Abstract
Increasing studies have indicated that hypoxia serves as a pivotal microenvironmental factor that facilitates activation of hepatic stellate cells (HSCs). However, the mechanism by which hypoxia activates HSCs is not clear. Here, we demonstrated that plasmacytoma variant translocation 1 (PVT1) and autophagy were overexpressed in liver fibrotic specimens. In primary mouse HSCs, both PVT1 and autophagy were induced by hypoxia. Further study showed that hypoxia-induced autophagy depended on expression of PVT1 and miR-152 in HSCs. Luciferase reporter assay indicated that autophagy-related gene 14 (ATG14) was a direct target of miR-152. In addition, inhibition of autophagy by 3‐methyladenine and Beclin-1 siRNA impeded activation of HSCs cultured in 1% O2. Taken together, autophagy induction via the PVT1-miR-152-ATG14 signaling pathway contributes to activation of HSCs under hypoxia condition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Liver fibrosis is the common outcome of all chronic liver diseases, eventually leading to liver cirrhosis and liver cancer. It is featured with excessive deposition of extracellular matrix (ECM) in the liver. The activation of hepatic stellate cells (HSCs) is the crucial step in the development of liver fibrosis [1]. During chronic liver injury, HSCs trans-differentiate into myofibroblast-like cells, which express plenty of α-SMA and synthesize abundant ECM [2]. Therefore, to clarify the detailed mechanism of HSC activation will contribute to develop a therapeutic target for liver fibrosis.
Hypoxia is a common phenomenon implicated in many pathophysiological conditions. Increasing studies have provided an important role for hypoxia in the progress of liver fibrosis [3, 4]. Hypoxia contributes to activation of HSCs, thereby resulting in ECM deposition [5]. However, the underlying mechanism of HSC activation induced by hypoxia is not completely elucidated.
Accumulating studies have suggested that hypoxia induces not only expression of protein-coding genes but also expression of noncoding RNAs [6]. Hypoxia is characteristic of tumor microenvironment. Plasmacytoma variant translocation 1 (PVT1) is overexpressed in renal carcinoma, uveal melanoma, colorectal cancer, glioma, gallbladder cancer, hepatocellular carcinoma [7,8,9,10,11,12]. Up-regulation of PVT1 predicts clinical progression, lymph node metastasis and unfavorable prognosis in cancers [13, 14]. PVT1 is closely correlated with cisplatin resistance in lung cancer, ovarian cancer cells, gastric and colorectal cancer [15,16,17,18]. In addition, PVT1 knockdown enhances radio-sensitivity via inducing miR-195 expression in lung cancer patients [19]. Our previous study indicated that PVT1 can promote activation of HSCs through competitively binding miR-152 [20]. However, little is known about the role of PVT1 in hypoxia-induced HSC activation.
Increasing studies have revealed that autophagy plays a pivotal role in liver fibrosis by modulating the activation of HSCs [21, 22]. In addition, hypoxia can activate autophagy via inducing the expression of autophagy-related gene 5 (ATG5), Beclin-1 and the conversion of LC3-I to LC3-II [23]. However, the molecular mechanisms underlying the induction of autophagy in response to hypoxia in HSCs remain incompletely understood.
In this study, we demonstrated that the PVT1-miR-152-ATG14 signaling pathway plays a key role in hypoxic-activated autophagy in HSCs. Our study sheds new light on the mechanisms of how hypoxia activates HSCs and proposes that PVT1 is a crucial player during the progression of liver fibrosis.
Materials and methods
Animals
All animal experiments were conducted in accordance with the Guide for the Care and Use of Laboratory Animals published by the US National Institutes of Health (NIH Publication, eighth Edition, 2011) and were approved by the Animal Ethics Committee of Wenzhou Medical University. Eight-week-old C57BL/6 J mice were bred in a clean room at a room temperature of 23 °C ± 2 °C (humidity: 50 ± 5%). They were given free access to food and water. For liver fibrosis model, mice received CCl4 (10% in olive oil) at 7 µL/g body weight two times weekly for 6 week. At the end of treatment, all mice were killed under anesthesia and the livers were collected for further examination.
HSC isolation and cell culture
The primary HSCs were obtained from male C57BL/6 J mice as previously detailed [24]. In brief, after the liver was in situ perfused with 2-step pronase–collagenase digestion, HSCs were isolated by density-gradient centrifugation using 11.5% OptiPrep (Axis-Shield, Oslo, Norway). HSCs were cultured in Dulbecco’s modified Eagle’s medium containing 10% fetal bovine serum and 1% penicillin/streptomycin at 37 °C with 5% CO2. Cell purity was evaluated by immunocytochemical staining for a‐SMA and was greater than 95%. To induce hypoxia, HSCs were incubated in a humidified modular incubator chamber (Thermo FisherScientific, Rochester, NY, USA) with a atmosphere of 1% O2 for 12 h at 37 °C. Control cells were incubated in a chamber with a humidified atmosphere of 21% O2 for equivalent periods.
Cell transfection
HSCs were plated in 6-well plates at a density of 2 × 105 cells per well. After cells reached approximately 70% confluency, they were transduced with adenoviral vector expressing PVT1 for 10 h and harvested 48 h following transduction. All small interfering RNAs (siRNAs) were synthesized by GenePharma Corporation (Shanghai, China). Cells were transfected with PVT1 siRNA, Beclin-1 siRNA or control siRNA using Lipofectamine 2000 reagent (Life Technologies, Carlsbad, CA, USA) at a final concentration of 100 nM for 5 h. Then, the medium was changed and the cells were cultured for another 48 h.
Immunofluorescence
Immunofluorescence staining was performed as we previously described [25]. After the nuclei were stained with 4′,6-diamidino-2-phenylindole in treated HSCs, all slides were examined by a Carl Zeiss LSM710 confocal microscope (Carl Zeiss AG, Jena, Germany) and representative images were presented.
Transmission electron microscopy (TEM)
Liver tissues were cut into tiny fragments and fixed with 2.5% glutaraldehyde. Then tissues were embedded in Epon, cut into thin sections and stained with toluidine blue. The cells were visualized by TEM (Hitachi, Japan).
Quantitative real time PCR (qRT-PCR)
After RNA was isolated from liver specimens and cells and reverse transcribed, qRT-PCR was performed using gene-specific primer sequences listed in Supplementary Table 1.
Luciferase reporter gene assay
The luciferase reporter assay was performed as previously detailed [26]. Briefly, cells were transfected with miR-152 or control miRNA plus pMIR-report luciferase plasmids containing wild-type or mutated miR-152 binding sites of the ATG14 3′UTR by Lipofectamine 2000. Twenty-four hours later, luciferase activity was analyzed by the dual-luciferase assay (Promega, Madison, WI, USA).
Cell proliferation assay
Cell viability was determined using 3‐(4,5‐cimethylthiazol‐2‐yl)‐2,5‐diphenyltetrazolium bromide (MTT) (Sigma, St Louis, MO) assay. HSCs plated in 96-well plates at a density of 1 × 104 cells per well under hypoxia were treated with 3‐methyladenosine (3-MA) or Beclin-1 siRNA for the indicated time. Then cell medium was replaced with fresh medium including 0.5 mg/mL of MTT and cultured for 5 h. Then the supernatant was removed and dimethyl sulfoxide (Sigma, St Louis, MO) was used to dissolve formazan. The absorbance at 570 nm was examined with a microplate reader (Bio-Rad, Hercules, CA, USA).
Western blot
The total protein was lysed in RIPA buffer and subjected to SDS-PAGE. After blocked with 5% non-fat dry milk for 1 h, the membranes were incubated with the primary antibodies against Beclin-1, LC3 I/II and P62 (Cell Signaling Technology, Boston, MA, USA), β‐actin, type I collagen, and α‐SMA (Abcam, Cambridge, MA, USA) overnight at 4 °C and then with the secondary antibodies for 1 h. Signals were analyzed using an Odyssey two-color infrared laser imaging system (LI-COR, Lincoln, NE, USA).
Statistical analysis
All the statistical analyses were performed using SPSS 13.0 (IBM, Armonk, NY, USA). The experimental data are expressed as the mean ± SD. The results were considered to be statistically significant at P < 0.05.
Results
Expression of PVT1 and LC3 in liver fibrosis
To explore whether PVT1 and autophagy are involved in liver fibrosis, we detected their expression in liver fibrotic specimens. Liver fibrosis stage was assessed by staining with Hematoxylin–Eosin and Sirius Red (Fig. 1a, b). qRT-PCR and western blot results showed that the expression of type I collagen and α-SMA in CCl4-treated mice was up-regulated compared with those in control mice (Fig. 1c–h). Consistently, we observed that PVT1 expression was higher in CCl4-treated mice than that in control mice (Fig. 1I). Then, western blot experiments were employed to examine the expression of autophagy marker LC3. As indicated in Fig. 1j, k, our results demonstrated that the up-regulation of LC3‐II protein was present in CCl4-treated mice relative to control mice. TEM is the gold standard method of autophagy monitoring. TEM results showed that there were more autophagosomes formation in CCl4-treated mice than those in control mice (Fig. 1l). To summarize, our data suggested that PVT1 and autophagy may participate in the pathological process of liver fibrosis.
PVT1 and LC3-II are up-regulated in liver fibrotic specimens. a CCl4-treated liver fibrosis model was evaluated by Hematoxylin–Eosin staining. Scale bars, 100 μm. b CCl4-treated liver fibrosis model was evaluated by Sirius Red staining. Scale bars, 100 μm. c The mRNA expression of Col1A1 was analyzed in livers of mice treated with CCl4. d, e The protein expression of type I collagen was analyzed in livers of mice treated with CCl4. f The mRNA expression of α-SMA was analyzed in livers of mice treated with CCl4. g, h The protein expression of α-SMA was analyzed in livers of mice treated with CCl4. i Expression of PVT1 was analyzed in livers of mice treated with CCl4. j, k The protein expression of LC3-II was analyzed in livers of mice treated with CCl4. l The number of autophagosomes was analyzed by TEM in livers of mice treated with CCl4. Results are expressed as mean ± SD, **P < 0.01
Hypoxia increases expression of PVT1 and induces activation of autophagy in HSCs
We firstly examined the expression of PVT1 in HSCs under hypoxia. As indicated in Fig. 2A, compared with HSCs cultured at room air, the increased expression of PVT1 was observed in HSCs cultured under 1% O2.
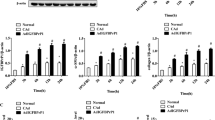
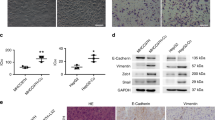
Hypoxia increases expression of PVT1 and induces activation of autophagy in HSCs. a Expression of PVT1 was analyzed in HSCs under 21% O2 or 1% O2. b–e HSCs were cultured under 21% O2 or 1% O2 for 12 h; and proteins were analyzed for western blotting of LC3, Beclin-1, and P62, with β-actin as the internal control. f, g Cells were cultured under 21% O2, chloroquine, or chloroquine with 1% O2 for 12 h; and the protein expression of LC3 was analyzed by western blotting. h The protein expression of vimentin was analyzed using an immunofluorescence staining method in HSCs under 21% O2 or 1% O2. Scale bars, 50 μm. i The protein expression of α-SMA was analyzed using an immunofluorescence staining method in HSCs under 21% O2 or 1% O2. Scale bars, 50 μm. Results are expressed as mean ± SD, **P < 0.01. CQ chloroquine
To further investigate whether hypoxia could induce autophagy in HSCs, we detected the expression of Beclin-1 and LC3‐II, the reliable autophagy-associated markers. As indicated in Fig. 2b–e, there was increased expression of Beclin-1 and LC3‐II protein in HSCs under hypoxia compared with normoxia. Conversely, there was the decreased expression of P62 protein. Then, we further confirmed whether the up-regulation of LC3-II came from autophagy induction or reduced autophagy flux. As shown in Fig. 2f, g, when compared with normoxia, chloroquine increased the expression of LC3-II, whereas chloroquine in combination with hypoxia resulted in higher expression of LC3-II than chloroquine, suggesting that the up-regulation of LC3-II is caused by autophagy induction.
We lastly examined the expression of vimentin and α-SMA in HSCs under hypoxia. The increased expression of vimentin was observed in hypoxic HSCs compared with normoxia (Fig. 2h). The expression of α-SMA was also more increased in HSCs under hypoxia, when compared with cells under normoxia (Fig. 2i). These results showed that HSCs are activated by hypoxia stress.
PVT1 modulates hypoxia-induced autophagy via miR-152 in HSCs
To explore the biological role of PVT1 in hypoxia-induced autophagy in HSCs, we successfully constructed a PVT1 siRNA. Our results demonstrated that the PVT1 siRNA efficiently decreased the expression of PVT1 compared with the control siRNA (Fig. 3a). Then we detected the changes of autophagy in the presence of hypoxia with or without PVT1 siRNA. We found that silencing of PVT1 markedly down-regulated the protein expression of Beclin-1 when compared with the control siRNA, suggesting that PVT1 contributes to hypoxia‐induced autophagy activation in HSCs (Fig. 3b, c).
PVT1 modulates hypoxia-induced autophagy via miR-152 in HSCs. a The silencing effect of PVT1 siRNA was analyzed by qRT-PCR. b, c The protein expression of Beclin-1 was analyzed by western blotting in HSCs under 1% O2, 1% O2 + si-Ctrl or 1% O2 + si-PVT1. d, e The protein expression of Beclin-1 was analyzed by western blotting in HSCs treated with Ctrl, PVT1 or PVT1 + miR-152. f A schematic representation of the putative miR-152 binding site in the 3′UTR of ATG14 mRNA. g The relative luciferase activities of luciferase reporters including the wild-type or mutant ATG14 were analyzed 48 h after transfection with miR-152 or miR-NC. Results are expressed as mean ± SD, **P < 0.01
Our previous study indicated that PVT1 can act as a sponge by binding miR-152, thereby contributing to activation of HSCs. Herein, we detected whether miR-152 was involved in the activation of autophagy induced by PVT1. Our results indicated that the increased expression of Beclin-1 by PVT1 overexpression was reversed in the presence of miR-152 (Fig. 3d, e), indicating that PVT1 activates autophagy via miR-152 in HSCs.
Lastly, we explored the mechanism underlying the down‐regulation of Beclin-1 induced by miR-152 in HSCs. Among the putative miR‐152 targets, we selected ATG14 because it can activate autophagy via deacetylating Beclin-1. Our data showed that ATG14 3′UTR has a target site of miR-152 according to TargetScan software (Fig. 3f). As shown in Fig. 3g, miR-152 decreased the luciferase activity of the pmirGLO‐ATG14‐wt reporter compared with the control miRNA. However, the luciferase activity of the pmirGLO‐ATG14‐mut was not changed by miR-152, confirming that ATG14 is a target of miR‐152.
Autophagy inhibition prevents activation of HSCs under hypoxia stress
Next, we explored whether autophagy induction can activate HSCs under hypoxia stress. MTT assay showed that 3‐MA and Beclin‐1 siRNA, autophagy inhibitors, decreased the cell viability of HSCs under hypoxia (Fig. 4a). Consistently, western blot data indicated that the protein expression of type 1 collagen and α-SMA was markedly decreased in HSCs under hypoxia with 3‐MA or Beclin-1 siRNA, when compared with hypoxia (Fig. 4b–d). These results suggested that autophagy is involved in the activation of HSCs under hypoxia conditions.
Autophagy inhibition prevents activation of HSCs under hypoxia condition. a Cell viability was analyzed by the MTT assay in HSCs under 1% O2, 1% O2 + 3-MA or 1% O2 + si-Beclin-1. b–d The protein expression of type I collagen and α-SMA was analyzed by western blotting in HSCs under 1% O2, 1% O2 + 3-MA or 1% O2 + si-Beclin-1. Results are expressed as mean ± SD, **P < 0.01
Discussion
In the present study, we showed that the expression of PVT1 and the autophagy marker LC3‐II was markedly increased in liver fibrotic samples. Furthermore, hypoxia dramatically increased the expression of PVT1, which induced autophagy via the miR-152/ATG14 signaling pathway, thus leading to activation of HSCs.
During the progression of liver fibrosis, hypoxia exists in the liver because of microvasculature distortion, normal hepatic blood flow reduction, and excess deposition of ECM in the sinusoidal space [27]. Many studies have indicated that hypoxia plays a pivotal role in the pathogenesis of liver fibrosis [28, 29]. However, the mechanism of hypoxia activating HSCs has not been fully understood. Hypoxia has been reported to regulate the expression of abundant protein-coding genes. Increasing evidence showed that many noncoding RNAs are also modulated by hypoxia [30, 31]. PVT1 is a highly overexpressed lncRNA in many cancers, relating with proliferation, metastasis, and prognosis [32, 33]. Here we showed that the level of PVT1 was markedly up-regulated in liver fibrotic samples and HSCs under hypoxia, compared with the control group. These data suggest that PVT1 is involved in the process of liver fibrosis.
Autophagy is a key catalytic process indispensable for cellular homoeostasis [34]. Many studies have indicated that hypoxia can activate autophagy in many cancers [35, 36]. Some studies have showed that autophagy can activate HSCs via degrading intracellular lipid droplets [37, 38]. When primary HSCs was exposed to 1% O2, we observed the increased expression of LC3-II, Beclin-1, α-SMA and vimentin as well as the decreased expression of P62. In addition, we found that the protein expression of α-SMA and type I collagen was down-regulated in HSCs under hypoxia as pretreated with autophagic inhibitors, which suggested that autophagy participates in activation of HSCs upon hypoxic exposure.
As we described above, hypoxia can induce the expression of PVT1 and autophagy. However, the relationship between PVT1 and autophagy in HSCs under hypoxia is still unknown. We showed that knockdown of PVT1 decreased the level of Beclin-1, indicating that PVT1 is involved in the induction of autophagy. It has been reported that lncRNA can act as competing endogenous RNA via miRNAs sponge [39]. Increasing studies have clarified the inter-play between PVT1 and autophagy [40, 41]. Yang et al. [12] showed that PVT1 induces autophagy as a decoy for miR-365 to target ATG3 in hepatocellular carcinoma. Additionally, PVT1 also activates autophagy by increasing the expression of Atg7 and Beclin-1 in glioma [42]. These results implied that PVT1 could also modulate autophagy in HSCs under 1% O2 in a similar way. Our previous data demonstrate that PVT1 can suppress PTCH1 via competitively binding miR-152, resulting in activation of Hh pathway and EMT process in liver fibrosis [20]. In the present study, we observed that the increased expression of Beclin-1 by PVT1 overexpression was rescued upon miR-152 treatment, which implies that miR-152 might target some ATGs to regulate autophagic activity. Among the putative miR-152 targets, we selected ATG14 due to its association with autophagy. Luciferase reporter assay confirmed that ATG14 is a target of miR-152, which enlarges the repertoire of miR-152 targets. These results suggest that targeting the PVT1-miR-152-ATG14 signaling pathway may become a new therapeutic application in liver fibrosis.
There are some limitations in the present study. PVT1 may sponge many miRNAs that have many targets. Therefore, “multiple-to-multiple” RNA interaction occurs in the real cellular context. Moreover, the biological function of PVT1 in vivo has not been studied.
Collectively, our results indicated that autophagy induction via the PVT1-miR-152-ATG14 signaling pathway contributes to activation of HSCs under hypoxia condition. Because HSCs play a key role in the development of liver fibrosis, it is reasonable to deduce that hypoxia-induced autophagy in vivo might lead to the progress of liver fibrosis, which will be studied in our future research.
References
Lai M, Afdhal NH (2019) Liver fibrosis determination. Gastroenterol Clin N Am 48:281–289. https://doi.org/10.1016/j.gtc.2019.02.002
Parola M, Pinzani M (2019) Liver fibrosis: pathophysiology, pathogenetic targets and clinical issues. Mol Aspects Med 65:37–55. https://doi.org/10.1016/j.mam.2018.09.002
Tian Y, Deng H, Han L, Hu S, Qi X (2018) Hypoxia-inducible factor may induce the development of liver fibrosis in Budd-Chiari syndrome by regulating CD248/endosialin expression: a hypothesis. J Transl Int Med 6:66–69. https://doi.org/10.2478/jtim-2018-0018
Roth KJ, Copple BL (2015) Role of hypoxia-inducible factors in the development of liver fibrosis. Cell Mol Gastroenterol Hepatol 1:589–597. https://doi.org/10.1016/j.jcmgh.2015.09.005
Zhang Q, Xiang S, Liu Q, Gu T, Yao Y, Lu X (2018) PPARgamma antagonizes hypoxia-induced activation of hepatic stellate cell through cross mediating PI3K/AKT and cGMP/PKG signaling. PPAR Res 2018:6970407. https://doi.org/10.1155/2018/6970407
Corrado C, Costa V, Giavaresi G, Calabrese A, Conigliaro A, Alessandro R (2019) Long non coding RNA H19: a new player in hypoxia-induced multiple myeloma cell dissemination. Int J Mol Sci. https://doi.org/10.3390/ijms20040801
Ren Y, Huang W, Weng G, Cui P, Liang H, Li Y (2019) LncRNA PVT1 promotes proliferation, invasion and epithelial-mesenchymal transition of renal cell carcinoma cells through downregulation of miR-16-5p. Onco Targets Ther 12:2563–2575. https://doi.org/10.2147/OTT.S190239
Huang XM, Shi SS, Jian TM, Tang DR, Wu T, Sun FY (2019) LncRNA PVT1 knockdown affects proliferation and apoptosis of uveal melanoma cells by inhibiting EZH2. Eur Rev Med Pharmacol Sci 23:2880–2887. https://doi.org/10.26355/eurrev_201904_17566
Gharib E, Anaraki F, Baghdar K, Ghavidel P, Sadeghi H, Nasrabadi PN, Peyravian N, Aghdaei HA, Zali MR, Mojarad EN (2019) Investigating the diagnostic performance of HOTTIP, PVT1, and UCA1 long noncoding RNAs as a predictive panel for the screening of colorectal cancer patients with lymph node metastasis. J Cell Biochem. https://doi.org/10.1002/jcb.28739
Han Y, Li X, Yan J, Ma C, Zheng X, Zhang J, Zhang D, Meng C, Zhang Z, Ji X, He F (2019) Knockdown of LncRNA PVT1 inhibits glioma progression by regulating miR-424 expression. Oncol Res. https://doi.org/10.3727/096504018X15424939990246
Chen J, Yu Y, Li H, Hu Q, Chen X, He Y, Xue C, Ren F, Ren Z, Li J, Liu L, Duan Z, Cui G, Sun R (2019) Long non-coding RNA PVT1 promotes tumor progression by regulating the miR-143/HK2 axis in gallbladder cancer. Mol Cancer 18:33. https://doi.org/10.1186/s12943-019-0947-9
Yang L, Peng X, Jin H, Liu J (2019) Long non-coding RNA PVT1 promotes autophagy as ceRNA to target ATG3 by sponging microRNA-365 in hepatocellular carcinoma. Gene 697:94–102. https://doi.org/10.1016/j.gene.2019.02.036
Yang Q, Yu Y, Sun Z, Pan Y (2018) Long non-coding RNA PVT1 promotes cell proliferation and invasion through regulating miR-133a in ovarian cancer. Biomed Pharmacother 106:61–67. https://doi.org/10.1016/j.biopha.2018.06.112
Liu C, Jin J, Liang D, Gao Z, Zhang Y, Guo T, He Y (2018) Long noncoding RNA PVT1 as a novel predictor of metastasis, clinicopathological characteristics and prognosis in human cancers: a meta-analysis. Pathol Oncol Res. https://doi.org/10.1007/s12253-018-0451-3
Chen L, Han X, Hu Z, Chen L (2019) The PVT1/miR-216b/Beclin-1 regulates cisplatin sensitivity of NSCLC cells via modulating autophagy and apoptosis. Cancer Chemother Pharmacol 83:921–931. https://doi.org/10.1007/s00280-019-03808-3
Liu E, Liu Z, Zhou Y, Mi R, Wang D (2015) Overexpression of long non-coding RNA PVT1 in ovarian cancer cells promotes cisplatin resistance by regulating apoptotic pathways. Int J Clin Exp Med 8:20565–20572
Zhang XW, Liu L, Zhang XZ, Bo P (2017) Kanglaite inhibits the expression of drug resistance genes through suppressing PVT1 in cisplatin-resistant gastric cancer cells. Exp Ther Med 14:1789–1794. https://doi.org/10.3892/etm.2017.4650
Ping G, Xiong W, Zhang L, Li Y, Zhang Y, Zhao Y (2018) Silencing long noncoding RNA PVT1 inhibits tumorigenesis and cisplatin resistance of colorectal cancer. Am J Transl Res 10:138–149
Wu D, Li Y, Zhang H, Hu X (2017) Knockdown of Lncrna PVT1 enhances radiosensitivity in non-small cell lung cancer by sponging Mir-195. Cell Physiol Biochem 42:2453–2466. https://doi.org/10.1159/000480209
Zheng J, Yu F, Dong P, Wu L, Zhang Y, Hu Y, Zheng L (2016) Long non-coding RNA PVT1 activates hepatic stellate cells through competitively binding microRNA-152. Oncotarget 7:62886–62897. https://doi.org/10.18632/oncotarget.11709
Ruart M, Chavarria L, Camprecios G, Suarez-Herrera N, Montironi C, Guixe-Muntet S, Bosch J, Friedman SL, Garcia-Pagan JC, Hernandez-Gea V (2019) Impaired endothelial autophagy promotes liver fibrosis by aggravating the oxidative stress response during acute liver injury. J Hepatol 70:458–469. https://doi.org/10.1016/j.jhep.2018.10.015
Tan S, Lu Y, Xu M, Huang X, Liu H, Jiang J, Wu B (2019) beta-Arrestin1 enhances liver fibrosis through autophagy-mediated Snail signaling. FASEB J 33:2000–2016. https://doi.org/10.1096/fj.201800828RR
Li R, Wang LZ, Du JH, Zhao L, Yao Y (2018) Autophagy activation and the mechanism of retinal microvascular endothelial cells in hypoxia. Int J Ophthalmol 11:567–574. https://doi.org/10.18240/ijo.2018.04.05
Chang W, Yang M, Song L, Shen K, Wang H, Gao X, Li M, Niu W, Qin X (2014) Isolation and culture of hepatic stellate cells from mouse liver. Acta Biochim Biophys Sin (Shanghai) 46:291–298. https://doi.org/10.1093/abbs/gmt143
Zhang K, Han X, Zhang Z, Zheng L, Hu Z, Yao Q, Cui H, Shu G, Si M, Li C, Shi Z, Chen T, Han Y, Chang Y, Yao Z, Han T, Hong W (2017) The liver-enriched lnc-LFAR1 promotes liver fibrosis by activating TGFbeta and Notch pathways. Nat Commun 8:144. https://doi.org/10.1038/s41467-017-00204-4
Han D, Li J, Wang H, Su X, Hou J, Gu Y, Qian C, Lin Y, Liu X, Huang M, Li N, Zhou W, Yu Y, Cao X (2017) Circular RNA circMTO1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression. Hepatology 66:1151–1164. https://doi.org/10.1002/hep.29270
Zhan L, Huang C, Meng XM, Song Y, Wu XQ, Yang Y, Li J (2015) Hypoxia-inducible factor-1alpha in hepatic fibrosis: a promising therapeutic target. Biochimie 108:1–7. https://doi.org/10.1016/j.biochi.2014.10.013
Ceni E, Mello T, Polvani S, Vasseur-Cognet M, Tarocchi M, Tempesti S, Cavalieri D, Beltrame L, Marroncini G, Pinzani M, Milani S, Galli A (2017) The orphan nuclear receptor COUP-TFII coordinates hypoxia-independent proangiogenic responses in hepatic stellate cells. J Hepatol 66:754–764. https://doi.org/10.1016/j.jhep.2016.11.003
Novo E, Povero D, Busletta C, Paternostro C, di Bonzo LV, Cannito S, Compagnone A, Bandino A, Marra F, Colombatto S, David E, Pinzani M, Parola M (2012) The biphasic nature of hypoxia-induced directional migration of activated human hepatic stellate cells. J Pathol 226:588–597. https://doi.org/10.1002/path.3005
Kim KM, Han CY, Kim JY, Cho SS, Kim YS, Koo JH, Lee JM, Lim SC, Kang KW, Kim JS, Hwang SJ, Ki SH, Kim SG (2018) Galpha12 overexpression induced by miR-16 dysregulation contributes to liver fibrosis by promoting autophagy in hepatic stellate cells. J Hepatol 68:493–504. https://doi.org/10.1016/j.jhep.2017.10.011
Nieminen T, Scott TA, Lin FM, Chen Z, Yla-Herttuala S, Morris KV (2018) Long non-coding RNA modulation of VEGF-A during hypoxia. Noncoding RNA. https://doi.org/10.3390/ncrna4040034
Huang F, Chen W, Peng J, Li Y, Zhuang Y, Zhu Z, Shao C, Yang W, Yao H, Zhang S (2018) LncRNA PVT1 triggers cyto-protective autophagy and promotes pancreatic ductal adenocarcinoma development via the miR-20a-5p/ULK1 Axis. Mol Cancer 17:98. https://doi.org/10.1186/s12943-018-0845-6
Tian Z, Cao S, Li C, Xu M, Wei H, Yang H, Sun Q, Ren Q, Zhang L (2019) LncRNA PVT1 regulates growth, migration, and invasion of bladder cancer by miR-31/CDK1. J Cell Physiol 234:4799–4811. https://doi.org/10.1002/jcp.27279
Wiegering A, Ruther U, Gerhardt C (2019) The role of primary cilia in the crosstalk between the ubiquitin(-)proteasome system and autophagy. Cells. https://doi.org/10.3390/cells8030241
Abdul Rahim SA, Dirkse A, Oudin A, Schuster A, Bohler J, Barthelemy V, Muller A, Vallar L, Janji B, Golebiewska A, Niclou SP (2017) Regulation of hypoxia-induced autophagy in glioblastoma involves ATG9A. Br J Cancer 117:813–825. https://doi.org/10.1038/bjc.2017.263
Chen Y, Henson ES, Xiao W, Huang D, McMillan-Ward EM, Israels SJ, Gibson SB (2016) Tyrosine kinase receptor EGFR regulates the switch in cancer cells between cell survival and cell death induced by autophagy in hypoxia. Autophagy 12:1029–1046. https://doi.org/10.1080/15548627.2016.1164357
Allaire M, Rautou PE, Codogno P, Lotersztajn S (2019) Autophagy in liver diseases: time for translation? J Hepatol 70:985–998. https://doi.org/10.1016/j.jhep.2019.01.026
Liu Y, Bi Y, Mo C, Zeng T, Huang S, Gao L, Sun X, Lv Z (2019) Betulinic acid attenuates liver fibrosis by inducing autophagy via the mitogen-activated protein kinase/extracellular signal-regulated kinase pathway. J Nat Med 73:179–189. https://doi.org/10.1007/s11418-018-1262-2
Song YX, Sun JX, Zhao JH, Yang YC, Shi JX, Wu ZH, Chen XW, Gao P, Miao ZF, Wang ZN (2017) Non-coding RNAs participate in the regulatory network of CLDN4 via ceRNA mediated miRNA evasion. Nat Commun 8:289. https://doi.org/10.1038/s41467-017-00304-1
Alessio E, Buson L, Chemello F, Peggion C, Grespi F, Martini P, Massimino ML, Pacchioni B, Millino C, Romualdi C, Bertoli A, Scorrano L, Lanfranchi G, Cagnin S (2019) Single cell analysis reveals the involvement of the long non-coding RNA Pvt1 in the modulation of muscle atrophy and mitochondrial network. Nucleic Acids Res 47:1653–1670. https://doi.org/10.1093/nar/gkz007
Li Z, Hao S, Yin H, Gao J, Yang Z (2016) Autophagy ameliorates cognitive impairment through activation of PVT1 and apoptosis in diabetes mice. Behav Brain Res 305:265–277. https://doi.org/10.1016/j.bbr.2016.03.023
Ma Y, Wang P, Xue Y, Qu C, Zheng J, Liu X, Ma J, Liu Y (2017) PVT1 affects growth of glioma microvascular endothelial cells by negatively regulating miR-186. Tumour Biol 39:1010428317694326. https://doi.org/10.1177/1010428317694326
Acknowledgements
The project was supported by the Shanghai Municipal Natural Science Foundation (Grant No. 17ZR1426100), the National Natural Science Foundation of China (Grant Nos. 81970527/H0317, 81500458/H0317, 81873576/H0317, 81000176/H0317), Zhejiang Provincial Natural Science Foundation of China (Grant No. LY19H030005), Wenzhou Municipal Science and technology Bureau (Grant No. Y20180138), the Key Project of the Science & Technology Development Fund of Nanjing Medical University (Grant No. 2016NJMUZD092), and Science & Technology Commission of Shanghai Songjiang (Grant No. 16SJGG38).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors confirm that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, F., Dong, B., Dong, P. et al. Hypoxia induces the activation of hepatic stellate cells through the PVT1-miR-152-ATG14 signaling pathway. Mol Cell Biochem 465, 115–123 (2020). https://doi.org/10.1007/s11010-019-03672-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-019-03672-y