Goñi FM et al (2008) Membranes: a meeting point for lipids, proteins and therapies. J Cell Mol Med. https://doi.org/10.1111/j.1582-4934.2008.00281.x
Ibarguren M, López DJ, Encinar JA et al (2013) Partitioning of liquid-ordered/liquid-disordered membrane microdomains induced by the fluidifying effect of 2-hydroxylated fatty acid derivatives. Biochim Biophys Acta-Biomembr 1828:2553–2563. https://doi.org/10.1016/j.bbamem.2013.06.014
Casares D, Escribá PV, Rosselló CA (2019) Membrane lipid composition: effect on membrane and organelle structure, function and compartmentalization and therapeutic avenues. Int J Mol Sci. https://doi.org/10.3390/ijms20092167
Zalba S, ten Hagen TLM (2017) Cell membrane modulation as adjuvant in cancer therapy. Cancer Treat Rev 52:48–57. https://doi.org/10.1016/j.ctrv.2016.10.008
Szlasa W, Zendran I, Zalesińska A, et al (2020) Lipid composition of the cancer cell membrane To cite this version: HAL Id: hal-03089978 Lipid composition of the cancer cell membrane
Preta G (2020) New insights into targeting membrane lipids for cancer therapy. Front Cell Dev Biol 8:1–10. https://doi.org/10.3389/fcell.2020.571237
Kubicek-Sutherland J, Vu D, Mendez H et al (2017) Detection of lipid and amphiphilic biomarkers for disease diagnostics. Biosensors. https://doi.org/10.3390/bios7030025
Escribá PV, Busquets X, Inokuchi JI et al (2015) Membrane lipid therapy: modulation of the cell membrane composition and structure as a molecular base for drug discovery and new disease treatment. Prog Lipid Res 59:38–53. https://doi.org/10.1016/j.plipres.2015.04.003
Escribá PV (2017) Membrane-lipid therapy: a historical perspective of membrane-targeted therapies—from lipid bilayer structure to the pathophysiological regulation of cells. Biochim Biophys Acta-Biomembr 1859:1493–1506. https://doi.org/10.1016/j.bbamem.2017.05.017
Rodrigues C, Milkovic L, Bujak IT et al (2019) Lipid profile and aquaporin expression under oxidative stress in breast cancer cells of different malignancies. Oxid Med Cell Longev. https://doi.org/10.1155/2019/2061830
Bhattarai S, Saini G, Gogineni K, Aneja R (2020) Quadruple-negative breast cancer: novel implications for a new disease. Breast Cancer Res. https://doi.org/10.1186/s13058-020-01369-5
Yin L, Duan J-J, Bian X-W, Yu S (2020) Triple-negative breast cancer molecular subtyping and treatment progress. Breast Cancer Res. https://doi.org/10.1186/s13058-020-01296-5
Gracià RS, Bezlyepkina N, Knorr RL et al (2010) Effect of cholesterol on the rigidity of saturated and unsaturated membranes: fluctuation and electrodeformation analysis of giant vesicles. Soft Matter 6:1472–1482. https://doi.org/10.1039/b920629a
Codini M, Garcia-Gil M, Albi E (2021) Cholesterol and sphingolipid enriched lipid rafts as therapeutic targets in cancer. Int J Mol Sci. https://doi.org/10.3390/ijms22020726
Vona R, Iessi E, Matarrese P (2021) Role of cholesterol and lipid rafts in cancer signaling: a promising therapeutic opportunity? Front Cell Dev Biol. https://doi.org/10.3389/fcell.2021.622908
Sezgin E, Levental I, Mayor S, Eggeling C (2017) The mystery of membrane organization: composition, regulation and roles of lipid rafts. Nat Rev Mol Cell Biol. https://doi.org/10.1038/nrm.2017.16
Pike LJ (2003) Lipid rafts: bringing order to chaos. J Lipid Res. https://doi.org/10.1194/jlr.R200021-JLR200
Grecco HE, Schmick M, Bastiaens PIH (2011) Signaling from the living plasma membrane. Cell. https://doi.org/10.1016/j.cell.2011.01.029
Landreh M, Robinson CV (2015) A new window into the molecular physiology of membrane proteins. J Physiol. https://doi.org/10.1113/jphysiol.2014.283150
Mollinedo F, Gajate C (2015) Lipid rafts as major platforms for signaling regulation in cancer. Adv Biol Regul. https://doi.org/10.1016/j.jbior.2014.10.003
Labilloy A, Youker RT, Bruns JR et al (2014) Altered dynamics of a lipid raft associated protein in a kidney model of Fabry disease. Mol Genet Metab. https://doi.org/10.1016/j.ymgme.2013.10.010
Sonnino S, Aureli M, Grassi S et al (2014) Lipid rafts in neurodegeneration and neuroprotection. Mol Neurobiol. https://doi.org/10.1007/s12035-013-8614-4
Badana AK, Chintala M, Gavara MM et al (2018) Lipid rafts disruption induces apoptosis by attenuating expression of LRP6 and survivin in triple negative breast cancer. Biomed Pharmacother 97:359–368. https://doi.org/10.1016/j.biopha.2017.10.045
Quinn PJ (2012) Lipid–lipid interactions in bilayer membranes: Married couples and casual liaisons. Prog Lipid Res. https://doi.org/10.1016/j.plipres.2012.01.001
Seddon JM, Templer RH (1995) Polymorphism of lipid-water systems. Elsevier, Amsterdam
Cullis PR, De Kruijff B (1979) Lipid polymorphism and the functional roles of lipids in biological membranes. Biochim Biophys Acta-Rev Biomembr. https://doi.org/10.1016/0304-4157(79)90012-1
Lladó V, López DJ, Ibarguren M et al (2014) Regulation of the cancer cell membrane lipid composition by NaCHOleate: effects on cell signaling and therapeutical relevance in glioma. Biochim Biophys Acta-Biomembr 1838:1619–1627. https://doi.org/10.1016/j.bbamem.2014.01.027
Yang L, Ding L, Huang HW (2003) New phases of phospholipids and implications to the membrane fusion problem †. Biochemistry. https://doi.org/10.1021/bi0344836
Goñi FM (2014) The basic structure and dynamics of cell membranes: an update of the Singer-Nicolson model. Biochim Biophys Acta-Biomembr. https://doi.org/10.1016/j.bbamem.2014.01.006
Escribá PV (2006) Membrane-lipid therapy: a new approach in molecular medicine. Trends Mol Med. https://doi.org/10.1016/j.molmed.2005.11.004
Fuller N, Rand RP (2001) The influence of lysolipids on the spontaneous curvature and bending elasticity of phospholipid membranes. Biophys J. https://doi.org/10.1016/S0006-3495(01)75695-0
Chernomordik LV, Kozlov MM (2008) Mechanics of membrane fusion. Nat Struct Mol Biol. https://doi.org/10.1038/nsmb.1455
Piomelli D, Astarita G, Rapaka R (2007) A neuroscientist’s guide to lipidomics. Nat Rev Neurosci. https://doi.org/10.1038/nrn2233
Holthuis JCM, Menon AK (2014) Lipid landscapes and pipelines in membrane homeostasis. Nature. https://doi.org/10.1038/nature13474
Bell RM, Ballas LM, Coleman RA (1981) Lipid topogenesis. J Lipid Res. https://doi.org/10.1016/S0022-2275(20)34952-X
van Meer G, Voelker DR, Feigenson GW (2008) Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol. https://doi.org/10.1038/nrm2330
Daum G (1985) Lipids of mitochondria. Biochim Biophys Acta-Rev Biomembr. https://doi.org/10.1016/0304-4157(85)90002-4
Koivuniemi A (2017) The biophysical properties of plasmalogens originating from their unique molecular architecture. FEBS Lett. https://doi.org/10.1002/1873-3468.12754
Messias MCF, Mecatti GC, Priolli DG, De Oliveira CP (2018) Plasmalogen lipids: functional mechanism and their involvement in gastrointestinal cancer. Lipids Health Dis 17:1–12. https://doi.org/10.1186/s12944-018-0685-9
Han X, Gross RW (1990) Plasmenylcholine and phosphatidylcholine membrane bilayers possess distinct conformational motifs. Biochemistry. https://doi.org/10.1021/bi00472a032
Lorent JH, Levental KR, Ganesan L et al (2020) Plasma membranes are asymmetric in lipid unsaturation, packing and protein shape. Nat Chem Biol. https://doi.org/10.1038/s41589-020-0529-6
Verkleij A, Zwaal RF, Roelofsen B et al (1973) The asymmetric distribution of phospholipids in the human red cell membrane. A combined study using phospholipases and freeze-etch electron microscopy. Biochim Biophys Acta-Biomembr. https://doi.org/10.1016/0005-2736(73)90143-0
Morrot G, Cribier S, Devaux PF et al (1986) Asymmetric lateral mobility of phospholipids in the human erythrocyte membrane. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.83.18.6863
Gupta A, Korte T, Herrmann A, Wohland T (2020) Plasma membrane asymmetry of lipid organization: fluorescence lifetime microscopy and correlation spectroscopy analysis. J Lipid Res. https://doi.org/10.1194/jlr.D119000364
Schachter D, Abbott RE, Cogan U, Flamm M (1983) Lipid fluidity of the individual hemileaflets of human erythrocyte membranes. Ann N Y Acad Sci. https://doi.org/10.1111/j.1749-6632.1983.tb31671.x
Sanchez SA, Tricerri MA, Gratton E (2012) Laurdan generalized polarization fluctuations measures membrane packing micro-heterogeneity in vivo. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.1118288109
Parasassi T, De Stasio G, Ravagnan G et al (1991) Quantitation of lipid phases in phospholipid vesicles by the generalized polarization of Laurdan fluorescence. Biophys J. https://doi.org/10.1016/S0006-3495(91)82041-0
Parasassi T, Ravagnan G, Rusch RM, Gratton E (1993) Modulation and dynamics of phase properties in phospholipid mixtures detected by Laurdan fluorescence. Photochem Photobiol. https://doi.org/10.1111/j.1751-1097.1993.tb02309.x
Parasassi T, Di Stefano M, Loiero M et al (1994) Influence of cholesterol on phospholipid bilayers phase domains as detected by Laurdan fluorescence. Biophys J. https://doi.org/10.1016/S0006-3495(94)80763-5
Parasassi T, Gratton E (1995) Membrane lipid domains and dynamics as detected by Laurdan fluorescence. J Fluoresc. https://doi.org/10.1007/BF00718783
Golfetto O, Hinde E, Gratton E (2015) The Laurdan spectral phasor method to explore membrane micro-heterogeneity and lipid domains in live cells. Methods Mol Biol 1232:273–90
Levi M, Wilson PV, Cooper OJ, Gratton E (1993) Lipid phases in renal brush border membranes revealed by Laurdan fluorescence. Photochem Photobiol. https://doi.org/10.1111/j.1751-1097.1993.tb02312.x
Sezgin E, Gutmann T, Buhl T et al (2015) Adaptive lipid packing and bioactivity in membrane domains. PLoS ONE. https://doi.org/10.1371/journal.pone.0123930
Kreutzberger AJB, Ji M, Aaron J et al (2019) Rhomboid distorts lipids to break the viscosity-imposed speed limit of membrane diffusion. Science (80- ). https://doi.org/10.1126/science.aao0076
Moon S, Yan R, Kenny SJ et al (2017) Spectrally resolved, functional super-resolution microscopy reveals nanoscale compositional heterogeneity in live-cell membranes. J Am Chem Soc. https://doi.org/10.1021/jacs.7b03846
Sameni S, Malacrida L, Tan Z, Digman MA (2018) Alteration in fluidity of cell plasma membrane in huntington disease revealed by spectral phasor analysis. Sci Rep. https://doi.org/10.1038/s41598-018-19160-0
Ammendolia DA, Bement WM, Brumell JH (2021) Plasma membrane integrity: implications for health and disease. BMC Biol. https://doi.org/10.1186/s12915-021-00972-y
Alves AC, Ribeiro D, Nunes C, Reis S (2016) Biophysics in cancer: the relevance of drug-membrane interaction studies. Biochim Biophys Acta-Biomembr 1858:2231–2244. https://doi.org/10.1016/j.bbamem.2016.06.025
Gu R-X, Baoukina S, Tieleman DP (2020) Phase separation in atomistic simulations of model membranes. J Am Chem Soc. https://doi.org/10.1021/jacs.9b11057
Klymchenko AS (2017) Solvatochromic and fluorogenic dyes as environment-sensitive probes: design and biological applications. Acc Chem Res. https://doi.org/10.1021/acs.accounts.6b00517
Steinkühler J, Sezgin E, Urbančič I et al (2019) Mechanical properties of plasma membrane vesicles correlate with lipid order, viscosity and cell density. Commun Biol 2:1–8. https://doi.org/10.1038/s42003-019-0583-3
Kumamoto Y, Harada Y, Takamatsu T, Tanaka H (2018) Label-free molecular imaging and analysis by Raman spectroscopy. ACTA Histochem Cytochem. https://doi.org/10.1267/ahc.18019
Zhanghao K, Liu W, Li M et al (2020) High-dimensional super-resolution imaging reveals heterogeneity and dynamics of subcellular lipid membranes. Nat Commun. https://doi.org/10.1038/s41467-020-19747-0
Heberle FA, Doktorova M, Scott HL et al (2020) Direct label-free imaging of nanodomains in biomimetic and biological membranes by cryogenic electron microscopy. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.2002200117
Yamaji-Hasegawa A, Tsujimoto M (2006) Asymmetric distribution of phospholipids in biomembranes. Biol Pharm Bull. https://doi.org/10.1248/bpb.29.1547
Skotland T, Sandvig K (2019) The role of PS 18:0/18:1 in membrane function. Nat Commun. https://doi.org/10.1038/s41467-019-10711-1
Bernardes N, Fialho A (2018) Perturbing the dynamics and organization of cell membrane components: a new paradigm for cancer-targeted therapies. Int J Mol Sci. https://doi.org/10.3390/ijms19123871
Nicolson GL (2014) The fluid—mosaic model of membrane structure: still relevant to understanding the structure, function and dynamics of biological membranes after more than 40years. Biochim Biophys Acta-Biomembr. https://doi.org/10.1016/j.bbamem.2013.10.019
Nagata S, Suzuki J, Segawa K, Fujii T (2016) Exposure of phosphatidylserine on the cell surface. Cell Death Differ 23:952–961. https://doi.org/10.1038/cdd.2016.7
Vallabhapurapu SD, Blanco VM, Sulaiman MK et al (2015) Variation in human cancer cell external phosphatidylserine is regulated by flippase activity and intracellular calcium. Oncotarget 6:34375–34388. https://doi.org/10.18632/oncotarget.6045
Voll RE, Herrmann M, Roth EA et al (1997) Immunosuppressive effects of apoptotic cells. Nature. https://doi.org/10.1038/37022
Voll RE, Roth EA, Girkontaite I et al (1997) Histone-specific Th0 and Th1 clones derived from systemic lupus erythematosus patients induce double-stranded DNA antibody production. Arthritis Rheum. https://doi.org/10.1002/art.1780401210
Cvetanovic M, Ucker DS (2004) Innate immune discrimination of apoptotic cells: repression of proinflammatory macrophage transcription is coupled directly to specific recognition. J Immunol. https://doi.org/10.4049/jimmunol.172.2.880
Birge RB, Boeltz S, Kumar S et al (2016) Phosphatidylserine is a global immunosuppressive signal in efferocytosis, infectious disease, and cancer. Cell Death Differ 23:962–978. https://doi.org/10.1038/CDD.2016.11
Wanderley JLM, DaMatta RA, Barcinski MA (2020) Apoptotic mimicry as a strategy for the establishment of parasitic infections: parasite- and host-derived phosphatidylserine as key molecule. Cell Commun Signal. https://doi.org/10.1186/s12964-019-0482-8
Bogdanov M, Dowhan W (1998) Phospholipid-assisted protein folding: phosphatidylethanolamine is required at a late step of the conformational maturation of the polytopic membrane protein lactose permease. EMBO J. https://doi.org/10.1093/emboj/17.18.5255
Patel D, Witt SN (2017) Ethanolamine and phosphatidylethanolamine: partners in health and disease. Oxid Med Cell Longev. https://doi.org/10.1155/2017/4829180
Pustylnikov S, Costabile F, Beghi S, Facciabene A (2018) Targeting mitochondria in cancer: current concepts and immunotherapy approaches. Transl Res. https://doi.org/10.1016/j.trsl.2018.07.013
Beloribi-Djefaflia S, Vasseur S, Guillaumond F (2016) Lipid metabolic reprogramming in cancer cells. Oncogenesis. https://doi.org/10.1038/oncsis.2015.49
Krawitz PM, Murakami Y, Rieß A et al (2013) PGAP2 mutations, affecting the GPI-anchor-synthesis pathway, cause hyperphosphatasia with mental retardation syndrome. Am J Hum Genet. https://doi.org/10.1016/j.ajhg.2013.03.011
Henriksen JR, Andresen TL, Feldborg LN et al (2010) Understanding detergent effects on lipid membranes: a model study of lysolipids. Biophys J. https://doi.org/10.1016/j.bpj.2010.01.037
Yeagle PL (2016) Biogenesis of Membrane Lipids. In: The Membranes of Cells. Elsevier
Gibellini F, Smith TK (2010) The Kennedy pathway-De novo synthesis of phosphatidylethanolamine and phosphatidylcholine. IUBMB Life. https://doi.org/10.1002/iub.337
Podo F, Paris L, Cecchetti S et al (2016) Activation of phosphatidylcholine-specific phospholipase C in breast and ovarian cancer: impact on MRS-detected choline metabolic profile and perspectives for targeted therapy. Front Oncol. https://doi.org/10.3389/fonc.2016.00171
Abalsamo L, Spadaro F, Bozzuto G et al (2012) Inhibition of phosphatidylcholine-specific phospholipase C results in loss of mesenchymal traits in metastatic breast cancer cells. Breast Cancer Res. https://doi.org/10.1186/bcr3151
Paris L, Podo F, Spadaro F et al (2017) Phosphatidylcholine-specific phospholipase C inhibition reduces HER2-overexpression, cell proliferation and in vivo tumor growth in a highly tumorigenic ovarian cancer model. Oncotarget. https://doi.org/10.18632/oncotarget.18992
Carrasco S, Mérida I (2007) Diacylglycerol, when simplicity becomes complex. Trends Biochem Sci. https://doi.org/10.1016/j.tibs.2006.11.004
Rauch C, Paine SW, Littlewood P (2013) Can long range mechanical interaction between drugs and membrane proteins define the notion of molecular promiscuity? Application to P-glycoprotein-mediated multidrug resistance (MDR). Biochim Biophys Acta-Gen Subj. https://doi.org/10.1016/j.bbagen.2013.06.038
Rivel T, Ramseyer C, Yesylevskyy S (2019) The asymmetry of plasma membranes and their cholesterol content influence the uptake of cisplatin. Sci Rep 9:1–14. https://doi.org/10.1038/s41598-019-41903-w
Peetla C, Bhave R, Vijayaraghavalu S et al (2010) Drug resistance in breast cancer cells: biophysical characterization of and doxorubicin interactions with membrane lipids. Mol Pharm 7:2334–2348. https://doi.org/10.1021/mp100308n
Munir R, Lisec J, Swinnen JV, Zaidi N (2019) Lipid metabolism in cancer cells under metabolic stress. Br J Cancer. https://doi.org/10.1038/s41416-019-0451-4
Snaebjornsson MT, Janaki-Raman S, Schulze A (2020) Greasing the wheels of the cancer machine: the role of lipid metabolism in cancer. Cell Metab 31:62–76. https://doi.org/10.1016/j.cmet.2019.11.010
Zaidi N, Lupien L, Kuemmerle NB et al (2013) Lipogenesis and lipolysis: the pathways exploited by the cancer cells to acquire fatty acids. Prog Lipid Res. https://doi.org/10.1016/j.plipres.2013.08.005
Ferreri C, Sansone A, Ferreri R et al (2020) Fatty acids and membrane lipidomics in oncology: a cross-road of nutritional, signaling and metabolic pathways. Metabolites 10:1–26. https://doi.org/10.3390/metabo10090345
Azordegan N, Fraser V, Le K et al (2013) Carcinogenesis alters fatty acid profile in breast tissue. Mol Cell Biochem 374:223–232. https://doi.org/10.1007/s11010-012-1523-4
Flavin R, Peluso S, Nguyen PL, Loda M (2010) Fatty acid synthase as a potential therapeutic target in cancer. Futur Oncol. https://doi.org/10.2217/fon.10.11
Igal RA (2016) Stearoyl CoA desaturase-1: new insights into a central regulator of cancer metabolism. Biochim Biophys Acta-Mol Cell Biol Lipids. https://doi.org/10.1016/j.bbalip.2016.09.009
Igal RA (2010) Stearoyl-CoA desaturase-1: a novel key player in the mechanisms of cell proliferation, programmed cell death and transformation to cancer. Carcinogenesis. https://doi.org/10.1093/carcin/bgq131
Bensaad K, Favaro E, Lewis CA et al (2014) Fatty acid uptake and lipid storage induced by HIF-1α contribute to cell growth and survival after hypoxia-reoxygenation. Cell Rep. https://doi.org/10.1016/j.celrep.2014.08.056
Samanta S, Sharma VM, Khan A, Mercurio AM (2012) Regulation of IMP3 by EGFR signaling and repression by ERβ: implications for triple-negative breast cancer. Oncogene. https://doi.org/10.1038/onc.2011.620
Schug ZT, Peck B, Jones DT et al (2015) Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell. https://doi.org/10.1016/j.ccell.2014.12.002
de Gonzalo-Calvo D, López-Vilaró L, Nasarre L et al (2015) Intratumor cholesteryl ester accumulation is associated with human breast cancer proliferation and aggressive potential: a molecular and clinicopathological study. BMC Cancer. https://doi.org/10.1186/s12885-015-1469-5
Yue S, Li J, Lee S-Y et al (2014) Cholesteryl ester accumulation induced by PTEN loss and PI3K/AKT activation underlies human prostate cancer aggressiveness. Cell Metab. https://doi.org/10.1016/j.cmet.2014.01.019
Vriens K, Christen S, Parik S et al (2019) Evidence for an alternative fatty acid desaturation pathway increasing cancer plasticity. Nature. https://doi.org/10.1038/s41586-019-0904-1
Scanferlato R, Bortolotti M, Sansone A et al (2019) Hexadecenoic fatty acid positional isomers and De Novo PUFA synthesis in colon cancer cells. Int J Mol Sci. https://doi.org/10.3390/ijms20040832
Leone RD, Amaravadi RK (2013) Autophagy: a targetable linchpin of cancer cell metabolism. Trends Endocrinol Metab 24:209–217. https://doi.org/10.1016/j.tem.2013.01.008
Pandey A, Yadav P, Shukla S (2021) Unfolding the role of autophagy in the cancer metabolism. Biochem Biophys Reports 28:101158. https://doi.org/10.1016/j.bbrep.2021.101158
Maan M, Peters JM, Dutta M, Patterson AD (2018) Lipid metabolism and lipophagy in cancer. Biochem Biophys Res Commun 504:582–589. https://doi.org/10.1016/j.bbrc.2018.02.097
Monaco ME (2017) Fatty acid metabolism in breast cancer subtypes. Oncotarget. https://doi.org/10.18632/oncotarget.15494
Cooke M, Orlando U, Maloberti P et al (2011) Tyrosine phosphatase SHP2 regulates the expression of acyl-CoA synthetase ACSL4. J Lipid Res. https://doi.org/10.1194/jlr.M015552
Zhao H, Agazie YM (2015) Inhibition of SHP2 in basal-like and triple-negative breast cells induces basal-to-luminal transition, hormone dependency, and sensitivity to anti-hormone treatment. BMC Cancer. https://doi.org/10.1186/s12885-015-1131-2
Sausgruber N, Coissieux M-M, Britschgi A et al (2015) Tyrosine phosphatase SHP2 increases cell motility in triple-negative breast cancer through the activation of SRC-family kinases. Oncogene. https://doi.org/10.1038/onc.2014.170
Sun X, Wang M, Wang M et al (2020) Metabolic reprogramming in triple-negative breast cancer. Front Oncol. https://doi.org/10.3389/fonc.2020.00428
Abramczyk H, Surmacki J, Kopeć M et al (2015) The role of lipid droplets and adipocytes in cancer. Raman imaging of cell cultures: MCF10A, MCF7, and MDA-MB-231 compared to adipocytes in cancerous human breast tissue. Analyst. https://doi.org/10.1039/C4AN01875C
He J, Zhang F, Rachel Tay LW et al (2017) Lipin-1 regulation of phospholipid synthesis maintains endoplasmic reticulum homeostasis and is critical for triple-negative breast cancer cell survival. FASEB J. https://doi.org/10.1096/fj.201601353R
Pucer A, Brglez V, Payré C et al (2013) Group X secreted phospholipase A2 induces lipid droplet formation and prolongs breast cancer cell survival. Mol Cancer. https://doi.org/10.1186/1476-4598-12-111
Han S, Huh J, Kim W et al (2014) Phospholipase D activates HIF-1-VEGF pathway via phosphatidic acid. Exp Mol Med. https://doi.org/10.1038/emm.2014.86
Herman MA, Peroni OD, Villoria J et al (2012) A novel ChREBP isoform in adipose tissue regulates systemic glucose metabolism. Nature 484:333–338. https://doi.org/10.1038/nature10986
Lei Y, Zhou S, Hu Q et al (2020) Carbohydrate response element binding protein (ChREBP) correlates with colon cancer progression and contributes to cell proliferation. Sci Rep 10:4233. https://doi.org/10.1038/s41598-020-60903-9
Iizuka K, Bruick RK, Liang G et al (2004) Deficiency of carbohydrate response element-binding protein (ChREBP) reduces lipogenesis as well as glycolysis. Proc Natl Acad Sci 101:7281–7286. https://doi.org/10.1073/pnas.0401516101
Bindesbøll C, Fan Q, Nørgaard RC et al (2015) Liver X receptor regulates hepatic nuclear O-GlcNAc signaling and carbohydrate responsive element-binding protein activity. J Lipid Res 56:771–785. https://doi.org/10.1194/jlr.M049130
Steffensen K, Jan-Åke K (2006) Liver X receptors: new drug targets to treat Type 2 diabetes? Future Lipidol 1:181–189. https://doi.org/10.2217/17460875.1.2.181
Guo D, Reinitz F, Youssef M et al (2011) An LXR agonist promotes glioblastoma cell death through inhibition of an EGFR/AKT/SREBP-1/LDLR–dependent pathway. Cancer Discov 1:442–456. https://doi.org/10.1158/2159-8290.CD-11-0102
Jeong D-W, Lee S, Chun Y-S (2021) How cancer cells remodel lipid metabolism: strategies targeting transcription factors. Lipids Health Dis 20:163. https://doi.org/10.1186/s12944-021-01593-8
Vedin L-L, Lewandowski SA, Parini P et al (2009) The oxysterol receptor LXR inhibits proliferation of human breast cancer cells. Carcinogenesis 30:575–579. https://doi.org/10.1093/carcin/bgp029
Matsubara T, Li F, Gonzalez FJ (2013) FXR signaling in the enterohepatic system. Mol Cell Endocrinol 368:17–29. https://doi.org/10.1016/j.mce.2012.05.004
Silva J, Dasgupta S, Wang G et al (2006) Lipids isolated from bone induce the migration of human breast cancer cells. J Lipid Res 47:724–733. https://doi.org/10.1194/jlr.M500473-JLR200
Peters JM, Hennuyer N, Staels B et al (1997) Alterations in lipoprotein metabolism in peroxisome proliferator-activated receptor α-deficient mice. J Biol Chem 272:27307–27312. https://doi.org/10.1074/jbc.272.43.27307
Her N-H, Jeong S-I, Cho K et al (2013) PPARδ promotes oncogenic redirection of TGF-β1 signaling through the activation of the ABCA1-Cav1 pathway. Cell Cycle 12:1521–1535. https://doi.org/10.4161/cc.24636
Yahagi N, Shimano H, Hasegawa K et al (2005) Co-ordinate activation of lipogenic enzymes in hepatocellular carcinoma. Eur J Cancer 41:1316–1322. https://doi.org/10.1016/j.ejca.2004.12.037
Min HK, Kong G, Moon MH (2010) Quantitative analysis of urinary phospholipids found in patients with breast cancer by nanoflow liquid chromatography–tandem mass spectrometry: II Negative ion mode analysis of four phospholipid classes. Anal Bioanal Chem. https://doi.org/10.1007/s00216-009-3292-9
Wang S, Chen X, Luan H et al (2016) Matrix-assisted laser desorption/ionization mass spectrometry imaging of cell cultures for the lipidomic analysis of potential lipid markers in human breast cancer invasion. Rapid Commun Mass Spectrom. https://doi.org/10.1002/rcm.7466
Kawashima M, Iwamoto N, Kawaguchi-Sakita N et al (2013) High-resolution imaging mass spectrometry reveals detailed spatial distribution of phosphatidylinositols in human breast cancer. Cancer Sci. https://doi.org/10.1111/cas.12229
Hosokawa Y, Masaki N, Takei S et al (2017) Recurrent triple-negative breast cancer (TNBC) tissues contain a higher amount of phosphatidylcholine (32:1) than non-recurrent TNBC tissues. PLoS ONE. https://doi.org/10.1371/journal.pone.0183724
Silva CL, Perestrelo R, Sousa-Ferreira I et al (2020) Lipid biosignature of breast cancer tissues by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Breast Cancer Res Treat. https://doi.org/10.1007/s10549-020-05672-9
Eiriksson FF, Nøhr MK, Costa M et al (2020) Lipidomic study of cell lines reveals differences between breast cancer subtypes. PLoS ONE 15:1–22. https://doi.org/10.1371/journal.pone.0231289
Kim H-Y, Lee K-M, Kim S-H et al (2016) Comparative metabolic and lipidomic profiling of human breast cancer cells with different metastatic potentials. Oncotarget. https://doi.org/10.18632/oncotarget.11560
Purwaha P, Gu F, Piyarathna D et al (2018) Unbiased lipidomic profiling of triple-negative breast cancer tissues reveals the association of sphingomyelin levels with patient disease-free survival. Metabolites. https://doi.org/10.3390/metabo8030041
Tabatabaei M, Tafazzoli-Shadpour M, Khani MM (2019) Correlation of the cell mechanical behavior and quantified cytoskeletal parameters in normal and cancerous breast cell lines. Biorheology. https://doi.org/10.3233/BIR-190214
Li QS, Lee GYH, Ong CN, Lim CT (2008) AFM indentation study of breast cancer cells. Biochem Biophys Res Commun. https://doi.org/10.1016/j.bbrc.2008.07.078
Luo Q, Kuang D, Zhang B, Song G (2016) Cell stiffness determined by atomic force microscopy and its correlation with cell motility. Biochim Biophys Acta-Gen Subj 1860:1953–1960. https://doi.org/10.1016/j.bbagen.2016.06.010
Yubero ML, Kosaka PM, San Paulo Á et al (2020) Effects of energy metabolism on the mechanical properties of breast cancer cells. Commun Biol. https://doi.org/10.1038/s42003-020-01330-4
Wirtz D, Konstantopoulos K, Searson P (2012) Mechanical forces in metastasis. Biomol Eng 11:512–522. https://doi.org/10.1038/nrc3080.The
Betapudi V, Licate LS, Egelhoff TT (2006) Distinct roles of nonmuscle myosin II isoforms in the regulation of MDA-MB-231 breast cancer cell spreading and migration. Cancer Res. https://doi.org/10.1158/0008-5472.CAN-05-4236
Smelser AM, Macosko JC, O’Dell AP et al (2015) Mechanical properties of normal versus cancerous breast cells. Biomech Model Mechanobiol 14:1335–1347. https://doi.org/10.1007/s10237-015-0677-x
Calzado-Martín A, Encinar M, Tamayo J et al (2016) Effect of actin organization on the stiffness of living breast cancer cells revealed by peak-force modulation atomic force microscopy. ACS Nano. https://doi.org/10.1021/acsnano.5b07162
Smolyakov G, Thiebot B, Campillo C et al (2016) Elasticity, adhesion, and tether extrusion on breast cancer cells provide a signature of their invasive potential. ACS Appl Mater Interfaces 8:27426–27431. https://doi.org/10.1021/acsami.6b07698
Onwudiwe K, Hu J, Obayemi J et al (2021) Actin cytoskeletal structure and the statistical variations of the mechanical properties of non-tumorigenic breast and triple-negative breast cancer cells. J Mech Behav Biomed Mater. https://doi.org/10.1016/j.jmbbm.2021.104505
Hui TH, Zhou ZL, Fong HW et al (2016) Characterizing the malignancy and drug resistance of cancer cells from their membrane resealing response. Sci Rep 6:1–8. https://doi.org/10.1038/srep26692
Bouvet F, Ros M, Bonedeau E et al (2020) Defective membrane repair machinery impairs survival of invasive cancer cells. Sci Rep. https://doi.org/10.1038/s41598-020-77902-5
Boye TL, Nylandsted J (2016) Annexins in plasma membrane repair. Biol Chem. https://doi.org/10.1515/hsz-2016-0171
Jaiswal JK, Nylandsted J (2015) S100 and annexin proteins identify cell membrane damage as the Achilles heel of metastatic cancer cells. Cell Cycle. https://doi.org/10.1080/15384101.2014.995495
Li M, Xi N, Wang Y, Liu L (2021) Atomic force microscopy for revealing micro/nanoscale mechanics in tumor metastasis: from single cells to microenvironmental cues. Acta Pharmacol Sin. https://doi.org/10.1038/s41401-020-0494-3
Ríos-Marco P, Marco C, Gálvez X et al (2017) Alkylphospholipids: an update on molecular mechanisms and clinical relevance. Biochim Biophys Acta-Biomembr 1859:1657–1667. https://doi.org/10.1016/j.bbamem.2017.02.016
Davis HW, Vallabhapurapu SD, Chu Z et al (2019) Enhanced phosphatidylserine-selective cancer therapy with irradiation and SapC-DOPS nanovesicles. Oncotarget. https://doi.org/10.18632/oncotarget.26615
Chang W, Fa H, Xiao D, Wang J (2020) Targeting phosphatidylserine for Cancer therapy: prospects and challenges. Theranostics. https://doi.org/10.7150/thno.45125
Belzile O, Huang X, Gong J et al (2018) Antibody targeting of phosphatidylserine for the detection and immunotherapy of cancer. ImmunoTargets Ther. https://doi.org/10.2147/ITT.S134834
Chalasani P, Marron M, Roe D et al (2015) A phase I clinical trial of bavituximab and paclitaxel in patients with HER2 negative metastatic breast cancer. Cancer Med. https://doi.org/10.1002/cam4.447
Desai TJ, Toombs JE, Minna JD et al (2016) Identification of lipid-phosphatidylserine (PS) as the target of unbiasedly selected cancer specific peptide-peptoid hybrid PPS1. Oncotarget. https://doi.org/10.18632/oncotarget.8929
Desai TJ, Udugamasooriya DG (2017) A comprehensive lipid binding and activity validation of a cancer-specific peptide-peptoid hybrid PPS1. Biochem Biophys Res Commun. https://doi.org/10.1016/j.bbrc.2017.03.083
De M, Ghosh S, Asad M et al (2020) Combining doxorubicin with stearylamine-bearing liposomes elicits Th1 cytokine responses and cures metastasis in a mouse model. Cancer Immunol Immunother. https://doi.org/10.1007/s00262-020-02578-9
De M, Ghosh S, Sen T et al (2018) A novel therapeutic strategy for cancer using phosphatidylserine targeting stearylamine-bearing cationic liposomes. Mol Ther-Nucleic Acids. https://doi.org/10.1016/j.omtn.2017.10.019
Hullin-Matsuda F, Makino A, Murate M, Kobayashi T (2016) Probing phosphoethanolamine-containing lipids in membranes with duramycin/cinnamycin and aegerolysin proteins. Biochimie 130:81–90. https://doi.org/10.1016/j.biochi.2016.09.020
Iwamoto K, Hayakawa T, Murate M et al (2007) Curvature-dependent recognition of ethanolamine phospholipids by duramycin and cinnamycin. Biophys J 93:1608–1619. https://doi.org/10.1529/biophysj.106.101584
Wang CK, Wacklin HP, Craik DJ (2012) Cyclotides insert into lipid bilayers to form membrane pores and destabilize the membrane through hydrophobic and phosphoethanolamine-specific interactions. J Biol Chem. https://doi.org/10.1074/jbc.M112.421198
Delmas D, Aires V, Colin DJ et al (2013) Importance of lipid microdomains, rafts, in absorption, delivery, and biological effects of resveratrol. Ann N Y Acad Sci. https://doi.org/10.1111/nyas.12177
Gao M, Zhou J, Su Z, Huang Y (2017) Bacterial cupredoxin azurin hijacks cellular signaling networks: protein-protein interactions and cancer therapy. Protein Sci. https://doi.org/10.1002/pro.3310
Huang F, Shu Q, Qin Z et al (2020) Anticancer actions of azurin and its derived peptide p28. Protein J. https://doi.org/10.1007/s10930-020-09891-3
Bernardes N, Garizo AR, Pinto SN et al (2018) Azurin interaction with the lipid raft components ganglioside GM-1 and caveolin-1 increases membrane fluidity and sensitivity to anti-cancer drugs. Cell Cycle. https://doi.org/10.1080/15384101.2018.1489178
Zhang J, Li Q, Wu Y et al (2019) Cholesterol content in cell membrane maintains surface levels of ErbB2 and confers a therapeutic vulnerability in ErbB2-positive breast cancer. Cell Commun Signal. https://doi.org/10.1186/s12964-019-0328-4
Penkauskas T, Zentelyte A, Ganpule S et al (2020) Pleiotropic effects of statins via interaction with the lipid bilayer: a combined approach. Biochim Biophys Acta-Biomembr. https://doi.org/10.1016/j.bbamem.2020.183306
Sahu SS, Sarkar P, Shrivastava S, Chattopadhyay A (2019) Differential effects of simvastatin on membrane organization and dynamics in varying phases. Chem Phys Lipids. https://doi.org/10.1016/j.chemphyslip.2019.104831
Sariisik E, Koçak M, Kucuk Baloglu F, Severcan F (2019) Interaction of the cholesterol reducing agent simvastatin with zwitterionic DPPC and charged DPPG phospholipid membranes. Biochim Biophys Acta-Biomembr. https://doi.org/10.1016/j.bbamem.2019.01.014
Beckwitt CH, Shiraha K, Wells A (2018) Lipophilic statins limit cancer cell growth and survival, via involvement of Akt signaling. PLoS ONE. https://doi.org/10.1371/journal.pone.0197422
Ahern TP, Pedersen L, Tarp M et al (2011) Statin prescriptions and breast cancer recurrence risk: a Danish nationwide prospective cohort study. JNCI J Natl Cancer Inst. https://doi.org/10.1093/jnci/djr291
Ahmadi Y, Karimian R, Panahi Y (2018) Effects of statins on the chemoresistance—the antagonistic drug-drug interactions versus the anti-cancer effects. Biomed Pharmacother. https://doi.org/10.1016/j.biopha.2018.09.122
Erazo-Oliveras A, Fuentes NR, Wright RC, Chapkin RS (2018) Functional link between plasma membrane spatiotemporal dynamics, cancer biology, and dietary membrane-altering agents. Cancer Metastas Rev 37:519–544. https://doi.org/10.1007/s10555-018-9733-1
Safwat S, Hathout RM, Ishak RA, Mortada ND (2017) Augmented simvastatin cytotoxicity using optimized lipid nanocapsules: a potential for breast cancer treatment. J Liposome Res. https://doi.org/10.3109/08982104.2015.1137313
Matusewicz L, Filip-Psurska B, Psurski M et al (2019) EGFR-targeted immunoliposomes as a selective delivery system of simvastatin, with potential use in treatment of triple-negative breast cancers. Int J Pharm. https://doi.org/10.1016/j.ijpharm.2019.118605
Silva-Cázares M, Saavedra-Leos M, Jordan-Alejandre E et al (2020) Lipid-based nanoparticles for the therapeutic delivery of non-coding RNAs in breast cancer (Review). Oncol Rep. https://doi.org/10.3892/or.2020.7791
Akshaya RL, Akshaya N, Selvamurugan N (2020) A computational study of non-coding RNAs on the regulation of activating transcription factor 3 in human breast cancer cells. Comput Biol Chem. https://doi.org/10.1016/j.compbiolchem.2020.107386
Vaidya AM, Sun Z, Ayat N et al (2019) Systemic delivery of tumor-targeting siRNA nanoparticles against an oncogenic LncRNA facilitates effective triple-negative breast cancer therapy. Bioconjug Chem. https://doi.org/10.1021/acs.bioconjchem.9b00028
Jin S-J, Jin M-Z, Xia B-R, Jin W-L (2019) Long non-coding RNA DANCR as an emerging therapeutic target in human cancers. Front Oncol. https://doi.org/10.3389/fonc.2019.01225
Saraiva SM, Gutiérrez-Lovera C, Martínez-Val J et al (2021) Edelfosine nanoemulsions inhibit tumor growth of triple negative breast cancer in zebrafish xenograft model. Sci Rep 11:1–13. https://doi.org/10.1038/s41598-021-87968-4
De Vita A, Liverani C, Molinaro R et al (2021) Lysyl oxidase engineered lipid nanovesicles for the treatment of triple negative breast cancer. Sci Rep. https://doi.org/10.1038/s41598-021-84492-3
Doll S, Proneth B, Tyurina YY et al (2017) ACSL4 dictates ferroptosis sensitivity by shaping cellular lipid composition. Nat Chem Biol. https://doi.org/10.1038/nchembio.2239
Li J, Cao F, Yin H et al (2020) Ferroptosis: past, present and future. Cell Death Dis. https://doi.org/10.1038/s41419-020-2298-2
Yang WS, SriRamaratnam R, Welsch ME et al (2014) Regulation of ferroptotic cancer cell death by GPX4. Cell. https://doi.org/10.1016/j.cell.2013.12.010
Yu H, Yang C, Jian L et al (2019) Sulfasalazine-induced ferroptosis in breast cancer cells is reduced by the inhibitory effect of estrogen receptor on the transferrin receptor. Oncol Rep. https://doi.org/10.3892/or.2019.7189
Hangauer MJ, Viswanathan VS, Ryan MJ et al (2017) Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature. https://doi.org/10.1038/nature24297
Chen M-S, Wang S-F, Hsu C-Y et al (2017) CHAC1 degradation of glutathione enhances cystine-starvation-induced necroptosis and ferroptosis in human triple negative breast cancer cells via the GCN2-eIF2α-ATF4 pathway. Oncotarget. https://doi.org/10.18632/oncotarget.23055
Guerra ÂR, Paulino AF, Castro MM et al (2020) Triple negative breast cancer and breast epithelial cells differentially reprogram glucose and lipid metabolism upon treatment with triterpenic acids. Biomolecules 10:1–18. https://doi.org/10.3390/biom10081163
Wright HJ, Hou J, Xu B et al (2017) CDCP1 drives triple-negative breast cancer metastasis through reduction of lipid-droplet abundance and stimulation of fatty acid oxidation. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.1703791114
Guardiola-Serrano F, Beteta-Göbel R, Rodríguez-Lorca R et al (2019) The triacylglycerol, hydroxytriolein, inhibits triple negative mammary breast cancer cell proliferation through a mechanism dependent on dihydroceramide and Akt. Oncotarget 10:2486–2507. https://doi.org/10.18632/oncotarget.26824
Bulbake U, Doppalapudi S, Kommineni N, Khan W (2017) Liposomal formulations in clinical use: an updated review. Pharmaceutics 9:12. https://doi.org/10.3390/pharmaceutics9020012
Beltrán-Gracia E, López-Camacho A, Higuera-Ciapara I et al (2019) Nanomedicine review: clinical developments in liposomal applications. Cancer Nanotechnol 10:11. https://doi.org/10.1186/s12645-019-0055-y
Bozzuto G, Molinari A (2015) Liposomes as nanomedical devices. Int J Nanomed. https://doi.org/10.2147/IJN.S68861
Miller K, Cortes J, Hurvitz SA et al (2016) HERMIONE: a randomized Phase 2 trial of MM-302 plus trastuzumab versus chemotherapy of physician’s choice plus trastuzumab in patients with previously treated, anthracycline-naïve, HER2-positive, locally advanced/metastatic breast cancer. BMC Cancer 16:352. https://doi.org/10.1186/s12885-016-2385-z
Harbeck N, Saupe S, Jäger E et al (2017) A randomized phase III study evaluating pegylated liposomal doxorubicin versus capecitabine as first-line therapy for metastatic breast cancer: results of the PELICAN study. Breast Cancer Res Treat 161:63–72. https://doi.org/10.1007/s10549-016-4033-3
Jehn CF, Hemmati P, Lehenbauer-Dehm S et al (2016) Biweekly pegylated liposomal doxorubicin (Caelyx) in heavily pretreated metastatic breast cancer: a phase 2 study. Clin Breast Cancer 16:514–519. https://doi.org/10.1016/j.clbc.2016.06.001
Chang AE, Wu QV, Jenkins IC et al (2018) Phase I/II trial of combined pegylated liposomal doxorubicin and cyclophosphamide in metastatic breast cancer. Clin Breast Cancer 18:e143–e149. https://doi.org/10.1016/j.clbc.2017.10.005
Basho RK, Gilcrease M, Murthy RK et al (2017) Targeting the PI3K/AKT/mTOR pathway for the treatment of mesenchymal triple-negative breast cancer. JAMA Oncol 3:509. https://doi.org/10.1001/jamaoncol.2016.5281
Awada A, Bondarenko IN, Bonneterre J et al (2014) A randomized controlled phase II trial of a novel composition of paclitaxel embedded into neutral and cationic lipids targeting tumor endothelial cells in advanced triple-negative breast cancer (TNBC). Ann Oncol 25:824–831. https://doi.org/10.1093/annonc/mdu025
Ahn HK, Jung M, Sym SJ et al (2014) A phase II trial of Cremorphor EL-free paclitaxel (Genexol-PM) and gemcitabine in patients with advanced non-small cell lung cancer. Cancer Chemother Pharmacol 74:277–282. https://doi.org/10.1007/s00280-014-2498-5
Acknowledgements
We acknowledge IIT Bombay for all central facilities.
Funding
Work in the SK lab is supported by the DST-SERB Grant EMR/2016/005414 and WEA/2020/000032, BRNS Grant 201805BRE01RP04922, Wadhwani Research Centre for Bioengineering (WRCB), DBT Ramalingaswami Fellowship, and the DST-Inspire Faculty Award. RD acknowledges funding from DST Women Scientist Scheme WOS-A/CS-16/2018.
Author information
Authors and Affiliations
Contributions
SK and RD conceived the idea, wrote the review, and made the figures. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dadhich, R., Kapoor, S. Lipidomic and Membrane Mechanical Signatures in Triple-Negative Breast Cancer: Scope for Membrane-Based Theranostics. Mol Cell Biochem 477, 2507–2528 (2022). https://doi.org/10.1007/s11010-022-04459-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-022-04459-4